Abstract
Dynamin 3 (DNM3) functions as a tumor suppressor in various malignancies. However, the underlying mechanism of DNM3 in cervical cancer remains to be elucidated. The present study aimed to indicate the function of DNM3 in cervical cancer. The expression of DNM3 in cervical tissues and cells was measured using bioinformatics analysis, immunohistochemistry and reverse transcription-quantitative PCR. The pcDNA3.1 plasmid was used to overexpress DNM3 in SiHa and C33A cells. The effects of DNM3 overexpression on cell proliferation, migration, invasion and apoptosis was detected by the CCK-8, clone formation, Transwell, flow cytometry and western blotting assays. In the present study, it was revealed that DNM3 was expressed at significantly low levels in cervical cancer tissues and cell lines compared with normal cervical tissues and cell lines. In addition, the low expression of DNM3 was significantly associated with high pathological grading of cervical cancer. The overall survival rate of patients with low DNM3 expression was significantly improved compared with patients with high DNM3 expression. In addition, the overexpression of DNM3 significantly inhibited the proliferation, migration and invasion of cervical carcinoma cells and induced cell apoptosis. The findings of the present study further revealed that the overexpression of DNM3 may inhibit cell migration and invasion by inactivating the epithelial mesenchymal transition process. In summary, the present study demonstrated that DNM3 was a tumor suppressor in cervical cancer progression and that it may serve as a potential prognostic biomarker for patients with cervical carcinoma.
Keywords: apoptosis, cervical cancer, dynamin 3, epithelial mesenchymal transition signaling pathway
Introduction
Cervical cancer is one of the most common malignancies, threatening women's physical and mental health worldwide, especially in developing countries (1). A total of ~570,000 cases of cervical cancer and 311,000 deaths from the disease occurred globally in 2018 (2). Cervical cancer is the fourth most common cancer among women, after breast cancer (2.1 million cases), colorectal cancer (0.8 million) and lung cancer (0.7 million) (2). China and India together contribute more than a third of the global cervical cancer burden, with 106,000 cases in China and 97,000 cases in India, and 48,000 deaths in China and 60,000 deaths in India (2). Cervical cancer is divided into squamous cell carcinoma, adenocarcinoma, adenosquamous carcinoma and neuroendorine carcinoma according to pathological characteristics (3). Almost all cervical cancer is induced by human papillomavirus infection (HPV) 16 and 18, which generally spreads through sexual activity and an impaired immune system (4,5). In recent years, regular Pap smear screening has detected cervical cancer at an earlier stage to reduce the number of associated deaths (6,7). However, advanced cervical cancer still has a high mortality rate, which results in 5-year survival rates ranging from 16% (stage IVA) to 58% (stage IIB) (8). Hence, it is of great importance to explore the molecular mechanisms, potential prediction and treatment targets of cervical cancer.
The dynamins family belongs to the guanylate triphosphatases superfamily, which is involved in the regulation of the pathogenesis of a variety of carcinomas (9). For example, dynamin-2 upregulation is associated with the progression of bladder cancer (10). Dynamin 1 and 2 can promote the proliferation and metastasis of cancer cells, whereas dynamin 3 (DNM3) is generally considered as a candidate tumor suppressor (11). The sequence analysis of the DNM3 promoter demonstrated that the DNM3 promoter is hypermethylated in hepatocellular carcinoma (HCC) tissues and DNM3 is expressed at significantly lower levels in these tissues (12). Patients with HCC with low expression of DNM3 generally have a poorer prognosis compared with those with high DNM3 expression (12). Furthermore, overexpression of DNM3 induces cell cycle arrest and reduces cell proliferation of HCC cell lines by activating p53 (13). In addition, DNM3 overexpression can activate nitric oxide (NO) synthases to generate NO, and the increased NO production can induce reactive oxygen species accumulation and activate cell apoptosis of HCC cells (14). However, the precise tumor suppressive mechanisms of DNM3 in cervical cancer remain unclear.
The present study aimed to investigate the expression pattern and biological functions of DNM3 in cervical cancer. The findings of this study may be beneficial for the prognosis or treatment of cervical cancer.
Materials and methods
Bioinformatics analysis
The mRNA expression level of DNM3 was analyzed in 306 cases of cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC) tissues and 13 cases of normal cervical tissues using the online Gene Expression Profiling Interactive Analysis (GEPIA) database (http://gepia.cancer-pku.cn/) (15). RNA sequencing data analyzed by GEPIA is generated by large consortium projects, such as The Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression project (GTex), using the output of a standard processing pipeline for RNA sequencing data (15). For survival analysis, GEPIA uses the log-rank test (Mantel-Cox test) for hypothesis evaluation (15).
