Abstract
Our goal is to estimate the true number of classes in a population, called the species richness. We consider the case where multiple frequency count tables have been collected from a homogeneous population and investigate a penalized maximum likelihood estimator under a negative binomial model. Because high probabilities of unobserved classes increase the variance of species richness estimates, our method penalizes the probability of a class being unobserved. Tuning the penalization parameter is challenging because the true species richness is never known, and so we propose and validate four novel methods for tuning the penalization parameter. We illustrate and contrast the performance of the proposed methods by estimating the strain-level microbial diversity of Lake Champlain over three consecutive years, and global human host-associated species-level microbial richness.
Keywords: Diversity, regularization, maximum likelihood, ecology, microbiome
1. Introduction
The species problem concerns estimating C, the number of classes that are present in a population. n individuals from the population can be sampled to find which classes they belong to, but only c classes are observed ( ). The problem is named for its origins in biological ecosystems, where C is species richness, or the total number of species. However, methods developed for the species problem can be applied to applications far removed from biology. For example, Efron and Thisted [9] estimated the number of words Shakespeare truly knew by modeling the frequencies of words in his published work, and Fegatelli and Tardella [10] estimated the number of cars covered by an insurer based on accident data where c cars had at least one accident.
In ecology, species richness is a quantitative measurement of ecosystem diversity. We focus on the specific application of estimating microbial diversity, that is, the number of strains of bacteria present in a population (e.g. a lake microenvironment or in an individual's oral cavity). Microbial diversity is often linked with ecosystem health, such as in the vaginal microbiome (where high diversity is associated with infection [20]) and in the gut microbiome (where high diversity is associated with healthy metabolism [17,18]). Microbial abundance data typically contain many species observed infrequently (rare species) and some which are observed a large number of times (abundant species). In data with both many rare and many abundant species, richness estimation methods often have high variance [28,30], motivating our regularized approach to estimation.
In this paper, we consider the penalized maximum likelihood approach of Wang and Lindsay [28] and investigate the open problem of selecting the penalization parameter. Because the true species richness C is never observed for any sample, common approaches to tuning parameter selection (e.g. cross-validation) cannot be employed. We propose using biological replicates to aid tuning. We find that replicate data is advantageous for tuning the required penalization parameter and provide a comparison of the performance of four proposed methods for tuning parameter selection.
This paper is organized as follows: We review species richness estimation in Section 2. In Section 3, we describe our extension to biological replicates, and in Section 4 we establish the utility of penalization via simulations. In Section 5, we propose several novel methods for tuning the penalization parameter, and in Section 6 we evaluate our proposals. In Section 7, we apply our methods to estimate microbial diversity in Lake Champlain, and Section 8 closes with a discussion of our results and suggested directions for future work. Software implementing the methods is available in the R package rre (regularized richness estimation), available at github.com/statdivlab/rre, and code to reproduce the simulation results is available at github.com/statdivlab/rre_sims.
2. Literature review
2.1. Poisson-mixture models
The classical model for estimating species abundance is a Poisson-mixture model. Under this model, the number of times we observe species i is where Λ is a random variable distributed according to some mixing distribution. Many mixing distributions have been proposed, including by [4,19,21] (see [5] for a comprehensive review). While our proposal may be easily generalized to other mixing distributions, in this paper, we consider the Gamma distribution for Λ, first considered by Greenwood and Yule [12] and Fisher et al. [11]. Note that we only observe , and therefore we are interested in estimating the parameters of a Poisson-mixture model using zero-truncated Poisson-mixed data.
Let be the number of classes observed k times, called the frequency counts. Then the set is a frequency count table, a common way to represent species abundance data. Because , the species problem can be framed as predicting given to obtain a species richness estimate .
Let with distribution function , and write . There exist both frequentist [6,11] and Bayesian [2,9] approaches to parameter estimation under this model. We will focus on frequentist maximum likelihood (penalized and unpenalized) in this paper.
2.2. Estimation and computation
The most straightforward maximum likelihood approach is to simultaneously maximize C and η using the full likelihood, which is known as the direct approach. A complication is that we have continuous parameters η as well as one discrete parameter C, and so derivative-based optimization methods are not appropriate. This issue has been studied by Lindsay and Roeder [14], who provide a discrete analog to the score function for this model. Two other approaches to likelihood maximization are the conditional and profile approaches. The conditional approach [25] involves writing the likelihood as the product
| (1) |
where
| (2) |
| (3) |
We see that the likelihood is the product of a binomial probability mass function Binomial , and a multinomial probability mass function Multinomial . This decomposition is convenient because is a function of η alone. Sanathanan [25]'s conditional approach to optimization involves first maximizing to obtain a conditional abundance estimate , then maximizing over C. The conditional estimate of C is then
| (4) |
where is the largest integer less than or equal to a. In the profile likelihood approach the expression in Equation (4) is substituted into the full likelihood, which gives us a function of η alone (see Wang and Lindsay [28]). The conditional and profile approaches have been shown to be asymptotically equivalent to the direct approach.
