Translation initiation regions in mRNAs that include the ribosome-binding site (RBS) and the start codon are often sequestered within a secondary structure. Therefore, to initiate protein synthesis, the mRNA secondary structure must be unfolded to allow the RBS to be accessible to the ribosome.
KEYWORDS: uORF, translation-inhibitory stem-loop structure, intergenic region, ribosome destabilization
ABSTRACT
Bacterial mRNAs often harbor upstream open reading frames (uORFs) in the 5′ untranslated regions (UTRs). Translation of the uORF usually affects downstream gene expression at the levels of transcription and/or translation initiation. Unlike other uORFs mostly located in the 5′ UTR, we discovered an 8-amino-acid ORF, designated mgtQ, in the intergenic region between the mgtC virulence gene and the mgtB Mg2+ transporter gene in the Salmonella mgtCBRU operon. Translation of mgtQ promotes downstream mgtB Mg2+ transporter expression at the level of translation by releasing the ribosome-binding sequence of the mgtB gene that is sequestered in a translation-inhibitory stem-loop structure. Interestingly, mgtQ Asp2 and Glu5 codons that induce ribosome destabilization are required for mgtQ-mediated mgtB translation. Moreover, the mgtQ Asp and Glu codons-mediated mgtB translation is counteracted by the ribosomal subunit L31 that stabilizes ribosome. Substitution of the Asp2 and Glu5 codons in mgtQ decreases MgtB Mg2+ transporter production and thus attenuates Salmonella virulence in mice, likely by limiting Mg2+ acquisition during infection.
INTRODUCTION
In bacteria, genes within an operon are transcribed as a single polycistronic mRNA and translated individually by allowing the ribosome access to the ribosome-binding sequence of each gene. However, translation of each gene within the operon is not fully independent, depending on the length of the intergenic region between the neighboring genes or the presence of specific regulatory mechanisms. Translation of the neighboring genes is usually tightly coupled when two genes are close to each other or overlapping because ribosome would simply reinitiate translation at the start codon of the following gene (1). However, given that many genes are separated by intergenic sequences ranging from a few to several hundred nucleotides (2, 3), additional regulatory mechanisms are often required to coordinate synthesis of proteins within the operon, which include interactions with proteins or small RNAs, endonucleolytic cleavage within mRNA, and translation of short open reading frames (ORFs) (4–7). Here, we identify a short ORF that is located in the intergenic region, translation of which promotes translation of the distal gene, thereby coupling protein synthesis between neighboring genes within the operon.
In the intracellular pathogen Salmonella enterica serovar Typhimurium, the mgtB gene encoding a Mg2+ transporting P-type ATPase lies between the virulence gene mgtC and the proteolysis-regulatory gene mgtR in the mgtCBRU operon. The mgtCBRU operon is transcribed from a single promoter located upstream of the first mgtC gene, and the MgtB Mg2+ transporter is produced from a part of >4-kb polycistronic mRNA (8). (Although there appeared to be a read-through transcript into the fifth gene cigR, we only focused on four genes here because the cigR gene is also transcribed from a constitutive promoter [9]). Because the MgtB protein imports Mg2+ ions at the expense of ATP (10), it is reasonable that the mgtCBRU operon is highly transcribed in low Mg2+ by the PhoP/PhoQ two-component system (11) to support bacterium’s growth in a Mg2+-limiting condition. However, although the mgtCBRU operon is transcribed as a single polycistronic message, the production of the MgtB Mg2+ transporter appears to be differentially regulated based on the following reasons. First, the first gene product, MgtC, is a virulence protein that inhibits FoF1 ATP synthase, thus decreasing ATP production (12). Given that one of the major biological functions of Mg2+ ions is neutralizing free ATP as well as other nucleotides (13, 14), the requirement of the MgtB protein importing Mg2+ ions may vary depending on the amount of the active MgtC protein in a given condition. Second, the intergenic region between the first mgtC and the second mgtB genes is 219 nucleotides (nt) long, which is relatively long considering the average length of the intergenic regions in bacteria (15, 16). Lastly, Mfold analysis predicted that the ribosome-binding site (RBS) of the mgtB gene is occluded by a potential base pairing with upstream sequences (Fig. 1) (17), suggesting that an additional regulatory element is required for mgtB translation.
FIG 1.
Regulation of the Salmonella mgtCBRU virulence operon by the mgtQ short ORF. (A) The phosphorylated PhoP response regulator binds to the mgtCBRU promoter and initiates transcription. Two short ORFs in the 5′ leader region, mgtM and mgtP, control transcription elongation in response to intracellular ATP and charged tRNAPro levels via transcription attenuation-like mechanisms. The 8-aa mgtQ ORF is located in the 219-nt intergenic region between mgtC and mgtB genes and harbors Asp and Glu codons at positions 2 and 5. The sequence overlapping the mgtQ ORF has the potential to adopt alternative RNA secondary structures (stem-loops 1:2 and 3:4 versus 2:3) and the mgtB RBS is occluded within the stem-loop 3:4 structure. The Asp and Glu codons at mgtQ appear to induce ribosome destabilization and promote the formation of stem-loop 2:3, thus releasing the RBS of the mgtB gene and enhancing mgtB translation. Positions of nucleotide substitutions used in the experiments presented in Fig. 4 are indicated. (B) Alignment of the deduced amino acid sequences of the mgtQ orthologs in the mgtC-mgtB intergenic regions from Salmonella enterica, Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans. Sequences in red correspond to Pro codons, and sequences in green correspond to Asp and Glu codons. Asterisks correspond to positions conserved in all listed species.
In this study, we discovered a short ORF, designated mgtQ, in the intergenic region between the mgtC and mgtB genes. Translation of mgtQ is required for downstream mgtB translation. Salmonella’s intrinsic ability to translate mgtQ is dependent on acidic residues in mgtQ, which are known to induce ribosomal destabilization and also require a specific ribosomal subunit encoded by the rpmE1 gene to counteract the acidic residue-mediated translation abortion. Therefore, translation efficiency of mgtQ is involved in controlling production of the MgtB Mg2+ transporter, thus contributing to the ability to import Mg2+ and Salmonella pathogenesis.
RESULTS
The intergenic region between the mgtC and mgtB genes includes a short ORF.
The mgtCBRU operon is transcribed as a polycistronic mRNA from the single promoter located upstream of the mgtC gene (8). Interestingly, the mgtC and mgtB genes are separated by the 219-nt intergenic region, which is unusually long for the operon (15, 16). The presence of the long intergenic region led us to examine whether it contains an additional regulatory element. We found a potential short ORF located 51 nt downstream of the mgtC stop codon, designated mgtQ. mgtQ is an 8-amino acid ORF preceded by a strong Shine-Dalgarno RBS (AGGAGG) (Fig. 1; see also Fig. S1 in the supplemental material). Although it varies in length, the presence of mgtQ is conserved in other bacteria, including Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans (Fig. 1; see also Fig. S1 and S2). The conservation of the mgtQ ORF in other bacteria raises the possibility that the mgtQ ORF is translated. To test this, we constructed a strain harboring a promoterless gfp plasmid translationally fused to the DNA fragment corresponding to nt 982 to 1054 (from the intergenic region to the last sense codon of the mgtQ ORF). As a control, we used an isogenic derivative in which the gfp gene was fused after the mgtQ stop codon (mgtQSTOP′-gfp) (Fig. 2A). The strain harboring mgtQ′-gfp produced high levels of fluorescence compared to control strains with the empty vector or mgtQSTOP′-gfp (Fig. 2B). This result indicates that mgtQ is translated in vivo.
FIG 2.
