Abstract
Recent case reports suggest that coronavirus disease 2019 (COVID-19) is associated with collapsing glomerulopathy in African Americans with apolipoprotein L1 gene (APOL1) risk alleles; however, it is unclear whether disease pathogenesis is similar to HIV-associated nephropathy. RNA sequencing analysis of a kidney biopsy specimen from a patient with COVID-19–associated collapsing glomerulopathy and APOL1 risk alleles (G1/G1) revealed similar levels of APOL1 and angiotensin-converting enzyme 2 (ACE2) messenger RNA transcripts as compared with 12 control kidney samples downloaded from the GTEx (Genotype-Tissue Expression) Portal. Whole-genome sequencing of the COVID-19–associated collapsing glomerulopathy kidney sample identified 4 indel gene variants, 3 of which are of unknown significance with respect to chronic kidney disease and/or focal segmental glomerulosclerosis. Molecular profiling of the kidney demonstrated activation of COVID-19–associated cell injury pathways such as inflammation and coagulation. Evidence for direct severe acute respiratory syndrome coronavirus 2 infection of kidney cells was lacking, which is consistent with the findings of several recent studies. Interestingly, immunostaining of kidney biopsy sections revealed increased expression of phospho-STAT3 (signal transducer and activator of transcription 3) in both COVID-19–associated collapsing glomerulopathy and HIV-associated nephropathy as compared with control kidney tissue. Importantly, interleukin 6–induced activation of STAT3 may be a targetable mechanism driving COVID-19–associated acute kidney injury.
Index Words: Collapsing glomerulopathy, COVID-19, FSGS, APOL1, STAT3, IL-6
Introduction
Acute kidney injury is a common occurrence in the setting of coronavirus disease 2019 (COVID-19) infection among hospitalized patients and is associated with poor overall prognosis.1 The exact mechanism of kidney injury is unknown. Earlier autopsy and biopsy series suggested the possibility of direct invasion of podocytes and renal tubular epithelial cells by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2),2, 3, 4 though more recently published data refute this hypothesis.5, 6, 7, 8, 9 Though renal tubular injury has been a nearly universal finding on autopsies and kidney biopsies, a range of glomerular pathologic states have also been described.5,6,10 Notably, collapsing glomerulopathy has been reported in several patients of African ancestry with COVID-19 infection.7,11, 12, 13
Collapsing glomerulopathy is associated with several viral infections, including HIV-1, parvovirus B19, cytomegalovirus, and Epstein-Barr virus.14 There is evidence for direct infection of podocytes by HIV-1 and possibly parvovirus B19,15,16 but collapsing glomerulopathy is also associated with elevated levels of circulating interferons.17 It is also well established that individuals with high-risk variants of the apolipoprotein L1 gene (APOL1), known as G1 and G2, have significantly increased risk for the development of HIV-associated nephropathy (HIVAN) and possibly parvovirus-associated collapsing glomerulopathy.16,18 In vitro and in vivo studies suggest a complex interplay between interferon and inflammatory mediators with APOL1 protein that may incite podocyte injury through defects in autophagy, mitochondrial dysfunction, and ion channel efflux, with enhancement in inflammatory cell death (pyroptosis).19, 20, 21, 22 However, little is known about APOL1 expression in the setting of human collapsing glomerulopathy.
We present a case of COVID-19–associated collapsing glomerulopathy in a patient homozygous for the G1 APOL1 allele. We used whole-genome sequencing and RNA sequencing technology in conjunction with standard microscopy and immunostaining to identify gene variants associated with focal segmental glomerulosclerosis and chronic kidney disease, examine gene transcript expression levels of APOL1 and ACE2 (angiotensin-converting enzyme 2), investigate the molecular pathways altered in COVID-19 infection, and test for direct invasion of kidney cells by SARS-CoV-2. Additionally, we evaluated protein expression levels of phosphorylated STAT3 (signal transducer and activator of transcription 3), a downstream target of interleukin 6 (IL-6) signaling that has been demonstrated to be important to the pathogenesis of HIVAN. Detailed methods are provided in Item S1.
