Abstract
Aberrantly activated Hedgehog (Hh) pathway is critical for driving the initiation and progression of multiple types of cancers, including medulloblastoma (MB) and basal cellular carcinoma (BCC). The majority of current Hh antagonist function by targeting the transmembrane domain of the oncoprotein Smoothened (Smo), a G-protein-coupled receptor-like receptor of Hh pathway. However, the primary and acquired resistance to current Smo inhibitors raise a critical need to develop next-generation of Smo inhibitors to improve their clinical efficacy. In this study, we identify that FDA approved drug ABT-199 significantly and selectively inhibits the Hh pathway. Mechanistically, ABT-199 acts as a competitive inhibitor of oxysterol by potentially targeting the cysteine rich domain (CRD) of Smo, rather as a BH3 mimetic. ABT-199 obviously inhibits the growth of Hh-driven tumors and possesses capacity of combating the primary and acquired resistance to Smo inhibitors caused by Smo mutations. Our data reposition ABT-199 as a Smo inhibitor for treating Hh-driven tumors, especially for those bearing Smo mutations and resistant to current Smo inhibitors. Meanwhile, our findings strengthen the argument that the CRD of Smo is a promising target for developing novel Smo inhibitors with capacity of combating the resistance to Smo inhibitors.
Keywords: Hedgehog, Smoothened, cysteine rich domain, medulloblastoma, basal cellular carcinoma
Introduction
The Hedgehog (Hh) signaling pathway is an evolutionarily conserved signal transduction cascade that is involved in embryonic patterning and tissue regeneration [1]. It is initiated by the binding of ligands (Shh, Dhh, and Ihh) to the receptor Patched 1 (Ptch1), a 12-transmembrane receptor protein. In the absence of ligands, Ptch1 suppresses the activity of the 7-transmembrane receptor protein Smoothened (Smo) and subsequent downstream signal transduction. The binding of ligands to Ptch1 relieves the suppressive effect of Ptch1, causing Smo to aggregate in the primary cilia of vertebrates, ultimately inducing the translocation of Gli proteins into the nucleus and the expression of Hh target genes, including Gli1 and Bcl-2 [2, 3].
Insufficient Hh activity is strongly implicated in human birth developmental defects, such as holoprosencephaly. However, its excessive activation during the postnatal period has been shown to be critically required for a variety of cancers, such as basal cell carcinoma (BCC) and medulloblastoma (MB) [2, 4]. Due to the critical roles of the Hh pathway in the initiation and progression of cancers, large-scale efforts have been devoted to developing Hh inhibitors for therapy of tumors addiction to the Hh pathway. The majority of current Hh inhibitors target Smo, an important oncoprotein and a member of the G-protein-coupled receptor (GPCR) subfamily. It harbors a conserved extracellular N-terminal cysteine rich domain (CRD), a heptahelical 7-transmembrane domain (7-TMD), and an intracellular C-terminal domain [5–8]. Among the reported Smo inhibitors, cyclopamine is the first identified naturally occurring antagonist of the Hh pathway. It has been shown to act by binding the 7-TMD of Smo, thereby stabilizing its inactive conformation [9]. To date, numerous small molecules have been developed and identified to modulate Hh activity by acting through the 7-TMD, including the specific Smo agonist SAG [10] and Smo antagonists, such as GDC-0449 (GDC) [11], NVP-LDE225 [12], and glasdegib [13]. Among these Smo antagonists, GDC and NVP-LDE225 have been approved for the treatment of locally advanced and metastatic BBC carrying ptch mutations [14], and glasdegib has been approved for the treatment of acute myeloid leukemia [15].
Unfortunately, similar to what has been observed for other mechanism-based anticancer drugs, tumors exhibit primary and acquired resistance to current Smo inhibitors. Studies have demonstrated that genetic alterations in the 7-TMD of Smo represent the most frequent mechanisms responsible for the primary and acquired resistance to current Smo inhibitors [16–18]. The genetic alterations in the 7-TMD of Smo may be classified into two types: (i) continuously constitutively activated Smo mutations that incur insensitivity to Smo inhibitors, such as Smo W535L, H231R, and W281C and (ii) spontaneous mutations in its drug binding pocket that interfere with the binding of Smo inhibitors, such as Smo D473H, F460L, and L412F [17, 18]. Hence, there is an urgent need to develop next-generation Smo inhibitors with alternative mechanisms to overcome resistance and improve clinical efficacy.
In addition to the 7-TMD, accumulating evidence has shown that the CRD represents an alternative binding site on Smo for ligands. Oxysterols, such as 20(S)-OHC, can engage the CRD and induce the accumulation of Smo in primary cilia, therefore functioning as specific Smo agonist [19–21]. Moreover, two Smo inhibitors, 20-keto-yne and 20(R)-yne [22], were found to function through the CRD and to be capable of attenuating in vitro Hh activity when Smo carries mutations in W535 (Smo M2; W535L) or D473 (D473H). This report indicates the possibility of using antagonists acting at the CRD to overcome resistance to current Smo inhibitors. However, observations on another CRD antagonist, 22-NHC, challenge this possibility, as it exhibits no inhibitory effect on Hh activity when Smo possesses a mutation in W535 (W535L) in vitro [20]. These data indicate that whether Smo inhibitors acting at the CRD of Smo might overcome the primary and acquired resistance to current Smo inhibitors remains to be better elucidated due to the discrepancy in in vitro efficacy results among CRD antagonists and the lack of in vivo evidence.
In this study, we found that ABT-199, a BH3 mimetic, suppressed Hh signaling through its function as a competitive inhibitor of oxysterol by likely targeting the CRD of Smo. This activity of ABT-199 was independent of its BH3 mimetic function. We further observed in vitro and in vivo that ABT-199 possessed ability to overcome resistance caused by mutations in Smo.
Materials and methods
Cell lines and reagents
NIH3T3, Light II, 293 T, and LS174T cells were obtained from American Type Culture Collection (ATCC; Manassas, VA, USA) and were routinely cultured according to the protocol provided by ATCC. Prostaglandin E2 (PGE2), 20-(S)-hydroxycholesterol(20(S)-OHC), SAG, ABT-199, GDC, GANT-61, cyclopamine, and JQ1 were purchased from Selleck Chemicals (Houston, TX, USA). Tumor necrosis factor α (TNF-α), BAY 11-8072, and H-89 were obtained from Beyotime (Suzhou, China). BODIPY-cyclopamine was purchased from BioVision (Milpitas, CA, USA).
ShhN-conditioned medium (ShhN CM) was collected as previously described [23]. Briefly, 293 T cells were transfected with the ShhN plasmid. Two days later, conditioned medium was collected and diluted prior to experiments.
