Data on the performance of saliva specimens for diagnosing coronavirus disease 2019 (COVID-19) in ambulatory patients are scarce and inconsistent. We assessed saliva-based specimens for detecting severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) by reverse transcriptase PCR (RT-PCR) in the community setting and compared three different collection methods.
KEYWORDS: COVID-19, saliva, SARS-CoV-2, PCR, diagnostics
ABSTRACT
Data on the performance of saliva specimens for diagnosing coronavirus disease 2019 (COVID-19) in ambulatory patients are scarce and inconsistent. We assessed saliva-based specimens for detecting severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) by reverse transcriptase PCR (RT-PCR) in the community setting and compared three different collection methods. This prospective study was conducted in three primary care centers. RT-PCR was performed on paired nasopharyngeal swabs (NPS) and saliva samples collected from outpatients with a broad clinical spectrum of illness. To assess differences in collection methods, saliva specimens were obtained in a different way in each of the participating centers: supervised collection (SVC), oropharyngeal washing (OPW), and self-collection (SC). Pairs of NPS and saliva samples from 577 patients (median age, 39 years; 44% men; 42% asymptomatic) were collected and tested, and 120 (20.8%) gave positive results. The overall agreement with NPS results and kappa coefficients (κ) for saliva samples obtained by SVC, OPW, and SC were 95% (κ = 0.85), 93.4% (κ = 0.76), and 93.3% (κ = 0.76), respectively. The sensitivities (95% confidence intervals [95% CI]) of the saliva specimens ranged from 86% (72.6% to 93.7%) for SVC to 66.7% (50.4% to 80%) for SC samples. Sensitivity was higher for samples with lower cycle threshold (CT) values. The best RT-PCR performance was observed for SVC, with sensitivities (95% CI) of 100% (85.9% to 100%) in symptomatic individuals and 88.9% (50.7% to 99.4%) in asymptomatic individuals at CT values of ≤30. We conclude that saliva is an acceptable specimen for the detection of SARS-CoV-2 in the community setting. Specimens collected under supervision perform comparably to NPS and can effectively identify individuals at higher risk of transmission under real-life conditions.
INTRODUCTION
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is usually detected by real-time reverse transcriptase PCR (RT-PCR) on RNA extracted from upper-respiratory-tract specimens, typically nasopharyngeal swabs (NPS). However, collection of NPS is uncomfortable for the patient and may induce coughing and sneezing, which may expose the health care provider to infectious aerosols. Therefore, alternative sampling has been investigated, including easy-to-obtain specimens with the potential for patient self-collection, such as saliva. Although initial studies investigating the use of saliva suggested that this specimen may be a good alternative sample to NPS, mixed results have been reported, with sensitivities in the range of 30% to 100% (1–12).
Most of the studies that have evaluated SARS-CoV-2 detection in saliva were conducted in patients admitted to hospital with known coronavirus disease 2019 (COVID-19) infection, and some of these studies were limited by the lack of simultaneous collection of NPS and saliva specimens and by the reduced composition of the cohorts, including mainly adults and symptomatic patients, all of which may limit the overall generalizability of the findings. Additionally, the procedures for collecting saliva specimens have differed substantially among the studies, from enhanced collection under direction or supervision by the clinician (1–5) to unsupervised self-collection by the participants (10–12). Differences in saliva sampling may explain the differing results of the published studies.
The use of saliva specimens in the ambulatory setting may be particularly appealing due to the ease of collection and reduced equipment required, but data are scarce and inconsistent. While findings from a study on 45 patients support the potential of these specimens for detecting SARS-CoV-2 from outpatients (5), reduced sensitivity relative to that of NPS has been reported in a recent community-based cohort, raising concerns about the use of saliva samples in this setting (12, 13). Therefore, to clarify the role of saliva as an alternate specimen type for the detection of SARS-CoV-2 in the community setting, larger clinical studies are needed.
The present study aimed to evaluate the performance of saliva-based specimens for detecting SARS-CoV-2 by RT-PCR in a prospective study of adults and children with suspected COVID-19. In this investigation, we performed RT-PCR on paired NPS and saliva samples collected from outpatients with a broad clinical spectrum of illness, including asymptomatic cases, undergoing SARS-CoV-2 testing, and we compared three different collection methods.
MATERIALS AND METHODS
Setting and study subjects.
This prospective, observational study was carried out by the COVID19-Elx-Rapid Diagnostic Tests Study Group at the Departments of Health 17 and 20 of the province of Alicante, Spain. The study was approved by the Hospital General Universitario de Elche COVID-19 Institutional Advisory Board. Patients enrolled in the study were those presenting for SARS-CoV-2 testing, as requested by their providers, at the facilities of three primary care centers (PCC). Both symptomatic patients and asymptomatic individuals who had been exposed to SARS-CoV-2 were invited to participate in the investigation by providing saliva samples immediately preceding the collection of the NPS. After written consent was obtained, demographic and clinical findings were recorded.