Case Report
An African American woman in her early 50s with a medical history of hypertension, hypothyroidism, depression, obstructive sleep apnea, and obesity (body mass index, 31 kg/m2) was hospitalized with nonoliguric acute kidney injury and nephrotic-range proteinuria in the setting of confirmed COVID-19 infection. Admission laboratory tests revealed elevated levels of inflammatory markers, including IL-6, IL-8, tumor necrosis factor α, C-reactive protein, and ferritin. Serologic workup for alternate causes of glomerulonephritis was negative (Table 1).
Table 1.
Laboratory Results on Admission
| Laboratory Test | Reference Range | Admission Result |
|---|---|---|
| Sodium, mEq/L | 135-145 | 140 |
| Potassium, mEq/L | 3.5-5.2 | 4.9 |
| Chloride, mEq/L | 96-108 | 101 |
| Bicarbonate, mEq/L | 22-30 | 28.3 |
| Serum urea nitrogen, mg/dL | 6-23 | 71 |
| Creatinine, mg/dL | 0.50-1.10 | 6.08 |
| Glucose, mg/dL | 60-100 | 88 |
| Calcium, mg/dL | 8.5-10.5 | 8.2 |
| Albumin, mg/dL | 3.5-4.9 | 2.2 |
| Hemoglobin A1c, % | 4.0-6.0% | 6.1 |
| White blood cell count, ×103/μL | 4.5-11 | 5.6 |
| Hemoglobin, g/dL | 11.7-15 | 13.6 |
| Platelets, ×103/μL | 150-450 | 356 |
| Urinary protein-creatinine ratio, g/g | 0.10-0.15 | 21.71 |
| Urinary RBCs, per high-power field | 0.0-3.0 | 4-10 |
| Sars-CoV-2 RT-PCR | Not detected | Detected |
| Lactate dehydrogenase, U/L | 100-220 | 591 |
| Creatine kinase, U/L | 25-175 | 71 |
| C-Reactive protein, mg/L | 0.0-5.0 | 19.5 |
| Ferritin, ng/mL | 15-150 | 221 |
| Interleukin 6, pg/mL | 0.0-5.0 | 23.2 |
| Interleukin 8, pg/mL | 0.0-5.0 | 111.0 |
| Tumor necrosis factor α, pg/mL | 0.0-22.0 | 83.3 |
| HIV-1/HIV-2 antigen/antibody | Nonreactive | Nonreactive |
| Hepatitis C virus antibody | Nonreactive | Nonreactive |
| Hepatitis B surface antigen | Nonreactive | Nonreactive |
| Hepatitis B surface antibody | Nonreactive | Nonreactive |
| Hepatitis B core antibody total | Nonreactive | Nonreactive |
| Phospholipase A2 receptor antibody, RU/mL | 0.0-19.9 | <1.8 |
| Anti-DNA (DS) antibody, IU/mL | 0-9 | <1.0 |
| Complement C3, mg/dL | 90-180 | 118 |
| Complement C4, mg/dL | 10-40 | 71 |
Note: Conversion factors for units: serum urea nitrogen in mg/dL to mmol/L, ×0.357; creatinine in mg/dL to μmol/L, ×88.4; glucose in mg/dL to mmol/L, x0.5551; calcium in mg/dL to mmol/L, ×0.2495.
Abbreviations: DS, double strand; RBC, red blood cell; RT-PCR, reverse transcriptase polymerase chain reaction; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
The patient was treated with a 5-day course of hydroxychloroquine without steroids, and treatment with renally dosed apixaban was initiated for anticoagulation. Her kidney function continued to worsen and she was initiated on acute peritoneal dialysis but then transitioned to hemodialysis due to a nonfunctional peritoneal catheter. Hemodialysis treatments were complicated by intradialytic hypotension and a malfunctioning dialysis catheter, necessitating catheter exchange and prolonging hospitalization. On hospital day 31, repeat urinary protein-creatinine ratio remained elevated at 12.5 g/g, while 2 repeat screening tests for SARS-CoV-2 were negative. The patient underwent kidney biopsy on hospital day 35. There were no complications following biopsy, and the patient was discharged the following day. She remains on outpatient hemodialysis treatments at the time of this report.
