Abstract
Animal domestication and artificial selection give rise to gradual changes at the genomic level in populations. Subsequent footprints of selection, known as selection signatures or selective sweeps, have been traced in the genomes of many animal livestock species by exploiting variation in linkage disequilibrium patterns and/or reduction of genetic diversity. Domestication of most aquatic species is recent in comparison with land animals, and salmonids are one of the most important fish species in aquaculture. Coho salmon (Oncorhynchus kisutch), cultivated primarily in Chile, has been subjected to breeding programs to improve growth, disease resistance traits, and flesh color. This study aimed to identify selection signatures that may be involved in adaptation to culture conditions and traits of productive interest. To do so, individuals of two domestic populations cultured in Chile were genotyped with 200 thousand SNPs, and analyses were conducted using iHS, XP-EHH and CLR. Several signatures of selection on different chromosomal regions were detected across both populations. Some of the identified regions under selection contained genes such anapc2, alad, chp2 and myn, which have been previously associated with body weight in Atlantic salmon, or sec24d and robo1, which have been associated with resistance to Piscirickettsia salmonis in coho salmon. Findings in our study can contribute to an integrated genome-wide map of selection signatures, to help identify the genetic mechanisms of phenotypic diversity in coho salmon.
Subject terms: Computational biology and bioinformatics, Evolution, Genetics
Introduction
The implementation of genetic improvement programs in domestic populations to enhance desirable production traits, combined with the natural adaptation to new environments, has led to a variety of phenotypic variation between domestic populations of plants and animals1 as well as a strong differentiation between wild and domestic populations. Both natural and artificial selection in domestic species have left footprints across the genome, known as selection signatures, which can point to regions harboring functionally important sequences of DNA2,3. Theoretically, a beneficial variant that has been under selection can show long-range linkage disequilibrium (LD) and high frequency over a long period of time4,5. Therefore, these patterns allow us to detect such selection signatures in genomes. When successful, selection signature identification can provide novel insights into mechanisms that create diversity across populations and contribute to mapping of genomic regions underlying selected traits or phenotypes6,7. Approaches for detecting selection signatures rely on scanning variants across the genome of interest, searching for regions in which (i) the allele frequency spectrum is shifted towards extreme (high or low) frequencies; (ii) there is an excess of homozygous genotypes; (iii) there are long haplotypes with high frequencies and (iv) there is an extreme differentiation among populations. Several statistical methods has been developed to search for selection signatures, such as extended haplotype homozygosity (EHH)8, integrated haplotype score (iHS)9, hapFLK10, Cross Population Extended Haplotype Homozogysity (XP-EHH)11, the composite likelihood ratio (CLR); runs of homozygosity (ROH)12 and FST statistics13.
In fishes, domestication is recent in comparison with other land animals14, although some evidence of fish farming dates back approximately 3,500 years ago15. An exponential development of aquaculture has occurred since 1960, relying on the domestication of a handful aquatic species14. Salmonid species in particular have seen very intensive production increases over the last four decades. Currently, the most important farmed salmonids species in the word are Atlantic salmon (Salmo salar), rainbow trout (Oncorhynchus mykiss) and coho salmon (Oncorhynchus kisutch). Coho salmon belong to the Oncorhynchus genus and are naturally distributed across Pacific North American and Asian watersheds16. About 90% of farmed coho salmon production is based in Chile17. Farmed populations of coho salmon were established in Chile at the end the 1970′s, with the importation of ova from the Kitimat river (British Columbia) and the US state of Oregon18. During the 1990´s, a variety of breeding programs have been implemented for this species in Chile18, mainly focused on improving growth, disease resistance traits and flesh color18. The implementation of such breeding programs have likely, shaped the genetic diversity and haplotype structure of these populations, and their present genomes may contain traceable signatures of selection.
Genomic scans for detecting selection signatures have been successfully applied to several domestic animals, including aquaculture species. For instance, selection signature scans have been carried out for Atlantic salmon19–24, Nile tilapia25,26; and channel catfish27. To date, there are no studies focusing on the identification of selection signatures in farmed coho salmon populations. To increase the knowledge of domestication and selection effects on the genome diversity in farmed populations of this species, we searched for selection signatures in two groups comprising farmed individuals of coho salmon using different statistical approaches based in haplotype structure and allele frequency spectrum. Tracing signatures of selection in this species could provide further insights on genomic regions responsible for important aquaculture features of domestic Pacific salmon.
Methods
Populations
In this study we used samples representing two different lines that come from a breeding program of coho salmon maintained in Chile, hereinafter pupulations A (Pop-A) and B (Pop-B): we used 89 individuals for A and 43 individuals for B. These breeding populations were established in 1997 and 1998, belonging to even and odd spawning year, respectively28. According to previous analyses (Ben Koop, unpublished data), it is very likely that these two populations correspond to a mixture of the two original broodstocks (Kitimat river and Oregon). Both populations have been selected to improve growth rate for around eight generations by using Best Linear Unbiased Prediction29,30 with genetic gains ranging between 6 to 13% per generation31. Sampling protocols were performed in accordance with the Comité de Bioética Animal, Facultad de Ciencias Veterinarias y Pecuarias, Universidad de Chile, Chile (certificate No. 08-2015). The study was carried out in compliance with the ARRIVE guidelines.
Genotyping
Genomic DNA was extracted from fin clips of each individual and genotyped with a 200 K Thermo Fisher Scientific Axiom myDesign Custom Array developed by the EPIC4 genome consortium32 (http://www.sfu.ca/epic4/).
Genotype calling was performed using Axiom Analysis Suite v3.1 (Thermo Fisher Scientific) following the Axiom Analysis user guide. After filtering, including removing markers with no position on the coho salmon reference genome (GCF_002021735.1) and markers that were identified as problematic (OTV, Call Rate Below Threshold, Other), 167,486 SNP were kept32. In this study, we removed markers positioned in scaffolds, which were not placed within chromosomes; therefore 136,500 markers were kept for downstream analyses. All these markers and samples passed the quality criteria (missing call rate ≤ 0.1).
Genetic diversity, linkage disequilibrium (LD) and population structure
A common subset of 72,616 SNPs with MAF ≥ 0.05 and no deviation from Hardy–Weinberg equilibrium in populations A and B were explored to describe the genetic diversity of each population. Three parameters, including the number of SNPs with MAF ≥ 0.05; observed heterozygosity (HO), and expected heterozygosity (HE) were calculated using PLINK33.
We used the squared correlation coefficient between SNP pairs (r2) to measure LD34, which was calculated for all syntenic marker pairs. To enable a clear presentation of results showing LD in relation to physical distance between markers, SNP pairs were put into bins of 100 kb, and mean values of r2 were calculated for each bin. The mean r2 for each of the distance bins was then plotted against the distance bin range (Mb). This analysis was carried out on a chromosome-by-chromosome basis.
