Abstract
Alzheimer’s disease (AD) is the most common form of dementia and is characterized by irreversible and progressive neurodegeneration. Cholinergic dysfunction has been reported in AD, and several cholinesterase inhibitors, including natural compounds and synthetic analogs, have been developed to treat the disease. However, there is currently no treatment for AD, as most drug-like compounds have failed in clinical trials. Acetylcholinesterase (AChE) is the target of most drugs used commercially to treat AD. This work focused on screening natural compounds obtained from the ZINC database (224, 205 compounds) against AChE to identify those possibly capable of enabling the management of AD. Indirubin and dehydroevodiamine were the best potential AChE inhibitors with free binding energies of −10.03 and −9.00 kcal/mol, respectively. The key residue (His447) of the active site of AChE was found to participate in complex interactions with these two molecules. Six H-bonds were involved in the ‘indirubin–AChE’ interaction and three H-bonds in the ‘dehydroevodiamine–AChE’ interaction. These compounds were predicted to cross the blood–brain barrier (BBB) and to exhibit high levels of intestinal absorption. Furthermore, ‘indirubin–AChE’ and ‘dehydroevodiamine–AChE’ complexes were found to be stable, as determined by root mean square deviation (RMSD) during a 50 ns molecular dynamics simulation study. Based on the free binding energies and stabilities obtained by simulation studies, we recommend that experimental studies be undertaken on indirubin and dehydroevodiamine with a view towards their potential use as treatments for AD.
Keywords: Alzheimer disease, molecular dynamics, neurotransmitters, ZINC database, pharmacokinetic
1. Introduction
Alzheimer’s disease (AD) is a typical neurodegenerative condition that affects more than 46 million individuals globally [1,2,3]. AD is a deadly neurodegenerative disease for which no preventative treatment is available [4,5,6,7]. In the US, the prevalence of AD is expected to increase three-fold by 2050 [8], and 121,404 deaths were attributed to AD during 2017, which made it the sixth most common cause of death and the fifth most common among Americans aged ≥65 years. In 2018, more than 16 million Americans and unpaid caregivers spent an estimated 18.5 billion hours caring for individuals with AD or another form of dementia. In 2019, total expenditure on health, long-term care, and hospice facilities for people age ≥65 years with dementia was projected to be $290 billion [9]. Disturbances in cholinergic neurotransmission contribute to the memory weakness that characterizes AD. Currently accessible treatments target cholinergic synapses to increase synaptic levels of acetylcholine (ACh) and relieve memory deficits [10]. The brain cholinergic neurotransmitter framework is critical for processing cognitive data [11], and neurotransmitters are fundamental components of the machinery articulated by neurons [12]. Cholinesterase inhibitors (ChEIs) have been approved for the symptomatic treatment of AD [13]. Tacrine was the first ChEI approved by the Food and Drug Administration for the treatment of AD [14,15], but unfortunately, its side effects, which include gastrointestinal problems and hepatotoxicity, have limited its use [16].
Plants have been known to have medicinal properties. Plant-derived drugs have been used to treat a variety of pathological disorders. Recent developments in computational methods have opened up new possibilities for processing complex natural products and using their structures to develop novel drugs [17]. Numerous natural and synthetic compounds have been investigated for effectiveness against AD [18,19,20]. In particular, indirubin plays significant roles in the treatments of numerous chronic sicknesses and has been reported to exhibit strong anti-inflammatory effects and antileukemic efficacy in chronic myelocytic leukemia [21,22]. Indirubin is a specific cyclin-dependent kinase (CDK) inhibitor that suppresses the activities of CDK1, CDK2, and CDK5 [23] and inhibits several eukaryotic cell-signaling pathways [24]. Dehydroevodiamine is a bioactive component of the Chinese herbal drug [25] and is used to treat cardiovascular and neuropharmacological diseases [26]. The goal of this research was to screen a wide range of natural compounds for anti-Alzheimer efficacy using an in silico approach with focus on AChE.
2. Results and Discussion
In this study, we selected the most active natural compounds from a library of 224, 205 compounds in the ZINC database. Of these, seven compounds were found to have greater binding energy (>−7.5 kcal/mol) with AChE than tacrine and to possess drug-like properties. Indirubin and dehydroevodiamine were focused on because of their higher free energy of binding with AChE receptor, as determined by pharmacokinetic analysis.
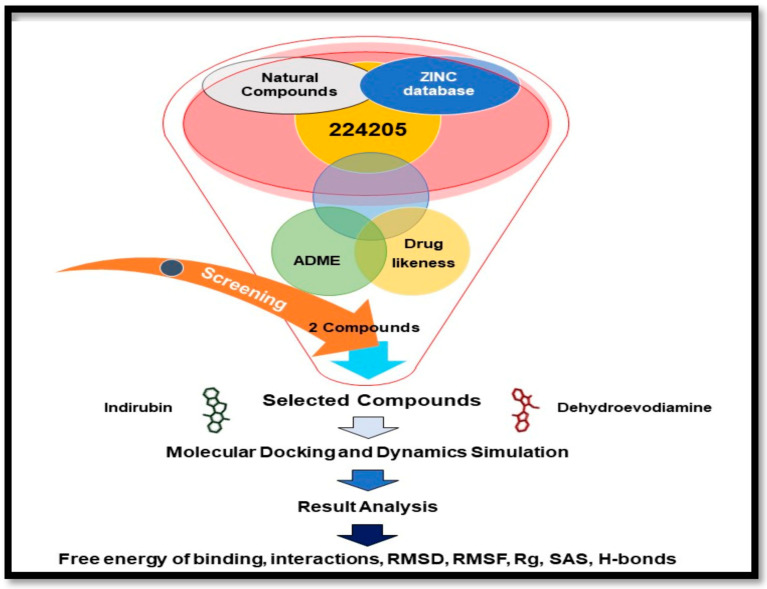
A schematic of the screening process used is shown in Figure 1.