On light microscopy, 2 of 18 total glomeruli demonstrated collapse of capillary loops with proliferation of overlying visceral epithelial cells, consistent with collapsing glomerulopathy (Fig 1A and B). In addition, 3 glomeruli showed prominence of visceral epithelial cells. Some glomeruli showed mild ischemic changes, and moderate to severe patchy interstitial fibrosis and tubular atrophy were seen throughout. There was diffuse acute tubular necrosis, characterized by attenuation of epithelial cells, loss of brush border, and sloughing of epithelial cells in the luminal tubules (Fig 1C). No intranuclear viral inclusions were seen. Immunofluorescence revealed no immune deposits. Electron microscopy showed diffuse effacement of podocyte foot processes associated with enlarged podocytes and tubulovillous transformation (Fig 1D). No immune-type electron-dense deposits, tubuloreticular inclusions, or viral particles were identified. In situ hybridization was negative for SARS-CoV-2 RNA using RNAscope (ACD Bio) in the biopsy specimen (Fig 1E and F).
Figure 1.
Kidney biopsy of coronavirus disease 2019 (COVID-19)–associated collapsing glomerulopathy case. (A) Representative light microscopy shows collapse of glomerular tufts with hyperplasia of epithelial cells (hematoxylin-eosin; original magnification, ×400). (B) Another glomerulus shows collapse of capillary loops with proliferation of overlying epithelial cells (Jones methenamine silver stain; original magnification, ×200). (C) Tubules show widespread attenuation of epithelial cells with drop-out nuclei and sloughing of epithelial cells inside the lumen of tubules (arrows) (hematoxylin-eosin; original magnification, ×100). (D) Electron microscopy reveals diffuse effacement of foot processes (arrow) and hypertrophy of podocytes with tubulovillous transformation (Transmission electron microscopy [TEM]; original magnification, ×2,000). (E) In situ hybridization for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) RNA is negative in COVID-19–associated collapsing glomerulopathy (original magnification, ×200). (F) In contrast, in situ hybridization is positive for SARS-CoV2 RNA in lung epithelia of another patient with COVID-19 infection (original magnification, ×200).
APOL1 genotyping revealed the patient to be homozygous for the G1 allele (G1/G1). We then performed whole-genome sequencing on genomic DNA from whole-blood samples. Whole-genome sequencing yielded 123,266.00 Mb raw bases. After removing low-quality reads, we obtained an average of 820,304,512 clean reads (123,045.68 Mb). The clean reads of each sample had high base calling rates (Q20 and Q30), indicating high sequencing quality. The average guanine-cytosine content was 41.05%. We then used a curated gene/variant list23 and scanned all variants. We detected 4 indel variants (Table S1), of which 3 (FOXC1, LFNG, and RTTN) were annotated as of uncertain significance and 1 (SALL1) was annotated as benign in ClinVar.
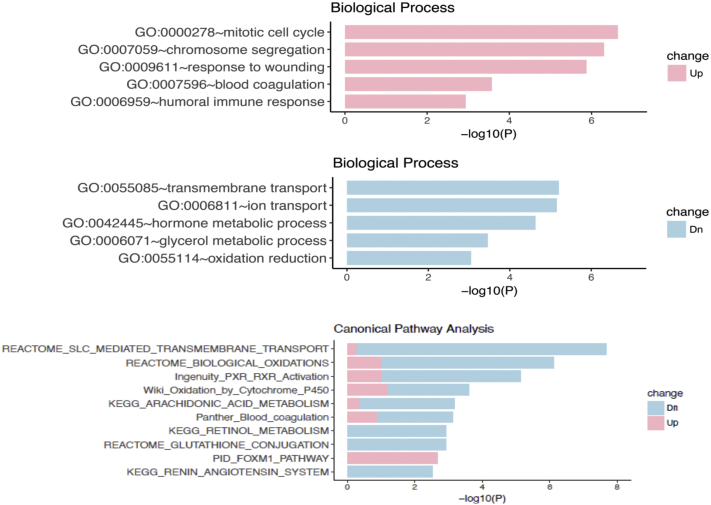
Using data from 12 normal kidney samples extracted from GTEx (Genotype-Tissue Expression) to serve as controls, RNA sequencing data revealed that in COVID-19–associated collapsing glomerulopathy, biological processes from upregulated genes were enriched for cell cycle, chromosome segregation, response to wounding, humoral immune response, and blood coagulation, suggesting that cell injury/regeneration, inflammatory response, and endothelial injury were the major disease processes involved (Fig 2). The biological processes from downregulated genes were enriched for ion transport, metabolic processes, and oxidation, likely secondary to severe tubular cell injury. Pathway analysis from both up- and downregulated genes showed enrichment of transmembrane transport, oxidation, and blood coagulation consistent with the Gene Ontology term enrichment. Upregulated genes (Fig 2; pink) were enriched only for the FOXM1 pathway, which was recently reported to promote tubular cell proliferation during injury repair.24 Additionally, genes related to the renin-angiotensin system were downregulated, but ACE2 expression did not differ from normal controls. Though COVID-19–infected patients display elevated levels of circulating cytokines and interferon can stimulate APOL1 expression,19,25 there were no differences in APOL1 messenger RNA levels between this patient and normal controls. Raw RNA sequencing reads were aligned to SARS-CoV-2 but no mapped reads were found, indicating the absence of SARS-CoV-2 in this biopsy sample.
