Abstract
R-loops, the byproduct of DNA–RNA hybridization and the displaced single-stranded DNA (ssDNA), have been identified in bacteria, yeasts, and other eukaryotic organisms. The persistent presence of R-loops contributes to defects in DNA replication and repair, gene expression, and genomic integrity. R-loops have not been detected at centromeric (CEN) chromatin in wild-type budding yeast. Here we used an hpr1∆ strain that accumulates R-loops to investigate the consequences of R-loops at CEN chromatin and chromosome segregation. We show that Hpr1 interacts with the CEN-histone H3 variant, Cse4, and prevents the accumulation of R-loops at CEN chromatin for chromosomal stability. DNA–RNA immunoprecipitation (DRIP) analysis showed an accumulation of R-loops at CEN chromatin that was reduced by overexpression of RNH1 in hpr1∆ strains. Increased levels of ssDNA, reduced levels of Cse4 and its assembly factor Scm3, and mislocalization of histone H3 at CEN chromatin were observed in hpr1∆ strains. We determined that accumulation of R-loops at CEN chromatin contributes to defects in kinetochore biorientation and chromosomal instability (CIN) and these phenotypes are suppressed by RNH1 overexpression in hpr1∆ strains. In summary, our studies provide mechanistic insights into how accumulation of R-loops at CEN contributes to defects in kinetochore integrity and CIN.
INTRODUCTION
Faithful chromosome segregation is important for proper growth and development because chromosomal instability (CIN) leads to aneuploidy, which is a hallmark of many cancers and a significant driver of tumorigenesis (Bakhoum and Swanton, 2014; Barra and Fachinetti, 2018). The kinetochore, which includes centromeric (CEN) DNA, evolutionarily conserved protein complexes, and a defined chromatin structure, is essential for faithful chromosome segregation (Diaz-Ingelmo et al., 2015; Bloom and Costanzo, 2017; Lawrimore and Bloom, 2019). The nucleotide composition and size of CEN range from ∼125 base pairs of DNA in budding yeast point centromeres to megabases of complex ordered arrays of DNA repeats, termed as regional centromeres, in other eukaryotes (McKinley and Cheeseman, 2016; Bloom and Costanzo, 2017). Despite the differences in CEN DNA, CEN identity is marked epigenetically by an evolutionarily conserved CEN-specific histone H3 variant (Cse4 in budding yeast, Cid in flies, CENP-A in humans) (Heun et al., 2006; McKinley and Cheeseman, 2016; Mishra and Basrai, 2019; Sharma et al., 2019). The recruitment of Cse4 to the CEN chromatin is mediated by a Cse4-specific assembly factor, Scm3 (HJURP in humans) (Camahort et al., 2007; Mizuguchi et al., 2007; Stoler et al., 2007). The CEN incorporation of Cse4 mediates the assembly of conserved kinetochore protein complexes, resulting in a highly ordered, unique, topologically distinct configuration of CEN chromatin, which is essential for faithful chromosome segregation (Mishra et al., 2013; Diaz-Ingelmo et al., 2015; Bloom and Costanzo, 2017; Lawrimore et al., 2018). Notably, the structural integrity and topological organization of CEN chromatin is an active area of research. For example, the formation of CEN loops, termed as C-loops (Bloom and Costanzo, 2017), is essential for the separation of sister CENs and kinetochore assembly in the absence of the Ctf19, Okp1, Mcm21, Ame1 (COMA) complex (Lawrimore et al., 2018; Lawrimore and Bloom, 2019). The dynamic configuration of CEN C-loops facilitates kinetochore biorientation for faithful chromosome segregation (Yeh et al., 2008; Lawrimore and Bloom, 2019). Moreover, under- and/or overwinding of DNA, referred to as DNA supercoiling, also regulates the topology of CEN chromatin (Furuyama and Henikoff, 2009; Henikoff and Furuyama, 2010). Notably, in vitro studies have shown that CEN DNA is positively supercoiled, and this configuration is mediated by Cse4 and Cid in budding yeast and flies, respectively (Furuyama and Henikoff, 2009; Huang et al., 2011). We have shown that the evolutionarily conserved protein Pat1 (PATL1 in humans) associates with CEN chromatin and regulates Cse4 levels and DNA supercoiling at CEN chromatin in vivo (Lawrimore et al., 2011; Haase et al., 2013; Mishra et al., 2013). The increase in the number of negative supercoils at CEN DNA contributes to errors in chromosome segregation in pat1∆ strains (Mishra et al., 2013, 2015), suggesting that positive supercoiling of CEN DNA is important for chromosome stability.
In addition to C-loops and DNA supercoiling, R-loops are a structural feature of DNA topology that has important roles in gene expression, DNA replication, DNA damage and repair, and genomic integrity (Santos-Pereira and Aguilera, 2015; Costantino and Koshland, 2018; Allison and Wang, 2019). R-loops are transcriptional byproducts, representing a three-stranded nucleic acid structure that contains a DNA–RNA hybrid and a displaced DNA strand (Santos-Pereira and Aguilera, 2015; Allison and Wang, 2019). R-loops are evolutionarily conserved nucleic acid structures and have been observed in bacteria, yeasts, and other eukaryotic organisms (Santos-Pereira and Aguilera, 2015; Allison and Wang, 2019). R-loops are preferentially enriched at highly expressed regions peaking at RNA polymerase II (RNAPII) start and end sites (Rondon and Aguilera, 2019). Accumulation of R-loops has been shown to be deleterious for genomic integrity as they act as physical barriers, which block the progression of replication forks, leading to DNA damage and genomic instability (Gan et al., 2011; Santos-Pereira and Aguilera, 2015; Costantino and Koshland, 2018; Allison and Wang, 2019; Crossley et al., 2019). Moreover, the displaced DNA strand in the R-loop is prone to DNA damage that predisposes cells to accumulate harmful mutations. If left unrepaired, these DNA mutations have been proposed to contribute to human diseases, including cancers (Groh and Gromak, 2014; Wells et al., 2019). R-loops have also been shown to promote RNAPII-mediated transcription of antisense long noncoding RNAs (lncRNAs) in human cells (Rondon and Aguilera, 2019; Tan-Wong et al., 2019) and facilitate efficient mRNA splicing of the second exon of the RPL28 gene in budding yeast (El Hage et al., 2014).
Recent studies have identified RNAPII-mediated CEN transcription in the form of lncRNAs in both sense and antisense orientations in budding yeast (Chen et al., 2019; Ling and Yuen, 2019). CEN lncRNA was detected in a wild-type (WT) strain arrested in S-phase of the cell cycle; increased CEN lncRNA was reported in cbf1∆ and htz1∆ strains when compared with a WT strain (Chen et al., 2019; Ling and Yuen, 2019). However, R-loops have not been detected at CEN chromatin in WT budding yeast, even though they are enriched at ribosomal DNA, retrotransposons, telomeres, and open reading frames (ORFs) (∼1490) that are transcribed primarily by RNAPII (Stirling et al., 2012; Chan et al., 2014; Costantino and Koshland, 2018). A genome-wide study for phosphorylation of histone H3 serine 10 (H3S10P) showed higher levels of H3S10P at ORFs, CENs, peri-CENs, and autonomously replicating sequence (ARS) regions in an hpr1∆ strain (Castellano-Pozo et al., 2013). DNA–RNA immunoprecipitation (DRIP) analysis showed that higher levels of H3S10P correlate with R-loops in budding yeast, Caenorhabditis elegans, and human cells (Castellano-Pozo et al., 2013). These studies confirmed the accumulation of R-loops at one of the CENs, namely CEN6 in cells arrested in G1-phase of the cell cycle in hpr1∆ strains (Castellano-Pozo et al., 2013). Hpr1, a component of the THO/TREX complex, regulates genomic integrity (Aguilera and Klein, 1990; Chavez and Aguilera, 1997; Luna et al., 2019). Increased DNA recombination, defects in mRNA export, and impairment of transcription elongation in the presence of the transcription elongation inhibitor 6-azauracil were observed in hpr1∆ strains (Fan et al., 1996; Chavez and Aguilera, 1997; Chavez et al., 2000; Zenklusen et al., 2002; Huertas et al., 2006). However, physiological consequences due to the accumulation of R-loops at CEN chromatin and its impact on kinetochore function have not been examined.
In this study, we show that accumulation of R-loops at CEN chromatin contributes to defects in the kinetochore integrity and CIN. Our results showed that Hpr1 interacts in vivo with Cse4 and that hpr1∆ causes accumulation of R-loops at CEN chromatin, correlating with increased single-stranded DNA (ssDNA) at CENs and reduced levels of Cse4 and its assembly factor Scm3 with a concomitant increase in histone H3 at CENs. We found that accumulation of R-loops at CEN chromatin contributes to defects in biorientation of sister chromatids during mitosis and increased frequency of chromosome loss in hpr1∆ strains, which were suppressed by RNH1 overexpression. In summary, our results show that persistence of R-loops at CEN chromatin contributes to defects in the kinetochore integrity, resulting in CIN.
RESULTS
Hpr1 associates with centromeric histone H3 variant Cse4
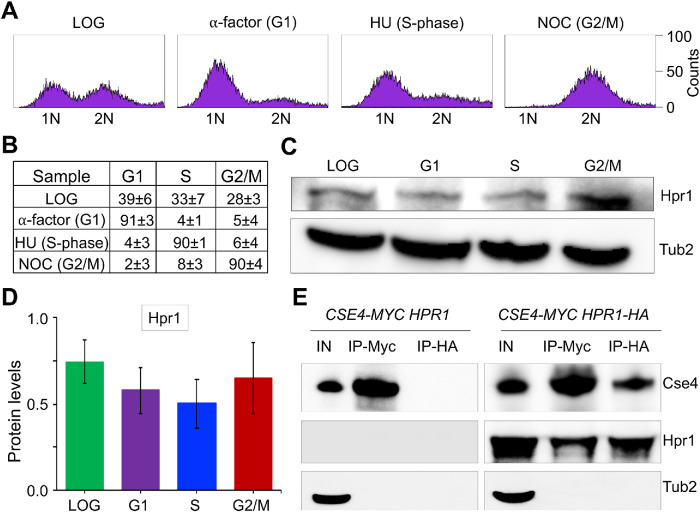
Hpr1 was detected in a high throughput affinity screen for proteins that interact with overexpressed Cse416KR (all lysines mutated to arginine) (Ranjitkar et al., 2010), which is highly stable and mislocalizes to non-CEN regions, leading to CIN (Au et al., 2008). We examined whether Hpr1 interacts with Cse4 in vivo using immunoprecipitation (IP) experiments with a WT strain that expresses HA-tagged Hpr1 and Myc-tagged Cse4 from their native promoters at the endogenous locus. The HA tag does not affect the function of Hpr1 as this strain does not exhibit a temperature-sensitive growth defect observed in a hpr1Δ strain (Supplemental Figure S1). We first examined whether the expression of Hpr1 is cell cycle regulated by Western blot analysis of extracts prepared from an Hpr1-HA strain grown to logarithmic phase, arrested in G1 with α-factor, in S-phase with hydroxyurea (HU), or in G2/M with nocodazole (NOC). Fluorescence activated cell sorting (FACS) examination of nuclear position and cell morphology confirmed the synchronization of cells in these cell cycle stages (Figure 1, A and B). Our results showed that the expression of Hpr1 is not significantly affected by the cell cycle stage (Figure 1, C and D). We next examined whether Hpr1 interacts with Cse4 by performing IP experiments in which protein extracts from logarithmically growing cultures were incubated with agarose beads conjugated with anti-HA (Hpr1) and anti-Myc (Cse4) antibodies. No signals were detected in control experiments performed using a Cse4-Myc strain without Hpr1-HA. We observed an interaction between Hpr1 and Cse4 in a Cse4-Myc strain carrying Hpr1-HA (Figure 1E). On the basis of these results, we conclude that Hpr1 interacts in vivo with Cse4 under normal physiological conditions.