The mgtQ ORF is translated in vivo. (A) Schematic representation of mgtQ′-gfp constructs. (B) Fluorescence produced by wild-type Salmonella (14028s) harboring the plasmid vector (ptGFP), or derivatives with a gfp translational fusion to the last mgtQ codon (pmgtQ′-gfp), or following the stop mgtQ codon (pmgtQ STOP′-gfp). Bacteria were grown for 4 h in N-minimal medium containing 10 mM Mg2+ as described in Materials and Methods. The means and standard deviations (SD) from three independent measurements are shown.
The intergenic regions between the mgtC and mgtB genes of other enteric bacteria harbor mgtQ orthologs. Alignment of the nucleotide sequences corresponding to part of the intergenic regions between the mgtC and mgtB genes from Salmonella enterica, Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans. Sequences in boldface correspond to mgtQ. Sequences in green, purple, blue, and red represent regions involved in stem-loop structures (SR1, SR2, SR3, and SR4, respectively) shown in Fig. 1. The predicted RBS for each ORF is underlined. Asterisks correspond to nucleotides conserved in all listed species. Download FIG S1, TIF file, 0.2 MB (187.4KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
The presence of mgtQ is conserved in enteric bacteria where the mgtC and mgtB genes are in the same operon. In Salmonella enterica, mgtQ is located in the mgtC-mgtB intergenic region. Similarly, the mgtC-mgtB intergenic regions from Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans include mgtQ-like ORFs. In Brucella melitensis, the mgtC and mgtB genes appear to be part of separate transcription units. An mgtQ-like sequence is found upstream of the mgtB gene. Mycobacterium tuberculosis harbors the mgtC gene but not mgtB or mgtQ. Download FIG S2, TIF file, 0.2 MB (170.9KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
mgtQ affects mgtB expression at the level of translation.
Because mgtQ is located upstream of the mgtB gene (Fig. 1), we hypothesized that mgtQ impacts downstream mgtB expression. To test mgtQ’s effect on mgtB expression, we constructed a plasmid with the gfp gene fused transcriptionally or translationally to the first 27 codons of the mgtB gene (Fig. 3A; see also Fig. S3). We also constructed an isogenic plasmid harboring the mgtB-fused gfp gene with the upstream mgtQ start codon substituted by the TAG stop codon (mgtQATG→TAG; Fig. 3A). Because both plasmids harbor a 1.2-kb DNA fragment containing the PhoP-dependent promoter (pmgtC), as well as the mgtC and mgtQ genes upstream of the mgtB-fused gfp gene, we grew Salmonella strains harboring the mgtB-fused gfp plasmids in PhoP-inducing (0.01 mM Mg2+) or PhoP-repressing (10 mM Mg2+) conditions. A Salmonella strain harboring the plasmid with the wild-type mgtQ produced high levels of mgtB-gfp fluorescence in 0.01 mM Mg2+, but low levels of fluorescence in 10 mM Mg2+ for both transcriptional and translational fusions (Fig. 3B and C). However, the introduction of the start to stop codon at mgtQ decreased mgtB-gfp fluorescence when fused translationally but not when fused transcriptionally (Fig. 3B and C). This indicates that mgtQ affects mgtB expression at translational level. As controls, strains harboring the empty vector or strains grown in the noninducing media exhibited low levels of fluorescence in all tested conditions (Fig. 3B and C).
FIG 3.
mgtQ translation affects mgtB expression at a translational level but not transcriptional level. (A) Schematic representation of mgtB-gfp constructs used in this experiment. (B and C) Fluorescence produced by wild-type Salmonella (14028s) harboring a plasmid with either a gfp gene translationally (B) or transcriptionally (C) fused to the mgtB gene with the wild-type mgtQ (mgtQWT) or an mgtQ mutant with the start codon substituted by the TAG stop codon (mgtQATG→TAG). Bacteria were grown for 5 h in N-minimal medium containing 0.01 mM (inducing) or 10 mM (noninducing) Mg2+ as described in Materials and Methods. The means and SD from three independent measurements are shown.
Schematic representation of reporter systems used in this study. (A) A partial sequence of pfpv25 plasmid, a plasmid for transcriptional fusion to a promoterless gfp gene. (B) A partial sequence of ptGFP, an engineered plasmid for translational fusion to a promoterless gfp gene. In ptGFP plasmid, a portion of sequence in pfpv25 (indicated in blue) was deleted to remove the ribosome binding site and the start codon of the gfp gene. Download FIG S3, TIF file, 0.09 MB (89.7KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
We also created a chromosomal mutant with the same start to stop codon substitution in mgtQ (see Fig. S4). Similarly to what we observed in gfp plasmids, the introduction of the TAG stop codon at mgtQ severely decreased MgtB protein levels in the PhoP-inducing condition, while the same substitution did not affect mRNA levels of the mgtC and mgtB genes (see Fig. S4). These results further demonstrate that mgtQ indeed affects mgtB expression at the translational level.
mgtQ translation increases MgtB protein levels but not mRNA levels. (A) Schematic representation of an mgtQ chromosomal mutant with the start codon substituted by the stop codon and the DNA fragments amplified by QRT-PCR in panels D and E. (B and C) Western blot analysis of crude extracts prepared from Salmonella strains with the wild-type mgtQ (mgtQWT) or the mgtQ derivative with the start to stop codon substitution (mgtQATG→TAG) grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+. The amounts of MgtB and Fur proteins were determined by anti-MgtB (B) and anti-Fur (C) antibodies. (D and E) Relative mRNA levels of the mgtC (D) and mgtB (E) coding regions produced by wild-type (mgtQWT) Salmonella and the mgtQ mutant with the start codon replaced by the stop codon (mgtQATG→TAG) grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+. The means and SD from three independent measurements are shown. Download FIG S4, TIF file, 0.1 MB (144.3KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Formation of alternative stem-loop structures overlapping the mgtQ ORF is involved in controlling mgtB translation.
We then wondered how mgtQ affects downstream mgtB translation. Sequence analyses of the mgtC-mgtB intergenic region revealed that the sequence overlapping with the mgtQ ORF has the potential to adopt two alternative stem-loop structures: stem-loops 1:2 and 3:4 or stem-loop 2:3 (Fig. 1 and Fig. 4A). The presence of alternative stem-loop structures neighboring the mgtQ ORF suggests that mgtQ translation might affect the formation of one of two alternative stem-loop structures, which eventually leads to an increase or decrease in downstream mgtB expression. Interestingly, among those stem-loop structures, stem-loop 3:4 appears to be a translation-inhibitory stem-loop structure because the formation of stem-loop 3:4 occludes the RBS of the mgtB gene (Fig. 4A). Given that the formations of stem-loop 2:3 and stem-loops 1:2/3:4 are mutually exclusive, the formation of stem-loop 2:3 is expected to increase mgtB translation by preventing the formation of stem-loop 3:4 and liberating the RBS of the mgtB gene (Fig. 4B). To explore this, we introduced substitutions in stem regions 1, 2, 3, or 4 of the mgtB-fused gfp plasmids to determine the effect of stem-loop formation on downstream mgtB expression. A nucleotide substitution in stem region 3 that is expected to hinder the formation of both stem-loop 3:4 and stem-loop 2:3 increased translationally fused mgtB-gfp expression (mgtB′-gfp), supporting the idea that stem-loop 3:4 is a translation-inhibitory stem-loop structure (Fig. 4B).
FIG 4.