Tissue collection
A total of 41 pairs of fresh cervical cancer tissues and normal adjacent tissues (<3 cm) were collected from patients diagnosed with cervical cancer who underwent surgical resection at Linyi Cancer Hospital (Linyi, China) between July 2018 and May 2019. The age range of the participants was 19–67 years, with a median age of 56 years. The exclusion criteria of the patients were: i) Had any other malignant tumors or serious lesions; and ii) received radiotherapy or any other preoperative treatment. The Ethics Committee of Linyi Cancer Hospital approved the study and all the participants provided written informed consent.
Immunohistochemistry (IHC)
Clinical samples that had been collected were fixed in formalin at 4°C for 12 h and then embedded in paraffin. Subsequently, the tissue was cut into 4-µm paraffin sections using a cryostat. The paraffin sections were dewaxed, rehydrated and blocked with 0.3% H2O2. Antigen retrieval was performed by microwaving in 0.01 M citrate buffer for 10 min. After blocking with 5% goat serum for 1 h at room temperature, the slides were exposed to rabbit anti-DNM3 (1:100; cat. no. 14737-1-AP; Proteintech Group, Inc.) for 2 h at room temperature and subsequently incubated with horseradish peroxidase (HRP)-labeled goat anti-mouse/rabbit IgG polymer (1:5,000; cat. no. 160101405L; Fuzhou Maixin Biotech Co., Ltd.) at room temperature for 20 min. Immune response was detected by the enhanced DAB chromogenic kit (cat. no. 1705252031; Fuzhou Maixin Biotech Co., Ltd.) at room temperature for 3 min, and hematoxylin was used for counterstaining at room temperature for 5 min. Finally, a light upright microscope system (Nikon Corporation) was used to obtain images.
The immunostaining score of DNM3 was the product of the score of the positive staining cells ratio (R) and the staining intensity score (S). R was divided into 4 levels: i) 0 (<5%, negative); ii) 1 (5-25%, sporadic); iii) 2 (25-50%, focus); and iv) 3 (>51%, diffuse). S was also divided into 4 levels: i) 0 (negative); ii) 1 (weak); iii) 2 (medium); and iv) 3 (strong). Finally, a total of 0–3 was considered to represent low expression and 4–9 was considered to represent high expression.
Cell culture and transfection
Human cervical cancer cell lines (Caski, SiHa, Hela and C33A) and normal cervical cell line H8 used in the present study were purchased from the ATCC. The cells were cultured in RPMI-1640 (Hyclone; GE Healthcare Life Sciences) supplemented with 10% fetal bovine serum (FBS) (Gibco; Thermo Fisher Scientific Inc.), 100 U/ml penicillin and 0.1 mg/ml streptomycin (Sigma-Aldrich; Merck KGaA). The cells were grown in a humidified incubator with 5% CO2 at 37°C. When the cells were in their exponential growth phase, they were washed with PBS and treated with trypsin-EDTA (Beijing Solarbio Science and Technology Co., Ltd.), and then the cells were resuspended and seeded into 6-well plates with a density of 5×104 for further experiments.
DNM3 cDNA was cloned into pcDNA3.1 plasmid (DNM3, 5 nM) (V87020; Thermo Fisher Scientific, Inc.). The empty pcDNA3.1 plasmid (Thermo Fisher Scientific, Inc.) was used as a negative control (NC, 5 nM). When the cell density reached 80% confluence, plasmid transfection was performed according to the manufacturer's instructions using Lipofectamine 2000® (Invitrogen; Thermo Fisher Scientific, Inc.) at 37 C for 6 h. After 6 h, medium was changed to complete medium, and the cells were cultured for a further 24 h prior to subsequent experimentation.
RNA extraction and reverse transcription-quantitative (RT-q) PCR
Total RNA from cells was extracted using a TRIzol kit (Thermo Fisher Scientific, Inc.) according to the manufacturer's instructions. The isolated RNAs were used as templates and reverse transcribed to obtain the first-strand cDNA using a HiFiScript cDNA Synthesis kit (CoWin Biosciences) according to the manufacturer's instructions. Quantitative real-time PCR analysis was performed using the Magic SYBR mixture (CoWin Biosciences). The cycling conditions were as follows: Denaturation at 95°C for 5 min, followed by 40 cycles of 95°C for 30 sec, 60°C for 45 sec and 72°C for 30 sec. The relative quantification was performed by the 2−∆∆Cq method (16). β-actin was used as the internal reference. The primer sequences were as follows: DNM3 forward, 5′-GCTCACCATCAGCAACATTGGC-3′ and reverse, 5′-CCGAACTTTCAGGTTGTCCAAGG-3′; and β-actin, forward, 5′-CCCGAGCCGTGTTTCCT-3′ and reverse, 5′-GTCCCAGTTGGTGACGATGC-3′.
