Abstract
Polygenic hazard score (PHS) models are associated with age at diagnosis of prostate cancer. Our model developed in Europeans (PHS46), showed reduced performance in men with African genetic ancestry. We used a cross-validated search to identify SNPs that might improve performance in this population. Anonymized genotypic data were obtained from the PRACTICAL consortium for 6,253 men with African genetic ancestry. Ten iterations of a ten-fold cross-validation search were conducted, to select SNPs that would be included in the final PHS46+African model. The coefficients of PHS46+African were estimated in a Cox proportional hazards framework using age at diagnosis as the dependent variable and PHS46, and selected SNPs as predictors. The performance of PHS46 and PHS46+African were compared using the same cross-validated approach. Three SNPs (rs76229939, rs74421890, and rs5013678) were selected for inclusion in PHS46+African. All three SNPs are located on chromosome 8q24. PHS46+African showed substantial improvements in all performance metrics measured, including a 75% increase in the relative hazard of those in the upper 20% compared to the bottom 20% (2.47 to 4.34) and a 20% reduction in the relative hazard of those in the bottom 20% compared to the middle 40% (0.65 to 0.53). In conclusion, we identified three SNPs that substantially improved the association of PHS46 with age at diagnosis of prostate cancer in men with African genetic ancestry to levels comparable to Europeans.
Keywords: Prostate cancer, health disparities, genome wide association study, polygenic risk, genomics, genotypic ancestry, African
Introduction
Polygenic models can provide personalized estimates of the risk of developing prostate cancer. In the context of survival analysis, these models can provide insight into age at diagnosis of prostate cancer, and thus could be used to guide decisions on whether and when to offer screening1. Studies of polygenic models have often included only individuals of European genetic ancestry, owing to greater availability of data from that population2,3. As a consequence, these models have been tailored to identify and estimate coefficients of genetic common variants for that particular population, while potentially missing variants that may hold value in other populations2. There is concern that using these European-focused models could actually exacerbate health disparities2-4.
As an example, our group recently published on the performance of a polygenic hazard score (PHS) originally developed using a European dataset, in a multi-ethnic dataset consisting of individuals of European, African, and Asian genetic ancestry5. The model (called here PHS46, referred to in the referenced manuscript as PHS2), includes 46 single nucleotide polymorphisms (SNPs) in its calculation and was strongly associated with age at diagnosis in all three genetic populations (p<10−16). However, the hazard ratio for prostate cancer between individuals in the upper 20th percentile to those in the lower 20th percentile of PHS46 was approximately half as large for those with African genetic ancestry (2.6) as it was for those with European (5.6) or Asian (4.6) ancestry. A similar pattern was observed for clinically significant prostate cancer and for death from prostate cancer.
In the current study, we attempt to bridge the apparent gap in model performance of PHS46 for individuals with African genetic ancestry. To this end, we used a machine learning approach to systematically search for SNPs that add statistical value to a base model of PHS46 among African men (PHS46+African). By including PHS46 as a covariate in our SNP search, we sought to identify those SNPs that may hold particular value for individuals with African genetic ancestry.
Material and Methods
Study dataset
We obtained genotype and phenotype data from the Prostate Cancer Association Group to Investigate Cancer Associated Alterations in the Genome (PRACTICAL)6 consortium for this study. Genotyping was performed using the OncoArray platform6,7 and had undergone quality assurance steps, as described previously8. The study dataset contains no overlap with that used to estimate model coefficients of PHS46. It is a subset of another dataset wherein the performance discrepancy of PHS46 between different genotypic ancestries was first observed5. All 46 SNPs of PHS46 were directly genotyped on the OncoArray platform.
The genotypic ancestry of each individual was also determined previously6,9. In total, the African dataset consisted of data from 6,253 men with African genotypic ancestry. Missing SNP calls were replaced with the mean of the genotyped data for that SNP in the African dataset. The percentage of individuals with missing SNP calls ranged from 0 to 5.9% across the SNPs, while the percentage of SNPs with missing calls ranged from 0.5% to 5.1% across the individuals. Individuals without prostate cancer were censored at age at last follow-up in the Cox proportional hazards models. A description of the PRACTICAL study groups that contributed data towards this analysis are described in Supplementary Table 1. PHS46 risk score for each individual in the African dataset was estimated as the sum of SNP allele counts (X) multiplied by their respective coefficients (β)5:
SNP-scan
A multi-step approach was used to select SNPs, from those directly genotyped on the OncoArray platform, that would improve the performance of PHS46 in the African dataset. Training and testing sets were generated using 10 iterations of a 10-fold cross-validation structure resulting in 100 total permutations. For each permutation, a multivariable logistic regression model using case/control status as the dependent variable was estimated using each genotyped SNP in turn, adjusting for PHS46 and four principal components based on genetic ancestry, determined previously9. SNPs with p-values less than 1x10−6 were considered for further analysis. In order of increasing p-value, each SNP was tested in a multiple Cox proportional hazards model, after adjusting for PHS46, four ancestral principal components, and previously selected SNPs. The Cox model in the SNP-scan used age at diagnosis of prostate cancer as the dependent variable. If the p-value of the coefficient of the tested SNP was less than 1x10−6, it was considered for the final model in that permutation. SNPs that reached this p-value threshold in more than 50% of the permutations were selected to construct the PHS46+African model, consisting of PHS46 and the newly identified SNPs.
Comparing performance between PHS46 and PHS46+African – Hazard Ratio
For each permutation of the previously described cross-validation structure, an PHS46+African Cox proportional hazards model was estimated in the training set using PHS46 and the selected SNPs as independent predictors. The PHS46+African risk score for each individual is then estimated using the corresponding PHS46 score, selected SNP allele counts (Y) and their respective coefficients (α):
The performance of the PHS46+African and PHS46 models was then determined in the cross-validation testing set, and the resulting hazard ratios (HR) were obtained, as previously described1. For each model, the PHS risk scores within the cross-validation testing set are assigned to quantile groups identified using the corresponding training set control values. The hazard ratio between two quantile groups, such as those in the top 20% to those in the bottom 20%, is estimated as the exponential of the difference in mean PHS values for each group. In this calculation, the PHS values are linearly scaled by a sample-weight correction factor, to account for case-control sampling1,5,10. Three HR were calculated: HR80/20 (top 20% to bottom 20%), HR98/50 (top 2% to middle 40%) and HR20/50 (bottom 20% to middle 40%). The average HR across permutations for both PHS46+African and PHS46 are reported.