2.3. Challenges with Poisson-mixture models
A perennial issue in the species problem, especially for microbial datasets, is the instability of species richness estimators [24,30]. A proposal which encourages stability of estimates under Poisson-mixture models is due to Wang and Lindsay [28]. Their proposal is to add a penalty term to the log-likelihood that penalizes the probability of observing a species zero times. They consider the penalized log-likelihood
| (5) |
where is the log-likelihood and is a penalization parameter. Let . Then, for [28, Theorem 1]. In particular, for the penalized maximum likelihood solution is less than or equal to , the maximum likelihood estimate. Furthermore, for a large enough λ, ; that is, the penalized maximum likelihood estimate shrinks to the observed richness c.
We note that and so the addition of the ‘penalty’ term in fact increases the objective function (5) over the (unpenalized) likelihood . While technically smaller adds a larger reward to the objective function (for a fixed λ), we refer to the term as a penalty to be consistent with the terminology of Wang and Lindsay [28]. Wang and Lindsay [28] also consider two other penalty functions, which we do not discuss here except to note that our tuning parameter selection methods would equally apply to these other penalty functions.
Wang and Lindsay [28] show that a trade-off exists: greater values of λ correspond to a more stable estimator, but at the potential cost of negative bias. The choice of penalization parameter implies a preference for lower variance or lower bias, adding subjectivity to the estimation procedure. Wang and Lindsay [28] note that a limitation of their proposal is that ‘one must select a penalty function and a tuning parameter, and it is nigh impossible to make convincing statements about why one choice should be uniformly superior to another'. Furthermore, we expect different data sets to require different λ values for optimal mean squared error. The goal of this paper is to propose and investigate data-adaptive methods to select λ.
3. Extension to biological replicates
We focus on the gamma-Poisson model and penalized log-likelihood of the form (5), and consider the case where we observe r independent frequency count tables from the population under study. We will assume that these r frequency count tables are biological replicates drawn independently from the same population, i.e. they have a common structure specified by the parameters C and η. A set of r frequency count tables from the same population will be called a sample, and a single frequency count table will be referred to as a replicate. Let the number of species observed k times in replicate j be , . Let be the observed richness in replicate j.
In Section 5, we will require an objective function defined in terms of a subset of indices of the frequency count tables, so we define it that way now. Let . J indicates the data being used in the evaluation of the objective function, with meaning we use all replicates. Let be the unpenalized log-likelihood for replicate j. Using the fact that the draws are independent, our objective function for a fixed is the sum of the log-likelihoods for each individual replicate. We define the penalized log-likelihood for multiple samples as
| (6) |
| (7) |
By fixing we obtain the unpenalized log-likelihood for the set J. We define to be the penalized maximum likelihood estimate of C based on tuning parameter λ, that is,
| (8) |
where is a nuisance parameter.
To maximize the likelihood, we will use the direct approach, rather than the profile and conditional approaches reviewed in Section 2. While it may be faster to use the profile or conditional approach, the results are not always the same in finite samples. Since our objective is to evaluate approaches to tuning, we place a priority on the accuracy of optimization over speed of optimization.
To implement a direct optimization approach, we search over candidate C values in a grid. A gradient-based search over η values is used at each C in the grid to find the maximum penalized likelihood .
4. Evaluating penalized species richness estimates
Before evaluating potential tuning parameter selection methods, we first establish that penalization can improve richness estimation. While Wang and Lindsay [28] established that penalization can improve estimates when r = 1, this needs to be investigated when r>1.
We use square root mean square error (RMSE) as the primary criterion for evaluating the performance of estimators. Let be the number of simulations completed for one choice of , and be the solution of Equation (8) obtained in simulation number s. We define
| (9) |
In this simulation, if there exists some for which RMSE( ) is less than RMSE( , we conclude that penalization is effective.
We simulated using C = 1000, and . We found that provided a good fit to the data of Walsh et al. [27], and provided a good fit to the data of Tromas et al. [26] (see Section 7). For completeness, we also investigated nearby η values: and . In Table 1, we provide summary measures for frequency count tables generated under each choice of η. This characterizes the η values by the proportion of unobserved, rare or abundant species we would expect to see. We return to a discussion of the different data structures in Section 6. For each combination of η and r, we performed 100 simulations. We investigated the grid , and found that this grid was sufficiently expansive to ensure that the RMSE-minimizing λ was not on the boundary of the grid (see Figures 1 and 2).
Table 1. The expected proportion of unobserved (k = 0), singleton (k = 1), rare (k = 1, 2, 3), and abundant ( ) species for 4 choices of η.
| η | ( , ) | ( , ) | ( , ) | ( , ) | ( , ) |
|---|---|---|---|---|---|
| Proportion unobserved ( ) | 0.787 | 0.501 | 0.316 | 0.891 | 0.933 |
| Proportion singletons ( ) | 0.072 | 0.050 | 0.032 | 0.009 | 0.009 |
| Proportion rare ( ) | 0.130 | 0.097 | 0.061 | 0.016 | 0.017 |
| Proportion abundant ( ) | 0.028 | 0.340 | 0.583 | 0.083 | 0.040 |
| Expected max abundance ( ) |
We also give the expected frequency count of the most abundant species when .
Figure 1.
Estimates of C and their root-MSE over λ when and C = 1000. Results are based on 100 simulations per λ.
Figure 2.
Estimates of C and their root-MSE over λ when and C = 1000. Results are based on 100 simulations per λ.