Formation of alternative stem-loop structures that include mgtQ affects downstream mgtB translation. (A) Schematic representation of a model showing predicted alternative stem-loop structures (1:2 and 3:4 versus 2:3) that are associated with the mgtQ ORF. (B) Fluorescence produced by wild-type Salmonella (14028s) harboring the plasmid vector (vector), or derivatives with a gfp translational fusion to the mgtB gene that includes the upstream intergenic region (wild-type), substitution mutations in the stem regions 1, 2, 3, or 4 that hinder stem-loop formation (1:2, 2:3, and/or 3:4), or compensatory mutations that recover the formation of stem-loops 1:2, 2:3, or 3:4. A strain harboring a plasmid with a gfp translational fusion derived from the plac1-6 promoter was used as a positive control. Bacteria were grown for 4 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. The means and SD from three independent measurements are shown. (C) Fluorescence produced by wild-type Salmonella (14028s) harboring the plasmid vector (vector), or derivatives with a gfp transcriptional fusion to the mgtB gene that includes the upstream intergenic region (wild-type), substitution mutations described in panel B. A strain harboring a plasmid with a gfp transcriptional fusion derived from the plac1-6 promoter was used as a positive control. Bacteria were grown for 4 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. The means and SD from three independent measurements are shown.
In contrast, the substitution of stem region 4 decreased mgtB′-gfp expression due to loss of the RBS of the mgtB gene (Fig. 4B). Substitution in stem region 2 that appears to disrupt the formation of stem-loop 2:3 and thus favor the formation of stem-loop 3:4, resulted in a decrease in mgtB′-gfp levels (Fig. 4B). Similarly, substitution in stem region 1 within mgtQ increased mgtB′-gfp levels because the substitution disrupts the formation of stem-loop 1:2 and favors the formation of stem-loop 2:3, hence releasing the RBS of the mgtB gene (Fig. 4B). The introduction of a compensatory mutation in either stem-loop 1:2 or 3:4 is expected to force the formation of stem-loop 3:4 and thus decreased mgtB′-gfp expression (Fig. 4B). In contrast, the introduction of a compensatory mutation in stem-loop 2:3 that possibly fails to occlude the RBS of the mgtB gene increased mgtB′-gfp expression. All substitutions in the stem-loop regions described above affect mgtB′-gfp expression only at translational level because isogenic substitutions did not have an effect on mgtB-gfp expression when fused transcriptionally (Fig. 4C). These data demonstrate that two alternative stem-loop structures could be formed and control mgtB translation by either masking or revealing the RBS of the downstream mgtB gene.
Stem-loop 1:2 formation but not mgtQ Pro codon is required for mgtB translation.
As described above, inhibition of mgtQ translation by introducing the mgtQ start codon mutation decreased mgtB translation, likely because stem-loops 1:2 and 3:4 are formed when mgtQ is not translated (Fig. 4A). Moreover, the substitution in stem region 1 within mgtQ increased mgtB translation by releasing the RBS from the inhibitory stem-loop structure 3:4. Because stem-loop 1:2 is also disrupted while mgtQ is translated, any condition that slows down mgtQ translation is expected to promote mgtB translation in cis by favoring the formation of stem-loop 2:3 (Fig. 4A).
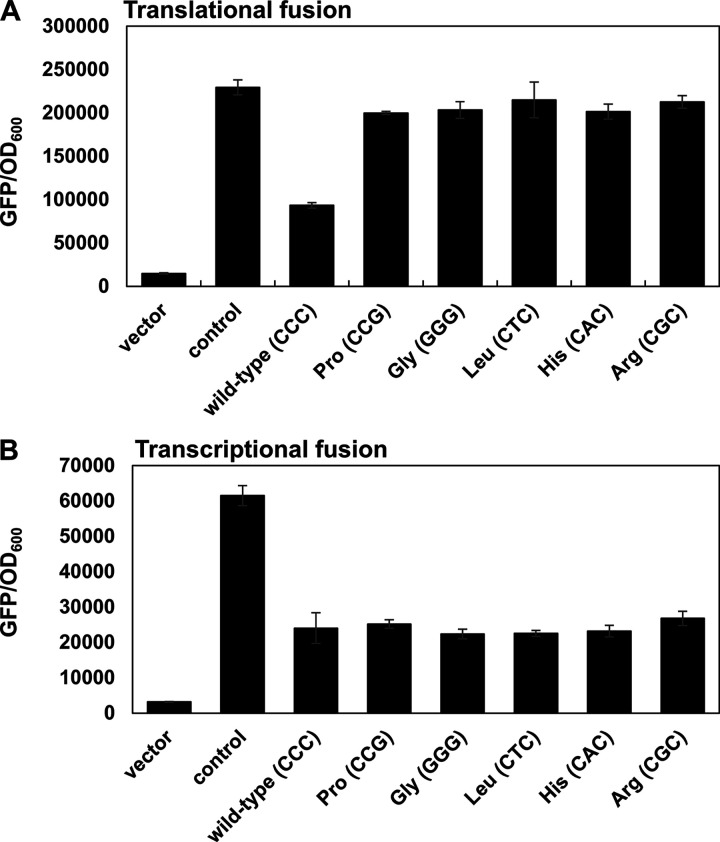
Given that the mgtQ ORF has a single conserved proline codon at the fourth position (Fig. 1B) and that the nucleotide sequence of the proline codon (CCC) is a part of the stem region 1 (Fig. 1), we wondered whether the conserved proline codon has a role to affect downstream mgtB translation. To test this, we substituted the Pro codon in mgtQ (CCC) with Gly (GGG), Leu (CTC), His (CAC), or Arg (CGC) codons and measured the downstream mgtB′-gfp levels. All of these mgtQ Pro substitutions to Gly, Leu, His, or Arg increased mgtB′-gfp levels compared to levels in the isogenic control with the wild-type mgtQ (Fig. 5A), suggesting that removal of the mgtQ Pro codon might have an effect on downstream mgtB expression. However, we found that the silent substitution of the Pro codon CCC to CCG still increased downstream mgtB′-gfp levels similar to those detected in other missense mutations (Fig. 5A). Thus, disruption of stem-loop 1:2 rather than removal of the Pro codon determines mgtB′-gfp expression by promoting the release of the mgtB RBS. It is interesting to note that, although the substitution of the Pro codon (CCC) with Leu codon (CTC) was expected to form stem-loop 1:2 by non-Watson Crick base-pairing (U:G), it seemed to fail to form stem-loop 1:2, because the substitution still increased mgtB′-gfp levels (Fig. 5A). As a control experiment, isogenic substitutions in transcriptionally fused mgtB-gfp plasmids did not have any effect on mgtB expression (Fig. 5B).
FIG 5.
Stem-loop formation but not mgtQ Pro codon is required for mgtQ-mediated mgtB translation. (A and B) Fluorescence produced by wild-type Salmonella (14028s) harboring the plasmid vector (vector), or derivatives with a gfp translational (A) or transcriptional (B) fusion to the mgtB gene that includes the wild-type mgtQ, nucleotide substitution mutations of the conserved proline codon replaced by a synonymous mutation (CCG) or nonsynonymous mutations, including Gly (GGG), Leu (CTC), His (CAC), and Arg (CGC). The strains harboring a plasmid with a gfp translational fusion derived from the plac1-6 promoter were used as positive controls. Bacteria were grown for 5 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. The means and SD from three independent measurements are shown.
Ribosome-destabilizing acidic residues in mgtQ are required for mgtB translation.
During the multiple rounds of translation, if the mgtQ-translating ribosome continues to occupy stem region 1, the transcribed mRNAs thermodynamically favor the formation of stem-loop 2:3 over the formation of stem-loop 3:4 (the former free energy is −16.4 kcal/mol and the latter is −12.26 kcal/mol), thereby increasing mgtB translation. The fact that the mgtQ start codon mutation decreased mgtB translation (Fig. 3; see also Fig. S4) suggests that the mgtQ-translating ribosome might be a determinant to disrupt stem-loop 1:2 formation, allowing the formation of stem-loop 2:3 and increasing mgtB expression. In addition, the mgtQ substitution that does not affect codons but disrupts stem-loop 1:2 formation exhibited higher levels of mgtB′-gfp than those with the wild-type mgtQ sequence (Fig. 5), suggesting that completion of mgtQ translation appears to provide a chance to reassociate stem-loop 1:2 and thus decreases mgtB translation unless stem-loop 1:2 formation is disrupted by the nucleotide substitutions.