To allow for comparisons with previously published results, the performance metrics for PHS46 and PHS46+African were also estimated for age at diagnosis of clinically significant prostate cancer. When estimating performance for clinically significant prostate cancer, controls and non-clinically significant cancers were censored at age of last follow-up and age of diagnosis, respectively. The previously used criteria for clinically significant cancer were any of: Gleason score >=7, stage T3-T4, PSA concentration >= 10ng/mL, pelvic lymph nodal metastasis, or distant metastasis1. Paired t-tests were used to test for statistically significant differences (α = 0.05) in HR between PHS46+African and PHS46.
Additionally, in each permutation, the performance of a Cox model consisting of PHS46 and SNPs that were considered in that permutation was also estimated. These results are provided within the Supplementary Table 2 and provide performance estimates that are not prone to information leakage from training to testing set.
Comparing performance between PHS46 and PHS46+African – C-index
In addition to the hazard ratio, the performances of PHS46 and PHS46+African were compared using Harrell’s c-index11. For each permutation of the aforementioned cross-validation structure, the c-index of PHS46 and PHS46+African scores were estimated in the testing fold using the “coxph” function in the R “survival” package. Paired t-tests were used to test for statistically significant differences (α = 0.05) between the two models.
Characterization of PHS46+African
Coefficients of the PHS46+African model, consisting of PHS46 and the SNPs selected in the SNP-scan, were estimated using 1000 bootstrapped samples of the African dataset.
Clinical utility of PHS46+African
As an example of the clinical utility of the PHS46+African risk score, the risk-equivalent age was estimated for those individuals in the upper 2 percentile of the distribution of PHS46+African risk scores. The risk-equivalent age, as defined previously12, is when an individual from a given PHS percentile has prostate cancer risk equivalent to the average 60-year-old man. The age-specific general cumulative incidence curve was generated using data from SEER*Explorer incidence rates by age at diagnosis, 2003-2017 for Black Americans13. The corresponding risk-adjusted incidence curve was estimated by multiplying the general cumulative incidence curve by the mean value of HR98/50 for PHS46+African obtained from the analysis of the age-of-diagnosis of prostate cancer. The risk-equivalent age was then calculated as the age at which the risk-adjusted cumulative incidence curve had the same value as the general cumulative incidence curve at age 60.
Results
Individual and OncoArray characteristics
In total, there were 3,013 men with (cases) and 3,240 men without (controls) prostate cancer in the African dataset. The mean [95% CI] ages of cases and controls were 62.4 [62.1, 62.7] and 61.8 [61.4, 62.1] years respectively. The OncoArray genotypic data, after the quality assurance process, included 444,323 SNPs.
Single Nucleotide Polymorphism (SNP)-scan
Across the 100 permutations of the cross-validation iterations, a total of twelve SNPs were considered for final selection (Supplementary Table 3). Three SNPs were selected in more than 50% of the permutations and included in the final PHS46+African model. By cross-referencing the chromosomal positions against dbSNP14, these variants were identified as rs7622993915, rs7442189016, and rs501367817. All three SNPs (Table 1) are located on chromosome 8q24, a region of the chromosome previously identified as containing common variants associated with prostate cancer18,19. An examination of the R2 (Supplementary Table 4) showed little association, ranging from 0.0027 to 0.0057, among genotype data from the three SNPs in the African dataset.
Table 1. Characteristics of PHS46+African SNPs.
RS-ID, chromosome and base-pair position (based on version 37), effect and reference alleles, bootstrap-estimated beta, and effect allele frequencies in aggregated Africans from 1000Genomes (referenced from dbSNP) of the three SNPs selected for addition to PHS46.
| RS number | Chromosome | Position | Effect | Ref | beta | Frequency (%) |
|---|---|---|---|---|---|---|
| rs76229939 | 8 | 128085394 | G | A | 0.441 | 4.8 |
| rs74421890 | 8 | 128096183 | A | G | 0.415 | 4.1 |
| rs5013678 | 8 | 128103979 | G | A | −0.260 | 8.1 |
Reference threshold (Supplementary Table 5) and mean (Supplementary Table 6) values for PHS46+African in the African dataset are presented in the Supplemental Data.
HR performance of PHS46+African
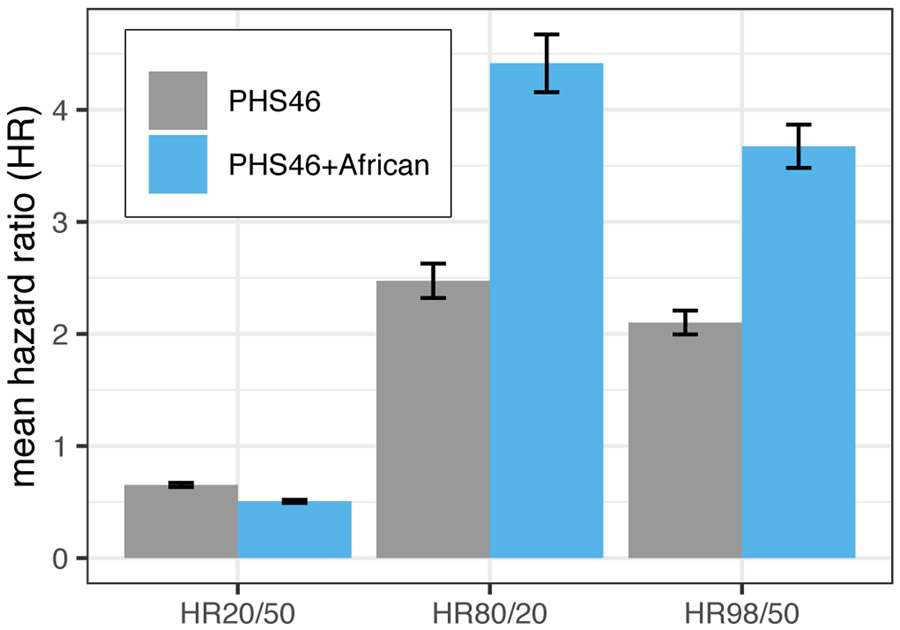
Figure 1 demonstrates the difference in HRs between PHS46+African and PHS46 within the African dataset using age at diagnosis of any prostate cancer (Supplementary Table 7). Overall, we observed an improvement in all the metrics calculated: a 75% increase in HR98/50 from 2.10 to 3.67; a 79% increase in HR80/20 from 2.47 to 4.42; and a 23% decrease in HR20/50 from 0.65 to 0.51. We also observed improvements in all performance metrics when using age at diagnosis of clinically significant prostate cancer: 103% increase in HR98/50 from 1.91 to 3.88, 113% improvement in HR80/20 from 2.21 to 4.71, and 29% improvement in HR20/50 from 0.70 to 0.50. All observed changes in HR were statistically significant (p < 1x10−16).