Specimen collection and processing.
Saliva specimens were collected into a 100-ml sterile empty container without transport medium. To assess differences in collection methods, saliva specimens were obtained in a different way in each of the participating PCC by random assignment. In center A, saliva specimens were obtained under the supervision of a health care worker (supervised collection [SVC]); in center B, saliva specimens were obtained after oropharyngeal washing (OPW), consisting of rinsing the mouth with 2 ml of saline solution and spitting the solution into the collection pot; and in center C, saliva specimens were collected independently by the individual providing the sample, following written instructions (self-collection [SC]). The instructions read as follows: “To collect saliva, bow your head forward to allow saliva to pool in the front of your mouth and spit up to a minimum of 1 ml of saliva (half a teaspoon) into the collection pot.” In the SVC center, subjects were instructed to produce and pool saliva in their mouths for 1 to 2 min, moving the mouth and passing the tip of the tongue across the cheeks and gums, and then to repeatedly spit a minimum of 1 ml saliva into the collection pot in the presence of a health care worker.
In the three PCC, nasopharyngeal samples were taken by qualified nurses following the same procedure and using the same kind of NPS. A flexible minitip swab was passed through the patient's nostril until the posterior nasopharynx was reached, left in place for several seconds to absorb secretions, and then slowly removed while rotating. Swabs were placed in 3 ml of sterile transport medium containing guanidine salt (Mole Bioscience, SUNGO Europe B.V., Amsterdam, Netherlands). To reduce variability in specimen transport, storage, and processing, the procedures were standardized in the three centers. Nasopharyngeal and saliva specimens were transported within 2 h of sample collection to the clinical microbiology laboratory for molecular analysis by RT-PCR. NPS samples were analyzed immediately, and saliva specimens were frozen (−20°C) and analyzed within 2 weeks after collection. Containers with saliva or OPW were thawed at room temperature; the sample was homogenized with the tip of an automatic pipette; and 300 μl was taken to the automated extraction plate.
SARS-CoV-2 detection.
Nucleic acid extraction was performed using 300 μl of specimen (NPS or saliva) on a chemagic 360 nucleic acid purification instrument (PerkinElmer España SL, Madrid, Spain). Then 10 μl of the eluate was used for a real-time RT-PCR assay targeting the E gene (LightMix Modular SARS-CoV (COVID19) E-gene kit; TIB Molbiol, Berlin, Germany; distributed by Roche). Testing was performed according to the manufacturer’s guidelines on a cobas z 480 analyzer (Roche, Basel, Switzerland). The number of cycles of amplification in RT-PCR (cycle threshold [CT] value) was assessed as a surrogate measure of the RNA concentration. According to the manufacturer, a CT value of ≤40 by PCR is considered a positive result.
Statistical methods.
Continuous variables were expressed as the median (Q1 to Q3) and categorical variables as percentages. The Wilcoxon test or Student's t test was used to compare two continuous variables; Kruskal-Wallis analysis of variance (ANOVA) was used for three or more continuous variables; and the chi-square test or Fisher's exact test was used for the comparison of categorical variables. The percentage of agreement (positive, negative, and overall) (PPA, NPA, and OPA) for saliva specimens obtained by SVC, OPW, and SC was calculated using the results of the RT-RCR for NPS as the reference standard. Performance agreement was evaluated using kappa coefficients (κ). Positive results of either NPS or saliva specimens were considered true-positive results for calculations of sensitivity. Patients with an undetermined result by RT-PCR for an NPS or saliva specimen were not considered for calculations of agreement. The performance of saliva specimens was also evaluated by stratifying by the presence of symptoms. Given the increasing evidence of a positive relationship between low CT values and the likelihood of culturing live virus and therefore infectious potential, we also evaluated the performance of saliva specimens according to CT values for NPS with different cutoff levels that have been associated with higher probabilities of positive viral culture (≤25 and ≤30) (14).
Statistical analyses were performed with R, version 4.0.3, software. The percentage of positive or negative agreement and kappa coefficients (κ) were calculated using the caret package (15). For graphical analysis, the ggplot2 package (16) was used.
We planned to include 133 patients per arm, assuming a sensitivity of 95% and a confidence level of 95%. With this sample size, the study would have a statistical power of 80% to detect a 10% difference in sensitivity between the different collection methods.
RESULTS
Patient characteristics and positivity rates.