Plasmids and lentivirus
The Gli1 luciferase reporter (8×GBS-luciferase) plasmid was a kindly gift from Dr. Hiroshi Sasaki. The luciferase reporter plasmids containing TCF/LEF, NF-κB, and TK-Renilla were purchased from Promega (Madison, WI, USA). The Gli2 (plasmid#17648), ShhN (plasmid#37680), and Cre-IRES (plasmid#30205) plasmids were obtained from Addgene (Cambridge, MA, USA). The Gli1 plasmid and human Smo plasmid were purchased from Origene (Rockville, MD, USA). The Smo mutant constructs were generated using a Quick Change Site-Directed Mutagenesis kit from Agilent (Santa Clara, CA, USA) and confirmed by DNA sequencing. Sufu-shRNA was purchased from Santa Cruz (Santa Cruz, CA, USA). Transient transfection was performed by Lipo2000 obtained from Thermo Fisher Scientific (Waltham, MA, USA) according to the manufacturer’s protocols.
Lentivirus infection
293 T cells were transfected with lentiviral pLKO.1 vector containing an shRNA to Sufu. Virus supernatant was collected 48 h after transfection. For virus infection, cells were coincubated with virus for 8 h, followed by selection for 72 h. Protein knockdown was confirmed by Western blot analysis.
Dual-luciferase reporter assay
Cells seeded in 96-well plates were treated with various compounds as indicated for 36 h. Dual-luciferase analysis was performed with a dual-luciferase reporter assay system (Promega) and a luminometer. After internal normalization of firefly luciferase with TK-Renilla luciferase for each well, relative luciferase values were obtained by normalization of Hh pathway-stimulated cells with unstimulated cells.
RT-qPCR
Total RNA was extracted with TRIzol obtained from Takara (Dalian, China) according to the manufacturer’s instructions. RNA reverse transcription and PCRs (Takara) were performed following standard protocols. Gene expression relative to GUSB expression was analyzed using the 2−ΔΔCt method. The primer sequences are listed in supporting information Table S1.
Fluorescent BODIPY-cyclopamine competition assays
293 T cells were seeded on glass coverslips and transfected with Smo plasmids. The cells were incubated with BODIPY-cyclopamine with or without additional compounds as indicated for 10 h. Cells were either subjected to flow cytometry assay or incubated with DAPI for microscopy analysis.
Sequencing transcriptomics
Gene expression in NIH3T3 cells after various treatments was measured by RNA-sequencing. Briefly, cells were harvested after 24 h of treatment for RNA-sequencing. Illumina sequencing was performed by OE Biotech Co., Ltd. (Shanghai, China) using the Illumina HiSeq 2500 (Illumina, Inc.). The data analyses were performed using edgeR software.
Western blot analysis
Cells were lysed in LDS buffer (Invitrogen) supplemented with a protease inhibitor cocktail, followed by centrifugation at 10,000 r.p.m. for 15 min at 4 °C and incubation at 95 °C for 10 min. The samples were subjected to SDS-PAGE and routinely immunoblotted with the indicated antibodies.
Tumor growth inhibition
Spontaneous primary MB containing a ptch+/− mutation was harvested from ptch+/−; p53−/− genetically engineered mice, which were obtained by crossing ptch+/− mice (Jackson Laboratory) with p53−/− mice (Jackson Laboratory). BCC models were established as previously reported [24]. Briefly, mice with K14-Cre-ER transgene (Jackson Laboratory), floxed p53 allele (Jackson Laboratory), and Ptch1+/− (Jackson Laboratory) alleles were crossed to generate K14-Cre-ER;Ptch1+/−; p53fl/fl mice, which were intraperitoneally treated with 100 μg·d−1 tamoxifen at the age of 6 weeks for three consecutive days. At the age of 8 weeks, K14-Cre-ER;Ptch1+/−; p53fl/fl mice were further exposed to 4 Gy of ionizing radiation. Visible BCCs usually appear at ~6 months of age [25]. For SmoA1 MB, spontaneous primary tumors were obtained from ND2:SmoA1 transgenic mice as previously described [26].
Male nude mice (4–5 weeks old) and NOD/SCID mice (4–5 weeks old) were purchased from Shanghai SLAC Laboratory Animal Co. and Shanghai Lingchang Biotechnology Co.(Shanghai, China), respectively. The spontaneous tumors described above were harvested and subcutaneously allografted into nude mice (MB) or NOD/SCID (BCC) mice. When the tumor volumes reached approximately 2000 mm3, they were harvested and further subcutaneously allografted into nude mice or NOD/SCID mice. When tumor volumes reached ~200 mm3, the mice were randomized into groups for treatments as indicated. Tumor size was measured with calipers every 3 d, and tumor volumes were calculated as follows: volume = 0.5 × width × length × length. Tumor tissues were harvested 4 h after the last dosage to examine Gli1 mRNA expression.
All in vivo efficacy studies were approved by and conformed to the policies and regulations of the Animal Care and Use Committee of Fudan University, China.
Statistical analysis
Data analysis was performed using Graph Pad Prism software. Significance values were assessed using one-way ANOVA or Student’s t-test. P values of less than 0.05 were considered significant difference (*P < 0.05; **P < 0.01; ***P < 0.001; nsP > 0.05).
Results
ABT-199 specifically inhibits Hh pathway activity
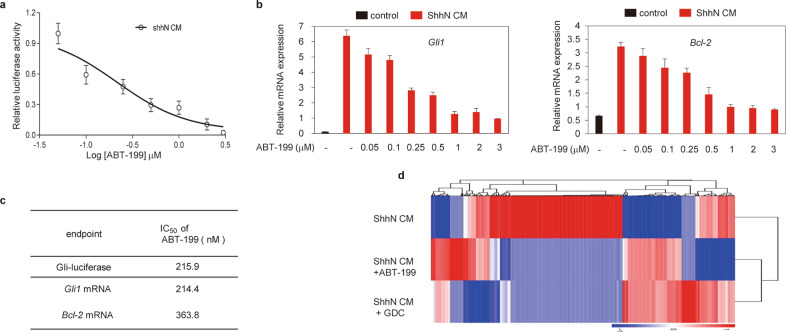
We previously reported that the Bcl-2 mimetic AT-101 inhibited the Hh pathway by targeting Smo [27]. We thus further tested the effect of all the other commercially available BH3 mimetics against Hh pathway activity using the Light II reporter cell line [28], a stable cloned mouse NIH3T3 cell line expressing Gli-luciferase and TK-Renilla. We found that ABT-199, a Bcl-2 inhibitor approved by the FDA for the treatment of chronic lymphocytic leukemia [29], potently suppressed Hh signaling induced by active amino-terminal variant of Shh (ShhN)-conditioned medium (ShhN CM) with an IC50 value of approximately 215.9 nM (Fig. 1a). To further confirm this ability of ABT-199, we examined its effect on the expression of Gli target genes Gli1 and Bcl-2 mRNA expression [30, 31]. Consistent with the observations in the Gli-luciferase assay, ABT-199 also blocked ShhN CM-induced Gli1 and Bcl-2 expression with IC50 values of 214.4 and 363.8 nM, respectively (Fig. 1b, c). To further confirm the inhibitory effect of ABT-199 on the Hh pathway, we used 3’-end polyadenylated RNA-sequencing analysis to examine the changes in global gene expression in NIH3T3 cells treated with ShhN CM in combination with ABT-199 or GDC (Fig. 1d). Prior to the RNA-sequencing analysis, we assessed the expression of Gli1 mRNA to determine the suitability of the samples (Supplementary Fig. S1a) and subjected the samples treated with ShhN CM, ShhN CM plus ABT-199 or ShhN CM plus GDC to RNA-sequencing analysis. We observed an obvious overlap in significantly differentially expressed genes, including Gli1, ptch1, and Bcl-2 (Fig. 1d, Supplementary material 1), between ShhN CM plus ABT-199-treated and ShhN CM plus GDC-treated cells compared with cells treated with ShhN CM. The inhibition of the Hh signaling pathway was not caused by cytotoxicity, as the viability of Light II cells showed no alterations when cells were exposed to ABT-199 at concentrations up to 10 μM for 72 h (data not shown). Collectively, these data demonstrate that ABT-199 is a potent and selective Hh inhibitor.