Figure 1. Comparison between PHS46 and PHS46+African.
Mean hazard ratio metrics plotted for PHS46 and PHS46+African models in the African dataset. Improvements were observed in all performance metrics investigated. Error bars represent 95% confidence interval
C-index of PHS46+African
The mean c-indices of PHS46 and PHS46+African across the cross-validation folds were estimated as 0.55 and 0.58 (p < 1x10−16), respectively.
Risk-equivalent age for PHS46+African
The risk-equivalent age for those individuals in the top 2 percentiles of the distribution of PHS46+African scores was estimated as 50 years old, suggesting that a man with a PHS46+African score in the top 2 percentile reached a prostate cancer detection risk equivalent to that of a standard 60-year-old roughly 10 years earlier, at an age of 50 years. The corresponding risk-equivalent age when using PHS46 scores was 54 years.
Discussion
Using a cross-validated search of a dataset made up entirely of men with African genetic ancestry, we were able to identify three SNPs that substantially improved the performance of PHS46 in this population to levels that are comparable to those observed in Europeans and Asians. Performance improvements were observed in hazard ratios tracking risk between PHS groups, concordance-indices tracking the overall utility of PHS as a continuous variable, and risk-equivalent age tracking the potential clinical utility of PHS. The three SNPs, rs76229939, rs74421890, and rs5013678, are all located on chromosome 8q24 – a region of the genome where variants have been associated with prostate cancer in both the general population and specifically in men with African genetic ancestry19,20. Despite the relative proximity of the three SNPs on chromosome 8, their genetic data was not strongly associated in our dataset, suggesting that each SNP provides non-redundant information for an individual’s genetic score.
Each of the three SNPs have been previously identified in the literature to be associated with prostate cancer: rs76229939 is an intron variant of the prostate-cancer-associated transcript 2 (PCAT2) gene, while rs74421890 and rs5013678 are both non-coding transcript variants of the prostate-cancer-associated non-coding RNA 1 (PRNCR1) gene. The minor allele frequencies of rs76229939 and rs74421890 in Europeans, as reported by dbSNP14, are approximately zero to three decimal places, which may explain why they were not selected in the original formulation of PHS46.
This study is not meant to be an exhaustive search for all possible SNPs that are associated with the age of diagnosis of prostate cancer in individuals with African genetic ancestry. Our study is also limited by the small number of available observations relative to those often found in many genome-wide association studies, which can have tens or hundreds of thousands of individuals. However, we were able to extract information that is likely robust by employing a cross-validated search for those SNPs that specifically add value to the performance of PHS46, and not simply independently associated with prostate cancer. Future analysis will include a more detailed analysis of the 8q24 region, including SNPs that are imputed using TOPMed reference panels. We also note that no SNP score, including PHS46 and PHS46+African, has been shown to discriminate men at risk of aggressive prostate cancer from those at risk of indolent prostate cancer. Finally, the performance metrics reported in this study may be biased by the leakage of information across cross-validated folds of the data when identifying those SNPs to include in the final African-PHS model. This bias is expected to be similar for all SNPs and should not have influenced selection of the three SNPs included in the final model over those not selected.
In conclusion, we identified three SNPs (rs76229939, rs74421890, and rs5013678) on 8q24 that substantially improved the performance of PHS46 in a dataset of men with African genetic ancestry. The addition of these SNPs to the polygenic risk score substantially improved its association with age at diagnosis of prostate cancer in Africans, to levels comparable with those found in Europeans.
Supplementary Material
Acknowledgments
This study was funded in part by a grant from the United States National Institute of Health/National Institute of Biomedical Imaging and Bioengineering (#K08EB026503), U.S. Department of Defense (#W81XWH-13-0391), the Research Council of Norway (#223273), KG Jebsen Stiftelsen, and South East Norway Health Authority.
Abbreviations
- HR
Hazard ratio
- PHS
Polygenic hazard score
- PHS46
Previously published polygenic hazard score model using 46 SNPs
- PHS46+African
Updated polygenic hazard score model for men with African genetic ancestry
- PRACTICAL
Prostate Cancer Association Group to Investigate Cancer Associated Alterations in the Genome consortium
- SNP
Single nucleotide polymorphism
Appendix
Appendix 1. Members of the PRACTICAL Consortium
Christopher A. Haiman1, Fredrick R. Schumacher2,3, Sara Benlloch4,5, Ali Amin Al Olama6,7, Sonja I. Berndt8, David V. Conti1, Fredrik Wiklund9, Stephen Chanock8, Susan M. Gapstur10, Victoria L. Stevens10, Jyotsna Batra11,12, Judith Clements11,12, APCB BioResource13,14, Henrik Grönberg15, Nora Pashayan16,17, Johanna Schleutker18,19, Demetrius Albanes8, Stephanie Weinstein8, Alicja Wolk20,21, Catharine West22, Lorelei Mucci23, Stella Koutros8, Karina Dalsgaard Sørensen24,25, Eli Marie Grindedal26, David E. Neal27,28,29, Freddie C. Hamdy30,31, Jenny L. Donovan32, Ruth C. Travis33, Robert J. Hamilton34,35, Sue Ann Ingles36, Barry S. Rosenstein37,38, Yong-Jie Lu39, Graham G. Giles40,41,42, Ana Vega43,44,45, Manolis Kogevinas46,47,48,49, Kathryn L. Penney50, Janet L. Stanford51,52, Cezary Cybulski53, Børge G. Nordestgaard54,55, Hermann Brenner56,57,58, Christiane Maier59, Jeri Kim60, Manuel R. Teixeira61,62, Susan L. Neuhausen63, Kim De Ruyck64, Azad Razack65, Lisa F. Newcomb51,66, Davor Lessel67, Radka Kaneva68, Nawaid Usmani69,70, Frank Claessens71, Paul A. Townsend72, Manuela Gago-Dominguez73,74, Monique J. Roobol75, Kay-Tee Khaw76, Lisa Cannon-Albright77,78, Hardev Pandha79, Stephen N. Thibodeau80, Peter Kraft81, Elio Riboli82
1Center for Genetic Epidemiology, Department of Preventive Medicine, Keck School of Medicine, University of Southern California/Norris Comprehensive Cancer Center, Los Angeles, CA 90015, USA
2Department of Population and Quantitative Health Sciences, Case Western Reserve University, Cleveland, OH 44106-7219, USA
3Seidman Cancer Center, University Hospitals, Cleveland, OH 44106, USA.