Immunofluorescence assay
SiHa and C33A cells (1,000 cells/well) were plated in 6-well chamber slides. After 24 h, cells were fixed in 4% paraformaldehyde for 30 min at room temperature and permeabilized with 0.1% Triton X-100. Cells were probed for DNM3 primary antibodies (1:1,000; cat. no. 14737-1-AP; Proteintech Group, Inc.) overnight at 4°C and FITC-labeled secondary antibody (1:200; cat. no. CL488-66484; Proteintech Group, Inc.) for 1 h at room temperature. Cell nuclei were stained with DAPI (Sigma-Aldrich; Merck KGaA) for 5 min in the dark at room temperature. Images were captured using a fluorescence microscope (IX2-SL; Olympus Corporation) (magnification, ×400).
Cell Counting Kit-8 assay
Following transfection, SiHa and C33A cells were cultured for 24 h. Next, the cells were digested by trypsin-EDTA solution, resuspended and counted. The cells (1,000/well) were seeded into 96-well plates, each well containing 100 µl RPMI-1640 medium (10% FBS). The plates were cultured in a humidified incubator with 5% CO2 at 37°C. Cell viability was measured every 24 h. For each measurement, 10 µl CCK-8 (Beijing Solarbio Science and Technology Co., Ltd.) reagent was added into the wells and incubated for 1.5 h. Finally, absorbance was measured by a microplate reader at 450 nm and the growth curve drawn.
Plate clone formation assay
Following transfection for 24 h, C33A and SiHa cells were resuspended and counted. The cells were seeded into 60-mm plates (500 cells/plate) containing 5 ml RPMI-1640 medium (10% FBS) and cultured for 2 weeks at 37°C. Finally, the cells were fixed with 4% paraformaldehyde for 30 min at room temperature and then stained with 0.1% crystal violet at room temperature for 30 min. Images of visible colonies were captured with a HP Scanjet G4010 scanner (HP Spectre) and counted manually.
Transwell migration and invasion assays
Cell migration and invasion detection was performed using Transwell chambers (EMD Millipore). The transfected C33A and SiHa cells were resuspended in serum-free medium. For the invasion assay, 1×105 cells in 100 µl serum-free medium were seeded into the top chamber coated with Matrigel (BD Biosciences). Complete medium (500 µl) containing 10% FBS was added to the bottom wells. After incubating for 12 h at 37°C, the non-invaded cells in the upper chamber were removed with a cotton swab. Subsequently, the invaded cells were stained with 0.1% crystal violet at room temperature for 10 min. Five fields per filter were counted using a light microscope (Nikon TE2000; Nikon Corporation) (magnification, ×40). For the migration assay, 5×103 cells were added to the top chambers. Except for the Matrigel coating, the migration assay was the same as the invasion assay. Matrigel (10 µl) (1:8) was added to the upper chamber for 4 h at 37°C.
Analysis of apoptosis by flow cytometry
Following transfection, C33A and SiHa cells were cultured in complete medium for 24 h at 37°C, and then cultured in serum-free medium for another 24 h at 37°C to induce apoptosis. The cells were digested with EDTA-free trypsin, and collected with the 1X binding buffer. The cell density was adjusted to 1×106/ml and then 5 µl Annexin V/FITC (Beijing 4A Biotech Co., Ltd.) was added to a 100 µl cell suspension and incubated at room temperature for 5 min. After that, 10 µl propidium iodide (PI) and 400 µl PBS buffer were added to the cell suspension. The analysis of apoptosis was used a FACSCalibur Flow Cytometer (BD Biosciences). Data were analyzed using FlowJo (v.4.5; TreeStar, Inc.).
Western blotting
After 48 h of transfection, C33A and SiHa cells were lysed with RIPA lysis buffer (CoWin Biosciences) with protease inhibitor cocktail (CoWin Biosciences) to collect total protein. The protein concentration was measured by using bicinchoninic acid (BCA) protein assay kit (CoWin Biosciences). The samples were boiled at 95°C for 5 min in LDS sample buffer (Invitrogen; Thermo Fisher Scientific, Inc.). Protein (20 µg) samples was loaded onto the 10% SDS-PAGE gel. After electrophoresis, the protein was transferred to a PVDF membrane (EMD Millipore). After transferring, the PVDF membrane was blocked with 5% skimmed milk for 1 h at room temperature. Then, the membranes were incubated with primary antibodies against DNM3 (1:1,000; cat. no. 14737-1-AP; Proteintech Group, Inc.), Bcl-2 (1:2,000; cat. no. 60178-1-Ig; ProteinTech Group, Inc.), Bax (1:1,000; cat. no. 50599-2-Ig; ProteinTech Group, Inc.), cleaved caspase3 (1:1,000; cat. no. 19677-1-AP; ProteinTech Group, Inc.); E-cadherin (1:1,000; cat. no. ab194982; Abcam), N-cadherin (1:1,000; cat. no. ab18203; Abcam), vimentin (1:1,000; cat. no. ab8978; Abcam) and GAPDH (1:500; cat. no. ab8245; Abcam) at 4°C overnight, followed by incubation with anti-rabbit IgG (1:2,000; cat. no. GTX300119; GeneTex, Inc.) or anti-mouse IgG (1:2,000; cat. no. GTX300120; GeneTex, Inc.) secondary antibodies for 1 h at room temperature. Finally, the protein bands were visualized by ECL reagents (Proteintech Group, Inc.). GAPDH was used as a protein-loading control. Data were analyzed by Quantity One software (v.4.6; Bio-Rad Laboratories, Inc.).