FIGURE 1:
Hpr1 is expressed throughout the cell cycle and interacts with Cse4 in vivo. (A) FACS profile of WT (YMB11085, CSE4-12MYC, HPR1-3HA) cells grown in YPD to logarithmic (LOG) phase at 30°C and synchronized in G1 with α-factor, S-phase with HU and G2/M with nocodazole. (B) Cell cycle categorization of samples from A representing the percentage of cells in G1, S, and G2/M. (C) Western blots showing expression of Hpr1 and Tub2 (loading control). (D) Hpr1 expressed throughout the cell cycle. Protein levels of Hpr1 in LOG-, G1-, S-, and G2/M-phases of the cell cycle. Values were normalized to Tub2. Average from three biological replicates ± SE. (E) Hpr1 interacts in vivo with Cse4. WT strains YMB11085 (CSE4-12MYC, HPR1-3HA) and JG595 (CSE4-12MYC, HPR1) were grown in YPD at 30°C to the logarithmic phase. Cell extracts were prepared, and IPs were performed using anti-HA (A2095; Sigma-Aldrich) and anti-Myc (A7470; Sigma Aldrich) agarose beads. Eluted proteins were analyzed by Western blotting with anti-Myc (Cse4; a-14, sc-789; Santa Cruz Biotechnology), anti-HA (Hpr1; H6908; Sigma Aldrich), and anti-Tub2 (loading control) antibodies. IN = input, and IP = immunoprecipitated samples.
Hpr1 prevents the accumulation of R-loops at CEN chromatin
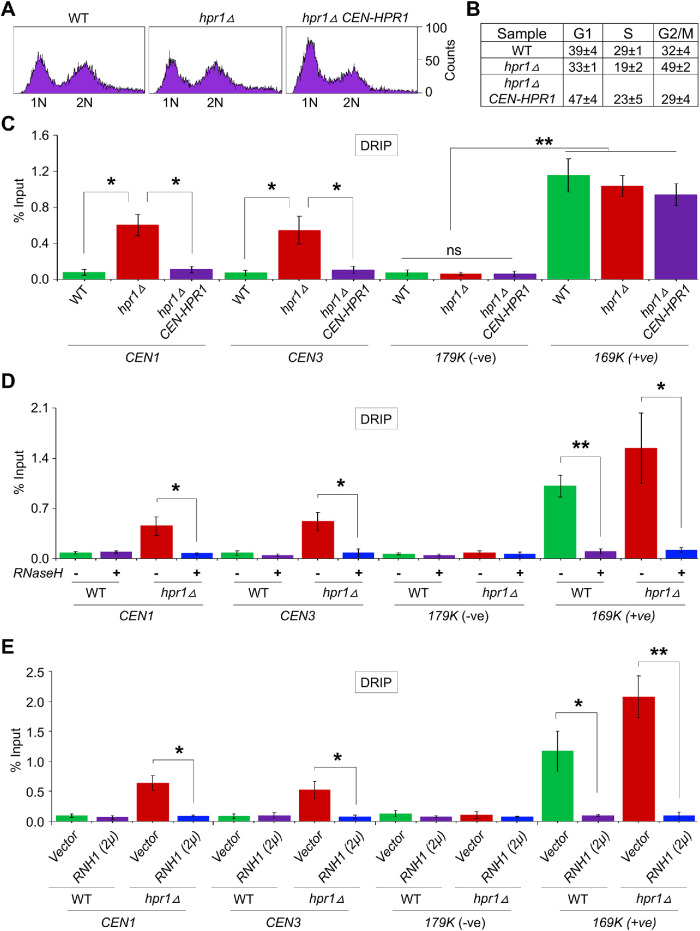
Our results for the interaction of Hpr1 with Cse4 prompted us to examine the role of Hpr1 in preventing the accumulation of R-loops at CENs. We performed three independent experiments to assay the presence of R-loops at CENs in WT and hpr1Δ strains. In the first experiment, we performed DRIP analysis using DNA from logarithmically grown WT and hpr1Δ strains with anti-DNA–RNA hybrid (S9.6) antibodies. FACS, nuclear position, and cell morphology analyses showed similar cell cycle profiles of WT and hpr1Δ strains (Figure 2, A and B). DRIP experiments were performed as described previously (Castellano-Pozo et al., 2013; Costantino and Koshland, 2018; Kabeche et al., 2018) followed by quantitative PCR (qPCR) to determine the levels of R-loops at the CENs (CEN1 and CEN3). R-loop–negative (179K) and –positive (169K) DNA regions identified in previous studies (Costantino and Koshland, 2018) were used as controls. DRIP–qPCR showed a significant accumulation of R-loops at CEN1 and CEN3 in an hpr1Δ strain when compared with that observed in the WT strain (Figure 2C). The accumulation of R-loops at the CEN chromatin is linked to the deletion of HPR1 because the hpr1∆ strain transformed with plasmid-based HPR1 resulted in a significant reduction in R-loops at CEN1 and CEN3, the levels of which were similar to those observed in a WT strain (Figure 2C). No significant accumulation of R-loops was detected at the negative control 179K region, whereas significant accumulation of R-loops was detected at the positive control 169K region in WT and hpr1∆ strains (Figure 2C).
FIGURE 2:
Hpr1 prevents the accumulation of R-loops at the CEN chromatin. (A) FACS profile of logarithmically growing cultures of WT (YMB11246), hpr1Δ (YMB11247), and hpr1Δ CEN-HPR1 (YMB11248) cells used for DRIP experiments. (B) Cell cycle categorization of strains from A representing the percentage of cells in G1, S, and G2/M. (C) Accumulation of R-loops at CEN chromatin in hpr1Δ and its suppression by CEN-HPR1. DRIP analysis of yeast strains from A using the S9.6 (anti–DNA-RNA hybrid) antibody. Levels of R-loops (% input) at CENs (CEN1 and CEN3), a negative control (179K), and a positive control (169K) were determined by DRIP-qPCR. Average from three biological replicates ± SE. **p value < 0.01, *p value < 0.05, ns = statistically not significant, Student’s t test. (D) Accumulation of R-loops at CEN chromatin in hpr1Δ cells with (+) or without (–) treatment with RNase H. DRIP analysis of WT (JG595) and hpr1Δ (YMB11097) using the S9.6 (anti–DNA-RNA hybrid) antibody. Levels of R-loops and statistical significance were determined as described in C. (E) Accumulation of R-loops at CEN chromatin in hpr1Δ cells with or without RNH1 overexpression. DRIP analysis using the S9.6 (anti–DNA-RNA hybrid) antibody was performed for a WT strain with vector (YMB11210) or 2μ-RNH1 (YMB11211) and an hpr1Δ strain with vector (YMB11212) or 2μ-RNH1 (YMB11213). Levels of R-loops and statistical significance as described in C.
In the second experiment, we performed DRIP analysis after treatment of DNA with the ribonuclease RNase-H, which hydrolyzes RNA in DNA–RNA hybrids, leading to elimination of R-loops (Allison and Wang, 2019). No detectable R-loops were observed at CEN1, CEN3, or the negative control region (179K) with or without RNase-H treatment in a WT strain (Figure 2D). However, the accumulation of R-loops at CEN1 and CEN3 was reduced significantly upon treatment with RNase-H in a hpr1∆ strain (Figure 2D). As expected, R-loops at the positive control region (169K) were not detected after treatment with RNase-H in WT and hpr1∆ strains (Figure 2D).
In the third experiment, we assayed R-loops in WT and hpr1∆ strains overexpressing RNH1, which specifically eliminates the RNA strand of an DNA–RNA hybrid, resulting in the degradation of the R-loops (Huertas and Aguilera, 2003; Castellano-Pozo et al., 2013; Garcia-Pichardo et al., 2017; Costantino and Koshland, 2018). Overexpression of RNH1 did not affect the levels of R-loops at CEN1 and CEN3 in a WT strain or at the negative control region (179K) in WT and hpr1∆ strains (Figure 2E). However, overexpression of RNH1 significantly reduced the accumulation of R-loops at CEN1 and CEN3 in an hpr1∆ strain and at the positive control region (169K) in WT and hpr1∆ strains (Figure 2E). Taken together, these results show that Hpr1 prevents the accumulation of R-loops at CENs.
CEN regions are enriched for ssDNA and R-loops during S-phase in an hpr1∆ strain
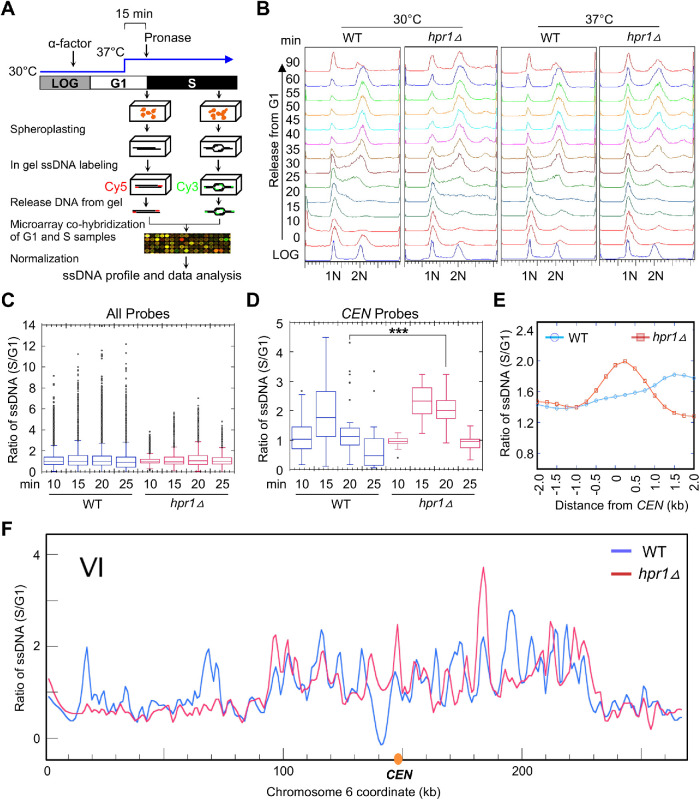
The enrichment of R-loops at CENs in hpr1∆ cells prompted us to examine the consequence of CEN R-loops on cell cycle progression in hpr1∆ cells. Previous studies have shown that hpr1∆ strains are temperature sensitive at 37°C and exhibit synthetic growth defects when combined with mutants of the S-phase checkpoint (Gomez-Gonzalez et al., 2009). WT and hpr1∆ cells were arrested in G1-phase and released at the permissive and restrictive temperatures, 30° and 37°C, respectively (Figure 3A). FACS analysis showed that while both WT and hpr1∆ cells progressed through S-phase similarly at 30°C, hpr1∆ cells showed a 10 to 15 min delay in the appearance of a mid–S-phase population at 37°C and never reached completion before the second S-phase initiated when compared to WT cells (Figure 3B). Next, we applied genome-wide mapping of ssDNA to monitor DNA replication dynamics (Feng et al., 2006). The original design of the method used HU, an inhibitor of the ribonucleotide reductase, to slow down replication fork and enrich for ssDNA formation (Feng et al., 2006). In the current experimental design, we instead used temperature sensitivity (37°C) to exert replication stress in cells synchronously entering S-phase in the absence of HU and sampled them throughout S-phase for ssDNA labeling (Figure 3A). By comparing ssDNA in hpr1∆ cells to WT cells, we asked if any specific regions of the chromosome experience replication fork delay and manifest in increased ssDNA formation after 10, 15, 20, and 25 min release from G1 arrest. We quantified the amount of ssDNA across the genome (all probes, Figure 3C) and within a 20 kb region centering on the CEN (Figure 3D). We did not observe a significant difference in enrichment of ssDNA across the genome between WT and hpr1∆ cells at all time points after release from G1 arrest (Figure 3C). However, specific enrichment of ssDNA at CENs at both 15 and 20 min postrelease from G1 arrest, with statistically significant enrichment at 20 min, was observed in hpr1∆ cells (Figure 3D). Meta-analysis of ssDNA across a 4 kb region centering on all CENs confirmed increased level of ssDNA in hpr1∆ cells at 20 min post–G1 release (Figure 3E). The enrichment levels of ssDNA at 20 min post–G1 release are shown along all of the 16 chromosomes for both WT and hpr1∆ strains (Figure 3F and Supplemental Figure S2). We next took a close-up view of ssDNA levels in a 4-kb window encompassing each of the 16 CENs for 15 and 20 min post–G1 release. Our analysis uncovered 13 CENs that showed higher levels of ssDNA in hpr1∆ than in the WT strain at 15 min post–G1 release (Supplemental Figure S3), and 12 such CENs at 20 min post–G1 release (Supplemental Figure S4). Overall, we found enrichment of ssDNA at 15 CENs (except CEN10) in hpr1∆ compared to the WT strain.
