Abstract
In modern biology, the correct identification of cell types is required for the developmental study of tissues and organs and the production of functional cells for cell therapies and disease modeling. For decades, cell types have been defined on the basis of morphological and physiological markers and, more recently, immunological markers and molecular properties. Recent advances in single-cell RNA sequencing have opened new doors for the characterization of cells at the individual and spatiotemporal levels on the basis of their RNA profiles, vastly transforming our understanding of cell types. The objective of this review is to survey the current progress in the field of cell-type identification, starting with the Human Cell Atlas project, which aims to sequence every cell in the human body, to molecular marker databases for individual cell types and other sources that address cell-type identification for regenerative medicine based on cell data guidelines.
Subject terms: Stem-cell research, Differentiation, Translational research, Classification and taxonomy, Stem-cell differentiation
Regenerative medicine: Precise cell identification will improve future therapies
The characterization of cells into more precise groups will improve the chances of selecting the best types of cells to use for regenerative medicine. Traditionally, cell types are defined by physiological and morphological markers and molecular properties. However, sequencing technologies are enabling researchers to classify cells into increasingly distinct subgroups. Wataru Fujibuchi at Kyoto University, Japan, and co-workers reviewed progress in cell-type identification to help guide regenerative medicine. They examined the insights gained from the Human Cell Atlas project, an international collaboration of over 1000 institutes across 71 countries. Participants in this project are sequencing cellular RNA and categorizing hundreds of thousands of individual cells, demonstrating that many assumptions about cells are too simplistic. Such data will inform regenerative therapies, for example, by selecting the “best” platelet-producing stem cells for blood donation.
The Human Cell Atlas
The Human Cell Atlas (HCA) project aims to characterize all cells by single-cell analytical techniques, specifically single-cell RNA sequencing (scRNA-seq) and assay for transposase accessible chromatin sequencing (scATAC-seq) and to link this information to classic knowledge, namely, location, lineage, and type, as well as cell states, state transitions, and cell−cell interactions. The HCA was initiated to unify scRNA-seq data in a manner similar to the data compilation achieved in the Human Genome Project1. As of February 2020, the HCA project has participation from 1027 institutes in 71 countries, from which 81 laboratories have already posted scRNA-seq data for 34 organs and tissues, including the liver2, lung3, blood and immune systems4, plasma cells5, human cortex6, colon7, and retina8, with the total number of sequenced cells reaching 4.5 million. The HCA is an open science project, and its standard operating protocols (SOPs) are available on the Web. A standardized data analysis service called the Data Coordination Platform (DCP) is provided to the community. In the long term, the HCA project aims to define not only normal cells but also cells in specific disease states1,9.
The HCA project is generating tremendous amounts of data. For example, there are currently 148 members in the immune system group, which is conducting approximately 100 projects and is expected to include more than 1000 projects over time. One project10 has produced scRNA-seq data for 530,000 cells. The challenge for the HCA project is to create a universal classification system for all of these projects. Adding to this challenge are regional HCAs, such as HCA Asia, that focus on regional diseases. To assign cell types to all these cells, exhaustive analysis methods are needed. Once this goal is accomplished, the HCA can shift to its second goal, which is to understand how normal cell states transition into disease states11.
Challenge of human cell-type classifications in the HCA
A typical scheme for assigning cell types to scRNA-seq data requires computational analysis and a workflow consisting of preprocessing and downstream analysis steps. Preprocessing is necessary to correct biological and technical noise caused by experimental errors and sample variabilities, thus contributing to quality control and normalization, whereas downstream analysis aims to cluster single-cell data on the basis of similarities in profiles and assign cell types using various computational tools depending on the data type12,13. Each cell type can be identified by a uniquely characteristic transcript pattern of gene expressions (i.e., RNA signature) or a uniquely characteristic transcript profile14. However, because the number of clusters depends on the algorithms and parameters, the results are not generally optimized for separating subtypes or states of the same cell type (e.g., cell cycle phase differences)15.
In this regard, several clustering approaches have been developed, including partition type (such as k-means and self-organizing maps), hierarchical clustering, graph-based (such as spectral clustering or clique detection), mixture model (such as Seurat), density-based, neural network type, ensemble, and affinity propagation type16–18. However, most of these approaches require parameter settings, and to complicate matters, some depend on random values. Consequently, the resulting clusters could vary in number and cluster members. Detailed reviews of these approaches can be found in recent papers15,19. In addition, a variety of software tools are available for clustering approaches, and the number is constantly growing.
Alternatively, the identification of a known cell type by its RNA signature can be performed using a cell−cell similarity search. In this case, researchers can identify known cell types and find similar or related unknown cell types by using hitherto unknown relationships. Software programs for cell−cell similarity search analyses include CellAtlasSearch20, CellSim21, and Cell BLAST22, all of which are tailored to different needs. For example, Cell BLAST has a “special tuning” mode for handling batch effects between a query and reference. CellSim calculates the similarity of different cells on the basis of cell ontology and molecular networks and has a feature that allows users to identify the cell type by entering a list of genes. Finally, CellAtlasSearch is tailored for handling ultra-large RNA-seq data through parallel screening of tens of thousands of single cells using efficient clustering methods. However, these methods are typically dependent on existing cell data and thus are effective for cell-type identification that demands large amounts of annotated cell-type entries.
Another approach for assigning cell types is the integration of gene expression information and spatial information of the individual cells to provide a molecular description of each cell type in the context of the tissue microenvironment. Recently, developed computational methods allow, in principle, the reconstruction of a spatial map of tissues using scRNA-seq data23–25. scRNA-seq-based maps have revealed cell-type-specific functions in the liver26, blastocyst27, and growth plate28. The expression of cell adhesion genes and specific gene functions defined by Gene Ontology (GO) terms is being used to develop new tools for single-cell 3D transcriptome analysis that enhance spatial prediction29,30. Although the identification of cell types in a spatial context is expected to yield more information relevant to the in vivo environment, these cutting-edge approaches are still at the elementary stage and need further improvement before they can be widely used.
