Abstract
The plant-derived sesquiterpene lactone micheliolide was recently found to possess promising antileukemic activity, including the ability to target and kill leukemia stem cells. Efforts toward improving the biological activity of micheliolide and investigating its mechanism of action have been hindered by the paucity of preexisting functional groups amenable for late-stage derivatization of this molecule. Here, we report the implementation of a probe-based P450 fingerprinting strategy to rapidly evolve engineered P450 catalysts useful for the regio- and stereoselective hydroxylation of micheliolide at two previously inaccessible aliphatic positions in this complex natural product. Via P450-mediated chemoenzymatic synthesis, a broad panel of novel micheliolide analogs could thus be obtained to gain structure–activity insights into the effect of C2, C4, and C14 substitutions on the antileukemic activity of micheliolide, ultimately leading to the discovery of “micheliologs” with improved potency against acute myelogenic leukemia cells. These late-stage C–H functionalization routes could be further leveraged to generate a panel of affinity probes for conducting a comprehensive analysis of the protein targeting profile of micheliolide in leukemia cells via chemical proteomics analyses. These studies introduce new micheliolide-based antileukemic agents and shed new light onto the biomolecular targets and mechanism of action of micheliolide in leukemia cells. More broadly, this work showcases the value of the present P450-mediated C–H functionalization strategy for streamlining the late-stage diversification and elucidation of the biomolecular targets of a complex bioactive molecule.
Short abstract
A P450-based chemoenzymatic C−H functionalization strategy was applied to discover improved analogs of the anticancer terpene micheliolide and to profile its target proteome in leukemia stem cells.
Introduction
Sesquiterpene lactones (SQLs) constitute a family of bioactive natural products isolated from various plants.1 Members of this family containing an α-methylene-γ-lactone moiety have recently attracted significant attention because of their promising anticancer properties, especially in the context of leukemias.2−4 Previous studies indeed showed that parthenolide (PTL) and semisynthetic derivatives thereof are capable of targeting and killing acute myeloid leukemia (AML) cells as well as leukemia stem cells (LSCs).5−10 Due to the role of LSCs in self-renewal, disease initiation, and disease propagation11,12 but low sensitivity to mainstay AML chemotherapeutic agents (e.g., daunorubicin and cytarabine),13,14 selective eradication of LSCs represents an important goal in the development of next-generation antileukemic drugs.15−19 Similar to parthenolide, the SQL micheliolide (MCL, 1; Scheme 1) was recently found to possess promising activity and selectivity against AML cells and LSCs.20,21 In addition to AML, this natural product was shown to possess promising in vitro and in vivo activity in the context of other forms of cancer.22−25 While showing lower antileukemic activity than PTL (∼3 fold higher LC50 in vitro), MCL exhibits several favorable features such as higher chemical stability, water solubility, and an improved pharmacokinetic profile in vivo,20 which make it a promising scaffold for the development of antileukemic agents. The anti-inflammatory and anticancer activities of SQLs have been linked to their reactive α-methylene-γ-lactone “warhead”, which can act as a Michael acceptor resulting in covalent inhibition of target proteins.26−28 While chemical modification of this reactive moiety provides a facile route for the functionalization of these molecules, modifications at this site typically result in significant reduction or loss of biological activity.7,20,29,30 Indeed, tolerated substitutions at this site in both PTL and MCL have been largely limited to dimethylamine adducts (Scheme 1A), which have improved oral bioavailability but show similar or lower anticancer activity than the parent natural product.6,20 In MCL, the hydroxyl group at C4 is also available for late-stage functionalization purposes (Scheme 1A) and C4-substituted analogs of MCL were investigated by Chen and co-workers.31 However, none of these C4-substituted analogs showed improved antileukemic activity compared to MCL, further highlighting the challenges of improving the anticancer potency of this natural product using currently available methods.
Scheme 1. Micheliolide (MCL) and Chemical vs Chemoenzymatic Synthesis of Micheliolide Analogs.
P450 enzymes constitute attractive biocatalysts for the selective oxyfunctionalization of organic molecules,32−34 and protein engineering has provided a means to expand the substrate scope of these enzymes to include non-native substrates, including natural products.9,35−42 Here, we report the successful application of engineered P450s and P450-mediated chemoenzymatic C–H functionalization9,38,43 as a strategy to access novel—and previously inaccessible sites—for the late-stage elaboration of MCL (Scheme 1B). As fine-tuning of the regio- and stereoselectivity of P450 enzymes remains a major challenge,32,33 we have implemented a novel probe-based fingerprinting approach for guiding the development of highly regio- and stereoselective P450 catalysts for the chemoenzymatic functionalization of this complex natural product. Using these selective P450 catalysts, a broad panel of semisynthetic MCL analogs could be obtained to illuminate structure–activity relationships of modifications at multiple sites within the MCL scaffold (Scheme 1B) and enable the development of MCL-based agents with improved antileukemic activity. In addition, using this chemoenzymatic strategy, a complementary set of MCL-based affinity reagents could be generated and leveraged to perform a comprehensive profiling of the MCL-targeted proteome in leukemia stem cells. Our results show that the present strategy was both effective and critical toward (i) enabling the identification of “hot spots” for potentiating the antileukemic activity of MCL, (ii) discovering improved MCL-based antileukemic agents, and (iii) permitting a comprehensive analysis of the MCL-targeted proteome in leukemia stem cells, including putative targets which would have been overlooked through derivatization of the parent natural product alone.
Results and Discussion
Micheliolide Oxidation via a Substrate Promiscuous P450BM3 Variant
In previous work, we found that FL#62, an engineered variant of the fatty acid monooxygenase P450BM3 (Bacillus megaterium),44,45 is able to oxidize parthenolide9 and a broad range of bulky terpenes.38,46 Based on these results, we envisioned that this P450 variant could also accept micheliolide (MCL, 1) as a substrate for oxidation. Indeed, FL#62 was found to support the oxidation of MCL with 280 turnovers (TON) in the presence of a NADPH cofactor regeneration system. From these enzymatic reactions, three mono-oxygenated products were produced in an 87:7:6 ratio, as determined by gas chromatography (GC) (Scheme 2). Structural elucidation of these compounds by NMR revealed that the major product corresponds to 2(R)-hydroxy-micheliolide (2, 87%), whereas the minor products correspond to 14-hydroxy-micheliolide (3, 7%) and 14-formyl-micheliolide (4, 6%), respectively. The R configuration of the C2 carbon in 2 was assigned based on NMR NOE experiments, which revealed a strong NOE between the 5(H) and the 2β(H) protons and the absence of an NOE between the 6(H) and 2α(H) protons (Figures S1–S2). The identity of compound 3 was further confirmed by X-ray crystallography (Scheme 2, Figure S3). Unlike FL#62, wild-type P450BM3 enzyme showed negligible oxidation activity on MCL (<1 TON).
Scheme 2. Enzymatic Oxidation of MCL (1) using P450BM3 Variant FL#62.
Product distribution: 87% 2; 7% 3; 6% 4. The X-ray crystal structure of 3 (CCDC 2033958) is also shown.
Interestingly, compounds 2, 3, and 4 are not naturally occurring derivatives of MCL, nor have they been previously obtained using synthetic means. The hydroxylation products 2(R)-hydroxy-MCL (2) and 14-hydroxy-MCL (3) were considered particularly valuable as they provide two new points of entry for late-stage functionalization of the MCL scaffold at two remote C(sp3)—H sites relative to the preexisting hydroxyl group at C4 and the reactive α-methylene-γ-lactone (C13 site). 14-Formyl-micheliolide (4) likely arises from a second P450-catalyzed oxidation of the C14 site in 3 to give a C14 gem-diol, which yields the aldehyde product via dehydration. Compound 4 can be reduced in the presence of NaBH4 in methanol to give the alcohol product 3 (data not shown). This notwithstanding, the poor regio- and chemoselectivity of FL#62 toward MCL oxidation was limiting, in particular for functionalization of the C14 site. Accordingly, FL#62 was chosen as a promising starting point for evolving P450 variants with improved site-selectivity for each of the targeted aliphatic C–H sites.
Probe-Guided Search of Regioselective P450 Catalysts for MCL Oxidation
In previous work, we have generated a collection of over 800 functionally diverse P450 variants via site-saturation mutagenesis of first-sphere active-site residues (74, 75, 78, 81, 82, 87, 180, 181, 184, 263, and 328; Figure 1b) in FL#62 (or derivative thereof), followed by high-throughput fingerprinting of these variants using a panel of structurally diverse chromogenic probes.9,38,46 As part of this work, we have reported methods for predicting P450 reactivity based on the analysis of their fingerprints via single component (SCA)9,46 and multicomponent analysis (MCA).9,38 In the interest of expanding the toolbox of high throughput methods for expediting the development of P450 catalysts with fine-tuned selectivity, we sought to develop a complementary fingerprint-based strategy (called “regio-fingerprinting”) that could concurrently relay information about the P450 oxidation activity on a target substrate (i.e., MCL) and the regioselectivity of the P450 enzymes on such substrate.
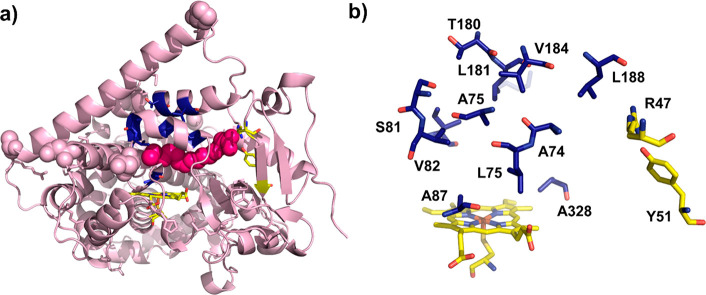
Figure 1.
(a) Crystal structure of the heme domain of P450BM3 in complex with N-palmitoyl glycine (PDB code 1JPZ). The heme and bound substrate are colored in yellow and magenta (space-filling model), respectively. (b) Model of the P450BM3 variant FL#62 active site with key residues around the heme cofactor (yellow) displayed as stick models.
As outlined in Figure 2a, this method relies on surrogate probes derived from the target substrate, in which a methoxy “reporter” group46 is installed at the C–H site of interest (i.e., C2 or C14 in MCL). In this approach, P450-dependent demethylation activity on the probe substrate, which is quantifiable via a Purpald-based colorimetric assay compatible with cell lysate and high-throughput screening,38,46 was expected to provide information about the oxidation activity of the P450 on the parent molecule (i.e., MCL) and its regioselectivity toward the corresponding C–H site in the unfunctionalized scaffold (Figure 2a). According to these design principles, we synthesized two MCL-based probes containing a methoxy (—OMe) group at either C2 (P1) or C14 (P2) position (Figure 2a) via methylation of 2 and 3, respectively. Next, we screened the library of ∼800 functionally diverse P450 variants against the two MCL-based probes P1 and P2, in parallel, using 96-well plates (E. coli DH5α cells). The enzymatic activity on each probe was quantified based on colorimetric measurement of the formaldehyde product generated upon demethylation of the reporter methoxy group. For each variant, the probe activity was normalized to that of a reference P450 (P450BM3(F87A)), which has low yet detectable activity on both probes (Figure 2b). A threshold of >20% relative activity compared to the reference P450BM3(F87A) was applied to discriminate MCL-active P450 variants from MCL-inactive ones. The probe-based regioselectivity of the P450 variants was then calculated based on their relative activity on P1 and P2 using the equation (AP1 × 100)/Atot or (AP2 × 100)/Atot, where AP1 and AP2 are the normalized demethylation activity of each variant on P1 and P2, respectively, and Atot is the total probe activity. A probe selectivity of >60% toward P1 (i.e., AP1/Atot > 0.6) or toward P2 (i.e., AP1/Atot < 0.4) was applied as a predictor for high regioselectivity toward the oxidation of aliphatic positions C2 and C14, respectively, in MCL.
Figure 2.