To visualize genetic differentiation between populations a Principal component analysis (PCA) was conducted using PLINK. In addition, ADMIXTURE35 was employed to analyze the population structure, which was run with 10,000 iterations using the correlated allele model with K value from 1 to 20 to choose the optimal number of clusters at the lowest cross-validation error. Results were plotted using R36.
Selection signatures
We combined three methods (XP-EHH, iHS and CLR) which have shown to have a power > 70% to detect selection signatures in comparison with other combinations of statistical tests5. Removing markers deviating from Hardy–Weinberg equilibrium might be obfuscating the important deviations such as those resulting from evolutionary selection37, therefore, we did not apply Hardy–Weinberg equilibrium filtering for selection signatures analyses. In addition, different MAF thresholds were used for each selection signatures test, which are described below.
XP-EHH. The XP-EHH (Cross Population Extended Haplotype Homozygosity) statistic compares the integrated extended haplotype homozygosity (EHH) profiles between two populations at the same SNP11. Therefore, the computation of EHH is required for each population. The XP-EHH statistic is then defined as ln(IPopA/IPopB), where IPopA is the integrated EHH for the population A and IPopB is the integrated EHH for population B. Negative XP-EHH scores suggest selection in B, whereas positive scores suggest selection acting in the A. A common subset of 89,715 SNPs with MAF > 0.01 was used for XP-EHH test, which was computed for each SNP using the R package rehh 3.1.138, with default options. As the XP-EHH values are normally distributed, a Z-test is applied to identify significant SNPs under selection. We used -log10(p value) = 3 (p value ≤ 0.001) as the threshold for considering XP-EHH score as significant evidence of selection and at least two SNPs ≤ 500 kb apart.
iHS. The iHS statistics was used to detect footprints of selection within the studied populations. This test is based on the standardized log ratio of integrals of the decay of the EHH, computed for both ancestral and derived alleles of the focal SNP. Selection induces hitchhiking (genetic draft), that leads to extended haplotypes for the selected allele and a slower fall-off of EHH on either side of the selected locus. Thus, a high iHS scoring SNP is typically associated with longer haplotypes and lower neighborhood diversity compared to the other SNPs39. Phased haplotypes were obtained using Beagle40. The ancestral and derived alleles of each SNP were inferred in two ways: (i) the SNP probes were compared with the genome of Atlantic salmon, which was used as an outgroup species; (ii) for SNPs that could not be compared, the ancestral allele were inferred as the most common allele in the total dataset, as suggested in other studies (e.g.41,42). SNPs with MAF < 0.05 were excluded from iHS analysis in accordance with9, therefore, we used 95,217 and 90,101 SNPs for populations A and B, respectively. The iHS score was computed for each autosomal SNP using the R package rehh 3.1.138, using default options. Similar to XP-EHH, iHS values are normally distributed, and a Z-test is applied to identify significant SNPs under selection. We also used -log10(p value) = 3 (p value ≤ 0.001) as the threshold for considering iHS score as significant evidence of selection and at least two SNPs ≤ 500 kb apart.
CLR: We implemented CLR test in the software SweeD V3.3.243, downloaded from https://cme.h-its.org/exelixis/web/software/sweed/. SweeD evaluates the variation in the site-frequency spectrum along the chromosome and implements the composite likelihood ratio (CLR) statistics44. CLR computes the ratio of the likelihood of a selective sweep at a given position to the likelihood of a null model without a selective sweep. Following other studies (e.i.45–47), SNPs with MAF < 0.05 were excluded from CLR analysis as well, therefore, the same subset of SNPs were used as for iHS. We calculated the CLR within each population and for each chromosome separately at grid points for every 20 kb. We used the 99.5th percentile of the distribution of CLR scores as threshold for the detection of outliers.
Candidate genes and functional analysis
A genomic region was considered as being under selection if it matched the criteria described above for iHS, XP-EHH and CLR. For each identified selective sweep region, we extended the region containing outlier scores by adding 250 kb to each end, to account for potential blocks of high linkage disequilibrium. Gene annotation was performed using GCF_002021735.1 coho salmon genome assembly (Accession PRJNA378663, Okis_V1; https://www.ncbi.nlm.nih.gov/assembly/GCF_002021735.1)48 from the NCBI Eukaryotic genome annotation pipeline.
The Database for Annotation, Visualization, and Integrated Discovery (DAVID) v6.8 tool49 was used to identify Gene Ontology (GO) terms and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways using a list of genes within significant regions based on iHS, XP-EHH, CLR values and the zebrafish annotation file as a reference genome.
Ethics approval and consent to participate
Coho salmon individuals and sampling procedures were approved by the Comité de Bioética Animal from the Facultad de Ciencias Veterinarias y Pecuarias, Universidad de Chile (certificate Nº No. 08-2015).
Consent for publication
Not applicable.
Results
Population B has a slightly greater HO = 0.39 (HE = 0.36) than Pop-A HO = 0.37 (HE = 0.36). The number of SNPs with MAF ≥ 0.05 was higher in Pop-A (95,217) than Pop-B (90,101). In addition LD presented a higher level in Pop-B (r 2= 0.14) than Pop-A (r2 = 0.10) (Fig. 1). LD decayed faster in Pop-A, reaching a level of 0.14 around 500 kb, while in Pop-B at the same distance LD reaches a value of 0.18. A similar pattern was observed for most of the chromosomes, except for, and Oki1, Oki21, Oki28, where LD decay was slightly higher in Pop-B and Oki-10, where both populations presented a similar pattern of LD decay (Supplementary Fig. S1).
Figure 1.
Decay of average linkage disequilibrium (r2) over distance across the two farmed populations. Different color lines represent populations: Pop-A = green, Pop-B = magenta.
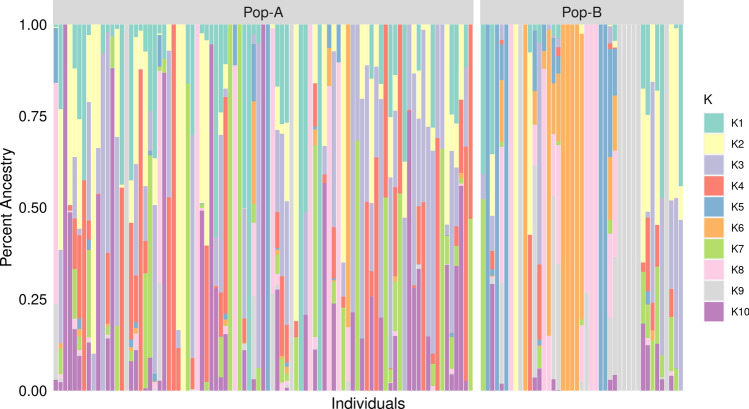
Principal component analysis (PCA) was employed to explore the clustering of individuals from the two populations. The PCA revealed two distinct clusters corresponding to population A and B. Principal components (PCs) 1, 2 and 3 jointly accounted for 39.7% of total variance, with PC 1 (26.1%) separating A and B; PC 2 (7.6%) and PC 3 (6%) not clearly discriminating populations. However, Pop-B produced a much more heterogeneous group with individuals spread along both PC 2 and 3 (Fig. 2). ADMIXTURE analysis revealed that the lowest cross-validation error was reached at K = 10 (Supplementary Fig. S2 and S3), showing 10 ancestral lineages for these two populations (Fig. 3). Pop-A individuals show high degree of mixed ancestry, being dominated by several ancestral lineages proportions among individuals. In contrast, Pop-B individuals tend to be dominated by single ancestral lineages among individuals, the represented lineages are quite different, which is consistent with a greater dispersion of Pop-B individuals in the PCA in comparison with Pop-A.