Figure 1.
Procedure used for the structure-based virtual screening of the initially identified 224, 205 compounds.
The Lead-likeness properties and Lipinski of the top seven selected compounds are provided in Table 1. The pharmacokinetic parameters of indirubin and dehydroevodiamine were checked and are detailed in Table 2. The values shown lie in acceptable ranges of Lead-likeness and Lipinski drug discovery pipeline rules.
Table 1.
Drug-like properties of top seven selected compounds.
| Compounds | ZINC ID | Molecular Weight | Lead-likeness Violations | Lipinski Violations | Binding Energy (kcal/mol) |
|---|---|---|---|---|---|
| Coronopilin | ZINC4026171 | 264.14 | 0 | 0 | −7.94 |
| Rutaecarpine | ZINC898237 | 287.11 | 0 | 0 | −8.09 |
| Chelerythrine | ZINC3872044 | 348.12 | 1 | 0 | −8.13 |
| Chelidonine | ZINC30727894 | 353.13 | 1 | 0 | −8.08 |
| Epiberberine | ZINC6017816 | 336.12 | 1 | 0 | −7.54 |
| Indirubin | ZINC13597821 | 262.07 | 0 | 0 | −10.03 |
| Dehydroevodiamine | ZINC13434330 | 301.12 | 0 | 0 | −9.00 |
| Tacrine | ZINC19014866 | 198.12 | 1 | 0 | −5.90 |
Table 2.
Pharmacokinetics properties of selected compounds.
| Compound Properties | Indirubin | Dehydroevodiamine | |
|---|---|---|---|
| Lipophilicity | Log Po/w (iLOGP) | 2.13 | 2.88 |
| Log Po/w (XLOGP3) | 2.73 | 2.11 | |
| Log Po/w (WLOGP) | 2.81 | 0.24 | |
| Log Po/w (MLOGP) | 1.70 | 2.78 | |
| Log Po/w (SILICOS-IT) | 4.10 | 3.45 | |
| Consensus Log Po/w | 2.69 | 2.29 | |
| Water Solubility | Log S (ESOL) | −3.67 (Soluble) | −3.55 (Soluble) |
| Log S (Ali) | −3.76 (Soluble) | −2.57 (Soluble) | |
| Log S (SILICOS-IT) | −5.70 (Moderately soluble) | −5.60 (Moderately soluble) | |
| Pharmacokinetics | Gastrointestinal absorption |
High | High |
| Blood–brain barrier permeant |
Yes | No | |
| P-gp substrate | No | No | |
| CYP1A2 inhibitor | Yes | Yes | |
| CYP2C19 inhibitor | No | No | |
| CYP2C9 inhibitor | No | No | |
| CYP2D6 inhibitor | Yes | No | |
| CYP3A4 inhibitor | Yes | Yes | |
| Log Kp (skin permeation) | −5.96 cm/s | −6.64 cm/s | |
| Druglikeness | Lipinski | Yes; 0 violation | Yes; 0 violation |
| Ghose | Yes | Yes | |
| Veber | Yes | Yes | |
| Egan | Yes | Yes | |
| Muegge | Yes | Yes | |
| Bioavailability Score | 0.55 | 0.55 | |
| Medicinal Chemistry | PAINS | 0 alert | 0 alert |
| Brenk | 0 alert | 0 alert | |
| Lead-likeness | Yes | Yes | |
| Synthetic accessibility | 2.84 | 3.83 | |
Free ADME-Tox Filtering Tool (FAF-Drugs4) [27] on the RPBS platform managed by the Mobyle Portal [28] was used to determine the properties of indirubin and dehydroevodiamine. In a recent analysis, it was reported that over 90% of candidate drug failures were due to hepatotoxicity and cardiovascular complications [29], and thus, in silico approaches have been utilized to predict some key absorption, distribution, metabolism, excretion and toxicity (ADME-Tox) properties [30] (Table 2). Drugs that affect the central nervous system (CNS) may serve as substrates, inhibitors, or inducers of enzymes that are encoded by metabolic genes. The drugs used for CNS are about 90% utilized by CYP enzymes as major metabolic pathways, and CNS drugs are major substrates of CYP3A4 [31]. The blood–brain barrier (BBB) is an important barrier for preserving the brain microenvironment and protecting the CNS from blood-borne neurotoxins. However, the BBB restricts medication therapeutic effectiveness in the CNS, making it difficult to manage brain diseases [32].
ADMET and other properties [33,34] (drug-like filter area, rule of 5, and toxicity) of indirubin and dehydroevodiamine are shown in Figure 2 and Figure 3, respectively, and are listed in Table 2.
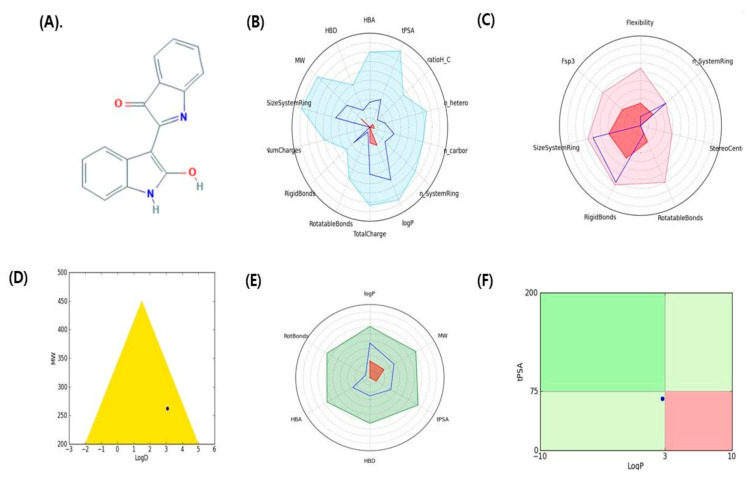
Figure 2.