To expedite methods for cell-type identification, the HCA project has multiple committees and working groups that coordinate the efforts of independent research groups and unify the results of scRNA-seq. Typically, the research groups perform all the data preparation, acquisition, and analysis, and the HCA committees provide the general framework, guidelines, and data repository space. The DCP team supplies vetted algorithms for data processing on the portal, and the research groups choose the algorithms for data processing, depositing the data into the HCA database under controlled pipelines. Hence, the cell-type classification, which is part of the RNA-seq data analysis, is performed separately by various groups, with the results processed by the cooperating DCP team according to HCA guidelines to ensure consistency in cell-type assignment and authentication.
History of cell-type classification
With the advances in stem cell research that have made it possible to engineer cells for cell therapies and drug discovery, the identification and authentication of cell types have become emerging priorities in the biological community31. The greatest problem in the authentication of human cells lies in the lack of integrated standard metrics for cell morphology, gene expression, and molecular markers32. Considering that scRNA-seq data are usually confounded by a high degree of noise33 and that current scientific communities use highly variable methodologies for scRNA-seq data analysis34, it is very difficult to standardize the data and harmonize it with classical morphology and marker maps. Accordingly, the HCA project has invested both time and resources to study ways of handling technical and biological noise affecting data reproducibility and the degradation of biopsy samples35.
Historically, the characterization of cell types was based on histological, i.e., anatomical, morphological, and functional, criteria36. For example, the earliest attempt to classify cells in the nervous system was based on histological characteristics, such as the locations from which the cells were obtained, cell morphology, and the presence of certain molecular markers37. Since then, the location (i.e., cerebral cortex, cardiac muscle, or stomach), cell morphology (i.e., fibroblast-like or epithelial-like), and molecular markers (such as CD75-positive cells, etc.) have been accepted by the scientific community as the three main pillars for defining cell types38–41. To accelerate cell-type identification, several cell marker databases are available (Table 1).
Table 1.
Online cell marker databases as of November 1, 2019.
| Database | No. of markers | Cell type | Species | Characteristic | PMID | Address |
|---|---|---|---|---|---|---|
| Labome | 226 | 7 major cell types and several others | Human | No distinction between diseased or healthy states. Few cell types are covered compared with other databases in this table. Source references are provided. No search engine. Only experimentally confirmed markers are included. This collection is most useful for searching verified markers. | N/A42 | https://www.labome.com/method/Cell-Markers.html |
| CellFinder | 553,905 | 3394 cell types and 50,951 cell lines | Human, mouse | Omics data are prioritized. In many cases, users must analyze raw data. No distinction between verified markers and ambiguous markers. This database is most useful for omics data exploration. | PMC3965082 43 | http://www.cellfinder.org/ |
| CellMarker | 22,753 | 467 human cell types and 389 mouse cell types | Human, mouse | A major portion of markers are for cancer cells. No distinction between experimentally confirmed markers and ones gathered through omics. No distinction between verified markers and ambiguous markers. References are provided. This database is recommended for a first search of information. | PMC6323899 44 | http://biocc.hrbmu.edu.cn/CellMarker/ |
| PanglaoDB | 8230 | 178 cell types | Human, mouse | Markers for healthy and diseased adult and embryonic cells. Source references are not provided for the markers. Contains single-cell sequencing data and automatically updates the list of papers on single-cell sequencing. Because of the relatively small number of cell types, this database is good for narrowing down the list of markers given by other databases. | PMC6450036 45 | https://panglaodb.se/markers.html |
| SHOGoiN | 2594 | 740 cell types | Human | Markers for healthy human cells including embryonic cells. Source references are provided. No distinction between verified markers and ambiguous markers. This database is most useful for a quick search by cell type. | PMC3204613 81 | https://stemcellinformatics.org/cell_marker/literatures |
Labome provides a list of 226 markers for epithelial, dendritic, glial, bone marrow, natural killer, and other cell types42. CellFinder was the first database website of molecular markers and now features information on 3394 cell types, 50,951 cell lines, and 553,905 protein expressions43. However, CellFinder data are diverse for species (mammals, fish, invertebrates, bacteria, viruses, plants, etc.), and the site includes other data, such as microscopic and anatomical images, whole-genome expression profiles from RNA-seq and microarrays, etc. In other words, it is not a cell marker database per se but a collection of various data of different cell types, which makes looking for markers relatively difficult compared to the search in other databases.
The first database compiled exclusively with markers of human and mouse cells was CellMarker44, which currently features 13,605 cell markers for 467 cell types in 158 human tissue and 9148 cell markers for 389 cell types in 81 mouse tissues. The gene expression data in CellMarker originate from scRNA-seq studies, experimental studies, and microarrays. At approximately the same time, PanglaoDB was published45, providing data on 8230 markers collected from human and mouse scRNA-seq experiments, along with other types of data. The markers are grouped by cell type (178 cell types, 4644 genes, and 29 tissues), and the cell types are subsequently grouped into 26 organs and 3 germ layers. A typical cell type has 28 (median) gene markers, but some, such as fibroblasts, have more than 100 markers. The major drawback of this database is that it lacks the source from which the marker information was obtained and information on how the marker was originally found and used, thus making information verification impossible.
Limitations of the current methods of cell-type classification
Despite the abovementioned databases, the new era of single-cell sequencing, which has revealed that markers can be expressed in different cells or at varying levels when cells are cultured in vitro, demands the reconsideration of marker-based cell-type classification methods. For example, the identification of mesenchymal stromal cells (MSCs) and the results of fate mapping in vivo have been problematic46. In addition, many markers often correspond to several cell types rather than a unique one, which may be related to the fact that MSCs encompass a number of different tissue-specific progenitor or stem cells. In other words, some markers are of mixed cell types47,48. Moreover, the expression levels of some markers make it difficult to discriminate among cell types49. For example, mature monocytes are usually characterized by the expression of CD33, CD11b, CD14, HLA-DR, and CD16, whereas granulocytes are characterized by the expression of CD33, CD11b, CD15, and CD66b. However, CD15 is expressed at low levels on monocytes in some anti-CD15 clones. Conversely, in disease states, CD14 can be variably expressed in neutrophils50. Notably, current markers all share the common feature that they are expressed on the cell surface. Intracellular markers such as microRNA (miRNA) could enhance the specific marker profile of a given cell type51. The combination of surface and intracellular markers for cell-type classification has yet to be fully developed in any database.