Fingerprint-guided prediction of P450 regioselectivity. (a) Overview of high-throughput “regio-fingerprinting” method using the MCL-based chromogenic probes P1 and P2. High P1- or P2-selectivity is used a predictor of C2- or C14-selectivity, respectively, for MCL hydroxylation. (b) Total probe activity with relative contribution from P1 and P2 activity for a representative set of P450 variants. Probe activity was normalized to the reference enzyme P450BM3(F87A), which was included in the screening. (c) Percentage of correct regiofingerprint-guided predictions for C2-selective variants (AP1/Atot > 0.6, C14-selective variants (AP2/Atot > 0.6), and unselective variants (0.6 > AP1/Atot > 0.4).
To experimentally test the predictions, we selected the top 15–20 variants favoring either probe (i.e., AP1/Atot > 0.6 or < 0.4), along with several nonselective variants (0.6 > AP1/Atot > 0.4) with variable total probe demethylation activity (Table S1). A total of 50 variants that met these criteria were expressed in 48-well plates and analyzed for their activity and regioselectivity toward MCL hydroxylation by GC. These experiments showed that the fingerprint-based substrate activity predictions were confirmed in all cases, indicating that both probes, combined, can relay reliable predictions about the ability of the corresponding P450 variant to accept MCL as a substrate for oxidation. More importantly, a very good correlation was also observed between the fingerprint-based site-selectivity predictions as derived from P1/Ptot activity and the enzyme regioselectivity in MCL hydroxylation (Table S1). In particular, the C2-selective P450 variants selected based on their high P1/Ptot probe activity ratio displayed the highest fraction of correct predictions (87%), whereas a rate of 69% and 78% correct predictions was observed for the C14-selective variants and the unselective variants, respectively (Figure 2c). Using this method, P450 variants featuring >90–95% regioselectivity toward hydroxylation of either C2 or C14 could be rapidly identified (Figure 3), thus demonstrating the functionality of regio-fingerprinting in guiding and expediting the search of regioselective P450 catalysts for MCL oxidation. The ability of the regio-fingerprints to capture differences in the enzyme active site configuration that favor C2 vs C14 hydroxylation (or vice versa) is remarkable also in view of the close spatial proximity of these C(sp3)—H sites within this molecule (Scheme 1). Another interesting observation emerging from these studies was the generally higher catalytic activity of the P1-selective P450 catalysts compared to the P2-selective variants (Figure 3). This result can be explained in view of the fact that the parent FL#62 inherently favors MCL oxidation at the C2 position—mimicked by P1 activity—and that dramatic changes in the regioselectivity of a P450 are often accompanied by a trade-off in catalytic activity.38,47−49
Figure 3.

Regioselectivity (blue/pink bars) and activity (black line) of representative P450 variants identified via regio-fingerprinting toward MCL oxidation. Conditions: cell lysate containing ∼0.1 mol % P450, 2 μM PTDH, 150 μM NADP+, 50 mM sodium phosphite in 50 mM potassium phosphate buffer (pH 8.0), r.t., 14 h. Activity was measured as percent total product conversion as determined via gas chromatography.
Identification of C2- and C14-Selective P450 Catalysts for MCL Hydroxylation
From the analyses above, a panel of candidate C2- and C14-selective enzymes were selected for further characterization as purified proteins (Table S2). Gratifyingly, these experiments revealed multiple variants featuring improved C2-selectivity compared to FL#62 (95–97% vs 87% C2-selectivity), such as V-F10 and V-E10 (Tables 1 and S2), both of which were predicted to be C2-selective based on their regiofingerprints (Figure S1). As an exception, II-G9 also exhibited high C2-selectivity (96%) despite its regiofingerprint indicating a preference for oxidation at the C14 site (Figure S1). Among these variants, V-F10 emerged as the best catalyst for C2-hydroxylation of MCL, enabling the synthesis of compound 2 with >97% regioselectivity, absolute R-stereoselectivity, and in near quantitative yield (97%) at a catalyst loading of merely 0.05 mol % (Table 1). This activity corresponds to 1930 catalytic turnovers (TON), i.e., nearly 7-fold higher TON than the parent enzyme FL#62. Notably, these variants were found to be all triple site mutants (vs FL#62) sharing a common Val → Ala mutation at position 82 and an aromatic residue (Phe or Tyr) at position 78. These positions cluster in the same region within the heme pocket of the enzyme (Figure 1b).
Table 1. Amino Acid Mutations and Catalytic Properties of Selected MCL-Oxidizing P450 Variantsa.
| amino
acid substitutionb |
(%)
product distribution |
||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| variant | 47 | 74 | 78 | 81 | 82 | 87 | 180 | 184 | 2 | 3 | 4 | [MCL] (μM) | mol %P450 | % yield (GC) | TONc |
| FL#62 | R | A | A | S | V | A | T | V | 87 | 4 | 9 | 500 | 0.1 | 28 | 280 ± 10 |
| II-G9 | F | A | A | 96 | 4 | 0 | 500 | 0.4 | >99 | >250 | |||||
| V-E10 | Y | Y | A | 95 | 5 | 0 | 500 | 0.4 | >99 | >250 | |||||
| V-F10 | F | V | A | 97 | 3 | 0 | 500 | 0.4 | >99 | >250 | |||||
| V-F10 | F | V | A | 97 | 3 | 0 | 1000 | 0.05 | 97 | 1,930 ± 20 | |||||
| V-D12 | L | Y | A | T | 51 | 49 | 0 | 500 | 0.2 | 9 | 42 ± 5 | ||||
| V-H10 | N | F | A | 36 | 64 | 0 | 500 | 0.2 | 51 | 256 ± 15 | |||||
| VIII-C4 | T | N | F | A | S | 30 | 70 | 0 | 500 | 0.2 | 15 | 74 ± 5 | |||
| VIII-B1 | T | N | F | F | A | S | 17 | 80 | 3 | 500 | 0.2 | 3 | 12 ± 1 | ||
| V-H10(A87L) | N | F | A | L | 13 | 87 | 0 | 500 | 0.1 | 9 | 90 ± 10 | ||||
| V-H10(A87I) | N | F | A | I | 3 | 97 | 0 | 500 | 0.1 | 10 | 95 ± 9 | ||||
| V-H10(R47C,A87I) | C | N | F | A | I | 3 | 97 | 0 | 500 | 0.1 | 26 | 258 ± 18 | |||
| V-H10(R47C,A87I)d | C | N | F | A | I | 1 | 99 | 0 | 500 | 0.1 | 72 | 720 ± 15 | |||
Reaction conditions: substrate and P450 at the indicated concentrations, 2 μM PTDH, 150 μM NADP+, 50 mM sodium phosphite in 50 mM KPi buffer (pH 8.0), r.t., 12 h.
Compared to P450BM3, FL#62 carries the following mutations: V78A, F81S, A82 V, F87A, P142S, T175I, A180T, A184V, A197V, F205C, S226R, H236Q, E252G, R255S, A290V, L353V.
Mean values and standard deviations are calculated from triplicate experiments.
Using 1 M potassium phosphate buffer (pH 8.0).
Importantly, a complementary set of P450 variants with high regioselectivity for C14-hydroxylation (64–80% C14-select. vs 4% for FL#62) was also successfully identified, which included VIII-B1, VIII-C4, and V-H10 (Table S2). The most C14-selective variant was VIII-B1, which produces 14-hydroxy-MCL (3) with 80% selectivity but in low yield (3%) and catalytic activity (12 TON; Table 1). A more promising catalyst for C14 hydroxylation of MCL was V-H10, which exhibits 64% regioselectivity toward formation of 3, no overoxidation to 4, and higher catalytic activity compared to VIII-B1 (256 TON; Table 1).
Using this variant, MCL was oxidized in 51% yield at a catalyst loading of 0.2 mol %. Sequencing analysis of the C14-selective variants revealed a series of “consensus” active site mutations corresponding to Ala78 → Asn and Ser81 → Phe (Tables 1 and S2), which appear to be critical for steering the enzyme site-selectivity toward favoring C14 hydroxylation. This mutational pattern is in stark contrast with that of the C2-selective variants, which feature a bulkier residue (Phe/Tyr) at position 78 and a small residue (i.e., Ala, Val, or Pro) at position 81. A subset of the C14-selective variants also shared mutations A74T, T180A, and V184S, but these substitutions did not appear to be critical for favoring C14 selectivity and they tended to be associated with low catalytic activities (<50 TON), as determined from comparison of structure–activity data for sequence-related variants (Table S2). In general, however, the C14-selective variants identified at this stage were ∼10-fold less efficient in MCL oxidation than the C2-selective variants (12–256 vs 250–1930 TON, respectively, Table 1), a result consistent with the trend observed from screening the P450 variants in cell lysate (Figure 3). Interestingly, some of the P2-selective variants, such as V-D12 (Table 1), exhibited a lower C14-regioselectivity as purified enzyme (49% C14-select.) than predicted based on their P2/Ptot activity ratio (56%; Table S1) and MCL oxidation reactivity in cell lysate (81% C14-select; Table S3). Control experiments involving reactions with the purified enzyme in a buffered solution spiked with E. coli cell lysate (50% v/v) reproduced the C14-selectivity observed during screening in cell lysate (Table S3). This phenomenon is not unprecedented,37 and it may arise from macromolecular crowding and/or allosteric effects in the presence of cell lysate. More importantly, these results confirmed that the fingerprinting method reports on the regioselectivity of these enzymes under the applied screening conditions (i.e., in cell lysate).
Further Refinement of C14 Regioselectivity
In order to develop an improved catalyst for C14 hydroxylation, a rational design approach was undertaken guided by the structure–activity data accrued from the C14-selective P450 variants identified via regio-fingerprinting (Table S2). Indeed, in addition to the crucial role of residues Asn78 and Phe81 as noted above, these data indicated that Ala87 → Ile/Leu mutations could potentially be beneficial for enhancing C14-selectivity, as suggested by the effect of a A87I mutation toward increasing C14-regioselectivity in XIII-A4 vs VIII-C4 (70 → 76% C14-select.) and that of a A87L mutation in disfavoring C2 hydroxylation over C14 hydroxylation in V-B2 vs V-E7 (10 → 39% C14-select.) (Table S2). The importance of position 87 in influencing C14-selectivity was also apparent from the improved C14-selectivity of VIII-C4 vs VIII-B1 (70% → 83% C14-select.), which differ only by an A87F mutation, although this mutation also caused a 6-fold reduction in activity (13 vs 74 TON) (Table 1). Based on these analyses, two derivatives of V-H10 that incorporate the mutation A87L or A87I were prepared. Gratifyingly, both V-H10-derived variants showed a noticeable improvement in C14 regioselectivity compared to V-H10 (64% → 87–97% C14-select.; Table 1). Specifically, V-H10(A87I) exhibited a nearly complete regioselectivity for C14 hydroxylation (97%) although this was accompanied by a 2.5-fold reduction in catalytic activity compared to the parent enzyme (95 vs 256 TON) and a corresponding reduction in product yield (10% vs 51%; Table 1).
In P450BM3, the Arg47 residue located at the opening of the substrate channel (Figure 1a) is known to facilitate binding of the fatty acids substrates.50 Mutation of this site has been found to facilitate recognition of non-native substrates by this enzyme.51−57 Accordingly, the R47C mutation was introduced into V-H10(A87I) to give V-H10(R47C,A87I). Gratifyingly, this variant showed improved catalytic activity (95 → 258 TON), while maintaining high C14-regioselectivity (97% C14-selectiv.; Table 1). Upon optimization of the reaction conditions (Table S4), and in particular in the presence of high concentration (1 M) phosphate buffer, the V-H10(R47C,A87I)-catalyzed reaction with MCL could be further optimized to yield 14-hydroxy-MCL (3) with excellent chemo- and regioselectivity (99% selectivity for 3), good yield (72%), and significantly improved catalytic activity (720 TON, Table 1). Overall, the P450 regiofingerprint strategy implemented in this work enabled us to rapidly isolate P450 variants with refined site-selectivity for each of the sites in MCL targeted by the parent enzyme FL#62. Furthermore, this method allowed for the acquisition of structure–reactivity data across multiple C2- and C14-selective variants (Table S2), which expedited the engineering of P450 catalysts with further improved regioselectivity and activity for selective C14 hydroxylation in MCL.