FIGURE 3:
Genome-wide ssDNA mapping shows accumulation of ssDNA at CENs in the hpr1Δ strain at the restrictive temperature (37°C). (A) Schematic presentation of genomic ssDNA mapping. Details of cell culture conditions, ssDNA labeling, and quantification are described in Materials and Methods. Briefly, cells synchronized in G1 by α-factor at 30°C were acclimatized to 37°C for 15 min before the release into S-phase by Pronase treatment. S-phase samples were collected every 5 min starting at 10 min postrelease, as shown in B. Select S-phase samples (10, 15, 20, and 25 min post–G1 release) were subject to ssDNA labeling, and each was cohybridized with a ssDNA labeled G1 control sample onto the microarray. (B) FACS profile of WT (BY4741) and hpr1Δ (BY4741 hpr1Δ) strains. Cells were collected as described in A. The positions of G1 and G2 cells are indicated by “1N” and “2N”, respectively. (C, D) Box plots for quantification of ssDNA from cells sampled at the indicated time points post–release from G1- into S-phase for all probes on the microarray (C) and for probes overlapping only with the CEN (D). Statistical analysis was performed by analysis of variance followed by Tukey’s multiple comparison. ***p value < 0.001. (E) Meta-analysis of ssDNA in a 4 kb window centering on the CEN for cells sampled at 20 min post–release from G1- into S-phase. (F) Example of a chromosomal plot of ssDNA on chromosome VI from cells sampled at 20 min at 37°C. For all chromosome plots, see Supplemental Figure S2.
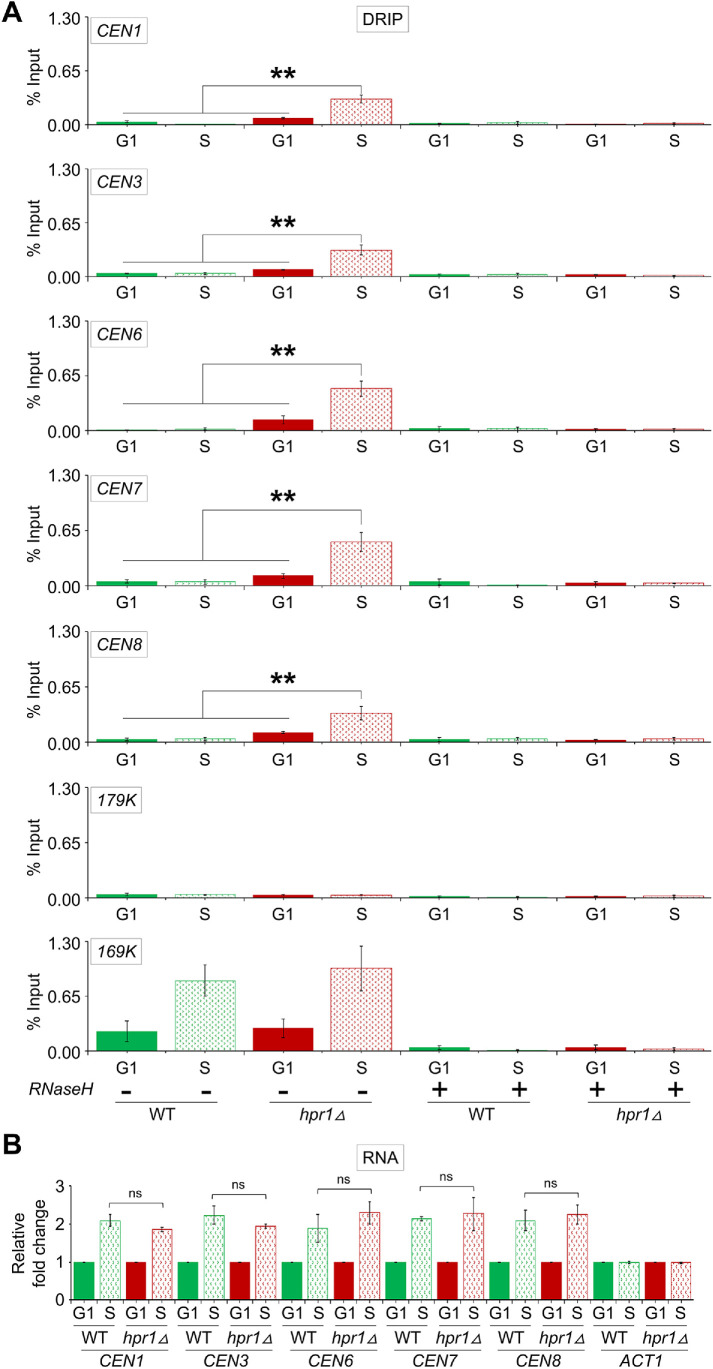
We hypothesized that R-loops at CEN chromatin contribute to the increased levels of CEN-associated ssDNA in hpr1∆ cells. The rationale for this is based on the fact that R-loop structure contains a displaced nontemplate ssDNA strand, opposite of the DNA:RNA hybrid on the template strand, and this ssDNA could serve as a template for labeling in our experiments. Hence, we performed DRIP using growth conditions as described above for ssDNA analysis. Experiments were done using WT and hpr1∆ strains arrested in G1-phase and at 20 min postrelease from G1 arrest at 37°C, referred to as S-phase in Figure 4 (FACS profiles are shown in Supplemental Figure S5). R-loops were not detected in G1- or S-phase cells at the negative control region (179K), but were enriched at the positive control region (169K) at both stages of the cell cycle, albeit higher in the S-phase cells in WT and hpr1∆ strains (Figure 4A). No significant enrichment of R-loops at CEN chromatin (CEN1, CEN3, CEN6, CEN7, and CEN8) was observed in G1- and S-phase in a WT strain. In agreement with a previous study that observed enrichment of R-loops at CEN6 (Castellano-Pozo et al., 2013), we observed higher levels of R-loops at CEN6 and other CENs (CEN1, CEN3, CEN7, and CEN8) in G1 cells of hpr1∆ strain when compared with WT strain. Higher levels of R-loops were observed at CENs (CEN1, CEN3, CEN6, CEN7, and CEN8) in S-phase cells than in G1-phase cells of the hpr1∆ strain (Figure 4A), coincident with the enrichment of ssDNA in this strain at the same time point in the cell cycle (Figure 3, D and E). These results suggest that accumulation of CEN R-loops contributes to the enrichment of ssDNA in the hpr1∆ strain.
FIGURE 4:
Accumulation of R-loops contributes to the enrichment of ssDNA at the CEN chromatin. (A) Accumulation of R-loops at CEN coincides with the enrichment of CEN-derived ssDNA in an hpr1Δ strain. DRIP-qPCR analysis of WT (BY4741) and hpr1Δ (BY4741 hpr1Δ) strains was performed using the S9.6 (anti–DNA-RNA hybrid) antibodies. Average from three biological replicates ± SE. **p value < 0.01, Student’s t test. (B) Expression of CEN lncRNA in S-phase is largely similar between WT and hpr1Δ strains. Total RNA was used in RT-qPCR to determine the expression level of CEN lncRNA (CEN1, CEN3, CEN6, CEN7, and CEN8) and ACT1 lncRNA (control). Average from three biological replicates ± SE. ns = statistically not significant, Student’s t test.
Recent studies have shown CEN lncRNA expression during S-phase of the cell cycle in a WT strain (Chen et al., 2019; Ling and Yuen, 2019). We asked whether increased levels of CEN lncRNA contribute to enrichment of R-loops at CENs. CEN lncRNA levels in G1- and S-phase cells of WT and hpr1∆ cells were examined by reverse transcription (RT)-qPCR. In agreement with previous reports (Chen et al., 2019; Ling and Yuen, 2019), we found enrichment of CEN lncRNA (CEN1, CEN3, CEN6, CEN7, and CEN8) in S-phase cells of a WT strain. However, there was no significant difference in the levels of CEN lncRNA between WT and hpr1∆ strains (Figure 4B). Together, our results revealed that CEN R-loops in hpr1∆ contribute to the enrichment of ssDNA and that the accumulation of CEN R-loops is not solely dependent on the amount of CEN lncRNA transcription.
Accumulation of R-loops at CEN chromatin affects kinetochore integrity and biorientation
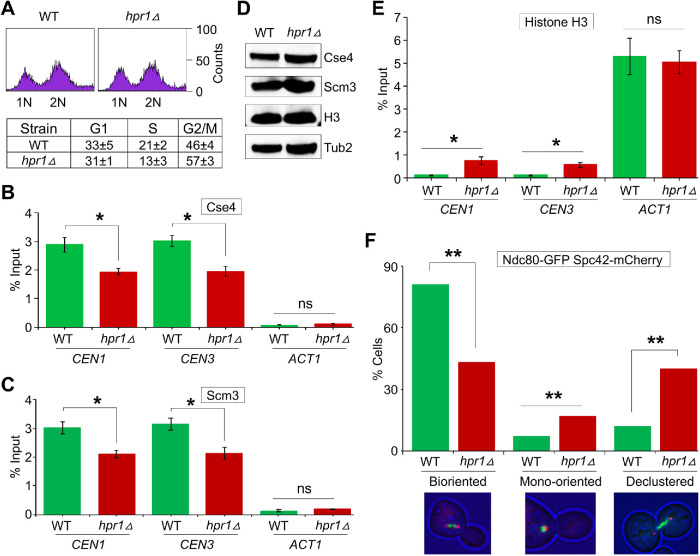
Our results for the interaction of Hpr1 with Cse4 and accumulation of R-loops at CEN in an hpr1∆ strain prompted us to examine the role of Hpr1 in the structural and functional integrity of the kinetochores. Hence, we performed chromatin IP (ChIP) experiments to determine the levels of Cse4 at the CEN chromatin in WT and hpr1∆ strains. FACS, microscopic examination of nuclear position, and cell morphology assays showed similar cell cycle profiles for WT and hpr1∆ strains used for these experiments (Figure 5A). CEN-associated Cse4 in the WT strain was significantly higher (∼2.9% of input at CEN1, 3.0% at CEN3), whereas levels of CEN-associated Cse4 were significantly reduced in the hpr1∆ strain (∼1.9% at CEN1 and CEN3) (Figure 5B). No significant enrichment of Cse4 was detected at a negative control ACT1 locus in either strain (Figure 5B).
FIGURE 5:
Accumulation of R-loops at CEN chromatin affects kinetochore integrity and biorientation. (A) FACS profile and cell cycle categorization of logarithmically growing cultures of WT (JG595) and hpr1Δ (YMB11097) strains. (B) CEN-associated Cse4 is reduced in an hpr1Δ strain. ChIP for Cse4 was performed using anti-Myc agarose beads (A7470; Sigma Aldrich). Cse4 enrichment at CEN1, CEN3, and a negative control (ACT1) was determined by qPCR and is presented as % input. Average from three biological replicates ± SE. *p value < 0.05, ns = statistically not significant, Student’s t test. (C) CEN-associated Scm3 is reduced in an hpr1Δ strain. ChIP for Scm3 was performed using anti-Flag agarose beads (A2220; Sigma Aldrich). Scm3 enrichment at CEN1, CEN3, and a negative control (ACT1) was determined by qPCR and is presented as % input. Average from three biological replicates ± SE. *p value < 0.05, ns = statistically not significant, Student’s t test. (D) Protein levels of Cse4 and Scm3 are not reduced in an hpr1Δ strain. Western blotting was done using the whole cell extracts from strains described in A. (E) Histone H3 associates with CEN chromatin in the hpr1Δ strain. ChIP for histone H3 was done using anti-H3 antibodies (Cat# ab176842; Abcam). Histone H3 enrichment at CEN1, CEN3, and ACT1 was determined by qPCR and is presented as % input. Average from three biological replicates ± SE. *p value < 0.05, ns = statistically not significant, Student’s t test. (F) Deletion of HPR1 causes defects in kinetochore biorientation. WT (KBY6380) and hpr1∆ (KBY6432) strains containing Ndc80-GFP and Spc42-mCherry were grown at room temperature (25°C). The cutoff for metaphase spindle length was 1.7 μm. Spc42-mCherry was used as a spindle pole marker. Representative images showing the position of Ndc80-GFP (green) and Spc42-mCherry (red) are shown. Percentage of cells showing bioriented, mono-oriented (large budded cell with unseparated SPBs), and declustered kinetochores are shown (n = 181 cells). Statistical significance was determined by χ2 test. **p value < 0.01.