Although the mgtQ Pro4 codon affected mgtB translation by participating in base pairing of stem-loop 1:2 (Fig. 5), Pro4 might not affect mgtQ translation because single proline codon is not known to affect ribosome stalling. We then wondered what other factors could affect mgtQ translation. In Escherichia coli, acidic residues such as Asp and Glu in the nascent peptide chain interact within the ribosome exit tunnel and destabilize the translating ribosome during elongation (18). Because the mgtQ sequence also contains Asp and Glu codons at positions 2 and 5, respectively, we investigated whether these residues affect mgtQ translation by introducing Ala codon substitutions (Fig. 6A). A strain harboring the mgtQ′-gfp plasmid with Asp2 and Glu5 codons substituted by Ala codons exhibited a 2.1-fold increase in fluorescence compared to those produced from mgtQWT′-gfp (Fig. 6C), indicating that Asp and Glu codons indeed decrease mgtQ translation. To detect mgtQ translation from its chromosomal location, we created chromosomal mutants with a C-terminal 8×myc tag fused to the mgtQWT or mgtQ2,5Ala genes (see Fig. S5). MgtQWT-8×myc fusion proteins were marginally detectable in the PhoP-inducing condition (see Fig. S5). However, when we substituted the Asp2 and Glu5 to Ala codons, MgtQ2,5Ala-8×myc fusion protein production was strongly enhanced in the same condition (see Fig. S5), indicating that the presence of the mgtQ Asp2 and Glu5 residues impedes mgtQ translation. This is also in agreement with the previous finding that acidic residues destabilize the translating ribosome (18), although it has not been demonstrated here.
FIG 6.
Asp2 and Glu5 codons in mgtQ are required for mgtB translation. (A) Schematic representation of mgtQ′-gfp constructs used in this experiment. (B) Schematic representation of mgtB′-gfp constructs used in this experiment. (C) Fluorescence produced by wild-type Salmonella (14028s) by rpmE1 (EN1119), rpmE2 (EN1120), or rpmE1 rpmE2 (EN1368) Salmonella isolates harboring the promoterless plasmid vector (ptGFP), or by derivatives with gfp translational fusions to the wild-type mgtQ (pmgtQ′-gfp) or Asp2, Glu5 to Ala-substituted mgtQ (pmgtQ2,5Ala′-gfp). (D) Fluorescence produced by wild-type Salmonella (14028s), by rpmE1 (EN1119), rpmE2 (EN1120), or rpmE1 rpmE2 double mutant (EN1368) Salmonella harboring the plasmid vector (vector), or by Salmonella derivatives with a gfp translational fusion to the mgtB gene that includes the wild-type mgtQ or nucleotide substitution mutation of the Asp2 and Glu5 codons replaced with Ala. Bacteria were grown for 4 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. The means and SD from three independent measurements are shown.
The Asp2 and Glu5 to Ala substitutions enhance detection of the C-terminally 8×myc-tagged MgtQ peptides. (A) Schematic representation of Salmonella strains used in this study. (B-C) Western blot analysis of crude extracts prepared from Salmonella strains with the C-terminally 8×myc-tag fused to the wild-type mgtQ (EN1469) or mgtQ2,5Ala (EN1471) genes probed with anti-Myc (B) or anti-Fur (C) antibodies to detect MgtQ or Fur proteins, respectively. MgtQ-8×myc showed a higher SDS-PAGE-displayed molecular weight (∼20 kDa) than its predicted size (10.4 kDa) due to high content of acidic amino acids (47.7% in MgtQWT-8×myc and 45.5% in MgtQ2,5Ala-8×myc) (Y. Guan, Q. Zhu, D. Huang, S. Zhao, L. Jan Lo, J. Peng, Sci Rep 5:13370, 2015, doi:10.1038/srep13370). Bacteria were grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+ as described in Materials and Methods. Download FIG S5, TIF file, 0.1 MB (119.8KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
We then tested the effect of the mgtQ Asp2 and Glu5 to Ala substitutions on mgtB translation by introducing the same mgtQ substitution to a plasmid harboring mgtB′-gfp expressed from the native pmgtC promoter (Fig. 6B). The mgtQ2,5Ala substitution decreased mgtB′-gfp levels compared to those with mgtQWT (Fig. 6D), while the same substitution increased mgtQ′-gfp levels (Fig. 6C). This demonstrates that mgtQ translation efficiency inversely correlates with that of the mgtB gene. Given that Asp2 and Glu5 codons decrease mgtQ expression, mgtQ Asp2 and Glu5 codons appear to induce ribosome destabilization at mgtQ, favoring the formation of stem-loop 2:3 and releasing the RBS of the mgtB gene for promoting translation.
One can argue that the mgtQ Asp2 and Glu5 to Ala (GCT) substitutions might affect mRNA secondary structure formation and thus mgtB expression. To exclude this possibility, we created a similar derivative of mgtQ′-gfp with Asp2 and Glu5 substituted by Gly2 (GGT) and Ala5 (GCT) codons that potentially maintain base-pairing between stem regions 1 and 2 (see Fig. S6). The Asp2-to-Gly (GGT) and Glu5-to-Ala (GCT) substitutions increased the mgtQ′-gfp levels, similar to those detected in the mgtQ Asp2 and Glu5 to Ala (GCT) substitutions. Moreover, when we introduced the same mgtQ substitution into the mgtB′-gfp plasmid, the Asp2-to-Gly (GGT) and Glu5-to-Ala (GCT) substitutions inversely decreased mgtB′-gfp levels (see Fig. S6), further supporting that acidic residues in mgtQ reduce mgtQ translation, thus increasing downstream mgtB expression at translational level.
Asp2 and Glu5 codons in mgtQ are required for mgtB translation independently of the ability to form stem-loop 1:2. (A) A possible secondary structure of mgtQ-harboring mRNA sequences. Substituted nucleotide sequences of Asp2 and Glu5 codons in mgtQ are indicated at the side of Asp2 and Glu5 sequences. (B-C) Schematic representation of mgtQ′-gfp (B) and mgtB′-gfp constructs (C) used in this experiment. (D) Fluorescence produced by wild-type (14028s) Salmonella harboring the promoterless plasmid vector (ptGFP), or derivatives with a gfp translational fusion to the wild-type mgtQ (pmgtQ′-gfp) or Asp2 (GAT) Glu5 (GAG) to Gly2 (GGT) Ala5 (GCT)-substituted mgtQ (pmgtQ 2Gly, 5Ala′-gfp). (E) Fluorescence produced by wild-type (14028s) Salmonella harboring the plasmid vector (vector), or derivatives with a gfp translational fusion to the mgtB gene that includes the wild-type mgtQ or the mgtQ mutant with the Asp2 and Glu5 substitutions as described previously (D). Bacteria were grown for 4 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. Means and SD from three independent measurements are shown. Download FIG S6, TIF file, 0.1 MB (150KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
L31 ribosomal protein encoded by the rpmE1 gene controls mgtQ translation via acidic residues.