Figure 2.
Principal components analysis (PCA) of genetic differentiation among individuals. Three first components are shown. Each point represents one individual, and different colors represent populations: Pop-A = green, Pop-B = magenta.
Figure 3.
Individual assignment probabilities generated with ADMIXTURE (K = 10). Each color represents a cluster, and the ratio of vertical lines is proportional to assignment probability of an individual to each cluster.
Genome-wide scanning for selection signatures
To detect signatures of selection in the two coho salmon breeding populations, we used three statistical methods. Two of them are based in extended haplotype homozygosity (EHH): iHS and XP-EHH, and one method is based on the variation of the site-frequency spectrum, the composite likelihood ratio (CLR) tests.
Inner circles of Figs. 4 and 5 show the genome-wide distribution of iHS values. The iHS statistics revealed 156 and 99 SNPs with significant iHS scores (p ≤ 0.001) representing 0.18% and 0.12% highest values in populations A and B respectively. We found 26 and 16 regions in these respective populations with at least two significant SNPs that are ≤ 500 kb apart. These genomic regions spanned 17.1 Mb across chromosomes Oki5, Oki6, Oki28 and Oki29 in Pop-A, and 7.5 Mb in Pop-B in Oki5, Oki14, Oki15, Oki17, Oki18, Oki22 and Oki26 (Table S1). The highest values were present in Oki5 and Oki28 for Pop-A, and Oki14 and Oki18 for Pop-B. XP-EHH revealed 43 SNPs, which surpassed the significance threshold (Figs. 4B, 5B), representing the 0.05% highest values, 41 with positive scores and indicating selection in Pop-A, and two with negative scores indicating selection in Pop-B (Table S2). According to the criteria that we defined, six genomic regions spanning 3.72 Mb were detected in Pop-A, and one region of 520.8 kb in Pop-B. Genomic regions with XP-EHH values above 3 were detected in Oki1, Oki10 and Oki28 for Pop-A. The highest XP-EHH estimate was on Oki1 (XP-EHH = 3.8030, -log10(p value) = 3.8448), and the lowest value was on Oki15 (XP-EHH = -3.373854, log10(p value) = 3.130043). As a complementary method to iHS and XP-EHH, we performed the composite likelihood ratio (CLR). Figures 4C and 5C illustrate the CLR statistics against the genome positions for coho salmon. Based on a conservative 99.9% outlier threshold (CLRA ≥ 4.81, CLRB ≥ 4.64), we detected 95 regions affected by selective sweeps, covering 28.24 Mb and 21.92 Mb in population A and B, respectively (Table S3). In both populations, the results provided evidence of selective sweeps in several chromosomes across the genome. In Pop-A the highest CLR values were located on Oki14 (CLR = 9.98) within a region of 180 kb and on Oki17 (CLR = 9.35) within a region of 400 kb. While, in Pop-B a clear evidence of selective sweep is observed on Oki15 (CLR = 10.75), with a cluster of extreme signals in a genomic region of 780 kb.
Figure 4.
Circos plot of the global distribution of selection signatures across the genome in Pop-A. The circles from inside to outside illustrate: (A) -log10(p-value) of iHS (Blue); (B) −log10(p value) of XP-EHH (Purple) and (C) CLR values (Green). Values for each test surpassing the threshold (iHS and XP-EHH ≥ 3, and CLR ≥ 4.81) are highlighted in red.
Figure 5.
Circos plot of the global distribution of selection signatures across the genome in Pop-B. The circles from inside to outside illustrate: (A) −log10(p value) of iHS (Blue); (B) −log10(p value) of XP-EHH (Purple) and (C) CLR values (Green). Values for each test surpassing the threshold (iHS and XP-EHH ≥ 3, and CLR ≥ 4.64) are highlighted in red.
Functional characterization of genomic regions
A total of 1,118 and 812 genes within the identified genomic regions putatively subjected to selection were retrieved for Pop-A and Pop-B, respectively. Gene annotation of selected regions revealed several functionally important genes, such as kdm6a involved in the regulation of anterior/posterior body axis development in zebrafish50 or sec24d and robo1 which were associated to resistance against Piscirickettsia salmonis in farmed coho salmon populations from Chile51. Genes in genomic regions with the highest scores for iHS, XP-EHH and CLR are shown in Tables 1, 2 and 3 while the complete list of the genes found is presented in Table S4.
Table 1.
Genomic regions and genes spanning the five strongest detected selection signatures by iHS in each population.