Statistical analysis
SPSS 18.0 (SPSS Inc.) software was used for statistical analysis. All assays were performed in triplicate. The data from individual experiments was presented as the mean ± SD. The difference between the two groups was determined by unpaired Student's t-test. The difference between multiple groups was determined by one-way analysis of variance (ANOVA) and a post hoc Scheffe's test. Table I was analyzed by the χ2 test, and Table II was analyzed by the Fisher's test. Kaplan-Meier survival analysis was used to evaluate the prognosis. The log-rank test was used to compare the survival curves. P<0.05 was considered to indicate a statistically significant difference.
Table I.
DNM3 expression in cervical cancer tissues (n=41) compared with normal adjacent tissue.
| DNM3 expression | |||
|---|---|---|---|
| Group | Low, n (%) | High, n (%) | P-value |
| Cervical cancer | 23 (56.1) | 18 (43.9) | 0.007a |
| Normal adjacent | 10 (24.4) | 31 (75.6) | |
P<0.01, χ2 test. DNM3, dynamin 3.
Table II.
DNM3 expression associated with clinicopathological parameters of patients with cervical cancer.
| Clinicopathological parameters | N | DNM3 Low, n (%) | DNM3 High, n (%) | P-value |
|---|---|---|---|---|
| Age, years | ||||
| <45 | 17 | 10 (0.588) | 7 (0.412) | 0.930 |
| ≥45 | 24 | 15 (0.625) | 9 (0.375) | |
| Tumor diameter, cm | ||||
| <3 | 15 | 9 (0.600) | 6 (0.400) | 0.956 |
| ≥3 | 26 | 14 (0.538) | 12 (0.462) | |
| Pathological grading | ||||
| I–II | 28 | 12 (0.429) | 16 (0.571) | 0.030a |
| II–III | 13 | 11 (0.846) | 2 (0.154) | |
P<0.05, Fisher's test. DNM3, dynamin 3.
Results
DNM3 is significantly expressed at low levels in human cervical cancer tissues and cells compared with normal cervical tissues and cells
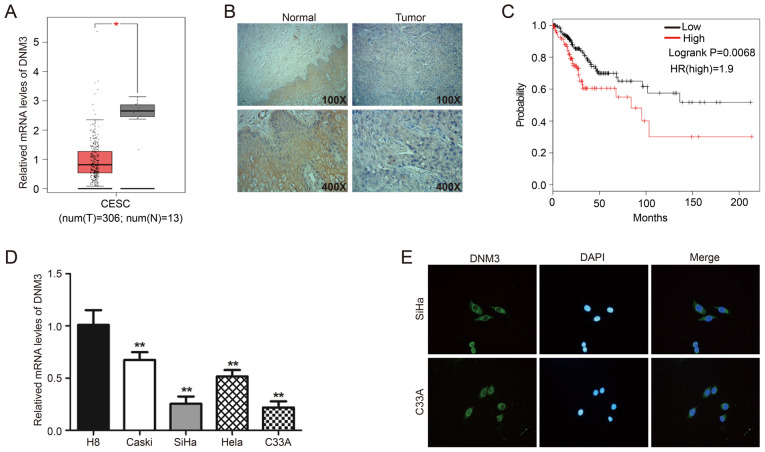
First, the GEPIA online database was used to analyze the mRNA level of DNM3 in CESC tissues (n=306) and normal cervical tissues (n=13). Expression analysis demonstrated that the mRNA expression of DNM3 in CESC tissue was significantly lower compared with that in normal cervical tissue (P<0.05; Fig. 1A). In addition, 41 clinical pairs of cervical cancer and corresponding adjacent normal tissues were used for IHC analysis of DNM3. The findings demonstrated that DNM3 was expressed at high levels in 75.6% (31/41) of adjacent tissues and 43.9% (18/41) of cervical cancer tissues (P=0.007; Fig. 1B; Table I). These data suggested that DNM3 was expressed at a significantly lower level in human cervical cancer tissues compared with that in normal tissues. In addition, the low expression of DNM3 was significantly associated with high pathological grading of cervical cancer (P=0.030; Table II). However, the expression level of DNM3 had no significant association with patient age (P=0.792) or tumor size (P=0.956; Table II). In further analysis of the association between DNM3 expression and the progression of cervical cancer, it was found that the survival of patients with low DNM3 expression was significantly improved compared with patients with high DNM3 expression (log-rank P=0.0068; Fig. 1C). The information regarding patients survival times were obtained from the GEPIA online database. The results of RT-qPCR demonstrated that the mRNA level of DNM3 in the cervical cancer Caski, SiHa, Hela and C33A cell lines was significantly lower compared with that of the normal cervical cell line H8 (P<0.01; Fig. 1D). Two cervical cancer cell lines, C33A and SiHa, with low DNM3 expression were selected for follow-up experiments. In addition, the results of the immunofluorescence assay demonstrated that DNM3 was mainly located in the cytoplasm (Fig. 1E).