Graphical representation of pharmacokinetic properties of indirubin. (A) 2D structure indirubin, (B) Physicochemical filter positioning of ligand, (C) Complexity of ligand, (D) Golden triangle rule (dot represents the position of compound), (E) Oral absorption, (F) Pfizer rule (dot represents the position of compound).
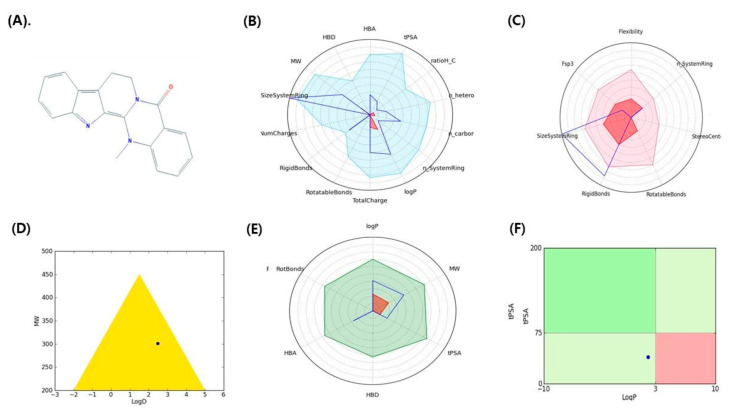
Figure 3.
Graphical representation of pharmacokinetic properties of dehydroevodiamine. (A) 2D structure dehydroevodiamine, (B) Physicochemical filter positioning of ligand, (C) Complexity of ligand, (D) Golden triangle rule (dot represents the position of compound), (E) Oral absorption, (F) Pfizer rule (dot represents the position of compound).
Molecular interaction studies are used to determine the binding orientations of small molecules and receptors and predict affinities and activities [35]. In the present study, the catalytic anionic site of human AChE was found to interact with indirubin through the amino acid residues Trp86, Tyr124, Tyr133, Glu202, Tyr337, Phe338, Tyr341, and His447; and with dehydroevodiamine through the amino acid residues Gln71, Tyr72, Asp74, Trp86, Asn87, Gly120, Gly121, Tyr124, Ser125, Gly126, Tyr133, Glu202, Ser203, Phe297, Tyr337, Phe338, Tyr341, His447, Gly448, and Ile451. Tacrine was also found to interact through Trp86, Gly120, Gly121, Gly122, Ser125, Gly126, Leu130, Tyr133, Glu202, Ser203, and Phe338. Interacting amino acid residues are presented in Table 3.
Table 3.
Interacting amino acid residues and hydrogen bonds form between selected compounds with AChE.
| Compounds | Hydrogen Bond | Hydrogen Bond Distance | Interacting Amino Acid Residues |
|---|---|---|---|
| Indirubin | Tyr133:OH-UNK1:C3 Tyr337:OH-UNK1:N12 His447:CD2-UNK1:O10 Tyr124:OH-UNK1 UNK1:H25-Trp86 UNK1:H26-Tyr337 |
3.279086 2.756860 3.239033 3.208647 4.083150 3.943169 |
Trp86, Tyr124, Tyr133, Glu202, Tyr337, Phe338, Tyr341, and His447 |
| Dehydroevodiamine | Tyr133:OH-UNK1:O6 UNK1:O10-His447:NE2 Tyr124:OH-UNK1 |
3.093207 3.296732 3.212351 |
Gln71, Tyr72, Asp74, Trp86, Asn87, Gly120, Gly121, Tyr124, Ser125, Gly126, Tyr133, Glu202, Ser203, Phe297, Tyr337, Phe338, Tyr341, His447, Gly448, and Ile451 |
| Tacrine | UNK1:H29-Glu202:OE1 | 2.30175 | Trp86, Gly120, Gly121, Gly122, Ser125, Gly126, Leu130, Tyr133, Glu202, Ser203, and Phe338 |
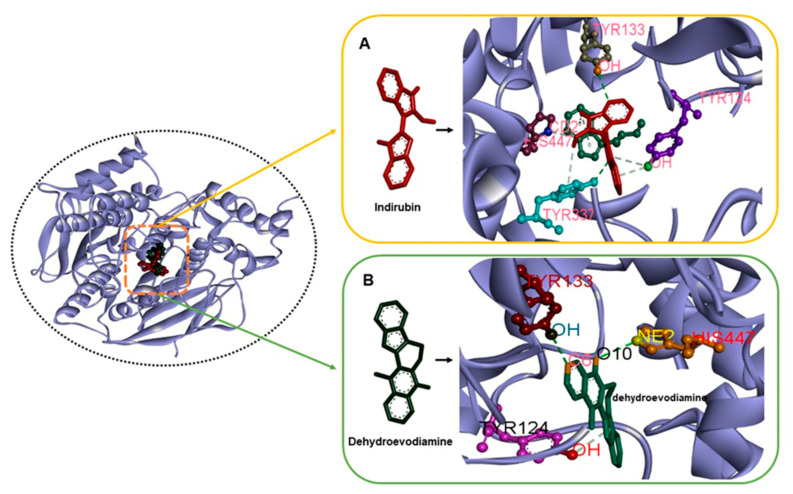
Based on these results, we concluded that, of the three residues that constitute the catalytic triad, namely, Ser203, His447, and Glu334 [36,37,38], His447 of AChE interacts with indirubin and dehydroevodiamine. We further investigated these interactions in the hope that this might aid the design of AChE inhibitors. The binding free energies and estimated inhibition constants of indirubin–AChE, dehydroevodiamine–AChE, and tacrine–AChE interactions were determined to be −10.03 kcal/mol and 4.36 μM, −9.00 kcal/mol and 4.25 μM, and −5.90 kcal/mol and 47.32 μM, respectively [39]. These results predicted that indirubin and dehydroevodiamine are more efficient AChE inhibitors than tacrine. Indirubin formed six hydrogen bonds (Tyr133:OH-UNK1:C3; Tyr337:OH-UNK1:N12; His447:CD2-UNK1:O10; Tyr124:OH-UNK1; UNK1:H25-Trp86; UNK1:H26-Tyr337), and dehydroevodiamine formed three hydrogen bonds (Tyr133:OH-UNK1:O6; UNK1:O10-His447:NE2; Tyr124:OH -UNK1) (Figure 4).