Figure 2.
RNA sequencing analysis of the kidney cortex in coronavirus disease 2019 (COVID-19)–associated collapsing glomerulopathy. Differentially expressed genes with 1.5-fold–magnitude changes between expression profiles of 1 patient with COVID-19–associated collapsing glomerulopathy and data from 12 control healthy individuals downloaded from GTEx (Genotype-Tissue Expression) database were used for Gene Ontology (GO) term enrichment (GO) terms and pathway analysis by Fisher exact test with P <0.05. Upregulated genes were enriched for cell cycle, chromosome segregation, response to wounding, blood coagulation, and humoral immune response, whereas downregulated genes were enriched for ion transport, metabolic processes, and oxidation. Pathway analysis from both up- and downregulated genes showed enrichment of transmembrane transport, oxidation, and blood coagulation consistent with the GO terms analysis. Upregulated genes were enriched only for FOXM1 pathway, while genes related to renin-angiotensin system were downregulated.
Immunostaining revealed that expression of phosphorylated STAT3, a downstream target of IL-6 signaling, was found to be significantly increased in glomeruli of COVID-19–associated collapsing glomerulopathy and HIVAN as compared with normal kidney tissue (Fig 3).
Figure 3.
Phospho-STAT3 (signal transducer and activator of transcription 3) expression in coronavirus disease 2019 (COVID-19)–associated collapsing glomerulopathy (CG) and HIV-associated nephropathy (HIVAN). Immunohistochemistry staining of kidney biopsy specimen reveals phospho-STAT3 expression is significantly increased in podocytes and proximal tubular cells in COVID-19–associated CG and HIVAN as compared with normal kidney tissue (scale bars, 50 μm).
Discussion
We present a case of collapsing glomerulopathy in a patient of African ancestry with acute COVID-19 infection who was subsequently identified as homozygous for the high-risk APOL1 G1 allele.11, 12, 13 These findings support a growing body of evidence for a strong association of APOL1 risk alleles with COVID-19–associated collapsing glomerulopathy. Though in vitro studies have demonstrated that interferon and other virally mediated inflammatory factors induce APOL1 expression in podocytes,19,20 we demonstrate comparable levels of APOL1 gene transcript between COVID-19–associated collapsing glomerulopathy and controls.
There is conflicting evidence as to whether SARS-CoV-2 can directly infect podocytes and renal tubular epithelial cells, as HIV-1 can.15,26,27 Two published autopsy series both demonstrated positive immunostaining for SARS-CoV-2 in podocytes and 1 of these additionally reported the isolation of SARS-CoV-2 RNA from kidney glomeruli.2,4 However, published in situ hybridization studies, including ours, have failed to detect SARS-CoV-2 RNA in glomeruli.5, 6, 7,9,12 In addition, we confirmed that viral RNA was absent from our patient’s biopsy specimen by RNA sequencing. Autopsy and biopsy series have also reported coronavirus-like particles with characteristic spikes in podocytes and tubular epithelial cells using electron microscopy, though concern has been raised that these particles may actually be small cellular vesicles instead of viral particles.2,3,11,28
Our study also demonstrated increased phosphorylation of STAT3 in COVID-19–associated collapsing glomerulopathy. We have previously shown that STAT3 activation is important in the pathogenesis of HIVAN and diabetic kidney disease.29,30 It is well-known that plasma IL-6 level elevations play a key role in the pathogenesis of COVID-19–induced organ damage25 and also that IL-6–induced activation of STAT3 upregulates inflammatory pathways. Our data suggest that STAT3 phosphorylation could be a major mechanism involved in COVID-19–associated kidney injury, and thus targeting STAT3 with available drug inhibitors31 could be a potential therapeutic strategy for patients with this disease.