Figure 1.
DNM3 is expressed at significantly low levels in human cervical cancer tissues and cells compared with normal cervical tissues and cells. (A) Bioinformatics analysis of the expression profile of DNM3 in cervical squamous cell carcinoma and endocervical adenocarcinoma and normal tissues. All data are from the GEPIA online database. (B) Immunohistochemistry staining of DNM3 protein in human cervical cancer tissue specimens. (C) Overall survival curve of the human cervical cancer patients with different DNM3 expression levels. All data are from the GEPIA online database. (D) The endogenous expression of DNM3 in cervical carcinoma cell lines (Caski, SiHa, Hela and C33A) was higher compared with that in the normal cervical cell line H8, as determined by RT-qPCR. (E) Immunofluorescence staining for DNM3 protein in C33A and SiHa cells (magnification, ×400). *P<0.05, **P<0.01. DNM3, dynamin 3; GEPIA, Gene Expression Profiling Interactive Analysis; RT-q, reverse transcription quantitative; t, tumor; n, normal; HR, hazard ratio.
DNM3 overexpression suppresses the proliferation of cervical carcinoma cells
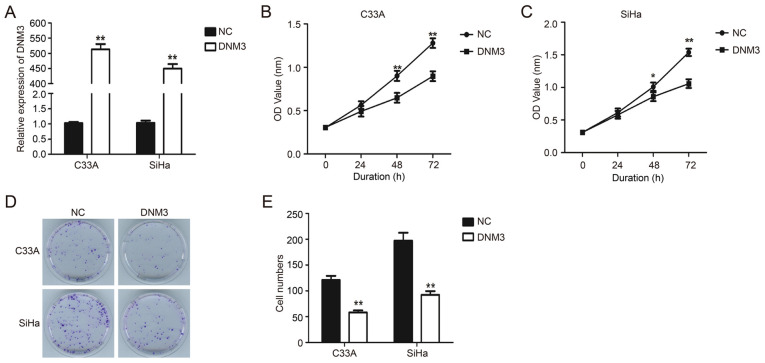
DNM3 expression plasmid was transfected into C33A and SiHa cells to upregulate the expression of DNM3 (P<0.01; Fig. 2A). Results of the CCK-8 experiment demonstrated that the proliferation of C33A and SiHa cells with DNM3 overexpression was significantly inhibited compared with that of the control group (P<0.01; Fig. 2B and C). Similarly, the results of plate clone formation demonstrated that DNM3 overexpression greatly reduced the number of cell clones compared with the control group in both cell lines (P<0.01; Fig. 2D and E). These data indicated that the overexpression of DNM3 suppressed the proliferation of human cervical carcinoma cells.
Figure 2.
DNM3 overexpression suppresses the proliferation of cervical cancer cells. (A) DNM3 expression in C33A and SiHa cells was determined by RT-qPCR after plasmid transfection. (B and C) Proliferation of C33A and SiHa cells was detected by CCK-8 assay after DNM3 expression plasmid or NC plasmid transfection. (D) Colony formation assay of C33A and SiHa cells transfected with NC or DNM3 expression plasmid. (E) Statistical analysis of cell numbers of colony formation assay. *P<0.05, **P<0.01. DNM3, dynamin 3; RT-q, reverse transcription quantitative; NC, negative control; OD, optical density.