Figure 4.
The complex structures of ACHE with indirubin and dehydroevodiamine (A) H-bond interactions in AChE–indirubin. (B) H-bond interactions in AChE–dehydroevodiamine.
On the other hand, tacrine formed only one hydrogen bond with Glu202 (UNK1:H29-Glu202:OE1) of AChE. These findings suggest that indirubin and dehydroevodiamine form more stable complexes with AChE than tacrine [40,41]. The lengths of these hydrogen bonds are provided in Table 3, and Van der Waals’, hydrogen bond, desolvation, and intermolecular energies are listed in Table 4.
Table 4.
Different energies obtained by docking between selected compounds and AChE.
| Compounds | Binding Energy (kcal/mol) |
Inhibition Constant (μM) |
Intermolecular Energy | Van der Waals’, ‘Hydrogen Bond’ and ‘Desolvation Energy’ | Electrostatic Energy |
|---|---|---|---|---|---|
| Indirubin | −10.03 | 4.36 | −7.31 | −7.33 | −0.02 |
| Dehydroevodiamine | −9.00 | 4.25 | −7.50 | −7.46 | −0.05 |
| Tacrine | −5.90 | 47.32 | −6.17 | −6.11 | −0.06 |
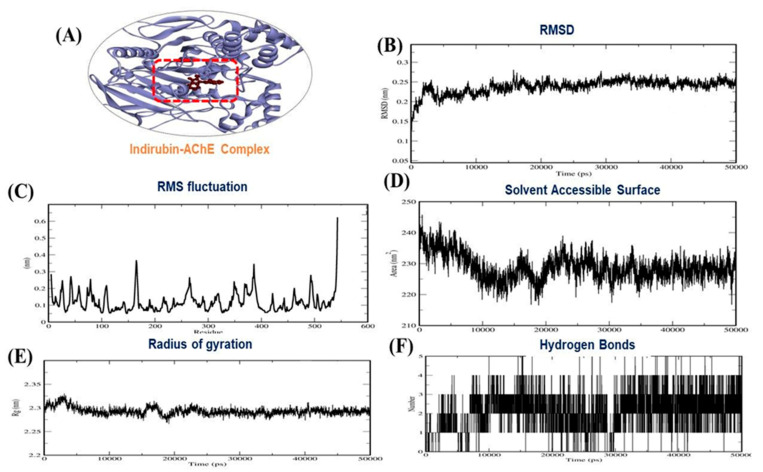
Several reports have checked best docking scores complex by molecular dynamics (MD) simulation [42,43,44,45,46]. Thus, we performed 50 ns MD simulations on indirubin and dehydroevodiamine to evaluate the stabilities of complexes and investigate possible ligand binding modes. Docked complexes in best docking conformations of AChE with indirubin and dehydroevodiamine were subjected to MD simulation to confirm their stabilities. MD trajectories were analyzed in a time-dependent manner and included root mean square deviations (RMSDs) and radii of gyration (Rg) of all backbone atoms. RMSDs of protein backbone atoms were plotted versus time to check stabilities throughout MD simulations. Rg values describe molecular dimensions, which are calculated as average root squares of distances between residues and the complex centers of gravity and provide measures of levels of molecular compaction. The plots of RMSD, RMSF, Rg, H-bond, and solvent-accessible surface area (SASA) for AChE–indirubin and –dehydroevodiamine complexes for 50 ns at 300 K are shown in Figure 5 and Figure 6.
Figure 5.
(A) 3D interaction of AChE–indirubin, (B) RMSD plot, (C) RMSF plot, (D), solvent accessible surface area (E) Radius of gyration plot and (F) H-bond interaction for AChE–indirubin complex.
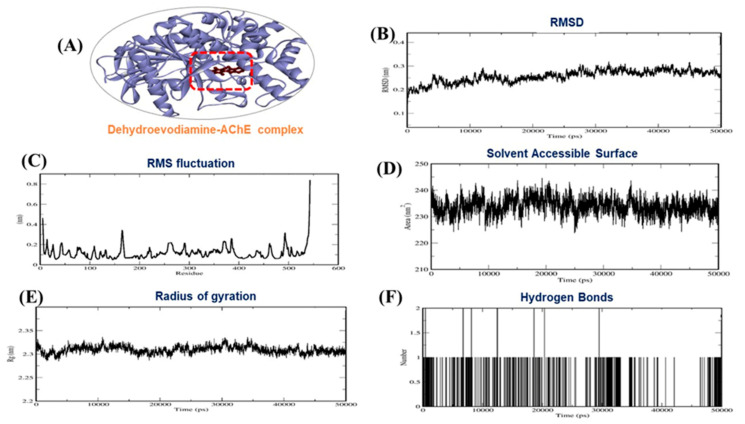
Figure 6.