| CHR | START | END | POP | Mean (-log p-value) | Max (-log p-value) | GENES |
|---|---|---|---|---|---|---|
| 5 | 25,314,578 | 26,652,908 | Pop-A | 3.7658 | 4.4176 | il17rd, appl1, hesx1, pacsin1b, lhfpl5a, si:ch211-86h15.1, atp6ap1lb, rps10, srpk1b, mapk14b, kif21b, LOC108182861, arhgef3, rps23, snrpc, zgc:101,577, grm4, nudt3b, b3galt6, spdef, tmem167b, adora1b, cpne5b, srpk1b |
| 6 | 24,927,655 | 28,923,100 | Pop-A | 3.6906 | 4.4797 | tsku, zswim7, timm22, LOC100004180, dscama, serpinh1b, ncor1, akap10, usp25, ei24, bhlha9, auts2a, stt3a, samsn1b, nrip1b, zgc:153,372, mmp13a, uvrag, spata22, acer3, specc1, wnt11r, umodl1, adora2b, ripk4, zgc:110,006, map6a, wscd1b, c2cd2, atg101, mmp30, rnft1, rps6kb1b, zgc:85,789, ptrh2, zbtb21, prdm15, mogat2, fuk, dgat2, cxadr, ints2, btg3, tmprss2, pcp4b, |
| 28 | 15,277,953 | 15,778,780 | Pop-A | 3.8221 | 4.1466 | aqp11, clns1a, fam168a, capn5b, elmod1, gdpd4a, rsf1b.1, gria4b, rps3, cul5b, LOC101885270 |
| 28 | 23,148,967 | 24,055,693 | Pop-A | 4.2248 | 4.4858 | rars, fgfr4, pgrmc1, wwc1, arl10, nop16, amot, uimc1, zgc:77,151, grk6, dbn1, pgk1, nsd1b, rab24, znf346, prr7 |
| 28 | 33,699,959 | 34,200,324 | Pop-A | 5.9758 | 5.9758 | LOC556987, myo18aa, cuedc1b, msi2b, akap1b, vezf1a, lgals9l3, ksr1a, LOC101886660 |
| 14 | 53,026,331 | 54,393,332 | Pop-B | 3.7398 | 4.5725 | zgc:113,201, rrh, mark3b, exoc3l4, cdca4, plcb3, tmx4, hao1, napba, cdc5l, supt3h, trmt61a, zgc:100,846, ckba, eif5, tnfaip2b, enpep, zgc:162,509, gabrb1, commd8, atp10d, metap1, eif4eb, sobpa, pdss2, bend3, apoba, hs1bp3, ypel5, LOC795051, xkr6a, kcnf1b, tmem151bb, ldah, gdf7, zgc:123,321, odc1, diexf, pqlc3, lamtor3, rock2a |
| 18 | 50,563,258 | 51,589,747 | Pop-B | 3.8271 | 4.8181 | dph1, dixdc1a, zbtb16b, hic1, tsr1, paf1, lrfn1, zgc:162,339, gmfg, socs9, tmprss5, pih1d2, zbtb16b, ncam1b |
| 18 | 53,846,890 | 54,476,564 | Pop-B | 3.6046 | 3.903 | rbm7, barx2, kirrel3b |
| 22 | 36,378,784 | 36,879,237 | Pop-B | 3.9302 | 4.2038 | glsl, garem, fanci, csnk1g1, nedd1, fam3c, LOC557301, LOC100002706, zgc:113,426, gnai1, zgc:114,188, rhcga, sh3gl3a, sycp3, actr6, fbxo31 |
| 26 | 17,677,558 | 18,183,422 | Pop-B | 3.7923 | 4.4897 | hdac4, LOC110440056, traf3ip1, twist2, asb1, adarb1b, agr1 |
Table 2.
Genome regions and genes spanning the detected selection signatures by XP-EHH in each population.
| CHR | START | END | POP | Mean (-log p-value) | Max (-logp-value) | GENES |
|---|---|---|---|---|---|---|
| 1 | 32,284,017 | 33,044,378 | Pop-A | 3.2336 | 3.4267 | hvcn1, myl2b, ddhd2, crybb3, loxl2b, tacc1, pxna, ela3l, golga7, cysltr3, cita, pnp4a, dnai1.2, letm2, rplp0, rem1, fam219ab, r3hcc1 |
| 1 | 33,174,438 | 33,714,143 | Pop-A | 3.4237 | 3.8448 | elmo2, tbc1d20, s100z, ssr1, rnf41, trib3, dgkaa, nrn1b, sys1, dnajc14, inpp1, nr4a1, arfgap1, acot19, maf1, rbck1, cs, cnpy2, nab2, ankrd52a |
| 1 | 35,604,181 | 36,229,121 | Pop-A | 3.1771 | 3.3292 | esyt1b, lrp1aa, zc3h10, lrrn1, sumf1, itpr1b, rbms2b |
| 1 | 36,926,728 | 37,662,405 | Pop-A | 3.109 | 3.2145 | srgap3, rad18, thumpd3, oxtr, kcnb1, grm2a, ptgis, cav3 |
| 10 | 31,318,086 | 31,853,386 | Pop-A | 3.1828 | 3.2101 | aqp4, cdh2, psma8, kdm4ab, kctd1, st3gal3b |
| 15 | 60,372,309 | 60,893,146 | Pop-B | 3.0711 | 3.13 | robo1 |
| 28 | 6,889,905 | 7,412,498 | Pop-A | 3.2376 | 3.473 | stk10, dpysl3, sh3pxd2b, stk32a |
Table 3.
Genome regions and genes spanning the five detected selection signatures by CLR in each population.
| CHR | START | END | POP | CLR | GENES |
|---|---|---|---|---|---|
| 14 | 55,748,684 | 56,248,684 | Pop-A | 8.928977 | |
| 14 | 55,808,732 | 56,308,732 | Pop-A | 9.977713 | spata7, spred1 |
| 17 | 39,471,248 | 39,971,248 | Pop-A | 9.266553 | cnbp, vgll4b, irf10, iqsec1b, raf1a |
| 17 | 39,491,264 | 39,991,264 | Pop-A | 9.349519 | syn2b, bmp7b, slc6a1a, tnnc1a, gp9, pparg, dusp7 |
| 21 | 28,486,708 | 28,986,708 | Pop-A | 8.221705 | cnstb |
| 15 | 65,822,888 | 66,322,888 | Pop-B | 9.704407 | |
| 15 | 65,842,896 | 66,342,896 | Pop-B | 10.53499 | |
| 15 | 65,862,900 | 66,362,900 | Pop-B | 9.326089 | aadac |
| 15 | 65,902,908 | 66,402,908 | Pop-B | 10.75308 | |
| 15 | 65,942,920 | 66,442,920 | Pop-B | 9.256851 | appbp2 |
We further investigated the functions associated with the putative genes undergoing positive selection by analyzing over-represented GO terms and pathways using DAVID. The identified GO terms and pathways are shown in Table S5. A total of 21 and 13 GO terms were observed for A and B, respectively. This includes 13 terms for biological process (BP), 3 for molecular function (MF), and 9 for cellular component (CC). The BP category is shown in Fig. 6; most of these terms are associated with basic metabolic processes, such as Developmental Process, Multicellular Organismal Process, Single-Organism Process, Response To Stimulus, Developmental Process, Presynaptic Process and Growth, among other. Additionally, 11 pathways were enriched (Table S5) such as Vascular smooth muscle contraction, Endocytosis, Regulation of actin cytoskeleton and TGF-beta signaling pathway.
Figure 6.
Functional annotation of candidate selective sweeps genes. Gene ontology (GO) analysis performed for Biological process. Color intensity of circle shows the significance of GO term and the size of the circle represents the number of genes associated with the GO term.
Discussion
Selective events are the major mechanism driving differentiation of populations. In this study, we assessed the genetic structure and detected selection signatures in two domestic populations of coho salmon by using three complementary statistical methods. First, we investigated basic population genetics statistics. Genetic diversity estimates, assessed by observed and expected heterozygosity, were in the upper limit of those reported in other salmon species introduced for farming purposes in Chile, such as Atlantic salmon, in which heterozygosity levels of these populations have been shown to range from 0.266 to 0.3721,22. In addition LD, which can be informative of population demography, decays similarly to what has been observed on previous studies in farmed coho salmon population from Chile32, and more rapidly than farmed Atlantic salmon52. Thus, the higher levels of genetic diversity and the lower levels of LD in farmed coho salmon populations suggest less impact of artificial selection on genome diversity, likely due to a shorter period of time of domestication in comparison with Atlantic salmon aquaculture populations from Chile. Genetic structure analyses revealed a high level of admixture in both populations. It is well known that demographic forces, such as admixture can mimic the patterns of genetic variation expected under selection53, 54. Therefore, we cannot discard the presence of false positives in the detection of selection signatures in this study, due to a high degree of mixed ancestry of these populations.