A total of 634 patients (103 children) were invited to take part in the study between 15 September and 29 October 2020. Fifty-four patients (median [Q1 to Q3] age, 5.8 [4 to 10] years; 44 children) were unable to provide saliva specimens, and 3 (0.5%) specimens had insufficient sample for laboratory analysis. A total of 577 pairs of samples (229 SVC, 140 OPW, 208 SC) were included in the analyses. A flow chart of the patients is depicted in Fig. S1 in the supplemental material. The demographic and clinical characteristics of the patients according to the collection method for saliva specimens are shown in Table 1. There were 120 (20.8%) positive results for SARS-CoV-2 by either NPS or saliva, 50 (21.8%) in the SVC, 28 (20%) in the OPW, and 42 (20.2%) in the SC group. NPS and saliva samples from 2 (0.3%) and 9 (1.6%) patients, respectively, generated invalid transcription-mediated amplification results due to internal-control failure.
TABLE 1.
Demographic and clinical characteristics of the patients according to the method of collection of saliva specimens
| Characteristic | Overall cohort |
Supervised collection |
Oropharyngeal washing |
Self-collection |
||||
|---|---|---|---|---|---|---|---|---|
| All patients | Patients with SARS-CoV-2-positive RNA | All SVC patientsa | SVC patients with SARS-CoV-2-positive RNA | All OPW patients | OPW patients with SARS-CoV-2-positive RNA | All SC patients | SC patients with SARS-CoV-2-positive RNA | |
| No. of patients (%) | 577 | 120 (20.8) | 229 | 50 (21.8) | 140 | 28 (20) | 208 | 42 (20.2) |
| No. male (%) | 251 (43.5) | 57 (47.5) | 91 (39.7) | 22 (44) | 66 (47.1) | 13 (46.4) | 94 (45.2) | 22 (52.4) |
| Median age (yr) (Q1–Q3) | 39 (24–51) | 42 (28–54) | 39 (21–48) | 42 (29–49) | 36 (23–51) | 41 (27–54) | 40 (27–54) | 41 (28–57) |
| No. (%) of patients aged: | ||||||||
| 0–13 yr | 59 (10.2) | 5 (4.2) | 28 (12.2) | 3 (6) | 16 (11.4) | 1 (3.6) | 15 (7.2) | 1 (2.4) |
| 14–49 yr | 368 (63.8) | 79 (65.8) | 152 (66.4) | 35 (70) | 87 (62.1) | 18 (64.3) | 129 (62) | 26 (61.9) |
| 50–64 yr | 105 (18.2) | 26 (21.7) | 40 (17.5) | 10 (20) | 26 (18.6) | 8 (28.6) | 39 (18.8) | 8 (19) |
| ≥65 yr | 45 (7.8) | 10 (8.3) | 9 (3.9) | 2 (4) | 11 (7.9) | 1 (3.6) | 25 (12) | 7 (16.7) |
| No. (%) with the following comorbidity: | ||||||||
| Hypertension | 42 (7.3) | 13 (10.8) | 17 (7.4) | 3 (6) | 12 (8.6) | 2 (7.1) | 13 (6.2) | 8 (19) |
| Dyslipidemia | 37 (6.4) | 15 (12.5) | 17 (7.4) | 5 (10) | 11 (7.9) | 5 (19.2) | 9 (4.3) | 5 (11.9) |
| Diabetes | 19 (3.3) | 9 (7.5) | 10 (4.4) | 3 (6) | 4 (2.9) | 3 (10.7) | 5 (2.4) | 3 (7.1) |
| Cardiomyopathy | 17 (2.9) | 3 (2.5) | 9 (3.9) | 0 (0) | 7 (5) | 2 (7.1) | 1 (0.5) | 1 (2.4) |
| Obesity | 25 (4.3) | 10 (8.3) | 9 (3.9) | 2 (4) | 8 (5.7) | 3 (10.7) | 8 (3.8) | 5 (11.9) |
| Presence of symptoms on the day of sample collection | ||||||||
| No. (%) symptomatic | 336 (58.2) | 86 (71.7) | 145 (63.3) | 37 (74) | 68 (48.6) | 17 (60.7) | 123 (59.1) | 32 (76.2) |
| Median (Q1–Q3) days with symptoms | 4 (3–6) | 5 (3–7) | 3 (2–5) | 4 (2–7) | 4 (3–5) | 4 (3–6) | 4 (3–6) | 5 (3–7) |
| Sensitivity of saliva specimens (% [95% CI]) | ||||||||
| Global | 76.7 (67.9–83.7) | 86 (72.6–93.7)* | 75 (54.8–88.6) | 66.7 (50.4–80) | ||||
| Symptomatic patients | 77.9 (67.4–85.9) | 89.2 (73.6–96.5)** | 70.6 (44–88.6) | 68.8 (49.9–83.3) | ||||
| Asymptomatic patients | 73.5 (55.3–86.5) | 76.9 (46–93.8) | 81.8 (47.8–96.8) | 60 (27.4–86.3) | ||||
| CT , ≤25b | 96.7 (87.5–99.4) | 100 (85.9–100) | 93.3 (66–99.7) | 93.3 (66–99.7) | ||||
| CT , ≤30 | 91.6 (82.9–96.3) | 97.4 (84.9–99.9) | 81 (57.4–93.7) | 91.3 (70.5–98.5) | ||||
| CT , >30 | 43.2 (27.5–60.4) | 45.5 (18.1–75.4) | 57.1 (20.2–88.2) | 36.8 (17.2–61.4) | ||||
Asterisks indicate significant differences between supervised collection and self-collection (*, P = 0.028; **, P = 0.035).