(A) 3D interaction of AChE–dehydroevodiamine, (B) RMSD plot, (C) RMSF plot, (D) solvent accessible surface area (E) Radius of gyration plot and (F) H-bond interaction for AChE–dehydroevodiamine complex.
Based on MD simulation studies, the binding free energies of indirubin and dehydroevodiamine against AChE were −146.00 and −126.70 kJ/mol, respectively. Other energy values for the complexes are provided in Table 5. Summarizing, our results show AChE–indirubin and –dehydroevodiamine complexes maintained structural integrity during MD simulations.
Table 5.
Different energies obtained by MD between selected compounds and AChE.
| S.No. | Energy (kJ/mol) | ‘Indirubin–AChE’ Complex | ‘Dehydroevodiamine–AChE’ Complex |
|---|---|---|---|
| 1. | Binding energy | −146.0+/−9.5 | −126.7+/−10.9 |
| 2. | Van der Waal energy | −177.0+/−8.4 | −159.4+/−9.1 |
| 3. | Electrostatic energy | −55.4+/−8.4 | −2.4+/−3.3 |
| 4. | Polar solvation energy | 101.8+/−10.1 | 50.4+/−7.6 |
| 5. | SASA energy | −15.4+/−0.71 | −15.3+/−0.84 |
3. Materials and Methods
3.1. Compound Library Preparation
Natural compounds were retrieved from the ZINC database (https://zinc.docking.org, accessed on 15 April 2021) by selecting ‘natural_products’ as a ‘subset’ under the ‘substances’ category. A total of 224,205 compounds were retrieved, downloaded in .sdf format, imported into Discovery Studio 2020, and processed using the ligand preparation tool.
3.2. Preparation of Receptor
A 3D structure of human AChE (PDB ID: 3LII) was obtained using RCSB-PDB (www.rcsb.org, accessed on 15 April 2021). The PDB file of AChE was cleaned, and heteroatoms were removed manually because they were non-standard. These heteroatoms included the atomic coordinates of cofactors, coenzymes, prosthetic groups, metal ions, sugars, drugs, peptides, heavy-atom derivatives, non-standard amino acid residues/nucleotides, and water molecules [47].
3.3. Structure-Based Virtual Screening
The prepared compound library was screened against the active site of AChE using AutoDock Vina (version 1.1.2) program. The compound library was converted from .sdf to.pdbqt format using the Open Babel tool. The XYZ axes of AChE were set as 90.81, 83.98, and −8.04, respectively. Top-ranked compounds obtained from AutoDock Vina were also processed for ADMET and drug-likeness filtration, and top-ranked compounds were subjected to in-depth docking analysis using AutoDock 4.2.
3.4. Drug-Likeness Study and ADMET Profiling
The drug-likeness properties were employed to the selected ligands. It included molecular mass (< = 500 Dalton), high lipophilicity (Log p < = 5), H-bond donors (< = 5), and H-bond acceptors (< = 10) [48,49].
3.5. Docking Simulations
The molecular interaction study was performed using Autodock version 4.2 suite and the Cygwin interface [50,51]. The docking protocol included receptor preparation, ligand preparation, and the plotting of a grid-box based on selected interaction sites around AChE (dimension 60 × 60 × 60) with grid centers at 90.81, 83.98, and −8.04 [52]. The Lamarckian genetic algorithm was applied at AChE with indirubin and dehydroevodiamine for adaptable docking calculations [53]. The search parameter was set to bind the AChE with indirubin and dehydroevodiamine. Docking experiments were performed using automated program (.exe) files of Autogrid and Autodock, which resulted in .glg and .dlg files. Docking outcomes in .dlg format were analyzed to obtain binding energies (kcal/mol) and inhibition constants (Ki value-μM/nM).
3.6. Molecular Dynamics
GROMACS 5.1.2 molecular dynamics (MD) [54] was used to analyze the structural stabilities of AChE–indirubin and –dehydroevodiamine complexes. Ligand topologies were generated using the PRODRG server [55]. In addition, complexes were solvated using the SPC216 water model in a triclinic-box of dimension 1.0 nm. Bond angles were constrained using LINCS [56]. Van der Waals and electrostatic long-range interactions were applied using fast Particle-Mesh Weald electrostatics (PME) [57]. Additionally, the Parrinello–Rahman [58] method was used to regulate pressure, and the modified weak Coupling Berendsen thermostat and Vrescale algorithm were used to regulate system temperature. The (constant number of particles, volume, and temperature) NVT and constant number of particles, pressure, and temperature (NPT) were used to monitor equilibration status. Finally, systems were simulated [59] using 50 ns MD runs. Binding energy calculations were performed [60].
4. Conclusions
Many compounds have already been reported to have potential activity against AChE, but none have been granted FDA approval due to failure to cross the BBB, toxicity, and other shortcomings. In terms of toxicity, natural compounds are generally accepted to be ‘safe’. Our in silico analysis predicted that indirubin and dehydroevodiamine both bind strongly to the active site residues of AChE and follow the drug-like properties. Further experimental evaluations of indirubin and dehydroevodiamine may eventually lead to exciting alternative AD therapies.
Abbreviations
| AD | Alzheimer’s disease |
| AChE | Acetylcholinesterase enzyme |
| ChEI | Cholinesterase inhibitor |
| MD | Molecular dynamics |
| RMSD | Root mean square deviation |
| RMSF | Root mean square fluctuation |
| Rg | Radius of gyration |
| BBB | Blood–brain barrier |
| CNS | Central nervous system |
Author Contributions
Conceptualization and original draft; S.S.A., K.A., E.-J.L. and I.C. methodology; S.S.A., M.B.K., K.A., J.-H.L. and S.S., review, editing and funding; I.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Korean Ministry of Education (Grant no. 2020R1A6A1A03044512) and by the NRF funded by the Korean government (MSIP: Grant Nos. NRF-2021R1A2C2004177 and NRF-2019R1C1C1006542).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors have no conflict of interest to declare.