Characterization of the Regioselective P450 Catalysts
The best variants for C2 and C14 oxidation in MCL were further characterized to gain insights into the effects of the amino acid mutations on their catalytic properties. All of these enzymes exhibited a small but detectable heme spin shift upon incubation with micheliolide (Figure S4a). Accordingly, difference spectra (Abs390–420) measured at varying MCL concentrations were used to estimate the enzyme binding affinity for the non-native substrate (Table 2, Figure S4b). Using this assay, the parent enzyme FL#62 was found to bind MCL with a KD of 61 μM. In comparison, the triple mutant V-F10 exhibits a comparable affinity for MCL (KD = 88 μM), whereas the C14-selective P450 variants show 6- to 20-fold higher KD values, indicating that the amino acid substitutions favoring C14 selectivity also result in a weakened enzyme–substrate interaction. Kinetic experiments further showed that FL#62 oxidizes MCL with an initial product formation rate (PFR) of 49 turnovers min–1 and a coupling efficiency of 36% (Table 2). Whereas the C2-selective V-F10 show similar kinetic properties to FL#62 (PFR: 28 turnovers min–1), optimization of C14 hydroxylation selectivity is accompanied by a more pronounced reduction in both product formation rate (2–13 turnovers min–1) and coupling efficiency (3–10%; Table 2) compared to the parent enzyme. These features correlate with the significantly lower substrate binding affinity (590–1250 vs 88 μM; Table 2) and catalytic activity (12–256 vs 1930 TON; Table 1) of these C14-selective variants compared to the C2-selective V-F10. This selectivity/activity trade-off is not unexpected considering the dramatic shift in regioselectivity exhibited by these enzymes compared to the parent enzyme and intrinsically lower reactivity of the methyl group in C14 compared to the secondary C–H bond in C2. In this context, noteworthy is the beneficial effect of the R47C mutation in V-H10(R47C,A87I), whose improved performance toward MCL hydroxylation (95 → 258 TON; Table 1) is associated with an ∼4-fold increase in binding affinity for MCL (KD: 1254 → 365 μM), product formation rate (3 → 13 turnovers min–1), and coupling efficiency (3 → 10%) when compared to V-H10(A87I) (Table 2).
Table 2. Substrate Binding Affinity and Kinetic Properties of Selected MCL-Oxidizing P450 Variantsa.
| variant | selectivity | KD (μM)b | product formation ratec | coupling efficiencyd |
|---|---|---|---|---|
| FL#62 | C2-selective | 61 ± 3 | 49 ± 1 | 36% |
| V-F10 | C2-selective | 88 ± 18 | 28 ± 5 | 18% |
| VIII-B1 | C14-selective | nd | 2 ± 0.1 | 3% |
| V-H10 | C14-selective | 590 ± 90 | 7 ± 1 | 7% |
| V-H10(A87I) | C14-selective | 1250 ± 54 | 3 ± 0.5 | 3% |
| V-H10 (R47C,A87I) | C14-selective | 365 ± 53 | 13 ± 3 | 10% |
| (128 ± 8)e | (22 ± 4)e | (15%)e |
Mean values and standard deviations are calculated from triplicate experiments conducted at room temperature.
Derived from heme spin shift experiments; nd = not determined.
Rates are measured over initial 60 s in 50 mM potassium phosphate (KPi) (pH 8.0) and expressed as mole product per mole P450 per minute.
Ratio between product formation rate and NADPH consumption rate in the presence of MCL.
Reactions in 1 M KPi buffer (pH 8.0).
These studies also offered insight into the important enhancement in catalytic activity observed for V-H10(R47C,A87I) in the presence of high-concentration (1 M) phosphate buffer. Indeed, these reaction conditions were found to further enhance the product formation rate, coupling efficiency, and substrate binding affinity of the enzyme (Table 2) compared to standard buffer conditions (50 mM phosphate), which in turn led to a significant improvement in catalytic activity (258 → 720 TON) and product yield (26 → 72%; Table 1). Control experiments ruled out that this effect is correlated with the buffering capacity of the solution (no observed changes in pH throughout the reaction) and/or to the production of hydrogen peroxide through uncoupling pathways, since the addition of catalase had no effect on these reactions (Table S5). It is worth noting that the high buffer concentration has a similar beneficial effect on V-H10(A87I) but negligible effect on variant V-F10, thus showing a certain P450 variant dependence (Table S5). To our knowledge, the beneficial effect of high phosphate buffer concentration on the catalytic activity of these P450BM3 variants was not documented before and this could prove to be a rather straightforward strategy to improve the performance of other P450s.
Large Scale Enzymatic Oxidation of Micheliolide
Based on these results above, V-F10 and V-H10(R47C,A87I) were selected as the best P450 catalysts for the functionalization of micheliolide at the C2 and C14 site, respectively. Accordingly, preparative scale (0.2 g) reactions were carried out using P450 variants V-F10 (0.1 mol %) and V-H10(R47C,A87I) (0.1 mol %) in the presence of a NADPH regeneration system to afford >150 mg of 2 and 3, respectively, in high isolated yield (70–90%; Scheme 3). These results demonstrated the utility of these engineered P450 enzymes to access two key intermediates for further late-stage elaboration of this complex natural product at a synthetically useful scale.
Scheme 3. Preparative-Scale Enzymatic Synthesis of 2(R)-Hydroxy-MCL (2) and 14-Hydroxy-MCL (3) with the Optimized P450 Catalysts.
Synthesis and Antileukemic Activity of Micheliolide Derivatives
The isolated 2(R)-hydroxy-MCL (2) and 14-hydroxy-MCL (3) were further processed via deoxofluorination and acylation chemistry to produce a diverse panel of semisynthetic MCL analogs functionalized at position C2, C14, and/or C4 in order to gain comprehensive structure–activity insights into the impact of these modifications on the antileukemic properties of micheliolide (Table 3). Among them, analogs with identical substituents installed at the three different sites across the molecule (C2, C14, C4) were prepared to examine the positional effect of these modifications and the relative potential of these sites toward potentiating the antileukemic activity of this natural product.
Table 3. Chemical Structures of MCL Analogs Prepared via Chemoenzymatic Functionalization of C2, C14, and C4 Sites.

Reagents and conditions: MeI, Ag2O, anhydrous DMF, r.t. to 50 °C, 24 h.
Reagents and conditions: DAST, anhydrous DCM, −78 °C → 0 °C.
Reagents and conditions: DMAP, Et3N, substituted benzoyl chloride, anhydrous DCM, r.t. to 40 °C, from 2 to 72 h.
The antileukemic activity of the MCL analogs was first tested using the leukemia cell line, M9-ENL1. These cells are derived from lineage depleted (Lin-) human cord blood cells transduced with a leukemogenic mixed-lineage leukemia (MLL)-11 19 leukemia (ENL) fusion gene and they develop a rapid and fatal pro-B cell acute lymphoblastic leukemia (ALL) disease in NOD/SCID mice.17,58 M9-ENL1 cells show an enrichment of embryonic and B-cell progenitor gene sets and exhibit the hallmarks and gene expression patters typical of leukemia stem cells.17 Cells were treated with a standard 20 μM dose of MCL analog for 24 h followed by measurement of cell viability using Annexin-V and 7-amino-actinomycin (7-AAD) stains (Figure 4). At this dose, MCL shows a 50% reduction in cell viability, which is in line with its LC50 value of 14.6 μM measured from dose-dependent curves using the same assay. Notably, the hydroxylated MCL derivatives show minimal activity (80–100% cell viability; Figure 4) at this dose indicating that the presence of a polar hydroxyl group at either site is deleterious to the antileukemic activity of the parent compound. The methoxy derivatives MCL-1 (P1) and MCL-2 (P2) and the C2- and C14-fluorinated derivatives MCL-3 and MCL-4 show comparable and slightly improved activity, respectively, relative to the parent natural product. In stark contrast, more than half of the MCL analogs derived from acylation of 2 or 3 exhibited significantly improved antileukemic activity compared to MCL (2–30% vs 50% cell viability at the fixed dose) (Figure 4).
Figure 4.
Viability of M9-ENL1 cells upon treatment (24 h) with MCL and the MCL analogs at 20 μM (* = 10 μM). The bars are color coded according to the site(s) of functionalization. UNT = untreated cells. VEH = cell treated with DMSO vehicle only.
The introduction of a benzoyl group at either the C2 (MCL-5) or C14 site (MCL-6) resulted in a loss (C14) or no change (C2) in antileukemic activity, while the same substitution at C4 (MCL-7) brought about a moderate (2-fold) improvement in antileukemic activity compared to MCL. Interestingly, the introduction of a fluorine substituent at the para position of the benzoyl group greatly improved the antileukemic activity of all three analogs (4–8% cell viability for MCL-8, -9, and -10). The beneficial effect of increasing the lipophilicity of the aryl group at either the C2 or C14 position was further supported by the substituent effect: 4-CF3-Bz ≈ 4-F-Bz ≫ 4-N(Me)2-Bz ≥ Bz observed across the series of C2-substituted analogs (MCL-13, -11, -8, and -5) and the C14-substituted counterparts MCL-14, -12, -9, and -6. Based on these results, the lipophilicity of the aryl group was further increased by installing two trifluoromethyl groups at the ortho, para positions (MCL-15, -16, -17) and meta, meta positions (MCL-18, -19, -20). The latter modification at the C2 or C14 site resulted in two very potent compounds, namely MCL-18 and MCL-19, capable of inducing nearly complete M9-ENL1 cell death (<2–3% cell viability) at the single dose of 20 μM. Interestingly, the same substitution at the C4 site (MCL-20) resulted in significant loss of activity, further revealing a strong positional effect of the aryl substituent toward influencing the activity of the molecule. Interestingly, a different structure–activity trend (C14 > C2 ≈ C4 vs C14 ≈ C2 ≫ C4) along with generally lower antileukemic activity (7–30% cell viability) was exhibited by the regioisomeric analogs incorporating the ortho,para-bis-trifluoromethyl-phenyl substituents (MCL-15, -16, -17). Altogether, these results point at a specific and position-dependent interaction of these substituents with the biomolecular targets of these compounds in leukemia cells. Along with the monosubstituted analogs, difunctionalized MCL derivatives obtained through modification of C2/C4 (MCL-21) and C14/C4 (MCL-22) were also tested. Both compounds, however, showed parentlike antileukemic activity and thus significantly reduced activity compared to the C2- and C14-monosubstituted counterparts.
Focusing on the modification of the C2 and C14 site, which showed most promise for potentiating the antileukemic activity of MCL based on the results above, additional analogs were prepared by introducing alternative aromatic, heteroaromatic and biaryl groups (i.e., naphthalene, indole, furan, pyrazole) at either of these positions (Table 3). Whereas all these MCL analogs show improved antileukemic activity compared to MCL (Figure 4), the naphthyl-substituted MCL-23 and MCL-24 as well as the phenyl-pyrazole containing analogs MCL-31 and MCL-32 showed very high cytotoxicity against M9-ENL1 cells, resulting in 3–7% cell viability after treatment with these compounds. Across the indolyl derivatives, the C14-substituted analog (MCL-28) emerged as the most promising analog, showing noticeably higher activity than the C2-substituted analog (4% vs 42% cell viability). In all cases, additional substitution at the C4 position caused a significant reduction (MCL-29 and MCL-30 vs MCL-28) or complete loss in activity (MCL-25 and MCL-26 vs MCL-24), further supporting the notion that introduction of bulky substituents at both sites is not well tolerated.