4Centre for Cancer Genetic Epidemiology, Department of Public Health and Primary Care, University of Cambridge, Strangeways Research Laboratory, Cambridge CB1 8RN, UK
5The Institute of Cancer Research, London, SM2 5NG, UK
6Centre for Cancer Genetic Epidemiology, Department of Public Health and Primary Care, University of Cambridge, Strangeways Research Laboratory, Cambridge, UK
7University of Cambridge, Department of Clinical Neurosciences, Stroke Research Group, R3, Box 83, Cambridge Biomedical Campus, Cambridge CB2 0QQ, UK
8Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, Maryland, 20892, USA
9Department of Medical Epidemiology and Biostatistics, Karolinska Institute, SE-171 77 Stockholm, Sweden
10Behavioral and Epidemiology Research Group, Research Program, American Cancer Society, 250 Williams Street, Atlanta, GA 30303, USA
11Australian Prostate Cancer Research Centre-Qld, Institute of Health and Biomedical Innovation and School of Biomedical Sciences, Queensland University of Technology, Brisbane QLD 4059, Australia
12Translational Research Institute, Brisbane, Queensland 4102, Australia
13Australian Prostate Cancer Research Centre-Qld, Queensland University of Technology, Brisbane; Prostate Cancer Research Program, Monash University, Melbourne; Dame Roma Mitchell Cancer Centre, University of Adelaide, Adelaide; Chris O'Brien Lifehouse and
14Translational Research Institute, Brisbane, Queensland, Australia
15Department of Medical Epidemiology and Biostatistics, Karolinska Institute, Stockholm, Sweden
16Department of Applied Health Research, University College London, London, WC1E 7HB, UK
17Centre for Cancer Genetic Epidemiology, Department of Oncology, University of Cambridge, Strangeways Laboratory, Worts Causeway, Cambridge, CB1 8RN, UK
18Institute of Biomedicine, Kiinamyllynkatu 10, FI-20014 University of Turku, Finland
19Department of Medical Genetics, Genomics, Laboratory Division, Turku University Hospital, PO Box 52, 20521 Turku, Finland
20Division of Nutritional Epidemiology, Institute of Environmental Medicine, Karolinska Institutet, SE-171 77 Stockholm, Sweden
21Department of Surgical Sciences, Uppsala University, 75185 Uppsala, Sweden
22Division of Cancer Sciences, University of Manchester, Manchester Academic Health Science Centre, Radiotherapy Related Research, The Christie Hospital NHS Foundation Trust, Manchester, M13 9PL UK
23Department of Epidemiology,Harvard T. H. Chan School of Public Health, Boston, MA 02115, USA
24Department of Molecular Medicine, Aarhus University Hospital, Palle Juul-Jensen Boulevard 99, 8200 Aarhus N, Denmark
25Department of Clinical Medicine, Aarhus University, DK-8200 Aarhus N
26Department of Medical Genetics, Oslo University Hospital, 0424 Oslo, Norway
27Nuffield Department of Surgical Sciences, University of Oxford, Room 6603, Level 6, John Radcliffe Hospital, Headley Way, Headington, Oxford, OX3 9DU, UK
28University of Cambridge, Department of Oncology, Box 279, Addenbrooke's Hospital, Hills Road, Cambridge CB2 0QQ, UK
29Cancer Research UK, Cambridge Research Institute, Li Ka Shing Centre, Cambridge UK
30Nuffield Department of Surgical Sciences, University of Oxford, Oxford, OX1 2JD, UK
31Faculty of Medical Science, University of Oxford, John Radcliffe Hospital, Oxford, UK
32Population Health Sciences, Bristol Medical School, University of Bristol, BS8 2PS, UK
33Cancer Epidemiology Unit, Nuffield Department of Population Health, University of Oxford, Oxford, OX3 7LF, UK
34Dept. of Surgical Oncology, Princess Margaret Cancer Centre, Toronto ON M5G 2M9, Canada
35Dept. of Surgery (Urology), University of Toronto, Canada
36Department of Preventive Medicine, Keck School of Medicine, University of Southern California/Norris Comprehensive Cancer Center, Los Angeles, CA 90015, USA
37Department of Radiation Oncology and Department of Genetics and Genomic Sciences, Box 1236, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY 10029, USA
38Department of Genetics and Genomic Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029-5674 , USA.
39Centre for Molecular Oncology, Barts Cancer Institute, Queen Mary University of London, John Vane Science Centre, Charterhouse Square, London, EC1M 6BQ, UK
40Cancer Epidemiology Division, Cancer Council Victoria, 615 St Kilda Road, Melbourne, VIC 3004, Australia
41Centre for Epidemiology and Biostatistics, Melbourne School of Population and Global Health, The University of Melbourne, Grattan Street, Parkville, VIC 3010, Australia
42Precision Medicine, School of Clinical Sciences at Monash Health, Monash University, Clayton, Victoria 3168, Australia
43Fundación Pública Galega Medicina Xenómica, Santiago De Compostela, 15706, Spain.
44Instituto de Investigación Sanitaria de Santiago de Compostela, Santiago De Compostela, 15706, Spain.
