Abstract
Dynamic modification of RNA affords proximal regulation of gene expression triggered by non-genomic or environmental changes. One such epitranscriptomic alteration in RNA metabolism is the installation of a methyl group on adenosine [N6-methyladenosine (m6A)] known to be the most prevalent modified state of messenger RNA (mRNA) in the mammalian cell. The methylation machinery responsible for the dynamic deposition and recognition of m6A on mRNA is composed of subunits that play specific roles, including reading, writing, and erasing of m6A marks on mRNA to influence gene expression. As a result, peculiar cellular perturbations have been linked to dysregulation of components of the mRNA methylation machinery or its cofactors. It is increasingly clear that neural tissues/cells, especially in the brain, make the most of m6A modification in maintaining normal morphology and function. Neurons in particular display dynamic distribution of m6A marks during development and in adulthood. Interestingly, such dynamic m6A patterns are responsive to external cues and experience. Specific disturbances in the neural m6A landscape lead to anomalous phenotypes, including aberrant stem/progenitor cell proliferation and differentiation, defective cell fate choices, and abnormal synaptogenesis. Such m6A-linked neural perturbations may singularly or together have implications for syndromic or non-syndromic neurological diseases, given that most RNAs in the brain are enriched with m6A tags. Here, we review the current perspectives on the m6A machinery and function, its role in brain development and possible association with brain disorders, and the prospects of applying the clustered regularly interspaced short palindromic repeats (CRISPR)–dCas13b system to obviate m6A-related neurological anomalies.
Keywords: mRNA methylation, mRNA metabolism, N6-methyladenosine (m6A), cortical development, neurological disorders, clustered regularly interspaced short palindromic repeats (CRISPR)–dCas13b, m6A editing
Introduction
Over 170 chemical modifications of RNA are known to exist in eukaryotes (Boccaletto et al., 2018). These RNA modifications, together referred to as the epitranscriptome, play essential roles in gene expression regulation via affecting RNA metabolism: RNA processing, decay, transport, and translation. N6-methyladenosine (m6A) is among the characterized adenosine methylations of messenger RNAs (mRNAs) (Engel and Chen, 2018) and the most occurring in mammalian cells (Roundtree et al., 2017a). The m6A mRNA methylome is dynamically regulated by factors that install, remove, or bind the m6A mark on mRNA. Such dynamism in the m6A landscape is known to critically regulate mRNA metabolism to influence gene expression. In essence, m6A modification is reported to modulate several biological events, including cell proliferation, differentiation, and embryonic development, and can also lead to disease conditions when dysregulated (Dominissini et al., 2012; Meyer et al., 2012; Ke et al., 2015; Linder et al., 2015; Zhao et al., 2017; Ries et al., 2019).
The impact of m6A modification on cell biological processes is notable in nervous tissues (Widagdo et al., 2016; Li et al., 2019). This is because neural cells are known to be enriched with m6A-tagged mRNAs. As a result, the developing and adult brain is reported to be ubiquitously enriched with m6A modifications (Chang et al., 2017; Zhang F. et al., 2018). The m6A level in the brain is temporally regulated in the course of its development such that the adult brain registers the highest level of m6A (Meyer et al., 2012). The massive prevalence of m6A in the developing and postnatal brain signifies the importance of m6A modification in regulating brain morphogenesis and function. Indeed, a chunk of the expanding knowledge indicating the phenomenal role of m6A in orchestrating neural development and function includes the proliferation of neural stem cells (NSCs) and other neural precursor cells, neuroprogenitor cell differentiation, gliogenesis, elaboration of neural processes, and synaptic transmission (reviewed in Widagdo and Anggono, 2018; Li et al., 2019; Chokkalla et al., 2020; Dermentzaki and Lotti, 2020; Livneh et al., 2020).
While maintenance of the general m6A homeostasis is indispensable for proper brain development and activity, selective hypomethylation and hypermethylation of gene transcripts are critical mechanistic modalities for normal brain neurodevelopment and functionality. Moreover, the targeted binding of m6A on transcription factor-encoding mRNAs and disease-risk gene transcripts in the developing brain (Zhang et al., 2020) reflects how m6A signaling is critical for brain structure and function in health. It also means that disturbance of the m6A RNA methylome can have implications for abnormal anatomy and physiology of the brain, leading to neurological disorders.
In this review, we present the role of the m6A methylation machinery in mRNA metabolism, with discussion focused on how the m6A landscape regulates brain development and function. Neurodevelopmental, neurodegenerative, and neuropsychiatric disorders of the brain attributable to m6A signaling dysregulation are highlighted.
The N6-Methyladenosine Modification Machinery
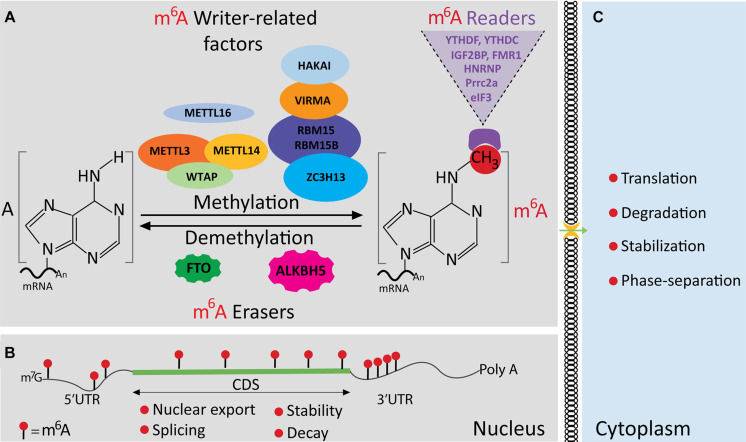
Early studies revealed that m6A is the most substantial posttranscriptional modification in eukaryotic mRNAs (Desrosiers et al., 1974; Perry et al., 1975a, b). It was proposed that nearly 7,000 mRNAs from human and mouse transcriptome contain m6A modification (Dominissini et al., 2012; Meyer et al., 2012). Currently, over 10,000 m6A-modified mRNA transcripts have been identified in yeast and mammalian cells (Wang and Zhao, 2016). m6A modification and recognition (binding) are achieved by three functional components of the m6A machinery, namely, m6A methyltransferases (“writers”), m6A demethylases (“erasers”), and m6A binding or interacting proteins (“readers”) (Figure 1A). Through the specific binding of m6A-interacting proteins, m6A mRNA methylation is able to play a central role in regulating several aspects of mRNA metabolism, such as transport, splicing, stability, translation, and phase separation (Figure 1). High-throughput data revealed that m6A modification typically occurs within the consensus sequence RRACH (R stands for G or A; H stands for A, C, or U) (Dominissini et al., 2012; Meyer et al., 2012). The consensus sequence was recently redefined as DRACH, where D stands for G, A, or U (Linder et al., 2015). As depicted in Figure 1B, m6A distribution along mRNA is asymmetric. In general, m6A sites are concentrated in the protein coding region (CDS) near stop codons, followed by the 3′ untranslated region (UTR), and in the 5′ UTR (Figure 1B) (Dominissini et al., 2012; Meyer et al., 2012; Ke et al., 2015; Linder et al., 2015).
FIGURE 1.
The N6-methyladenosine (m6A) machinery and modification of messenger RNA (mRNA). (A) An illustration showing the m6A machinery. It is made of factors that can functionally be categorized as writers, erasers, and readers of m6A. The m6A marks are deposited by the methylation complex (writers), including METTL3, METTL14, and WTAP, which is counteracted by the m6A demethylases (FTO and ALKBH5), leading to m6A removal. The recognition of m6A marks is done by the readers or binding proteins as indicated. (B) Diagram showing a typical m6A distribution in regions of an mRNA located in the nucleus. (C) The m6A readout affects mRNA fates, including trafficking, stability, decay, translation, and localization.
N6-Methyladenosine Writers
The installation of m6A is carried out by ∼1 MDa m6A writer complex composed of the methyltransferase-like protein 3 (METTL3) and METTL14, which heterodimerize (METTL3-METTL14) to function as the enzymatic core of the writer complex (Bokar et al., 1994; Bujnicki et al., 2002; Liu et al., 2014; Iyer et al., 2016). Additionally, other factors are known to interact with the m6A writer complex. These include Wilms tumor 1-associated protein (WTAP) (Ping et al., 2014), VIRMA/KIA1429 (Yue et al., 2018), RNA-binding protein 15 (RBM15) (Patil et al., 2016; Huang and Yin, 2018), ZC3H13 (Knuckles et al., 2018), and HAKAI (Yue et al., 2018) (Figure 1A). These cofactors are regulated by the binding of RNA and the catalytic activity of the enzymatic core of the m6A writer complex (Bujnicki et al., 2002; Liu et al., 2014; Ping et al., 2014; Iyer et al., 2016; Yue et al., 2018).
METTL3 and METTL14
Bokar et al. (1994) partially purified the m6A writer complex using an in vitro methylation system and identified MT-70, a 70-kDa sub-complex possessing S-adenosylmethionine-binding methyltransferase capacity (Bokar et al., 1994). Later, it was renamed METTL3 (Narayan and Rottman, 1988; Bokar et al., 1997). Knockout of METTL3 in cells effectively blocks m6A modification of mRNAs (Zhong et al., 2008; Agarwala et al., 2012; Geula et al., 2015). On the other hand, METTL14 forms a stable heterodimer with METTL3 to form the methyltransferase core of the m6A methylation machinery (Liu et al., 2014; Wang Y. et al., 2014). METTL14, however, lacks enzymatic function and instead acts as an RNA-binding scaffold to augment the enzyme activity of METTL3 by directing the location of SAM methyl group required for the reaction (Śledź and Jinek, 2016; Wang P. et al., 2016; Wang X. et al., 2016). Therefore, METTL3 is the primary enzyme responsible for m6A installation on mRNA.