CT, cycle threshold of RT-PCR for nasopharyngeal swabs (or saliva specimens in cases with a negative nasopharyngeal swab and positive saliva).
Sensitivities of the different specimens for SARS-CoV-2 detection and concordance between saliva specimens and NPS.
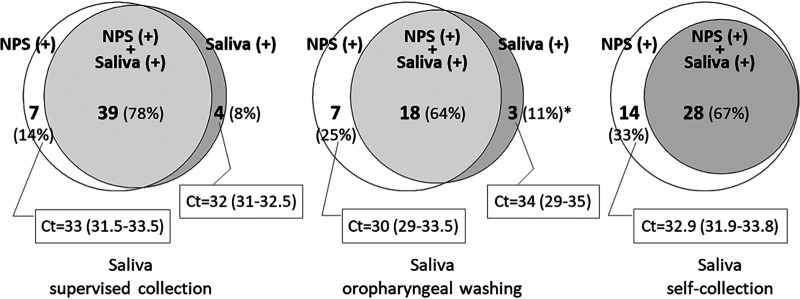
Table 2 shows the qualitative positive results for SARS-Cov-2 RNA obtained from NPS and saliva specimens according to the collection method, and Fig. 1 shows the concordance of positive test results between NPS and the different saliva specimen types. The sensitivity for NPS specimens was 95% (95% confidence interval [95% CI], 88.9% to 97.9%). Among saliva specimens, SVC showed the best case detection rate (43 of 50 infected patients), with significantly higher sensitivity than SC samples (86% [95% CI, 72.6% to 93.7%] versus 66.7% [95% CI, 50.4% to 80%]; P = 0.027). The OPW method detected 21 of 28 individuals (sensitivity, 75% [95% CI, 54.8% to 88.6%]) (Table 1). The greatest sensitivity was obtained by combining NPS sampling with saliva collected under supervision (sensitivity, 97.5% [95% CI, 92.3% to 99.3%]). Table 3 shows the agreement of the three different kinds of saliva specimens with NPS. The best agreement with NPS was found for the specimens of the SVC group, with a kappa coefficient of 0.85. For the OPW and SC groups, the kappa coefficient was 0.76.
TABLE 2.
Comparison of qualitative results from nasopharyngeal swabs and saliva specimens according to the collection method
| Nasopharyngeal swab resulta | No. of saliva samples with the indicated result, taken by: |
||||||||
|---|---|---|---|---|---|---|---|---|---|
| Supervised collection (n = 229) |
Oropharyngeal washing (n = 140) |
Self-collection (n = 208) |
|||||||
| Pos | Neg | Und | Pos | Neg | Und | Pos | Neg | Und | |
| Pos | 39 | 7 | 0 | 18 | 7 | 0 | 28 | 14 | 0 |
| Neg | 4 | 171 | 7 | 2 | 110 | 2 | 0 | 166 | 0 |
| Und | 0 | 1 | 1 | 0 | 0 | 0 | |||
Pos, positive; Neg, negative; Und, undetermined.
FIG 1.
Positivity for SARS-CoV-2 RNA in nasopharyngeal swabs (NPS) and in the different saliva specimens, including cycle threshold values of discordant specimens. The asterisk indicates one positive saliva specimen for which a negative NPS was confirmed 6 days later.
TABLE 3.
Agreement of the different kinds of saliva specimens with the nasopharyngeal swabs
| Saliva type | % agreement (95% CI) |
Performance agreement (κ) (95% CI) | ||
|---|---|---|---|---|
| Positive | Negative | Overall | ||
| Supervised collection | 84.8 (70.5–93.2) | 97.7 (93.9–99.3) | 95 (91–97.4) | 0.85 (0.76–0.93) |
| Oropharyngeal washing | 72 (50.4–87.1) | 98.2 (93.1–99.7) | 93.4 (87.5–96.8) | 0.76 (0.61–0.91) |
| Self-collection | 66.7 (50.4–80) | 100 (97.2–100) | 93.3 (88.7–96.1) | 0.76 (0.64–0.88) |
Performance of the different saliva specimens for SARS-CoV-2 detection by cycle threshold values and the presence of symptoms.