The results of the simulation are shown in Table 2. For every parameter choice a reduction in RMSE( ) was found for some . Depending on simulation parameters the reduction varied from 22% to 70% compared to RMSE( ) for the best λ. These results show that for a variety of choices, penalization improves species richness estimation. We also see that the optimal λ value varies considerably over , ranging from 10 to 70. This suggests that there is not a universally appropriate λ choice. Therefore, tuning λ to the sample appears desirable. We note that the optimal choice of λ is not consistent either for fixed r and variable η, nor for fixed η and variable r.
Table 2. The penalized maximum likelihood estimate of C has lower RMSE for all investigated choices of η and r under a zero-truncated Gamma-mixed Poisson model for species abundances based on 100 simulations for each choice of η and r.
| r | RMSE ( ) | RMSE ( ) | ||
|---|---|---|---|---|
| 6 | 796.90 | 10 | 326.50 | |
| 10 | 527.53 | 15 | 286.27 | |
| 14 | 337.31 | 10 | 235.51 | |
| 6 | 735.95 | 15 | 516.37 | |
| 10 | 700.21 | 20 | 456.04 | |
| 14 | 666.55 | 20 | 470.35 | |
| 6 | 200.09 | 20 | 156.29 | |
| 10 | 213.22 | 55 | 142.64 | |
| 14 | 243.42 | 55 | 148.26 | |
| 6 | 401.17 | 55 | 147.13 | |
| 10 | 283.91 | 25 | 137.04 | |
| 14 | 415.19 | 70 | 126.72 |
is the value of λ which produced the lowest RMSE. is the estimate of C when , and is the estimate of C when .
To illustrate the bias-variance trade-off in this problem, we show the effect on of increasing λ in Figure 1 (when ) and in Figure 2 (when ). When , for small values of λ we observe lower variance in the estimates while incurring some positive bias, and the RMSE improves. However, as λ continues to increase, the bias term becomes large and negative and the RMSE increases. Similarly, when , we observe that the bias becomes large and negative and the RMSE increases. However, even for small values of λ, the estimate is not positively biased, and the larger RMSE is attributable to higher variance.
5. Methods for tuning λ
We have established that penalization can improve richness estimates, but different abundance structures (η) require different values of λ to minimize RMSE. We therefore develop methods to tune λ based on a sample. We propose several novel methods in this section. Each is evaluated in Section 6.
5.1. Method 0: no penalization
We compare all proposed methods to the unpenalized MLE using :
| (10) |
This method is fast and simple, making it an ideal baseline for comparison.
For all of the remaining methods (1)–(4), we generate estimates over , where is user-specified (e.g., in Section 4).
5.2. Method 1: minimum subset variance
Since large variance in C is a major concern in species richness estimation, ideally an estimator will have low variance. If this is the case, there should be low variance in estimates from equally sized subsets of J. For Method 1, we exploit the fact that we have replicate data by repeatedly partitioning the replicates into two subsets and calculating two estimates. We then select the λ, which yields the lowest between-subset variance. This partitioning is repeated p times to average out the arbitrary choice of subsets.
Let be the first subset of the lth partition and be the second subset of the lth partition, for . That is, , and . For each , let
| (11) |
| (12) |
We now have a corresponding to each in each subset of each partition. Our goal in Method 1 is to use these estimates to select the λ value which gave us the lowest average variance over all partitions, denoted by . The overall estimate for Method 1, , is a simple average of the estimates produced under :
| (13) |
| (14) |
In our simulations, we chose an equal split of the indices for each partition: . The subsets of each partition are selected at random, and we sample with replacement. We partition a total 10 times (p = 10). For example, if r = 4, then and would be valid subsets. This approach to partitioning is also used in Methods 2 and 4.
5.3. Method 2: cross-validated likelihood
In Method 2, we propose to repeatedly partition the data into subsets and evaluate the estimates based on the ‘training’ subset using the likelihood based on the ‘evaluation’ subset. We partition the data into two subsets p times, calling them for the training subset of the lth partition, and for the evaluation subset of the lth partition. For each , let
| (15) |
that is, is the penalized maximum likelihood estimate evaluated on the training subset. The λ value which maximizes the unpenalized likelihood calculated using the evaluation subset is selected, and the average species richness estimate at is the estimated richness from Method 2:
| (16) |
| (17) |
5.4. Method 3: goodness of fit
Method 3 uses goodness of fit of the fitted frequency counts to select an optimal tuning parameter. Given that Binomial and Multinomial , we have that
| (18) |
Therefore, given estimates and , we consider a plug-in estimate for the expected frequency counts: . The usual goodness of fit statistic is then
| (19) |
where is the largest k such that . Equation (19) is useful in software implementation as we can make use of precomputed tail probabilities .
In Method 3, we make use of this goodness of fit metric by first generating estimates using all replicates. For each , let
| (20) |
is the λ value with the best-fitting estimates of C and η:
| (21) |
and is the estimated value of C at this choice of λ:
| (22) |
An advantage of Method 3 that is not shared by Methods 1, 2 and 4 is that it can be used when r = 1, that is, when no repeated measurements are available.