METTL3-METTL14-Associated Adaptors: WTAP, VIRMA (KIAA1429), RBM15/15B, ZC3H13 (KIAA0853), and HAKAI
The core m6A writer complex METTL3-METTL4 interacts with other adaptor proteins. It was found that FIP37 (the plant homolog of WTAP) co-localized with MTA (Arabidopsis homolog of METTL3) in the nucleus through physical interaction (Zhong et al., 2008). Similar interaction between WTAP and METTL3 was observed in mammalian cells (Liu et al., 2014; Ping et al., 2014; Schwartz et al., 2014). WTAP is key in keeping the METTL3-METTL4 heterodimer in nuclear speckles (Ping et al., 2014). Loss of WTAP leads to the depletion of m6A modification in mRNA, indicating that WTAP may orient METTL3-METTL14 onto targets (Ping et al., 2014). However, the detailed mechanism remains elusive. Of note, it was demonstrated that two classes of m6A sites exist: WTAP-dependent and WTAP-independent sites (Schwartz et al., 2014). VIRMA is known to also interact with the WTAP-METTL3-METTL4 complex (Figure 1A; Schwartz et al., 2014) and indicates its essentiality for the m6A writer complex functionality. Indeed, VIRMA deletion in human cells leads to a significant reduction in mRNA methylation levels, although not as intense as that resulting from WTAP knockdown (Schwartz et al., 2014). Biochemical studies from Yue et al. (2018) demonstrated that VIRMA preferentially regulates m6A modification in the 3’ UTR proximal to stop codon.
Through proteomic studies, Horiuchi et al. (2013) observed that RBM15 and its paralog RBM15B, together with ZC3H13/KIAA0853, and MTA70 (METTL3), associate with WTAP (Horiuchi et al., 2013; Patil et al., 2016), which raises the possibility that RBM15 and RBM15B may also play role(s) in m6A modification. Indeed, silencing of RBM15 and RBM15B led to a demonstrable decrease in m6A levels of mRNA (Patil et al., 2016). Based on Individual-nucleotide resolution UV crosslinking and immunoprecipitation (iCLIP) data, it was proposed that RBM15/15B recruit the m6A methylation machinery to perform m6A modification through binding to uridine-rich regions near DRACH sites. That notwithstanding, it is not always the case that uridine-rich regions exist near m6A sites; therefore, other methylation complex adaptors may mediate the complex binding to such variant m6A sites (Patil et al., 2016; Meyer and Jaffrey, 2017).
ZC3H13/KIAA0853 is also an interactor of the m6A machinery, and it is demonstrated to be crucial in linking RBM15/15B to WTAP (Horiuchi et al., 2013; Knuckles et al., 2018; Wen et al., 2018). Knockdown of ZC3H13 shifts the localization of the m6A adaptors WTAP, Virilizer, and Hakai from nucleus to cytosol in embryonic stem cells and leads to a significant total reduction in m6A level on mRNA (Wen et al., 2018). This reflects an essential role of ZC3H13 in the deposition of m6A on mRNAs. The E3 ubiquitin ligase HAKAI (CBLL1) is another notable factor that interacts with the m6A machinery (Figure 1A; Horiuchi et al., 2013; Rùžièka et al., 2017). However, its function in m6A modification of mRNA in mammals is yet to be established.
Erasers (Demethylases) of N6-Methyladenosine
m6A modification is believed to be a reversible dynamic process premised on the identification of two demethylases: fat mass and obesity-associated protein (FTO) (Jia et al., 2011) and α-ketoglutarate-dependent dioxygenase alkB homolog 5 (ALKBH5) (Figure 1A). However, this important concept has been in controversy due to various supporting data from various studies (Mauer et al., 2017; Darnell et al., 2018; Wei et al., 2018) as discussed below.
Fat Mass and Obesity-Associated Protein
Following an in vitro assay, which demonstrated that FTO erases m6A methylation of mRNA (Jia et al., 2008), it was further shown that downregulation (knockdown) of FTO in HeLa or 293FT cells caused reduction in m6A methylation of mRNA (Jia et al., 2011). In support of this observation, it was identified that a small proportion of m6A peaks of the whole transcriptome increased in Fto knockout mouse (Hess et al., 2013). These evidence consolidates the concept that m6A modification can be reversed by FTO functionality. However, this idea was challenged by another study, in which no significant increase in m6A level was observed in Fto knockout cells (Mauer et al., 2017). Instead, they noticed that FTO exhibits much higher catalytic capacity against m6Am than m6A. These studies indicate that the preferred substrate of FTO may be m6Am (Meyer and Jaffrey, 2017). What could be the explanation behind the discrepancy between these findings? It is worth pointing out that several independent groups have reported that the Kcat/Km of FTO against m6A is in the range of 0.6−0.7 min–1 μM–1 (Jia et al., 2011; Zhu and Yi, 2014; Zou et al., 2016), whereas that from the study of Mauer et al. (2017) is only 0.06 min–1 μM–1, indicating that most likely there is a technique issue behind quantification of the Kcat/Km of FTO against m6A. Additionally, both investigations used different methods to determine the level of m6A, noting that the RNase T1 treatment of mRNA combined with thin-layer chromatography can only measure the m6A in the case of RGACH, but not RAACH (Mauer et al., 2017). Moreover, a recent study further demonstrates that FTO not only demethylates internal m6A but also caps m6Am (Wei et al., 2018). The subcellular distribution of FTO varies among cultured cell lines, which indicates that the pattern of FTO demethylation of m6A in cytosol or nucleus could be cell lineage-dependent. Consistent with the above studies, it was found that FTO plays a vital role in cell cycle and mitosis regulation in an m6A demethylation-dependent manner during spermatogenesis (Huang T. et al., 2018).
Structural studies uncovered that FTO prefers m6A-modified nucleobase, and its demethylase activity can be influenced by the primary and the tertiary structure of target RNA (Zhang X. et al., 2019), thus shedding light on the molecular mechanism behind the demethylation function of FTO. Recent findings show that the transcription of FTO is regulated by a transcriptional factor Zfp217 during adipogenesis, and Zfp217 is critical for FTO to associate with m6A sites, albeit through competition with YTHDF2 for binding sites (Wei et al., 2019).
ALKBH5
ALKBH5 is another m6A factor with demethylase capacity (Jia et al., 2011; Zheng et al., 2013). Manipulating ALKBH5 expression level leads to a slight but significant change in m6A levels in the poly(A) region of mRNA. Compared with FTO, which demethylates m6Am and m6A, ALKBH5 shows specificity for m6A demethylation (Wei et al., 2018). Importantly, m6A-mediated conformational change facilitates distinction of substrates with minor sequence by FTO and ALKBH5 (Zou et al., 2016). As a nuclear protein, ALKBH5 is proposed to only erase the m6A methylation in the nucleus (Meyer and Jaffrey, 2017). The demethylation capacity of ALKBH5 plays important roles in mRNA splicing, transport, stability, and processing. For instance, spermatogenic transcripts with increased m6A levels exhibit increased splicing events in Alkbh5 KO mice (Tang et al., 2018). Recently, it was reported that METTL3 and ALKBH5 counteractively modulate the m6A methylation of TFEB transcript to effect regulation of autophagy (Song et al., 2019). The demethylation activity of ALKBH5 can be regulated by DEAD-Box RNA helicase through physical interaction (Shah et al., 2017).
Readers (Binding Proteins) of N6-Methyladenosine
The functional significance of m6A modification also depends on m6A-binding proteins also referred to as m6A readers. As described below, we categorize the readers in mammalian cells into two groups: YTH domain-containing proteins, including YTHDC1, YTHDC2, and YTHDF1−3 (Hazra et al., 2019), and Non YTH domain-containing proteins, including eIF3 (Meyer et al., 2015), IGF2BPs (Huang H. et al., 2018), HuR (Dominissini et al., 2012; Wang Y. et al., 2014), FMRP (Zhang F. et al., 2018), hnRNPA2/B1/C (Dominissini et al., 2012; Alarcón et al., 2015), and METTLs (Wang et al., 2015).
YTH Domain Containing N6-Methyladenosine-Binding Proteins
This group contains YTHDC1, YTHDC2, and YTHDF1−3 families in mammals. The common YTH domain defines members of this group of m6A binding and determines the nature of m6A reading (Zhang et al., 2010). However, they are not paralogs. This is because of the non-similarity of other aspects of proteins apart from the common YTH domain (Hazra et al., 2019).
YTHDC1 (YT521-B) is the first identified m6A reader, which was found as a protein associated with splicing factors (Imai et al., 1998; Harfmann et al., 1999; Xiao et al., 2016; Hazra et al., 2019). Interestingly, human YTHDC1 shows much greater binding affinity for the m6A-modified mRNA region in the context of Gm6AC (five-fold to six-fold difference) than Am6AC, although the distribution of m6A modification found in the consensus sequence is Gm6AC (70%) and Am6AC (30%) (Xu et al., 2014, 2015). It localizes in various subnuclear bodies close to the nuclear splicing factor (SF) compartments and plays a role in mRNA splicing through physical interaction with splicing factor SRSF3 and SRSF10 (Figure 2A; Xiao et al., 2016). Furthermore, YTHDC1 works together with NXF1 and SRSF3 to regulate m6A-modified mRNA nuclear export (Figure 2B; Roundtree et al., 2017b). Moreover, YTHDC1 has been reported to bind m6A-modified MAT2A mRNA. The m6A modification results in the degradation of MAT2A mRNA, although the detailed mechanism is not known (Shima et al., 2017).
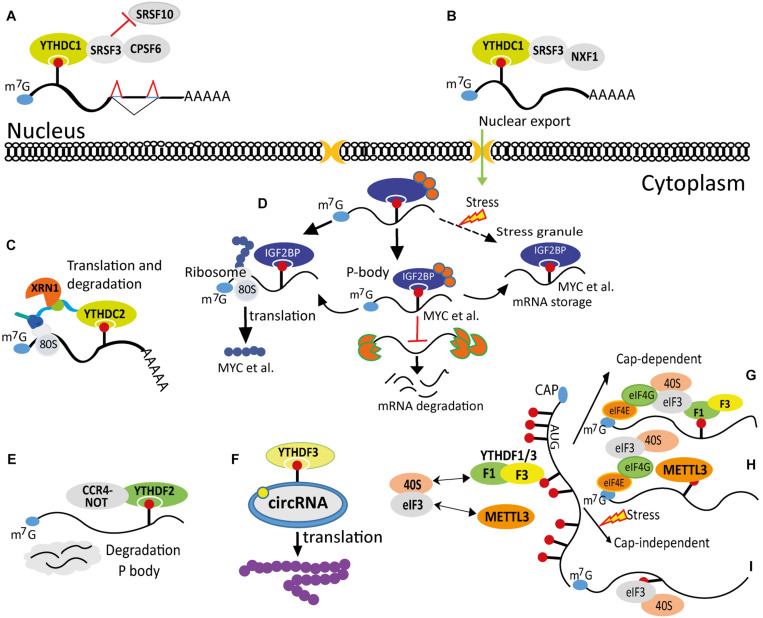
FIGURE 2.