The kinetochore protein Scm3 interacts with Cse4, both in vivo and in vitro, and Scm3 is required for association of Cse4 with CEN chromatin (Camahort et al., 2007; Mizuguchi et al., 2007; Stoler et al., 2007). We reasoned that a defect in CEN-association of Scm3 might contribute to the reduced levels of Cse4 at the CEN in hpr1∆ strains. ChIP-qPCR revealed a significant reduction in enrichment of Scm3 at CENs in the hpr1∆ strain (∼30% reduction at CEN1 and CEN3) when compared with the levels observed in a WT strain (Figure 5C). No significant enrichment of Scm3 was observed at a negative control ACT1 locus (Figure 5C). Western blot analysis showed that the reduction in CEN-associated Cse4 and Scm3 observed in the hpr1∆ strain was not due to reduced expression of these proteins (Figure 5D). Previous studies have shown that Hpr1 plays a role in gene transcription and is preferentially required for the transcription of either GC-rich or long transcriptional units (Chavez and Aguilera, 1997; Chavez et al., 2001). Hence, we examined if transcription of CSE4, SCM3, and other genes involved in kinetochore assembly, protein modifications, DNA replication, and chromatin organization (Chavez et al., 2001; Ciftci-Yilmaz et al., 2018; Mishra et al., 2018) was affected in the hpr1∆ strain. In agreement with previous studies (Chavez and Aguilera, 1997; Chavez et al., 2001), we observed significant down-regulation of ADR1 and GAL83 and up-regulation of SLD2 and RRP12 in the hpr1∆ strain. However, no significant differences in transcription of CSE4 or SCM3 were observed between WT and hpr1∆ strains (Supplemental Figure S6). On the basis of these results, we conclude that reduced association of Cse4 and Scm3 to CENs is not due to defects in transcription of CSE4 or SCM3 in hpr1∆ strain.
The reduced Cse4 at CEN prompted us to examine whether histone H3 replaces Cse4 at CEN chromatin in an hpr1∆ strain. Moreover, a previous study reported accumulation of H3S10P at CENs upon deletion of HPR1 (Castellano-Pozo et al., 2013). ChIP-qPCR showed a similar enrichment of histone H3 at the ACT1 locus in WT and hpr1∆ strains (∼5% of input, Figure 5E). The enrichment of histone H3 at CEN chromatin in the hpr1∆ strain was about sixfold higher than in the WT strain (Figure 5E). However, the levels of histone H3 at CEN chromatin in the hpr1∆ strain were about 10- to 12-fold lower than those observed at the ACT1 locus (Figure 5E).
Based on the reduced levels of Cse4 and Scm3 at CEN chromatin, we postulated that accumulation of R-loops at CEN chromatin would contribute to functional defects of the kinetochores in an hpr1∆ strain. Moreover, previous studies have shown that depletion of CEN-associated Scm3 leads to defects in sister kinetochore biorientation (Camahort et al., 2007). Hence, we examined kinetochore biorientation in mitotic cells of WT and hpr1∆ strains by monitoring the localization of Ndc80-GFP (outer kinetochore protein that facilitates kinetochore–microtubule interactions) and Spc42-mCherry (protein that regulates the architecture of the spindle pole) expressed from their native promoters at the endogenous locus. In the WT strain, 81% of the cells had bioriented kinetochores, 7% mono-oriented, and 12% declustered kinetochores whereas, in the hpr1∆ strain, 43% of the cells had bioriented kinetochores, 17% mono-oriented, and 40% declustered kinetochores (Figure 5F). The defect in kinetochore biorientation in the hpr1∆ strain was about threefold higher than in the WT strain (Figure 5F), similar to that reported for mutants defective in kinetochore structure and function, such as Dam1, Mcm21, Stu2, and Spc29 (Cheeseman et al., 2002; Yoder et al., 2005; Ng et al., 2009; Miller et al., 2019). Kinetochore declustering is diagnostic of weakened or altered kinetochores, due to the inability to maintain microtubule attachment during dynamic cycles of microtubule growth or shortening, or defective interactions between adjacent kinetochores that contribute to their spatial clustering. The results showing defects in kinetochore biorientation in the hpr1∆ strain together with the reduced levels of CEN-associated Cse4 and Scm3 suggest that the accumulation of R-loops at CEN chromatin affects the structural and functional integrity of kinetochores.
Accumulation of R-loops at CEN contributes to CIN
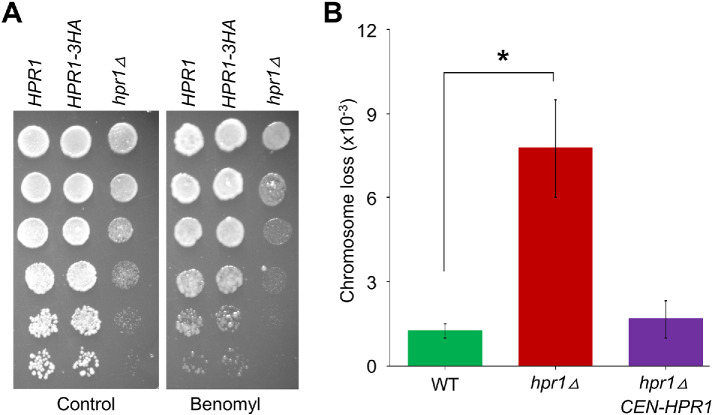
Our results for accumulation of R-loops at CEN, reduced levels of CEN-associated Cse4 and Scm3, and defects in kinetochore biorientation in the hpr1Δ strain led us to examine whether these strains exhibit a CIN phenotype. We first examined the growth phenotype of WT and hpr1Δ strains exposed to benomyl. Benomyl treatment causes depolymerization of mitotic spindles, and benomyl sensitivity is a feature commonly observed in mutants with defects in chromosome segregation (Spencer et al., 1990; Hyland et al., 1999). Our results showed that hpr1Δ strains exhibit increased sensitivity to benomyl when compared to a WT and HPR1-3HA strain (Figure 6A). Notably, a previous study has observed increased frequency of loss for chromosoms V and XV in a homozygous diploid hpr1Δ strain using fluctuation analysis on populations of cells (Santos-Rosa and Aguilera, 1994). To examine the chromosome transmission fidelity, we constructed a haploid hpr1Δ strain carrying a reporter chromosome fragment (CF) and determined the frequency of CF loss using a highly sensitive, visible colony color assay that is based on plating of individual cells (Spencer et al., 1990). The frequency of CF loss in hpr1Δ strains is about sixfold higher than in the WT strain (Figure 6B), which is similar to the CF loss in kinetochore mutants, such as ndc10-1 and mad1∆ strains (Kastenmayer et al., 2005; Ma et al., 2012). The CF loss phenotype of hpr1∆ strain is complemented by a plasmid-based HPR1 expressed from its native promoter (Figure 6B). On the basis of these results, we conclude that accumulation of R-loops at CEN chromatin contributes to CIN.
FIGURE 6:
Accumulation of R-loops at CEN chromatin contributes to CIN. (A) An hpr1Δ strain displays sensitivity to benomyl. Serial (fivefold) dilutions of HPR1 (JG595), HPR1-3HA (YMB11085), and hpr1Δ (YMB11097) strains were plated on YPD plates with or without benomyl (10 μg/ml) and grown for 3 d at 25°C. (B) Errors in chromosome segregation are increased in hpr1Δ strain. Frequency of CF loss in WT (YPH1018), hpr1Δ (YMB11087), and hpr1Δ CEN-HPR1 (YMB11251) strains were measured by a colony color assay as detailed in Materials and Methods. Average from three biological experiments ± SE. *p value < 0.05, Student’s t test.
Overexpression of RNH1 suppresses CIN and kinetochore biorientation defects in an hpr1∆ strain
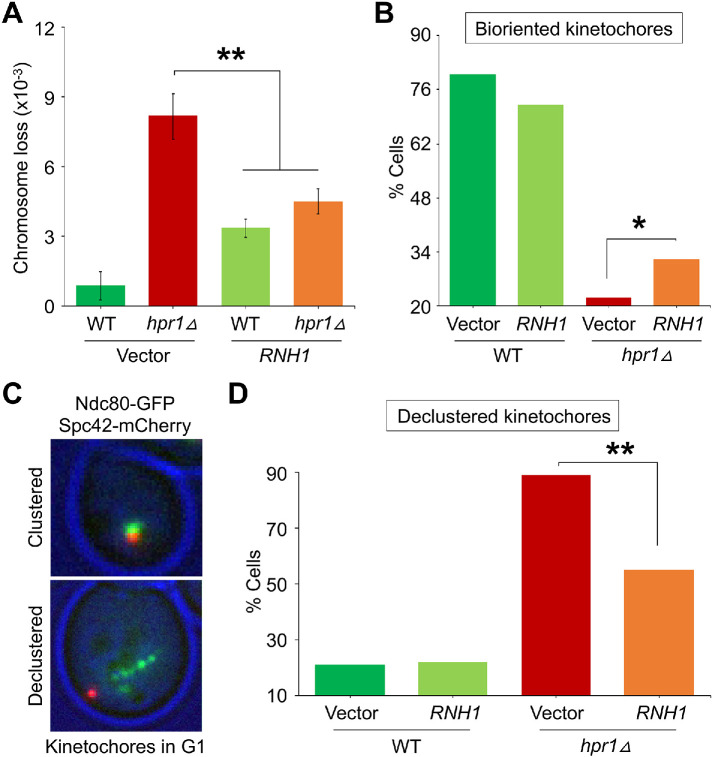
To further confirm that accumulation of R-loops at CEN chromatin contributes to CIN and defects in kinetochore biorientation in hpr1∆ strains, we examined whether these phenotypes would be suppressed upon overexpression of RNH1. We assayed CIN in WT and hpr1Δ strains overexpressing RNH1. As noted previously (Figure 6B), we observed a significantly higher frequency of CF loss in hpr1Δ with vector than in the WT with vector (Figure 7A). The frequency of CF loss in the hpr1∆ strain was significantly suppressed by overexpression of RNH1 when compared to the hpr1∆ strain containing vector alone (Figure 7A). The CF loss phenotype was largely similar between WT and hpr1∆ strains overexpressing RNH1, although overexpression of RNH1 in the WT strain showed slightly higher CF loss than the WT with vector alone (Figure 7A).
FIGURE 7:
Overexpression of RNH1 suppresses CIN, kinetochore biorientation, and kinetochore clustering defects in hpr1∆ strains. (A) RNH1 overexpression suppresses the frequency of chromosome segregation errors in the hpr1Δ strain. Frequencies of CF loss in WT with vector (YMB11206) or 2μ-RNH1 (YMB11207) and hpr1Δ with vector (YMB11208) or 2μ-RNH1 (YMB11209) strains were measured by a colony color assay as detailed in Materials and Methods. Average from three biological experiments ± SE. **p value < 0.01, Student’s t test. (B) RNH1 overexpression suppresses kinetochore biorientation defects in metaphase cells of hpr1∆ strains. WT with vector (YMB11494; n = 135 cells) or 2μ-RNH1 (YMB11495; n = 123 cells) and hpr1∆ with vector (YMB11496; n = 134 cells) or 2μ-RNH1 (YMB11497; n = 216 cells) strains containing Ndc80-GFP and Spc42-mCherry were grown at room temperature (25°C). The cutoff for metaphase spindle length was 1.7 μm. Spc42-mCherry was used as a spindle pole marker. Percentages of cells showing bioriented kinetochores are shown. Statistical significance was determined by χ2 test. *p value < 0.05. (C) Representative images of G1 cells depicting clustered and declustered kinetochores based on the position of Ndc80-GFP (green) and Spc42-mCherry (red). (D) RNH1 overexpression suppresses kinetochore clustering defects of hpr1∆ cells in G1. WT with vector (YMB11494; n = 104 cells) or 2μ-RNH1 (YMB11495; n = 103 cells) and hpr1∆ with vector (YMB11496; n = 100 cells) or 2μ-RNH1 (YMB11497; n = 292 cells) carrying Ndc80-GFP and Spc42-mCherry were grown at room temperature (25°C) to LOG phase, and G1 cells were selected based on the cell morphology (single-celled, no bud). Percentage of cells showing declustered kinetochores in G1 are shown. Statistical significance was determined by χ2 test. **p value < 0.01.