Several methods have been developed for detecting selection signatures in the genome. These methods can be grouped into three categories based on the type of genetic information exploited: population differentiation, site-frequency spectrum and linkage disequilibrium approaches18. In the present study we implemented three different tests (iHS, XP-EHH and CLR) to comprehensively identify candidate regions of positive selection in coho salmon. As is shown in Fig. 7, different genomic regions were detected by these methods. CLR have the largest overall number of signals detected to be underlying selection, followed by iHS and XP-EHH. No overlap was observed among the three methods, however CLR and iHS (Fig. 7), showed five overlapping regions on Oki5, Oki6 and Oki14 spanning a total of 2.02 Mb. Low overlap among methods based on allele frequencies compared to those based on haplotype patterns has been observed in previous studies in Atlantic salmon21,23. Discrepancies between loci under putative selection detected by different methods are expected, given that they benefit from different sources of information from genome variation. For instance, iHS test has advantage in detecting selective sweep with variants at intermediate frequencies11, while XP-EHH is more powerful at detecting complete or nearly complete selective sweeps11.
Figure 7.
Genomic distribution of selection signatures detected by iHS, XP-EHH, CLR and per populations on all chromosomes.
On the other hand, a low overlap between the two populations was observed (Fig. 7). Four chromosomes showed common regions harboring selection signatures: Oki5 (1.86 Mb); Oki14 (946 kb), Oki16 (41.5 kb) and Oki26 (57.3 kb). We suggest this low overlap among populations might be the effect of breeding program goals, which predominantly are focused on improving polygenic traits (e.g., growth), therefore, different genomics regions on different populations are affected by selection across the genome. The population-specific signals suggest that selection may have acted upon different genes, which has been already shown in other species and can be interpreted as a polygenic basis of parallel selection traits55. Several studies in fish have suggested that the same phenotype may arise from different genetic pathways among populations21,23,56–59. Evolution of complex parallel phenotypes can indeed emerge from different evolutionary processes and this is likely to happen when inbreeding and genetic drift play a greater role than selection21. On the other hand, it is worth mentioning that a low coverage of the genome using a SNP array might have caused lacking of information of some important genomic regions harboring selection signatures. This issue might be addressed with the use of whole genome sequencing, which could strongly enhance the accuracy of selection signatures detection by increasing the density of SNPs covering a greater area of the genome60,61.
One of the goals of this study was to identify putative candidates genes involved in the domestication process and artificial selection of coho salmon. In accordance with the primary genetic improvement goal, where growth rate has been the main focus, we found a series of genes potentially relevant for this trait, including anapc2; alad; bdkrb2; fam98b; chp2; myn that were previously associated with body weight in a genome-wide association study in Atlantic salmon (GWAS)62 and sytl5, txnrd1 genes associated with growth traits in juvenile farmed Atlantic salmon63. The chp2 gene was also found to be putatively under selection in an Atlantic salmon domestic population selected for fast growth22. We also identified the kdm6a gene, which has been involved in the regulation of anterior/posterior body axis development in zebrafish50 and has been related to growth-related traits in pigs64. Furthermore, we identified the biological processes “Developmental Process” and “Growth”, within the significant terms associated with the genes harbored in the identified genomic regions, suggesting that loci controlling growth and development are most likely under selection in these farmed coho salmon populations.
On the other hand, host–pathogen interactions lead to strong selection in the genome of vertebrate species65. We suggest that adaptation to farming environment has imposed selection on genomic regions related to the immune system in coho salmon. Here we found the cnpy2, synrg and med10 genes, which were previously shown to be affected by parasite-driven selection in Atlantic salmon66. More importantly, we detected the genes sec24d and robo1 that have been associated with disease resistance in the face of a bacterial disease (Piscirickettsia salmonis) in coho salmon populations farmed in Chile51.
Artificial selection may be applied either inadvertently (unconsciously) or intentionally (consciously)67. For most livestock species, it is thought that the early stages of domestication involved unconscious selection for behavioral traits (e.g., for tameness and reduced aggression), followed by selection focused on breeding objectives68. In fish, behavioral traits such as swimming capacities, foraging, social interactions, reproduction, or personality and cognitive abilities, are also modified by domestication69. In the present study we identified the genes robo1 and dcdc2 which are associated to complex cognitive acquired skills, including spoken and written language in human70. We also identified the gtf2ird1 associated to aggression and social interaction in mice71 and recently shown to be under selection in Nile tilapia25. We suggest that such genes could be associated with behavioral traits in coho salmon as well.
Sexual maturation and reproductive traits have profound importance from both evolutionary and economic perspective. In fish farming, maturation is frequently delayed by exposing fish to continuous light or light regimes, which are different to those, found in natural environment, affecting the perception of seasonality and circannual rhythms72. In this study, we identified the picalmb and tsku genes with evidence of selection. These genes were recently associated to maturation traits in Atlantic salmon73. Furthermore, we also detected the vgll4b, which has previously shown signs of selection in Atlantic salmon farmed populations22. It is important to mention that a gene from the VGLL (vgll3) family is associated with age to maturity in natural populations of Atlantic salmon74,75.
Conclusion
This study reported the identification of selection signatures in the genome of aquaculture lines of coho salmon. Our analyses of two domestic populations revealed putative selection signatures of genomic regions containing genes involved in growth, immune system, behavior and maturation traits. As expected, these findings are congruent with the breeding goal for the populations studied here, which includes the most highly selected trait, growth rate. In addition, the effect of domestication and adaptation to captive environment, most likely affecting loci controlling traits that have not been directly part of the breeding objectives, including immunity and, behavior. These results provide further insights into the genome diversity changes driven by domestication and selection mechanisms in farmed coho salmon populations.
Supplementary Information
Acknowledgements
We thank the Government of Canada through Genome Canada, Genome British Columbia, and Genome Quebec for supporting this research that was carried out in conjunction with the project EPIC4 (Enhanced Production in Coho: Culture, Community, Catch). We also would like to acknowledge Pesquera Antares S.A., which provided the fish used in this study. JMY is supported by Núcleo Milenio INVASAL funded by Chile’s government program, Iniciativa Científica Milenio from Ministerio de Economía, Fomento y Turismo.
Author contributions
M.E.L. and J.M.Y. conceived and designed the study. M.E.L. performed the analyses and wrote the the manuscript. M.I.C. contributed with data analyses. M.I.C., E.B.R., B.F.K. and J.M.Y. contributed to discussion and writing. All authors have reviewed and approved the final manuscript.