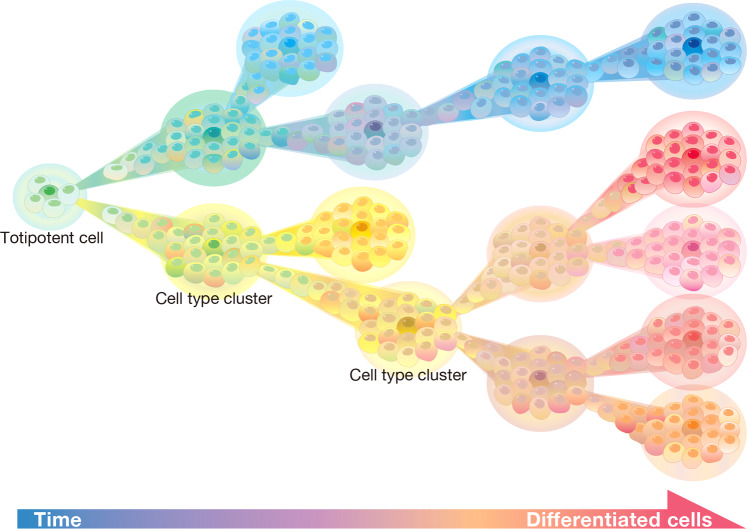
Another problem is the heterogeneity of cell states. In some cases, heterogeneity, such as that during states of immaturity or senescence, blurs discrete states, leaving a continuous spectrum that greatly complicates cell-type classification (Fig. 1). In such cases, cell types may seem to have hundreds of variants52,53. This lack of clarity is a major issue in clinical applications using cell therapies, for which cells are cultured in vitro to generate and stabilize a specific cell type. For example, for Parkinson’s disease, the differentiation of pluripotent stem cells (PSCs) into specific neural cell types for cell therapies demands extremely precise markers54. As with their use as intercellular markers, the application of miRNA would be helpful. Another important problem involves changes in cell behavior and cell composition under different conditions or even within the same culture. These changes compromise stable cell phenotypes across cell populations over the course of an experiment, leading to confusing experimental interpretations55.
Fig. 1. Renewed concept of cell types that takes into account the cell state continuum and diversity, as well as the state stability, of a cell type.
Classically, the continuum was thought to be unidirectional, but cell reprogramming induces cells to regress to a previous state and/or take a different trajectory.
Emerging concepts for cell-type classification
Accordingly, several new concepts for cell-type authentication have been proposed. Evolutionary biologists have developed a classification on the basis of the evolution of gene expression states56. In this proposition, the cell type is defined by a set of changes in the “core regulatory complex” (CoRC) of transcription factors that regulate cell-type-specific traits. From this point of view, cell types are defined by evolutionary units that differ according to their evolutionary lineages rather than their phenotypic similarities and are characterized by their ability to evolve gene expression states independently of each other. Thus, a gene regulatory network that defines the cell type would include “master” transcription factors that control downstream effector genes57–59. A classic example is the transformation of fibroblasts into skeletal muscle cells by the forced expression of the myoblast determination protein (MYOD) transcription factor60. The discovery of induced pluripotent stem cells (iPSCs) cemented the notion that master transcription factors can determine cell type because, through reprogramming to iPSCs and their subsequent differentiation, nearly any cell type can be converted into any other cell type61,62. Later works in neuronal lineages63, the transcription factor competition observed in embryonic stem cells64, and pancreatic cell transdifferentiation65 support this idea.
Another evolutionary approach for cell-type classification is the construction of a hierarchy to describe the relationships between cells, which is analogous to how taxonomy hierarchies are created to describe the relationships between species66. Based on this approach, we proposed a “periodic table” for cell types67. This proposal aims to distinguish cell types from cell states, in which the periods and groups correspond to developmental trajectories and stages of differentiation68. scRNA-seq has paved the way for new interpretations of cell states. For example, in the epigenetic landscapes described by Waddington69, it was originally assumed that cell states follow continuous trajectories that branch at cell-fate decision points, but these decision points have since been refined into transition states70. Whereas Waddington assumed that the decision points were deterministic, the transition states in the modernized version are stochastic and the related signaling networks are probabilistic. This concept is meant to accommodate the gene expression heterogeneity encountered in real-world single-cell data. Indeed, the identification and characterization of cell transition states is one of the biggest challenges in single-cell transcriptomics71. Dimension reduction techniques, such as principal components analysis (PCA)72 and t-distributed stochastic neighbor embedding (tSNE)73, and graph and community detection algorithms, such as consensus clustering74, SNN-Cliq75, and Seurat18, have been developed and utilized to identify cell transition states. Fully comprehending the influence of cell transition states on cell types at the single-cell level will require both new tools and parameters76. Finally, although the results are still in the rudimentary stage, recent research has made automated cluster annotation available for unbiased cell-type identification. This type of annotation is rapid and allows investigators to forgo the manual clustering step by combining annotation and clustering. Numerous tools based on various algorithms have been developed to facilitate automated cell identification methods (for a detailed review, see Abdelaal et al.77).