5.5. Method 4: cross-validated goodness of fit
In Method 4, we return to the partitioning scheme of Method 2, but rather than using the likelihood in the evaluation step, we hypothesize that the goodness of fit metric may be a better choice. We partition the data p times, indexing the partitions by l. For each partition we have a training set and a evaluation set . For all and , we generate estimates exactly as in Method 2:
| (23) |
To select λ we evaluate the goodness of fit metric using only the evaluation subset data, . The λ which produces the best fitting estimates on the evaluation subset is :
| (24) |
and is the mean estimate of C at , averaged all partitions l:
| (25) |
Similar to Method 2, this method generates training set-based estimates , however, it evaluates these estimates using goodness of fit rather than likelihood maximization. Compared to Method 3, this method uses each replicate to either generate an estimate or evaluate the fit, while in Method 3 all replicates are used in both steps.
6. Comparison of methods for selecting λ
We have proposed four tuning methods motivated by properties which would be desirable in an estimator of C. In this section, we compare the performance of each estimator. We simulate zero-truncated gamma-mixed Poisson data using the same parameters used in Section 4. Based on the results of Section 4, we know that estimation can be improved through penalization, at least for the choices of η that we propose to simulate from. The purpose of this section is to determine if any method can reliably select a λ that reduces RMSE compared to unpenalized maximum likelihood estimation. In each simulation below, we select the largest value in by doubling the optimal value of λ found in Table 2.
6.1. Initial comparison of methods' performance
In this simulation, we let C = 1000, and simulate 100 times over all combinations of , . Recall these η are the values chosen based on our motivating examples. We know that the optimal λ choice for these r and η are between 10 and 20, so we chose for this simulation. Methods 0–4 are all evaluated over the same random draws for each parameter choice using the R package simulator [3].
In Table 3, we show the RMSE over all simulations for each method and each combination of η and r. Over all parameter choices, Method 3 performs at least as well as Method 0, with an RMSE which was between 0 and 23% lower. Method 3 is the only method which performs at least as well as Method 0 for all parameter choices.
Table 3. RMSE for Methods 0–4 based on a zero-truncated gamma-mixed Poisson data generating process.
| r = 6 | r = 10 | r = 14 | r = 6 | r = 10 | r = 14 | |
|---|---|---|---|---|---|---|
| Method 0: MLE (no penalization) | 709 | 326 | 339 | 787 | 775 | 716 |
| Method 1: Minimum subset variance | 689 | 630 | 578 | 763 | 821 | 796 |
| Method 2: Cross-validated likelihood | 797 | 521 | 492 | 602 | 658 | 617 |
| Method 3: Goodness of fit | 707 | 326 | 339 | 781 | 663 | 554 |
| Method 4: Cross-validated g.o.f. | 812 | 571 | 533 | 738 | 787 | 679 |
C = 1000 is constant for all simulations. Under each combination, 100 simulations were run. Methods with RMSE better than Method 0 have a grey highlighting, and the best method for each combination is bolded.
We note some differences between the performance of the methods for different η. When , Method 2 also outperforms Method 0 for , while Method 2 does not outperform Method 0 for any r when . Method 4 outperforms Method 0 for when , but never when . We conjecture that the differing performance across η is due to the differing rare species structures implied by the different η's. For example, has more abundant species and larger expected maximum abundance , but relatively few rare species (Table 1).
In Figure 3, we display the C estimates for each method. Recall that as we increase λ, monotonically decreases. Therefore, we can discuss whether a method is on average over-penalizing (selecting which is larger than optimal) or under-penalizing (selecting which is smaller than optimal): if , then the method has under-penalized while if the method has over-penalized.
Figure 3.
Simulation results for all proposed methods when , and when .
We see from Figure 3 that Method 1 over-penalizes for all parameter choices, as all of the estimates are far below the truth. We can understand this result given Figure 1: for very high values of λ the estimates are equal to , and so have low variance. However, the observed richness is severely negatively biased for C. Given its large and consistent negative bias and high RMSE, we do not discuss Method 1 further.
Method 2 has the opposite behavior: the estimates tend to be too high, especially when . This is similarly the case for Method 0, though we see that Method 2 has a slightly lower bias compared to Method 0. Even for , where the Method 0 bias is smaller, we see that Method 2 has poor performance (Table 3). Recall that in each partition of Method 2, we use only half the data to generate the estimates . As a consequence Method 2 has slightly higher variance when compared with Method 0, as evidenced by the interquartile range in Figure 3 for . In conclusion, Method 2 does not appear to be effectively tuning λ, and sample splitting to generate estimates may be leading to high variance.
Methods 3 and 4, which are both based on goodness of fit criteria, display different patterns depending on the η and r values of the simulation. Method 4 outperforms Method 0 when (Table 3), but under the RMSE for Method 4 is 15–75% greater than for Method 0. In comparison, Method 3 is at least as good as Method 0 with respect to RMSE for all parameter combinations tested. In addition, Method 3 is simpler and faster than Method 4. We conclude that Method 3 is the most promising method based on this simulation. We propose to conduct a secondary simulation to further investigate Method 3, testing whether it remains superior to Method 0 over a wider range of η values.
6.2. Performance over wider range of parameter values
In this simulation, we will vary , and perform 100 simulations for three additional choices of η. We let and only consider Methods 0 and 3, based on the results of the previous section. To account for the fact that a larger λ was optimal for these η values (see Table 2) we use . In Table 4, we display the RMSE for both methods and each combination of C, r and η.