45Centro de Investigación en Red de Enfermedades Raras (CIBERER), Spain
46ISGlobal, Barcelona, Spain
47IMIM (Hospital del Mar Medical Research Institute), Barcelona, Spain
48Universitat Pompeu Fabra (UPF), Barcelona, Spain
49CIBER Epidemiología y Salud Pública (CIBERESP), Madrid, Spain
50Channing Division of Network Medicine, Department of Medicine, Brigham and Women's Hospital/Harvard Medical School, Boston, MA 02184, USA
51Division of Public Health Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington, 98109-1024, USA
52Department of Epidemiology, School of Public Health, University of Washington, Seattle, Washington 98195, USA
53International Hereditary Cancer Center, Department of Genetics and Pathology, Pomeranian Medical University, Szczecin, Poland
54Faculty of Health and Medical Sciences, University of Copenhagen, 2200 Copenhagen, Denmark
55Department of Clinical Biochemistry, Herlev and Gentofte Hospital, Copenhagen University Hospital, Herlev, 2200 Copenhagen, Denmark
56Division of Clinical Epidemiology and Aging Research, German Cancer Research Center (DKFZ), D-69120, Heidelberg, Germany
57German Cancer Consortium (DKTK), German Cancer Research Center (DKFZ), D-69120 Heidelberg, Germany
58Division of Preventive Oncology, German Cancer Research Center (DKFZ) and National Center for Tumor Diseases (NCT), Im Neuenheimer Feld 460 69120 Heidelberg, Germany
59Humangenetik Tuebingen, Paul-Ehrlich-Str 23, D-72076 Tuebingen, Germany
60The University of Texas M. D. Anderson Cancer Center, Department of Genitourinary Medical Oncology, 1515 Holcombe Blvd., Houston, TX 77030, USA
61Department of Genetics, Portuguese Oncology Institute of Porto (IPO-Porto), Porto, Portugal
62Biomedical Sciences Institute (ICBAS), University of Porto, Porto, Portugal
63Department of Population Sciences, Beckman Research Institute of the City of Hope, 1500 East Duarte Road, Duarte, CA 91010, 626-256-HOPE (4673)
64Ghent University, Faculty of Medicine and Health Sciences, Basic Medical Sciences, Proeftuinstraat 86, B-9000 Gent
65Department of Surgery, Faculty of Medicine, University of Malaya, 50603 Kuala Lumpur, Malaysia
66Department of Urology, University of Washington, 1959 NE Pacific Street, Box 356510, Seattle, WA 98195, USA
67Institute of Human Genetics, University Medical Center Hamburg-Eppendorf, D-20246 Hamburg, Germany
68Molecular Medicine Center, Department of Medical Chemistry and Biochemistry, Medical University of Sofia, Sofia, 2 Zdrave Str., 1431 Sofia, Bulgaria
69Department of Oncology, Cross Cancer Institute, University of Alberta, 11560 University Avenue, Edmonton, Alberta, Canada T6G 1Z2
70Division of Radiation Oncology, Cross Cancer Institute, 11560 University Avenue, Edmonton, Alberta, Canada T6G 1Z2
71Molecular Endocrinology Laboratory, Department of Cellular and Molecular Medicine, KU Leuven, BE-3000, Belgium
72Division of Cancer Sciences, Manchester Cancer Research Centre, Faculty of Biology, Medicine and Health, Manchester Academic Health Science Centre, NIHR Manchester Biomedical Research Centre, Health Innovation Manchester, Univeristy of Manchester, M13 9WL
73Genomic Medicine Group, Galician Foundation of Genomic Medicine, Instituto de Investigacion Sanitaria de Santiago de Compostela (IDIS), Complejo Hospitalario Universitario de Santiago, Servicio Galego de Saúde, SERGAS, 15706, Santiago de Compostela, Spai
74University of California San Diego, Moores Cancer Center, La Jolla, CA 92037, USA
75Department of Urology, Erasmus University Medical Center, 3015 CE Rotterdam, The Netherlands
76Clinical Gerontology Unit, University of Cambridge, Cambridge, CB2 2QQ, UK
77Division of Epidemiology, Department of Internal Medicine, University of Utah School of Medicine, Salt Lake City, Utah, USA
78George E. Wahlen Department of Veterans Affairs Medical Center, Salt Lake City, Utah 84148, USA
79The University of Surrey, Guildford, Surrey, GU2 7XH
80Department of Laboratory Medicine and Pathology, Mayo Clinic, Rochester, MN 55905, USA
81Program in Genetic Epidemiology and Statistical Genetics, Department of Epidemiology, Harvard School of Public Health, Boston, MA, USA
82Department of Epidemiology and Biostatistics, School of Public Health, Imperial College London, SW7 2AZ, UK
Appendix 2. Funding sources for the PRACTICAL consortium
CRUK and PRACTICAL consortium
This work was supported by the Canadian Institutes of Health Research, European Commission's Seventh Framework Programme grant agreement n° 223175 (HEALTH-F2-2009-223175), Cancer Research UK Grants C5047/A7357, C1287/A10118, C1287/A16563, C5047/A3354, C5047/A10692, C16913/A6135, and The National Institute of Health (NIH) Cancer Post-Cancer GWAS initiative grant: No. 1 U19 CA 148537-01 (the GAME-ON initiative).
We would also like to thank the following for funding support: The Institute of Cancer Research and The Everyman Campaign, The Prostate Cancer Research Foundation, Prostate Research Campaign UK (now Prostate Action), The Orchid Cancer Appeal, The National Cancer Research Network UK, The National Cancer Research Institute (NCRI) UK. We are grateful for support of NIHR funding to the NIHR Biomedical Research Centre at The Institute of Cancer Research and The Royal Marsden NHS Foundation Trust.
The Prostate Cancer Program of Cancer Council Victoria also acknowledge grant support from The National Health and Medical Research Council, Australia (126402, 209057, 251533, , 396414, 450104, 504700, 504702, 504715, 623204, 940394, 614296,), VicHealth, Cancer Council Victoria, The Prostate Cancer Foundation of Australia, The Whitten Foundation, PricewaterhouseCoopers, and Tattersall’s. EAO, DMK, and EMK acknowledge the Intramural Program of the National Human Genome Research Institute for their support.
Genotyping of the OncoArray was funded by the US National Institutes of Health (NIH) [U19 CA 148537 for ELucidating Loci Involved in Prostate cancer SuscEptibility (ELLIPSE) project and X01HG007492 to the Center for Inherited Disease Research (CIDR) under contract number HHSN268201200008I].