We next examined the effect of RNH1 overexpression on kinetochore biorientation and kinetochore clustering in WT and hpr1∆ strains. The hpr1∆ strain with vector showed significant defects in kinetochore biorientation when compared to the WT strain with vector alone (Figure 7B). Overexpression of RNH1 suppressed kinetochore biorientation defects in the hpr1∆ strain with no significant effect in the WT strain (Figure 7B). Kinetochores in budding yeast form a single cluster close to the spindle pole bodies (SPB) from G1- through late S-phase where all 16 CENs are clustered (Jin et al., 1998, 2000; Joglekar et al., 2008). Hence, we examined the phenotype of kinetochore clustering in G1 cells of the WT or hpr1∆ overexpressing RNH1 (Figure 7C). In the WT strain with vector alone about 20% of cells had declustered or detached kinetochores and this was not affected by overexpression of RNH1 (Figure 7D), whereas, in hpr1∆ with vector alone, 89% of the cells had declustered kinetochores, and this defect was suppressed by overexpressing RNH1 (Figure 7D). Remarkably, there was about 40% suppression in defects in kinetochore clustering in the hpr1∆ strain overexpressing RNH1 (Figure 7D). Taken together, these results show that accumulation of R-loops at CEN contributes to CIN phenotype and RNH1 overexpression suppresses CIN, kinetochore biorientation, and kinetochore clustering defects in hpr1Δ strains.
DISCUSSION
R-loops are not detected at CENs in a WT budding yeast strain (Costantino and Koshland, 2018), and the physiological consequences due to accumulation of R-loops at CEN chromatin have not been investigated. We examined the effect of R-loops in kinetochore function and chromosome segregation using deletion of HPR1, which causes accumulation of R-loops at CEN chromatin (Huertas and Aguilera, 2003). We herein report that accumulation of R-loops at CENs contributes to defects in kinetochore integrity and CIN. Our results show an in vivo interaction of Hpr1 with Cse4 and accumulation of R-loops and ssDNA at CENs in hpr1∆ strains. Accumulation of R-loops at CEN chromatin contributes to reduced levels of Cse4 and Scm3, with a concomitant increase in histone H3 at CENs. Defects in kinetochore biorientation and CIN phenotype in hpr1∆ strains are suppressed by overexpression of RNH1. Our studies provide mechanistic insights into how accumulation of R-loops at CEN contributes to defects in kinetochore integrity and CIN.
Our results from three independent in vivo experiments showed accumulation of R-loops at CEN chromatin in hpr1∆ strains. Treatment with RNase H, overexpression of RNH1, and transformation with a plasmid containing WT HPR1 reduced the accumulation of R-loops at CENs in hpr1∆ strains. Notably, treatment with RNase H and overexpression of RNH1, both of which specifically degrade RNA engaged in R-loops, have been used to validate the presence of R-loops (Huertas and Aguilera, 2003; Castellano-Pozo et al., 2013; Garcia-Pichardo et al., 2017; Costantino and Koshland, 2018; Allison and Wang, 2019). Although transcription is a prerequisite for R-loop formation (Allison and Wang, 2019; Rondon and Aguilera, 2019), our data show that increased transcription of CEN lncRNA alone is not sufficient for accumulation of R-loops at CEN chromatin. For example, WT cells arrested in S-phase show higher levels of CEN lncRNA than cells arrested in G1-phase; however, R-loops were not detected at CEN chromatin in either of these cells. Furthermore, even though the levels of CEN lncRNA are similar in WT and hpr1∆ strains arrested in S-phase, significantly higher enrichment of R-loops at CEN chromatin was observed in hpr1∆ S-phase cells. On the basis of these observations, we propose that small amounts of RNA are sufficient for the seeding and/or initiation of R-loops at CENs, and once established, Hpr1 and the underlying chromatin environment determine the fate and stability of R-loops. We detected significant enrichment of ssDNA at CEN chromatin in hpr1∆ cells. We posit that the CEN-associated ssDNA in hpr1∆ cells is the result of R-loop formation because we have detected R-loops at CEN chromatin in these strains.
We observed reduced levels of CEN-specific histone H3 variant Cse4 and its assembly factor Scm3 with a concomitant increase in histone H3 at CEN chromatin, defects in kinetochore biorientation, and clustering resulting in CIN in hpr1∆ strains. We have previously reported that overexpression of histone H3 contributes to reduced levels of Cse4 at CEN chromatin, increased chromosome loss in WT strains, and synthetic dosage lethality in kinetochore mutants (Au et al., 2008). Our results for increased association of histone H3 at CEN chromatin in hpr1∆ strains are consistent with previous studies showing a link between R-loops and enrichment of H3S10P, a mark for chromatin condensation, at CEN6 in hpr1∆ G1 cells (Castellano-Pozo et al., 2013). Notably, the recruitment of Cse4 to kinetochores occurs during a brief window in early S-phase (∼10 min post–bud emergence) of the cell cycle, and once recruited, it remains stably associated with kinetochores throughout the cell cycle (Pearson et al., 2004). Depletion of Cse4 during S-phase causes phenotypes reminiscent of a defective kinetochore that are manifested in cells undergoing mitosis (Stoler et al., 1995; Collins et al., 2005). Furthermore, defects in kinetochore structure in mutants corresponding to Dam1, Mcm21, Stu2, and Spc29 lead to defects in kinetochore biorientation (Cheeseman et al., 2002; Yoder et al., 2005; Ng et al., 2009; Miller et al., 2019) such as those that we observed in hpr1∆ strains. We propose that the presence of histone H3 and R-loops at CEN chromatin in G1 cells creates a chromatin environment that is not conducive for the Scm3-mediated recruitment of Cse4 in early S-phase, and that the reduced levels of CEN-associated Cse4 and other proteins contribute to mitotic defects in hpr1∆ cells. Our results for the in vivo interaction of Hpr1 with Cse4 and other experimental evidence support this hypothesis. For example, we have shown that hpr1∆ strains exhibit 1) reduced levels of Cse4 and Scm3 and increased levels of histone H3 at CEN chromatin, 2) defects in kinetochore biorientation and kinetochore clustering, and 3) the CIN phenotype.
Overexpression of RNH1 suppressed defects in kinetochore biorientation, kinetochore clustering, and CIN in hpr1∆ strains even though they were not restored to levels observed in the WT strain with vector alone. Furthermore, we observed higher chromosome loss due to overexpression of RNH1 in WT strains. Since RNH1 overexpression does not specifically remove R-loops at CEN chromatin alone, we propose that a threshold of R-loops throughout the genome is needed for faithful chromosome segregation and that accumulation of R-loops at CEN chromatin affects kinetochore function. The defects in kinetochore biorientation and kinetochore clustering observed at a single-cell level provide the strongest evidence for how accumulation of R-loops at CEN chromatin contributes to defects in the integrity of the kinetochore, leading to defects in mitosis in hpr1∆ strains.
Our results from budding yeast have begun to provide mechanistic insights into how accumulation of R-loops at CEN chromatin perturbs kinetochore integrity and contributes to CIN. In maize, R-loops at CENs correlate with lower levels of C-loops and reduced localization of CENH3 (Cse4 homolog) to centromeres (Liu et al., 2020); however, the physiological consequences of R-loops have not been defined. In human cells, R-loops in S-phase interfere with DNA replication, resulting in genomic instability, whereas R-loops in mitosis activate the mitosis-specific ATR checkpoint pathway to prevent errors in chromosome segregation (Kabeche et al., 2018). Notably, ATR associates with human CENs in mitosis mediated by Aurora-A and CENP-F (centromere protein F) (Kabeche et al., 2018). ATR activity at mitotic CENs facilitates the recruitment of replication protein A–coated CEN R-loops, leading to the stimulation of Aurora-B kinase, which in turn prevents the formation of lagging chromosomes (Kabeche et al., 2018). Our results show increased chromosome loss due to accumulation of CEN R-loops in the hpr1∆ strain. The core-CEN in budding yeast is small in size (∼125 base pairs) and occupied by a single Cse4 nucleosome (Furuyama and Biggins, 2007; Bloom and Costanzo, 2017), whereas human CENs are megabases in size and contain multiple CENP-A nucleosomes interspersed with canonical histone H3 nucleosomes (McKinley and Cheeseman, 2016; Andronov et al., 2019). Furthermore, there are multiple kinetochore–microtubule (KT-MT) attachments in higher eukaryotes for each CEN as opposed to a single KT-MT attachment for each CEN in budding yeast (Furuyama and Biggins, 2007; McKinley and Cheeseman, 2016; Bloom and Costanzo, 2017; Andronov et al., 2019). The presence of R-loops at complex CEN chromatin in mitosis may provide an additional surveillance mechanism to monitor multiple KT-MT attachments for faithful chromosome segregation in human cells.
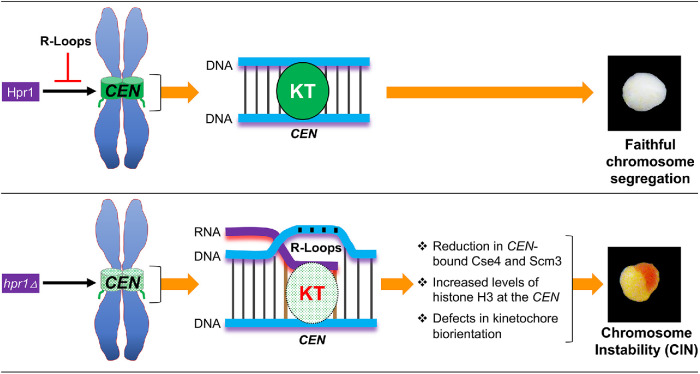
We propose a model in which accumulation of R-loops at CEN chromatin contributes to defects in kinetochore integrity and promotes CIN (Figure 8). Consistent with this model, we observed that RNH1 overexpression, which degrades the RNA moiety of R-loops, suppressed CIN, kinetochore biorientation, and kinetochore clustering defects in hpr1∆ strains. Taken together, we conclude that R-loops and increased levels of histone H3 at the CEN chromatin interfere with the assembly and function of the kinetochore. On the microtubule side of the kinetochore, it has long been appreciated that kinetochores are able to maintain constant interaction with microtubule plus ends that are constantly switching states from growth to shortening. On the DNA side of the kinetochore, we are starting to appreciate the involvement of active transcription. There is an optimal level of RNA transcription required for centromere/kinetochore function, and levels above or below lead to failures in segregation (Hill and Bloom, 1987; Ohkuni and Kitagawa, 2012). The accumulation of R-loops and H3 at CENs may be one of the key mechanisms of CEN dysfunction. This may act as a physical barrier and interfere with the structural configuration and geometric oscillation of kinetochores required for the localization of kinetochore proteins and chromosome stability. Future studies should help us understand how Hpr1 prevents accumulation of R-loops and the implications of R-loops in geometric oscillation of kinetochores during the cell cycle and chromosome segregation. Moreover, R-loops have been implicated in a number of human diseases, including cancers (Groh and Gromak, 2014; Wells et al., 2019). Hence, it will be of interest to examine the physiological consequences of R-loops at CEN chromatin in other systems including human cancers.