DNM3 overexpression suppresses the migration and invasion of cervical carcinoma cells
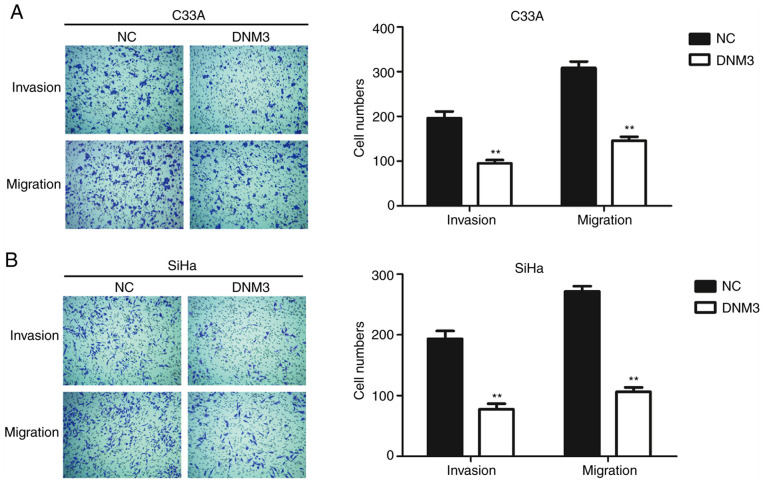
Cancer cells are generally characterized by aberrant motility (17). In order to further investigate the role of DNM3 in regulating cancer cell motility, Transwell assays were performed. The invasion and migration abilities of C33A and SiHa cells with high expression of DNM3 were inhibited significantly compared with those of the control group (P<0.01; Fig. 3).
Figure 3.
DNM3 overexpression suppresses the migration and invasion of C33A and SiHa cells. (A and B) C33A and SiHa cells were transfected with NC or DNM3 expression plasmid. Cell migration and invasion were analyzed by Transwell assays. Statistical analysis of cell numbers of migration and invasion assay are presented (magnification, ×40). **P<0.01. DNM3, dynamin 3; NC, negative control.
DNM3 overexpression induces apoptosis of cervical carcinoma cells
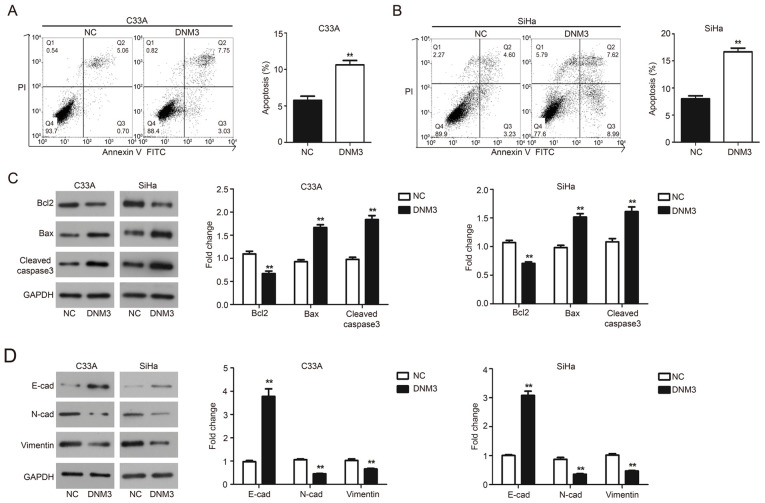
Flow cytometry and western blotting were performed to evaluate the effect of DNM3 overexpression on the apoptosis of C33A and SiHa cells. The results of flow cytometry demonstrated that DNM3 overexpression significantly promoted the apoptosis of C33A (P<0.01; Fig. 4A) and SiHa (P<0.01; Fig. 4B) cells compared with the negative control (NC) group, respectively. These findings were further validated by western blotting. Bcl2 is an antiapoptotic protein, Bax is a proapoptotic protein and caspase-3 is an apoptotic executioner (18). The aforementioned proteins serve an important role in the process of apoptosis (19,20). The results demonstrated that following DNM3 overexpression, the level of Bcl2 was downregulated, while the expression of Bax and cleaved caspase-3 was upregulated in both cell lines compared with the NC group (P<0.01; Fig. 4C). These data indicated that DNM3 promoted the apoptosis of cervical cancer cells.
Figure 4.
DNM3 overexpression promotes cell apoptosis and inhibits epithelial mesenchymal transition of C33A and SiHa cells. (A and B) C33A and SiHa cells were transfected with NC or DNM3 expression plasmid, and then cell apoptosis was determined using double staining with Annexin V/PI by flow cytometry. Statistical analysis of cell numbers after flow cytometry was presented. (C) C33A and SiHa cells were transfected with NC or DNM3 expression plasmid for 24 h. Next, the protein expression of Bcl2, Bax, cleaved caspase-3 and GAPDH was assessed by western blotting. Relative amounts of proteins were normalized to GAPDH in C33A and SiHa cells. (D) Protein expression of E-cadherin, N-cadherin, vimentin and GAPDH by western blotting. Relative amounts of proteins were normalized to GAPDH in C33A and SiHa cells. **P<0.01. DNM3, dynamin 3; NC, negative control; PI, propidium iodide; E-cad, E-cadherin; N-cad, N-cadherin.