Sample Availability
Not applicable.
Footnotes
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Ahmad S.S., Khan H., Danish Rizvi S.M., Ansari S.A., Ullah R., Rastrelli L., Mahmood H.M., Siddiqui M.H. Computational Study of Natural Compounds for the Clearance of Amyloid-Betaeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease. Molecules. 2019;24:3233. doi: 10.3390/molecules24183233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kandimalla R., Reddy P.H. Therapeutics of Neurotransmitters in Alzheimer’s Disease. J. Alzheimers Dis. 2017;57:1049–1069. doi: 10.3233/JAD-161118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Jan A.T., Azam M., Rahman S., Almigeiti A.M.S., Choi D.H., Lee E.J., Haq Q.M.R., Choi I. Perspective Insights into Disease Progression, Diagnostics, and Therapeutic Approaches in Alzheimer’s Disease: A Judicious Update. Front Aging Neurosci. 2017;9:356. doi: 10.3389/fnagi.2017.00356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jack C.R., Jr., Albert M.S., Knopman D.S., McKhann G.M., Sperling R.A., Carrillo M.C., Thies B., Phelps C.H. Introduction to the recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011;7:257–262. doi: 10.1016/j.jalz.2011.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dunkin J.J., Anderson-Hanley C. Dementia caregiver burden: A review of the literature and guidelines for assessment and intervention. Neurology. 1998;51:S53–S60. doi: 10.1212/WNL.51.1_Suppl_1.S53. [DOI] [PubMed] [Google Scholar]
- 6.Hurd M.D., Martorell P., Delavande A., Mullen K.J., Langa K.M. Monetary costs of dementia in the United States. N Engl. J. Med. 2013;368:1326–1334. doi: 10.1056/NEJMsa1204629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Langa K.M., Chernew M.E., Kabeto M.U., Herzog A.R., Ofstedal M.B., Willis R.J., Wallace R.B., Mucha L.M., Straus W.L., Fendrick A.M. National estimates of the quantity and cost of informal caregiving for the elderly with dementia. J. Gen. Intern. Med. 2001;16:770–778. doi: 10.1111/j.1525-1497.2001.10123.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hebert L.E., Weuve J., Scherr P.A., Evans D.A. Alzheimer disease in the United States (2010–2050) estimated using the 2010 census. Neurology. 2013;80:1778–1783. doi: 10.1212/WNL.0b013e31828726f5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Association A.s. 2019 Alzheimer’s disease facts and figures. Alzheimer’s Dement. 2019;15:321–387. doi: 10.1016/j.jalz.2019.01.010. [DOI] [Google Scholar]
- 10.Bales K.R., Tzavara E.T., Wu S., Wade M.R., Bymaster F.P., Paul S.M., Nomikos G.G. Cholinergic dysfunction in a mouse model of Alzheimer disease is reversed by an anti-A beta antibody. J. Clin. Investig. 2006;116:825–832. doi: 10.1172/JCI27120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Perry E., Walker M., Grace J., Perry R. Acetylcholine in mind: A neurotransmitter correlate of consciousness? Trends Neurosci. 1999;22:273–280. doi: 10.1016/S0166-2236(98)01361-7. [DOI] [PubMed] [Google Scholar]
- 12.Spitzer N.C. Activity-dependent neurotransmitter respecification. Nat Rev Neurosci. 2012;13:94–106. doi: 10.1038/nrn3154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Han J.Y., Besser L.M., Xiong C., Kukull W.A., Morris J.C. Cholinesterase Inhibitors May Not Benefit Mild Cognitive Impairment and Mild Alzheimer Disease Dementia. Alzheimer Dis. Assoc. Disord. 2019;33:87–94. doi: 10.1097/WAD.0000000000000291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Summers W.K. Tacrine, and Alzheimer’s treatments. J. Alzheimers Dis. 2006;9:439–445. doi: 10.3233/JAD-2006-9S350. [DOI] [PubMed] [Google Scholar]
- 15.Horak M., Holubova K., Nepovimova E., Krusek J., Kaniakova M., Korabecny J., Vyklicky L., Kuca K., Stuchlik A., Ricny J., et al. The pharmacology of tacrine at N-methyl-d-aspartate receptors. Prog. Neuropsychopharmacol. Biol. Psychiatry. 2017;75:54–62. doi: 10.1016/j.pnpbp.2017.01.003. [DOI] [PubMed] [Google Scholar]
- 16.Korabecny J., Spilovska K., Benek O., Musilek K., Soukup O., Kuca K. [Tacrine and its derivatives in the therapy of Alzheimers disease] Ceska Slov. Farm. 2012;61:210–221. [PubMed] [Google Scholar]
- 17.Thomford N.E., Senthebane D.A., Rowe A., Munro D., Seele P., Maroyi A., Dzobo K. Natural Products for Drug Discovery in the 21st Century: Innovations for Novel Drug Discovery. Int. J. Mol. Sci. 2018;19:1578. doi: 10.3390/ijms19061578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Dubey T., Chinnathambi S. Brahmi (Bacopa monnieri): An ayurvedic herb against the Alzheimer’s disease. Arch. Biochem. Biophys. 2019;676:108153. doi: 10.1016/j.abb.2019.108153. [DOI] [PubMed] [Google Scholar]