Fig. 1. ABT-199 specifically inhibits Hh pathway activity.
a Dual-luciferase reporter analysis of the inhibitory effect of ABT-199 on Hh activity in Light II cells exposed to ShhN CM with or without various concentrations of ABT-199 as indicated for 36 h. Data represent the mean ± SD (n = 3). b RT-qPCR analysis of the effect of ABT-199 on the mRNA expression of Gli1 (left) and Bcl-2 (right) in NIH3T3 cells exposed to ShhN CM with or without various concentrations of ABT-199 as indicated for 36 h. Data shown represent the mean ± SD (n = 3). c IC50 values of ABT-199 for the inhibition of Hh activity as determined in Fig. 1b and b. IC50 values were determined by nonlinear regression dose response fit. d Heatmap of significantly altered transcripts in NIH3T3 cells treated with ShhN CM with or without ABT-199 (1 μM) or GDC (100 nM) for 24 h. ShhN CM: ShhN-conditioned medium, GDC: GDC-0449.
To further determine the specific effect of ABT-199 against Hh activity, we explored the effect of ABT-199 on other unrelated signaling pathways, such as the NF-κB pathway. In contrast to the specific NF-κB inhibitor BAY 11-7082, we observed that ABT-199 failed to suppress the activation of NF-κB luciferase reporter activity induced by TNF-α (Supplementary Fig. S1b) at a concentration that completely inhibited Hh activity. Our laboratory has found that PGE2 may noncanonically induce Gli activity in a Smo-independent manner (data to be published), so we then asked whether ABT-199 influences noncanonical Gli activity in response to PGE2. The Gli-luciferase assay showed that the Gli inhibitor GANT-61 [32], but no tABT-199, significantly inhibited the Gli-luciferase activity stimulated by PGE2 (Supplementary Fig. S1c). We also found that ABT-199 failed to block TCF/LEF reporter activity stimulated by PGE2, with H-89 serving as a positive control (Supplementary Fig. S1d). These observations suggest that ABT-199 selectively inhibits Hh activity. To further strengthen this argument, we next examined whether ABT-199 may influence the transcript output of nontarget genes of Gli transcription factors, such as the critical components of the Hh pathway Smo and Sufu, as well as key regulators of the Wnt pathway, such as Dkk1, Axin, and Lgr5 [33]. Real-time quantitative polymerase chain reaction (RT-qPCR) analysis revealed that ABT-199, in contrast to the DNA transcription and replication inhibitor actinomycin D, exhibited no appreciable influence on the transcript output of the Hh pathway components Smo and Sufu (Supplementary Fig. S1e) or regulators of the Wnt pathway, such as Dkk1, Axin, and Lgr5 (Supplementary Fig. S1f). Taken together, these data suggest that ABT-199 selectively inhibits Hh activity.
ABT-199 inhibits the Hh pathway by acting at a level upstream of Sufu
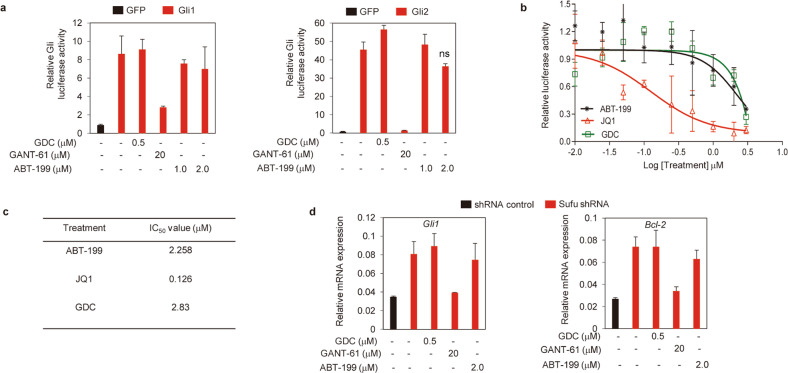
Having identified ABT-199 as a specific Hh inhibitor, we next set out to map its putative molecular target. Given that Gli1 and Gli2 are two positive transcription factors responsible for the output of the Hh pathway [34], we exogenously expressed Gli1 and Gli2 in Light II cells and investigated the effect of ABT-199 on Hh activity induced by the ectopic expression of Gli1 and Gli2 (Supplementary Fig. S2a). ABT-199 failed to inhibit Gli-luciferase activity induced by Gli1and Gli2 at concentrations that would completely inhibit Hh pathway activity in response to ShhN CM (Fig. 2a). This activity profile was similar to that of the Smo inhibitor GDC, whereas it contrasted with that of the specific Gli inhibitor GANT-61 (Fig. 2a), thus indicating that ABT-199 acts upstream of the transcription factor Gli.
Fig. 2. ABT-199 inhibits the Hh pathway by acting at a level upstream of Sufu.
a Dual-luciferase reporter analysis of Hh activity in Light II cells transfected with a Gli1 (left) or Gli2 (right) plasmid and treated with ABT-199, GDC, or GANT-61 as indicated for 36 h. nsP > 0.05. The data shown represent the mean ± SD (n = 3). b Dual-luciferase reporter analysis of Hh activity in Light II cells infected with lentivirus harboring Sufu-shRNA and exposed to various concentrations of ABT-199, GDC, or JQ1 as indicated for 36 h. The data shown represent the mean ± SD (n = 3). c IC50 values of the treatments in Fig. 2b. d RT-qPCR analysis of mRNA expression of Hh target genes (Gli1, Bcl-2) in Light II cells infected with Sufu-shRNA lentivirus after treatment with ABT-199, GDC, or JQ1 as indicated for 36 h. Data shown represent the mean ± SD (n = 3).