Antileukemic Activity against Patient-Derived AML Specimens
From these activity tests, several MCL analogs were discovered that exhibit significantly improved antileukemic activity compared to the parent molecule, MCL (Figure 4). MCL analogs causing ≥93% lethality of M9-ENL1 cells after 24-h incubation at 20 μM were further characterized by determining their half-maximal lethal concentration (LC50) value from dose–response curves (Table 4). In these experiments, MCL was determined to exhibit a LC50 of 15.4 μM (Table 4). MCL-9, which bears a para-fluorobenzoyl moiety at the C14 position, showed a 3-fold enhanced activity (LC50 = 4.8 μM) compared to MCL and the C2-substituted counterpart MCL-8 (LC50 = 14.7 μM). Notably, MCL-13, which differs from MCL-8 by substitution of the fluorine group with a trifluoromethyl group, exhibited high antileukemic activity with a LC50 of 1.8 μM; this corresponding to an 8-fold improvement compared to the parent molecule (Table 4). The o,p- and m,m-ditrifluoromethyl-substituted aryl derivatives MCL-16 and MCL-18, MCL-19 as well as the pyrazole-containing analogs MCL-31 and MCL-32 were also found to possess improved antileukemic activity (2- to 3-fold) compared to MCL, with LC50 values ranging from 4.1 μM to 9.5 μM. In contrast, the napthoyl- and indolyl-substituted analogs MCL-24 and MCL-28, respectively, show no significantly improved LC50 values compared to MCL (LC50 ≈ 13–14 μM; Table 4).
Table 4. LC50 Values of Micheliolide (MCL) and Its Derivatives against M9-ENL1 Cells and Four Primary AML Specimens Derived from Patients with Relapsed AMLa.
| compound | M9-ENL1 LC50 (μM) | AML01 LC50 (μM) | AML02 LC50 (μM) | AML03 LC50 (μM) | AML04 LC50 (μM) |
|---|---|---|---|---|---|
| MCL | 15.4 | 18.8 | 20.3 | 15.7 | 22.1 |
| MCL-8 | 14.7 | nd | nd | nd | nd |
| MCL-9 | 4.8 | 9.0 | 16.9 | 13.1 | 9.5 |
| MCL-13 | 1.8 | 8.2 | 6.6 | 4.3 | 4.3 |
| MCL-16/SG-8 | 5.2 | 8.3 | 12.5 | 9.1 | 7.4 |
| MCL-18 | 7.0 | 25.6 | 5.9 | 5.8 | 4.7 |
| MCL-19 | 4.1 | 3.2 | 2.5 | 2.5 | 2.3 |
| MCL-24 | 13.5 | nd | nd | nd | nd |
| MCL-28 | 13.6 | nd | nd | nd | nd |
| MCL-31 | 9.5 | 19.9 | 15.8 | 10.2 | 19.0 |
| MCL-32 | 8.6 | 18.5 | 16.1 | 9.5 | 19.5 |
nd = not determined. Standard errors are within 20%.
The most promising MCL analogs identified through these studies were then tested for their activity against acute myelogenous leukemia (AML) specimens derived from patients with relapsed AML. Whereas compounds MCL-9, MCL-13, MCL-16, and MCL-18 all showed 2- to 3-fold enhanced activity against primary AML specimens compared to the parent natural product, compound MCL-19 emerged as the most potent antileukemic agent within this group, exhibiting a LC50 of ∼2–3 μM across all the four primary AML specimens tested (Table 4). This activity corresponds to approximately an order of magnitude higher potency compared to MCL (LC50 ∼ 19–22 μM).
Altogether, these results demonstrated that C2 and C14 constitute two “hot spots” for potentiating the antileukemic activity of MCL, with selective chemoenzymatic functionalization of these sites enabling the development of MCL analogs with significantly enhanced antileukemic properties compared to the parent natural product. Furthermore, as shown by the data in Table 3, systematic structure–activity relationship (SAR) data across three distinct sites within this complex molecule could be obtained by combining this approach with functionalization of the pre-existing hydroxyl group at position C4, thus enabling direct comparison of positional effect of a given substituent on the activity of the molecule against leukemia stem cells and patient-derived AML specimens. Of note, the functionalization pattern at position C14 which resulted in the most potent compound identified in this study, MCL-19, causes a decrease in antileukemic activity when installed at position C4, further highlighting the value of the present P450-mediated late-stage functionalization strategy with engineered P450 with tailored site-selectivity.
Multiprobe Profiling of Micheliolide-Targeted Proteome in Leukemia Cells
Pull-down experiments in combination with proteomic analyses constitute a powerful approach for the identification of the cellular target(s) of bioactive molecules, including natural products.59−65 Conventionally, this approach relies on the use of a single affinity probe generated through functionalization of the bioactive molecule of interest at a permissible site.66 Especially for bioactive molecules with a complex mechanism of action and/or multitarget specificity, however, this single-probe approach bears the risk of providing only partial or biased information due to a potentially differential effect of the affinity tag in the recognition of the molecule by its biomolecular targets. Leveraging the regiodivergent P450 catalysts described above, we thus sought to obtain a set of multiple MCL-based affinity probes for a multiplexed analysis of the biomolecular targets of MCL in leukemia cells. Accordingly, two MCL-based affinity probes were prepared by conjugating a biotin moiety, via a flexible linker, to the C2 or the C14 position of MCL by chemoenzymatic means (2-biot-MCL and 14-biot-MCL, respectively; Figure 5a; see Scheme S1 and the SI for details). A third biotin-conjugated probe was prepared through functionalization of the pre-existing OH group at C4 (4-biot-MCL; Scheme S1). Upon testing them in M9-ENL1 cells, the three biotinylated MCL-based probes showed a LC50 value comparable to that of MCL (i.e., LC50 = 13–17 μM; Figure 5a), indicating that, in each case, the affinity tag did not have any significant impact on the overall antileukemic properties of the molecule. Therefore, all three probes were deemed viable affinity reagents for examining proteome targeted by MCL in leukemia cells via pull-down experiments.
Figure 5.
Multiprobe analysis of MCL-targeted proteome in leukemia cells. (a) Chemical structures of the biotinylated MCL-based affinity probes and their antileukemic activity (LD50) in M9-ENL1 cells (n = 6). (b) Overview of the competitive pull-down experiments for analysis of MCL-targeted proteins in acute myeloid leukemia cells (M9-ENL1 cells) (n = 3 per probe). (c) Volcano plots of protein hits identified using the biotinylated MCL-based probes. The top three highly enriched/scoring proteins (= enrichment ratio >3-fold; P < 0.05) are labeled and highlighted in red. See Figures S5–S7 for additional data. (d) Venn diagram showing the overlap (or not) of the most representative putative target proteins identified using the different biotinylated MCL-based probes. The total number of significant hits (>1.25-fold enrichment; P < 0.05 in ≥1 probes) for each probe is displayed in brackets. See Tables S7 and S8 for additional data. (e) Dose-dependent induction of heme oxygenase 1 (HO-1) in primary AML specimens by MCL, PTL, and MCL analogs MCL-13 and MCL-19.
As outlined in Figure 5b, these experiments were carried out by treating M9-ENL1 cell lysates with each of the three MCL-based probes both with and without pretreatment with MCL. This competitive setup allows identification of hits based on the fold enrichment over the MCL-treated control, minimizing background noise due to unspecific protein binding to the affinity matrix. The probe-bound proteins were captured with neutravidin-coated beads and analyzed via LC-MS/MS and bioinformatic tools. Upon comparison of the sample groups (pretreated with vehicle only) versus controls (pretreated with MCL), a total of 47 putative target proteins were identified using the entire panel of MCL-based probes (Figures 5c; Table S7). While various hits were shared among the probes (vide infra), the total number of significant hits identified with each probe varied, corresponding to 14 for C2-biot-MCL, 24 for C14-biot-MCL, and 33 for C4-biot-MCL (Figure 5d; Table S7). As summarized in Tables 5 and S8, putative target proteins showing the highest enrichment (>3-fold) with one or more probes included proteins/enzymes involved in redox homeostasis (TXNDC12), metabolism (PAPSS1, DHFR, DCXR), and signal transduction (RAB2A, GIT2, ITPR2). Insightfully, these experiments revealed the MCL features a complex protein-targeting profile in leukemia cells, suggesting the engagement of multiple protein targets in these cells. These results are in contrast with the previously proposed single-target specificity and high selectivity of MCL toward targeting pyruvate kinase M2 (PKM2), as determined using HL60 cells and a C4-biotinylated probe similar to 4-biot-MCL.67 At the same time, our results are consistent with the behavior of parthenolide, which shares with MCL an α-methylene-γ-lactone moiety as an electrophilic warhead critical for anticancer activity and which was determined to target multiple proteins and pathways in leukemia cells.68 While the specific proteins targeted by MCL and PTL differ from each other, functionally related proteins include proteins implicated in redox homeostasis (i.e., GPX1 and TXN for PTL).68 Similar to PTL,28 MCL was reported to suppress the NF-κB pathway in cancer and noncancer cells, a mechanism associated with its anti-inflammatory activity.21,69,70 Interestingly, while PTL is known to covalently inhibit NF-κB,27 NF-κB was not among the hits for any of the MCL-based probes. Thus, these results suggest that, in stark contrast to PTL, the inhibitory effect of MCL on the NF-κB pathway does not stem from a direct interaction of the natural product with NF-κB.
Table 5. Most Significant Putative Targets of MCL in Leukemia Cells (M9-ENL1) as Determined Using the Three Biotinylated Probesa.
| gene name | protein | function | enrichment ratio | probe |
|---|---|---|---|---|
| EXOC1 | exocyst complex component 1 | vesicle transport | 5.1 | C2 |
| GIT2 | ARF GTPase-activating protein GIT2 | signal transduction | 5.8 | C14 |
| ITPR2 | inositol 1,4,5-trisphosphate receptor type 2 | signal transduction | 4.0 | C14 |
| DHFR | dihydrofolate reductase | metabolism | 3.0 | C14 |
| RAB2A | Ras-related protein Rab-2A | signal transduction | 7.0 | C4 |
| DCXR | l-xylulose reductase | metabolism | 3.1–1.8 | C2/C4 |
| PAPSS1 | 3′-phosphoadenosine 5′-phosphosulfate synthase 1 | metabolism | 7.6–1.4 | C14/C4 |
| TXNDC12 | thioredoxin domain containing protein 12 | redox homeostasis | 7.0–1.5 | C14/C4 |
| LPXN | leupaxin | cell signaling in hematopoietic cells | 2.5–1.4 | C14/C4 |
| RAP1B | Ras-related protein Rap-1b | signal transduction | 1.7–1.2 | C14/C4 |
| CNBP | cellular nucleic acid-binding protein | gene transcription regulation | 3.2–1.5 | C2/C14/C4 |
| APRT | adenine phosphoribosyl-transferase | metabolism | 1.6–1.2 | C2/C14/C4 |
| PRDX1 | peroxiredoxin-1 | redox homeostasis | 1.3–1.1 | C2/C14/C4 |
The protein function, gene name, fold-enrichment (vs MCL-treated cells), and probe(s) used for detection are indicated. See Table S8 for additional information.
Other interesting insights could be obtained from analysis of the overlapping and nonoverlapping hits identified using the three MCL-based probes, as summarized by the Venn diagram in Figure 5d. Indeed, while a large fraction of the hits (22/47 = 47%) were shared among the affinity probes, many others (25/47 = 53%) were uniquely enriched by the different probes (3 with 2-biot-MCL; 13 with 4-biot-MCL; 9 with 14-biot-MCL). For example, the exocytic vesicle protein EXOC1 was a major target solely identified using 2-biot-MCL, whereas the signal transduction proteins ARF GTPase-activating protein GIT2 and Ras-related protein Rab-2A were major targets identified using 14-biot-MCL and 2-biot-MCL, respectively (Figure 5d, Tables S7–S8). These results can be rationalized considering the complex, multitarget specificity of MCL as emerging from our chemical proteomics experiments and the fact that installation of the affinity tag at different positions within the MCL molecule is expected to influence its interaction with the different target proteins in different ways. Most importantly, these results showed that a much more comprehensive picture of the MCL-targeted proteome in leukemia cells could be achieved using the multiprobe set than possible using any single one of these probes. In turn, these results highlight another key benefit of the present approach based on chemoenzymatic C–H functionalization at multiple sites of the bioactive molecule for target identification purposes.