This study would not have been possible without the contributions of the following: Coordination team, bioinformatician and genotyping centers: Genotyping at CCGE, Cambridge: Caroline Baines and Don Conroy
Funding for the iCOGS infrastructure came from: the European Community's Seventh Framework Programme under grant agreement n° 223175 (HEALTH-F2-2009-223175) (COGS), Cancer Research UK (C1287/A10118, C1287/A 10710, C12292/A11174, C1281/A12014, C5047/A8384, C5047/A15007, C5047/A10692, C8197/A16565), the National Institutes of Health (CA128978) and Post-Cancer GWAS initiative (1U19 CA148537, 1U19 CA148065 and 1U19 CA148112 - the GAME-ON initiative), the Department of Defence (W81XWH-10-1-0341), the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer, Komen Foundation for the Cure, the Breast Cancer Research Foundation, and the Ovarian Cancer Research Fund.
This study would not have been possible without the contributions of the following: Per Hall (COGS); Douglas F. Easton, Paul Pharoah, Kyriaki Michailidou, Manjeet K. Bolla, Qin Wang (BCAC), Andrew Berchuck (OCAC), Rosalind A. Eeles, Douglas F. Easton, Ali Amin Al Olama, Zsofia Kote-Jarai, Sara Benlloch (PRACTICAL), Georgia Chenevix-Trench, Antonis Antoniou, Lesley McGuffog, Fergus Couch and Ken Offit (CIMBA), Joe Dennis, Alison M. Dunning, Andrew Lee, and Ed Dicks, Craig Luccarini and the staff of the Centre for Genetic Epidemiology Laboratory, Javier Benitez, Anna Gonzalez-Neira and the staff of the CNIO genotyping unit, Jacques Simard and Daniel C. Tessier, Francois Bacot, Daniel Vincent, Sylvie LaBoissière and Frederic Robidoux and the staff of the McGill University and Génome Québec Innovation Centre, Stig E. Bojesen, Sune F. Nielsen, Borge G. Nordestgaard, and the staff of the Copenhagen DNA laboratory, and Julie M. Cunningham, Sharon A. Windebank, Christopher A. Hilker, Jeffrey Meyer and the staff of Mayo Clinic Genotyping Core Facility
Additional funding and acknowledgments from studies in PRACTICAL:
Information of the PRACTICAL consortium can be found at http://practical.icr.ac.uk/
BioVU
The dataset(s) used for the analyses described were obtained from Vanderbilt University Medical Center’s BioVU, which is supported by institutional funding and by the National Center for Research Resources, Grant UL1 RR024975-01 (which is now at the National Center for Advancing Translational Sciences, Grant 2 UL1 TR000445-06).
CPDR
Uniformed Services University for the Health Sciences HU0001-10-2-0002 (PI: David G. McLeod, MD)
EPICAP
The EPICAP study was supported by grants from Ligue Nationale Contre le Cancer; Institut National du Cancer (INCa); Fondation ARC; Fondation de France; Agence Nationale de sécurité sanitaire de l’alimentation, de l’environnement et du travail (ANSES); Ligue départementale du Val de Marne. The EPICAP study group would like to thank all urologists, Antoinette Anger and Hasina Randrianasolo (study monitors), Anne-Laure Astolfi, Coline Bernard, Oriane Noyer, Marie-Hélène De Campo, Sandrine Margaroline, Louise N'Diaye, Sabine Perrier-Bonnet (Clinical Research nurses)
KARUPROSTATE
The Karuprostate study was supported by the the Frech National Health Directorate, the Association pour la Recherche sur le Cancer, la Ligue Nationale contre le Cancer, the French Agency for Environmental and Occupational Health Safety (ANSES) and by the Association pour la Recherche sur les Tumeurs de la Prostate. We would like to thank Séverine Ferdinand for valuable contributions to the study.
MOFFITT
The Moffitt group was supported by the US National Cancer Institute (R01CA128813, PI: J.Y. Park).
NMHS
Funding for the Nashville Men's Health Study (NMHS) was provided by the National Institutes of Health Grant numbers: RO1CA121060
PCaP
The North Carolina - Louisiana Prostate Cancer Project (PCaP) and the Health Care Access and Prostate Cancer Treatment in North Carolina (HCaP-NC) study are carried out as collaborative studies supported by the Department of Defense contract DAMD 17-03-2-0052 and the American Cancer Society award RSGT-08-008-01-CPHPS, respectively.
The authors thank the staff, advisory committees and research subjects participating in the PCaP and HCaP-NC studies for their important contributions.
PROtEuS
PROtEuS was supported financially through grants from the Canadian Cancer Society [13149, 19500, 19864, 19865] and the Cancer Research Society, in partnership with the Ministère de l'enseignement supérieur, de la recherche, de la science et de la technologie du Québec, and the Fonds de la recherche du Québec - Santé.PROtEuS would like to thank its collaborators and research personnel, and the urologists involved in subjects recruitment. We also wish to acknowledge the special contribution made by Ann Hsing and Anand Chokkalingam to the conception of the genetic component of PROtEuS.
SABOR
The SABOR research is supported by NIH/NCI Early Detection Research Network, grant U01 CA0866402-18. Also supported by the Cancer Center Support Grant to the Mays Cancer Center from the National Cancer Institute (US) P30 CA054174
SCCS
SCCS is funded by NIH grant R01 CA092447, and SCCS sample preparation was conducted at the Epidemiology Biospecimen Core Lab that is supported in part by the Vanderbilt-Ingram Cancer Center (P30 CA68485). Data on SCCS cancer cases used in this publication were provided by the Alabama Statewide Cancer Registry; Kentucky Cancer Registry, Lexington, KY; Tennessee Department of Health, Office of Cancer Surveillance; Florida Cancer Data System; North Carolina Central Cancer Registry, North Carolina Division of Public Health; Georgia Comprehensive Cancer Registry; Louisiana Tumor Registry; Mississippi Cancer Registry; South Carolina Central Cancer Registry; Virginia Department of Health, Virginia Cancer Registry; Arkansas Department of Health, Cancer Registry, 4815 W. Markham, Little Rock, AR 72205. The Arkansas Central Cancer Registry is fully funded by a grant from National Program of Cancer Registries, Centers for Disease Control and Prevention (CDC). Data on SCCS cancer cases from Mississippi were collected by the Mississippi Cancer Registry which participates in the National Program of Cancer Registries (NPCR) of the Centers for Disease Control and Prevention (CDC). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of the CDC or the Mississippi Cancer Registry.
SCPCS
SCPCS is funded by CDC grant S1135-19/19, and SCPCS sample preparation was conducted at the Epidemiology Biospecimen Core Lab that is supported in part by the Vanderbilt-Ingram Cancer Center (P30 CA68485).