Effects of N6-methyladenosine (m6A) methylation on messenger RNA (mRNA) fate. (A) m6A modification regulates mRNA splicing and polyadenylation via YTHDC1 and its associating factors SRSF3, SRSF10, and CPSF6. (B) m6A modulates mRNA nuclear export through YTHDC1, SRSF3, and NXF1. (C) m6A regulates mRNA translation and stability via YTHDC2-mediated recruitment of the ribosome and the XRN1 exoribonuclease. (D) m6A marks are bound by IGFBPs, which can regulate a subset of mRNA translation, decay in P-body, and storage in stress granules. (E) m6A modification regulates mRNA degradation in P-body through associating with the YTHDF2-CCR4-NOT complex. (F) m6A marks on circRNA modulate its translation via recruiting YTHDF3. (G) m6A marks recruit YTHDF1/YTHDF3 to enhance translation in a Cap-dependent manner. (H) METTL3 serves as an m6A reader and increases translation via recruiting translation initiation complex independent of its methyltransferase activity. (I) m6A directly binds to eIF3 and increases translation in a Cap-independent manner.
On the other hand, YTHDC2 is a multi-domain protein and mainly localized in the cytoplasm, but it is also highly expressed in perinuclear compartment. It prefers to bind m6A-containing RNAs through the YTH domain and enhances RNA degradation. Meanwhile, it also enhances m6A-modified mRNA translation efficiency (Kretschmer et al., 2018).
Human YTHDF1−3 proteins contain a YTH domain in the C-terminus and a low-complexity domain in the N-terminus. These three members of the YTHDF family share high sequence identity and similarity (65%−80%) (Li et al., 2014; Wang X. et al., 2014; Hazra et al., 2019). As a characterized m6A modification reader, the human YTHDF2 binds over 3,000 transcripts primarily in their 3′ UTRs and around the stop codon. The binding of YTHDF2 leads to degradation of the bound mRNAs in cytoplasmic processing bodies (P-bodies). Knockdown of YTHDF2 leads to an accumulation of m6A-containing mRNAs (Wang X. et al., 2014). YTHDF2 was also found to associate with CNOT1, the scaffolding component of the CCR4-NOT mRNA deadenylation complex (Figure 2E). This interaction is required for YTHDF2 to localize in P-bodies (Du et al., 2016). Therefore, the main function of YTHDF2 is to control the degradation of m6A-modified mRNAs (Kang et al., 2014; Hazra et al., 2019).
Unlike YTHDF2, YTHDF1 does not induce the degradation of associated m6A-containing mRNAs. Instead, but arguably, YTHDF1 increases the translation efficiency of associated mRNAs (about 1,200) in an m6A-dependent fashion (Wang et al., 2015). This function of YTHDF1 is further supported by the work of Wu et al. (2019), who showed that YTHDF1 targets m6A-modified Jak2 and regulates its translation (Wu et al., 2019). Recently, another cytoplasmic m6A reader protein YTHDF3 was found to interact with YTHDF1 to promote translation, whereas YTHDF3 interacts with YTHDF2 to reinforce mRNA decay (Li A. et al., 2017; Shi et al., 2017). Furthermore, biochemical studies showed that YTHDF3 shares greater than 50% of common m6A-modified mRNA targets with YTHDF1 and also with YTHDF2 (Li A. et al., 2017; Shi et al., 2017). In addition, YTHDF3 might also function as m6A-modification reader independent of YTHDF1 and YTHDF2 under certain conditions. Oxidative stress induces specific m6A modifications in a set of transcripts, and the binding of YTHDF3 to the modifications triggers the mRNA–YTHDF3 complex localization in the stress granules, but without much influence on YTHDF1 and YTHDF2 (Anders et al., 2018). Importantly, YTHDF3 can also enhance translation independent of METTL3-mediated m6A modification. For example, YTHDF3 functions together with eIF4G2 and Poly(A)-binding protein 1 (PABP1) to promote the translation of forkhead box protein O3 (FOXO3) (Zhang Y. et al., 2019).
Interestingly, very recent studies have shown evidence indicating functional redundancy of the YTHDFs during mRNA degradation and cellular differentiation. As such, it is only when all three YTHDF homologs (YTHDF1−3) are ablated that mRNA stability and cell differentiation regulation become evident (Kontur et al., 2020; Zaccara and Jaffrey, 2020). This may partly stem from the observations that all three YTHDFs are similar in sequence characteristics and usually have common mRNA binding targets (Zaccara and Jaffrey, 2020). Yet, it has been reported that probably due to variation in its expression, YTHDF2 dominates the m6A reader function of all the YTHDFs (Lasman et al., 2020). It was also unraveled that YTHDFs are unable to induce translation in HeLa cells (Zaccara and Jaffrey, 2020). While these new findings present a unified model seeking to define the regulatory functions of YTHDFs in m6A modification, they provoke questions that need to be addressed to reconcile the discrepancy between the recent findings and previous observations with respect to the precise role of YTHDFs in mRNA translation.
Non YTH Domain Containing N6-Methyladenosine Readers (eIF3, IGF2BPs, HuR, FMRP, HNRNP Proteins, and PRRC2A)
Meyer et al. (2015) characterized the function of eIF3 as an m6A reader. eIF3 is preferentially recruited by the m6A-modified mRNA over unmethylated mRNA (Meyer et al., 2015). It was shown that about 35% of m6A marks in the 5′ UTR are also eIF3-binding sites. Depletion of m6A through METTL3 loss-of-function decreased the translation of m6A-modified mRNA in the 5′ UTR, but not the mRNAs bearing m6A marks elsewhere (Meyer et al., 2015). Notably, one of the two modes of m6A-mediated Cap-independent translation is through direct association of m6A in the 5′ UTR and eIF3 (Figure 2I; Meyer et al., 2015), while the other mode involves YTHDF1 association with m6A mark followed by delivery of eIF3 to the 5′ UTR (Figure 2G; Wang et al., 2015). This indicates the correlation between the 5′ UTR m6A and translation and highlights the involvement of eIF3 in the regulation of mRNA translation. Currently, it is not known what the detailed mechanisms are in determining the mode of eIF3−5′ UTR association.
Insulin-like growth factor-2-binding proteins (IGFBPs), including IGFBP1−3, have been reported as RNA-binding proteins (Bell et al., 2013). Recently, it was demonstrated that IGFBP1−3 bind m6A-modified mRNAs with a three-fold to four-fold greater affinity than the m6A-unmodified mRNAs (Huang H. et al., 2018). By means of RIP-Seq or PAR-CLIP-Seq, it was found that IGFBP1−3 share 55%−70% RNA targets with preference for binding to the “UGGAC” consensus motif, e.g., MYC, FSCN1, and TK1 (Huang H. et al., 2018). Knockdown of METTL14, a critical component of the methylation machinery, dramatically undermined IGFBP binding. Interestingly, knockdown of IGF2BPs reduces mRNA stability (Huang H. et al., 2018). Consistently, IGFBPs were found to associate with three mRNA stabilizing factors, including HuR, MATR3, and PABP1, which can support IGFBPs in stabilizing their mRNA targets (Huang H. et al., 2018).
HuR is an RNA-binding protein with multiple molecular functions. It was first described as a stabilizer of ARE-containing mRNAs (Fan and Steitz, 1998; Peng et al., 1998). It is also known to enhance translation, although it can also exert translation suppression (Hinman and Lou, 2008; Abdelmohsen and Gorospe, 2010). This portrays HuR as both a reader and anti-reader of m6A (Dominissini et al., 2012; Wang Y. et al., 2014). However, the underlying mechanism that makes m6A modification sites to recruit or block HuR binding is unknown. We think that a sequence-dependent context may be at play in determining the function of HuR in m6A interaction. This speculation remains to be investigated.
FMR1 (also known as FMRP1) is an RNA-binding protein and known to associate with hundreds of transcripts to decrease their translation. It binds to m6A-modified mRNA in an RNA sequence context-dependent manner. FMR1 selectively binds to the m6A marks associated with GGACU RNA sequence (Edupuganti et al., 2017). Bioinformatic analysis revealed that FMR1 and YTHDF1 shared an abundant set of common m6A-modified mRNAs, indicating that FMR1 might compete with YTHDF1 for binding of m6A-modified mRNAs to downregulate translation (Ascano et al., 2012; Wang et al., 2015). It is possible that the mechanism may underlie the previously reported regulatory function on the translation of mRNA targets.
Heterogeneous nuclear ribonucleoproteins (hnRNPs: hnRNPA2/B1, hnRNPC, and hnRNPG) are RNA-binding proteins that play important roles in pre-RNA processing (Dominissini et al., 2012; Alarcón et al., 2015; Liu et al., 2015, 2017; Xiao et al., 2016). Alarcón et al. (2015) discovered that hnRNPA2B1 interacts with a group of m6A-modified RNAs in the nucleus and regulates their splicing in a comparable pattern as for METTL3. However, the binding of hnRNPA2/B1 to m6A is likely indirect and may require an hnRNPC-mediated switch mechanism to do so (Wu et al., 2018). hnRNPC can read m6A-modified hairpin and m6A-containing RNAs. m6A-modification leads to a change in the regional RNA structure and increases the binding of hnRNPC (Liu et al., 2015). Consistently, general reduction in m6A marks due to METTL3/L14 knockdown eliminates the association of hnRNPC to the aforementioned m6A-mediated RNA structural modification (Liu et al., 2015). Furthermore, HNRNPG is known to bind m6A-modified lncRNA through its C-terminal low-complexity domain (LCD), indicating that LCD domain might be used by some other readers to bind to m6A modification (Liu et al., 2017).
Recently, PRRC2a was reported as an m6A modification reader (Wu et al., 2019). Through RIP-seq and m6A-seq, it was identified that PRRC2a binding peaks within over 2,800 genes in brain samples, and PRRC2a mainly binds to the consensus motif UGGAC in m6A-modified transcripts (Wu et al., 2019). PRRC2A was found to be associated with YTHDF2 in granule-like organelles, which may be involved in the regulation of PRRC2A involvement in Olig2 mRNA stability (Wu et al., 2019). However, since PRRC2A has low tissue expression specificity, it is unclear whether PRRC2A serves as an m6A modification reader in other tissues.
Reader Function of METTLs
Besides its role as an m6A writer, METTL3 can also bind to m6A-modified mRNAs to act as a reader. It was found that METTL3 regulates the translation of some oncogenic m6A-modified mRNAs independent of its methyltransferase activity through eIF3 recruitment to the translation initiation complex (Lin et al., 2016). A study from the same group identified a physical interaction between m6A-bound METTL3 near the stop codon and eIF3h, providing a mechanism to explain how METTL3 can enhance translation (Choe et al., 2018). The methyltransferase METTL16 also serves as an m6A reader in a certain context. When SAM concentrations become low, METTL16 remains bound to m6A-modified MAT2A in its 3′ UTR hp1 site to enhance MAT2A splicing, resulting in increased MAT2A levels in the cytosol. On the contrary, when SAM levels are high, METTL16 methylates MAT2A and facilitates intron retention (Pendleton et al., 2017).