Median CT values were significantly lower in NPS than in their paired saliva specimens (P = 0.035), although in 19 (15.8%) patients, the saliva showed a lower CT value than the corresponding NPS. Median CT values in the three saliva specimen types were not significantly different from each other (P = 0.962) (Fig. 2). Figure 1 displays the concordance for SARS-CoV-2 detection between paired NPS and saliva samples, and the CT values of the discordant positive results. Median (Q1 to Q3) CT values for NPS-positive-only or saliva-positive-only specimens were 33 (31 to 34) and 32 (29 to 33.5), respectively.
FIG 2.
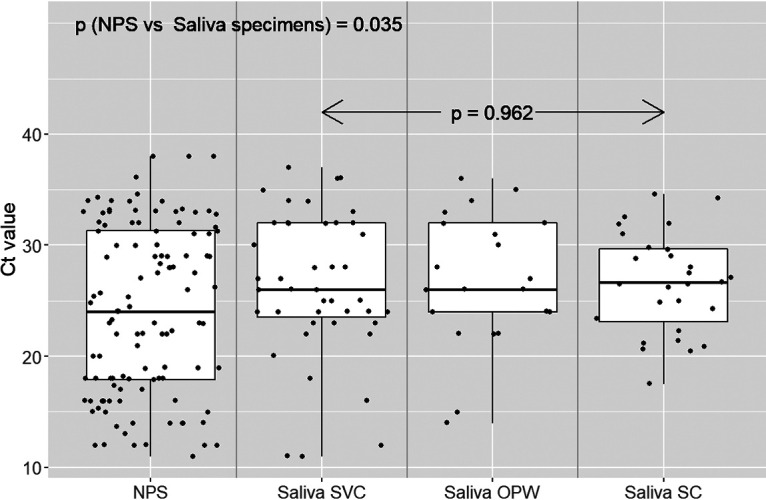
Cycle threshold values for pairs of nasopharyngeal swabs (NPS) and saliva specimens from all positive individuals. Each dot represents the CT value (in RT-PCR) for a positive specimen: NPS (n = 113), saliva obtained under supervised collection (SVC) (n = 43), saliva obtained after oropharyngeal washing (OPW) (n = 21), or saliva obtained by self-collection (SC) (n = 28).
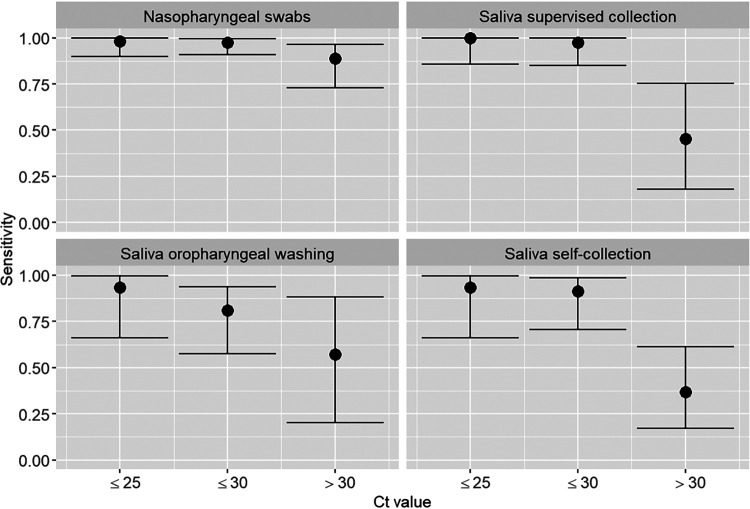
The sensitivities of the different specimens for SARS-CoV-2 detection according to CT values are depicted in Fig. 3. The sensitivities of the saliva specimens were higher at lower CT values. For CT values of ≤25, the median (95% CI) sensitivities of SVC, OPW, and SC sampling reached 100% (85.9% to 100%), 93.3% (66% to 99.7%), and 93.3% (66% to 99.7%), respectively, and decreased only minimally, to 97.4% (84.9% to 99.9%), 81% (57.4% to 93.7%), and 91.3% (70.5% to 98.5%), respectively, for CT values of ≤30. Figure S2 shows the sensitivity by sample type according to CT values and the presence of symptoms.
FIG 3.
Sensitivities of the different specimens for SARS-CoV-2 detection according to the CT value with nasopharyngeal swabs. In cases with a negative nasopharyngeal swab and positive saliva, the CT value of the saliva specimen was taken.
There were no significant differences in sensitivity between patients with and without active symptoms for the same CT values across the different specimens. The best RT-PCR performance was observed for NPS, closely followed by supervised collected saliva, with sensitivities (95% CI) of 98.4% (90.2% to 99.9%) and 100% (85.9% to 100%), respectively, in symptomatic individuals, and 95.2% (74.1% to 99.8%) and 88.9% (50.7% to 99.4%), respectively, in asymptomatic individuals at CT values of ≤30.