SFPCS
SFPCS was funded by California Cancer Research Fund grant 99-00527V-10182
SWOG-PCPT / SWOG-SELECT
PCPT and SELECT are funded by Public Health Service grants U10CA37429 and 5UM1CA182883 from the National Cancer Institute. The authors thank the site investigators and staff and, most importantly, the participants from PCPT who donated their time to this trial.
UKGPCS
UKGPCS would also like to thank the following for funding support: The Institute of Cancer Research and The Everyman Campaign, The Prostate Cancer Research Foundation, Prostate Research Campaign UK (now Prostate Action), The Orchid Cancer Appeal, The National Cancer Research Network UK, The National Cancer Research Institute (NCRI) UK. We are grateful for support of NIHR funding to the NIHR Biomedical Research Centre at The Institute of Cancer Research and The Royal Marsden NHS Foundation Trust. UKGPCS should also like to acknowledge the NCRN nurses, data managers and Consultants for their work in the UKGPCS study.UKGPCS would like to thank all urologists and other persons involved in the planning, coordination, and data collection of the study. KM and AL were in part supported from the NIHR Manchester Biomedical Research Centre
WUGS
WUGS would like to thank the following for funding support: The Anthony DeNovi Fund, the Donald C. McGraw Foundation, and the St. Louis Men’s Group Against Cancer.
Footnotes
Conflict of Interest:
All authors declare no personal or financial conflicts of interest for the submitted work except as follows. CCF is a scientific consultant for CorTechs Labs, Inc. RE reports honorarium as a speaker for GU-ASCO meeting in San Francisco Jan 2016, support from Janssen, and honorarium as speaker for RMH-FR meeting Nov 2017. She reports honorarium as a speaker at the University of Chicago invited talk May 2018, and an educational honorarium by Bayer & Ipsen to attend GU Connect “Treatment sequencing for mCRPC patients within the changing landscape of mHSPC” at ESMO Barcelona, Sep 2019. She reports member of external Expert Committee on the Prostate Dx Advisory Panel. OAA received speaker’s honorarium from Lundbeck, and is a consultant for Healthlytix. AMD reports that he was a founder and holds equity in CorTechs Labs Inc., and serves on its Scientific Advisory Board. He is a member of the Scientific Advisory Board of Human Longevity, Inc., and the Mohn Medical Imaging and Visualization Centre. He received funding through research grants from GE Healthcare to UCSD. The terms of these arrangements have been reviewed by and approved by UCSD in accordance with its conflict of interest policies. TMS reports honoraria, outside of the present work, from: University of Rochester, Varian Medical Systems, Multimodal Imaging Servcies Corporation; and WebMD. He reports research funding from NIH/NBIB, U.S. Department of Defense, Radiological Society of North America, American Society for Radiation Oncology, and Varian Medical Systems.
Ethics Statement
All contributing studies were approved by the relevant ethics committees; written informed consent was obtained from the study participants21. The present analyses used de-identified data from the PRACTICAL consortium and have been approved by the review board at the corresponding authors’ institution.
Novelty and Impact
We identified three SNPs (rs76229939, rs74421890, and rs5013678), using cross-validation, that significantly improved performance of a previously published polygenic hazard model (PHS46) in men of African genetic ancestry. The performance of the novel PHS46+African score improved by as much as 79% after incorporation of rs76229939, rs74421890, and rs5013678. PHS46+African significantly improved association between polygenic risk with age at diagnosis of prostate cancer in Africans, a population generally under-served in genome-wide studies, to levels comparable with Europeans.
Data Availability Statement
The data used in this work were obtained from the Prostate Cancer Association Group to Investigate Cancer Associated Alterations in the Genome (PRACTICAL) consortium.
Readers who are interested in accessing the data must first submit a proposal to the Data Access Committee. If the reader is not a member of the consortium, their concept form must be sponsored by a principal investigator (PI) of one of the PRACTICAL consortium member studies. If approved by the Data Access Committee, PIs within the consortium, each of whom retains ownership of their data submitted to the consortium, can then choose to participate in the specific proposal. In addition, portions of the data are available for request from dbGaP (database of Genotypes and Phenotypes) which is maintained by the National Center for Biotechnology Information (NCBI): https://www.ncbi.nlm.nih.gov/gap/?term=Icogs+prostatehttps://www.ncbi.nlm.nih.gov/gap/?term=Icogs+prostate.
Anyone can apply to join the consortium. The eligibility requirements are listed here: http://practical.icr.ac.uk/blog/?page_id=9. Joining the consortium would not guarantee access, as a proposal for access would still be submitted to the Data Access Committee, but there would be no need for a separate member sponsor. Readers may find information about application by using the contact information below:
Rosalind Eeles
Principal Investigator for PRACTICAL
Professor of Oncogenetics
Institute of Cancer Research (ICR)
Sutton, UK
PRACTICAL@icr.ac.uk
URL: http://practical.icr.ac.uk
Tel: ++44 (0)20 8722 4094
References