Sufu is a negative regulatory component of Hh pathway activity, which is located between Smo and Gli in the Hh signaling cascade. Limiting its expression may lead to elevation of Hh pathway activity [35]. Gli-luciferase analysis showed that ABT-199 exhibited a much weaker ability to inhibit Hh pathway activity induced by reducing the expression of Sufu with lentivirus containing short hairpin RNA targeting Sufu (Sufu-shRNA), as reflected by the much higher IC50 value when compared to that against Hh activity in response to ShhN CM (Figs. 2b, c, 1c, Supplementary Fig. S2b). This activity characteristic of ABT-199 mimicked that of GDC, whereas it was contrary to that of the Gli inhibitor JQ1 [36], when compared to their previously reported IC50 values against Hh activity stimulated by ShhN CM [11, 36] (Fig. 2b, c). We further confirmed these observations using transcripts of Gli1 and Bcl-2 as readouts of Hh pathway activity (Fig. 2d). Thus, these data suggest that ABT-199 predominantly functions by acting on a component upstream of Sufu.
ABT-199 does not engage the cyclopamine binding site
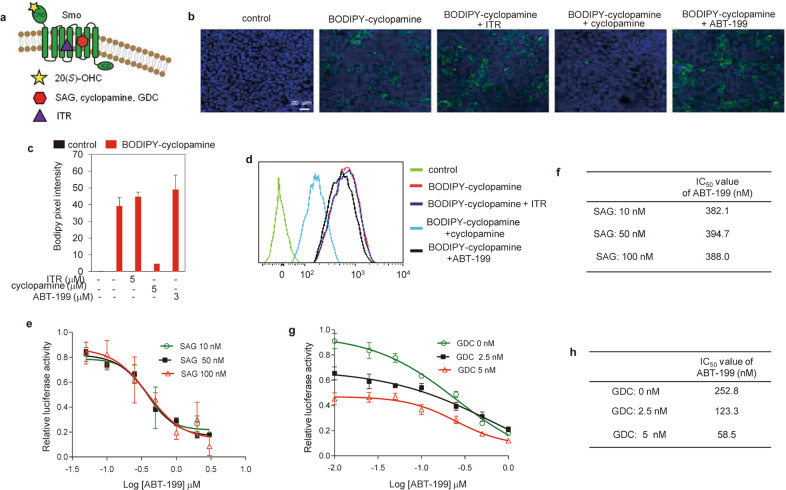
Having determined that ABT-199 inhibits the Hh pathway by acting at a level upstream of Sufu, we then set out to show whether it targets Smo, which is the target of the majority of current Hh inhibitors [8, 37]. Considering that the vast majority of Smo antagonists and agonists compete with cyclopamine, the first identified Smo inhibitor acting at the 7-TMD of Smo [8, 9] (hereafter referred to as cyclopamine binding site), we sought to determine whether ABT-199 engages the cyclopamine binding site (Fig. 3a). Using BODIPY-cyclopamine, a fluorescent analog of cyclopamine [9], as a molecular probe, we showed that cyclopamine dramatically reduced the binding of BODIPY-cyclopamine to Smo (Fig. 3b, c). Conversely, ABT-199 lacked the ability to interfere with the binding of BODIPY-cyclopamine, behaving like itraconazole (ITR), which is a Smo inhibitor with a distinct binding pocket on the 7-TMD from that of cyclopamine [38] (Fig. 3a–c). These observations were further confirmed by FACS analysis to detect the binding of BODIPY-cyclopamine to Smo (Fig. 3d). Therefore, our data indicate that ABT-199 does not share the same binding site as cyclopamine.
Fig. 3. ABT-199 does not engage the cyclopamine binding site.
a Schematic model depicting distinct binding sites of Smo ligands on Smo. b, c Representative images (b) and corresponding quantification (c) of the binding of BC to Smo in 293T cells. Cells were transfected with Smo and exposed to BODIPY-cyclopamine (1 μM) alone or in the presence of ABT-199 (3 μM), ITR (5 μM), or cyclopamine (5 μM) as indicated for 12 h. Scale bars = 20 μm. d Flow cytometry analysis of the binding of BODIPY-cyclopamine to Smo in 293T cells. e Dual-luciferase reporter analysis of the inhibitory effect of ABT-199 on Hh activity in Light II cells exposed to distinct concentrations of SAG with or without various concentrations of ABT-199 as indicated for 36 h. Data represent the mean ± SD (n = 3). f IC50 values of the effect of ABT-199 on Hh activity measured in Fig. 3e. g Dual-luciferase reporter analysis of the inhibitory effect of ABT-199 on Hh activity in Light II cells exposed to ShhN CM with or without various concentrations of ABT-199 combined with different concentrations of GDC as indicated for 36 h. Data represent the mean ± SD. h IC50 values of the inhibitory effect of ABT-199 on Hh activity measured in Fig. 3g. ITR: itraconazole.
To further reinforce this notion, we tested the ability of ABT-199 to inhibit Hh activity in response to distinct concentrations of SAG, a specific Smo agonist whose binding site on Smo overlaps with that of cyclopamine (Fig. 3a) [10]. We observed that ABT-199 obviously inhibited Hh activity in response to SAG at concentrations of 10, 50, and 100 nM, with no appreciable alterations in the IC50 values of ABT-199 (Fig. 3e, f). These observations suggest that ABT-199 functions as a noncompetitive inhibitor of Hh activation induced by SAG. Moreover, we measured the IC50 values of ABT-199 against Hh activity in response to ShhN CM in combinations of distinct concentrations of GDC, which shares the same binding pocket with cyclopamine (Fig. 3a) [8, 37]. We found that increasing concentrations of GDC enhanced the inhibitory effect of ABT-199 on Hh activity, as evidenced by the decreased IC50 values of ABT-199 (Fig. 3g, h). These data suggest that GDC possesses a distinct binding pocket from that of ABT-199 and that GDC has a synergistic inhibitory effect with ABT-199 on Hh activity in response to ShhN CM. Taken together, these data demonstrate that the cyclopamine binding site on Smo is not the drug binding pocket of ABT-199.