Deposition of N6-Methyladenosine Modification During Transcription
Mechanistically, how m6A modification of transcripts is carried out needs elucidation. A recent study uncovered an insightful detail in the installation of m6A. Specifically, it was found that H3K36me3 cooperates with METTL3/METTL14 to deposit m6A on mRNA (Huang et al., 2019). The study showed that H3K36me3 physically interacts with METT14, thus recruits the m6A methylation machinery to RNA Pol II, and allows the m6A methylation machinery to effect m6A modification during transcription. Decreasing the level of H3K36me3 through loss-of-function of SETD2, the specific enzyme that converts H3K36me2 or H3K36me0 to H3K36me3, significantly led to the reduction in m6A level on RNAs, mimicking the impact of depletion of individual m6A writer complex components (Huang et al., 2019).
Impact of N6-Methyladenosine Modification on Gene Regulation
The reversible modification of m6A exerts functional impact on several aspects of mRNA metabolism, including nuclear export, polyadenylation, splicing, degradation, and translation (Figure 2). By these means, the m6A methylome affords an additional level of gene expression regulation to sculpt the transcriptome (Fu et al., 2014).
N6-Methyladenosine Modification Regulates mRNA Splicing
Some factors involved in m6A modification of mRNA are known to interact with pre-mRNA splicing factors (SRSFs), indicating a possible role for m6A in mRNA splicing (Zhao et al., 2014; Xiao et al., 2016). It has been demonstrated that enrichment of m6A modification promotes recruitment of SRSF2 and leads to enhanced exon inclusion of target mRNA (Zhao et al., 2014). It has been further suggested that the m6A reader YTHDC1 regulates the association of m6A and SRSFs. Indeed, m6A-bound YTHDC1 enhances the recruitment of SRSF3 that favors exon inclusion but blocks the recruitment of SRSF10, an exon skipping-related splicing factor (Xiao et al., 2016). Moreover, hnRNPs may also be involved in the regulation of RNA splicing (Liu et al., 2015, 2017). For example, the modification of m6A on pre-mRNA favors the binding of hnRNPC (Liu et al., 2015), which could further facilitate splicing through its known function in repressing exon inclusion (Zarnack et al., 2013). Therefore, it is possible that perturbation of the m6A machinery components can impair mRNA alternative splicing. This idea is especially supported by the observation that knockdown of METTL3 can antagonize the association of SRSF2 or SRSF3 with m6A-modified pre-mRNAs (Zhao et al., 2014; Xiao et al., 2016), and facilitates the expression of the long isoform of MyD88 (MyD88L) via exon skipping attenuation (Feng et al., 2018). Additional evidence is also based on the essential role played by METTL16 in MAT2A-mediated pre-mRNA alternative splicing (Pendleton et al., 2017).
N6-Methyladenosine Controls Alternative Polyadenylation
Ke et al. (2015) found that m6A modification peaks in the 3′ UTR, especially for transcripts that use alternative polyadenylation (APA), and longer last exons exhibit a higher m6A density. By comparing the m6A density of thousands of mRNA UTRs from liver and brain tissues, it was observed that greater amount m6A marks in the last exons are linked to the usage of more distal polyA sites. Indeed, global reduction of m6A levels via triple knockdown of METTL3, METTL14, and WTAP changed the polyA sites of one-sixth of the examined 661 mRNAs and promoted the usage of proximal APA sites, indicating that some m6A marks inhibit proximal polyadenylation (Ke et al., 2015).
Recently, a mechanism through which m6A controls alternative polyadenylation was proposed. VIRMA (Figure 1A) was found to interact with polyA cleavage factors F5 and CPSF6 (Yue et al., 2018). Consistent with an earlier report, knockdown of METTL3 or VIRMA was found to encourage the usage of distal APA sites, thus lengthening the 3′ UTR of m6A-rich mRNAs. In contrast, CPSF5 knockdown elicits an opposite effect on the length of the 3′ UTR of m6A-marked mRNAs (Yue et al., 2018).
N6-Methyladenosine Promotes Nuclear Export
Considerable amount of nuclear export of mRNAs is regulated by the THO/TREX complex and the nuclear export factor heterodimer NXF1/P15 (Lesbirel and Wilson, 2019). Evidence is accumulating for the role of m6A modification in mRNA nuclear export. Knockdown of METTL3 resulted in delayed nuclear export of specific mRNAs of clock genes (Fustin et al., 2013), indicating the requirement of m6A methylation for specific mRNA nuclear export. Conversely, knockdown of ALKBH5 increased the cytoplasmic accumulation of polyA mRNAs (Zheng et al., 2013). Moreover, VIRMA was observed to interact with the mRNA export factor ALYREF, and its downregulation led to defective mRNA export (Masuda et al., 2005).
Interestingly, several TREX components associate with the components of the core m6A machinery (METTL3-METTL14-WTAP-VIRMA), and TREX also enhances the association of m6A reader YTHDC1 with the mRNA. Knockdown of YTHDC1 also resulted in reduced nuclear export of specific mRNAs (Lesbirel et al., 2018). Taken together, the abovementioned literature demonstrates that m6A modification factors promote mRNA nuclear transport through physical interaction with the mRNA transport machinery.
N6-Methyladenosine Enhances mRNA Degradation
Numerous recent studies suggest that impaired m6A writer complex function reduces m6A modification levels and raises mRNA stability, indicating that m6A methylation drives mRNA degradation (Batista et al., 2014; Schwartz et al., 2014; Wang X. et al., 2014; Wang Y. et al., 2014; Park et al., 2019). Mechanistically, m6A-containing mRNA recruits YTHDF2, which is followed by the translocation of the YTHDF2–mRNA complex from the translation machinery to P-bodies, leading to the degradation of YTHDF2-targeted mRNA. As a result, mRNA targets have increased half-life following YTHDF2 knockdown (Wang X. et al., 2014). It has been clearly demonstrated that YTHDF2 enhances m6A-modified mRNA decay through recruiting CCR4-NOT deadenylase complex via the N-terminus of YTHDF2 and reveals an underlying mechanism by which YTHDF2 regulates degradation of m6A-modified mRNAs (Du et al., 2016).
In a recent study, it was reported that some m6A-modified mRNAs interact with YTHDF2 to undergo decay in an RNase P/MRP-dependent manner and in which HRSP12 serves as a bridge between YTHDF2 and RNase P/MRP (Park et al., 2019). The interaction of human YTHDF2 and HRSP12 was first hinted by the association between their respective yeast homologs Pho92 and Mmf1 (Krogan et al., 2006). It was found in an immunoprecipitation experiment that HRSP12 links YTHDF2 and RNase P/MRP and that the N-terminus of YTHDF2 is required to interact with HRSP12 (Park et al., 2019). Of note, the subset of m6A-modified mRNAs, whose decay depends on YTHDF2–HRSP12–RNase P/MRP complex, contains a specific HRSP12-binding motif proximally upstream of the YTHDF2-binding motif, while the RNase P/MRP cleavage site is downstream and close to the YTHDF2-binding motif (Park et al., 2019). Therefore, this study discloses at least two mechanisms involved in the degradation of YTHDF2-associated m6A RNAs: HRSP12-RNase P/MRP-dependent and CCR4-NOT complex-dependent.
N6-Methyladenosine Modulates Translation
The m6A reader YTHDF1 enhances translation efficiency via interaction with eIF3A/eIF3B, and the YTHDF1-regulated translation likely hinges on eIF4G-dependent loop formation (Wang et al., 2015). According to Meyer et al. (2015), 5′ UTR m6A elevates cap-independent translation through recruiting the 43S complex to form 48S initiation complex in the absence of the cap-associating complex, eIF4F. This mechanism is important for cells to bypass 5′ cap-binding factors to enhance translation under stress conditions (Meyer et al., 2015). Moreover, heat stress-induced cytoplasmic-to-nuclear translocation of YTHDF2 is required for maintaining 5′ UTR m6A levels via competing for binding of the demethylase FTO to m6A sites, which further promotes cap-independent translation (Zhou et al., 2015). YTHDF1 preferentially binds to m6A marks in 3′ UTR of the oncogene CDCP1 mRNA and promotes translation by increasing the amount of polysome-bound (translationally active) CDCP1 transcripts (Yang et al., 2019).
Of note, METTL3 is also involved in m6A-enhanced mRNA translation through its role as an m6A reader in several ways. It promotes mRNA translation via physical association with the translation initiation complex (Lin et al., 2016). It was found that promoter-associated METTL3 regulates m6A methylation inside the coding region and improves mRNA translation through relief of ribosome stalling (Barbieri et al., 2017).
Besides promoting translation efficiency, m6A modification also plays an important role in regulating alternative translation (Zhou J. et al., 2018). It has been reported that widespread alternative translation occurs under various nutrient conditions, but the underlying mechanism is unclear (Gao et al., 2015). Recently, Zhou J. et al. (2018) found that m6A modification in the 5′ UTR modulates the selection of start codon globally, hence driving alternative translation. As representative examples, Atf4 depends on decreased m6A modification of the upstream open reading frame 2 (uORF2) to improve the translation of the major isoform, and Gadd45g heightens the translation of the major isoform by lowering the m6A modification of the 5′ UTR (Zhou J. et al., 2018).
N6-Methyladenosine Methylation Increases the Phase Separation Capacity of mRNA
Only until recently has it become clearer how m6A modification drives mRNA fate and why the consequence of m6A modifications can vary in various scenarios. According to Ries et al. (2019), the m6A readers, YTHDF1−3, experience liquid–liquid phase separation (LLPS). The mRNAs with multiple m6A marks serve as a scaffold to bind with YTHDF readers via their low-complexity regions (LCRs). The mRNA–YTHDF complexes are then transported into various phase separators, like P-bodies, stress granule, and neuronal granules. The study suggests that the number and allocation of m6A modifications in mRNAs remodel the transcriptome of different phase-separated compartments. The efficacy of m6A modification-dependent modulation of an mRNA is likely governed by signals regulating the ability of YTHDF protein involved in LLPS formation (Ries et al., 2019).