Chadani et al. (18) previously reported that E. coli L31 intersubunit bridge counteracts ribosome destabilization mediated by acidic residues in the nascent peptide. Because Salmonella enterica contains two types of L31 proteins encoded by rpmE1 and rpmE2 genes, respectively, we constructed chromosomal mutant strains deleted the rpmE1 and/or rpmE2 genes. The rpmE1 mutant exhibited decreased mgtQ′-gfp levels, and thus further increased mgtB′-gfp levels (Fig. 6C and D). Alterations of mgtQ′-gfp and mgtB′-gfp levels in the rpmE1 mutant were Asp2 and Glu5 dependent because the rpmE1 mutation had no further effect on Asp2- and Glu5-substituted mgtQ′-gfp levels or mgtQ2,5Ala-containing mgtB′-gfp levels (Fig. 6C and D). These data suggest that the L31 ribosomal protein encoded by the rpmE1 gene appears to be involved in resolving the acidic residue-mediated ribosomal destabilization at mgtQ, thereby decreasing mgtB translation. Interestingly, the rpmE2 mutation further increased mgtQ′-gfp levels, implicating that the L31 ribosomal protein encoded by the rpmE2 gene might have an opposing role in ribosomal destabilization, that is, to induce a stronger translation abortion at mgtQ than that observed in wild type. The proposed function of the rpmE2 gene seemed not to be exerted via Asp2 and Glu5 residues in mgtQ because the rpmE2 mutation further increased the Asp2- and Glu5-substituted mgtQ′-gfp compared to the wild-type mgtQ′-gfp (Fig. 6C). The facilitated mgtQ translation of both the wild-type and 2,5Ala-substituted mgtQ in the rpmE2 mutant further decreased mgtB′-gfp levels, supporting the finding that mgtQ and mgtB translation are inversely correlated. Finally, the effect of the rpmE1 mutation on determining mgtQ translation efficiency was dominant over that of the rpmE2 mutation, because the rpmE1 rpmE2 double mutant behaved just like the rpmE1 mutant (Fig. 6C).
The data described above suggest that availability of the L31 ribosomal protein could control MgtB Mg2+ transporter production via mgtQ ORF translation. To further confirm whether we could tune MgtB production by controlling the L31 ribosomal protein levels, we used the chromosomal rpmE1 deletion mutant and provided a plasmid with the rpmE1 gene expressed from an arabinose-inducible promoter (see Fig. S7). Similar to those detected in mgtB′-gfp levels, the MgtB protein levels were lower in the rpmE1 deletion mutant than those in the wild type (see Fig. S7). However, as we increased the L31 protein levels by adding arabinose, MgtB protein levels started to be restored to those of wild type at 1 mM arabinose and increased further at 10 mM arabinose (see Fig. S7), indicating that MgtB Mg2+ transporter production is indeed controlled by the cellular availability of the L31 ribosomal proteins.
Tuning L31 protein production restores MgtB production. (A and B) Western blot analysis of crude extracts prepared from wild-type or the rpmE1 mutant (EN1119) Salmonella harboring the rpmE1 gene from an arabinose inducible promoter (pBAD-rpmE1) probed with anti-MgtB (A) or anti-Fur (B) antibodies to detect MgtB or Fur proteins, respectively. Bacteria were grown for 5 h in N-minimal media containing 0.01 mM Mg2+ in the presence of the indicated concentration of arabinose. Download FIG S7, TIF file, 0.09 MB (97.3KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Acidic residues in mgtQ are required for promoting MgtB Mg2+ transporter production in low Mg2+.
AS described above, in the wild type, mgtQ-translating ribosomes are likely to be destabilized at Asp and Glu codons in mgtQ and allow the formation of stem-loop 2:3, thereby releasing the RBS of the mgtB gene to be translated. The degree of ribosome destabilization could be greater in ribosomes lacking the rpmE1-encoded L31 subunit, thus further promoting mgtB translation.
To understand the physiological role of the translation-aborting mgtQ ORF, we created a chromosomal mgtQ mutant with Asp2 and Glu5 substituted to Ala in the wild-type, rpmE1, rpmE2, or rpmE1 rpmE2 deletion backgrounds. In wild-type Salmonella, MgtB proteins were detected when grown in low Mg2+ to activate transcription from the PhoP-dependent promoter (19) (Fig. 7A). However, the Asp2-to-Ala and Glu5-to-Ala substitutions in mgtQ severely decreased MgtB production (Fig. 7A). This could be ascribed to the efficient mgtQ translation mediated by the Asp2-to-Ala and Glu5-to-Ala substitutions (Fig. 6C; see also Fig. S5) because efficient mgtQ translation likely favors the formation of stem-loops 1:2 and 3:4 instead of stem-loop 2:3. rpmE1 deletion seems to strengthen the ribosomal destabilization at mgtQ because it further increased MgtB protein levels (Fig. 7A). In contrast, rpmE2 deletion completely abolished MgtB production (Fig. 7A) by promoting mgtQ translation (Fig. 6C). Combining rpmE1 and rpmE2 mutations restored MgtB protein levels similar to those in the wild type, suggesting that two types of L31 ribosomal proteins have opposing effects on ribosomal destabilization at mgtQ (Fig. 7A). The Asp2 and Glu5 codons at mgtQ are critical for determining mgtQ translation efficiency and thus control mgtB translation because when we substituted the Asp2 and Glu5 in mgtQ to Ala codons, MgtB protein levels were low, regardless of the rpmE1 or rpmE2 mutations (Fig. 7A). As a control experiment, Fur protein levels were unaffected in all tested conditions (Fig. 7B).
FIG 7.
mgtQ translation promotes MgtB production and is required for Salmonella virulence in mice. (A and B) Western blot analysis of crude extracts prepared from wild-type, rpmE1, rpmE2, or rpmE1 rpmE2 Salmonella strains with either the wild-type mgtQ (14028s, EN1119, EN1120, or EN1368, respectively) or mgtQ2,5Ala (EN1389, EN1408, EN1409, or EN1414, respectively) gene probed with anti-MgtB (A) or anti-Fur (B) antibodies to detect MgtB or Fur proteins, respectively. Bacteria were grown for 5 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. (C) The mgtQ2,5Ala substitution attenuates Salmonella virulence in mice. Survival of C3H/HeN mice inoculated intraperitoneally with approximately 3000 CFU of wild-type (14028s), mgtQ2,5Ala mutant (EN1389), and mgtB deletion mutant (EN481) Salmonella strains.
The Asp2 and Glu5 codons at mgtQ are required for Salmonella virulence by ensuring MgtB production.
The mgtB gene is required for Salmonella virulence in Nramp1 (natural resistance-associated macrophage protein 1)-expressing mice (20). Given that the Asp2 and Glu5 to Ala substitution in mgtQ decreased MgtB protein levels (Fig. 7A), we tested whether the Asp2 and Glu5 codons in mgtQ would be required for Salmonella virulence by controlling MgtB protein production. When approximately 3,000 CFU of Salmonella were injected intraperitoneally in C3H/HeN Nramp1+/+ mice, the Salmonella strain with the Asp2 and Glu5 to Ala substitution in mgtQ was attenuated for virulence in mice compared to wild type (Fig. 7C). As a control, a strain lacking the mgtB gene was completely defective for virulence in mice (Fig. 7C). Therefore, these data indicate that the Asp2- and Glu5-mediated ribosomal destabilization at mgtQ guarantees MgtB Mg2+ transporter production, which might be critical for Salmonella pathogenesis by acquiring Mg2+ ions during infection.
DISCUSSION
Here, we discovered mgtQ, an 8-amino-acid (aa) ORF in the mgtCBRU operon in S. enterica. The mgtQ ORF is located between the mgtC and mgtB genes and is required for downstream MgtB Mg2+ transporter production. The mgtQ ORF harbors Asp and Glu codons at positions 2 and 5, respectively, which intrinsically impede translation (see Fig. S5). The Asp- and Glu-mediated decrease in mgtQ translation reflects that mgtQ-translating ribosomes might be destabilized at mgtQ. In turn, this decrease in mgtQ translation efficiency promotes MgtB production possibly because it favors the formation of a stem-loop structure that suppresses the formation of a mutually exclusive translation-inhibitory stem-loop structure near the RBS of the mgtB gene. Thus, the substitution of Asp and Glu codons in mgtQ decreased MgtB Mg2+ transporter production and attenuated virulence in mice by affecting MgtB-mediated Salmonella’s ability to import Mg2+ ions during infection.