DNM3 overexpression suppresses the EMT of cervical carcinoma cells
EMT is a developmental program and related to the progression and metastasis of cancer (21). The present study investigated the involvement of EMT in cervical cancer tumor progression. Epithelial and mesenchymal markers were assessed by western blotting and the results demonstrated that C33A and SiHa cells overexpressing DNM3 exhibited a significant upregulation of E-cadherin expression; meanwhile the expression of mesenchymal markers N-cadherin and vimentin was significantly downregulated compared with the NC group (P<0.01; Fig. 4D).
Discussion
DNM3 serves a vital role in the regulation of tumor progression (22). Numerous emerging studies have demonstrated that DNM3 exhibits decreased expression in some types of tumor tissues, such as colon cancer (22) and hepatocellular carcinoma (10). By contrast, one study demonstrated that the expression of DNM3 is upregulated in glioblastoma multiforme (GBM). In GBM orthotopic xenograft models, DNM3 expression was increased in original tumors and exosomes. The mRNA levels of DNM3 also increased in recurrence tumors and exosomes from recurrent tumor xenografts (23). To the best of our knowledge, there are no previous studies on the association between DNM3 expression and cervical cancer. The present study evaluated the expression of DNM3 in cervical tumors. The expression of DNM3 decreased in cervical cancerous tissues compared with that in normal adjacent tissues. The present study also examined the mRNA expression profile of DNM3 in four different cervical carcinoma cell lines (Caski, SiHa, Hela and C33A) and one normal cell line (H8) by RT-qPCR. The results demonstrated that the expression of DNM3 was downregulated in cervical carcinoma cells compared with that in normal cervical cells. In addition, using immunofluorescence analysis in the present study, it was demonstrated that DNM3 was located in the cytoplasm. In the present study, the methylation status of the DNM3 promoter region was not assessed in cervical cancer tissues and cells. Promoter hypermethylation may be related to decreased expression. This should be investigated by future studies.
In the present study, the function of DNM3 in cervical cancer cells was assessed by function analyses, such as proliferation, apoptosis, migration and invasion of SiHa and C33A cells. The results of CCK-8 and clone formation assays demonstrated that DNM3 overexpression significantly inhibited the proliferation of SiHa and C33A cells compared with NC groups. Cervical cancer generally develops distant metastases through the hematogenous route. In the present study, the findings of the Transwell assay demonstrated that the overexpression of DNM3 reduced cell migration and invasion of SiHa and C33A cells compared with NC groups. Flow cytometry analysis revealed that the overexpression of DNM3 promoted the apoptosis of SiHa and C33A cells.
The present study demonstrated that the overexpression of DNM3 inhibits the EMT process. EMT is a crucial process in the regulation of cell migration and invasion (21). Combined with the results of the Transwell assay, the present study revealed that the overexpression of DNM3 may inhibit the migration and invasion of cancer cells by suppressing the EMT process. However, the association between DNM3 and EMT needs further investigation.
In conclusion, the present study demonstrated the functions of DNM3 in cervical carcinoma cells. The expression of DNM3 was downregulated in cervical cancer tissues and cells compared with normal cervical tissues and cells. Meanwhile, the overexpression of DNM3 inhibited the proliferation, migration and invasion of cervical cancer cells and promoted apoptosis. Based on the findings of the present study, DNM3 may be a new potential prognostic biomarker and therapeutic target for cervical cancer.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Author's contributions
JF designed the study, performed experiments, analyzed data and wrote the manuscript. The author read and approved the final manuscript.
Ethics approval and consent to participate
The present study was approved by The Ethics Committee of Linyi Cancer Hospital (Linyi, China). All patients provided written informed consent for participation in the present study.
Patient consent for publication
Not applicable.
Competing interests
The author declares that they have no competing interests.