N6-Methyladenosine Modification Prominently Regulates Brain Development and Function
Evidence for the role of m6A signaling in modulating the development of the brain and its functions has accumulated in recent years, and the quest for extending the frontier is of great interest. Several investigations have revealed that the various factors that come together to form the m6A methylation machinery exert notable effect(s) on specific aspects of brain morphogenesis to permit optimal neural function, as summarized in Table 1. Conversely, the dysregulation of the m6A methylation machinery is known to elicit perturbations in the neural transcriptome, which have implications for defective development and dysfunction of the brain. The integrity of the m6A machinery functionality is of high priority in cells to the extent that simply ablating its cofactors can have significant consequences for brain development disturbance, as exemplified by the importance of Exosc10-mediated regulation of mRNA stability in forebrain development (Ulmke et al., 2021). The sections below discuss how specific factors associated with the m6A methylation machinery drive neural development, functional adaptation, and plasticity of the brain (Figure 3).
TABLE 1.
m6A mRNA methylation factors and their role in brain development and function.
| Effector | Experimental manipulation | Phenotype | Mechanisms | References |
| Neurogenesis | ||||
| METTL3 | Mettl3fl/fl; Nestin-Cre | Prolongation of the cell cycle of RGCs and protraction of embryonic cortical neurogenesis | m6A depletion caused increased stability of NSC transcripts | Yoon et al. (2017) |
| METTL14 | Mettl14fl/fl; Nestin-Cre | Reduced NSC proliferation and precocious NSC differentiation; loss of late-born neurons during cortical neurogenesis | Stabilization of CBP and p300 transcripts; H3K27me3-mediated transcription suppression of NSC proliferation genes; upregulation of H3K27ac in differentiation-related genes | Wang Y. et al. (2018) |
| RBM15 | OE of RBM15 | NSC delamination | Suppression of BAF155-dependent gene expression | Xie et al. (2019) |
| FTO | KO of FTO | Decrease in adult NSC proliferation and defective hippocampal neurogenesis | Impairment of BDNF and MAPK signaling | Li L. et al. (2017); Spychala and Ruther (2019) |
| YTHDF2 | Ythdf2 fl/fl; Cre (ubiquitously) | Decreased proliferation and differentiation capabilities of NSCs; less Tbr2+ bIPs; Reduced CP thickness | Promotion of m6A-dependent degradation of neurodevelopment-related transcripts | Li M. et al. (2018) |
| FMRP | KO of Fmr1 | Nuclear retention of neurogenic mRNAs; prolonged cell cycle progression in the postnatal mouse brain | Unknown | Edens et al. (2019) |
| Exosc10 | Exosc10fl/fl; Foxg1-Cre; Emx1-Cre | Apoptosis-mediated cortical agenesis | Mediates degradation of Bbc3 and Aen mRNAs, which are effectors of apoptosis | Ulmke et al. (2021) |
| Gliogenesis | ||||
| METTL14 | Mettl14fl/fl; Olig2-Cre | Decrease in oligodendrocytes maturation; cortical hypomyelination | Alters alternative splicing and expression of Nfasc155 | Xu et al. (2020) |
| METTL14 | Mettl14; Nestin-Cre | Reduced number of s100β+ astrocytes | Unknown | Yoon et al. (2017) |
| FTO | FTOfl/fl; Olig2-Cre; Nestin-Cre | Loss of OPCs and Sox10+ cells; cortical hypomyelination | Promotes Olig2 mRNA degradation | Wu et al. (2019) |
| PRRC2A | Prrc2a; Nestin-Cre; Olig2-Cre | Loss of OPCs and mature oligodendrocytes; cortical hypomyelination | Gene targeting of Prrc2a by olig2 mRNA | Wu et al. (2019) |
| PRRC2A | Prrc2a; Nestin-Cre; | Reduced proliferation capacity and number of astrocytes | Competitive expression with YTHDF2 | Wu et al. (2019) |
| Axonogenesis, dendritogenesis, synaptic plasticity | ||||
| YTHDF1 YTHDF3 | KD of Ythdf1 and Ythdf3 | Abnormal dendritic spine morphology | Inhibition of Apc mRNA translation | Merkurjev et al. (2018) |
| METTL14 | Mettl14fl/fl; D1R-Cre | Abnormal excitability of striatal neurons | Unknown | Koranda et al. (2018) |
| FTO | KO of FTO | Defective synaptic plasticity | Unknown | Hess et al. (2013) |
| YTHDF1 YTHDF3 | KD of Ythdf1 and Ythdf3; CRISPR/Cas9-based KO of Ythdf1 | Suppression of neuronal excitability | Not clear | Merkurjev et al. (2018); Shi et al. (2018) |
| Learning and behavior | ||||
| METTL3 | OE of METTL3 | Improved long-term memory consolidation | Unknown | Zhang Z. Y. et al. (2018) |
| METTL14 | Mettl14fl/fl; D1R-Cre | Impaired striatum-mediated behavior patterns | Unknown | Koranda et al. (2018) |
| FTO | CRISPR/Cas9 or shRNA-mediated KD of FTO | Learning disabilities; defective memory processing and verbal fluency | Not clear | Ho et al. (2010); Benedict et al. (2011); Widagdo et al. (2016); Li L. et al. (2017); Walters et al. (2017); Sun et al. (2019) |
| PRRC2A | Prrc2a; Nestin-Cre; Olig2-Cre | Cognitive defects due to cortical hypomyelination | Unknown | Wu et al. (2019) |
| YTHDF1 | CRISPR/Cas9-based KO of Ythdf1; KD of Ythdf1 | Defective long-term potentiation and synaptic transmission in hippocampus; behavioral defects | Unknown | Shi et al. (2018) |
| Circadian clock | ||||
| METTL3 | KD of Mettl3 | Elongation of circadian period | Defective processing of Per2 and Arntl (clock genes) mRNAs | Fustin et al. (2013) |
| Stress response | ||||
| FTO | FTOfl/fl; Camk2a-Cre; Nex-CreERT2 | Reduced ability to cope with stress | Unknown | Engel et al. (2018) |
| METTL3 | Mettl14fl/fl; Camk2a-Cre; Nex-CreERT2 | Reduced ability to cope with stress | Unknown | Engel et al. (2018) |
KO, knockout; KD, knockdown; OE, overexpression; fl/fl, double floxed; RGCs, radial glial cells; OPCs, oligodendrocyte precursor cells; NSCs, neural stem cells; CRISPR, clustered regularly interspaced short palindromic repeats; m6A, N6-methyladenosine.
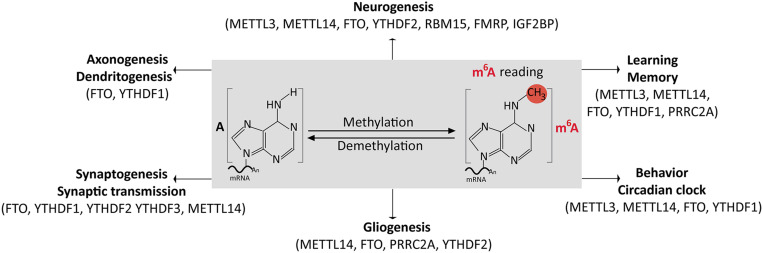
FIGURE 3.
An illustration summarizing the role of N6-methyladenosine (m6A) in brain development and function. The functions of the various m6A-related factors involved in the proliferation and differentiation of neural precursors, neuronal maturation, production of glia, synapse formation during brain development, and common brain physiology are shown.
N6-Methyladenosine Modification Is Indispensable for Neurogenesis in the Brain
Neurons are produced through the process of neurogenesis, which entails specification and proliferation of NSCs, and the differentiation of such neural progenitors into neuroblasts, which undergo maturation to become functional neurons. It has been shown that the dynamic addition of m6A to gene transcripts in the multipotent NSCs greatly influences cortical neuroprogenitor competence and the generation of neurons during brain development (Yao et al., 2016; Boles and Temple, 2017; Yoon et al., 2018; Zhou H. et al., 2018; Rockwell and Hongay, 2019). Dysregulation of writers, erasers, and readers of m6A has been reported to cause notable perturbations in the cell cycle progression, proliferation, and differentiation of NSCs in the developing and adult brain.
Effect of N6-Methyladenosine Writers on Neurogenesis in the Brain
So far, it has been shown that ablation of the m6A writer METTL3 or its cofactor METTL14 in cortical neuroepithelium or isolated cortical NSC results in prolonged cell cycle dynamics of cortical neuroprogenitors and their precocious differentiation into neuronal or neurogenic cells (Batista et al., 2014; Yoon et al., 2017; Wang Y. et al., 2018). Detailed analysis through m6A sequencing revealed that gene transcripts involved in the cell cycle of neural cells, production of neurons, and neuronal differentiation are enriched with m6A tags. Interestingly, the decay of such mRNAs is promoted in the absence of METTL3 and METTL14 (Yoon et al., 2017), meaning that METTL3 and METTL14 are key players in driving neurogenesis via the stabilization of gene transcripts critical for neurogenesis in the brain. For example, loss of m6A due to deletion of METTL3 in mouse cerebellum resulted in overt hypoplasia partly attributable to apoptosis of cerebellar granule cells (Wang C. X. et al., 2018). Key downstream effects of m6A on genes important for neurogenesis include the modulation of histone modification in the promoter environment of NSC proliferation- and differentiation-related gene loci (Wang Y. et al., 2018). In the absence of METTL14, the transcription-suppressing histone mark H3K27me3 is upregulated on genes involved in cell proliferation, whereas differentiation-related genes show an increase in the transcription activation histone mark H3K27ac when METTL14 is deficient (Wang Y. et al., 2018). Lack of METTL3 in the developing brain can also cause the aforementioned histone alterations, at least in terms of H3K27me3 enhancement, which can cause transcription repression (Chen J. et al., 2019). This is possible because in the absence of METTL3, which leads to a reduction in m6A levels, the polycomb repressor complex becomes hyperactive due to derepression of its core methlytransferase factor Ezh2 (Chen J. et al., 2019).
RBM15, a core component of the m6A writer complex (Figure 1A), is a potential regulator of cortical neurogenesis due to its distinctive expression in the cortical germinative zone and cortical plate of the developing mouse cortex (Xie et al., 2019). Knockdown of RBM15 in neurons in vitro promoted endogenous expression of the chromatin remodeling factor BAF155 (Xie et al., 2019), which is a known key regulator of cortical development (Nguyen et al., 2016, 2018; Narayanan et al., 2018). This profound effect can be linked to a significant reduction in cellular levels of m6A due to the inactivation of RBM15 (Knuckles et al., 2018). However, overexpression of RBM15 in vivo was found to promote delamination of radial glial cells in the cortical ventricular zone by suppressing the expression BAF155 and, hence, BAF155-dependent gene expression program supportive for cortical development (Xie et al., 2019). The role of RBM15 in cortical neurogenesis further highlights the contribution of m6A methyltransferase in brain development.