ABT-199 acts as a competitive inhibitor of oxysterol
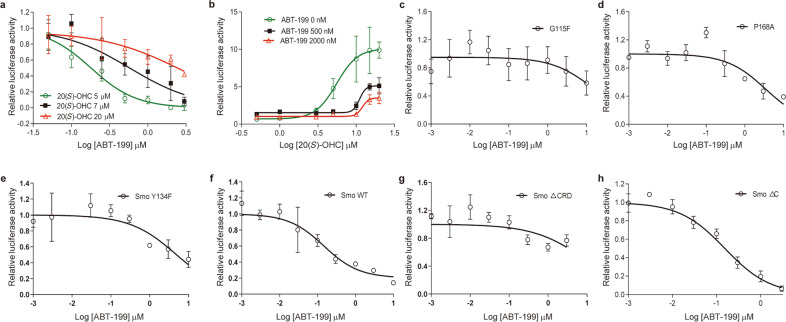
Considering that the oxysterol binding site on the CRD of Smo is an additional ligand binding pocket distinct from that of cyclopamine and itraconazole [19–21], we set out to explore whether ABT-199 inhibits the Hh pathway by engaging the oxysterol binding site. We first examined the ability of ABT-199 to inhibit Hh activity in response to 20(S)-OHC, which is a Smo agonist targeting the CRD of Smo (Fig. 3a) [20, 21]. ABT-199 drastically suppressed the Hh activity induced by 20(S)-OHC; however, increasing concentrations of 20(S)-OHC resulted in progressively higher IC50 values of ABT-199 (Fig. 4a, Supplementary Fig. S3a). These data indicate that ABT-199 acts as a competitive inhibitor of 20(S)-OHC and that the binding site of ABT-199 on the target protein potentially overlaps with that of 20(S)-OHC. To further validate this result, we tested the EC50 of 20(S)-OHC on stimulating Hh pathway activity in combination with different concentrations of ABT-199. 20(S)-OHC strongly increased the Hh pathway activity with an EC50 value of 5.54 μM when no ABT-199 was included. However, the addition of ABT-199 at concentrations of 0.5 and 2 μM increased the EC50 of 20(S)-OHC to 11.06 and 12.03 μM, respectively (Fig. 4b, Supplementary Fig. S3b). These observations further substantiate the observation that ABT-199 is a competitive inhibitor of 20(S)-OHC and that ABT-199 and 20(S)-OHC likely share the same drug binding pocket at the CRD of Smo.
Fig. 4. ABT-199 acts as a competitive inhibitor of oxysterol.
a Dual-luciferase reporter analysis of the activity of ABT-199 against Hh activity in Light II cells exposed to different concentrations of 20(S)-OHC with or without various concentrations of ABT-199 as indicated for 36 h. Data represent the mean ± SD (n = 3). b Dual-luciferase reporter analysis of the influence of different concentrations of ABT-199 on the ability of 20(S)-OHC to promote Hh activity. Light II cells were exposed to various concentrations of 20(S)-OHC with or without different concentrations of ABT-199 as indicated for 36 h. Data represent the mean ± SD (n = 3). c–h Dual-luciferase reporter analysis of the influence of distinct Smo mutants on the ability of ABT-199 to inhibit Hh activity in response to SAG. Light II cells were transfected with GFP or plasmids encoding Smo G115F (c), Smo P168A (d), Smo Y134F (e), Smo WT (f), Smo ΔCRD (g), or Smo ΔC (h), followed by exposure to SAG alone or in the presence of different concentrations of ABT-199 for 36 h as indicated. Data represent the mean ± SD (n = 3). 20(S)-OHC: 20-(S)-hydroxycholesterol.
Multiple amino acid residues located in the CRD of Smo have been identified to be critical for ligands binding to CRD, such as G115, P168, and Y134 [22]. We then asked whether these amino acid residues were essential for the inhibitory effect of ABT-199 on Hh activity. We found that exogenously expressed Smo constructs carrying mutations in G115 (Smo G115F), P168 (Smo P168A), and Y134 (Smo Y134F) in Light II cells largely reduced the ability of ABT-199 to inhibit Hh activity in response to SAG, as reflected by the increased IC50 values of ABT-199 when compared to that obtained by transfection of the wild-type Smo (Smo WT) construct (Fig. 4c–f, Supplementary Fig. S3c). The influence of Smo G115F, Smo P168A, and Smo Y134F on the inhibitory effect of ABT-199 on Hh activity is similar to that of Smo constructs with deletion of the CRD (Smo ΔCRD; Fig. 4g, Supplementary Fig. S3c). However, the Smo construct with deletion of the carboxyl terminal (Smo ΔC) had no effect on the efficacy of ABT-199 (Fig. 4h, Supplementary Fig. S3c). These data reinforce the conclusion that ABT-199 inhibits Hh activity as a competitive inhibitor of oxysterol by likely acting on the same drug binding pocket on CRD as oxysterol.
ABT-199 suppresses the growth of Hh-driven MB and BCC in vivo
To translate the in vitro ability of ABT-199 to inhibit Hh activity into an in vivo antitumor efficacy on Hh-dependent tumors, we established a spontaneous primary MB model from genetically engineered ptch+/−; p53−/− mice, a frequently used Hh-dependent tumor model driven by a ptch mutation [39]. ABT-199 at a dosage of 50 mg·kg−1 twice a day, comparable to the dosage for the treatment of chronic lymphoid leukemia in vivo [40], showed a remarkable inhibition of tumor growth compared to the vehicle control (Fig. 5a, b). Body weight loss was not observed during the treatment. Moreover, the inhibition of tumor growth paralleled the reduction in Hh activity, as evidenced by alterations in the expression of Gli1 mRNA (Fig. 5c). To further confirm the effect of ABT-199 on the growth inhibition of tumors associated with the Hh pathway, we also examined its impact on the growth of BCCs in K14-Cre-ER; ptch+/−; p53fl/fl mice, which is another commonly used model for evaluating the antitumor efficacy of Hh inhibitors [24, 38]. Consistent with the observations obtained from the MB model, ABT-199 treatment obviously delayed the growth of BCC tumors (Fig. 5d, e), concomitant with a reduction in Gli1 mRNA expression (Fig. 5f). These results demonstrate that ABT-199 may inhibit the growth of tumors dependent on Hh activity by suppressing Hh activity.
Fig. 5. ABT-199 suppresses the growth of Hh-driven MB and BCC in vivo.
a–c Effect of ABT-199 on MB growth as determined by tumor volume (a), growth inhibition rate (b), and Gli1 mRNA expression (c). Nude mice with MB allografts from Ptch+/−; p53−/− mice were treated with vehicle control (n = 6) or ABT-199 (n = 7) as indicated by gavage twice a day. Data on tumor volumes represent the mean ± SEM. d–f Effect of ABT-199 on BCC growth as demonstrated by tumor volume (d), growth inhibition rate (e), and Gli1 mRNA expression (f). NOD/SCID mice with BCC generated from K14-Cre-ER; Ptch1+/−; p53fl/fl mice were treated with vehicle or ABT-199 as indicated by gavage twice a day. Data on tumor volumes represent the mean ± SEM (n = 6). *P < 0.05, **P < 0.01, ***P < 0.001, nsP > 0.05. BCC: basal cell carcinoma, MB: medulloblastoma.
ABT-199 possesses the ability to overcome resistance to Smo inhibitors caused by Smo mutations
Smo mutations within its ligand binding pockets (LBP), for example, D473H, H231R, and W281C, as well as mutations in its important structural regions, such as W535L, L412F, and F460L, represent the predominant mechanisms underlying resistance to current Smo inhibitors [16–18]. Having demonstrated that ABT-199 functions as a competitive inhibitor of oxysterol, we thus continued to test its in vitro ability to overcome resistance to current Smo inhibitors caused by Smo mutations. ABT-199 obviously suppressed SAG-stimulated Hh activity in Light II cells exogenously expressing Smo constructs carrying mutations in either LBP, namely, D473H, H231R, and W281C (Fig. 6a–c), or in important structural regions, namely, W535L, L412F, and F460L (Fig. 6d–f). The potency of ABT-199 paralleled its potency against Hh activity elicited by transfection of Smo WT constructs (Fig. 6g) as well as that against Hh activity stimulated by ShhN CM, as reflected by the lack of significant differences between their IC50 values (Supplementary Fig. S4, Fig. 1c). Taken together, these data suggest that ABT-199 possesses an in vitro ability to overcome resistance to current Smo inhibitors caused by Smo mutations.