Table 4. RMSE for Methods 0 and 3 when counts are drawn from a gamma-mixed Poisson distribution with parameter η.
| Method | C | η | r = 6 | r = 10 | r = 14 | r = 30 | r = 50 |
|---|---|---|---|---|---|---|---|
| Method 0: MLE (no penalization) | 500 | 287 | 244 | 216 | 133 | 73 | |
| Method 3: Goodness of fit | 500 | 312 | 249 | 248 | 139 | 83 | |
| Method 0: MLE (no penalization) | 1000 | 418 | 266 | 277 | 158 | 127 | |
| Method 3: Goodness of fit | 1000 | 427 | 268 | 273 | 187 | 153 | |
| Method 0: MLE (no penalization) | 2000 | 463 | 419 | 372 | 367 | 346 | |
| Method 3: Goodness of fit | 2000 | 513 | 439 | 338 | 413 | 386 | |
| Method 0: MLE (no penalization) | 500 | 266 | 230 | 211 | 208 | 173 | |
| Method 3: Goodness of fit | 500 | 301 | 241 | 229 | 230 | 192 | |
| Method 0: MLE (no penalization) | 1000 | 485 | 430 | 429 | 365 | 372 | |
| Method 3: Goodness of fit | 1000 | 498 | 446 | 445 | 393 | 455 | |
| Method 0: MLE (no penalization) | 2000 | 908 | 775 | 781 | 719 | 787 | |
| Method 3: Goodness of fit | 2000 | 999 | 863 | 833 | 906 | 945 | |
| Method 0: MLE (no penalization) | 500 | 314 | 207 | 305 | 295 | 84 | |
| Method 3: Goodness of fit | 500 | 41 | 89 | 9 | 8 | 7 | |
| Method 0: MLE (no penalization) | 1000 | 701 | 438 | 500 | 375 | 227 | |
| Method 3: Goodness of fit | 1000 | 35 | 218 | 387 | 246 | 16 | |
| Method 0: MLE (no penalization) | 2000 | 1648 | 1143 | 766 | 743 | 998 | |
| Method 3: Goodness of fit | 2000 | 796 | 690 | 542 | 538 | 74 |
Results are based on 100 draws.
We find that for , Method 0 has a lower RMSE for 28 out of 30 combinations of C and r. However, on average over these 30 combinations, the RMSE is only 8% lower. Similarly, when , we also observe that Method 0 has a lower RMSE for 30 out of 30 combinations of C and r, but the RMSE is only 11% lower. We find that in 59% of simulations when , and in 22% of simulations when . We conclude that for these choices of η, maximum likelihood estimates outperform the goodness of fit-based penalization method, but that the advantage of not penalizing is marginal and the chosen value of λ is often zero.
When , Method 3 significantly outperforms Method 0 in 30 out of 30 combinations of C and r. The RMSE is 914% lower for Method 3 on average over these combinations. We also compare these results with those from Table 3, where we found superior performance of Method 3 when . Taken together, these results suggest that Method 3 outperforms Method 0 for small values of δ, but that the methods become comparable as δ increases.
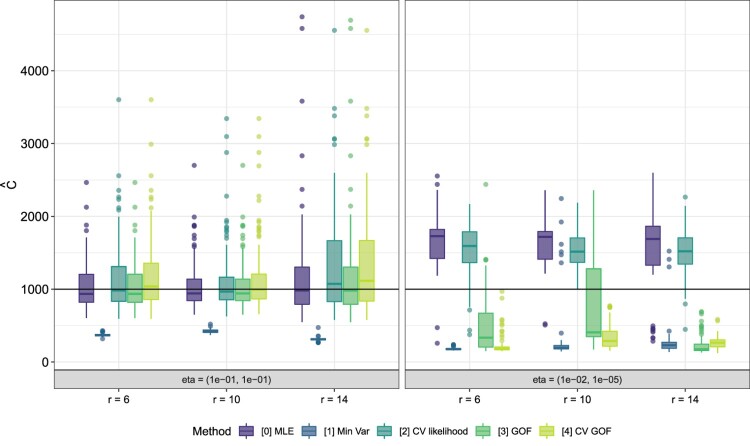
In Figure 4, we display boxplots of the C estimates over r when for and . When we note that is very large for a few simulations, especially for smaller values of r. We recall from Table 1 that this η choice has the highest proportion of abundant species. This is an example of a simulation structure which will cause the instability problem in maximum likelihood estimation discussed in Section 2. Method 3 outperforms Method 0 on this η by selecting lower estimates. In contrast, while Table 4 indicates that Method 3 has larger RMSE than Method 0 when , Figure 4 suggests that the estimates produced by the methods are generally very similar.
Figure 4.
Simulation results for Methods 0 and 3 when , and . The distribution of is shown over 100 draws. The true value of C is indicated with a solid horizontal line.
As a result of these simulations, we conclude that no method (including Method 0) is best for all simulation settings, but Method 3 has advantages in many settings. We found that when and , Method 0 slightly outperformed Method 3 (Table 4). However, when and , Method 3 outperformed Method 0 (Tables 3 and 4). Both methods were about the same when (Table 3). This is consistent with Method 3 having improved performance compared to Method 0 when there are highly abundant species present in the data (see Table 1), in which case Method 0 can be unstable (see Figure 4).