Proposing a data-driven cell-type definition
To help consolidate the many opinions about cell-type classification and provide data-related guidelines for cell-type authentication for clinical application, the International Cell Type Authentication Committee (ICTAC) was created (https://cell-type.org/). The ICTAC was launched to establish criteria and processes for defining, determining, and authenticating all human cell types. Its mission is to help scientific communities identify cell types and provide systematic information on cell classifications. The ICTAC originated from the International Stem Cell Banking Initiative (ISCBI) (https://www.iscbi.org/)78, an organization that focuses on practical issues in cell banking and regenerative medicine. More than 300 stem cell and policy professionals from 28 countries are part of the ISCBI community and are working together to advance stem cell research and biobanking along with developing regulations and public policy. The ISCBI is managed by an executive board, with delegates and steering group members of the community closely collaborating with the International Human Pluripotent Stem Cell Registry (hPSCreg)79,80. The first task of the ICTAC is to integrate existing cell databases, such as SHOGoiN81, CellFinder43, and Cell Ontology82, to comprehend information on existing cell-type definitions. One of the key functions of the ICTAC is to provide a tool that can process the massive amount of accumulating single-cell data to classify new cell types that do not fit into existing definitions or cannot be identified by cell-matching software, such as CellSim, CellAtlasSearch, or Cell BLAST.
The ICTAC proposed the concept of reference cell types (RCTs), which are defined by an integrative examination of core properties (species, physiological system, source age, and markers) and additional attributes (functions including potency, morphology, developmental origin, omics, and environmental conditions) and are constantly updated by experts of relevant tissues and organs. RCTs provide a framework for identifying and authenticating new and known cell types. Ultimately, RCTs are designed to support cell-type classifications in various communities, including stem cell banking initiatives and massive-scale single-cell sequencing projects.
Cell type authentication for regenerative medicine
In the field of regenerative medicine, PSCs such as the aforementioned iPSCs have tremendous potential because of their capacity to differentiate into most cell types in the human body. Moreover, recent technological progress has made it possible to produce PSCs on a large scale83 and generate significant resources for a large number of well-characterized and documented PSCs (e.g., https://fujifilmcdi.com/the-cirm-ipsc-bank, https://ebisc.org84). However, in addition to scalability, the clinical translation of PSCs depends on fast and reliable ways to assess quality and safety in terms of three important properties: cellular identification, differentiation potency, and malignant potential85.
For example, the establishment of PSC-derived platelets as a substitute for primary donor cells can compensate for anticipated donor shortages and are useful against platelet transfusion refractoriness86. In this system, PSCs need to be differentiated into megakaryocytes, which are then cultured in bioreactors to shed platelets87. However, platelet production by iPSC-derived megakaryocytes is heterogeneous, and many biochemical and biophysical approaches have been attempted to enhance and homogenize their production88. Ultimately, the characterization of the best megakaryocytes is lacking. In other words, the existing classification of the megakaryocytes is too broad to identify the cells that are optimal for platelet production, thereby resulting in inefficient platelet production. This problem could be solved by identifying markers associated with platelet-producing megakaryocytes and developing differentiation protocols aimed at the selection of the relevant cell types.
Differentiation efficacy depends on PSC quality. There are several methods for ensuring high PSC quality that are based on assessing the potency of PSC differentiation into cells of the three germ layers. The gold standard is the teratoma assay89,90, in which the differentiation capacity of a PSC line in vivo is assessed by grafting the cells into immunodeficient mice. In addition to differentiation capacity, this assay enables the assessment of viability, histotypic organization, and carcinogenicity at the same time. However, the assay is lengthy and laborious, and it requires experts in pathological assessment and the use of experimental animals91,92. Furthermore, in a comparison of the methods used for and the results obtained from teratoma assays performed at 18 centers worldwide, the ISCBI found that both the test methods and test results varied substantially among expert centers93. Another quality check method is the embryoid body (EB) assay, which enables the monitoring of differentiation capacity in vitro. EBs are cell aggregates that spontaneously differentiate into three developmental germ layers when cultured in suspension94. This approach is more standardized than the teratoma assay and considered more robust by some researchers95. Contributing to its favorability, an EB assay can be combined relatively easily with a bioinformatics analysis of gene expression profiles96–98.
Finally, the detection of pluripotency-specific markers, such as alkaline phosphatase99, Nanog, and Oct4, as well as other mRNAs and proteins, is another way to perform a quality check of PSC characteristics. The detection of these markers is usually performed using flow cytometry, RT-qPCR, and cell-staining techniques100. A number of markers can be used to identify PSC types, such as the naive PSC state or the high-yield expansion PSC state101–104, and to measure the quality of the PSCs105. At the single-cell level, pluripotency can be assessed with scRNA-seq and bioinformatics tools. For example, a PluriTest® assay is used to distinguish typical PSCs from other PSC-like populations through machine learning that is based on the transcriptomes of ideal cell lines and control PSC lines106. However, functional differentiation assays are still required to exclude false-positive results. Other tools, such as SLICE107, SCENT108, and Epi-Pluri-Score109, allow researchers to quantify cell potency and cellular differentiation by entropy analysis. However, a recent multinational study of a range of pluripotency assays concluded that the demonstration of at least some capacity for in vitro or in vivo differentiation was important for the veracity of the results85.
Guidelines for big data generation, storage, and management
As described above, numerous groups are working internationally to generate human omics data not only for tissues and bulk cells but also for millions of individual cells. The gathered data have the potential to greatly advance experimentation practices and quality checks for cells intended for clinical applications. However, for this goal to be realized, the data must be structured, and new algorithms that can efficiently curate the data are needed. Accordingly, members of the ISCBI have proposed Minimum Information About a Cellular Assay for Regenerative Medicine (MIACARM) guidelines, which are directed first and foremost at stem cell banks, although MIACARM can also be used to structure data formats of cellular assays in general110. MIACARM is based on the Minimum Information About a Cellular Assay (MIACA), the first attempt at creating a reporting format for describing functional research on cell lines111. Unlike MIACA, MIACARM sets guidelines for human cells used in medical applications or single-cell analysis, including omics. The proposed guidelines have the potential to enhance information flow in stem cell research that aims to produce clinical-grade cells for therapies. As an extension of MIACARM, which currently targets source cell characterization (MIACARM-I) and stem cell characterization (MIACARM-II), the ISCBI community is engaged in plans to provide guidelines for characterizing differentiated cells (MIACARM-III) using ICTAC proposals for cell authentication based on the framework developed for RCTs.