DISCUSSION
We confirmed that saliva is an acceptable specimen for the molecular detection of SARS-CoV-2 in the community setting and can effectively identify individuals with the highest risks of transmission under real-life conditions. This study revealed that the collection method may be critical for improving sensitivity. Saliva specimens obtained under supervision outperform self-collected samples and show higher sensitivity in symptomatic and asymptomatic patients.
As expected, the sensitivity of the saliva specimens increased in samples with low CT values. Indeed, supervised collected specimens performed almost as well as nasopharyngeal samples, with sensitivities well above 90% for patients with low CT values, who are considered to have the greatest potential to spread the virus (14, 17). For CT values of ≤25, the sensitivities of SVC, OPW, and SC saliva specimens reached 100%, 93%, and 93%, respectively, and for patients with CT values of ≤30, they were 97%, 81%, and 91%, respectively. Among NPS-only-positive individuals, we found only one case of saliva collected under supervision with a CT value of ≤30. Therefore, the use of saliva specimens in general, particularly when obtained under supervision, allowed the detection of SARS-CoV-2 in the vast majority of the patients with significant risks of transmission.
In contrast to most of the previous studies, which have been largely focused on inpatient populations, in the present investigation we included outpatients with a broad clinical spectrum of the illness, comprising children and asymptomatic cases. Like Williams et al. (11), we evaluated casual saliva specimens without previous fasting or enhancement techniques such as strong sniffing or coughing, used in other studies (4). Prior investigations comparing different samples for the molecular detection of SARS-CoV-2 in the community setting have reported lower sensitivity rates with saliva specimens than with NPS, ranging from 30% to 85% (11–13). The reduced performance has been attributed to the milder symptoms in outpatients, with viral loads lower than those in more-severe cases (18, 19), and to differences in the temporal dynamics of viral shedding between upper respiratory locations and saliva (20, 21), with lower viral loads in saliva samples (11). We did not find prominent differences in performance between samples from individuals with and without active symptoms, but, in line with other studies (8, 22, 23), we detected higher CT values, indicating lower viral loads, in saliva specimens than in the corresponding NPS, suggesting differences in viral shedding between the two compartments. In addition, the significant differences in the performances of the specimen types evaluated in our study suggest that variation in saliva sampling may have contributed to the disparities in sensitivity observed in previous investigations.
This study has limitations. The investigation focused on comparing three specific methods for the collection of saliva samples and was powered to detect rather large differences among groups. The sample size does not allow one to draw conclusions on the performance in particular subgroups, including children and patients tested at different time points of illness. Other potential weaknesses include the possibility that differences in performance among the collection methods could be confounded by differences in the populations and sampling among the three PCC. However, the populations were comparable in terms of gender, age, and the prevalence of SARS-CoV-2 RNA positivity, and to reduce variability in specimen transport, storage, and processing, the procedures were standardized in the three participating centers. Noteworthy, a substantial proportion of the children recruited, most of them very young children, were unable to provide the minimum of 1 ml of saliva sample after the procedure was explained, suggesting that this specimen type might be less suitable for this group. In addition, we used a particular detection system (cobas z 480 analyzer); other platforms may have yielded different results. The strengths of the study are that it was population based and was carried out under real-life conditions, enrolling consecutive outpatients of all ages presenting for testing due to symptoms and asymptomatic people who had come into contact with confirmed cases.
In conclusion, our results indicate that adequate collection of samples may be essential for the molecular diagnosis of COVID-19 when one is using saliva specimens. Saliva specimens obtained under supervision perform comparably to NPS and should be considered reliable samples for diagnosis of both symptomatic and asymptomatic individuals in the community setting, particularly for detecting individuals with significant risks of transmission. Although self-collected saliva would be the most advantageous way of sampling if mass testing were considered, these specimens had less sensitivity in our study. Further research is needed to determine whether other strategies of instruction, for example, videos or telecommunications, can substitute for the direct supervision of a health professional.
Supplementary Material
ACKNOWLEDGMENTS
This work was supported by the RD16/0025/0038 project as a part of the Plan Nacional Research + Development + Innovation (R+D+I) and was cofinanced by the Instituto de Salud Carlos III - Subdirección General de Evaluación y Fondo Europeo de Desarrollo Regional and Instituto de Salud Carlos III (Fondo de Investigaciones Sanitarias [grants PI16/01740, PI18/01861, CM 19/00160, and COV20-00005]).
All authors declare no conflict of interest.