While further studies are warranted to correlate the role of the various MCL-targeted proteins (Tables 5 and S8) with the antileukemic activity of the natural product, another advantage of the present multiprobe chemical proteomics approach is to guide initial focusing of these efforts toward hits lying within the overlapping regions of the Venn diagram of Figure 5d. Among these, peroxiredoxin-1 (PRDX1) and thioredoxin domain containing protein 12 (TXNDC12) (Table 5) are of particular interest, as they play a role in cell protection against oxidative stress.71,72 By targeting PRDX1 and TXNDC12, MCL is thus expected to cause an unbalance of cell redox homeostasis in leukemia cells, resulting in an oxidative stress induced cytotoxicity. To probe this hypothesis, we investigated the induction of Heme oxygenase (HO-1) in primary AML specimens upon treatment with MCL and the more active analogs MCL-13 and MCL-19 (Figure 5e). HO-1 is an inducible form of heme oxygenase that responds to oxidative stress in cells.73,74 These experiments showed a dose-dependent induction of HO-1 by MCL compared to the vehicle-only control (Figure 5e). This feature is shared by PTL and the more potent MCL analogs, which were found to cause a comparatively larger induction of HO-1 at lower doses, thus indicating a potentiation of this activity (Figure 5e). Altogether, and in agreement with the chemical proteomics experiments, these results point at oxidative stress induction as one of the cellular mechanisms underlying the antileukemic activity of MCL and its improved analogs. As noted above, further studies are warranted to investigate other potential mechanisms of action of MCL in leukemia cells, as suggested by its complex protein targeting profile as emerging from the present studies (Figure 5). For example, other interesting putative targets lying at the intersection of the probe-targeted profiles (Figure 5d) include the multifunctional nuclear acid-binding protein CNBP, which was reported to control the transcription of multiple oncogenes,75 and leupaxin (LXPN), whose downregulation (by siRNA) was found to suppress the proliferation and invasion of human acute monocytic leukemia cells in vivo.76
Conclusion
In summary, we have introduced a novel probe-based approach for guiding the development of site-selective P450 oxidation catalysts. This strategy complements and expands the toolbox of fingerprint-based approaches for P450 function prediction,9,38,46 providing an efficient solution for accelerating the identification of engineered P450 enzymes with tailored regioselectivity for the oxyfunctionalization of a complex molecule. Using this strategy, we were able to obtain two highly regio- and stereoselective P450 catalysts for the late-stage chemoenzymatic C–H functionalization of two previously inaccessible aliphatic positions (C2 and C14) in the antileukemic natural product micheliolide. Along with functionalization of the C4 position, these chemoenzymatic routes have enabled the synthesis and evaluation of a broad panel of C2-, C14-, and disubstituted MCL analogs, providing comprehensive structure–activity data on the effect of substitutions at multiple sites across the complex natural products on its antileukemic properties. Importantly, these studies revealed that both C2 and C14 constitute key sites for potentiating the antileukemic activity of MCL, resulting in the discovery of MCL analogs such as MCL-13 and MCL-19 with significantly enhanced antileukemic activity against both leukemia stem cells (M9-ENL1) and patient-derived primary AML cells. These findings and methods pave the way to the development of MCL-based compounds as antileukemic and anticancer agents. Finally, these chemoenzymatic routes have also enabled the generation of a set of affinity reagents for investigation of the MCL-targeted proteins in leukemia cells via pull-down experiments and proteomics analysis. To our knowledge, this is the first example of exploiting engineered P450 enzymes for the selective incorporation of an affinity tag into a bioactive molecule. In contrast to a previous report, our studies revealed that MCL has a complex protein targeting profile, involving multiple target enzymes implicated in regulation of cellular redox homeostasis, signal transduction and metabolism. Our results further support the notion that induction of oxidative stress in leukemia cells through targeting PRDX1 and TXNDC12 represents one of the key mechanisms of actions underlying the antileukemic activity of MCL. More broadly, these studies demonstrated how the present strategy permitted a comprehensive analysis of the proteome targeted by a bioactive natural product, as made possible through a complementary set of affinity probes generated through the site-selective functionalization of multiple C–H positions across the molecule’s scaffold. In particular, this approach enabled unveiling of multiple targets engaged by MCL in leukemia cells which would have been missed or overlooked using a conventional, single probe-based strategy. We anticipate that this overall approach can prove valuable toward enabling the optimization of the biological activity as well as the investigation of the mechanism of action of other bioactive molecules and natural products.
Experimental Details
Expression and Fingerprinting of the P450 Library
P450s were expressed from pCWori vectors containing the P450 gene under the control of a ptac promoter. 96-Deep-well plates containing 400 μL of LB medium (100 mg ampicillin L–1) per well were inoculated with single variants from the P450 library and incubated overnight at 37 °C (200 rpm, 80% humidity). Eight wells in each plate were set to contain P450BM3(F87A) as a reference variant for data normalization. After overnight growth, 50 μL from each well was used to inoculate a second set of 96-deep-well plates containing 900 μL of Terrific Broth (TB) medium (100 mg ampicillin L–1). The plates were incubated at 37 °C (200 rpm, 80% humidity) for 2–3 h until the OD600 reached 1.0. The cultures were then induced with β-d-1-thiogalactopyranoside (IPTG) and δ-aminolevulinic acid (ALA) at final concentrations of 0.5 and 0.35 mM, respectively. The cultures were incubated at 27 °C for 20 h (150 rpm, 80% humidity) and harvested by centrifugation at 4000 rpm. The pellets were separated from the supernatant and frozen at −80 °C. Cell lysates were prepared by adding 400 μL of lysis solution (4 U deoxyribonuclease I, 0.8 mg/mL lysozyme, 10 mM MgCl2, 50 mM phosphate buffer pH 8.0) to each well. After incubation at 37 °C for 60 min (150 rpm), clarified lysates were obtained by centrifugation at 4000 rpm for 20 min. P450 demethylation activity on probes P1 and P2 was measured in parallel reactions with the aid of a Beckman Coulter Multimek 96 automated pipettor and a TECAN Infinity plate reader. Reactions were carried out by mixing 50 μL of clarified cell lysate with 150 μL of 50 mM phosphate buffer (pH 8.0) containing 0.7 mM probe (solubilized in 2% DMSO) and a phosphite dehydrogenase (PTDH)-based cofactor regeneration system (1.8 μM PTDH, 50 mM sodium phosphite, 150 μM NADP+). After incubation for 12 h at room temperature (80 rpm), 50 μL of 2 M NaOH containing 150 mM Purpald reagent was added to each well, followed by measurement of the optical density at 550 nm after 30 min.
Regiofingerprint Analysis
The P450 regiofingerprints were analyzed as follows. The normalized activity of each P450 variant on probe P1 (AP1) and probe P2 (AP2) was calculated using eqs 1 and 2, respectively:
| 1 |
| 2 |
where x1 is the absorbance value measured from the reaction of the P450 with probe P1 in the purpald assay; x2 is the absolute absorbance value measured from the reaction of the P450 with probe P2 in the purpald assay; RP1 is the mean value (n = 8) for the reaction of reference P450BM3(F87A) with probe P1; RP2 is the mean value (n = 8) for the reaction of reference P450BM3(F87A) with probe P2. The probe activity of the reference P450 were averaged to eliminate bias in AP1 and AP2 calculations due to P450BM3(F87A)’s inherent preference for C2-hydroxylation (65% C2-selectivity). P450 variants with normalized P1 (AP1) or normalized P2 activity (AP2) lower than 0.2 were considered MCL-inactive. Upon calculation of AP1 and AP2, the fingerprint-based regioselectivity of the screened P450 variants was calculated using (AP1 × 100)/Atot for P1 selectivity and (AP2 × 100)/Atot for P2 selectivity, where Atot = AP1 + AP2.
Cell Lysate Reactions
Selected P450 variants identified via regio-fingerprinting were expressed in 48-well plates as described above. For each variant, the clarified lysate (400 μL) was transferred to a 48-deep well plate and incubated with 0.7 mM MCL (solubilized in 2% DMSO) and the PTDH-based cofactor regeneration system. After incubation for 12 h at room temperature (80 rpm), 500 μM guaiacol (in EtOH) was added as internal standard. The reaction mixture were extracted with DCM, followed by gas chromatography (GC) analysis using a Shimadzu GC2010 equipped with an FID detector and a Restek RTX-5 column (15 m × 0.25 mm × 0.25 μm film), and the following program: 200 °C inlet, 300 °C detector, 130 °C oven, 12 °C/min ramp to 150 °C, 36 °C/min ramp to 200 °C, 12 °C/min ramp to 250 °C, 24 °C/min ramp to 290 °C for a total of 8.89 min. P450 concentration in the cell lysate was determined via a CO-binding assay. Product conversion and catalytic turnovers (TON) were calculated based on GC peak areas corresponding to 1 (ret. time: 5.72 min), 2 (rt: 6.82 min), 3 (rt: 7.38 min), and 4 (rt: 7.06 min). Mean and standard deviation values reported were calculated from experiments performed at least in duplicate.
Mutagenesis
The V-H10-derived variants were prepared using pCWori_V-H10 (or pCWori_V-H10(87I)) as template, primers BamHI_2_fwd (5′-GGAAACAGGATCCATCGATGC-3′) and SacI_2_rev (5′- AATATCGAGCTCGTAGTTTGTATGATC-3′) as megaprimers, and the oligonucleotides in Table S6 as mutagenizing primers. The target gene product (1.4 Kbp) was prepared by PCR overlap extension mutagenesis and digested with BamH I-HF and Sac I-HF restriction enzymes at 37 °C for 2 h. The digested gene was ligated to a BamH I-HF/Sac I-HF double-digested pCWori_P450BM3(WT) vector using T4 DNA ligase. The ligation mixtures were transformed in chemically competent DH5α cells and transformants were selected on LB agar plates containing ampicillin (100 mg L–1). Mutations were confirmed by Sanger sequencing.
Protein Expression and Purification
The P450 enzymes were expressed from pCWori-based vectors and purified by ion-exchange chromatography as described previously.46 P450 concentration was determined from CO binding difference spectra (ε450–500 = 91 000 M–1 cm–1). A vector for the expression of a thermostable phosphite dehydrogenase (PTDH) was kindly provided by the Zhao group.77 PTDH was expressed and purified via Ni-affinity chromatography as described previously.46
Determination of TON and Regioselectivity for MCL Oxidation
Analytical-scale reactions (1 mL) were carried out by dissolving micheliolide (0.5–1 mM) in potassium phosphate buffer (50 mM, pH 8.0) containing 2% (v/v) DMSO. This solution was added with purified P450 enzyme (0.5–1 μM), followed by the NADPH cofactor regeneration system (2 μM PTDH, 150 μM NADP+, 50 mM sodium phosphite). The mixture was stirred (100 rpm) for 12 h at room temperature followed by the addition of 0.5 mM guaiacol as internal standard. The crude products were extracted with DCM and analyzed by gas chromatography as described above. Product yield and TON values were calculated using calibration curves generated with isolated products 2–4.
Product Formation Rates and Coupling Efficiency
Initial product formation rates were measured using 1 mM MCL, 1 μM P450, and 200 μM NADPH in potassium phosphate buffer (50 mM, pH 8.0). After 60 s, the reaction was quenched by addition and rapid mixing with 300 μL DCM. The reaction was analyzed by GC as described above. NADPH oxidation rate in the presence of micheliolide was measured by monitoring NADPH depletion at 340 nm (ε = 6.22 mM–1 cm–1) in reactions containing 1 mM MCL, 1 μΜ purified P450, and 200 μM NADPH. The coupling efficiency was determined as the ratio between the product formation rate and the NADPH consumption rate in the presence of the substrate. Reported mean and standard deviation values were calculated from experiments performed at least in triplicate.
KD Measurements
Binding experiments were performed using 3 μM purified P450 in potassium phosphate buffer (50 mM, pH 8.0) by titrating increasing amounts of micheliolide (5–2000 μM) from an ethanol stock solution (50 mM). At each concentration, a difference spectrum from 350 to 500 nm was recorded and binding curves were generated by plotting the change in absorbance at 390 and 420 nm corresponding to the high-spin and low-spin state of the enzyme, respectively, against MCL concentration. KD values were calculated using Graph Pad Prism via nonlinear fitting of the experimental binding curves to an equation describing a standard 1:1 binding interaction. Reported mean and standard deviation values were calculated from experiments performed at least in triplicate.