N6-Methyladenosine Erasers Regulate Neurogenesis in the Brain
Erasers of the m6A mark (FTO and ALKBH5) can also exert a regulatory effect on the process of neurogenesis given their prominent expression in neurons (Li L. et al., 2017; Yoon et al., 2017; Spychala and Ruther, 2019; Du et al., 2020). Whereas FTO displays the highest expression level late in brain neurogenesis (Li L. et al., 2017; Yoon et al., 2017), ALKBH5 expression decreases in the course of brain development (Du et al., 2020). This may have implications for their roles in the spatiotemporal regulation of neurogenesis during brain development. Indeed, it was reported that FTO deficiency in the adult mouse brain induces signal transducer and activator of transcription (STAT)3 pathway activation via its modulators platelet-derived growth factor receptor (PDGFR)α and suppressor of cytokine signaling (SOCS)5 in an m6A-dependent manner (Cao et al., 2020). As a result, a transient increase in the proliferation and differentiation of adult NSCs was observed in the FTO mutant brain, with implications for adult neurogenesis inhibition in the long term (Cao et al., 2020). It was also observed that FTO deletion in adult mouse brain impairs brain-derived neurotrophic factor (BDNF) and mitogen-activated protein kinase (MAPK) signaling pathways, leading to a reduction in adult NSC proliferation and neurogenesis in the hippocampal formation (Li L. et al., 2017; Spychala and Ruther, 2019). Although these studies report diverging effects of FTO loss on adult NSC proliferation, they both show a resultant effect of adult neurogenesis reduction. We think that, while being mindful of the low level of FTO expression in the early developing cortex, conducting an investigation on how FTO regulates corticogenesis in the course of development may lend clarity to how it mechanistically impacts cortical neurogenesis.
Notable N6-Methyladenosine Readers in Cortical Neurogenesis
Protein factors that act as readers of the m6A mark have also been shown to have a profound effect on neurogenesis in the brain. For instance, the m6A reader YTHDF2 has been reported to be indispensable for corticogenesis in mouse. Conditional knockout of YTHDF2 in the mouse neocortical neuroepithelium resulted in a reduction in the proliferation and differentiation of the Ythdf2–/– neuroprogenitor cells (Li M. et al., 2018). This phenotype may have mechanistic underpinnings, including abnormal upregulation of genes that inhibit the JAK–STAT signaling pathway, due to increased stability of such gene transcripts in the absence of YTHDF2 (Li M. et al., 2018). Yet, it seems that the induction of neural fate in pluripotent stem cells requires downregulation of YTHDF2, leading to the stability and expression of neural gene transcripts (Heck et al., 2020). We are of the opinion that the functional consequence of the m6A reading by YTHDF2 may be contextually variable along the cortical development axis such that reduced dosage may support neural cell fate specification, whereas its increased activity/expression is necessary for later cortical neurodevelopment.
Another m6A reader, FMRP, was identified to be critical for neural progenitor cell proliferation. Mice lacking Fmr1 displayed prolonged cell cycle progression. As a result, proliferation of neural progenitors extended into postnatal stages of brain development (Edens et al., 2019). Of note, it was observed that nuclear export of m6A-modified neurogenic mRNAs readable by FMRP is defective, leading to retention of such neurodifferentiation gene transcripts in the nucleus of the Fmr1-deficient neural progenitor cells (Edens et al., 2019). Lastly, the m6A reader protein Imp (IGF2BP) was identified as a key regulator of NSC proliferation rate through the stabilization of Myc mRNA in Drosophila brain neuroblasts (Samuels et al., 2020).
Together, the m6A machinery has been identified to play critical roles in brain morphogenesis by regulating the proliferation of neural progenitor cells and the production of neurons. As such, hypomethylation due to METTL3 or METTL14 deficiency and aberrant m6A reading or erasure in the embryonic or adult brain can precipitate phenotypes, including defective transcriptional prepatterning, abnormal neuroprogenitor pool, impaired neurogenesis, and cortical hypoplasia (Yoon et al., 2017), which can engender deficits in brain structure and function.
N6-Methyladenosine Signaling Is Essential for Gliogenesis in the Brain
The process of generating glial cells constitutes gliogenesis. Brain neuroglia include astrocytes and oligodendrocytes, which are derived from the neuroepithelium. During cortical development, a switch from neurogenesis to gliogenesis coincides with a decrease in m6A modification of proneural genes (Donega et al., 2018). Although m6A enrichment in glial cells is less than that observed in neurons (Chang et al., 2017), a few studies have uncovered the importance of the m6A methylome in brain gliogenesis, at least for astrocyte production (astrogenesis) and oligodendrocyte generation (oligodendrogenesis).
Regulation of Glia Production in the Brain by an N6-Methyladenosine Writer-Related Factor
It was observed that loss of METTL14-mediated m6A writing in the mouse cortex leads to hypomyelination that can be linked to a reduction in the number of (mature) oligodendrocytes (Xu et al., 2020). The loss of oligodendrocytes caused by the absence of METTL14 in the brain likely did not emanate from abnormal specification or proliferation of oligodendrocyte precursor cells (OPCs) (Xu et al., 2020). Notably, the transcriptome of OPCs and oligodendrocytes is altered following METTL14 deletion, with possible impact on gene expression programs critical for oligodendrocyte lineage progression (Xu et al., 2020). Lack of METTL14 has also been reported to disrupt astrogenesis. Indeed, s100β-expressing astrocytic progenitors were found to be reduced in the METTL14 knockout mouse cortex at postnatal stage 5 (Yoon et al., 2017). It would be interesting to investigate whether other m6A writer-related factors, including METTL3, have roles to play in cortical gliogenesis.
The N6-Methyladenosine Eraser FTO Regulates Glia Production in the Brain
m6A-mediated RNA methylation dynamics under the guild of FTO is known to influence oligodendrogenesis via modulation of the half-life of Olig2 mRNA (Wu et al., 2019). Olig2 is a central factor indispensable for oligodendrocyte lineage progression (Liu et al., 2007). Specifically, FTO was reported to regulate the degradation of Olig2 transcripts via removal of m6A tags installed on the Olig2 mRNA. The stability of Olig2 transcripts in OPCs deficient in FTO was thus seen to increase. In effect, the white matter in FTO mutant mouse brain was characterized by hypomyelination (Wu et al., 2019).
Involvement of N6-Methyladenosine Readers in Glia Production in the Brain
The m6A reader PRRC2A is known to be essential for oligodendrogenesis. It prominently regulates the specification, proliferation, and differentiation of oligodendroglia and the ability of oligodendrocytes to carry out myelination in the brain (Wu et al., 2019). More specifically, abolishing PRRC2A function in cortical NSCs or precisely in oligodendroglial lineage caused significant loss of OPCs (PDGFRα+ cells), Sox10+ cells, and mature oligodendrocytes (CC1+Oilg2+), which culminated in hypomyelination in the PRRC2A mutant brain (Wu et al., 2019). Interestingly, deletion of PRRC2A also affects astrogenesis, although slightly. Deficiency of PRRC2A in mouse brain caused a reduction in the proliferative capacity of astrocytes, leading to a reduced number of astrocytes in the mutant mouse brain (Wu et al., 2019). The additional role of PRRC2A in regulating the production of astrocytes in the brain during development may hinge on its interaction with YTHDF2, another m6A-binding protein, such that lack of either m6A reader augments the expression of the other to influence gliogenesis (Wu et al., 2019).
The competitive relationship between PRRC2A and YTHDF2 makes it complex to explain or reconcile the observation that glial fibrillary acidic protein (GFAP) expression, which can indicate astrocytic cells, was found to be dramatically reduced in neurospheres derived from the E14.5 Ythdf2–/– forebrain NSC. Such GFAP+ Ythdf2–/– cells also displayed abnormally branched processes (Li M. et al., 2018). Thus, further investigation is required to elucidate the role of YTHDF2 in brain gliogenesis and how the function of PRRC2A features in the regulatory pathway.
N6-Methyladenosine Effectors Regulate the Formation of Neural Processes and Synapses
The developing and adult brain is characterized by the outgrowth of dendrites and axons of neurons known to form neural connections called synapses. Interestingly, synapses are enriched with m6A, which modulates dendrite formation (dendritogenesis), axonogenesis, and synaptic growth (synaptogenesis) and activity (reviewed in Li et al., 2019; Dermentzaki and Lotti, 2020). m6A-based transcriptome profiling of the mouse brain (cortex and cerebellum) showed enrichment of m6A modification linked to dendrite and dendritic spine, axon and axon guidance, and synaptogenesis and synaptic transmission (Chang et al., 2017).
Distinctive localization of the YTHDFs, FTO, and METTL14 in dendrites of hippocampal neurons in culture and cortical neurons suggests the involvement of these m6A-regulatory factors in the development of neural dendrites. Indeed, Ythdf1 and Ythdf3 knockdown in such cultured neurons resulted in abnormal dendritic spine (Merkurjev et al., 2018). Axons are also enriched with FTO, which can be translated locally. As such, FTO ablation in axons resulted in upregulation of m6A levels, leading to a reduction in Gap-43 mRNA translation in axons of cultured dorsal root ganglion neurons (Yu et al., 2018). Yet, GAP-43 is a key factor involved in axon growth in neural tissues (Skene et al., 1986). In effect, the neurons lacking FTO displayed axon elongation repression (Yu et al., 2018). The m6A reader YTHDF1 was also reported to influence axon formation by binding and promoting the translation of the axon guidance receptor Robo3.1, which directs spinal commissural axons in crossing the midline, in an m6A modification-dependent manner (Zhuang et al., 2019). Together, these observations may have implications for perturbed axonogenesis in the brain lacking optimal m6A modification due to ablation of FTO or YTHDF1. At least in the case of the m6A-regulatory protein PRRC2A, it was found that axons that form the corpus callosum, a brain midline structure, are hypomyelinated and appeared hypoplastic in the PRRC2A-deleted mouse brain (Wu et al., 2019).