Conservation of mgtQ among several bacteria harboring the mgtC and mgtB genes in the same transcription unit (see Fig. S2) suggests that mgtQ might be required for a condition in which both the MgtC virulence protein and MgtB Mg2+ transporter act together. However, the genomic distribution of the mgtQ ORF in other bacteria also suggests that the mgtQ ORF is linked to the mgtB gene. In Brucella melitensis, the mgtB gene precedes the mgtC gene and is also separated by two other genes. In this case, we found an mgtQ-like ORF upstream of the mgtB gene (see Fig. S2). Mycobacterium tuberculosis contains the mgtC gene, but not the mgtB gene. As expected, we could not find an mgtQ ortholog in this strain (see Fig. S2). Together with our findings showing that mgtQ affects translation of the mgtB gene, the distribution suggests that mgtQ could be a cis-acting regulatory element that guarantees MgtB Mg2+ transporter production.
Translation initiation regions in mRNAs are often folded into a secondary structure that occludes ribosome binding and inhibits translation. To initiate protein synthesis, such a ribosome-inaccessible secondary structure needs to be destabilized to allow the ribosome to bind the Shine-Dalgarno sequence and/or the start codon. This could be achieved by a variety of mechanisms that release the RBS/start codon. RNA-binding proteins and/or small regulatory RNAs can bind the anti-RBS sequence to release the RBS site or an endonucleolytic cleavage could remove upstream anti-RBS sequence that was base-paired to the RBS and allow the ribosome to access the RBS (4, 6). Small ligands could bind to the mRNA and promote to form an alternative RNA secondary structure that releases the RBS from the translation-inhibitory stem-loop structure (6, 21). In this example, translation of the short ORF mgtQ also contributes to destabilizing a translation-inhibitory stem-loop structure of the downstream mgtB gene, similarly to a transcription attenuation mechanism found in the E. coli trp operon (22). However, the mgtQ-mediated translational control differs from that of trpL in the trp operon because the mgtQ ORF and its neighboring sequence are associated with a translation-inhibitory stem-loop structure, while the trpL leader peptide gene is associated with a Rho-independent transcriptional terminator (22). Moreover, mgtQ is located in the intergenic region of the first mgtC gene and the second mgtB gene within the mgtCBRU operon. In contrast, most known transcription attenuators, including the trp operon, are found in the 5′ leader regions of the operons (22, 23).
During mgtQ translation, the mgtQ-translating ribosome occupies an mRNA sequence overlapping mgtQ (region 1) and allows the formation of stem-loop structure 2:3, which sequesters the anti-RBS sequence and releases the RBS of the mgtB gene. The mgtQ-mediated mgtB RBS release is further enhanced by the codon composition of the mgtQ ORF because mgtQ contains Asp and Glu codons that are known to induce ribosome destabilization (18). Given that the ribosomal subunit L31 could counteract the Asp and Glu codon-mediated ribosome destabilization (18), mgtQ’s effect on mgtB translation could be maximized in a strain lacking the rpmE1 gene (Fig. 6 and 7) or a condition that decreases the availability of the L31 subunit encoded by the rpmE1 gene.
A translating ribosome can stall or be destabilized on mRNA depending on mRNA sequences. Two or more proline codons on mRNA induce ribosome stalling due to the intrinsic difficulty in forming a peptide bond when peptidyl-prolyl-tRNA at the P-site transfers a peptidyl-prolyl group to an incoming prolyl-tRNA at the A-site (24, 25). In this case, elongation factor EF-P rescues the stalled ribosome to continue translation (24, 25). When cellular free tryptophan levels are high, expression of the tna operon encoding tryptophanase in E. coli is induced by a mechanism that also involves yet another type of ribosome stalling (26). A ribosome translating the TnaC leader peptide in the tna operon stalls when the ribosome reaches the last Pro sense codon in the P-site and UGA stop codon in the A-site (27). In this case, Trp codon at position 12 of tnaC is required for the ribosome stalling because Trp12 in the nascent peptide binds to the ribosome exit tunnel and blocks the nascent peptide release at the tnaC UGA stop codon and ribosome dissociation. This stalling prevents Rho loading and thus Rho-dependent transcription termination, thus increasing tna operon expression. In addition, the stalling can be eventually relieved by release factor RF-3 and ribosome recycling factor RRF (26). Repeated acidic Asp and Glu codons on mRNA also induce ribosome destabilization and subsequent translation abortion because Asp and Glu residues on a nascent peptide interact with the interior of the exit tunnel (18). In this case, the ribosomal subunit L31 counteracts the destabilized ribosome to continue translation on mRNA. Interestingly, Salmonella harbors two L31 subunits encoded by the rpmE1 and rpmE2 genes, respectively. Based on our results that rpmE1 and rpmE2 mutations showed opposing effects on mgtQ′-gfp (Fig. 6) and reports from others that one of two L31 subunits is expressed in Zn2+-depleted media (28), the functional roles of two L31 subunits might differ.
Finally, the mgtCBRU operon contains several small ORFs including the third mgtR and fourth mgtU genes encoding 30- and 28-aa regulatory peptides that guide MgtC and MgtB to the FtsH-mediated proteolysis, respectively (29, 30). The 12-aa mgtM and 16-aa mgtP ORFs are located in the 5′ leader region of the operon and mediate ATP- and charged tRNAPro-responsive transcriptional control of the mgtCBRU operon, respectively (31, 32). The 8-aa mgtQ is located in the mgtC-mgtB intergenic region and controls mgtB translation via a mechanism similar to acidic residue-mediated ribosomal destabilization. Although it is not clear why the mgtCBRU operon harbors multiple small ORFs to control expression of the entire operon or a part of the operon, Salmonella seems to have evolved to fine-tune the levels of proteins within this virulence operon in response to different signals, including those from the host environment.
MATERIALS AND METHODS
Bacterial strains, plasmids, oligodeoxynucleotides, and growth conditions.
Bacterial strains and plasmids used in this study are listed in Table S1 in the supplemental material. All S. enterica serovar Typhimurium strains were derived from the wild-type strain 14028s (33) and were constructed by one-step gene inactivation method (34) and/or P22-mediated transduction as previously described (35). DNA oligonucleotides are listed in Table S2. Bacteria were grown at 37°C in Luria-Bertani broth, N-minimal media (36) supplemented with 0.1% Casamino Acids and 38 mM glycerol, and the indicated concentrations of MgCl2. E. coli DH5α was used as the host for preparing plasmid DNA. Ampicillin was used at 50 μg ml−1, chloramphenicol was used at 20 μg ml−1, kanamycin was used at 20 μg ml−1, tetracycline was used at 10 μg ml−1, fusaric acid (37) was used at 12 μg ml−1, and l-arabinose was used at 0.2% (wt/vol). See Text S1 for more information.
Bacterial strains and plasmids used in this study. Download Table S1, DOCX file, 0.04 MB (44.5KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Primers used in this study. Download Table S2, DOCX file, 0.03 MB (30.1KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Data availability.
All other relevant data are available from the corresponding author upon reasonable request.
Supplemental materials and methods. Download Text S1, DOCX file, 0.04 MB (37KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
ACKNOWLEDGMENTS
This study was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT and Future Planning (NRF-2019R1A2C2003460 and NRF-2020M3A9H5104235 to E.-J.L. and NRF-2018R1D1A1B07043844 to E.C.), and a KU-FRG grant from Korea University.
E.-J.L. designed the research, analyzed the data, and wrote the manuscript. E.C. performed the experiments and wrote the manuscript. Y.H. performed the experiments. S.P. and H.K. performed mouse experiments. J.-S.L. analyzed the data and edited the manuscript.