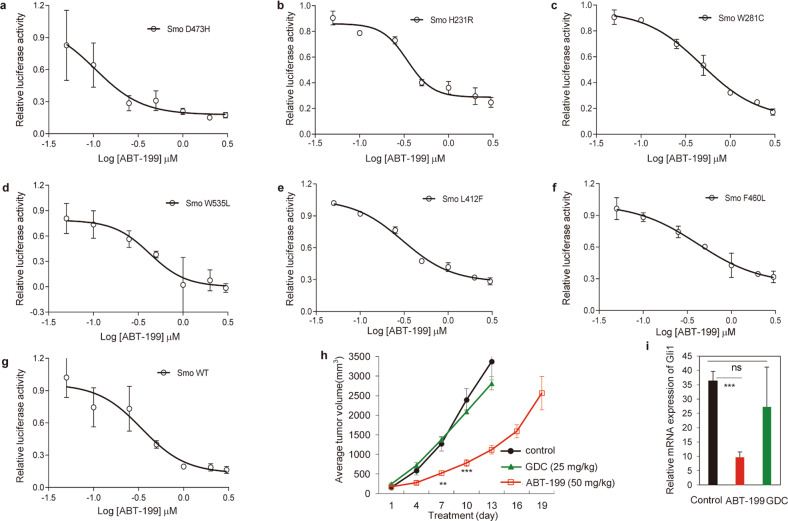
Fig. 6. ABT-199 possesses the ability to overcome the resistance to Smo inhibitors caused by Smo mutations.
a–g Dual-luciferase reporter analysis of the influence of distinct Smo mutants on the ability of ABT-199 to inhibit Hh activity in response to SAG. Light II cells were transfected with Smo D473H (a), Smo H231R (b), Smo W281C (c), Smo W535L (d), Smo L412F (e), Smo F460L (f), or Smo WT (g), followed by treatment with increasing concentrations of ABT-199 for 36 h. Data represent the mean ± SD (n = 3). h, i Effect of ABT-199 on SmoA1 MB tumor growth (h) and Gli1 mRNA expression (i) compared to vehicle control. Nude mice with SmoA1 MB allografts from ND2:SmoA1 transgenic mice were treated with vehicle control (n = 5), GDC (n = 6), or ABT-199 (n = 6) as indicated by gavage twice a day. Data on tumor growth represent the mean ± SEM. **P < 0.01, ***P < 0.001, nsP > 0.05.
To further test its in vivo capacity to overcome the resistance to Smo inhibitors caused by Smo mutations, we used spontaneous MB from ND2:SmoA1 transgenic mice, which contain the W539L mutation in the important structural regions of Smo [26]. ABT-199 treatment at 50 mg·kg−1 twice a day resulted in remarkable growth inhibition of SmoA1 tumors, whereas GDC exhibited no effect at a dosage of 25 mg·kg−1 twice a day, a dosage of GDC that would elicit 100% growth inhibition of tumors sensitive to Smo inhibitors [41] (Fig. 6h). Moreover, RT-qPCR analysis of the Gli1 mRNA levels in tumor tissues revealed that the effect of tumor growth inhibition of ABT-199 and GDC was consistent with their respective reductions of Gli1 mRNA (Fig. 6i), indicating that the tumor growth inhibitory effect of ABT-199 was caused by its inhibition of Hh activity. Collectively, these results show that ABT-199 possesses the ability to overcome resistance to current Smo inhibitors caused by Smo mutations.
Discussion
ABT-199 canonically acts as a Bcl-2 inhibitor through its BH3 mimetic function, thereby promoting apoptosis by inhibiting the prosurvival function of Bcl-2 [42]. ABT-199 has been approved by the FDA for the treatment of chronic lymphocytic leukemia with the 17p deletion and dependence on Bcl-2 [29]. In this study, we determined that ABT-199 significantly inhibited the Hh signaling pathway, with an IC50 value of ~300 nM. Moreover, the RNA-seq results and its inability to inhibit Wnt-, NF-κB-, and PGE2-induced noncanonical Gli activity showed that ABT-199 possesses selectivity against the Hh pathway. Mechanistically, ABT-199 acted as a competitive inhibitor of oxysterol rather than through its BH3 mimetic function. Furthermore, we found that ABT-199 possessed the ability to overcome resistance to current Smo inhibitors caused by Smo mutations in vitro and in vivo. Given that the tumor models used in this study were both driven by aberrant Hh activity, as well as the similarity between the tumor growth inhibition and the inhibitory effect on Hh activity elicited by ABT-199, we conclude that ABT-199 inhibits the growth of Hh-driven tumors shown in this study by its effect against Hh activity, rather by functioning as a BH3 mimetic. Hence, this study indicates the possibility of using ABT-199 for the treatment of tumors driven by the Hh pathway, especially for Smo inhibitor-resistant tumors caused by Smo mutations. Moreover, this study reinforces the argument that Smo inhibitors targeting Smo CRD possess the potential to overcome resistance to Smo inhibitors.
Smo is the most predominant target for developing anticancer drugs for the treatment of Hh-driven tumors. This GPCR-like receptor is comprised of an extracellular N-terminal region harboring a CRD, a 7-TMD and an intracellular C-terminal tail. The majority of Smo inhibitors are developed to target the 7-TMD of Smo and share the cyclopamine binding site [8, 43]. To our knowledge, there are only three Smo inhibitors targeting the CRD of Smo to date, 22-NHC, 20(R)-yne, and 20-keto-yne [20, 22]. In this study, we present evidence that ABT-199 acts as a competitive inhibitor of oxysterol, likely by engaging the CRD of Smo. Due to the large molecular weight of ABT-199, we failed to obtain direct evidence of it engaging the CRD using ABT-199 labeled with a radioactive isotope. In this context, we could not rule out the possibility that ABT-199 acts by indirectly targeting Smo, such as through a potential interaction with EVC proteins to influence Smo activity [44]. Compared to those of the three previously reported CRD inhibitors, the action profile of ABT-199 is most similar to that of 20(R)-yne and 20-keto-yne, especially in terms of the obvious inhibitory effect on Hh activity in response to either SAG or Smo mutants (SmoW539L), as 22-NHC does not affect Hh activity stimulated by SAG and SmoW539L [20, 22].