Conclusion
It is obvious that the body constitutes a myriad of different cell types that have specific functions. Advances in microscopy, histology, and, now, omics technologies have made it clear that the list of cell types is much longer than originally imagined and that our understanding of molecular expressions in systems both in vitro and in vivo is still at the nascent stage. Moreover, the concept of cell reprogramming has taught us that cells can be in constant flux, oscillating between cell types. The ability to define these different cell types is crucial for understanding natural development, including the development of diseased states, and for producing cells for clinical therapies. Critical points are a consensus definition of cell types and the data required for their authentication. In addition, it is vital to ensure accurate and traceable links between precious resources of biological materials and the associated data sets to make full use of both biological and electronic resources and promote reproducibility in scientific data. The many existing databases and the massive data already accumulated affirm the need for the scientific community to work together in creating a universal standard.
Acknowledgements
This work was partially supported by the Core Center for iPS Cell Research, Research Center Network for Realization of Regenerative Medicine (16bm0104001h0004) and the Formulation of Regenerative Medicine National Consortium, which Renders Nation-wide Assistance to Clinical Researches, Project to Build Foundation for Promoting Clinical Research of Regenerative Medicine (19bk0204001h0004), Japan Agency for Medical Research and Development, Grant-in-Aid for Scientific Research on Innovative Areas (17H06392), Ministry of Education, Culture, Sports, Science and Technology (MEXT), and German Academic Exchange Service (DAAD) PPP grant.
Conflict of interest
The authors declare that they have no conflict of interest.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Pullen LC. Human cell atlas poised to transform our understanding of organs. Am. J. Transplant. 2018;18:1–2. doi: 10.1111/ajt.14605. [DOI] [PubMed] [Google Scholar]
- 2.MacParland SA, et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 2018;9:4383. doi: 10.1038/s41467-018-06318-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Reyfman PA, et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 2019;199:1517–1536. doi: 10.1164/rccm.201712-2410OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Popescu D-M, et al. Decoding human fetal liver haematopoiesis. Nature. 2019;574:365–371. doi: 10.1038/s41586-019-1652-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ledergor G, et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 2018;24:1867–1876. doi: 10.1038/s41591-018-0269-2. [DOI] [PubMed] [Google Scholar]
- 6.Hodge RD, et al. Conserved cell types with divergent features in human versus mouse cortex. Nature. 2019;573:61–68. doi: 10.1038/s41586-019-1506-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Smillie CS, et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell. 2019;178:714–730.e22. doi: 10.1016/j.cell.2019.06.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lukowski SW, et al. A single-cell transcriptome atlas of the adult human retina. EMBO J. 2019;38:e100811. doi: 10.15252/embj.2018100811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Regev A, et al. The human cell atlas. Elife. 2017;6:e27041. doi: 10.7554/eLife.27041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rosa, F., Kurochkin, I., Pires, C. & Pereira, F. HCA census of immune cells. https://data.humancellatlas.org/explore/projects/116965f3-f094-4769-9d28-ae675c1b569c (2019).
- 11.Ponting CP. The Human Cell Atlas: making “cell space” for disease. Dis. Model. Mech. 2019;12:dmm037622. doi: 10.1242/dmm.037622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Prakadan SM, Shalek AK, Weitz DA. Scaling by shrinking: empowering single-cell “omics” with microfluidic devices. Nat. Rev. Genet. 2017;18:345–361. doi: 10.1038/nrg.2017.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mezger A, et al. High-throughput chromatin accessibility profiling at single-cell resolution. Nat. Commun. 2018;9:3647. doi: 10.1038/s41467-018-05887-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Saelens W, Cannoodt R, Saeys Y. A comprehensive evaluation of module detection methods for gene expression data. Nat. Commun. 2018;9:1090. doi: 10.1038/s41467-018-03424-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Luecken MD, Theis FJ. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 2019;15:e8746. doi: 10.15252/msb.20188746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li P, et al. The developmental dynamics of the maize leaf transcriptome. Nat. Genet. 2010;42:1060–1067. doi: 10.1038/ng.703. [DOI] [PubMed] [Google Scholar]
- 17.Reeb PD, Bramardi SJ, Steibel JP. Assessing dissimilarity measures for sample-based hierarchical clustering of RNA sequencing data using plasmode datasets. PLoS ONE. 2015;10:e0132310. doi: 10.1371/journal.pone.0132310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018;36:411–420. doi: 10.1038/nbt.4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Petegrosso, R., Li, Z. & Kuang, R. Machine learning and statistical methods for clustering single-cell RNA-sequencing data. Brief. Bioinformatics10.1093/bib/bbz063 (2019). [DOI] [PubMed]
- 20.Srivastava D, Iyer A, Kumar V, Sengupta D. CellAtlasSearch: a scalable search engine for single cells. Nucleic Acids Res. 2018;46:W141–W147. doi: 10.1093/nar/gky421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li L, et al. CellSim: a novel software to calculate cell similarity and identify their co-regulation networks. BMC Bioinform. 2019;20:111. doi: 10.1186/s12859-019-2699-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cao, Z.-J., Wei, L., Lu, S., Yang, D.-C. & Gao, G. Cell BLAST: searching large-scale scRNA-seq database via unbiased cell embedding. BioRxiv10.1101/587360 (2019). [DOI] [PMC free article] [PubMed]
- 23.Satija R, Farrell JA, Gennert D, Schier AF, Regev A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 2015;33:495–502. doi: 10.1038/nbt.3192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Achim K, et al. High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin. Nat. Biotechnol. 2015;33:503–509. doi: 10.1038/nbt.3209. [DOI] [PubMed] [Google Scholar]