The members of the COVID19-Elx-Rapid Diagnostic Tests Study Group are as follows: Félix Gutiérrez, Mar Masiá, Sergio Padilla, Guillermo Telenti, Lucia Guillen, María Andreo, Fernando Lidón, Vladimir Ospino, José López, Marta Fernández, Vanesa Agulló, Gabriel Estañ, Javier García, Cristina Martínez, Leticia Alonso, Joan Sanchís, Ángela Botella, Paula Mascarell, María Selene Falcón, Sandra Ruiz, José Carlos Asenjo, Carolina Ding, Mar Carvajal, Inmaculada Candela, Jorge Guijarro, Cristina la Moneda, Cristina Jara, Raquel Mora, Juan Manuel Quinto, Sergio Ros, Daniel Canal, Pascual Pérez, Carolina Garrido, Manuel Sánchez, Jaime Sastre, Carlos de Gregorio, Francisco Carrasco, Juan Navarro, Andrés Navarro, Nieves Gonzalo, Clara Pérez, Adoración Alcalá, José Luis Rincón, Montserrat Ruiz, and José Antonio Gutiérrez.
REFERENCES
- 1.To KK, Tsang OT, Leung WS, Tam AR, Wu TC, Lung DC, Yip CC, Cai JP, Chan JM, Chik TS, Lau DP, Choi CY, Chen LL, Chan WM, Chan KH, Ip JD, Ng AC, Poon RW, Luo CT, Cheng VC, Chan JF, Hung IF, Chen Z, Chen H, Yuen KY. 2020. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis 20:565–574. doi: 10.1016/S1473-3099(20)30196-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.To KK, Tsang OT, Yip CC, Chan KH, Wu TC, Chan JM, Leung WS, Chik TS, Choi CY, Kandamby DH, Lung DC, Tam AR, Poon RW, Fung AY, Hung IF, Cheng VC, Chan JF, Yuen KY. 2020. Consistent detection of 2019 novel coronavirus in saliva. Clin Infect Dis 71:841–843. doi: 10.1093/cid/ciaa149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hanson KE, Barker AP, Hillyard DR, Gilmore N, Barrett JW, Orlandi RR, Shakir SM. 2020. Self-collected anterior nasal and saliva specimens versus health care worker-collected nasopharyngeal swabs for the molecular detection of SARS-CoV-2. J Clin Microbiol 58:e01824-20. doi: 10.1128/JCM.01824-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Procop GW, Shrestha NK, Vogel S, Van Sickle K, Harrington S, Rhoads DD, Rubin BP, Terpeluk P. 2020. A direct comparison of enhanced saliva to nasopharyngeal swab for the detection of SARS-CoV-2 in symptomatic patients. J Clin Microbiol 58:e01946-20. doi: 10.1128/JCM.01946-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kojima N, Turner F, Slepnev V, Bacelar A, Deming L, Kodeboyina S, Klausner JD. 2020. Self-collected oral fluid and nasal swab specimens demonstrate comparable sensitivity to clinician-collected nasopharyngeal swab specimens for the detection of SARS-CoV-2. Clin Infect Dis doi: 10.1093/cid/ciaa1589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Jamal AJ, Mozafarihashjin M, Coomes E, Powis J, Li AX, Paterson A, Anceva-Sami S, Barati S, Crowl G, Faheem A, Farooqi L, Khan S, Prost K, Poutanen S, Taylor M, Yip L, Zhong XZ, McGeer AJ, Mubareka S, Toronto Invasive Bacterial Diseases Network COVID-19 Investigators. 2020. Sensitivity of nasopharyngeal swabs and saliva for the detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Clin Infect Dis doi: 10.1093/cid/ciaa848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Leung EC, Chow VC, Lee MK, Lai RW. 2020. Deep throat saliva as an alternative diagnostic specimen type for the detection of SARS-CoV-2. J Med Virol doi: 10.1002/jmv.26258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.McCormick-Baw C, Morgan K, Gaffney D, Cazares Y, Jaworski K, Byrd A, Molberg K, Cavuoti D. 2020. Saliva as an alternate specimen source for detection of SARS-CoV-2 in symptomatic patients using Cepheid Xpert Xpress SARS-CoV-2. J Clin Microbiol 58:e01109-20. doi: 10.1128/JCM.01109-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Khurshid Z, Zohaib S, Joshi C, Moin SF, Zafar MS, Speicher DJ. 2020. Saliva as a non-invasive sample for the detection of SARS-CoV-2: a systematic review. medRxiv 10.1101/2020.05.09.20096354. [DOI]