Wu et al. reported that Sufu, a critical negative regulator of Hh activity, harbors a Bcl-2 homology (BH) domain, by which the antiapoptotic Bcl-2 protein engages Sufu [45]. This engagement results in liberating Gli from its interaction with Sufu and ultimately activating the Hh pathway. They further found that ABT-199 might inhibit the Hh signaling pathway by disrupting the engagement of Bcl-2 to the BH domain of Sufu, acting as a BH3 mimetic. Hence, our current study, together with Wu’s report, suggests that ABT-199 might inhibit Hh activity by dual modes of action: a BH3 mimetic-dependent manner and BH3 mimetic-independent manner. It should be noted that the potency against Hh activity of ABT-199 as a competitive inhibitor of oxysterol, namely, in a BH3 mimetic-independent manner, is much stronger than that of its action as a BH3 mimetic, as evidenced by our observations that ABT-199 exhibited much weaker inhibition of Hh activity induced by limiting Sufu expression and ectopic expression of Gli proteins.
Supplementary information
Acknowledgements
This work was financially supported by grants from National Natural Science Foundation of China (No. 81773767, 81573452). We also appreciate Fudan-SIMM Joint Research Fund (No. FU-SIMM20181005), MHHFDU-SPFDU Joint Research Fund (No. RO-MY201805), and the fund from State Key Laboratory of Drug Research (No. SIMM1903KF-12).
Author contributions
WFT, JW, YZ, TMH, and WGT conceived and designed the project and wrote the manuscript. JW, YZ, WJH, and JY performed experiments and analyzed experiment results. WFT supervised the study.
Competing interests
The authors declare no competing interests.
Supplementary information
The online version of this article (10.1038/s41401-020-00504-4) contains supplementary material, which is available to authorized users.
References
- 1.Lee RT, Zhao Z, Ingham PW. Hedgehog signalling. Development. 2016;143:367–72. doi: 10.1242/dev.120154. [DOI] [PubMed] [Google Scholar]
- 2.Briscoe J, Therond PP. The mechanisms of Hedgehog signalling and its roles in development and disease. Nat Rev Mol Cell Biol. 2013;14:416–29. doi: 10.1038/nrm3598. [DOI] [PubMed] [Google Scholar]
- 3.Liu X, Ding C, Tan W, Zhang A. Medulloblastoma: molecular understanding, treatment evolution, and new developments. Pharmacol Ther. 2020;210:107516. 10.1016/j.pharmthera.2020.107516. [DOI] [PubMed]
- 4.Jiang J, Hui CC. Hedgehog signaling in development and cancer. Dev Cell. 2008;15:801–12. doi: 10.1016/j.devcel.2008.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Quaglio D, Infante P, Di Marcotullio L, Botta B, Mori M. Hedgehog signaling pathway inhibitors: an updated patent review (2015-present) Expert Opin Ther Pat. 2020;30:235–50.. doi: 10.1080/13543776.2020.1730327. [DOI] [PubMed] [Google Scholar]
- 6.Xin M, Ji X, De La Cruz LK, Thareja S, Wang B. Strategies to target the Hedgehog signaling pathway for cancer therapy. Med Res Rev. 2018;38:870–913. doi: 10.1002/med.21482. [DOI] [PubMed] [Google Scholar]
- 7.Rimkus TK, Carpenter RL, Qasem S, Chan M, Lo HW. Targeting the sonic Hedgehog signaling pathway: review of Smoothened and GLI inhibitors. Cancers (Basel). 2016;8:22. 10.3390/cancers8020022. [DOI] [PMC free article] [PubMed]
- 8.Sharpe HJ, Wang W, Hannoush RN, de Sauvage FJ. Regulation of the oncoprotein Smoothened by small molecules. Nat Chem Biol. 2015;11:246–55. doi: 10.1038/nchembio.1776. [DOI] [PubMed] [Google Scholar]
- 9.Chen JK, Taipale J, Cooper MK, Beachy PA. Inhibition of Hedgehog signaling by direct binding of cyclopamine to Smoothened. Genes Dev. 2002;16:2743–8. doi: 10.1101/gad.1025302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen JK, Taipale J, Young KE, Maiti T, Beachy PA. Small molecule modulation of Smoothened activity. Proc Natl Acad Sci USA. 2002;99:14071–6. doi: 10.1073/pnas.182542899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Robarge KD, Brunton SA, Castanedo GM, Cui Y, Dina MS, Goldsmith R, et al. GDC-0449-a potent inhibitor of the hedgehog pathway. Bioorg Med Chem Lett. 2009;19:5576–81. doi: 10.1016/j.bmcl.2009.08.049. [DOI] [PubMed] [Google Scholar]
- 12.Pan S, Wu X, Jiang J, Gao W, Wan Y, Cheng D, et al. Discovery of NVP-LDE225, a potent and selective smoothened antagonist. ACS Med Chem Lett. 2010;1:130–4. doi: 10.1021/ml1000307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hoy SM. Glasdegib: first global approval. Drugs. 2019;79:207–13.. doi: 10.1007/s40265-018-1047-7. [DOI] [PubMed] [Google Scholar]
- 14.Sekulic A, Von Hoff D. Hedgehog pathway inhibition. Cell. 2016;164:831. doi: 10.1016/j.cell.2016.02.021. [DOI] [PubMed] [Google Scholar]
- 15.Norsworthy KJ, By K, Subramaniam S, Zhuang L, Del Valle PL, Przepiorka D, et al. FDA approval summary: Glasdegib for newly diagnosed acute myeloid leukemia. Clin Cancer Res. 2019;25:6021–5. doi: 10.1158/1078-0432.CCR-19-0365. [DOI] [PubMed] [Google Scholar]
- 16.Metcalfe C, de Sauvage FJ. Hedgehog fights back: mechanisms of acquired resistance against Smoothened antagonists. Cancer Res. 2011;71:5057–61. doi: 10.1158/0008-5472.CAN-11-0923. [DOI] [PubMed] [Google Scholar]
- 17.Sharpe HJ, Pau G, Dijkgraaf GJ, Basset-Seguin N, Modrusan Z, Januario T, et al. Genomic analysis of smoothened inhibitor resistance in basal cell carcinoma. Cancer Cell. 2015;27:327–41. doi: 10.1016/j.ccell.2015.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Atwood SX, Sarin KY, Whitson RJ, Li JR, Kim G, Rezaee M, et al. Smoothened variants explain the majority of drug resistance in basal cell carcinoma. Cancer Cell. 2015;27:342–53. doi: 10.1016/j.ccell.2015.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Huang P, Nedelcu D, Watanabe M, Jao C, Kim Y, Liu J, et al. Cellular cholesterol directly activates Smoothened in Hedgehog signaling. Cell. 2016;166:1176–87.e14. doi: 10.1016/j.cell.2016.