- 19.Ahmad S.S., Sinha M., Ahmad K., Khalid M., Choi I. Study of Caspase 8 Inhibition for the Management of Alzheimer’s Disease: A Molecular Docking and Dynamics Simulation. Molecules. 2020;25:2071. doi: 10.3390/molecules25092071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ahmad S.S., Khalid M., Kamal M.A., Younis K. Study of Nutraceuticals and Phytochemicals for the Management of Alzheimer’s Disease: A Review. Curr. Neuropharmacol. 2021 doi: 10.2174/1570159X19666210215122333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lai J.L., Liu Y.H., Liu C., Qi M.P., Liu R.N., Zhu X.F., Zhou Q.G., Chen Y.Y., Guo A.Z., Hu C.M. Indirubin Inhibits LPS-Induced Inflammation via TLR4 Abrogation Mediated by the NF-kB and MAPK Signaling Pathways. Inflammation. 2017;40:1–12. doi: 10.1007/s10753-016-0447-7. [DOI] [PubMed] [Google Scholar]
- 22.Eisenbrand G., Hippe F., Jakobs S., Muehlbeyer S. Molecular mechanisms of indirubin and its derivatives: Novel anticancer molecules with their origin in traditional Chinese phytomedicine. J. Cancer Res. Clin. Oncol. 2004;130:627–635. doi: 10.1007/s00432-004-0579-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Absalon S., Kochanek D.M., Raghavan V., Krichevsky A.M. MiR-26b, upregulated in Alzheimer’s disease, activates cell cycle entry, tau-phosphorylation, and apoptosis in postmitotic neurons. J. Neurosci. 2013;33:14645–14659. doi: 10.1523/JNEUROSCI.1327-13.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wongsaroj L., Sallabhan R., Dubbs J.M., Mongkolsuk S., Loprasert S. Cloning of Toluene 4-Monooxygenase Genes and Application of Two-Phase System to the Production of the Anticancer Agent, Indirubin. Mol. Biotechnol. 2015;57:720–726. doi: 10.1007/s12033-015-9863-4. [DOI] [PubMed] [Google Scholar]
- 25.Loh S.H., Tsai Y.T., Lee C.Y., Chang C.Y., Tsai C.S., Cheng T.H., Lin C.I. Antiarrhythmic effects of dehydroevodiamine in isolated human myocardium and cardiomyocytes. J. Ethnopharmacol. 2014;153:753–762. doi: 10.1016/j.jep.2014.03.043. [DOI] [PubMed] [Google Scholar]
- 26.Schramm A., Hamburger M. Gram-scale purification of dehydroevodiamine from Evodia rutaecarpa fruits, and a procedure for selective removal of quaternary indoloquinazoline alkaloids from Evodia extracts. Fitoterapia. 2014;94:127–133. doi: 10.1016/j.fitote.2014.02.005. [DOI] [PubMed] [Google Scholar]
- 27.Hinsen K. Analysis of domain motions by approximate normal mode calculations. Proteins. 1998;33:417–429. doi: 10.1002/(SICI)1097-0134(19981115)33:3<417::AID-PROT10>3.0.CO;2-8. [DOI] [PubMed] [Google Scholar]
- 28.Tetko I.V., Gasteiger J., Todeschini R., Mauri A., Livingstone D., Ertl P., Palyulin V.A., Radchenko E.V., Zefirov N.S., Makarenko A.S., et al. Virtual computational chemistry laboratory—Design and description. J. Comput. Aided Mol. Des. 2005;19:453–463. doi: 10.1007/s10822-005-8694-y. [DOI] [PubMed] [Google Scholar]
- 29.Schuster D., Laggner C., Langer T. Why drugs fail—A study on side effects in new chemical entities. Curr. Pharm. Des. 2005;11:3545–3559. doi: 10.2174/138161205774414510. [DOI] [PubMed] [Google Scholar]
- 30.Di L., Kerns E.H. Drug-Like Properties: Concepts, Structure Design and Methods from ADME to Toxicity Optimization. Academic Press; Cambridge, MA, USA: 2015. [Google Scholar]
- 31.Cacabelos R. Pharmacogenomics of Cognitive Dysfunction and Neuropsychiatric Disorders in Dementia. Int. J. Mol. Sci. 2020;21:3059. doi: 10.3390/ijms21093059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Fu B.M. Transport Across the Blood-Brain Barrier. Adv. Exp. Med. Biol. 2018;1097:235–259. doi: 10.1007/978-3-319-96445-4_13. [DOI] [PubMed] [Google Scholar]
- 33.Daina A., Michielin O., Zoete V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017;7:42717. doi: 10.1038/srep42717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Daina A., Michielin O., Zoete V. iLOGP: A simple, robust, and efficient description of n-octanol/water partition coefficient for drug design using the GB/SA approach. J. Chem. Inf. Model. 2014;54:3284–3301. doi: 10.1021/ci500467k. [DOI] [PubMed] [Google Scholar]
- 35.Seeliger D., de Groot B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010;24:417–422. doi: 10.1007/s10822-010-9352-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Silman I., Sussman J.L. Acetylcholinesterase: ‘classical’ and ‘non-classical’ functions and pharmacology. Curr. Opin. Pharmacol. 2005;5:293–302. doi: 10.1016/j.coph.2005.01.014. [DOI] [PubMed] [Google Scholar]
- 37.Silman I., Sussman J.L. Acetylcholinesterase: How is structure related to function? Chem. Biol. Interact. 2008;175:3–10. doi: 10.1016/j.cbi.2008.05.035. [DOI] [PubMed] [Google Scholar]
- 38.Sussman J.L., Harel M., Frolow F., Oefner C., Goldman A., Toker L., Silman I. Atomic structure of acetylcholinesterase from Torpedo californica: A prototypic acetylcholine-binding protein. Science. 1991;253:872–879. doi: 10.1126/science.1678899. [DOI] [PubMed] [Google Scholar]