- 1.Seibert TM, Fan CC, Wang Y, Zuber V, Karunamuni R, Parsons JK, Eeles RA, Easton DF, Kote-Jarai Z, Al Olama AA, Garcia SB, Muir K, Grönberg H, Wiklund F, Aly M, Schleutker J, Sipeky C, Tammela TLJ, Nordestgaard BG et al. Polygenic hazard score to guide screening for aggressive prostate cancer: Development and validation in large scale cohorts. BMJ 2018;360:1–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Duncan L, Shen H, Gelaye B, Meijsen J, Ressler K, Feldman M, Peterson R, Domingue B. Analysis of polygenic risk score usage and performance in diverse human populations. Nat Commun [Internet] 2019;10. Available from: 10.1038/s41467-019-11112-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Popejoy A, Fullerton S. Genomics is falling on diversity. Nature 2016;538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet [Internet] 2019;51:584–91. Available from: 10.1038/s41588-019-0379-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Huynh-Le M-P, Chieh Fan C, Karunamuni R, Martinez ME, Eeles RA, Kote-Jarai Z, Muir K, Collaborators U, Schleutker J, Pashayan N, Batra J, APCB (Australian Prostate Cancer Bioresource), Grönberg H, Neal DE, Donovan JL, Hamdy FC, Martin RM, Nielsen SF, Nordestgaard BG et al. Polygenic hazard score is associated with prostate cancer in multi-ethnic populations. medRxiv 2019; [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Amos CI, Dennis J, Wang Z, Byun J, Schumacher FR, Gayther SA, Casey G, Hunter DJ, Sellers TA, Gruber SB, Dunning AM, Michailidou K, Fachal L, Doheny K, Spurdle AB, Li Y, Xiao X, Romm J, Pugh E et al. The oncoarray consortium: A network for understanding the genetic architecture of common cancers. Cancer Epidemiol Biomarkers Prev 2017;26:126–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Infinium OncoArray-500K BeadChip ∣ Research cancer predisposition and risk [Internet]. [cited 2020 Jul 21];Available from: https://www.illumina.com/products/by-type/microarray-kits/infinium-oncoarray-500k.html
- 8.Leitzmann MF, Rohrmann S. Risk factors for the onset of prostatic cancer: Age, location, and behavioral correlates. Clin Epidemiol 2012;4:1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li Y, Byun J, Cai G, Xiao X, Han Y, Cornelis O, Dinulos JE, Dennis J, Easton D, Gorlov I, Seldin MF, Amos CI. FastPop: A rapid principal component derived method to infer intercontinental ancestry using genetic data. BMC Bioinformatics [Internet] 2016;17:1–8. Available from: 10.1186/s12859-016-0965-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Therneau TM, Li H. Computing the Cox Model for Case Cohort Designs. Lifetime Data Anal 1999;5:99–112. [DOI] [PubMed] [Google Scholar]
- 11.Harrell FE, Califf RM, Pryor DB, Lee KL, Rosati RA. Evaluating the Yield of Medical Tests. JAMA J Am Med Assoc 1982;247:2543–6. [PubMed] [Google Scholar]
- 12.Huynh-Le M-P, Fan CC, Karunamuni R, Walsh EI, Turner EL, Lane JA, Martin RM, Neal DE, Donovan JL, Hamdy FC, Parsons JKK, Eeles RA, Easton DF, Kote-Jarai Z, Amin Al Olama A, Benlloch Garcia S, Muir K, Grönberg H, Wiklund F et al. A genetic risk score to personalize prostate cancer screening, applied to population data. Cancer Epidemiol Biomarkers Prev 2020;cebp.1527.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.SEER*Explorer: An interactive website for SEER cancer statistics Surveillance Research Program, National Cancer Institute; [Internet]. [cited 2020 Jul 21];Available from: https://seer.cancer.gov/explorer/ [Google Scholar]
- 14.Sherry ST, Ward M, Sirotkin K. dbSNP - database for single nucleotide polymorphisms and other classes of minor genetic variation. Genome Res 1999;9:677–9. [PubMed] [Google Scholar]
- 15.rs76229939 RefSNP Report - dbSNP - NCBI [Internet]. [cited 2020 Jul 14];Available from: https://www.ncbi.nlm.nih.gov/snp/rs76229939 [Google Scholar]
- 16.rs74421890 RefSNP Report - dbSNP - NCBI [Internet]. [cited 2020 Jul 14];Available from: https://www.ncbi.nlm.nih.gov/snp/rs74421890 [Google Scholar]
- 17.rs5013678 RefSNP Report - dbSNP - NCBI [Internet]. [cited 2020 Jul 14];Available from: https://www.ncbi.nlm.nih.gov/snp/rs5013678 [Google Scholar]
- 18.Yeager M, Orr N, Hayes RB, Jacobs KB, Kraft P, Wacholder S, Minichiello MJ, Fearnhead P, Yu K, Chatterjee N, Wang Z, Welch R, Staats BJ, Calle EE, Feigelson HS, Thun MJ, Rodriguez C, Albanes D, Virtamo J et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet 2007;39:645–9. [DOI] [PubMed] [Google Scholar]
- 19.Haiman CA, Patterson N, Freedman ML, Myers SR, Pike MC, Waliszewska A, Neubauer J, Tandon A, Schirmer C, McDonald GJ, Greenway SC, Stram DO, Le Marchand L, Kolonel LN, Frasco M, Wong D, Pooler LC, Ardlie K, Oakley-Girvan I et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat Genet 2007;39:638–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Han Y, Rand KA, Hazelett DJ, Ingles SA, Kittles RA, Strom SS, Rybicki BA, Nemesure B, Isaacs WB, Stanford JL, Zheng W, Schumacher FR, Berndt SI, Wang Z, Xu J, Rohland N, Reich D, Tandon A, Pasaniuc B et al. Prostate cancer susceptibility in men of African ancestry at 8q24. J Natl Cancer Inst 2016;108:1–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kote-Jarai Z, Easton DF, Stanford JL, Ostrander EA, Schleutker J, Ingles SA, Schaid D, Thibodeau S, Dörk T, Neal D, Cox A, Maier C, Vogel W, Guy M, Muir K, Lophatananon A, Kedda MA, Spurdle A, Steginga S et al. Multiple novel prostate cancer predisposition loci confirmed by an international study: The PRACTICAL consortium. Cancer Epidemiol Biomarkers Prev 2008;17:2052–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data used in this work were obtained from the Prostate Cancer Association Group to Investigate Cancer Associated Alterations in the Genome (PRACTICAL) consortium.
Readers who are interested in accessing the data must first submit a proposal to the Data Access Committee. If the reader is not a member of the consortium, their concept form must be sponsored by a principal investigator (PI) of one of the PRACTICAL consortium member studies. If approved by the Data Access Committee, PIs within the consortium, each of whom retains ownership of their data submitted to the consortium, can then choose to participate in the specific proposal. In addition, portions of the data are available for request from dbGaP (database of Genotypes and Phenotypes) which is maintained by the National Center for Biotechnology Information (NCBI): https://www.ncbi.nlm.nih.gov/gap/?term=Icogs+prostatehttps://www.ncbi.nlm.nih.gov/gap/?term=Icogs+prostate.
Anyone can apply to join the consortium. The eligibility requirements are listed here: http://practical.icr.ac.uk/blog/?page_id=9. Joining the consortium would not guarantee access, as a proposal for access would still be submitted to the Data Access Committee, but there would be no need for a separate member sponsor. Readers may find information about application by using the contact information below:
Rosalind Eeles
Principal Investigator for PRACTICAL
Professor of Oncogenetics
Institute of Cancer Research (ICR)
Sutton, UK
PRACTICAL@icr.ac.uk
URL: http://practical.icr.ac.uk
Tel: ++44 (0)20 8722 4094