LETTER
Reverse transcription-PCR (RT-PCR) is the gold standard for the diagnosis of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection; however, testing has been complicated by supply shortages and long turnaround times. A major limiting factor early in the coronavirus disease 2019 (COVID-19) pandemic was the lack of ready availability of the reagents for RNA extraction, which remains an expensive and time-consuming part of some testing modalities. Of great interest, extraction-free methods for use with both nasopharyngeal and saliva sources have been investigated in pilot studies (1–6). In this study, we assessed the detection of SARS-CoV-2 in two clinical laboratories from 144 nasopharyngeal specimens, utilizing an extraction-free method and RT-PCR using CDC primers/probe (Fig. 1A). We compared this method (Direct N2) with the widely used Cepheid Xpert Xpress SARS-CoV-2 assay, which has FDA emergency use authorization (EUA).
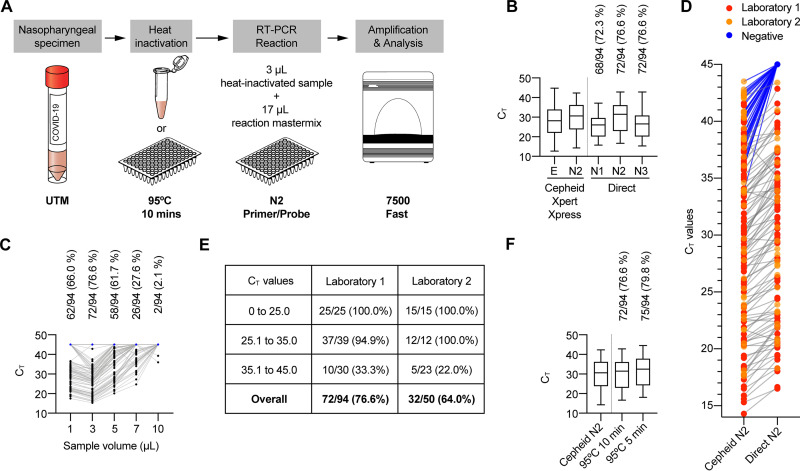
FIG 1.
Extraction-free SARS-CoV-2 N2 screening. (A) Generalized direct PCR procedure. (B) The performance of the N2 primer/probe pair was most comparable to that seen with the Cepheid N2 target. (C) Optimization of sample volume in reaction mixture. (D) Direct N2 CT values compared to initial Cepheid N2 CT value (negative = CT value of 45). (E) Performance of Direct N2 screening by CT value and performing laboratory. (F) Shorter heat inactivation duration modestly increased performance.
Both of the laboratories used previously frozen nasopharyngeal swab specimens in viral transport media (BD Universal viral transport media [UTM] at laboratory 1 and Medical Diagnostic Laboratories viral transport media or institution-produced viral transport media [following CDC standard operating procedure {SOP} no. DSR-052-05] at laboratory 2) which had been previously tested using the Cepheid assay described above. Laboratory 1 subjected 100-μl aliquots of 94 specimens to heat inactivation at 95°C for 10 min in a PCR thermocycler. Laboratory 2 subjected 500-μl aliquots of 50 thawed specimens in cryotubes to heat inactivation in an incubator set to 95°C for 10 min. Samples were brought to room temperature before 3 μl was added to 17 μl of master mix (1.5 μl, CDC 2019-nCoV primer/probe; 10.5 μl, nuclease-free water; 5.0 μl, TaqPath 1-Step reverse transcription-quantitative PCR [RT-qPCR] master mix [4×]). Synthetic “nCoVPC” was used for the positive control in laboratory 1, and whole viral SARS-CoV-2 RNA (kind gift from Scott Weaver, University of Texas Medical Branch) was used in laboratory 2. PCR cycling was performed using CDC parameters on a 7500 Fast real-time PCR instrument and SDS software (version 1.4.1; Applied Biosystems). The N2 primers/probe performed better than N1 and exhibited a mean and range of threshold cycle (CT) values more similar to those determined for the Cepheid N2 target than to those seen with the N3 target (Fig. 1B). The 3-μl sample input volume outperformed all other sample volumes tested (Fig. 1C); it is possible that inhibitory substances impair performance at higher volumes. The mean CT values from Direct N2 screening were expectedly higher than the corresponding CT values from the original Cepheid N2 assay (Fig. 1D), although results of comparisons of CT values across different assays must be interpreted with caution. Most of the specimens with Cepheid N2 CT values of <35 were detected with Direct N2 (89/91, 98%), whereas Direct N2 performed markedly less well for specimens with Cepheid N2 CT values of >35 (15/53, 28%) (Fig. 1E). Heat inactivating in plates for 5 min instead of 10 min led to a modest improvement in sensitivity (Fig. 1F).