To our knowledge, this is the first report of whole-genome sequencing and RNA sequencing data from a patient with COVID-19–associated kidney disease. Although the clinical significance and potential interaction with APOL1 of the gene variants identified by whole-genome sequencing remain unclear, Foxc1 and Sall1 are known to regulate the integrity of the glomerular filtration barrier in experimental models.32,33 Our RNA sequencing results also reveal several interesting findings: (1) we confirmed that SARS-CoV-2 RNA was absent in the kidney, (2) we found that ACE2 and APOL1 gene expression did not differ between COVID-19–infected and normal kidneys, and (3) we identified several injury pathways in the COVID-19–infected kidney that have been associated with SARS-CoV-2–induced cell injury. Our findings in addition to a recent NanoString analysis of 6 COVID-19–associated collapsing glomerulopathy biopsy cases showing upregulation of chemokine gene expression and altered expression of genes related to tubular injury add to the growing body of literature that seeks to elucidate the pathogenic mechanisms of COVID-19–associated kidney injury.
Our study and associated analysis have limitations. First, we have reported just 1 case of COVID-19–associated collapsing glomerulopathy and are only able to describe fold changes without formal statistical analysis. Second, the kidney biopsy for our patient was delayed until 35 days after her initial presentation of acute COVID-19 infection and thus the gene expression profile captured may not accurately represent acute COVID-19–induced kidney injury pathways. Additionally, it is possible that APOL1 gene transcript expression was modulated by the anti-inflammatory effects of the hydroxychloroquine administered. Furthermore, we were not able to separate glomeruli from tubulointerstitial compartments before RNA sequencing. Still, these findings offer important additional insight into the complex interplay between genetic susceptibility, viral infection, and glomerular disease progression associated with COVID-19 infection. Further studies involving additional patients with COVID-19–associated collapsing glomerulopathy who undergo biopsy earlier in the course of their disease will need to be conducted.
Article Information
Authors’ Full Names and Academic Degrees
Kristin Meliambro, MD, Xuezhu Li, MD, Fadi Salem, MD, Zhengzi Yi, MS, Zeguo Sun, MS, Lili Chan, MSCR, Miriam Chung, MD, Jorge Chancay, MD, Ha My T. Vy, PhD, Girish Nadkarni, MD, MS, Jenny S. Wong, PhD, Jia Fu, MD, Kyung Lee, PhD, Weijia Zhang, PhD, John C. He, PhD, and Kirk N. Campbell, MD.
Author Contributions
KM and XL contributed equally to this work.
Support
KM is supported by DK113281. KNC is supported by DK103022 and DK122807.
Financial Disclosure
Dr He reports advisory board fees from Renalytix AL and grants from Shangpharma outside the submitted work. Dr Campbell reports consulting agreements with Retrophin, Aurinia, Calliditas, and Vertex and grants from Goldfinch and Mallinckrodt Pharmaceuticals outside the submitted work. Dr Nadkarni reports grants, personal fees, and nonfinancial support from Renalytix AI, nonfinancial support from Pensieve Health, and personal fees from AstraZeneca, BioVie, and GLG Consulting outside the submitted work. The remaining authors declare that they have no relevant financial interests.
Acknowledgements
We thank Stuart Scott, Yoshinori Seki, and Ping Lu of the Sema4 lab for performing APOL1 genotyping on this patient.
Patient Protections
The authors declare that they have obtained consent from the patient reported in this article for publication of the information about her that appears within this Case Report and any associated supplementary material.
Peer Review
Received November 4, 2020, as a submission to the expedited consideration track with 2 external peer reviews. Direct editorial input from an Associate Editor and the Editor-in-Chief. Accepted in revised form February 15, 2021.