- 39.Shakil S., Khan A. Interaction of 2009 CTX-M variants with drugs and inhibitors: A molecular modelling and docking study. J. Proteom. Bioinform. 2010;3:130–134. doi: 10.4172/jpb.1000131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Steiner T., Koellner G. Hydrogen bonds with pi-acceptors in proteins: Frequencies and role in stabilizing local 3D structures. J. Mol. Biol. 2001;305:535–557. doi: 10.1006/jmbi.2000.4301. [DOI] [PubMed] [Google Scholar]
- 41.Weiss M.S., Brandl M., Suhnel J., Pal D., Hilgenfeld R. More hydrogen bonds for the (structural) biologist. Trends Biochem. Sci. 2001;26:521–523. doi: 10.1016/S0968-0004(01)01935-1. [DOI] [PubMed] [Google Scholar]
- 42.Chen Y.C. Beware of docking! Trends Pharmacol. Sci. 2015;36:78–95. doi: 10.1016/j.tips.2014.12.001. [DOI] [PubMed] [Google Scholar]
- 43.Hung T.C., Lee W.Y., Chen K.B., Chan Y.C., Lee C.C., Chen C.Y. In silico investigation of traditional Chinese medicine compounds to inhibit human histone deacetylase 2 for patients with Alzheimer’s disease. Biomed. Res. Int. 2014;2014:769867. doi: 10.1155/2014/769867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Chen K.C., Chen H.Y., Chen C.Y. Potential Protein Phosphatase 2A Agents from Traditional Chinese Medicine against Cancer. Evid Based Complement Alternat. Med. 2014;2014:436863. doi: 10.1155/2014/436863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Huang H.J., Lee C.C., Chen C.Y. Lead discovery for Alzheimer’s disease related target protein RbAp48 from traditional Chinese medicine. Biomed. Res. Int. 2014;2014:764946. doi: 10.1155/2014/764946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Baig M.H., Ahmad K., Roy S., Ashraf J.M., Adil M., Siddiqui M.H., Khan S., Kamal M.A., Provaznik I., Choi I. Computer Aided Drug Design: Success and Limitations. Curr. Pharm. Des. 2016;22:572–581. doi: 10.2174/1381612822666151125000550. [DOI] [PubMed] [Google Scholar]
- 47.Guruprasad K., Savitha S., Babu A.V. Computational tools for the analysis of heteroatom groups and their neighbours in protein tertiary structure. Int. J. Biol. Macromol. 2005;37:35–41. doi: 10.1016/j.ijbiomac.2005.08.002. [DOI] [PubMed] [Google Scholar]
- 48.Lipinski C.A. Lead- and drug-like compounds: The rule-of-five revolution. Drug Discov. Today Technol. 2004;1:337–341. doi: 10.1016/j.ddtec.2004.11.007. [DOI] [PubMed] [Google Scholar]
- 49.Rehman A., Akhtar S., Siddiqui M.H., Sayeed U., Ahmad S.S., Arif J.M., Khan M.K. Identification of potential leads against 4-hydroxytetrahydrodipicolinate synthase from Mycobacterium tuberculosis. Bioinformation. 2016;12:400–407. doi: 10.6026/97320630012400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Morris G.M., Goodsell D.S., Halliday R.S., Huey R., Hart W.E., Belew R.K., Olson A.J. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 1998;19:1639–1662. doi: 10.1002/(SICI)1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B. [DOI] [Google Scholar]
- 51.Rarey M., Kramer B., Lengauer T., Klebe G. A fast flexible docking method using an incremental construction algorithm. J. Mol. Biol. 1996;261:470–489. doi: 10.1006/jmbi.1996.0477. [DOI] [PubMed] [Google Scholar]
- 52.Alam A., Shaikh S., Ahmad S.S., Ansari M.A., Shakil S., Rizvi S.M., Shakil S., Imran M., Haneef M., Abuzenadah A.M., et al. Molecular interaction of human brain acetylcholinesterase with a natural inhibitor huperzine-B: An enzoinformatics approach. CNS Neurol. Disord. Drug Targets. 2014;13:487–490. doi: 10.2174/18715273113126660163. [DOI] [PubMed] [Google Scholar]
- 53.Goodsell D.S., Morris G.M., Olson A.J. Automated docking of flexible ligands: Applications of AutoDock. J. Mol. Recognit. 1996;9:1–5. doi: 10.1002/(SICI)1099-1352(199601)9:1<1::AID-JMR241>3.0.CO;2-6. [DOI] [PubMed] [Google Scholar]
- 54.Abraham M., van der Spoel D., Lindahl E., Hess B., atGd t. GROMACS User Manual Version 5.1.2. GROMACS Development Team; Uppsala, Sweden: 2016. [Google Scholar]
- 55.Schuttelkopf A.W., van Aalten D.M. PRODRG: A tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 2004;60:1355–1363. doi: 10.1107/S0907444904011679. [DOI] [PubMed] [Google Scholar]
- 56.Hess B., Bekker H., Berendsen H.J., Fraaije J.G. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997;18:1463–1472. doi: 10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H. [DOI] [Google Scholar]
- 57.Bussi G., Donadio D., Parrinello M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007;126:014101. doi: 10.1063/1.2408420. [DOI] [PubMed] [Google Scholar]
- 58.Parrinello M., Rahman A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981;52:7182–7190. doi: 10.1063/1.328693. [DOI] [Google Scholar]
- 59.Van Der Spoel D., Lindahl E., Hess B., Groenhof G., Mark A.E., Berendsen H.J. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005;26:1701–1718. doi: 10.1002/jcc.20291. [DOI] [PubMed] [Google Scholar]
- 60.Kumari R., Kumar R., Open Source Drug Discovery C., Lynn A. g_mmpbsa—A GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. Model. 2014;54:1951–1962. doi: 10.1021/ci500020m. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.