- 25.Durruthy-Durruthy R, Gottlieb A, Heller S. 3D computational reconstruction of tissues with hollow spherical morphologies using single-cell gene expression data. Nat. Protoc. 2015;10:459–474. doi: 10.1038/nprot.2015.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Halpern KB, et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature. 2017;542:352–356. doi: 10.1038/nature21065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Durruthy-Durruthy J, et al. Spatiotemporal reconstruction of the human blastocyst by single-cell gene-expression analysis informs induction of naive pluripotency. Dev. Cell. 2016;38:100–115. doi: 10.1016/j.devcel.2016.06.014. [DOI] [PubMed] [Google Scholar]
- 28.Li J, et al. Systematic reconstruction of molecular cascades regulating GP development using single-cell RNA-seq. Cell Rep. 2016;15:1467–1480. doi: 10.1016/j.celrep.2016.04.043. [DOI] [PubMed] [Google Scholar]
- 29.Mori T, et al. Development of 3D tissue reconstruction method from single-cell RNA-seq data. Genomics Comput. Biol. 2017;3:53. [Google Scholar]
- 30.Mori T, Takaoka H, Yamane J, Alev C, Fujibuchi W. Novel computational model of gastrula morphogenesis to identify spatial discriminator genes by self-organizing map (SOM) clustering. Sci. Rep. 2019;9:12597. doi: 10.1038/s41598-019-49031-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Masters JR. Cell-line authentication: end the scandal of false cell lines. Nature. 2012;492:186. doi: 10.1038/492186a. [DOI] [PubMed] [Google Scholar]
- 32.Clevers H, et al. What is your conceptual definition of “cell type” in the context of a mature organism? Cell Syst. 2017;4:255–259. doi: 10.1016/j.cels.2017.03.006. [DOI] [PubMed] [Google Scholar]
- 33.Brennecke P, et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods. 2013;10:1093–1095. doi: 10.1038/nmeth.2645. [DOI] [PubMed] [Google Scholar]
- 34.Zappia L, Phipson B, Oshlack A. Exploring the single-cell RNA-seq analysis landscape with the scRNA-tools database. PLoS Comput. Biol. 2018;14:e1006245. doi: 10.1371/journal.pcbi.1006245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hon C-C, Shin JW, Carninci P, Stubbington MJT. The Human Cell Atlas: technical approaches and challenges. Brief. Funct. Genomics. 2018;17:283–294. doi: 10.1093/bfgp/elx029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Junqueira, L. C., Carneiro, J. & Kelly, R. O. Histologi Dasar (Basic Histology) (EGC Penebrit Buku Kedokteran, 1980).
- 37.Ramon y Cajal, S. Histologie du système nerveux de l’homme & des vertébrés (Maloine, 1909).
- 38.Molyneaux BJ, Arlotta P, Menezes JRL, Macklis JD. Neuronal subtype specification in the cerebral cortex. Nat. Rev. Neurosci. 2007;8:427–437. doi: 10.1038/nrn2151. [DOI] [PubMed] [Google Scholar]
- 39.Klausberger T, Somogyi P. Neuronal diversity and temporal dynamics: the unity of hippocampal circuit operations. Science. 2008;321:53–57. doi: 10.1126/science.1149381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.DeFelipe J, et al. New insights into the classification and nomenclature of cortical GABAergic interneurons. Nat. Rev. Neurosci. 2013;14:202–216. doi: 10.1038/nrn3444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sugino K, et al. Molecular taxonomy of major neuronal classes in the adult mouse forebrain. Nat. Neurosci. 2006;9:99–107. doi: 10.1038/nn1618. [DOI] [PubMed] [Google Scholar]
- 42.Yakimchuk K. Cell markers. Mater. Methods. 2013;3:183. [Google Scholar]
- 43.Stachelscheid H, et al. CellFinder: a cell data repository. Nucleic Acids Res. 2014;42:D950–D958. doi: 10.1093/nar/gkt1264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhang X, et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 2019;47:D721–D728. doi: 10.1093/nar/gky900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Franzén O, Gan L-M, Björkegren JLM. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database (Oxford) 2019;2019:baz046. doi: 10.1093/database/baz046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhou BO, Yue R, Murphy MM, Peyer JG, Morrison SJ. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell. 2014;15:154–168. doi: 10.1016/j.stem.2014.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Debnath S, et al. Discovery of a periosteal stem cell mediating intramembranous bone formation. Nature. 2018;562:133–139. doi: 10.1038/s41586-018-0554-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhang J, Link DC. Targeting of mesenchymal stromal cells by Cre-Recombinase transgenes commonly used to target osteoblast lineage cells. J. Bone Miner. Res. 2016;31:2001–2007. doi: 10.1002/jbmr.2877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gustafson MP, et al. A method for identification and analysis of non-overlapping myeloid immunophenotypes in humans. PLoS ONE. 2015;10:e0121546. doi: 10.1371/journal.pone.0121546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wagner C, et al. Expression patterns of the lipopolysaccharide receptor CD14, and the FCgamma receptors CD16 and CD64 on polymorphonuclear neutrophils: data from patients with severe bacterial infections and lipopolysaccharide-exposed cells. Shock. 2003;19:5–12. doi: 10.1097/00024382-200301000-00002. [DOI] [PubMed] [Google Scholar]
- 51.Miki K, et al. Efficient detection and purification of cell populations using synthetic microrna switches. Cell Stem Cell. 2015;16:699–711. doi: 10.1016/j.stem.2015.04.005. [DOI] [PubMed] [Google Scholar]
- 52.Markram H, et al. Interneurons of the neocortical inhibitory system. Nat. Rev. Neurosci. 2004;5:793–807. doi: 10.1038/nrn1519. [DOI] [PubMed] [Google Scholar]
- 53.Parra P, Gulyás AI, Miles R. How many subtypes of inhibitory cells in the hippocampus? Neuron. 1998;20:983–993. doi: 10.1016/s0896-6273(00)80479-1. [DOI] [PubMed] [Google Scholar]
- 54.Takahashi J. Stem cells and regenerative medicine for neural repair. Curr. Opin. Biotechnol. 2018;52:102–108. doi: 10.1016/j.copbio.2018.03.006. [DOI] [PubMed] [Google Scholar]
- 55.Greenblatt MB, Ono N, Ayturk UM, Debnath S, Lalani S. The unmixing problem: a guide to applying single-cell RNA sequencing to bone. J. Bone Miner. Res. 2019;34:1207–1219. doi: 10.1002/jbmr.3802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Arendt D, et al. The origin and evolution of cell types. Nat. Rev. Genet. 2016;17:744–757. doi: 10.1038/nrg.2016.127. [DOI] [PubMed] [Google Scholar]