The direct detection method used in this study is simple, economical, and robust, with an approximate hands-on time of 1 h and an instrument run time of ∼80 min to result per 96-well plate. Discrepant results occurred for specimens with Cepheid N2 CT values of approximately >35. Further, CT values from Cepheid testing were obtained using fresh specimens, whereas Direct N2 was performed after a freeze/thaw cycle, which may have contributed to increased CT values. Direct N2 also uses a lower specimen volume, which was likely the primary cause of reduced sensitivity. Overall, the process is high throughput as described here and is amenable to automation. In conclusion, while moderately less sensitive than conventional RT-PCR-based SARS-CoV-2 assays, direct methods may represent a viable high-throughput diagnostic approach.
REFERENCES
- 1.Barza R, Patel P, Sabatini L, Singh K. 2020. Use of a simplified sample processing step without RNA extraction for direct SARS-CoV-2 RT-PCR detection. J Clin Virol 132:104587. doi: 10.1016/j.jcv.2020.104587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hasan MR, Mirza F, Al-Hail H, Sundararaju S, Xaba T, Iqbal M, Alhussain H, Yassine HM, Perez-Lopez A, Tang P. 2020. Detection of SARS-CoV-2 RNA by direct RT-qPCR on nasopharyngeal specimens without extraction of viral RNA. PLoS One 15:e0236564. doi: 10.1371/journal.pone.0236564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Merindol N, Pepin G, Marchand C, Rheault M, Peterson C, Poirier A, Houle C, Germain H, Danylo A. 2020. SARS-CoV-2 detection by direct rRT-PCR without RNA extraction. J Clin Virol 128:104423. doi: 10.1016/j.jcv.2020.104423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bruce EA, Huang ML, Perchetti GA, Tighe S, Laaguiby P, Hoffman JJ, Gerrard DL, Nalla AK, Wei Y, Greninger AL, Diehl SA, Shirley DJ, Leonard DGB, Huston CD, Kirkpatrick BD, Dragon JA, Crothers JW, Jerome KR, Botten JW. 2020. Direct RT-qPCR detection of SARS-CoV-2 RNA from patient nasopharyngeal swabs without an RNA extraction step. PLoS Biol 18:e3000896. doi: 10.1371/journal.pbio.3000896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wyllie AL, Fournier J, Casanovas-Massana A, Campbell M, Tokuyama M, Vijayakumar P, Warren JL, Geng B, Muenker MC, Moore AJ, Vogels CBF, Petrone ME, Ott IM, Lu P, Venkataraman A, Lu-Culligan A, Klein J, Earnest R, Simonov M, Datta R, Handoko R, Naushad N, Sewanan LR, Valdez J, White EB, Lapidus S, Kalinich CC, Jiang X, Kim DJ, Kudo E, Linehan M, Mao T, Moriyama M, Oh JE, Park A, Silva J, Song E, Takahashi T, Taura M, Weizman OE, Wong P, Yang Y, Bermejo S, Odio CD, Omer SB, Dela Cruz CS, Farhadian S, Martinello RA, Iwasaki A, Grubaugh ND, Ko AI. 2020. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N Engl J Med 383:1283–1286. doi: 10.1056/NEJMc2016359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Smyrlaki I, Ekman M, Lentini A, Rufino de Sousa N, Papanicolaou N, Vondracek M, Aarum J, Safari H, Muradrasoli S, Rothfuchs AG, Albert J, Hogberg B, Reinius B. 2020. Massive and rapid COVID-19 testing is feasible by extraction-free SARS-CoV-2 RT-PCR. Nat Commun 11:4812. doi: 10.1038/s41467-020-18611-5. [DOI] [PMC free article] [PubMed] [Google Scholar]