Given the enrichment of m6A marks and related proteins in neural processes, it is not surprising that synapses are endowed with m6A-modified mRNAs, especially postsynaptic transcripts in the mouse brain (Chang et al., 2017). The high localization of m6A-modified mRNAs in synapses reflects the possible impact of the m6A epitranscriptome on the structure, maturation, and function of synapses (Chang et al., 2017; Merkurjev et al., 2018; Yu et al., 2018; Zhuang et al., 2019). As a result, selective ablation of YTHDF1 and YTHDF3 in the cultured hippocampal neurons caused excitatory synaptic transmission suppression (Merkurjev et al., 2018; Shi et al., 2018). In addition, synapses formed by neurons lacking YTHDF2 appeared abnormal (Li M. et al., 2018), and synaptic transmission-related transcripts are hypermethylated in dopaminergic neurons with defective synaptic plasticity implication in the FTO-deficient mouse midbrain (Hess et al., 2013). Another indication of synapse malformation and synaptic plasticity impairment due to m6A dysregulation was observed in METTL14-deleted striatal neurons, in which METTL14 abrogation resulted in aberrant neuronal excitability (Koranda et al., 2018). Given that Nito, the Drosophila version of RBM15, also regulates synaptic growth through regulation of axonogenesis (Gu et al., 2017), it would be interesting to investigate whether indeed RBM15 is involved in synaptogenesis in the mammalian brain.
Cognition and Behavior Are Modulated by N6-Methyladenosine Signaling
The brain’s ability to process and store information and form or control behavior patterns has been shown to be greatly regulated by posttranscriptional modification of mRNA involved in brain development (reviewed in Jung and Goldman, 2018; Leighton et al., 2018; Noack and Calegari, 2018). Prominently emerging among these new (epitranscriptomic) levels of brain function regulation is m6A modification of mRNA in the brain. Various studies in mouse models have revealed the involvement of the m6A machinery-related factors in cognition and behavior (reviewed in Nainar et al., 2016; Chokkalla et al., 2020). The role of m6A in the regulation of learning and behavior may be partly explained by the previously discussed role of m6A in synaptogenesis and synaptic transmission (Weng et al., 2018).
N6-Methyladenosine Writers Involved in Memory and Behavior
In a recent study by Zhang F. et al. (2018), it was found that the enrichment of METTL3 in the mouse hippocampus is supportive for memory consolidation via the promotion of neuronal early-response gene translation. Therefore, mice lacking METTL3 in the hippocampus displayed impaired long-term potentiation with attendant reduced ability to consolidate memory. Interestingly, long-term memory consolidation is demonstrably augmented following METTL3 overexpression in the dorsal hippocampus of the wild-type mouse brain (Zhang Z. Y. et al., 2018). The m6A writer function of METTL14 is reported to be important for learning and behavior mediated by the striatum. Without affecting the number or morphology of striatal neurons, loss of METTL14 in striatopallidal and striatonigral neurons caused alterations in the transcriptome, eliciting increased neuronal excitability and spike frequency adaptation reduction, which possibly culminated in impairment of striatum-dependent behavior patterns (Koranda et al., 2018).
The N6-Methyladenosine Eraser FTO Regulates Learning and Behavior
Accumulation of m6A in the brain can affect its learning capacity and behavior. By regulating adult neurogenesis in the mouse hippocampus, FTO has been identified to play a pivotal role in learning (Li L. et al., 2017). Hypermethylation in the mouse brain or hippocampus caused by FTO functional loss was observed to call forth learning disabilities in mice, including increased fear memory consolidation (Widagdo et al., 2016; Walters et al., 2017). Additional evidence indicating the role of FTO in learning and behavior includes a study in which mice deficient in FTO were reported to exhibit behaviors consistent with depression and anxiety (Sun et al., 2019). Moreover, available data show that memory processing and verbal fluency may be affected in individuals with FTO ablation in the brain (Ho et al., 2010; Benedict et al., 2011).
Readers of N6-Methyladenosine Modulate Learning and Memory
Cognitive deficits have been implicitly linked to lack of function of the m6A reader PRRC2A, whose absence caused hypomyelination, leading to the cognitive anomalies in the mouse brain (Wu et al., 2019). Evidence indicating a more direct importance of an m6A reader in learning and memory was obtained when YTHDF1 was deleted in the adult mouse brain. It was found that neuronal stimuli can evoke translation of gene transcripts readable by YTHDF1 to facilitate learning and memory (Shi et al., 2018). Hence, silencing of YTHDF1 in the mouse hippocampus resulted in defective long-term potentiation and impaired synaptic transmission in the hippocampus, which did not allow normal learning and memory processing, and the defects were rescuable by YTHDF1 re-expression in the YTHDF1 mutant brain (Shi et al., 2018).
Stress Response Is Regulated by Factors of the N6-Methyladenosine Machinery
The brain plays a central role in stress response. In responding to stress, a host of gene expression programs is activated in the brain, leading to the secretion of several neuropeptides (de Kloet et al., 2005). Vulnerability to stressful stimuli and the response mechanism can have implications for neuropsychiatric anomalies under abnormal regulatory conditions. Thus, the transcriptomic stress response system is particularly crucial in maintaining homeostasis following exposure to stress.
Epigenetic mechanisms are known to play central roles in stress response (McEwen et al., 2015), and the epitranscriptome is an emerging gene expression regulation domain for stress modulation (Harvey et al., 2017). A putative role for m6A in the regulation of stress response is evidenced by the presence of glucocorticoid response elements upstream the transcription start site of genes that encode for enzymes involved in m6A modification (Engel et al., 2018). Additionally, nuclear localization of YTHDF2 precipitated by heat stress results in dynamic methylation of the 5′ UTR of newly synthesized mRNAs (Zhou et al., 2015). By limiting FTO, YTHDF2 is able to preserve methylation in the 5′ UTR of heat stress-induced mRNAs (Zhou et al., 2015).
In chick, upregulation of FTO in the brain (hypothalamus) may be a mechanism to afford thermoregulation in heat stress conditions (Kisliouk et al., 2020). However, following acute restraint stress, the mouse prefrontal cortex and amygdala displayed m6A hypomethylation and hypermethylation, respectively (Engel et al., 2018). Fear-induced stress can cause downregulation of FTO, leading to elevation of m6A in the prefrontal cortex and hippocampus of the mouse (Walters et al., 2017). Mice lacking METTL3 or FTO are unable to cope with stress (Engel et al., 2018). A general effect that may be caused by stress-induced alteration in m6A modification is the suppression of mRNAs involved in synaptic plasticity and brain morphogenesis (Engel et al., 2018). Together, the above observations indicate a putative role for m6A modulation in the human brain during stressful insults.
Neurological Disorders Attributable to Defective N6-Methyladenosine Modification in the Brain
Emerging evidence shows that a number of syndromic and non-syndromic neurological disturbances can be linked to m6A methylome dysregulation in the brain (Engel and Chen, 2018). This is not surprising, given the previously discussed extensive role of m6A in brain neurodevelopment (Figure 3). The m6A ubiquity in the brain implies that neural perturbations due to m6A dysregulation are likely to be complex and multifactorial in terms of downstream causatives. Neurologic problems so far identified to be caused by genetic variants of m6A modification factors can be broadly characterized as neurodevelopmental, neurodegenerative, or neuropsychiatric. Specifically, these include Parkinson’s disease (PD), Alzheimer’s disease (AD), autism, Smith–Magenis syndrome, schizophrenia, and depression (Table 2). The following subsections discuss the role of m6A and associated factors in neurological disorders of the brain.
TABLE 2.
Brain disorders associated with m6A dysregulation.
| Neurological disorders | Experimental system | m6A factor(s) implicated | References |
| Neurodevelopmental disorders | |||
| Microcephaly | GWAS; KO mice | Fto deletion; METTL5 frameshift | Richard et al. (2019) |
| Fragile X Syndrome | GWAS | SNP in FMRP | Verkerk et al. (1991); Dictenberg et al. (2008) |
| Cerebellar ataxia | KO mice; KO Drosophila | Deletion of Ythdc1, Mettl3, Alkbh5 | Fernandez-Funez et al. (2000); Ma et al. (2018); Wang C. X. et al. (2018) |
| Smith–Magenis syndrome | Genetic analysis in mouse | Alkbh5 deletion | Ricard et al. (2010) |
| Intellectual disability | GWAS | METTL5 frameshift | Richard et al. (2019) |
| Autism spectrum disorder | GWAS | Mutations in FMR1 | Reddy (2005); Edupuganti et al. (2017); Kaufmann et al. (2017) |
| Neurodegenerative disorders | |||
| Parkinson’s disease | 6-OHDA treatment of PC12 cells and rats; KO mice | Fto deletion or inhibition | Hess et al. (2013); Chen X. C. et al. (2019) |
| Alzheimer’s disease | GWAS | SNP in FTO | Ho et al. (2010); Keller et al. (2011); Reitz et al. (2012) |
| Amyotrophic lateral sclerosis | GWAS | SNP in FTO; SNP in HNRNP (A2B1 and A1); SNP in RBM15 | Kim et al. (2013); Cooper-Knock et al. (2017); Mitropoulos et al. (2017) |
| Cerebellar ataxia | KO mice; KO Drosophila | Deletion of Ythdc1, Mettl3, Alkbh5 | Fernandez-Funez et al. (2000); Ma et al. (2018); Wang C. X. et al. (2018) |
| Multiple sclerosis | GWAS | SNP in METTL1 | Mo et al. (2019) |
| Neuropsychiatric disorders | |||
| Major depressive disorder | GWAS | SNP in ALKBH5; SNP in FTO | Samaan et al. (2013); Milaneschi et al. (2014); Du et al. (2015) |
| Schizophrenia | GWAS | SNP in ZC3H13 | Oldmeadow et al. (2014) |
| Attention-deficit/hyperactivity disorder | GWAS | SNP in FTO | Velders et al. (2012); Choudhry et al. (2013) |
GWAS, genome-wide association studies; 6-OHDA, 6-hydroxydopamine; SNP, single-nucleotide polymorphism; KO, knockout; m6A, N6-methyladenosine.
Fragile X Syndrome
It has been identified that Fragile X syndrome (FXS) is the most common cause of inherited intellectual disorders and usually co-occurs with autism spectrum disorder (ASD). Patients present with features such as poor language development, abnormal behavior, and seizures, which are mainly clinical manifestations of neuronal excitation–inhibition imbalance (Hagerman et al., 2017; Kaufmann et al., 2017). Silencing of the FMR1 gene, which leads to lack of FMRP expression, is the cause of FXS (Brown et al., 2001). The role of FMRP in multiple gene expression programs partly accounts for the syndromic nature of FXS (Hagerman et al., 2017). Synaptic abnormalities or loss of neuroplasticity caused by FMRP loss-of-function and perhaps YTHFC2 deficiency is a critical underlying mechanism that contributes to the etiology of FXS and associated ASD (reviewed in Liu et al., 2016; Bagni and Zukin, 2019).