Footnotes
Complete author and article information provided before references.
Item S1: Detailed Methods
Table S1: Indel variants detected with whole genome sequencing
Contributor Information
John C. He, Email: Cijiang.he@mssm.edu.
Kirk N. Campbell, Email: kirk.campbell@mssm.edu.
Supplementary Material
Item S1, Table S1
References
- 1.Chan L., Chaudhary K., Saha A. AKI in hospitalized patients with COVID-19. J Am Soc Nephrol. 2021;32(1):151–160. doi: 10.1681/ASN.2020050615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Su H., Yang M., Wan C. Renal histopathological analysis of 26 postmortem findings of patients with COVID-19 in China. Kidney Int. 2020;98(1):219–227. doi: 10.1016/j.kint.2020.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Farkash E.A., Wilson A.M., Jentzen J.M. Ultrastructural evidence for direct renal infection with SARS-CoV-2. J Am Soc Nephrol. 2020;31(8):1683–1687. doi: 10.1681/ASN.2020040432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Puelles V.G., Lutgehetmann M., Lindenmeyer M.T. Multiorgan and renal tropism of SARS-CoV-2. N Engl J Med. 2020;383(6):590–592. doi: 10.1056/NEJMc2011400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sharma P., Uppal N.N., Wanchoo R. COVID-19-associated kidney injury: a case series of kidney biopsy findings. J Am Soc Nephrol. 2020;31(9):1948–1958. doi: 10.1681/ASN.2020050699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kudose S., Batal I., Santoriello D. Kidney biopsy findings in patients with COVID-19. J Am Soc Nephrol. 2020;31(9):1959–1968. doi: 10.1681/ASN.2020060802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wu H., Larsen C.P., Hernandez-Arroyo C.F. AKI and collapsing glomerulopathy associated with COVID-19 and APOL 1 high-risk genotype. J Am Soc Nephrol. 2020;31(8):1688–1695. doi: 10.1681/ASN.2020050558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Golmai P., Larsen C.P., DeVita M.V. Histopathologic and ultrastructural findings in postmortem kidney biopsy material in 12 patients with AKI and COVID-19. J Am Soc Nephrol. 2020;31(9):1944–1947. doi: 10.1681/ASN.2020050683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Santoriello D., Khairallah P., Bomback A.S. Postmortem kidney pathology findings in patients with COVID-19. J Am Soc Nephrol. 2020;31(9):2158–2167. doi: 10.1681/ASN.2020050744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Jhaveri K.D., Meir L.R., Flores Chang B.S. Thrombotic microangiopathy in a patient with COVID-19. Kidney Int. 2020;98:509–512. doi: 10.1016/j.kint.2020.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kissling S., Rotman S., Gerber C. Collapsing glomerulopathy in a COVID-19 patient. Kidney Int. 2020;98(1):228–231. doi: 10.1016/j.kint.2020.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Peleg Y., Kudose S., D’Agati V. Acute kidney injury due to collapsing glomerulopathy following COVID-19 infection. Kidney Int Rep. 2020;5(6):940–945. doi: 10.1016/j.ekir.2020.04.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Larsen C.P., Bourne T.D., Wilson J.D., Saqqa O., Sharshir M.A. Collapsing glomerulopathy in a patient with coronavirus disease 2019 (COVID-19) Kidney Int Rep. 2020;5(6):935–939. doi: 10.1016/j.ekir.2020.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chandra P., Kopp J.B. Viruses and collapsing glomerulopathy: a brief critical review. Clin Kidney J. 2013;6:1–5. doi: 10.1093/ckj/sft002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cohen A.H., Sun N.C., Shapshak P., Imagawa D.T. Demonstration of human immunodeficiency virus in renal epithelium in HIV-associated nephropathy. Mod Pathol. 1989;2:125–128. [PubMed] [Google Scholar]