Funding
This research was partially funded by Government of Canada through EPIC4 (Enhanced Production in Coho: Culture, Community, Catch) from Genome Canada, Genome British Columbia, and Genome Quebec.
Data availability
Genotype data available from Figshare repository, 10.6084/m9.figshare.14230430
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-021-86154-w.
References
- 1.Sánchez-Villagra MR, Geiger M, Schneider RA. The taming of the neural crest: a developmental perspective on the origins of morphological covariation in domesticated mammals. R. Soc. Open Sci. 2016;3:160107. doi: 10.1098/rsos.160107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bertolini F, Servin B, Talenti A, Rochat E, Kim ES, Oget C, Palhière I, Crisà A, Catillo G, Steri R, Amills M, Colli L, Marras G, Milanesi M, Nicolazzi E, Rosen BD, Van Tassell CP, Guldbrandtsen B, Sonstegard TS, Tosser-Klopp G, Stella A, Rothschild MF, Joost S, Crepaldi P, Adaptmap C. Signatures of selection and environmental adaptation across the goat genome post-domestication. Genet. Sel. Evol. GSE. 2018;50:57–57. doi: 10.1186/s12711-018-0421-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Qanbari S, Simianer H. Mapping signatures of positive selection in the genome of livestock. Livest. Sci. 2014;166:133–143. doi: 10.1016/j.livsci.2014.05.003. [DOI] [Google Scholar]
- 4.Grossman S, Shylakhter I, Karlsson E, Byrne E, Morales S, Frieden G, Hostetter E, Angelino E, Garber M, Zuk O, Lander E, Schaffner S, Sabeti P. A composite of multiple signals distinguishes causal variants in regions of positive selection. Science. 2010;327:883–886. doi: 10.1126/science.1183863. [DOI] [PubMed] [Google Scholar]
- 5.Ma Y, Ding X, Qanbari S, Weigend S, Zhang Q, Simianer H. Properties of different selection signature statistics and a new strategy for combining them. Heredity. 2015;115:426. doi: 10.1038/hdy.2015.42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Andersson L, Georges M. Domestic-animal genomics: deciphering the genetics of complex traits. Nat. Rev. Genet. 2004 doi: 10.1038/nrg1294. [DOI] [PubMed] [Google Scholar]
- 7.Oleksyk TK, Smith MW, O’brien SJ. Genome-wide scans for footprints of natural selection. Philos. Trans. R. Soc. B Biol. Sci. 2010;365:185–205. doi: 10.1098/rstb.2009.0219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sabeti P, Reich D, Higgins J, Levine H, Richter D, Schaffner S, Gabriel S, Platko J, Patterson N, Mcdonald G, Ackerman H, Campbell S, Altshuler D, Cooper R, Kwiatkowski D, Ward R, Lander E. Detecting recent positive selection in the human genome from haplotype structure. Nature. 2002;419:832–837. doi: 10.1038/nature01140. [DOI] [PubMed] [Google Scholar]
- 9.Voight B, Kudaravalli S, Wen X, Pritchard J. A map of recent positive selection in the human genome. PLoS Biol. 2006;4:e72. doi: 10.1371/journal.pbio.0040072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Fariello MI, Boitard S, Naya H, Sancristobal M, Servin B. Detecting signatures of selection through haplotype differentiation among hierarchically structured populations. Genetics. 2013;193:929–941. doi: 10.1534/genetics.112.147231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sabeti P, Varilly P, Fry B, Lohmueller J, Hostetter E, Cotsapas C, Xie X, Byrne E, Mccarroll S, Gaudet R, Schaffner S, Lander E, Frazer K, Ballinger D, Cox D, Hinds D, Stuve L, Gibbs R, Belmont J, Boudreau A, Hardenbol P, Leal S, Pasternak S, Wheeler D, Willis T, Yu F, Yang H, Zeng C, Gao Y, Hu H. Genome-wide detection and characterization of positive selection in human populations. Nature. 2007;449:913–918. doi: 10.1038/nature06250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ceballos FC, Joshi PK, Clark DW, Ramsay M, Wilson JF. Runs of homozygosity: windows into population history and trait architecture. Nat. Rev. Genet. 2018;19:220. doi: 10.1038/nrg.2017.109. [DOI] [PubMed] [Google Scholar]
- 13.Weir B, Cardon L, Anderson A, Nielsen D, Hill W. Measures of human population structure show heterogeneity among genomic regions. Genome Res. 2005;15:1468–1476. doi: 10.1101/gr.4398405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Teletchea, F. Fish Domestication: An Overview. Animal Domestication (2018).
- 15.Balon EK. About the oldest domesticates among fishes. J. Fish Biol. 2004;65:1–27. doi: 10.1111/j.0022-1112.2004.00563.x. [DOI] [Google Scholar]
- 16.Groot C, Margolis L. Pacific Salmon Life Histories. Vancouver: UBC Press; 1991. [Google Scholar]
- 17.FAO. Fisheries and aquaculture information and statistical branch. (Rome: FAO). (2016).
- 18.Neira, R. et al. Evolution of coho salmon (Oncorhynchus kisutch) breeding programs. In Proceedings, 10th World Congress of Genetics Applied to Livestock Production (2014).