We declare there are no conflicts of interest.
Footnotes
Citation Choi E, Han Y, Park S, Koo H, Lee J-S, Lee E-J. 2021. A translation-aborting small open reading frame in the intergenic region promotes translation of a Mg2+ transporter in Salmonella Typhimurium. mBio 12:e03376-20. https://doi.org/10.1128/mBio.03376-20.
REFERENCES
- 1.Govantes F, Andujar E, Santero E. 1998. Mechanism of translational coupling in the nifLA operon of Klebsiella pneumoniae. EMBO J 17:2368–2377. doi: 10.1093/emboj/17.8.2368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Koonin EV, Wolf YI. 2008. Genomics of bacteria and archaea: the emerging dynamic view of the prokaryotic world. Nucleic Acids Res 36:6688–6719. doi: 10.1093/nar/gkn668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Thorpe HA, Bayliss SC, Hurst LD, Feil EJ. 2017. Comparative analyses of selection operating on nontranslated intergenic regions of diverse bacterial species. Genetics 206:363–376. doi: 10.1534/genetics.116.195784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.McCarthy JE, Gualerzi C. 1990. Translational control of prokaryotic gene expression. Trends Genet 6:78–85. doi: 10.1016/0168-9525(90)90098-q. [DOI] [PubMed] [Google Scholar]
- 5.Desnoyers G, Bouchard MP, Masse E. 2013. New insights into small RNA-dependent translational regulation in prokaryotes. Trends Genet 29:92–98. doi: 10.1016/j.tig.2012.10.004. [DOI] [PubMed] [Google Scholar]
- 6.Geissmann T, Marzi S, Romby P. 2009. The role of mRNA structure in translational control in bacteria. RNA Biol 6:153–160. doi: 10.4161/rna.6.2.8047. [DOI] [PubMed] [Google Scholar]
- 7.Oppenheim DS, Yanofsky C. 1980. Translational coupling during expression of the tryptophan operon of Escherichia coli. Genetics 95:785–795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lee EJ, Groisman EA. 2010. An antisense RNA that governs the expression kinetics of a multifunctional virulence gene. Mol Microbiol 76:1020–1033. doi: 10.1111/j.1365-2958.2010.07161.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yeom J, Pontes MH, Choi J, Groisman EA. 2018. A protein that controls the onset of a Salmonella virulence program. EMBO J 37. doi: 10.15252/embj.201796977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Smith DL, Tao T, Maguire ME. 1993. Membrane topology of a P-type ATPase: the MgtB magnesium transport protein of Salmonella Typhimurium. J Biol Chem 268:22469–22479. doi: 10.1016/S0021-9258(18)41553-0. [DOI] [PubMed] [Google Scholar]
- 11.Soncini FC, Garcia Vescovi E, Solomon F, Groisman EA. 1996. Molecular basis of the magnesium deprivation response in Salmonella Typhimurium: identification of PhoP-regulated genes. J Bacteriol 178:5092–5099. doi: 10.1128/jb.178.17.5092-5099.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee EJ, Pontes MH, Groisman EA. 2013. A bacterial virulence protein promotes pathogenicity by inhibiting the bacterium’s own F1Fo ATP synthase. Cell 154:146–156. doi: 10.1016/j.cell.2013.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Maguire ME, Cowan JA. 2002. Magnesium chemistry and biochemistry. Biometals 15:203–210. doi: 10.1023/a:1016058229972. [DOI] [PubMed] [Google Scholar]
- 14.Wacker WE. 1969. The biochemistry of magnesium. Ann N Y Acad Sci 162:717–726. doi: 10.1111/j.1749-6632.1969.tb13003.x. [DOI] [PubMed] [Google Scholar]
- 15.Molina N, van Nimwegen E. 2008. Universal patterns of purifying selection at noncoding positions in bacteria. Genome Res 18:148–160. doi: 10.1101/gr.6759507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tsai CH, Liao R, Chou B, Palumbo M, Contreras LM. 2015. Genome-wide analyses in bacteria show small-RNA enrichment for long and conserved intergenic regions. J Bacteriol 197:40–50. doi: 10.1128/JB.02359-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zuker M. 2003. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415. doi: 10.1093/nar/gkg595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chadani Y, Niwa T, Izumi T, Sugata N, Nagao A, Suzuki T, Chiba S, Ito K, Taguchi H. 2017. Intrinsic ribosome destabilization underlies translation and provides an organism with a strategy of environmental sensing. Mol Cell 68:528–539 e5. doi: 10.1016/j.molcel.2017.10.020. [DOI] [PubMed] [Google Scholar]
- 19.Zwir I, Shin D, Kato A, Nishino K, Latifi T, Solomon F, Hare JM, Huang H, Groisman EA. 2005. Dissecting the PhoP regulatory network of Escherichia coli and Salmonella enterica. Proc Natl Acad Sci U S A 102:2862–2867. doi: 10.1073/pnas.0408238102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Choi E, Choi S, Nam D, Park S, Han Y, Lee JS, Lee EJ. 2017. Elongation factor P restricts Salmonella’s growth by controlling translation of a Mg2+ transporter gene during infection. Sci Rep 7:42098. doi: 10.1038/srep42098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Breaker RR. 2018. Riboswitches and translation control. Cold Spring Harb Perspect Biol 10:a032797. doi: 10.1101/cshperspect.a032797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Merino E, Yanofsky C. 2005. Transcription attenuation: a highly conserved regulatory strategy used by bacteria. Trends Genet 21:260–264. doi: 10.1016/j.tig.2005.03.002. [DOI] [PubMed] [Google Scholar]
- 23.Naville M, Gautheret D. 2010. Transcription attenuation in bacteria: theme and variations. Brief Funct Genomics 9:178–189. doi: 10.1093/bfgp/elq008. [DOI] [PubMed] [Google Scholar]
- 24.Ude S, Lassak J, Starosta AL, Kraxenberger T, Wilson DN, Jung K. 2013. Translation elongation factor EF-P alleviates ribosome stalling at polyproline stretches. Science 339:82–85. doi: 10.1126/science.1228985. [DOI] [PubMed] [Google Scholar]
- 25.Doerfel LK, Wohlgemuth I, Kothe C, Peske F, Urlaub H, Rodnina MV. 2013. EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science 339:85–88. doi: 10.1126/science.1229017. [DOI] [PubMed] [Google Scholar]
- 26.Gong F, Ito K, Nakamura Y, Yanofsky C. 2001. The mechanism of tryptophan induction of tryptophanase operon expression: tryptophan inhibits release factor-mediated cleavage of TnaC-peptidyl-tRNAPro. Proc Natl Acad Sci U S A 98:8997–9001. doi: 10.1073/pnas.171299298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gong F, Yanofsky C. 2002. Instruction of translating ribosome by nascent peptide. Science 297:1864–1867. doi: 10.1126/science.1073997. [DOI] [PubMed] [Google Scholar]
- 28.Nanamiya H, Akanuma G, Natori Y, Murayama R, Kosono S, Kudo T, Kobayashi K, Ogasawara N, Park SM, Ochi K, Kawamura F. 2004. Zinc is a key factor in controlling alternation of two types of L31 protein in the Bacillus subtilis ribosome. Mol Microbiol 52:273–283. doi: 10.1111/j.1365-2958.2003.03972.x. [DOI] [PubMed] [Google Scholar]
- 29.Alix E, Blanc-Potard AB. 2008. Peptide-assisted degradation of the Salmonella MgtC virulence factor. EMBO J 27:546–557. doi: 10.1038/sj.emboj.7601983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yeom J, Shao Y, Groisman EA. 2020. Small proteins regulate Salmonella survival inside macrophages by controlling degradation of a magnesium transporter. Proc Natl Acad Sci U S A 117:20235–20243. doi: 10.1073/pnas.2006116117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lee EJ, Groisman EA. 2012. Control of a Salmonella virulence locus by an ATP-sensing leader messenger RNA. Nature 486:271–275. doi: 10.1038/nature11090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lee EJ, Groisman EA. 2012. Tandem attenuators control expression of the Salmonella mgtCBR virulence operon. Mol Microbiol 86:212–224. doi: 10.1111/j.1365-2958.2012.08188.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fields PI, Swanson RV, Haidaris CG, Heffron F. 1986. Mutants of Salmonella Typhimurium that cannot survive within the macrophage are avirulent. Proc Natl Acad Sci U S A 83:5189–5193. doi: 10.1073/pnas.83.14.5189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Datsenko KA, Wanner BL. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645. doi: 10.1073/pnas.120163297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Davis RW, Bolstein D, Roth JR. 1980. Advanced bacterial genetics. Cold Spring Harbor Laboratories, Cold Spring Harbor, NY. [Google Scholar]