FIGURE 8:
Schematic model proposing that accumulation of R-loops at CEN chromatin contributes to CIN. In a WT strain, the presence of Hpr1 prevents the accumulation of R-loops at CEN chromatin leading to faithful chromosome segregation and white colonies due to retention of the reporter CF. In an hpr1Δ strain, R-loops accumulate at the CEN chromatin and this contributes to reduced levels of Cse4 and Scm3, increased levels of histone H3 at CEN chromatin, defects in kinetochore biorientation, benomyl sensitivity, and CIN as depicted by half red/half white colonies due to loss of reporter CF.
MATERIALS AND METHODS
Yeast strains, plasmids, and growth conditions
Budding yeast strains, plasmid constructs, and PCR primers used in this study are listed in Table 1. Strains were grown in yeast peptone dextrose medium (1% yeast extract, 2% Bacto-peptone, 2% glucose; YPD) or in synthetic medium with supplements to allow for the selection of plasmids.
TABLE 1:
Budding yeast strains, plasmids, and primers used in this study.
| Saccharomyces cerevisiae | |||
|---|---|---|---|
| Strain | Genotype | Reference | |
| BY4741 | MATa ura3Δ0 leu2Δ0 his3Δ1 met15Δ0 | Open Biosystems | |
| BY4741 hpr1Δ | MATa ura3Δ0 leu2Δ0 his3Δ1 met15Δ0 hpr1Δ::KAN | Open Biosystems | |
| YPH1018 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 CFIII (CEN3L.YPH278) HIS3 SUP11 | Phil Hieter, University of British Columbia, Vancouver, BC, Canada | |
| JG595 | MATa ura3-1 leu2,3-112 his3-1 trp1-1 ade2-1 can1-100 Δbar1 CSE4-12Myc::URA3 SCM3-3Flag::KANMX | Camahort et al., 2007 | |
| YMB11085 | MATa ura3-1 leu2,3-112 his3-1 trp1-1 ade2-1 can1-100 Δbar1 CSE4-12Myc::URA3 SCM3-3Flag::KANMX HPR1-3HA::HIS3 | This study | |
| YMB11087 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 hpr1∆::TRP1 CFIII (CEN3L.YPH278) HIS3 SUP11 | This study | |
| YMB11097 | MATa ura3-1 leu2,3-112 his3-1 trp1-1 ade2-1 can1-100 Δbar1 CSE4-12Myc::URA3 SCM3-3Flag::KANMX hpr1∆::TRP1 | This study | |
| YMB11210 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 URA3-Vector (pRS426) | This study | |
| YMB11211 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| YMB11212 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 hpr1∆::TRP1 URA3-Vector (pRS426) | This study | |
| YMB11213 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 hpr1∆::TRP1 pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| YMB11246 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 | This study | |
| YMB11247 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 hpr1∆::TRP1 | This study | |
| YMB11248 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3∆200 leu2∆1 hpr1∆::TRP1 CEN-HPR1::URA3 (pMB1979) | This study | |
| YMB11251 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 hpr1∆::TRP1 CFIII (CEN3L.YPH278) HIS3 SUP11 CEN-HPR1::URA3 (pMB1979) | This study | |
| KBY6380 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN | Kerry Bloom, University of North Carolina, Chapell Hill, NC | |
| KBY6432 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN hpr1∆::HIS3 | This study | |
| YMB11206 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 CFIII (CEN3L.YPH278) HIS3 SUP11 URA3-Vector (pRS426) | This study | |
| YMB11207 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 CFIII (CEN3L.YPH278) HIS3 SUP11 pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| YMB11208 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 hpr1∆::TRP1 CFIII (CEN3L.YPH278) HIS3 SUP11 URA3-Vector (pRS426) | This study | |
| YMB11209 | MATα ura3-52 lys2-801 ade2-101 trp1∆63 his3-∆200 leu2∆1 hpr1∆::TRP1 CFIII (CEN3L.YPH278) HIS3 SUP11 pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| YMB11494 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN URA3-Vector (pRS426) | This study | |
| YMB11495 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| YMB11496 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN hpr1∆::HIS3 URA3-Vector (pRS426) | This study | |
| YMB11497 | MATa trp1∆63 ura3-52 his3 lys2 SPC42mCherry::NAT Ndc80-GFP::KAN hpr1∆::HIS3 pGPD-RNH1-HA::URA3 (pBB39) | This study | |
| Plasmids | |||
|---|---|---|---|
| Plasmid | Description | Reference | |
| pBB39 | pGPD-RNH1-HA::URA3, 2-μ | Bak et al., 2013 | |
| pRS426 | URA3 Vector, 2-μ | Christianson et al., 1992 | |
| pMB1979 | CEN-HPR1::URA3 | Dharmacon | |
| Primers | |||
|---|---|---|---|
| Locus | Forward (5′–3′) | Reverse (5′ –3′) | Reference |
| CEN1 | CTCGATTTGCATAAGTGTGCC | GTGCTTAAGAGTTCTGTACCAC | Choy et al., 2011 |
| CEN3 | GATCAGCGCCAAACAATATGG | AACTTCCACCAGTAAACGTTTC | Paul Megee, Oakland University, Rochester, MI |
| CEN6 | ACCTTGAAGACTATATTTCTTTT CATCACGTG | GGTCGTCCAATATCATCGTAAACGTG | John Choy, National Cancer Institute, Bethesda, MD (Basrai lab) |
| CEN7 | AAGAGCTTCGATAAATTTGAAAATTAATTGTACTAC | CCTTGCATTATAATTATCCAA TACTTTGTCGTC | John Choy, National Cancer Institute, Bethesda, MD (Basrai lab) |
| CEN8 | CCTTGAATGTACAGCTCTAATTACACAC | GATAATGTCTTAACCAATTTT CTAAGTTCGGAAC | John Choy, National Cancer Institute, Bethesda, MD (Basrai lab) |
| ACT1 | ACAACGAATTGAGAGTTGCCCCAG | AATGGCGTGAGGTAGAGAGAAACC | Crotti and Basrai, 2004 |
| 169K | CATTATCAATCCTTGCGTTTCA | GCTCGAGTAATACCGGAGTGTC | Costantino and Koshland, 2018 |
| 179K | AAAGCTTTGTTTGCGGTATGTT | CCCGGATTACAAAGTCACTACC | Costantino and Koshland, 2018 |
| 310K | TCTCGGAATTTATCATGACCCAT | AAACCCTGCACACATTTCGT | Laloraya et al., 2000 |
| ADR1 | AAGCCAGATAGCGGCAACT | CACTCACAGCTGGCATTAACA | Ciftci-Yilmaz et al., 2018 |
| GAL83 | TTCATCGTCATCATCGTCGT | AAGGTCGTTTCCTGCACAAT | Ciftci-Yilmaz et al., 2018 |
| CSE4 | CCATTTGCAAGGCTAGTGAAAG | CCAATAATCCTACCAGATACGC | Sultan Ciftci-Yilmaz, Lars Boeckmann, National Cancer Institute, Bethesda, MD (Basrai lab) |
| SCM3 | ATGAAAACCAATAAGAAAATTTCTAAAAG | TCCTTTTTGGTCTTCCGTTTTTCG | This study |
| TUB2 | ATTGACGGCAATTGGCTCT | ACCAGTGCAAGAAAGCTTTTC | Carole Carter, National Cancer Institute, Bethesda, MD (Basrai lab) |
| SLD2 | AGCTGAAAACATGGGAGCAT | TCGCCTTACCGTAAACTTGG | Ciftci-Yilmaz et al., 2018 |
| RRP12 | CATTTCTTCTGGATTGGCTGA | ACATTGAGAGGCAGCTTGGT | Ciftci-Yilmaz et al., 2018 |
IP and Western blotting
Extracts were prepared from yeast cells grown to logarithmic phase at 30°C in YPD medium. IP experiments were done with anti-HA (A2095; Sigma Aldrich) and anti-Myc (A7470; Sigma Aldrich) agarose antibodies following the procedure as described (Mishra et al., 2011, 2018). Total proteins were extracted with the trichloroacetic acid (TCA) procedure (Kastenmayer et al., 2005) and quantified using the Bio-Rad DC protein quantitation assay (Bio-Rad Laboratories, Hercules, CA). Protein samples for Western blotting were size separated on SDS–polyacrylamide gels and transferred to a 0.45-µm nitrocellulose membrane. Primary antibodies used were anti-Myc (a-14, sc-789; Santa Cruz Biotechnology), anti-HA (H6908; Sigma Aldrich), and anti-Tub2 (Au et al., 2008; Mishra et al., 2016). Secondary antibodies were horseradish peroxidase (HRP)-conjugated sheep anti-mouse immunoglobulin G (IgG) (NA931V) and HRP-conjugated sheep anti-rabbit IgG (NA934V), both obtained from Amersham Biosciences.
ChIP and qPCR
ChIP experiments were performed with three independent biological replicates using the approach as described previously (Mishra et al., 2007, 2011). Antibodies used to capture DNA-protein complexes were anti-HA agarose (A2095; Sigma Aldrich), anti-Myc agarose (A7470; Sigma Aldrich), anti-Flag agarose (A2220; Sigma Aldrich), and anti-histone H3 (ab176842; Abcam). ChIP-qPCR was done using Fast SYBR Green Master Mix in 7500 Fast Real Time PCR System (Applied Biosystems, Foster City, CA) following conditions described previously (Mishra et al., 2015). The enrichment as presented as percent input was determined using the ΔΔCT method (Livak and Schmittgen, 2001).
Flow cytometry and cell morphology analysis
For FACS analysis, cells were fixed in fixation buffer (70% ethanol, 0.2 M Tris, pH 7.4), washed twice in 0.2 M Tris-buffer (pH 7.4), and then treated with RNase A (0.2 M Tris-buffer [pH 7.4]; 0.1 mg/ml RNase A) at 37°C for 3 h followed by proteinase K treatment (final concentration of 0.2 mg/ml) at 50°C for 1 h. Cells were washed twice in 0.2 M Tris-buffer (pH 7.4) and stained with propidium iodide solution (0.2 M Tris-buffer [pH 7.4], 15 µg/ml propidium iodide) overnight at 4°C. Sample were processed using a FACS Calibur flow cytometer and analyzed using Cell Quest Pro software (BD Biosciences). Cells were examined under the microscope (Axioskop 2; Zeiss), and cell cycle stages were determined based on nuclear position and cell morphology as described previously (Mishra et al., 2011).
DRIP-qPCR experiments
DRIP experiments were performed as described previously (Castellano-Pozo et al., 2013; Costantino and Koshland, 2018; Kabeche et al., 2018) with some minor modifications. Briefly, cells were dissolved in spheroplasting buffer (20 mM HEPES, pH 7.4, 1.2 M sorbitol, 0.5 mM phenylmethylsulfonyl fluoride [PMSF]), containing Zymolyase 100T (0.04 mg/ml; MP Biomedicals) and β-mercaptoethanol (5 μl/ml; Sigma Aldrich), and incubated at 30°C for spheroplast preparation. Spheroplasting was monitored by OD800 measurements in 1% SDS, and reactions were stopped when >90% spheroplasting was achieved by washing in postspheroplasting buffer (20 mM PIPES, pH 6.8, 1.2 M sorbitol, 1 mM MgCl2, 1 mM PMSF). DNA extractions were done using the QIAamp DNA Blood Mini Kit (Cat# 51104; Qiagen). Spheroplasts were resuspended in lysis buffer AL and sonicated to obtain an average fragment size of ∼500 base pairs. DNA was extracted from sonicated spheroplasts using Qiagen columns following manufacturer’s recommendations (Cat# 51104; Qiagen), and eluted in nuclease-free AE buffer. DNA was treated with RNase H (Cat# M0297S; New England Biolabs) before DRIP experiments as described previously (El Hage et al., 2010). Control (untreated) and RNase H–treated DNA was dissolved in 1× FA buffer (50 mM HEPES-Na, pH 7.6, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate, and 1× protease inhibitor cocktail [Sigma]), and DRIP was performed by adding 5 μg of S9.6 antibodies (Kera Fast) conjugated to magnetic Protein-A Dynabeads (Invitrogen) and incubating overnight at 4°C. Beads were then washed at room temperature for 5 min with 1× FA buffer (three times), 1× FA with 500 mM NaCl (once), RIPA (250 mM LiCl, 0.5% NP-40, 0.5% sodium deoxycholate, 1 mM EDTA, and 10 mM Tris-HCl) buffer (once), and finally twice with 1× TE buffer. DNA was then eluted from the beads with 100 μl of elution buffer (0.5% SDS, 10 mM EDTA, 25 mM Tris-HCl, pH 7.6), treated with proteinase K, and purified using MinElute columns (Cat# 28004; Qiagen). The percentage of the hybrid signal was quantified using qPCR on DNA from IP and total input with the Fast SYBR-Green Master Mix (Applied Biosystems) using PCR primers from CENs (CEN1, CEN3, CEN6, CEN7, and CEN8) and control regions: R-loop–positive region (169K), and R-loop–negative region (179K), for which primers were designed based on a previous study (Costantino and Koshland, 2018). The amplification conditions were initial denaturation at 95°C for 20 s followed by cycling of 95°C for 3 s and 60°C for 30 s (data acquisition step) in a 7500 Fast-Real Time PCR System (Applied Biosystems). Three independent biological replicates were performed for each sample, and data are presented as percent input calculated using the ΔΔCT method (Livak and Schmittgen, 2001).