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nedelcu D, Liu J, Xu Y, Jao C, Salic A. Oxysterol binding to the extracellular domain of Smoothened in Hedgehog signaling. Nat Chem Biol. 2013;9:557–64. doi: 10.1038/nchembio.1290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nachtergaele S, Mydock LK, Krishnan K, Rammohan J, Schlesinger PH, Covey DF, et al. Oxysterols are allosteric activators of the oncoprotein Smoothened. Nat Chem Biol. 2012;8:211–20. doi: 10.1038/nchembio.765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nachtergaele S, Whalen DM, Mydock LK, Zhao Z, Malinauskas T, Krishnan K, et al. Structure and function of the Smoothened extracellular domain in vertebrate Hedgehog signaling. eLife. 2013;2:e01340. doi: 10.7554/eLife.01340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Maity T, Fuse N, Beachy PA. Molecular mechanisms of Sonic hedgehog mutant effects in holoprosencephaly. Proc Natl Acad Sci USA. 2005;102:17026–31. doi: 10.1073/pnas.0507848102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tang JY, Xiao TZ, Oda Y, Chang KS, Shpall E, Wu A, et al. Vitamin D3 inhibits hedgehog signaling and proliferation in murine Basal cell carcinomas. Cancer Prev Res (Philos) 2011;4:744–51. doi: 10.1158/1940-6207.CAPR-10-0285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Xiao TZ, Tang JY, Wu A. Hedgehog signaling of BCC is inhibited by Vitamin D: implication for a chemopreventive agent against BCC carcinogenesis. J Invest Dermatol. 2009;129:S32. [Google Scholar]
- 26.Hatton BA, Villavicencio EH, Tsuchiya KD, Pritchard JI, Ditzler S, Pullar B, et al. The Smo/Smo model: Hedgehog-induced medulloblastoma with 90% incidence and leptomeningeal spread. Cancer Res. 2008;68:1768–76. doi: 10.1158/0008-5472.CAN-07-5092. [DOI] [PubMed] [Google Scholar]
- 27.Wang J, Peng Y, Liu Y, Yang J, Huang M, Tan W. AT-101 inhibits Hedgehog pathway activity and cancer growth. Cancer Chemother Pharmacol. 2015;76:461–9. doi: 10.1007/s00280-015-2812-x. [DOI] [PubMed] [Google Scholar]
- 28.Taipale J, Chen JK, Cooper MK, Wang B, Mann RK, Milenkovic L, et al. Effects of oncogenic mutations in Smoothened and Patched can be reversed by cyclopamine. Nature. 2000;406:1005–9. doi: 10.1038/35023008. [DOI] [PubMed] [Google Scholar]
- 29.Deeks ED. Venetoclax: first global approval. Drugs. 2016;76:979–87. doi: 10.1007/s40265-016-0596-x. [DOI] [PubMed] [Google Scholar]
- 30.Dai P, Akimaru H, Tanaka Y, Maekawa T, Nakafuku M, Ishii S. Sonic Hedgehog-induced activation of the Gli1 promoter is mediated by GLI3. J Biol Chem. 1999;274:8143–52. doi: 10.1074/jbc.274.12.8143. [DOI] [PubMed] [Google Scholar]
- 31.Bigelow RL, Chari NS, Unden AB, Spurgers KB, Lee S, Roop DR, et al. Transcriptional regulation of bcl-2 mediated by the sonic hedgehog signaling pathway through gli-1. J Biol Chem. 2004;279:1197–205. doi: 10.1074/jbc.M310589200. [DOI] [PubMed] [Google Scholar]
- 32.Lauth M, Bergstrom A, Shimokawa T, Toftgard R. Inhibition of GLI-mediated transcription and tumor cell growth by small-molecule antagonists. Proc Natl Acad Sci U S A. 2007;104:8455–60. doi: 10.1073/pnas.0609699104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nusse R, Clevers H. Wnt/beta-catenin signaling, disease, and emerging therapeutic modalities. Cell. 2017;169:985–99.. doi: 10.1016/j.cell.2017.05.016. [DOI] [PubMed] [Google Scholar]
- 34.Hui CC, Angers S. Gli proteins in development and disease. Annu Rev Cell Dev Biol. 2011;27:513–37. doi: 10.1146/annurev-cellbio-092910-154048. [DOI] [PubMed] [Google Scholar]
- 35.Kogerman P, Grimm T, Kogerman L, Krause D, Unden AB, Sandstedt B, et al. Mammalian suppressor-of-fused modulates nuclear-cytoplasmic shuttling of Gli-1. Nat Cell Biol. 1999;1:312–9. doi: 10.1038/13031. [DOI] [PubMed] [Google Scholar]
- 36.Tang Y, Gholamin S, Schubert S, Willardson MI, Lee A, Bandopadhayay P, et al. Epigenetic targeting of Hedgehog pathway transcriptional output through BET bromodomain inhibition. Nat Med. 2014;20:732–40. doi: 10.1038/nm.3613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Frank-Kamenetsky M, Zhang XM, Bottega S, Guicherit O, Wichterle H, Dudek H, et al. Small-molecule modulators of Hedgehog signaling: identification and characterization of Smoothened agonists and antagonists. J Biol. 2002;1:10. doi: 10.1186/1475-4924-1-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kim J, Tang JY, Gong R, Kim J, Lee JJ, Clemons KV, et al. Itraconazole, a commonly used antifungal that inhibits Hedgehog pathway activity and cancer growth. Cancer Cell. 2010;17:388–99. doi: 10.1016/j.ccr.2010.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Romer JT, Kimura H, Magdaleno S, Sasai K, Fuller C, Baines H, et al. Suppression of the Shh pathway using a small molecule inhibitor eliminates medulloblastoma in Ptc1(+/−)p53(−/−) mice. Cancer Cell. 2004;6:229–40. doi: 10.1016/j.ccr.2004.08.019. [DOI] [PubMed] [Google Scholar]
- 40.Cang S, Iragavarapu C, Savooji J, Song Y, Liu D. ABT-199 (venetoclax) and BCL-2 inhibitors in clinical development. J Hematol Oncol. 2015;8:129. doi: 10.1186/s13045-015-0224-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Petricci E, Manetti F. Targeting the Hedgehog signaling pathway with small molecules from natural sources. Curr Med Chem. 2015;22:4058–90. doi: 10.2174/0929867322666150904111042. [DOI] [PubMed] [Google Scholar]
- 42.Souers AJ, Leverson JD, Boghaert ER, Ackler SL, Catron ND, Chen J, et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat Med. 2013;19:202–8. doi: 10.1038/nm.3048. [DOI] [PubMed] [Google Scholar]
- 43.Pak E, Segal RA. Hedgehog signal transduction: key players, oncogenic drivers, and cancer therapy. Dev Cell. 2016;38:333–44. doi: 10.1016/j.devcel.2016.07.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dorn KV, Hughes CE, Rohatgi R. A Smoothened-Evc2 complex transduces the Hedgehog signal at primary cilia. Dev Cell. 2012;23:823–35. doi: 10.1016/j.devcel.2012.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wu X, Zhang LS, Toombs J, Kuo YC, Piazza JT, Tuladhar R, et al. Extra-mitochondrial prosurvival BCL-2 proteins regulate gene transcription by inhibiting the SUFU tumour suppressor. Nat Cell Biol. 2017;19:1226–36.. doi: 10.1038/ncb3616. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.