- 36.Snavely MD, Miller CG, Maguire ME. 1991. The mgtB Mg2+ transport locus of Salmonella Typhimurium encodes a P-type ATPase. J Biol Chem 266:815–823. doi: 10.1016/S0021-9258(17)35246-8. [DOI] [PubMed] [Google Scholar]
- 37.Maloy SR, Nunn WD. 1981. Selection for loss of tetracycline resistance by Escherichia coli. J Bacteriol 145:1110–1111. doi: 10.1128/JB.145.2.1110-1111.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The intergenic regions between the mgtC and mgtB genes of other enteric bacteria harbor mgtQ orthologs. Alignment of the nucleotide sequences corresponding to part of the intergenic regions between the mgtC and mgtB genes from Salmonella enterica, Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans. Sequences in boldface correspond to mgtQ. Sequences in green, purple, blue, and red represent regions involved in stem-loop structures (SR1, SR2, SR3, and SR4, respectively) shown in Fig. 1. The predicted RBS for each ORF is underlined. Asterisks correspond to nucleotides conserved in all listed species. Download FIG S1, TIF file, 0.2 MB (187.4KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
The presence of mgtQ is conserved in enteric bacteria where the mgtC and mgtB genes are in the same operon. In Salmonella enterica, mgtQ is located in the mgtC-mgtB intergenic region. Similarly, the mgtC-mgtB intergenic regions from Erwinia tasmaniensis, Yersinia pestis, Yersinia enterocolitica, and Serratia proteamaculans include mgtQ-like ORFs. In Brucella melitensis, the mgtC and mgtB genes appear to be part of separate transcription units. An mgtQ-like sequence is found upstream of the mgtB gene. Mycobacterium tuberculosis harbors the mgtC gene but not mgtB or mgtQ. Download FIG S2, TIF file, 0.2 MB (170.9KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Schematic representation of reporter systems used in this study. (A) A partial sequence of pfpv25 plasmid, a plasmid for transcriptional fusion to a promoterless gfp gene. (B) A partial sequence of ptGFP, an engineered plasmid for translational fusion to a promoterless gfp gene. In ptGFP plasmid, a portion of sequence in pfpv25 (indicated in blue) was deleted to remove the ribosome binding site and the start codon of the gfp gene. Download FIG S3, TIF file, 0.09 MB (89.7KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
mgtQ translation increases MgtB protein levels but not mRNA levels. (A) Schematic representation of an mgtQ chromosomal mutant with the start codon substituted by the stop codon and the DNA fragments amplified by QRT-PCR in panels D and E. (B and C) Western blot analysis of crude extracts prepared from Salmonella strains with the wild-type mgtQ (mgtQWT) or the mgtQ derivative with the start to stop codon substitution (mgtQATG→TAG) grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+. The amounts of MgtB and Fur proteins were determined by anti-MgtB (B) and anti-Fur (C) antibodies. (D and E) Relative mRNA levels of the mgtC (D) and mgtB (E) coding regions produced by wild-type (mgtQWT) Salmonella and the mgtQ mutant with the start codon replaced by the stop codon (mgtQATG→TAG) grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+. The means and SD from three independent measurements are shown. Download FIG S4, TIF file, 0.1 MB (144.3KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
The Asp2 and Glu5 to Ala substitutions enhance detection of the C-terminally 8×myc-tagged MgtQ peptides. (A) Schematic representation of Salmonella strains used in this study. (B-C) Western blot analysis of crude extracts prepared from Salmonella strains with the C-terminally 8×myc-tag fused to the wild-type mgtQ (EN1469) or mgtQ2,5Ala (EN1471) genes probed with anti-Myc (B) or anti-Fur (C) antibodies to detect MgtQ or Fur proteins, respectively. MgtQ-8×myc showed a higher SDS-PAGE-displayed molecular weight (∼20 kDa) than its predicted size (10.4 kDa) due to high content of acidic amino acids (47.7% in MgtQWT-8×myc and 45.5% in MgtQ2,5Ala-8×myc) (Y. Guan, Q. Zhu, D. Huang, S. Zhao, L. Jan Lo, J. Peng, Sci Rep 5:13370, 2015, doi:10.1038/srep13370). Bacteria were grown for 5 h in N-minimal media containing 10 or 0.01 mM Mg2+ as described in Materials and Methods. Download FIG S5, TIF file, 0.1 MB (119.8KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Asp2 and Glu5 codons in mgtQ are required for mgtB translation independently of the ability to form stem-loop 1:2. (A) A possible secondary structure of mgtQ-harboring mRNA sequences. Substituted nucleotide sequences of Asp2 and Glu5 codons in mgtQ are indicated at the side of Asp2 and Glu5 sequences. (B-C) Schematic representation of mgtQ′-gfp (B) and mgtB′-gfp constructs (C) used in this experiment. (D) Fluorescence produced by wild-type (14028s) Salmonella harboring the promoterless plasmid vector (ptGFP), or derivatives with a gfp translational fusion to the wild-type mgtQ (pmgtQ′-gfp) or Asp2 (GAT) Glu5 (GAG) to Gly2 (GGT) Ala5 (GCT)-substituted mgtQ (pmgtQ 2Gly, 5Ala′-gfp). (E) Fluorescence produced by wild-type (14028s) Salmonella harboring the plasmid vector (vector), or derivatives with a gfp translational fusion to the mgtB gene that includes the wild-type mgtQ or the mgtQ mutant with the Asp2 and Glu5 substitutions as described previously (D). Bacteria were grown for 4 h in N-minimal media containing 0.01 mM Mg2+ as described in Materials and Methods. Means and SD from three independent measurements are shown. Download FIG S6, TIF file, 0.1 MB (150KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Tuning L31 protein production restores MgtB production. (A and B) Western blot analysis of crude extracts prepared from wild-type or the rpmE1 mutant (EN1119) Salmonella harboring the rpmE1 gene from an arabinose inducible promoter (pBAD-rpmE1) probed with anti-MgtB (A) or anti-Fur (B) antibodies to detect MgtB or Fur proteins, respectively. Bacteria were grown for 5 h in N-minimal media containing 0.01 mM Mg2+ in the presence of the indicated concentration of arabinose. Download FIG S7, TIF file, 0.09 MB (97.3KB, tif) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Bacterial strains and plasmids used in this study. Download Table S1, DOCX file, 0.04 MB (44.5KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Primers used in this study. Download Table S2, DOCX file, 0.03 MB (30.1KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Supplemental materials and methods. Download Text S1, DOCX file, 0.04 MB (37KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.
Data Availability Statement
All other relevant data are available from the corresponding author upon reasonable request.
Supplemental materials and methods. Download Text S1, DOCX file, 0.04 MB (37KB, docx) .
Copyright © 2021 Choi et al.
This content is distributed under the terms of the Creative Commons Attribution 4.0 International license.