Enzymatic Hydroxylation of MCL
Detailed procedures for the enzymatic synthesis, isolation, and characterization of 2, 3, and 4 are provided as Supporting Information.
Synthesis of MCL and Probes
Detailed procedures for the synthesis of MCL (from parthenolide), fingerprint probes P1–P2, biotinylated probes, and the MCL analogs described in Table 3 are provided as Supporting Information.
Cell Cultures and Cell Viability Assays
For all cell culture studies, cells were kept in a 37 °C humidified incubator with 5% CO2. Human acute myeloid leukemia samples were obtained after informed consent from volunteer donors. Live mononuclear cells were isolated by subjection to Ficoll-Paque (GE Healthcare Bio-Sciences) density gradient and either cryopreserved (leukemia samples) or cultured without cryopreservation (umbilical cord samples). The viability of cryopreserved leukemia samples after thawing was 40–70%. The cell viability assays with M9-ENL-1 cells were carried out by plating M9-ENL1 cells at a density of 106 cells/mL in αMEM culture media (Invitrogen) supplemented with 5% human plasma, 20% FBS, and the cytokines SCF, IL-3, IL-7, FLT3 ligand (Peprotech), and penicillin/streptomycin. MCL and the MCL analogs were diluted into the culture media from a DMSO stock solution and added to the cells at a fixed dose (i.e., 20 μM) or at different concentrations (typically, between 0.5 and 100 μM) for LC50 determination. After 24 h of drug exposure, cells were collected for flow cytometry analysis. Cells were stained with Annexin V and 7-aminoactinomycin D (7-AAD) to quantify the percentage of nonapoptotic (negative for both stains) cells. Analyses were conducted in triplicate. LC50 values for the biotinylated probes 2-biot-MCL, 14-biot-MCL, and 4-biot-MCL were determined using M9-ENL1 suspension cells seeded in conical 96-well plates (Nunc) at a density of 30 000 cells per well in 90 μL of αMEM culture media (Invitrogen) supplemented with 5% human plasma, 20% FBS, and the cytokines SCF, IL-3, IL-7, FLT3 ligand (Peprotech), and penicillin/streptomycin. The plates were incubated in a humidified incubator for 4 h prior to treatment. The biotinylated probes were diluted from a 10 mM DMSO stock solution into complete cell culture medium and added to the cells to give a final concentration of 2–100 μM and 1% (v/v) DMSO (n = 6). Following a 24-h incubation, 20 μL of a 5 mg/mL Thiazolyl Blue tetrazolium bromide solution was added directly to the culture media. After incubation at 37 °C for 3 h, the plates were centrifuged (4000 rpm, 10 min), the media was removed, and 100 μL of DMSO was added to solubilize the formazan product. The resulting OD was measured at 550 nm using a multiwell plate reader (Tecan).
Pull-down Experiments
M9-ENL1 cells were harvested by centrifugation (500g, 10 min) and washed 5× with cold phosphate buffered saline to remove the culture medium. The cell pellet was lysed in cold RIPA buffer containing a protease inhibitor (pi) cocktail for 10 min at 4 °C followed by sonication. The lysate was centrifuged (10 000g, 10 min) to remove cell debris and the protein concentration was determined using the BCA reagent per manufacturer’s instruction (ThermoFisher). M9-ENL1 cell lysates (2 mg protein per sample) were incubated with 10 μM MCL (MCL-treated control) or just DMSO (sample) for 1 h on ice with gentle shaking. The samples were then treated with the appropriate biotinylated MCL-based probe at a final concentration of 10 μM (control and noncontrol groups). The lysate was incubated on ice for 2 h with gentle shaking and transferred to a 3 kDa centrifugal filter unit. The samples were washed with cold PBS (+ pi) three times to remove any unreacted probe and incubated with 50 μL of ultrahigh capacity neutravidin beads (100 μL of slurry, Pierce) for 60 min at 4 °C with gentle shaking. Cold phosphate buffer saline (PBS) (+ pi) was added to the mixture and the samples were centrifuged for 2 min at 10 000 rpm (2×). Beads were successively washed with 1 mL of PBS containing pi and 2% SDS and the supernatant was removed via centrifugation (2 min, 10 000 rpm, 4×). A 40 μL portion of 1× laemli sample buffer was added to the beads, and then beads were heated at 95 °C for 10 min to elute the neutravidin bound proteins. The pull-down experiments were performed in triplicate for each probe.
LC-MS/MS and Bioinformatic Analysis
Samples were run into a 4–12% SDS-PAGE gel to create a ∼10 mm length region, allowing total protein to be evaluated in a single gel digest. After staining with SimplyBlue SafeStain (Invitrogen), these regions were excised, cut into 1 mm cubes, destained, then reduced and alkylated with DTT and IAA, respectively (Sigma). Gel pieces were dehydrated with acetonitrile. Aliquots of trypsin (Promega) were reconstituted to 10 ng/μL in 50 mM ammonium bicarbonate and added so that the solution was just covering the dehydrated gel pieces. After half an hour at room temp, additional ammonium bicarbonate was added until the gel pieces were completely submerged and placed at 37 °C overnight. Peptides were extracted the next day by addition of 50% acetonitrile, 0.1% TFA, then dried down in a CentriVap concentrator (Labconco). Peptides were desalted with homemade C18 spin columns, dried again, and reconstituted in 0.1% TFA. Peptides were injected onto a homemade 30 cm C18 column with 1.8 um beads (Sepax), with an Easy nLC-1000 HPLC (Thermo Fisher), connected to a Q Exactive Plus mass spectrometer (Thermo Fisher): solvent A 0.1% formic acid in water, solvent B 0.1% formic acid in acetonitrile, gradient 3% B for 2 min, then ramp to 30% B over 41 min, then to 70% over 3 min, then 70% for 4 min, then re-equilibrated to 3%, for a total run time of 60 min. The Q Exactive Plus was operated in data-dependent mode, with a full MS1 scan followed by 10 data-dependent MS2 scans. The full scan was done over a range of 400–1400 m/z, with a resolution of 70 000 at m/z of 200, an AGC target of 1e6, and a maximum injection time of 50 ms. The MS2 scans were performed at 17 500 resolution, with an AGC target of 5e4 and a maximum injection time of 120 ms. The isolation width was 1.5 m/z, with an offset of 0.3 m/z, and a normalized collision energy of 27. Raw data was searched using the SEQUEST search engine within the Proteome Discoverer software platform, version 2.2 (Thermo Fisher), using the SwissProt human database. Trypsin was selected as the enzyme allowing up to 2 missed cleavages, with an MS1 mass tolerance of 10 ppm, and an MS2 mass tolerance of 25 mmu. Carbamidomethyl was set as a fixed modification, while oxidation of methionine was set as a variable modification. Minora was used to determine relative protein abundance between samples. Normalization was performed using the total peptide amount option in the precursor ions quantifier node. Percolator was used as the FDR calculator, filtering out peptides which had a q-value greater than 0.01. Missing values were imputed from the normal distribution using Perseus software. The identified proteins were evaluated using an unpaired t test between the MCL treated (control) and DMSO treated (sample) groups with a p-value cutoff of 0.05. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE78 partner repository with the data set identifier PXD024455.
Acknowledgments
This research was supported by the U.S. National Institute of Health (NIH) R01 grant GM098628 (R.F.), NIH R35 grant R35 GM119502 (S.G.), and U.S. National Science Foundation (NSF) grant CHE-1609550 (R.F.). C.T.J. is supported by the Nancy Carroll Allen Chair in Hematology. MS instrumentation at the University of Rochester was supported by the National Science Foundation grants CHE-0840410 and CHE-0946653 and by the NIH grant S10 OD025242. The authors are grateful to Dr. William Brennessel for assistance with crystallographic analyses. X-ray instrumentation is supported by the U.S. NSF grant CHE-1725028.
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acscentsci.0c01624.
Author Contributions
○ H.A. and S.G. contributed equally to this work.
The authors declare the following competing financial interest(s): A patent application was filed by the University of Rochester that covers methods, enzymes and/or compounds described in this work.
Supplementary Material
References
- Breitmaier E.Terpenes: Flavors, Fragrances, Pharmaca, Pheromones; Wiley-VCH Verlag GmbH, Weinheim, 2006. [Google Scholar]
- Ghantous A.; Gali-Muhtasib H.; Vuorela H.; Saliba N. A.; Darwiche N. What made sesquiterpene lactones reach cancer clinical trials?. Drug Discovery Today 2010, 15, 668–678. 10.1016/j.drudis.2010.06.002. [DOI] [PubMed] [Google Scholar]
- Merfort I. Perspectives on Sesquiterpene Lactones in Inflammation and Cancer. Curr. Drug Targets 2011, 12, 1560–1573. 10.2174/138945011798109437. [DOI] [PubMed] [Google Scholar]
- Orofino Kreuger M. R.; Grootjans S.; Biavatti M. W.; Vandenabeele P.; D’Herde K. Sesquiterpene lactones as drugs with multiple targets in cancer treatment: focus on parthenolide. Anti-Cancer Drugs 2012, 23, 883–896. 10.1097/CAD.0b013e328356cad9. [DOI] [PubMed] [Google Scholar]
- Guzman M. L.; Rossi R. M.; Karnischky L.; Li X.; Peterson D. R.; Howard D. S.; Jordan C. T. The sesquiterpene lactone parthenolide induces apoptosis of human acute myelogenous leukemia stem and progenitor cells. Blood 2005, 105, 4163–4169. 10.1182/blood-2004-10-4135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guzman M. L.; Rossi R. M.; Li X. J.; Corbett C.; Hassane D. C.; Bushnell T.; Carroll M.; Sullivan E.; Neelakantan S.; Crooks P. A.; Jordan C. T. A novel orally available parthenolide analog selectively eradicates AML stem and progenitor cells.. Blood 2006, 108, 237. 10.1182/blood.V108.11.237.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neelakantan S.; Nasim S.; Guzman M. L.; Jordan C. T.; Crooks P. A. Aminoparthenolides as novel anti-leukemic agents: Discovery of the NF-kappaB inhibitor, DMAPT (LC-1). Bioorg. Med. Chem. Lett. 2009, 19, 4346–4349. 10.1016/j.bmcl.2009.05.092. [DOI] [PubMed] [Google Scholar]