- 16.Besse W., Mansour S., Jatwan K., Nast C.C., Brewster U.C. Collapsing glomerulopathy in a young woman with APOL1 risk alleles following acute parvovirus B19 infection: a case report investigation. BMC Nephrol. 2016;17:125. doi: 10.1186/s12882-016-0330-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Markowitz G.S., Nasr S.H., Stokes M.B., D’Agati V.D. Treatment with IFN-{alpha}, -{beta}, or -{gamma} is associated with collapsing focal segmental glomerulosclerosis. Clin J Am Soc Nephrol. 2010;5:607–615. doi: 10.2215/CJN.07311009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kopp J.B., Nelson G.W., Sampath K. APOL1 genetic variants in focal segmental glomerulosclerosis and HIV-associated nephropathy. J Am Soc Nephrol. 2011;22:2129–2137. doi: 10.1681/ASN.2011040388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nichols B., Jog P., Lee J.H. Innate immunity pathways regulate the nephropathy gene apolipoprotein L1. Kidney Int. 2015;87:332–342. doi: 10.1038/ki.2014.270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Beckerman P., Bi-Karchin J., Park A.S. Transgenic expression of human APOL1 risk variants in podocytes induces kidney disease in mice. Nat Med. 2017;23:429–438. doi: 10.1038/nm.4287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Olabisi O.A., Zhang J.Y., VerPlank L. APOL1 kidney disease risk variants cause cytotoxicity by depleting cellular potassium and inducing stress-activated protein kinases. Proc Natl Acad Sci U S A. 2016;113:830–837. doi: 10.1073/pnas.1522913113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ma L., Chou J.W., Snipes J.A. APOL1 renal-risk variants induce mitochondrial dysfunction. J Am Soc Nephrol. 2017;28:1093–1105. doi: 10.1681/ASN.2016050567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Steers N.J., Li Y., Drace Z. Genomic mismatch at LIMS1 locus and kidney allograft rejection. N Engl J Med. 2019;380:1918–1928. doi: 10.1056/NEJMoa1803731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chang-Panesso M., Kadyrov F.F., Lalli M. FOXM1 drives proximal tubule proliferation during repair from acute ischemic kidney injury. J Clin Invest. 2019;129:5501–5517. doi: 10.1172/JCI125519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.McGonagle D., Sharif K., O’Regan A., Bridgewood C. Re: Patiently waiting for the results of anti-IL 6 therapy in severe COVID-19 infection [letter] Autoimmun Rev. 2020;19:102560. doi: 10.1016/j.autrev.2020.102560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lu T.C., He J.C., Wang Z.H. HIV-1 Nef disrupts the podocyte actin cytoskeleton by interacting with diaphanous interacting protein. J Biol Chem. 2008;283:8173–8182. doi: 10.1074/jbc.M708920200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Winston J.A., Bruggeman L.A., Ross M.D. Nephropathy and establishment of a renal reservoir of HIV type 1 during primary infection. N Engl J Med. 2001;344:1979–1984. doi: 10.1056/NEJM200106283442604. [DOI] [PubMed] [Google Scholar]
- 28.Smith K.D., Akilesh S., Alpers C.E., Nicosia R.F. Am I a coronavirus? Kidney Int. 2020;98(2):506–507. doi: 10.1016/j.kint.2020.05.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.He J.C., Husain M., Sunamoto M. Nef stimulates proliferation of glomerular podocytes through activation of Src-dependent Stat3 and MAPK1,2 pathways. J Clin Invest. 2004;114:643–651. doi: 10.1172/JCI21004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lu T.C., Wang Z.H., Feng X. Knockdown of Stat3 activity in vivo prevents diabetic glomerulopathy. Kidney Int. 2009;76:63–71. doi: 10.1038/ki.2009.98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Brosius F.C., Tuttle K.R., Kretzler M. JAK inhibition in the treatment of diabetic kidney disease. Diabetologia. 2016;59:1624–1627. doi: 10.1007/s00125-016-4021-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Motojima M., Kume T., Matsusaka T. Foxc1 and Foxc2 are necessary to maintain glomerular podocytes. Exp Cell Res. 2017;352:265–272. doi: 10.1016/j.yexcr.2017.02.016. [DOI] [PubMed] [Google Scholar]
- 33.Hosoe-Nagai Y., Hidaka T., Sonoda A. Re-expression of Sall1 in podocytes protects against adriamycin-induced nephrosis. Lab Invest. 2017;97:1306–1320. doi: 10.1038/labinvest.2017.69. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Item S1, Table S1