- 57.Graf T, Enver T. Forcing cells to change lineages. Nature. 2009;462:587–594. doi: 10.1038/nature08533. [DOI] [PubMed] [Google Scholar]
- 58.Saint-André V, et al. Models of human core transcriptional regulatory circuitries. Genome Res. 2016;26:385–396. doi: 10.1101/gr.197590.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Mullen AC, et al. Master transcription factors determine cell-type-specific responses to TGF-β signaling. Cell. 2011;147:565–576. doi: 10.1016/j.cell.2011.08.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Davis RL, Weintraub H, Lassar AB. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell. 1987;51:987–1000. doi: 10.1016/0092-8674(87)90585-x. [DOI] [PubMed] [Google Scholar]
- 61.Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–676. doi: 10.1016/j.cell.2006.07.024. [DOI] [PubMed] [Google Scholar]
- 62.Nakagawa M, et al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat. Biotechnol. 2008;26:101–106. doi: 10.1038/nbt1374. [DOI] [PubMed] [Google Scholar]
- 63.Hobert O. Regulatory logic of neuronal diversity: terminal selector genes and selector motifs. Proc. Natl. Acad. Sci. USA. 2008;105:20067–20071. doi: 10.1073/pnas.0806070105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Sokolik C, et al. Transcription factor competition allows embryonic stem cells to distinguish authentic signals from noise. Cell Syst. 2015;1:117–129. doi: 10.1016/j.cels.2015.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.van der Meulen T, Huising MO. Role of transcription factors in the transdifferentiation of pancreatic islet cells. J. Mol. Endocrinol. 2015;54:R103–R117. doi: 10.1530/JME-14-0290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zeng H, Sanes JR. Neuronal cell-type classification: challenges, opportunities and the path forward. Nat. Rev. Neurosci. 2017;18:530–546. doi: 10.1038/nrn.2017.85. [DOI] [PubMed] [Google Scholar]
- 67.Sakurai K, Fujibuchi W. Close relationships between iPS cells and Periodic Table of chemical elements. Trans. Res. Inst. Oceanochem. 2016;29:17–23. [Google Scholar]
- 68.Xia B, Yanai I. A periodic table of cell types. Development. 2019;146:dev169854. doi: 10.1242/dev.169854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Waddington CH. Canalization of development and the inheritance of acquired characters. Nature. 1942;150:563–565. doi: 10.1038/1831654a0. [DOI] [PubMed] [Google Scholar]
- 70.Moris N, Pina C, Arias AM. Transition states and cell fate decisions in epigenetic landscapes. Nat. Rev. Genet. 2016;17:693–703. doi: 10.1038/nrg.2016.98. [DOI] [PubMed] [Google Scholar]
- 71.Moignard V, et al. Decoding the regulatory network of early blood development from single-cell gene expression measurements. Nat. Biotechnol. 2015;33:269–276. doi: 10.1038/nbt.3154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Trapnell C. Defining cell types and states with single-cell genomics. Genome Res. 2015;25:1491–1498. doi: 10.1101/gr.190595.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Weinreb C, Wolock S, Klein AM. SPRING: a kinetic interface for visualizing high dimensional single-cell expression data. Bioinformatics. 2018;34:1246–1248. doi: 10.1093/bioinformatics/btx792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kiselev VY, et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods. 2017;14:483–486. doi: 10.1038/nmeth.4236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Xu C, Su Z. Identification of cell types from single-cell transcriptomes using a novel clustering method. Bioinformatics. 2015;31:1974–1980. doi: 10.1093/bioinformatics/btv088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Zheng, X., Jin, S., Nie, Q. & Zou, X. scRCMF: identification of cell subpopulations and transition states from single cell transcriptomes. IEEE Trans. Biomed. Eng. 10.1109/TBME.2019.2937228 (2019). [DOI] [PMC free article] [PubMed]
- 77.Abdelaal T, et al. A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol. 2019;20:194. doi: 10.1186/s13059-019-1795-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Crook JM, Hei D, Stacey GN. The International Stem Cell Banking Initiative (ISCBI): raising standards to bank on. Vitr. Cell Dev. Biol. Anim. 2010;46:169–172. doi: 10.1007/s11626-010-9301-7. [DOI] [PubMed] [Google Scholar]
- 79.Seltmann S, et al. hPSCreg-the human pluripotent stem cell registry. Nucleic Acids Res. 2016;44:D757–D763. doi: 10.1093/nar/gkv963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Kurtz A, et al. A standard nomenclature for referencing and authentication of pluripotent stem cells. Stem Cell Rep. 2018;10:1–6. doi: 10.1016/j.stemcr.2017.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Hatano A, et al. CELLPEDIA: a repository for human cell information for cell studies and differentiation analyses. Database. 2011;2011:bar046. doi: 10.1093/database/bar046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Diehl AD, et al. The Cell Ontology 2016: enhanced content, modularization, and ontology interoperability. J. Biomed. Semant. 2016;7:44. doi: 10.1186/s13326-016-0088-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Jenkins MJ, Farid SS. Human pluripotent stem cell-derived products: advances towards robust, scalable and cost-effective manufacturing strategies. Biotechnol. J. 2015;10:83–95. doi: 10.1002/biot.201400348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.De Sousa PA, et al. Rapid establishment of the European Bank for induced Pluripotent Stem Cells (EBiSC)—the Hot Start experience. Stem Cell Res. 2017;20:105–114. doi: 10.1016/j.scr.2017.03.002. [DOI] [PubMed] [Google Scholar]
- 85.International Stem Cell Initiative. Assessment of established techniques to determine developmental and malignant potential of human pluripotent stem cells. Nat. Commun. 2018;9:1925. doi: 10.1038/s41467-018-04011-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sugimoto N, Eto K. Platelet production from induced pluripotent stem cells. J. Thromb. Haemost. 2017;15:1717–1727. doi: 10.1111/jth.13736. [DOI] [PubMed] [Google Scholar]
- 87.Ito Y, et al. Turbulence activates platelet biogenesis to enable clinical scale ex vivo production. Cell. 2018;174:636–648.e18. doi: 10.1016/j.cell.2018.06.011. [DOI] [PubMed] [Google Scholar]
- 88.Karagiannis, P., Sugimoto, N. & Eto, K. in Platelets, 4th edn (ed. Michelson, A. D.) 1173–1189 (Academic Press, 2019).