Genome-wide microarray-based analysis of ssDNA
Cells grown exponentially at 30°C were synchronized in G1 by addition of 3 µM α-factor. Fifteen minutes before the release from G1- into S-phase by addition of 0.3 mg/ml Pronase, half of the cells were shifted to 37°C and used for ssDNA mapping. As control (for FACS analysis), the other half of the culture was maintained at 30°C and released into S-phase at 30°C. Following the release from G1- into S-phase, cells were sampled at 10, 15, 20, and 25 min and subjected to ssDNA labeling. A G1 control sample was collected before Pronase addition and also labeled for ssDNA. Each S-phase sample was paired with the G1 control, differentially labeled with Cy3- and Cy5-dUTP, respectively, and cohybridized onto the same microarray slide (Agilent 4 × 44 k ChIP-to-chip microarray). ssDNA labeling and quantification were performed as previously described (Feng et al., 2006). The genome-wide data are available from the National Center for Biotechnology Information (NCBI) GEO repository under the accession reference number GSE151849.
RNA extraction and RT-qPCR
Total RNA was extracted from the yeast cells using the Qiagen RNeasy kit and subjected to rigorous DNase treatment according to the manufacturer’s instruction (Cat# 74106; Qiagen). DNase-treated RNA was reverse transcribed using the Access RT-PCR System (Promega Corporation, Madison, WI) in a 10-μl reaction with the primers listed in Table 1. Expression levels of CENs (CEN1, CEN3, CEN6, CEN7, and CEN8) and genes (ACT1, ADR1, CSE4, GAL83, RRP12, SCM3, SLD2, and TUB2) were determined by real-time qPCR using 7500 Fast SYBR Green Master Mix (Applied Biosystems). RNA samples processed through the RT step without reverse transcriptase enzyme were used as negative controls. The CT values, which denote the number of PCR cycles needed to cross the threshold fluorescence in the amplification curve (t½ of the exponential region of amplification), were determined for each sample, and the corresponding LOG transcription was deduced from a standard curve (Supplemental Figure S7) prepared under identical conditions based on quantitation of a 323 base pair product with increasing amounts (102, 103, 104, 105, and 106 copies) of control mRNA (Promega Corporation). For CEN lncRNA experiments, the relative fold changes were calculated after normalization to the G1 cells, where G1 was set to a value of 1. ACT1, which is expressed constitutively with transcription levels similar in G1- and S-phase, was used as a positive control. Three independent biological replicates for each sample were performed, and the statistical significance of the data was determined as described previously (Ling and Yuen, 2019).
Microscopy and kinetochore biorientation assay
Yeast cells containing the SPB protein Spc42-mCherry and the outer kinetochore protein Ndc80-GFP were imaged at 25°C on a Nikon Eclipse Ti wide-field inverted microscope with a 100×Apo total internal reflection fluorescence 1.49 numerical aperture objective (Nikon, Melville, NY) and Andor Clara charge-coupled device camera (Andor, South Windsor, CT). Images were acquired using Nikon Elements imaging software. Images were taken in transilluminated light and GFP and RFP fluorescence illumination. Metaphase spindles are in the length range of 1.4–1.7 µm.
Dilution plating assays
WT and hpr1Δ strains were grown in YPD medium at 25°C, and cell suspensions were prepared in water to obtain an OD600 = 2.0. Then 5 μl of the cell suspension was plated on YPD plates in serial fivefoldl dilutions. Cells were incubated on plates at 25° and 37°C to examine the temperature sensitivity phenotype. Benomyl sensitivity was determined at 25°C on YPD plates, with or without 10 μg/ml benomyl (Cat# 17804-35-2; Aldrich Chemicals).
Chromosome transmission fidelity assay
The frequency of chromosome segregation was measured by a colony color assay in which the loss of a reporter CF results in red-colored sectors in an otherwise white colony (Spencer et al., 1990). WT and hpr1Δ strains containing the CF were grown to early logarithmic phase in medium selective for the CF and the plasmids being used. Cells were then plated on synthetic dextrose agar medium with limiting adenine at 25°C. The frequency of CF loss represents the number of colonies that were at least half red (indicative of CF loss in the first cell division) over the total number of colonies counted. About 1000 colonies from three independent transformants were examined for each strain.
Supplementary Material
Acknowledgments
We appreciate technical support for ssDNA mapping from Jie Peng. We thank Andres Aguilera, Sue Biggins, Jennifer Gerton, and Vincent Guacci for strains and plasmids; Kathy McKinnon of the National Cancer Institute Vaccine Branch FACS Core for technical assistance; and the members of the Basrai laboratory for helpful discussions. P.K.M. and M.A.B. were supported by the Intramural Research Program of the National Cancer Institute, National Institutes of Health (NIH). E. Y. and K. B. were supported by NIH grant R37 GM32238; W. F. and A. C. were supported by NIH grant GM118799-01A1.
Abbreviations used:
- C-loop
centromere loop
- CEN
centromere
- CF
chromosome fragment
- ChIP
chromatin immunoprecipitation
- CIN
chromosomal instability
- DRIP
DNA-RNA immunoprecipitation
- FACS
fluorescence-activated cell sorting
- GFP
green fluorescent protein
- IP
immunoprecipitation
- KT-MT
kinetochore-microtubule
- lncRNA
long non-coding RNA
- ORF
open reading frame
- qPCR
quantitative PCR
- RNAPII
RNA polymerase II
- SPB
spindle pole body
- ssDNA
single-strand DNA.
Footnotes
This article was published online ahead of print in MBoC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E20-06-0379) on November 4, 2020.
REFERENCES
- Aguilera A, Klein HL (1990). HPR1, a novel yeast gene that prevents intrachromosomal excision recombination, shows carboxy-terminal homology to the Saccharomyces cerevisiae TOP1 gene. Mol Cell Biol 10, 1439–1451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allison DF, Wang GG (2019). R-loops: formation, function, and relevance to cell stress. Cell Stress 3, 38–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andronov L, Ouararhni K, Stoll I, Klaholz BP, Hamiche A (2019). CENP-A nucleosome clusters form rosette-like structures around HJURP during G1. Nat Commun 10, 4436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Au WC, Crisp MJ, DeLuca SZ, Rando OJ, Basrai MA (2008). Altered dosage and mislocalization of histone H3 and Cse4p lead to chromosome loss in Saccharomyces cerevisiae. Genetics 179, 263–275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bakhoum SF, Swanton C (2014). Chromosomal instability, aneuploidy, and cancer. Front Oncol 4, 161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balk B, Maicher A, Dees M, Klermund J, Luke-Glaser S, Bender K, Luke B (2013). Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat Struct Mol Biol 20, 1199–1205. [DOI] [PubMed] [Google Scholar]
- Barra V, Fachinetti D (2018). The dark side of centromeres: types, causes and consequences of structural abnormalities implicating centromeric DNA. Nat Commun 9, 4340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bloom K, Costanzo V (2017). Centromere structure and function. Prog Mol Subcell Biol 56, 515–539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camahort R, Li B, Florens L, Swanson SK, Washburn MP, Gerton JL (2007). Scm3 is essential to recruit the histone h3 variant cse4 to centromeres and to maintain a functional kinetochore. Mol Cell 26, 853–865. [DOI] [PubMed] [Google Scholar]
- Castellano-Pozo M, Santos-Pereira JM, Rondon AG, Barroso S, Andujar E, Perez-Alegre M, Garcia-Muse T, Aguilera A (2013). R loops are linked to histone H3 S10 phosphorylation and chromatin condensation. Mol Cell 52, 583–590. [DOI] [PubMed] [Google Scholar]
- Chan YA, Aristizabal MJ, Lu PY, Luo Z, Hamza A, Kobor MS, Stirling PC, Hieter P (2014). Genome-wide profiling of yeast DNA:RNA hybrid prone sites with DRIP-chip. PLoS Genet 10, e1004288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chavez S, Aguilera A (1997). The yeast HPR1 gene has a functional role in transcriptional elongation that uncovers a novel source of genome instability. Genes Dev 11, 3459–3470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chavez S, Beilharz T, Rondon AG, Erdjument-Bromage H, Tempst P, Svejstrup JQ, Lithgow T, Aguilera A (2000). A protein complex containing Tho2, Hpr1, Mft1 and a novel protein, Thp2, connects transcription elongation with mitotic recombination in Saccharomyces cerevisiae. EMBO J 19, 5824–5834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chavez S, Garcia-Rubio M, Prado F, Aguilera A (2001). Hpr1 is preferentially required for transcription of either long or G+C-rich DNA sequences in Saccharomyces cerevisiae. Mol Cell Biol 21, 7054–7064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheeseman IM, Anderson S, Jwa M, Green EM, Kang J, Yates JR 3rd, Chan CS, Drubin DG, Barnes G (2002). Phospho-regulation of kinetochore-microtubule attachments by the Aurora kinase Ipl1p. Cell 111, 163–172. [DOI] [PubMed] [Google Scholar]
- Chen CF, Pohl TJ, Chan A, Slocum JS, Zakian VA (2019). Saccharomyces cerevisiae centromere RNA is negatively regulated by Cbf1 and its unscheduled synthesis impacts CenH3 binding. Genetics 213, 465–479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choy JS, Acuna R, Au WC, Basrai MA (2011). A role for histone H4K16 hypoacetylation in Saccharomyces cerevisiae kinetochore function. Genetics 189, 11–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christianson TW, Sikorski RS, Dante M, Shero JH, Hieter P (1992). Multifunctional yeast high-copy-number shuttle vectors. Gene 110, 119–122. [DOI] [PubMed] [Google Scholar]
- Ciftci-Yilmaz S, Au WC, Mishra PK, Eisenstatt JR, Chang J, Dawson AR, Zhu I, Rahman M, Bilke S, Costanzo M, et al. (2018). A genome-wide screen reveals a role for the HIR histone chaperone complex in preventing mislocalization of budding yeast CENP-A. Genetics 210, 203–218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins KA, Castillo AR, Tatsutani SY, Biggins S (2005). De novo kinetochore assembly requires the centromeric histone H3 variant. Mol Biol Cell 16, 5649–5660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Costantino L, Koshland D (2018). Genome-wide map of R-loop-induced damage reveals how a subset of R-loops contributes to genomic instability. Mol Cell 71, 487–497.e483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crossley MP, Bocek M, Cimprich KA (2019). R-loops as cellular regulators and genomic threats. Mol Cell 73, 398–411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crotti LB, Basrai MA (2004). Functional roles for evolutionarily conserved Spt4p at centromeres and heterochromatin in Saccharomyces cerevisiae. EMBO J 23, 1804–1814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz-Ingelmo O, Martinez-Garcia B, Segura J, Valdes A, Roca J (2015). DNA topology and global architecture of point centromeres. Cell Rep 13, 667–677. [DOI] [PubMed] [Google Scholar]