6.3. Performance under model misspecification
We now investigate the performance of Methods 0 and 3 when the model is misspecified. Specifically, we simulate species frequencies under two additional (non-gamma–Poisson) models and compare the performance of the methods.
We first investigate the effect of zero-inflation on species richness estimation by simulating data from the mixture distribution where is the probability mass function of a gamma-mixed Poisson distribution with parameters η. We investigated (where Method 0 outperformed Method 3) and (where Method 3 outperformed Method 0) and fixed C = 1000. We investigate and perform 100 simulations. We find that in 16 out of 18 combinations of p and r, Method 0 outperforms Method 3 when , and that Method 3 outperforms Method 0 in 16 out of 18 combinations of p and r when (Table 5). Unsurprisingly, we find that increased zero-inflation adversely affects both methods. We conclude that neither method is robust to model misspecification via zero-inflation, and that the value of η is more important than the zero-inflation parameter in determining the relative performance of the two methods.
Table 5. RMSE for Methods 0 and 3 when counts are drawn according to a zero-inflated gamma-mixed Poisson distribution with C = 1000.
| Method | η | p | r = 6 | r = 10 | r = 14 | r = 30 | r = 50 |
|---|---|---|---|---|---|---|---|
| Method 0: MLE (no penalization) | 0.1 | 246 | 315 | 146 | 237 | 142 | |
| Method 3: Goodness of fit | 0.1 | 293 | 327 | 163 | 247 | 163 | |
| Method 0: MLE (no penalization) | 0.2 | 288 | 203 | 209 | 194 | 227 | |
| Method 3: Goodness of fit | 0.2 | 331 | 239 | 211 | 209 | 218 | |
| Method 0: MLE (no penalization) | 0.3 | 393 | 375 | 342 | 290 | 298 | |
| Method 3: Goodness of fit | 0.3 | 439 | 397 | 334 | 291 | 312 | |
| Method 0: MLE (no penalization) | 0.1 | 292 | 466 | 224 | 171 | 299 | |
| Method 3: Goodness of fit | 0.1 | 130 | 121 | 145 | 195 | 216 | |
| Method 0: MLE (no penalization) | 0.2 | 566 | 320 | 250 | 286 | 222 | |
| Method 3: Goodness of fit | 0.2 | 244 | 241 | 233 | 204 | 199 | |
| Method 0: MLE (no penalization) | 0.3 | 446 | 422 | 344 | 297 | 299 | |
| Method 3: Goodness of fit | 0.3 | 320 | 307 | 323 | 319 | 291 |
Results are based on 100 draws from the distribution where is a gamma-mixed Poisson distribution with parameters η.
We also investigate the effect of data draws from a different non-gamma-mixed Poisson distribution on the estimation error of Methods 0 and 3. We simulate data according to a shifted logarithmic distribution with probability mass function for and . We investigate , , and . Note that p = 0.9937 is the maximum likelihood estimate of p for the data described in Section 7.2. For each combination of C, r and p, we performed 50 simulations. We found that, without exception, for every single draw. That is, for each of simulations from a logarithmic distribution, Methods 0 and 3 produced identical estimates. Correspondingly, both methods have identical RMSE for all C, r and p. We found that the median was zero for 14 out of 18 combinations of C, r and p, and that for 59% of the 900 simulations. Note that was not always chosen to be zero, but even for a nonzero , the same value of C maximized the regularized likelihood function. We therefore conclude that regularization does not change the estimated species richness when the data generating process is misspecified and drawn according to a logarithmic distribution, and neither method has an advantage over the other in this setting.
7. Data analysis
7.1. Estimating microbial richness in Lake Champlain
To illustrate the performance of our methods on ecological data, we estimate strain-level microbial diversity in Lake Champlain, a large eutrophic lake in Canada. We analyze data from Tromas et al. [26], considering samples from the littoral zone in the summer season of the same year as replicates. This gives us 8 replicates from 2009, 6 replicates from 2010 and 6 replicates from 2011. Given our results from 6, we focus on Methods 0 and 3.
Method 3 produces lower estimates of C than Method 0, as expected. The 2009 and 2010 estimates were approximately 3.5 times lower for Method 3, while the 2011 estimate was approximately 1.5 times lower for Method 3. We see that the estimates of δ are comparable across the two methods, but that the estimates of α differ, and may be higher or lower depending on the dataset (Table 6).
Table 6. Diversity estimates from the Lake Champlain data analysis from 2009 (r = 8), 2010 (r = 6) and 2011 (r = 6) using our proposed methods.
| 2009 | 2010 | 2011 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Method | ||||||||||||
| [0] Unpenalized MLE | 73,404 | — | 0.00088 | 0.00180 | 47,631 | — | 0.00185 | 0.00253 | 57,686 | — | 0.00161 | 0.00140 |
| [3] Goodness of fit | 20,160 | 550 | 0.00323 | 0.00174 | 13,156 | 225 | 0.00685 | 0.00257 | 40,040 | 230 | 0.00231 | 0.00137 |
7.2. Estimating global human host-associated microbial richness
We also applied our method to estimate the species-level diversity of human host-associated microbes. Pasolli et al. [22] assembled c = 4930 species-level genome bins (SGBs) using publicly available shotgun metagenomic data, but we expect that many SGBs were not observed due to undersampling and challenges in genome assembly. The frequency counts of each SGB are available at Pasolli et al. [22, Table S4].