- 89.Gropp M, et al. Standardization of the teratoma assay for analysis of pluripotency of human ES cells and biosafety of their differentiated progeny. PLoS ONE. 2012;7:e45532. doi: 10.1371/journal.pone.0045532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.International Stem Cell Banking Initiative. Consensus guidance for banking and supply of human embryonic stem cell lines for research purposes. Stem Cell Rev. 2009;5:301–314. doi: 10.1007/s12015-009-9085-x. [DOI] [PubMed] [Google Scholar]
- 91.Buta C, et al. Reconsidering pluripotency tests: do we still need teratoma assays? Stem Cell Res. 2013;11:552–562. doi: 10.1016/j.scr.2013.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Ellis J, et al. Alternative induced pluripotent stem cell characterization criteria for in vitro applications. Cell Stem Cell. 2009;4:198–199. doi: 10.1016/j.stem.2009.02.010. [DOI] [PubMed] [Google Scholar]
- 93.Andrews PW, et al. Points to consider in the development of seed stocks of pluripotent stem cells for clinical applications: International Stem Cell Banking Initiative (ISCBI) Regen. Med. 2015;10:1–44. doi: 10.2217/rme.14.93. [DOI] [PubMed] [Google Scholar]
- 94.Kurosawa H. Methods for inducing embryoid body formation: in vitro differentiation system of embryonic stem cells. J. Biosci. Bioeng. 2007;103:389–398. doi: 10.1263/jbb.103.389. [DOI] [PubMed] [Google Scholar]
- 95.Sheridan SD, Surampudi V, Rao RR. Analysis of embryoid bodies derived from human induced pluripotent stem cells as a means to assess pluripotency. Stem Cells Int. 2012;2012:738910. doi: 10.1155/2012/738910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Ng ES, Davis RP, Azzola L, Stanley EG, Elefanty AG. Forced aggregation of defined numbers of human embryonic stem cells into embryoid bodies fosters robust, reproducible hematopoietic differentiation. Blood. 2005;106:1601–1603. doi: 10.1182/blood-2005-03-0987. [DOI] [PubMed] [Google Scholar]
- 97.Bock C, et al. Reference Maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell. 2011;144:439–452. doi: 10.1016/j.cell.2010.12.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Avior Y, Biancotti JC, Benvenisty N. TeratoScore: assessing the differentiation potential of human pluripotent stem cells by quantitative expression analysis of teratomas. Stem Cell Rep. 2015;4:967–974. doi: 10.1016/j.stemcr.2015.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.O’Connor MD, et al. Alkaline phosphatase-positive colony formation is a sensitive, specific, and quantitative indicator of undifferentiated human embryonic stem cells. Stem Cells. 2008;26:1109–1116. doi: 10.1634/stemcells.2007-0801. [DOI] [PubMed] [Google Scholar]
- 100.O’Connor MD, Kardel MD, Eaves CJ. Functional assays for human embryonic stem cell pluripotency. Methods Mol. Biol. 2011;690:67–80. doi: 10.1007/978-1-60761-962-8_4. [DOI] [PubMed] [Google Scholar]
- 101.Collier AJ, Rugg-Gunn PJ. Identifying human naïve pluripotent stem cells—evaluating state-specific reporter lines and cell-surface markers. Bioessays. 2018;40:e1700239. doi: 10.1002/bies.201700239. [DOI] [PubMed] [Google Scholar]
- 102.Messmer T, et al. Transcriptional heterogeneity in naive and primed human pluripotent stem cells at single-cell resolution. Cell Rep. 2019;26:815–824.e4. doi: 10.1016/j.celrep.2018.12.099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ghimire S, et al. Comparative analysis of naive, primed and ground state pluripotency in mouse embryonic stem cells originating from the same genetic background. Sci. Rep. 2018;8:5884. doi: 10.1038/s41598-018-24051-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Lipsitz YY, Woodford C, Yin T, Hanna JH, Zandstra PW. Modulating cell state to enhance suspension expansion of human pluripotent stem cells. Proc. Natl. Acad. Sci. USA. 2018;115:6369–6374. doi: 10.1073/pnas.1714099115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Ungrin, M., O’Connor, M., Eaves, C. & Zandstra, P. W. Phenotypic analysis of human embryonic stem cells. Curr. Protoc. Stem Cell Biol. Chapter 1, Unit 1B.3 (2007). [DOI] [PubMed]
- 106.Müller F-J, et al. A bioinformatic assay for pluripotency in human cells. Nat. Methods. 2011;8:315–317. doi: 10.1038/nmeth.1580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Guo M, Bao EL, Wagner M, Whitsett JA, Xu Y. SLICE: determining cell differentiation and lineage based on single cell entropy. Nucleic Acids Res. 2017;45:e54. doi: 10.1093/nar/gkw1278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Teschendorff AE, Enver T. Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome. Nat. Commun. 2017;8:15599. doi: 10.1038/ncomms15599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Lenz M, et al. Epigenetic biomarker to support classification into pluripotent and non-pluripotent cells. Sci. Rep. 2015;5:8973. doi: 10.1038/srep08973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Sakurai K, Kurtz A, Stacey GN, Sheldon M, Fujibuchi W. First proposal of minimum information about a cellular assay for regenerative medicine. Stem Cells Transl. Med. 2016;5:1345–1361. doi: 10.5966/sctm.2015-0393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Wiemann, S., Mehrle, A. & Hahne, F. MIACA—minimum information about a cellular assay, and the cellular assay object model http://miaca.sourceforge.net/ (2015)