- 19.Gutierrez AP, Yáñez JM, Davidson WS. Evidence of recent signatures of selection during domestication in an Atlantic salmon population. Mar. Genomics. 2016;26:41–50. doi: 10.1016/j.margen.2015.12.007. [DOI] [PubMed] [Google Scholar]
- 20.Liu L, Ang KP, Elliott JaK, Kent MP, Lien S, Macdonald D, Boulding EG. A genome scan for selection signatures comparing farmed Atlantic salmon with two wild populations: testing colocalization among outlier markers, candidate genes, and quantitative trait loci for production traits. Evolut. Appl. 2016;10:276–296. doi: 10.1111/eva.12450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.López ME, Benestan L, Moore JS, Perrier C, Gilbey J, Di Genova A, Maass A, Diaz D, Lhorente JP, Correa K, Neira R, Bernatchez L, Yáñez JM. Comparing genomic signatures of domestication in two Atlantic salmon (Salmo salar L) populations with different geographical origins. Evol. Appl. 2018 doi: 10.1111/eva.12689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.López ME, Linderoth T, Norris A, Lhorente JP, Neira R, Yáñez JM. Multiple selection signatures in farmed atlantic salmon adapted to different environments across hemispheres. Front. Genet. 2019 doi: 10.3389/fgene.2019.00901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mäkinen H, Vasemägi A, Mcginnity P, Cross TF, Primmer CR. Population genomic analyses of early-phase Atlantic Salmon (Salmo salar) domestication/captive breeding. Evol. Appl. 2014;8:93–107. doi: 10.1111/eva.12230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Naval-Sanchez M, Mcwilliam S, Evans B, Yáñez JM, Houston RD, Kijas JW. Changed patterns of genomic variation following recent domestication: selection sweeps in farmed Atlantic salmon. Front. Genet. 2020 doi: 10.3389/fgene.2020.00264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cadiz MI, Lopez ME, Diaz-Dominguez D, Caceres G, Yoshida G, Gomez-Uchida D. Yanez J (2019) Whole genome re-sequencing reveals recent signatures of selection in three strains of farmed Nile tilapia (Oreochromis niloticus) Sci. Rep. 2020;10:11514. doi: 10.1038/s41598-020-68064-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Xia JH, Bai Z, Meng Z, Zhang Y, Wang L, Liu F, Jing W, Wan ZY, Li J, Lin H. Signatures of selection in tilapia revealed by whole genome resequencing. Sci. Rep. 2015;5:14168. doi: 10.1038/srep14168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nam B-H, Yoo D, Kim Y-O, Park JY, Shin Y, Shin G-H, Park C-I, Kim H, Kwak W. Whole genome sequencing reveals the impact of recent artificial selection on red sea bream reared in fish farms. Sci. Rep. 2019;9:6487. doi: 10.1038/s41598-019-42988-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yáñez JM, Bassini LN, Filp M, Lhorente JP, Ponzoni RW, Neira R. Inbreeding and effective population size in a coho salmon (Oncorhynchus kisutch) breeding nucleus in Chile. Aquaculture. 2014 doi: 10.1016/j.aquaculture.2013.05.028. [DOI] [Google Scholar]
- 29.Dufflocq P, Lhorente JP, Bangera R, Neira R, Newman S, Yáñez JM. Correlated response of flesh color to selection for harvest weight in coho salmon (Oncorhynchus kisutch) Aquaculture. 2017;472:38–43. doi: 10.1016/j.aquaculture.2016.08.037. [DOI] [Google Scholar]
- 30.Yáñez JM, Bangera R, Lhorente JP, Barría A, Oyarzún M, Neira R, Newman S. Negative genetic correlation between resistance against Piscirickettsia salmonis and harvest weight in coho salmon (Oncorhynchus kisutch) Aquaculture. 2016;459:8–13. doi: 10.1016/j.aquaculture.2016.03.020. [DOI] [Google Scholar]
- 31.Lhorente JP, Araneda M, Neira R, Yáñez JM. Advances in genetic improvement for salmon and trout aquaculture: the Chilean situation and prospects. Rev. Aquac. 2019;11:340–353. doi: 10.1111/raq.12335. [DOI] [Google Scholar]
- 32.Barria A, Christensen KA, Yoshida G, Jedlicki A, Leong JS, Rondeau EB, Lhorente JP, Koop BF, Davidson WS, Yáñez JM. Whole genome linkage disequilibrium and effective population size in a coho salmon (Oncorhynchus kisutch) breeding population using a high-density SNP array. Front. Genet. 2019;10:498. doi: 10.3389/fgene.2019.00498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, Bakker PI, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007 doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hill W, Robertson A. Linkage disequilibrium in finite populations. Theor. Appl. Genet. 1968;38:226–231. doi: 10.1007/BF01245622. [DOI] [PubMed] [Google Scholar]
- 35.Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009;19:1655–1664. doi: 10.1101/gr.094052.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.R Core Team . R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2019. [Google Scholar]
- 37.Hao W, Storey JD. Extending tests of Hardy–Weinberg equilibrium to structured populations. Genetics. 2019;213:759. doi: 10.1534/genetics.119.302370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gautier M, Klassmann A, Vitalis R. rehh 2.0: a reimplementation of the R package rehh to detect positive selection from haplotype structure. Mol. Ecol. Resour. 2017;17:78–90. doi: 10.1111/1755-0998.12634. [DOI] [PubMed] [Google Scholar]
- 39.Wagh K, Bhatia A, Alexe G, Reddy A, Ravikumar V, Seiler M, Boemo M, Yao M, Cronk L, Naqvi A, Ganesan S, Levine AJ, Bhanot G. Lactase persistence and lipid pathway selection in the Maasai. PLoS ONE. 2012;7:e44751. doi: 10.1371/journal.pone.0044751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Browning B, Browning S. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 2009;84:210–223. doi: 10.1016/j.ajhg.2009.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bahbahani H, Clifford D, Wragg MN, Mbole-Kariuki CV, Tassell TS, et al. Signatures of positive selection in East African Shorthorn Zebu: a genome-wide single nucleotide polymorphism analysis. Sci. Rep. 2015;5:11729. doi: 10.1038/srep11729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hacia JG, et al. Determination of ancestral alleles for human single-nucleotide polymorphisms using high-density oligonucleotide arrays. Nat. Genet. 1999;22:164–167. doi: 10.1038/9674. [DOI] [PubMed] [Google Scholar]
- 43.Pavlidis P, Živkovic D, Stamatakis A, Alachiotis N. SweeD: likelihood-based detection of selective sweeps in thousands of genomes. Mol. Biol. Evol. 2013;30:2224–2234. doi: 10.1093/molbev/mst112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Nielsen R, Williamson S, Kim Y, Hubisz MJ, Clark AG, Bustamante C. Genomic scans for selective sweeps using SNP data. Genome Res. 2005;15:1566–1575. doi: 10.1101/gr.4252305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen M, Wang J, Wang Y, Wu Y, Fu J, et al. Genome-wide detection of selection signatures in Chinese indigenous Laiwu pigs revealed candidate genes regulating fat deposition in muscle. BMC Genet. 2018;19:31. doi: 10.1186/s12863-018-0622-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hartmann FE, McDonald BA, Croll D. Genome-wide evidence for divergent selection between populations of a major agricultural pathogen. Mol. Ecol. 2018;27:2725–2741. doi: 10.1111/mec.14711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wang K, Wu P, Yang Q, Chen D, Zhou J, et al. Detection of selection signatures in Chinese Landrace and Yorkshire pigs based on genotyping-by-sequencing data. Front. Genet. 2018;9:119–119. doi: 10.3389/fgene.2018.00119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kim J-H, Leong JS, Koop BF, Devlin RH. Multi-tissue transcriptome profiles for coho salmon (Oncorhynchus kisutch), a species undergoing rediploidization following whole-genome duplication. Mar. Genomics. 2016;25:33–37. doi: 10.1016/j.margen.2015.11.008. [DOI] [PubMed] [Google Scholar]