References
- 1.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 2.Arbyn M, Weiderpass E, Bruni L, de Sanjosé S, Saraiya M, Ferlay J, Bray F. Estimates of incidence and mortality of cervical cancer in 2018: A worldwide analysis. Lancet Glob Health. 2020;8:e191–e203. doi: 10.1016/S2214-109X(19)30482-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Vinh-Hung V, Bourgain C, Vlastos G, Cserni G, De Ridder M, Storme G, Vlastos AT. Prognostic value of histopathology and trends in cervical cancer: A SEER population study. BMC Cancer. 2007;7:164. doi: 10.1186/1471-2407-7-164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Denny L, Adewole I, Anorlu R, Dreyer G, Moodley M, Smith T, Snyman L, Wiredu E, Molijn A, Quint W, et al. Human papillomavirus prevalence and type distribution in invasive cervical cancer in sub-Saharan Africa. Int J Cancer. 2014;134:1389–1398. doi: 10.1002/ijc.28425. [DOI] [PubMed] [Google Scholar]
- 5.Xu XX, Zhou JS, Yuan SH, Yu H, Lou HM. Distribution of HPV genotype in invasive cervical carcinoma and cervical intraepithelial neoplasia in Zhejiang province, southeast China: Establishing the baseline for surveillance. Int J Environ Res Public Health. 2015;12:10794–10805. doi: 10.3390/ijerph120910794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.McDowell M, Pardee DJ, Peitzmeier S, Reisner SL, Agénor M, Alizaga N, Bernstein I, Potter J. Cervical cancer screening preferences among trans-masculine individuals: Patient-collected human papillomavirus vaginal swabs versus provider-administered pap tests. LGBT Health. 2017;4:252–259. doi: 10.1089/lgbt.2016.0187. [DOI] [PubMed] [Google Scholar]
- 7.Bosgraaf RP, Ketelaars PJ, Verhoef VM, Massuger LF, Meijer CJ, Melchers WJ, Bekkers RL. Reasons for non-attendance to cervical screening and preferences for HPV self-sampling in Dutch women. Prev Med. 2014;64:108–113. doi: 10.1016/j.ypmed.2014.04.011. [DOI] [PubMed] [Google Scholar]
- 8.Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30. doi: 10.3322/caac.21166. [DOI] [PubMed] [Google Scholar]
- 9.Heymann JA, Hinshaw JE. Dynamins at a glance. J Cell Sci. 2009;122((Pt 19)):3427–3431. doi: 10.1242/jcs.051714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Raja SA, Shah STA, Tariq A, Bibi N, Sughra K, Yousuf A, Khawaja A, Nawaz M, Mehmood A, Khan MJ, Hussain A. Caveolin1 and dynamin2 overexpression is associated with the progression of bladder cancer. Oncol Lett. 2019;18:219–226. doi: 10.3892/ol.2019.10310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Meng J. Distinct functions of dynamin isoforms in tumorigenesis and their potential as therapeutic targets in cancer. Oncotarget. 2017;8:41701–41716. doi: 10.18632/oncotarget.16678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Inokawa Y, Nomoto S, Hishida M, Hayashi M, Kanda M, Nishikawa Y, Takeda S, Fujiwara M, Koike M, Sugimoto H, et al. Dynamin 3: A new candidate tumor suppressor gene in hepatocellular carcinoma detected by triple combination array analysis. Onco Targets Ther. 2013;6:1417–1424. doi: 10.2147/OTT.S51913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhang Z, Chen C, Guo W, Zheng S, Sun Z, Geng X. DNM3 attenuates hepatocellular carcinoma growth by activating P53. Med Sci Monit. 2016;22:197–205. doi: 10.12659/MSM.896545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gu C, Yao J, Sun P. Dynamin 3 suppresses growth and induces apoptosis of hepatocellular carcinoma cells by activating inducible nitric oxide synthase production. Oncol Lett. 2017;13:4776–4784. doi: 10.3892/ol.2017.6057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102. doi: 10.1093/nar/gkx247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 17.Trepat X, Chen Z, Jacobson K. Cell migration. Compr Physiol. 2012;2:2369–2392. doi: 10.1002/cphy.c110012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Dietrich JB. Apoptosis and anti-apoptosis genes in the Bcl-2 family. Arch Physiol Biochem. 1997;105:125–135. doi: 10.1076/apab.105.2.125.12927. (In French) [DOI] [PubMed] [Google Scholar]
- 19.Brambilla E, Negoescu A, Gazzeri S, Lantuejoul S, Moro D, Brambilla C, Coll JL. Apoptosis-related factors p53, Bcl2, and Bax in neuroendocrine lung tumors. Am J Pathol. 1996;149:1941–1952. [PMC free article] [PubMed] [Google Scholar]
- 20.Levesley J, Steele L, Bruning-Richardson A, Davison A, Zhou J, Ding C, Lawler S, Short SC. Selective BCL-XL inhibition promotes apoptosis in combination with MLN8237 in medulloblastoma and pediatric glioblastoma cells. Neuro Oncol. 2018;20:203–214. doi: 10.1093/neuonc/nox134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Suarez-Carmona M, Lesage J, Cataldo D, Gilles C. EMT and inflammation: Inseparable actors of cancer progression. Mol Oncol. 2017;11:805–823. doi: 10.1002/1878-0261.12095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ma Y, Guan L, Han Y, Zhou Y, Li X, Liu Y, Zhang X, Zhang W, Li X, Wang S, Lu W. siPRDX2-elevated DNM3 inhibits the proliferation and metastasis of colon cancer cells via AKT signaling pathway. Cancer Manag Res. 2019;11:5799–5811. doi: 10.2147/CMAR.S193805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yang JK, Yang JP, Tong J, Jing SY, Fan B, Wang F, Sun GZ, Jiao BH. Exosomal miR-221 targets DNM3 to induce tumor progression and temozolomide resistance in glioma. J Neurooncol. 2017;131:255–265. doi: 10.1007/s11060-016-2308-5. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.