- Nasim S.; Pei S.; Hagen F. K.; Jordan C. T.; Crooks P. A. Melampomagnolide B: a new antileukemic sesquiterpene. Bioorg. Med. Chem. 2011, 19, 1515–1519. 10.1016/j.bmc.2010.12.045. [DOI] [PubMed] [Google Scholar]
- Kolev J. N.; O’Dwyer K. M.; Jordan C. T.; Fasan R. Discovery of potent parthenolide-based antileukemic agents enabled by late-stage P450-mediated C-H functionalization. ACS Chem. Biol. 2014, 9, 164–173. 10.1021/cb400626w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kempema A. M.; Widen J. C.; Hexum J. K.; Andrews T. E.; Wang D.; Rathe S. K.; Meece F. A.; Noble K. E.; Sachs Z.; Largaespada D. A.; Harki D. A. Synthesis and antileukemic activities of C1-C10-modified parthenolide analogues. Bioorg. Med. Chem. 2015, 23, 4737–4745. 10.1016/j.bmc.2015.05.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lapidot T.; Sirard C.; Vormoor J.; Murdoch B.; Hoang T.; Cacerescortes J.; Minden M.; Paterson B.; Caligiuri M. A.; Dick J. E. A Cell Initiating Human Acute Myeloid-Leukemia after Transplantation into Scid Mice. Nature 1994, 367, 645–648. 10.1038/367645a0. [DOI] [PubMed] [Google Scholar]
- Hope K. J.; Jin L.; Dick J. E. Acute myeloid leukemia originates from a hierarchy of leukemic stem cell classes that differ in self-renewal capacity. Nat. Immunol. 2004, 5, 738–743. 10.1038/ni1080. [DOI] [PubMed] [Google Scholar]
- Guzman M. L.; Neering S. J.; Upchurch D.; Grimes B.; Howard D. S.; Rizzieri D. A.; Luger S. M.; Jordan C. T. Nuclear factor-kappaB is constitutively activated in primitive human acute myelogenous leukemia cells. Blood 2001, 98, 2301–2307. 10.1182/blood.V98.8.2301. [DOI] [PubMed] [Google Scholar]
- Costello R. T.; Mallet F.; Gaugler B.; Sainty D.; Arnoulet C.; Gastaut J. A.; Olive D. Human acute myeloid leukemia CD34+/CD38- progenitor cells have decreased sensitivity to chemotherapy and Fas-induced apoptosis, reduced immunogenicity, and impaired dendritic cell transformation capacities. Cancer Res. 2000, 60, 4403–4411. [PubMed] [Google Scholar]
- Pollyea D. A.; Gutman J. A.; Gore L.; Smith C. A.; Jordan C. T. Targeting acute myeloid leukemia stem cells: a review and principles for the development of clinical trials. Haematologica 2014, 99, 1277–1284. 10.3324/haematol.2013.085209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guzman M. L.; Li X.; Corbett C. A.; Rossi R. M.; Bushnell T.; Liesveld J. L.; Hebert J.; Young F.; Jordan C. T. Rapid and selective death of leukemia stem and progenitor cells induced by the compound 4-benzyl, 2-methyl, 1,2,4-thiadiazolidine, 3,5 dione (TDZD-8). Blood 2007, 110, 4436–4444. 10.1182/blood-2007-05-088815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDermott S. P.; Eppert K.; Notta F.; Isaac M.; Datti A.; Al-Awar R.; Wrana J.; Minden M. D.; Dick J. E. A small molecule screening strategy with validation on human leukemia stem cells uncovers the therapeutic efficacy of kinetin riboside. Blood 2012, 119, 1200–1207. 10.1182/blood-2011-01-330019. [DOI] [PubMed] [Google Scholar]
- Lagadinou E. D.; Sach A.; Callahan K.; Rossi R. M.; Neering S. J.; Minhajuddin M.; Ashton J. M.; Pei S.; Grose V.; O’Dwyer K. M.; Liesveld J. L.; Brookes P. S.; Becker M. W.; Jordan C. T. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 2013, 12, 329–341. 10.1016/j.stem.2012.12.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartwell K. A.; Miller P. G.; Mukherjee S.; Kahn A. R.; Stewart A. L.; Logan D. J.; Negri J. M.; Duvet M.; Jaras M.; Puram R.; Dancik V.; Al-Shahrour F.; Kindler T.; Tothova Z.; Chattopadhyay S.; Hasaka T.; Narayan R.; Dai M.; Huang C.; Shterental S.; Chu L. P.; Haydu J. E.; Shieh J. H.; Steensma D. P.; Munoz B.; Bittker J. A.; Shamji A. F.; Clemons P. A.; Tolliday N. J.; Carpenter A. E.; Gilliland D. G.; Stern A. M.; Moore M. A.; Scadden D. T.; Schreiber S. L.; Ebert B. L.; Golub T. R. Niche-based screening identifies small-molecule inhibitors of leukemia stem cells. Nat. Chem. Biol. 2013, 9, 840–848. 10.1038/nchembio.1367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Q.; Lu Y.; Ding Y.; Zhai J.; Ji Q.; Ma W.; Yang M.; Fan H.; Long J.; Tong Z.; Shi Y.; Jia Y.; Han B.; Zhang W.; Qiu C.; Ma X.; Li Q.; Shi Q.; Zhang H.; Li D.; Zhang J.; Lin J.; Li L. Y.; Gao Y.; Chen Y. Guaianolide sesquiterpene lactones, a source to discover agents that selectively inhibit acute myelogenous leukemia stem and progenitor cells. J. Med. Chem. 2012, 55, 8757–8769. 10.1021/jm301064b. [DOI] [PubMed] [Google Scholar]
- Ji Q.; Ding Y. H.; Sun Y.; Zhang Y.; Gao H. E.; Song H. N.; Yang M.; Liu X. L.; Zhang Z. X.; Li Y. H.; Gao Y. D. Antineoplastic effects and mechanisms of micheliolide in acute myelogenous leukemia stem cells. Oncotarget 2016, 7, 65012–65023. 10.18632/oncotarget.11342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu L. L.; Chen W. C.; Tang Q. S.; Ji K. Y. Micheliolide Inhibits Liver Cancer Cell Growth Via Inducing Apoptosis And Perturbing Actin Cytoskeleton. Cancer Manage. Res. 2019, 11, 9203–9212. 10.2147/CMAR.S216870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- An Y. H.; Guo W. J.; Li L. N.; Xu C. W.; Yang D. X.; Wang S. S.; Lu Y. X.; Zhang Q.; Zhai J. D.; Fan H. X.; Qiu C. J.; Qi J.; Chen Y.; Yuan S. J. Micheliolide Derivative DMAMCL Inhibits Glioma Cell Growth In Vitro and In Vivo. PLoS One 2015, 10, e0116202. 10.1371/journal.pone.0116202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu N.; Hua Z. Y.; Ba G.; Zhang S. M.; Liu Z. H.; Thiele C. J.; Li Z. J. The anti-tumor growth effect of a novel agent DMAMCL in rhabdomyosarcoma in vitro and in vivo. J. Exp. Clin. Cancer Res. 2019, 10.1186/s13046-019-1107-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao S. N.; Ye J. P.; Yin M. Q.; Yu R. DMAMCL exerts antitumor effects on hepatocellular carcinoma both in vitro and in vivo. Cancer Lett. 2020, 483, 87–97. 10.1016/j.canlet.2020.04.003. [DOI] [PubMed] [Google Scholar]
- Kwok B. H.; Koh B.; Ndubuisi M. I.; Elofsson M.; Crews C. M. The anti-inflammatory natural product parthenolide from the medicinal herb Feverfew directly binds to and inhibits IkappaB kinase. Chem. Biol. 2001, 8, 759–766. 10.1016/S1074-5521(01)00049-7. [DOI] [PubMed] [Google Scholar]
- Garcia-Pineres A. J.; Castro V.; Mora G.; Schmidt T. J.; Strunck E.; Pahl H. L.; Merfort I. Cysteine 38 in p65/NF-kappaB plays a crucial role in DNA binding inhibition by sesquiterpene lactones. J. Biol. Chem. 2001, 276, 39713–39720. 10.1074/jbc.M101985200. [DOI] [PubMed] [Google Scholar]
- Garcia-Pineres A. J.; Lindenmeyer M. T.; Merfort I. Role of cysteine residues of p65/NF-kappaB on the inhibition by the sesquiterpene lactone parthenolide and N-ethyl maleimide, and on its transactivating potential. Life Sci. 2004, 75, 841–856. 10.1016/j.lfs.2004.01.024. [DOI] [PubMed] [Google Scholar]
- Nasim S.; Crooks P. A. Antileukemic activity of aminoparthenolide analogs. Bioorg. Med. Chem. Lett. 2008, 18, 3870–3873. 10.1016/j.bmcl.2008.06.050. [DOI] [PubMed] [Google Scholar]
- Han C.; Barrios F. J.; Riofski M. V.; Colby D. A. Semisynthetic derivatives of sesquiterpene lactones by palladium-catalyzed arylation of the alpha-methylene-gamma-lactone substructure. J. Org. Chem. 2009, 74, 7176–7179. 10.1021/jo901533e. [DOI] [PubMed] [Google Scholar]
- Ma W. W.; Shi Q. Q.; Ding Y. H.; Long J.; Zhang Q.; Chen Y. Synthesis of micheliolide derivatives and their activities against AML progenitor cells. Molecules 2013, 18, 5980–5992. 10.3390/molecules18055980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fasan R. Tuning P450 Enzymes as Oxidation Catalysts. ACS Catal. 2012, 2, 647–666. 10.1021/cs300001x. [DOI] [Google Scholar]
- Reetz M. T. Biocatalysis in Organic Chemistry and Biotechnology: Past, Present, and Future. J. Am. Chem. Soc. 2013, 135, 12480–12496. 10.1021/ja405051f. [DOI] [PubMed] [Google Scholar]
- Lewis J. C.; Coelho P. S.; Arnold F. H. Enzymatic functionalization of carbon-hydrogen bonds. Chem. Soc. Rev. 2011, 40, 2003–2021. 10.1039/C0CS00067A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rentmeister A.; Arnold F. H.; Fasan R. Chemo-enzymatic fluorination of unactivated organic compounds. Nat. Chem. Biol. 2009, 5, 26–28. 10.1038/nchembio.128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sowden R. J.; Yasmin S.; Rees N. H.; Bell S. G.; Wong L. L. Biotransformation of the sesquiterpene (+)-valencene by cytochrome P450(cam) and P450(BM-3). Org. Biomol. Chem. 2005, 3, 57–64. 10.1039/b413068e. [DOI] [PubMed] [Google Scholar]
- Kille S.; Zilly F. E.; Acevedo J. P.; Reetz M. T. Regio- and stereoselectivity of P450-catalysed hydroxylation of steroids controlled by laboratory evolution. Nat. Chem. 2011, 3, 738–743. 10.1038/nchem.1113. [DOI] [PubMed] [Google Scholar]
- Zhang K. D.; Shafer B. M.; Demars M. D.; Stern H. A.; Fasan R. Controlled Oxidation of Remote sp(3) C-H Bonds in Artemisinin via P450 Catalysts with Fine-Tuned Regio- and Stereoselectivity. J. Am. Chem. Soc. 2012, 134, 18695–18704. 10.1021/ja3073462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le-Huu P.; Heidt T.; Claasen B.; Laschat S.; Urlacher V. B. Chemo-, Regio-, and Stereoselective Oxidation of the Monocyclic Diterpenoid beta-Cembrenediol by P450 BM3. ACS Catal. 2015, 5, 1772–1780. 10.1021/cs5020404. [DOI] [Google Scholar]
- Loskot S. A.; Romney D. K.; Arnold F. H.; Stoltz B. M. Enantioselective Total Synthesis of Nigelladine A via Late-Stage C-H Oxidation Enabled by an Engineered P450 Enzyme. J. Am. Chem. Soc. 2017, 139, 10196–10199. 10.1021/jacs.7b05196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowell A. N.; DeMars M. D.; Slocum S. T.; Yu F. A.; Anand K.; Chemler J. A.; Korakavi N.; Priessnitz J. K.; Park S. R.; Koch A. A.; Schultz P. J.; Sherman D. H. Chemoenzymatic Total Synthesis and Structural Diversification of Tylactone-Based Macrolide Antibiotics through Late-Stage Polyketide Assembly, Tailoring, and C-H Functionalization. J. Am. Chem. Soc. 2017, 139, 7913–7920. 10.1021/jacs.7b02875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Acevedo-Rocha C. G.; Gamble C. G.; Lonsdale R.; Li A. T.; Nett N.; Hoebenreich S.; Lingnau J. B.; Wirtz C.; Fares C.; Hinrichs H.; Deege A.; Mulholland A. J.; Nov Y.; Leys D.; McLean K. J.; Munro A. W.; Reetz M. T. P450-Catalyzed Regio- and Diastereoselective Steroid Hydroxylation: Efficient Directed Evolution Enabled by Mutability Landscaping. ACS Catal. 2018, 8, 3395–3410. 10.1021/acscatal.8b00389. [DOI] [Google Scholar]