In this dataset r = 1, and so it is not possible to sample split replicate frequency count tables. Therefore, only Methods 0 and Method 3 can be applied. We found that with . In contrast, with and . We therefore find Method 3 to produce an estimate approximately 2.5 times lower than the estimate produced by Method 0. Similar to our analysis of the [26] dataset, we find comparable 's across the two methods but different 's.
8. Discussion
8.1. Conclusions
In this paper, we outlined an extension of the penalized maximum likelihood procedure of Wang and Lindsay [28] for species richness estimation to data with biological replicates, and proposed several methods for tuning the penalization parameter. We demonstrated that penalization can reduce estimation error when analyzing replicate data. We found that tuning the penalization parameter is challenging, but that a tuning method based on goodness of fit (Method 3) has similar or better performance than the unpenalized MLE in many of the settings we analyzed. On two datasets, we found that it reduces the magnitude of species richness estimates. Since species richness estimates can be unstable, we find the reduction in estimates appealing. While we cannot conclude that our proposed goodness of fit tuning parameter selection method provides more reliable estimates than unregularized estimation, the performance of the goodness of fit approach on simulated data is encouraging.
Our investigation highlights the challenges of selecting tuning parameters in the absence of ground truth, since we never observe C. However, even in the absence of information with which to calibrate λ, we showed that with a parametric model for species abundance data we can use goodness of fit in conjunction with maximum likelihood to select λ. We conjecture that the goodness of fit method performs well because it employs a combination of likelihood and goodness of fit metrics calculated on the full dataset to select the tuning parameter, unlike methods that only rely on the likelihood (e.g. Method 2) or split the data (e.g. Method 4).
8.2. Limitations and future work
The approach of Wang and Lindsay [28] is considerably more general than the gamma–Poisson model. The gamma–Poisson model is common for modeling microbiome data [13,15], and for this reason, we focused on it in our investigation. However, our goodness of fit method is also amenable to other parametric models. We leave the investigation of tuning parameter selection under different models to future work.
Our results from Section 3 hint at a possible positive correlation between r and the optimal choice of λ. We investigated whether the optimal choice of λ remained constant for penalties of the form (instead of ; see Equation (7)). We tested and , but found that the trend in optimal λ is not so simple. We leave further investigation into how the optimal λ varies with r to future work. Understanding of would allow us to consider unequally sized evaluation and training partitions for Methods 2 and 4.
For our investigations, we intentionally chose a likelihood optimization algorithm which was stable and exhaustive. We also did not construct standard errors for our estimates, and the long computation times precluded the consideration of a bootstrapping approach. Refining the optimization algorithm would be a valuable extension, and a faster optimization algorithm would facilitate a resample-based variance estimation procedure.
9. Code references
An R package implementing our methods is available at github.com/statdivlab/rre. Code to reproduce our figures and simulations can be found at github.com/statdivlab/rre_sims. We are also grateful to the R Core Team [23] and authors of the packages tidyverse [29], magrittr [1], breakaway [31], foreach [16], Rcpp [8] and data.table [7], which were used for constructing the figures and running the analyses in this paper.
Acknowledgments
The authors of this manuscript are grateful to Jim Hughes for many helpful suggestions that improved content and exposition. We also thank two anonymous referees and the Associate Editor for their very constructive comments regarding the simulation study.
Funding Statement
This work was supported in part by the National Institute of General Medical Sciences (NIGMS) of the NIH under grant number [R35 GM133420].
Disclosure statement
No potential conflict of interest was reported by the author(s).
References
- 1.Bache S.M. and Wickham H., magrittr: A Forward-Pipe Operator for R, R package version 1.5, 2014.
- 2.Barger K. and Bunge J., Objective Bayesian estimation for the number of species, Bayesian Anal. 5 (2010), pp. 765–785. doi: 10.1214/10-BA527 [DOI] [Google Scholar]
- 3.Bien J., The simulator: An engine to streamline simulations, preprint (2016). Available at http://www.arxiv.org/1607.00021.
- 4.Bulmer M.G., On fitting the Poisson lognormal distribution to species-abundance data, Biometrics 30 (1974), pp. 101–110. doi: 10.2307/2529621 [DOI] [Google Scholar]
- 5.Bunge J. and Fitzpatrick M., Estimating the number of species: A review, J. Am. Stat. Assoc. 88 (1993), pp. 364–373. [Google Scholar]
- 6.Chao A. and Bunge J., Estimating the number of species in a stochastic abundance model, Biometrics 58 (2002), pp. 531–539. doi: 10.1111/j.0006-341X.2002.00531.x [DOI] [PubMed] [Google Scholar]
- 7.Dowle M. and Srinivasan A., data.table: Extension of ‘data.frame’, R package version 1.12.2, 2019.
- 8.Eddelbuettel D. and François R., Rcpp: Seamless R and C++ integration, J. Stat. Softw. 40 (2011), pp. 1–18. [Google Scholar]
- 9.Efron B. and Thisted R., Estimating the number of unseen species: How many words did Shakespeare know? Biometrika 63 (1976), pp. 435–447. [Google Scholar]
- 10.Fegatelli D.A. and Tardella L., Moment-based Bayesian Poisson mixtures for inferring unobserved units, preprint (2018). Available at http://www.arxiv.org/1806.06489.