- 10.Wyllie AL, Fournier J, Casanovas-Massana A, Campbell M, Tokuyama M, Vijayakumar P, Warren JL, Geng B, Muenker MC, Moore AJ, Vogels CBF, Petrone ME, Ott IM, Lu P, Venkataraman A, Lu-Culligan A, Klein J, Earnest R, Simonov M, Datta R, Handoko R, Naushad N, Sewanan LR, Valdez J, White EB, Lapidus S, Kalinich CC, Jiang X, Kim DJ, Kudo E, Linehan M, Mao T, Moriyama M, Oh JE, Park A, Silva J, Song E, Takahashi T, Taura M, Weizman OE, Wong P, Yang Y, Bermejo S, Odio CD, Omer SB, Dela Cruz CS, Farhadian S, Martinello RA, Iwasaki A, Grubaugh ND, Ko AI. 2020. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N Engl J Med 383:1283–1286. doi: 10.1056/NEJMc2016359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Williams E, Bond K, Zhang B, Putland M, Williamson DA. 2020. Saliva as a noninvasive specimen for detection of SARS-CoV-2. J Clin Microbiol 58:e00776-20. doi: 10.1128/JCM.00776-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Becker D, Sandoval E, Amin A, De Hoff P, Diets A, Leonetti N, Lim YW, Elliott C, Laurent L, Grzymski J, Lu JT. 2020. Saliva is less sensitive than nasopharyngeal swabs for COVID-19 detection in the community setting. medRxiv 10.1101/2020.05.11.20092338. [DOI]
- 13.Torres M, Collins K, Corbit M, Ramirez M, Winters CR, Katz L, Ross M, Relkin N, Zhou W. 2020. Comparison of saliva and nasopharyngeal swab SARS-CoV-2 RT-qPCR testing in a community setting. J Infect doi: 10.1016/j.jinf.2020.11.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jefferson T, Spencer EA, Brassey J, Heneghan C. 2020. Viral cultures for COVID-19 infectious potential assessment—a systematic review. Clin Infect Dis doi: 10.1093/cid/ciaa1764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kuhn M. 2020. caret: classification and regression training. R package, version 6.0-86. https://CRAN.R-project.org/package=caret.
- 16.Wickham H, Sievert C. 2016. ggplot2: elegant graphics for data analysis. Springer-Verlag, New York, NY. [Google Scholar]
- 17.Basile K, McPhie K, Carter I, Alderson S, Rahman H, Donovan L, Kumar S, Tran T, Ko D, Sivaruban T, Ngo C, Toi C, O’Sullivan MV, Sintchenko V, Chen SC, Maddocks S, Dwyer DE, Kok J. 2020. Cell-based culture of SARS-CoV-2 informs infectivity and safe de-isolation assessments during COVID-19. Clin Infect Dis doi: 10.1093/cid/ciaa1579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Liu Y, Yan LM, Wan L, Xiang TX, Le A, Liu JM, Peiris M, Poon LLM, Zhang W. 2020. Viral dynamics in mild and severe cases of COVID-19. Lancet Infect Dis 20:656–657. doi: 10.1016/S1473-3099(20)30232-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chau NVV, Lam VT, Dung NT, Yen LM, Minh NNQ, Hung LM, Ngoc NM, Dung NT, Man DNH, Nguyet LA, Nhat LTH, Nhu LNT, Ny NTH, Hong NTT, Kestelyn E, Dung NTP, Xuan TC, Hien TT, Thanh Phong N, Tu TNH, Geskus RB, Thanh TT, Thanh Truong N, Binh NT, Thuong TC, Thwaites G, Tan LV, OUCRU COVID-19 research group. 2020. The natural history and transmission potential of asymptomatic SARS-CoV-2 infection. Clin Infect Dis 71:2679–2687. doi: 10.1093/cid/ciaa711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.He X, Lau EHY, Wu P, Deng X, Wang J, Hao X, Lau YC, Wong JY, Guan Y, Tan X, Mo X, Chen Y, Liao B, Chen W, Hu F, Zhang Q, Zhong M, Wu Y, Zhao L, Zhang F, Cowling BJ, Li F, Leung GM. 2020. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med 26:672–675. doi: 10.1038/s41591-020-0869-5. [DOI] [PubMed] [Google Scholar]
- 21.Zou L, Ruan F, Huang M, Liang L, Huang H, Hong Z, Yu J, Kang M, Song Y, Xia J, Guo Q, Song T, He J, Yen HL, Peiris M, Wu J. 2020. SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med 382:1177–1179. doi: 10.1056/NEJMc2001737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen JH, Yip CC, Poon RW, Chan KH, Cheng VC, Hung IF, Chan JF, Yuen KY, To KK. 2020. Evaluating the use of posterior oropharyngeal saliva in a point-of-care assay for the detection of SARS-CoV-2. Emerg Microbes Infect 9:1356–1359. doi: 10.1080/22221751.2020.1775133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pasomsub E, Watcharananan SP, Boonyawat K, Janchompoo P, Wongtabtim G, Suksuwan W, Sungkanuparph S, Phuphuakrat A. 2020. Saliva sample as a non-invasive specimen for the diagnosis of coronavirus disease 2019: a cross-sectional study. Clin Microbiol Infect doi: 10.1016/j.cmi.2020.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.