- 49.Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13. doi: 10.1093/nar/gkn923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lan F, Bayliss PE, Rinn JL, Whetstine JR, Wang JK, Chen S, Iwase S, Alpatov R, Issaeva I, Canaani E. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature. 2007;449:689. doi: 10.1038/nature06192. [DOI] [PubMed] [Google Scholar]
- 51.Barría A, Christensen KA, Yoshida GM, Correa K, Jedlicki A, Lhorente JP, Davidson WS, Yáñez JM. Genomic Predictions and Genome-Wide Association Study of Resistance Against Piscirickettsia salmonis in Coho Salmon (Oncorhynchus kisutch) Using ddRAD Sequencing. G3: Genes|Genomes|Genetics. 2018;8:1183. doi: 10.1534/g3.118.200053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Barria A, López ME, Yoshida G, Carvalheiro R, Lhorente JP, Yáñez JM. Population genomic structure and genome-wide linkage disequilibrium in farmed atlantic salmon (Salmo salar L.) using dense SNP genotypes. Front. Genet. 2018 doi: 10.3389/fgene.2018.00649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jensen JD, Kim Y, DuMont VB, Aquadro CF, Bustamante CD. Distinguishing between selective sweeps and demography using DNA polymorphism data. Genetics. 2005;170:1401–1410. doi: 10.1534/genetics.104.038224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Tajima F. The effect of change in population size on DNA polymorphism. Genetics. 1989;123:597–601. doi: 10.1093/genetics/123.3.597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zhang S, Zhang K, Peng X, Zhan H, Lu J, et al. Selective sweep analysis reveals extensive parallel selection traits between large white and Duroc pigs. Evol. Appl. 2020;13:2807–2820. doi: 10.1111/eva.13085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Elmer KR, Fan S, Kusche H, Luise Spreitzer M, Kautt AF, et al. Parallel evolution of Nicaraguan crater lake cichlid fishes via non-parallel routes. Nat. Commun. 2014;5:5168. doi: 10.1038/ncomms6168. [DOI] [PubMed] [Google Scholar]
- 57.Perrier C, Bourret V, Kent MP, Bernatchez L. Parallel and nonparallel genome-wide divergence among replicate population pairs of freshwater and anadromous Atlantic salmon. Mol. Ecol. 2013;22:5577–5593. doi: 10.1111/mec.12500. [DOI] [PubMed] [Google Scholar]
- 58.Pujolar J, Ferchaud A, Bekkevold D, Hansen M. Non-parallel divergence across freshwater and marine three-spined stickleback Gasterosteus aculeatus populations. J. Fish Biol. 2017;91:175–194. doi: 10.1111/jfb.13336. [DOI] [PubMed] [Google Scholar]
- 59.Vasemägi, A. et al. Screen for footprints of selection during domestication/captive breeding of Atlantic salmon. Comp. Funct. Genomics2012, 1–14 (2012). [DOI] [PMC free article] [PubMed]
- 60.Rubin C-J, Megens H-J, Barrio AM, Maqbool K, Sayyab S, et al. Strong signatures of selection in the domestic pig genome. Proc. Natl. Acad. Sci. 2012;109:19529–19536. doi: 10.1073/pnas.1217149109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rubin C-J, Zody MC, Eriksson J, Meadows JRS, Sherwood E, et al. Whole-genome resequencing reveals loci under selection during chicken domestication. Nature. 2010;464:587–591. doi: 10.1038/nature08832. [DOI] [PubMed] [Google Scholar]
- 62.Yoshida GM, Lhorente JP, Carvalheiro R, Yáñez JM. Bayesian genome-wide association analysis for body weight in farmed Atlantic salmon (Salmo salar L.) Anim. Genet. 2017;48:698–703. doi: 10.1111/age.12621. [DOI] [PubMed] [Google Scholar]
- 63.Tsai, H. Y. et al Genome wide association and genomic prediction for growth traits in juvenile farmed Atlantic salmon using a high density SNP array. BMC genomics, 16(1), 1–9 (2015). [DOI] [PMC free article] [PubMed]
- 64.Guo X, Li M, Gao P, Cao G, Zhang W, Liu M, Wang H, Qin B, Liu J, Wang L. Identification of candidate genes of growth traits in pigs using RNA-sequencing. Ital. J. Anim. Sci. 2019;18:279–286. doi: 10.1080/1828051X.2018.1507628. [DOI] [Google Scholar]
- 65.Early AM, Clark AG. Genomic signatures of local adaptation in the Drosophila immune response. Fly. 2017;11:277–283. doi: 10.1080/19336934.2017.1337612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zueva KJ, Lumme J, Veselov AE, Kent MP, Lien S, Primmer CR. Footprints of directional selection in wild Atlantic salmon populations: evidence for parasite-driven evolution? PLoS ONE. 2014;9:e91672. doi: 10.1371/journal.pone.0091672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Price, E. O. Animal Domestication and Behavior (2002).
- 68.Gregory TR. Artificial selection and domestication: modern lessons from Darwin’s enduring analogy. Evol. Educ. Outreach. 2009;2:5–27. doi: 10.1007/s12052-008-0114-z. [DOI] [Google Scholar]
- 69.Pasquet, A. Effects of Domestication on Fish Behaviour. Animal Domestication (2018)
- 70.Mozzi A, Forni D, Clerici M, Pozzoli U, Mascheretti S, Guerini FR, Riva S, Bresolin N, Cagliani R, Sironi M. The evolutionary history of genes involved in spoken and written language: beyond FOXP2. Sci. Rep. 2016;6:1–12. doi: 10.1038/srep22157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Young E, Lipina T, Tam E, Mandel A, Clapcote S, Bechard A, Chambers J, Mount H, Fletcher P, Roder J. Reduced fear and aggression and altered serotonin metabolism in Gtf2ird1-targeted mice. Genes Brain Behav. 2008;7:224–234. doi: 10.1111/j.1601-183X.2007.00343.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Taranger GL, Carrillo M, Schulz RW, Fontaine P, Zanuy S, Felip A, et al. Control of puberty in farmed fish. Gen. Comp. Endocrinol. 2010;165:483–515. doi: 10.1016/j.ygcen.2009.05.004. [DOI] [PubMed] [Google Scholar]
- 73.Mohamed AR, Verbyla KL, Al-Mamun HA, Mcwilliam S, Evans B, King H, Kube P, Kijas JW. Polygenic and sex specific architecture for two maturation traits in farmed Atlantic salmon. BMC Genomics. 2019;20:139. doi: 10.1186/s12864-019-5525-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Ayllon F, Kjærner-Semb E, Furmanek T, Wennevik V, Solberg MF, Dahle G, Taranger GL, Glover KA, Almén MS, Rubin CJ, Edvardsen RB, Wargelius A. The vgll3 locus controls age at maturity in wild and domesticated atlantic salmon (Salmo salar L.) males. PLoS Genet. 2015;11:e1005628. doi: 10.1371/journal.pgen.1005628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Barson NJ, Aykanat T, Hindar K, Baranski M, Bolstad GH, Fiske P, Jacq C, Jensen AJ, Johnston SE, Karlsson S, Kent M, Moen T, Niemelä E, Nome T, Næsje TF, Orell P, Romakkaniemi A, Sægrov H, Urdal K, Erkinaro J, Lien S, Primmer CR. Sex-dependent dominance at a single locus maintains variation in age at maturity in salmon. Nature. 2015;528:405–408. doi: 10.1038/nature16062. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Genotype data available from Figshare repository, 10.6084/m9.figshare.14230430