- 11.Fisher R.A., Corbett A.S., and Williams C.B., The relationship between the number of species and the number of individuals in a random sample of an animal population, J. Anim. Ecol. 12 (1943), pp. 42–58. doi: 10.2307/1411 [DOI] [Google Scholar]
- 12.Greenwood M. and Yule G.U., An inquiry into the nature of frequency distributions representative of multiple happenings with particular reference to the occurrence of multiple attacks of disease or of repeated accidents, J. R. Stat. Soc. 83 (1920), pp. 255–279. doi: 10.2307/2341080 [DOI] [Google Scholar]
- 13.S.,Holmes and W.,Huber, Modern Statistics for Modern Biology, Cambridge University Press, Cambridge, 2018. [Google Scholar]
- 14.Lindsay B.G. and Roeder K., A unified treatment of integer parameter models, J. Am. Stat. Assoc. 82 (1987), pp. 758–764. doi: 10.1080/01621459.1987.10478496 [DOI] [Google Scholar]
- 15.Love M.I., Huber W., and Anders S., Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2, Genome Biol. 15 (2014), p. 550. doi: 10.1186/s13059-014-0550-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Microsoft and Weston S., foreach: Provides Foreach Looping Construct, R package version 1.4.7, 2019.
- 17.Minot S.S. and Willis A.D., Clustering co-abundant genes identifies components of the gut microbiome that are reproducibly associated with colorectal cancer and inflammatory bowel disease, Microbiome 7 (2019), p. 110. doi: 10.1186/s40168-019-0722-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Morgan X.C., Tickle T.L., Sokol H., Gevers D., Devaney K.L., Ward D.V., Reyes J.A., Shah S.A., LeLeiko N., Snapper S.B., Bousvaros A., Korzenik J., Sands B.E., R.J., Xavier, and C., Huttenhower, Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment, Genome Biol. 13 (2012), p. R79. doi: 10.1186/gb-2012-13-9-r79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Norris J.L. and Pollock K.H., Non-parametric MLE for Poisson species abundance models allowing for heterogeneity between species, Environ. Ecol. Stat. 5 (1998), pp. 391–402. doi: 10.1023/A:1009659922745 [DOI] [Google Scholar]
- 20.Oakley B.B., Fiedler T.L., Marrazzo J.M., and Fredricks D.N., Diversity of human vaginal bacterial communities and associations with clinically defined bacterial vaginosis, Appl. Environ. Microbiol. 74 (2008), pp. 4898–4909. doi: 10.1128/AEM.02884-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ord J.K. and Whitmore G.A., The Poisson-inverse Gaussian distribution as a model for species abundance, Commun. Stat.-Theory Methods 15 (1986), pp. 853–871. doi: 10.1080/03610928608829156 [DOI] [Google Scholar]
- 22.Pasolli E., Asnicar F., Manara S., Zolfo M., Karcher N., Armanini F., Beghini F., Manghi P., Tett A., Ghensi P., Collado M.C., Rice B.L., DuLong C., Morgan X.C., Golden C.D., Quince C., Huttenhower C., and Segata N., Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle, Cell 176 (2019), pp. 649–662. doi: 10.1016/j.cell.2019.01.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.R Core Team , R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing, Vienna, Austria, 2018.
- 24.Rocchetti I., Bunge J., and Bohning D., Population size estimation based upon ratios of recapture probabilities, Ann. Appl. Stat. 5 (2011), pp. 1512–1533. doi: 10.1214/10-AOAS436 [DOI] [Google Scholar]
- 25.Sanathanan L., Estimating the size of a truncated sample, J. Am. Stat. Assoc. 72 (1977), pp. 669–672. doi: 10.1080/01621459.1977.10480634 [DOI] [Google Scholar]
- 26.Tromas N., Fortin N., Bedrani L., Terrat Y., Cardoso P., Bird D., Greer C.W., and Shapiro B.J., Characterising and predicting cyanobacterial blooms in an 8-year amplicon sequencing time course, ISME J. 11 (2017), pp. 1746. doi: 10.1038/ismej.2017.58 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Walsh F., Smith D.P., Owens S.M., Duffy B., and Frey J.E., Restricted streptomycin use in apple orchards did not adversely alter the soil bacteria communities, Front. Microbiol. 4 (2014), pp. 383. doi: 10.3389/fmicb.2013.00383 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang J.-P.Z. and Lindsay B.G., A penalized nonparametric maximum likelihood approach to species richness estimation, J. Am. Stat. Assoc. 100 (2005), pp. 942–959. doi: 10.1198/016214504000002005 [DOI] [Google Scholar]
- 29.Wickham H., tidyverse: Easily Install and Load the ‘Tidyverse’, R package version 1.2.1, 2017.
- 30.Willis A. and Bunge J., Estimating diversity via frequency ratios, Biometrics 71 (2015), pp. 1042–1049. doi: 10.1111/biom.12332 [DOI] [PubMed] [Google Scholar]
- 31.Willis A., Martin B.D., Trinh P., Barger K., and Bunge J., breakaway: Species Richness Estimation and Modeling, R package version 4.6.10, 2018.