- Tyagi V.; Alwaseem H.; O’Dwyer K. M.; Ponder J.; Li Q. Y.; Jordan C. T.; Fasan R. Chemoenzymatic synthesis and antileukemic activity of novel C9-and C14-functionalized parthenolide analogs. Bioorg. Med. Chem. 2016, 24, 3876–3886. 10.1016/j.bmc.2016.06.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munro A. W.; Leys D. G.; McLean K. J.; Marshall K. R.; Ost T. W.; Daff S.; Miles C. S.; Chapman S. K.; Lysek D. A.; Moser C. C.; Page C. C.; Dutton P. L. P450 BM3: the very model of a modern flavocytochrome. Trends Biochem. Sci. 2002, 27, 250–257. 10.1016/S0968-0004(02)02086-8. [DOI] [PubMed] [Google Scholar]
- Whitehouse C. J.; Bell S. G.; Wong L. L. P450(BM3) (CYP102A1): connecting the dots. Chem. Soc. Rev. 2012, 41, 1218–1260. 10.1039/C1CS15192D. [DOI] [PubMed] [Google Scholar]
- Zhang K.; El Damaty S.; Fasan R. P450 Fingerprinting Method for Rapid Discovery of Terpene Hydroxylating P450 Catalysts with Diversified Regioselectivity. J. Am. Chem. Soc. 2011, 133, 3242–3245. 10.1021/ja109590h. [DOI] [PubMed] [Google Scholar]
- Kubo T.; Peters M. W.; Meinhold P.; Arnold F. H. Enantioselective epoxidation of terminal alkenes to (R)- and (S)-epoxides by engineered cytochromes P450 BM-3. Chem. - Eur. J. 2006, 12, 1216–1220. 10.1002/chem.200500584. [DOI] [PubMed] [Google Scholar]
- Tang W. L.; Li Z.; Zhao H. Inverting the enantioselectivity of P450pyr monooxygenase by directed evolution. Chem. Commun. 2010, 46, 5461–5463. 10.1039/c0cc00735h. [DOI] [PubMed] [Google Scholar]
- Kolev J. N.; Zaengle J. M.; Ravikumar R.; Fasan R. Enhancing the efficiency and regioselectivity of P450 oxidation catalysts by unnatural amino acid mutagenesis. ChemBioChem 2014, 15, 1001–1010. 10.1002/cbic.201400060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H.; Poulos T. L. The structure of the cytochrome p450BM-3 haem domain complexed with the fatty acid substrate, palmitoleic acid. Nat. Struct. Biol. 1997, 4, 140–146. 10.1038/nsb0297-140. [DOI] [PubMed] [Google Scholar]
- Noble M. A.; Miles C. S.; Chapman S. K.; Lysek D. A.; Mackay A. C.; Reid G. A.; Hanzlik R. P.; Munro A. W. Roles of active site residues in flavocytochrome P450 BM3. Biochem. J. 1999, 339, 371–379. 10.1042/bj3390371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ost T. W.; Miles C. S.; Murdoch J.; Cheung Y.; Reid G. A.; Chapman S. K.; Munro A. W. Rational re-design of the substrate binding site of flavocytochrome P450 BM3. FEBS Lett. 2000, 486, 173–177. 10.1016/S0014-5793(00)02267-5. [DOI] [PubMed] [Google Scholar]
- Carmichael A. B.; Wong L. L. Protein engineering of Bacillus megaterium CYP102. The oxidation of polycyclic aromatic hydrocarbons. Eur. J. Biochem. 2001, 268, 3117–3125. 10.1046/j.1432-1327.2001.02212.x. [DOI] [PubMed] [Google Scholar]
- Kuehnel K.; Maurer S. C.; Galeyeva Y.; Frey W.; Laschat S.; Urlacher V. B. Hydroxylation of dodecanoic acid and (2R,4R,6R,8R)-tetramethyldecanol on a preparative scale using an NADH-dependent CYP102A1 mutant. Adv. Synth. Catal. 2007, 349, 1451–1461. 10.1002/adsc.200700054. [DOI] [Google Scholar]
- van Vugt-Lussenburg B. M. A.; Damsten M. C.; Maasdijk D. M.; Vermeulen N. P. E.; Commandeur J. N. M. Heterotropic and homotropic cooperativity by a drug-metabolising mutant of cytochrome P450BM3. Biochem. Biophys. Res. Commun. 2006, 346, 810–818. 10.1016/j.bbrc.2006.05.179. [DOI] [PubMed] [Google Scholar]
- Nazor J.; Schwaneberg U. Laboratory evolution of P450BM-3 for mediated electron transfer. ChemBioChem 2006, 7, 638–644. 10.1002/cbic.200500436. [DOI] [PubMed] [Google Scholar]
- Fasan R.; Meharenna Y. T.; Snow C. D.; Poulos T. L.; Arnold F. H. Evolutionary history of a specialized P450 propane monooxygenase. J. Mol. Biol. 2008, 383, 1069–1080. 10.1016/j.jmb.2008.06.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barabe F.; Kennedy J. A.; Hope K. J.; Dick J. E. Modeling the initiation and progression of human acute leukemia in mice. Science 2007, 316, 600–604. 10.1126/science.1139851. [DOI] [PubMed] [Google Scholar]
- Taunton J.; Hassig C. A.; Schreiber S. L. A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 1996, 272, 408–411. 10.1126/science.272.5260.408. [DOI] [PubMed] [Google Scholar]
- Statsuk A. V.; Bai R. L.; Baryza J. L.; Verma V. A.; Hamel E.; Wender P. A.; Kozmin S. A. Actin is the primary cellular receptor of bistramide A. Nat. Chem. Biol. 2005, 1, 383–388. 10.1038/nchembio748. [DOI] [PubMed] [Google Scholar]
- Yang P. Y.; Liu K.; Ngai M. H.; Lear M. J.; Wenk M. R.; Yao S. Q. Activity-Based Proteome Profiling of Potential Cellular Targets of Orlistat - An FDA-Approved Drug with Anti-Tumor Activities. J. Am. Chem. Soc. 2010, 132, 656–666. 10.1021/ja907716f. [DOI] [PubMed] [Google Scholar]
- Li J.; Cisar J. S.; Zhou C. Y.; Vera B.; Williams H.; Rodriguez A. D.; Cravatt B. F.; Romo D. Simultaneous structure-activity studies and arming of natural products by C-H amination reveal cellular targets of eupalmerin acetate. Nat. Chem. 2013, 5, 510–517. 10.1038/nchem.1653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreuzer J.; Bach N. C.; Forler D.; Sieber S. A. Target discovery of acivicin in cancer cells elucidates its mechanism of growth inhibition. Chem. Sci. 2015, 6, 237–245. 10.1039/C4SC02339K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brand S.; Roy S.; Schroder P.; Rathmer B.; Roos J.; Kapoor S.; Patil S.; Pommerenke C.; Maier T.; Janning P.; Eberth S.; Steinhilber D.; Schade D.; Schneider G.; Kumar K.; Ziegler S.; Waldmann H. Combined Proteomic and In Silico Target Identification Reveal a Role for 5-Lipoxygenase in Developmental Signaling Pathways. Cell Chem. Biol. 2018, 25, 1095. 10.1016/j.chembiol.2018.05.016. [DOI] [PubMed] [Google Scholar]
- Llabani E.; Hicklin R. W.; Lee H. Y.; Motika S. E.; Crawford L. A.; Weerapana E.; Hergenrother P. J. Diverse compounds from pleuromutilin lead to a thioredoxin inhibitor and inducer of ferroptosis. Nat. Chem. 2019, 11, 521–532. 10.1038/s41557-019-0261-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leslie B. J.; Hergenrother P. J. Identification of the cellular targets of bioactive small organic molecules using affinity reagents. Chem. Soc. Rev. 2008, 37, 1347–1360. 10.1039/b702942j. [DOI] [PubMed] [Google Scholar]
- Li J.; Li S. S.; Guo J. S.; Li Q. Y.; Long J.; Ma C.; Ding Y. H.; Yan C. L.; Li L. W.; Wu Z. G.; Zhu H.; Li K. K.; Wen L. Q.; Zhang Q.; Xue Q. Q.; Zhao C. L.; Liu N.; Ivanov I.; Luo M.; Xi R. M.; Long H. B.; Wang P. G.; Chen Y. Natural Product Micheliolide (MCL) Irreversibly Activates Pyruvate Kinase M2 and Suppresses Leukemia. J. Med. Chem. 2018, 61, 4155–4164. 10.1021/acs.jmedchem.8b00241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pei S. S.; Minhajuddin M.; D’Alessandro A.; Nemkov T.; Stevens B. M.; Adane B.; Khan N.; Hagen F. K.; Yadav V. K.; De S.; Ashton J. M.; Hansen K. C.; Gutman J. A.; Pollyea D. A.; Crooks P. A.; Smith C.; Jordan C. T. Rational Design of a Parthenolide-based Drug Regimen That Selectively Eradicates Acute Myelogenous Leukemia Stem Cells. J. Biol. Chem. 2016, 291, 21984–22000. 10.1074/jbc.M116.750653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang X. R.; Wang Y. L.; Qin Y. F.; He W. G.; Benlahrech A.; Zhang Q. W.; Jiang X.; Lu Z. H.; Ji G.; Zheng Y. J. Micheliolide provides protection of mice against Staphylococcus aureus and MRSA infection by down-regulating inflammatory response. Sci. Rep. 2017, 10.1038/srep41964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin X.; Jiang X.; Jiang X.; Wang Y.; Miao Z.; He W.; Yang G.; Lv Z.; Yu Y.; Zheng Y. Micheliolide inhibits LPS-induced inflammatory response and protects mice from LPS challenge. Sci. Rep. 2016, 6, 23240. 10.1038/srep23240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Netto L. E. S.; Antunes F. The Roles of Peroxiredoxin and Thioredoxin in Hydrogen Peroxide Sensing and in Signal Transduction. Mol. Cells 2016, 39, 65–71. 10.14348/molcells.2016.2349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang S. W.; Chae H. Z.; Seo M. S.; Kim K. H.; Baines I. C.; Rhee S. G. Mammalian peroxiredoxin isoforms can reduce hydrogen peroxide generated in response to growth factors and tumor necrosis factor-alpha. J. Biol. Chem. 1998, 273, 6297–6302. 10.1074/jbc.273.11.6297. [DOI] [PubMed] [Google Scholar]
- Nitti M.; Piras S.; Marinari U. M.; Moretta L.; Pronzato M. A.; Furfaro A. L. HO-1 Induction in Cancer Progression: A Matter of Cell Adaptation. Antioxidants 2017, 6, 29. 10.3390/antiox6020029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guzman M. L.; Rossi R. M.; Neelakantan S.; Li X.; Corbett C. A.; Hassane D. C.; Becker M. W.; Bennett J. M.; Sullivan E.; Lachowicz J. L.; Vaughan A.; Sweeney C. J.; Matthews W.; Carroll M.; Liesveld J. L.; Crooks P. A.; Jordan C. T. An orally bioavailable parthenolide analog selectively eradicates acute myelogenous leukemia stem and progenitor cells. Blood 2007, 110, 4427–4435. 10.1182/blood-2007-05-090621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee E.; Lee T. A.; Yoo H. J.; Lee S.; Park B. CNBP controls tumor cell biology by regulating tumor-promoting gene expression. Mol. Carcinog. 2019, 58, 1492–1501. 10.1002/mc.23030. [DOI] [PubMed] [Google Scholar]
- Zhu G. H.; Dai H. P.; Shen Q.; Zhang Q. Downregulation of LPXN expression by siRNA decreases the malignant proliferation and transmembrane invasion of SHI-1 cells. Oncol. Lett. 2018, 17, 135–140. 10.3892/ol.2018.9605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McLachlan M. J.; Johannes T. W.; Zhao H. Further improvement of phosphite dehydrogenase thermostability by saturation mutagenesis. Biotechnol. Bioeng. 2008, 99, 268–274. 10.1002/bit.21546. [DOI] [PubMed] [Google Scholar]
- Perez-Riverol Y.; Csordas A.; Bai J. W.; Bernal-Llinares M.; Hewapathirana S.; Kundu D. J.; Inuganti A.; Griss J.; Mayer G.; Eisenacher M.; Perez E.; Uszkoreit J.; Pfeuffer J.; Sachsenberg T.; Yilmaz S.; Tiwary S.; Cox J.; Audain E.; Walzer M.; Jarnuczak A. F.; Ternent T.; Brazma A.; Vizcaino J. A. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. 10.1093/nar/gky1106. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.