- El Hage A, French SL, Beyer AL, Tollervey D (2010). Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev 24, 1546–1558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El Hage A, Webb S, Kerr A, Tollervey D (2014). Genome-wide distribution of RNA-DNA hybrids identifies RNase H targets in tRNA genes, retrotransposons and mitochondria. PLoS Genet 10, e1004716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan HY, Cheng KK, Klein HL (1996). Mutations in the RNA polymerase II transcription machinery suppress the hyperrecombination mutant hpr1 delta of Saccharomyces cerevisiae. Genetics 142, 749–759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng W, Collingwood D, Boeck ME, Fox LA, Alvino GM, Fangman WL, Raghuraman MK, Brewer BJ (2006). Genomic mapping of single-stranded DNA in hydroxyurea-challenged yeasts identifies origins of replication. Nat Cell Biol 8, 148–155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furuyama S, Biggins S (2007). Centromere identity is specified by a single centromeric nucleosome in budding yeast. Proc Natl Acad Sci USA 104, 14706–14711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furuyama T, Henikoff S (2009). Centromeric nucleosomes induce positive DNA supercoils. Cell 138, 104–113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gan W, Guan Z, Liu J, Gui T, Shen K, Manley JL, Li X (2011). R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes Dev 25, 2041–2056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia-Pichardo D, Canas JC, Garcia-Rubio ML, Gomez-Gonzalez B, Rondon AG, Aguilera A (2017). Histone mutants separate R loop formation from genome instability induction. Mol Cell 66, 597–609.e595. [DOI] [PubMed] [Google Scholar]
- Gomez-Gonzalez B, Felipe-Abrio I, Aguilera A (2009). The S-phase checkpoint is required to respond to R-loops accumulated in THO mutants. Mol Cell Biol 29, 5203–5213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groh M, Gromak N (2014). Out of balance: R-loops in human disease. PLoS Genet 10, e1004630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haase J, Mishra PK, Stephens A, Haggerty R, Quammen C, Taylor RM 2nd, Yeh E, Basrai MA, Bloom K (2013). A 3D map of the yeast kinetochore reveals the presence of core and accessory centromere-specific histone. Curr Biol 23, 1939–1944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S, Furuyama T (2010). Epigenetic inheritance of centromeres. Cold Spring Harb Symp Quant Biol 75, 51–60. [DOI] [PubMed] [Google Scholar]
- Heun P, Erhardt S, Blower MD, Weiss S, Skora AD, Karpen GH (2006). Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev Cell 10, 303–315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hill A, Bloom K (1987). Genetic manipulation of centromere function. Mol Cell Biol 7, 2397–2405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang CC, Chang KM, Cui H, Jayaram M (2011). Histone H3-variant Cse4-induced positive DNA supercoiling in the yeast plasmid has implications for a plasmid origin of a chromosome centromere. Proc Natl Acad Sci USA 108, 13671–13676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huertas P, Aguilera A (2003). Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol Cell 12, 711–721. [DOI] [PubMed] [Google Scholar]
- Huertas P, Garcia-Rubio ML, Wellinger RE, Luna R, Aguilera A (2006). An hpr1 point mutation that impairs transcription and mRNP biogenesis without increasing recombination. Mol Cell Biol 26, 7451–7465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyland KM, Kingsbury J, Koshland D, Hieter P (1999). Ctf19p: a novel kinetochore protein in Saccharomyces cerevisiae and a potential link between the kinetochore and mitotic spindle. J Cell Biol 145, 15–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin Q, Trelles-Sticken E, Scherthan H, Loidl J (1998). Yeast nuclei display prominent centromere clustering that is reduced in nondividing cells and in meiotic prophase. J Cell Biol 141, 21–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin QW, Fuchs J, Loidl J (2000). Centromere clustering is a major determinant of yeast interphase nuclear organization. J Cell Sci 113(Pt 11), 1903–1912. [DOI] [PubMed] [Google Scholar]
- Joglekar AP, Salmon ED, Bloom KS (2008). Counting kinetochore protein numbers in budding yeast using genetically encoded fluorescent proteins. Methods Cell Biol 85, 127–151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kabeche L, Nguyen HD, Buisson R, Zou L (2018). A mitosis-specific and R loop-driven ATR pathway promotes faithful chromosome segregation. Science 359, 108–114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kastenmayer JP, Lee MS, Hong AL, Spencer FA, Basrai MA (2005). The C-terminal half of Saccharomyces cerevisiae Mad1p mediates spindle checkpoint function, chromosome transmission fidelity and CEN association. Genetics 170, 509–517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laloraya S, Guacci V, Koshland D (2000). Chromosomal addresses of the cohesin component Mcd1p. J Cell Biol 151, 1047–1056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrimore J, Bloom K (2019). The regulation of chromosome segregation via centromere loops. Crit Rev Biochem Mol Biol 54, 352–370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrimore J, Bloom KS, Salmon ED (2011). Point centromeres contain more than a single centromere-specific Cse4 (CENP-A) nucleosome. J Cell Biol 195, 573–582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrimore J, Doshi A, Friedman B, Yeh E, Bloom K (2018). Geometric partitioning of cohesin and condensin is a consequence of chromatin loops. Mol Biol Cell 29, 2737–2750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ling YH, Yuen KWY (2019). Point centromere activity requires an optimal level of centromeric noncoding RNA. Proc Natl Acad Sci USA 116, 6270–6279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Su H, Zhang J, Liu Y, Feng C, Han F (2020). Back-spliced RNA from retrotransposon binds to centromere and regulates centromeric chromatin loops in maize. PLoS Biol 18, e3000582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T))Method. Methods 25, 402–408. [DOI] [PubMed] [Google Scholar]
- Luna R, Rondon AG, Perez-Calero C, Salas-Armenteros I, Aguilera A (2019). The THO complex as a paradigm for the prevention of cotranscriptional R-loops. Cold Spring Harb Symp Quant Biol 84, 105–114. [DOI] [PubMed] [Google Scholar]
- Ma L, Ho K, Piggott N, Luo Z, Measday V (2012). Interactions between the kinetochore complex and the protein kinase A pathway in Saccharomyces cerevisiae. G3 (Bethesda) 2, 831–841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKinley KL, Cheeseman IM (2016). The molecular basis for centromere identity and function. Nat Rev Mol Cell Biol 17, 16–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller MP, Evans RK, Zelter A, Geyer EA, MacCoss MJ, Rice LM, Davis TN, Asbury CL, Biggins S (2019). Kinetochore-associated Stu2 promotes chromosome biorientation in vivo. PLoS Genet 15, e1008423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra PK, Au WC, Choy JS, Kuich PH, Baker RE, Foltz DR, Basrai MA (2011). Misregulation of Scm3p/HJURP causes chromosome instability in Saccharomyces cerevisiae and human cells. PLoS Genet 7, e1002303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra PK, Basrai MA (2019). Protein kinases in mitotic phosphorylation of budding yeast CENP-A. Curr Genet 65, 1325–1332. [DOI] [PubMed] [Google Scholar]
- Mishra PK, Baum M, Carbon J (2007). Centromere size and position in Candida albicans are evolutionarily conserved independent of DNA sequence heterogeneity. Mol Genet Genomics 278, 455–465. [DOI] [PubMed] [Google Scholar]
- Mishra PK, Ciftci-Yilmaz S, Reynolds D, Au WC, Boeckmann L, Dittman LE, Jowhar Z, Pachpor T, Yeh E, Baker RE, et al. (2016). Polo kinase Cdc5 associates with centromeres to facilitate the removal of centromeric cohesin during mitosis. Mol Biol Cell 27, 2286–2300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra PK, Guo J, Dittman LE, Haase J, Yeh E, Bloom K, Basrai MA (2015). Pat1 protects centromere-specific histone H3 variant Cse4 from Psh1-mediated ubiquitination. Mol Biol Cell 26, 2067–2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra PK, Ottmann AR, Basrai MA (2013). Structural integrity of centromeric chromatin and faithful chromosome segregation requires Pat1. Genetics 195, 369–379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra PK, Thapa KS, Chen P, Wang S, Hazbun TR, Basrai MA (2018). Budding yeast CENP-A(Cse4) interacts with the N-terminus of Sgo1 and regulates its association with centromeric chromatin. Cell Cycle 17, 11–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizuguchi G, Xiao H, Wisniewski J, Smith MM, Wu C (2007). Nonhistone Scm3 and histones CenH3-H4 assemble the core of centromere-specific nucleosomes. Cell 129, 1153–1164. [DOI] [PubMed] [Google Scholar]
- Ng TM, Waples WG, Lavoie BD, Biggins S (2009). Pericentromeric sister chromatid cohesion promotes kinetochore biorientation. Mol Biol Cell 20, 3818–3827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohkuni K, Kitagawa K (2012). Role of transcription at centromeres in budding yeast. Transcription 3, 193–197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearson CG, Yeh E, Gardner M, Odde D, Salmon ED, Bloom K (2004). Stable kinetochore-microtubule attachment constrains centromere positioning in metaphase. Curr Biol 14, 1962–1967. [DOI] [PubMed] [Google Scholar]
- Ranjitkar P, Press MO, Yi X, Baker R, MacCoss MJ, Biggins S (2010). An E3 ubiquitin ligase prevents ectopic localization of the centromeric histone H3 variant via the centromere targeting domain. Mol Cell 40, 455–464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rondon AG, Aguilera A (2019). R-loops as promoters of antisense transcription. Mol Cell 76, 529–530. [DOI] [PubMed] [Google Scholar]
- Santos-Pereira JM, Aguilera A (2015). R loops: new modulators of genome dynamics and function. Nat Rev Genet 16, 583–597. [DOI] [PubMed] [Google Scholar]
- Santos-Rosa H, Aguilera A (1994). Increase in incidence of chromosome instability and non-conservative recombination between repeats in Saccharomyces cerevisiae hpr1 delta strains. Mol Gen Genet 245, 224–236. [DOI] [PubMed] [Google Scholar]
- Sharma AB, Dimitrov S, Hamiche A, Van Dyck E (2019). Centromeric and ectopic assembly of CENP-A chromatin in health and cancer: old marks and new tracks. Nucleic Acids Res 47, 1051–1069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spencer F, Gerring SL, Connelly C, Hieter P (1990). Mitotic chromosome transmission fidelity mutants in Saccharomyces cerevisiae. Genetics 124, 237–249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stirling PC, Chan YA, Minaker SW, Aristizabal MJ, Barrett I, Sipahimalani P, Kobor MS, Hieter P (2012). R-loop-mediated genome instability in mRNA cleavage and polyadenylation mutants. Genes Dev 26, 163–175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stoler S, Keith KC, Curnick KE, Fitzgerald-Hayes M (1995). A mutation in CSE4, an essential gene encoding a novel chromatin-associated protein in yeast, causes chromosome nondisjunction and cell cycle arrest at mitosis. Genes Dev 9, 573–586. [DOI] [PubMed] [Google Scholar]
- Stoler S, Rogers K, Weitze S, Morey L, Fitzgerald-Hayes M, Baker RE (2007). Scm3, an essential Saccharomyces cerevisiae centromere protein required for G2/M progression and Cse4 localization. Proc Natl Acad Sci USA 104, 10571–10576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan-Wong SM, Dhir S, Proudfoot NJ (2019). R-loops promote antisense transcription across the mammalian genome. Mol Cell 76, 600–616.e606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells JP, White J, Stirling PC (2019). R loops and their composite cancer connections. Trends Cancer 5, 619–631. [DOI] [PubMed] [Google Scholar]
- Yeh E, Haase J, Paliulis LV, Joglekar A, Bond L, Bouck D, Salmon ED, Bloom KS (2008). Pericentric chromatin is organized into an intramolecular loop in mitosis. Curr Biol 18, 81–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoder TJ, McElwain MA, Francis SE, Bagley J, Muller EG, Pak B, O’Toole ET, Winey M, Davis TN (2005). Analysis of a spindle pole body mutant reveals a defect in biorientation and illuminates spindle forces. Mol Biol Cell 16, 141–152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zenklusen D, Vinciguerra P, Wyss JC, Stutz F (2002). Stable mRNP formation and export require cotranscriptional recruitment of the mRNA export factors Yra1p and Sub2p by Hpr1p. Mol Cell Biol 22, 8241–8253. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.