Parkinson’s Disease
Parkinson’s disease is a complex progressive neurodegenerative disorder mainly associated with death of dopamine-producing neurons in the midbrain (substantia nigra pars compacta) and aggregation of Lewy bodies in various brain regions. The main symptoms of PD include tremor and bradykinesia. Until now, the cause of PD is unknown, as many genetic and environmental risks are involved, making definitive diagnosis and treatment challenging (Kalia and Lang, 2015; Hayes, 2019).
Interestingly, m6A methylation deregulation caused by FTO abrogation, in the midbrain or in dopaminergic neurons, has been implicated in PD pathogenesis via impairment of neuronal activity and behavior response dependent on dopamine receptor types 2 and 3 (Hess et al., 2013). mRNAs involved in dopaminergic signaling are hypermethylated in the FTO-deficient mouse midbrain and striatum, leading to their decreased translation (Hess et al., 2013). It was found in another study that m6A may play a role in loss of dopaminergic neurons, which characterizes PD (Chen X. C. et al., 2019). The study reported that PC12 cells treated with 6-hydroxydopamine (6-OHDA) and the striatum of rat brain with 6-OHDA-induced PD display m6A modification downregulation, which is capable of inducing N-methyl-D-aspartate (NMDA) receptor 1 expression, alongside elevated oxidative stress and influx of Ca2+, culminating in cell death of dopaminergic neurons. Notably, FTO inhibition, and perhaps inhibition of ALKBH5, can attenuate 6-OHDA-induced PC12 cells apoptosis (Chen X. C. et al., 2019).
Alzheimer’s Disease
The commonest cause of dementia worldwide is AD. It is mainly characterized by progressive (age-dependent) neurodegeneration in brain regions (especially in the temporal and frontal lobes), with key clinical features, including memory loss, behavior abnormalities, and eventual cognitive decline (reviewed in Weller and Budson, 2018; Soria Lopez et al., 2019). Errors in RNA metabolism can have implications for AD. As discussed further, studies in human populations and in mouse models have shown that specific dysregulations in m6A mRNA methylation contribute to AD pathogenesis.
Typically, m6A levels in various brain regions increase with aging, and this disposition was shown to likely have relevance for AD development (Shafik et al., 2021). Interestingly, while METTL3 is downregulated in AD brain (hippocampus), it was observed to have accumulated in the postmortem AD brain at levels comparable to the insoluble Tau protein therein (Huang H. et al., 2020). Immunohistochemistry of the entorhinal cortex of patients with AD showed selective deficiency in the expression of another m6A factor hnRNP-A/B, which probably underscores the alteration in alternative splicing in the AD brain (Berson et al., 2012). Moreover, FTO mis-expression is implicated in the development of AD. Carriers of the FTO variant rs9939609 were reported to display systematic deficits in brain volume consistent with brain atrophy in the elderly (Ho et al., 2010). Indeed, a population-based study found an association between the FTO variant rs9939609 and increased risk of AD (Keller et al., 2011). Reitz et al. (2012) reported an increased risk caused by some polymorphisms (rs11075997, rs11075996, rs17219084) in the FTO gene in AD cases among some investigated Caribbean Hispanics and Caucasians (Reitz et al., 2012). Reduced verbal fluency in obese and overweight elderly men, with unaffected general cognitive function, was attributed to bearing of the FTO A allele. Thus, the (dys)functional effect of FTO A allele mainly manifests in the frontal lobe of the brain to constitute AD (Benedict et al., 2011). These observations indicate perturbation of m6A signaling as a notable underlying factor in the pathophysiology of AD in humans.
In vitro and in vivo experimentations using mouse models have yielded results that further support the involvement of m6A mRNA methylation in AD. In one study, it was observed that knockdown of hnRNP A/B impaired alternative splicing in cultured neurons, which resulted in loss of dendrites, and caused memory impairment in mice that can be ascribed to aberrance in the cortical connectome (Berson et al., 2012). The level of hnRNP A/B increases with cholinergic excitation, whereas loss of cholinergic signaling was found to induce AD-like reduction in hnRNP levels in the cortex (Berson et al., 2012).
The AD brain of the APP/PS1 transgenic mouse has elevated levels of m6A in the hippocampus and cortex, which may be due to the increased expression of METTL3 and concurrent downregulation of FTO expression (Han et al., 2020). However, the expression of FTO was identified to be increased in the brain of the triple transgenic AD mouse (Li H. et al., 2018). This gives an impression of the complex nature of the mechanism through which FTO or other m6A-associated factors may drive the development of AD. In the case of FTO, a proposed mechanism is that it may promote the phosphorylation of Tau protein by encouraging a methylation scheme leading to stabilization of tuberous sclerosis complex 1 (TSC1) mRNA, which activates the kinase activity of the mammalian target of rapamycin (mTOR) (Li H. et al., 2018).
Interestingly, cognition in a mouse model of AD was observed to improve when FTO was conditionally deleted in neurons in the mouse brain with AD (Li H. et al., 2018). This makes FTO a prospective therapeutic candidate worth further investigation for its potential in slowing down the progression of AD or in remedying related symptoms.
Amyotrophic Lateral Sclerosis
Amyotrophic lateral sclerosis (ALS) is a debilitating neurodegenerative disorder hallmarked by loss of motor neurons leading to skeletal muscle dysfunction and other clinical features, including psychological disorders and respiratory distress (Rowland and Shneider, 2001). It is believed to be idiopathic, with a greater percentage (∼90%) of cases being sporadic, while 5%−10% of cases are familial or inheritable (Kiernan et al., 2011). Studies have revealed the prominent role played by pathogenic mutation of factors associated with the RNA methylation machinery (Kim et al., 2013; Cooper-Knock et al., 2017; Mitropoulos et al., 2017).
By means of whole-genome sequencing, it became evident that m6A may be involved in the pathogenesis of ALS through FTO function alteration (Mitropoulos et al., 2017). Variants of FTO gene were thus associated with sporadic cases of ALS, which appears to be a founder effect among Greeks (Mitropoulos et al., 2017). In another key study, mutation in the prion-like domain of the m6A reader HNRNP (A2B1 and A1) was implicated in the pathogenesis of a familial ALS case (Kim et al., 2013). The work of Cooper-Knock et al. (2017) supports the involvement of RNA-binding protein mutations in ALS. Deleterious variants of RBM15 gene or its paralog RMB15B were found to contribute to the pathogenesis of ALS (Cooper-Knock et al., 2017).
Major Depressive Disorder
Major depressive disorder (MDD) is a common neuropsychiatric condition that is considered a biobehavioral syndrome with clinical characteristics including depressed mood, cognitive dysfunction, neurovegetative disturbance, and diminished interests. Females are known to be more affected by MDD than males. Multiple factors are known to cause MDD. Notable underlying causatives include genetic and environmental factors leading to alteration in the volume of the hippocampus and aberrant brain circuitry (Fava and Kendler, 2000; Flint and Kendler, 2014; Otte et al., 2016).
The m6A RNA methylome plays a role in the development of MDD (Engel et al., 2018). Genetic variants of FTO have been implicated in MDD, although heterogeneity in the associated phenotype is noteworthy (Milaneschi et al., 2014). In particular, it was found in a genome-wide association study that the FTO rs9939609 A variant is associated with a reduced risk of MDD (Samaan et al., 2013). A single-nucleotide polymorphism (rs12936694) in ALKBH5 was also found to likely be the cause of MDD among the Chinese Han population in an association study (Du et al., 2015). Interestingly, by blocking the translocation of ALKBH5 into the nucleus, it was possible to attenuate depression-like behavior in the mouse due to attendant hypermethylation and subsequent degradation of fatty acid amide hydrolase mRNA in astrocytes (Huang R. R. et al., 2020).
Therapeutic Prospects of CRISPR–Cas13-Mediated RNA Methylation Regulation in N6-Methyladenosine-Related Neurological Disease Treatment
While it seems intuitive that a simple strategy of traditional knockdown or overexpression of dysfunctional m6A factors in the epitranscriptome can correct pathologic alterations in the RNA methylation program, heterogeneity of the m6A methylome and, in some cases, the possible functional duplication or duality of the m6A writers, erasers, and readers, possess a challenge for the applicability of such solutions. To circumvent the aforementioned constraints, a system or tool capable of targeting defective m6A sites with high specificity should be considered. Such targeted approach to reversing disease-causing m6A modification can have therapeutic application if perfected.
The discovery of the Cas13 family of proteins, which are able to target endogenous RNA, has opened up avenues to deliver specific effectors at single sites on gene transcripts (Abudayyeh et al., 2016). By associating clustered regularly interspaced short palindromic repeats (CRISPR) with a catalytically inactive form of Cas13 protein (dCas13), but having preserved RNA binding ability, (m)RNA can be targeted at specific nucleic acid loci with such designed programmable CRISPR–dCas13 system (Figure 4; Wang et al., 2019; Burmistrz et al., 2020). Here, we discuss various salient in vitro applications of the CRISPR–dCas13 system to achieve m6A editing (Table 3).
FIGURE 4.
Schema showing application of clustered regularly interspaced short palindromic repeats (CRISPR)–dCas13 for N6-methyladenosine (m6A) editing. The m6A editing system is made by fusing deactivated Cas13b (dCas13b) with a guide RNA (gRNA) that can specifically target abnormally methylated messenger RNA (mRNA) (pathogenic mRNA) and coupling of CRISPR–dCas13 to an m6A factor to effect desired changes in m6A modification. Depending on the effector used, it is possible to induce m6A deposition, removal, or recognition (binding/reading), leading to the induction of degradation, translation enhancement, or increased stability of the target mRNA.
TABLE 3.
Applications based on CRISPR–dCas13b system for targeted manipulation of m6A modification and fate of m6A-tagged mRNA.
| System | Effect | Targeted mRNA | Reference |
| Targeting m6A installation | |||
| dCas13b-METTL3 dCas13b-METTL14 | Induces effective m6A incorporation in endogenous transcripts with increased specificity; regulates m6A-dependent mechanism for controlling transcript abundance; induces alternative splicing | Actb, Gapdh, Foxm1, Sox2, Brd8, Znf638, | Wilson et al. (2020) |
| dCas13b-CIBN CRY2PHR-METTL3-METTL14 | Effect photoactivatable RNA m6A level upregulation | TPT1, ACTB, TUG1 | Zhao et al. (2020) |
| Targeting m6A removal | |||
| dCas13b-FTO | Demethylates m6A of targeted mRNAs to enhance their stability | MALAT1, ASB2, HBV, HXB2 | Mo et al. (2020) |
| dCas13b-CIBN CRY2PHR-FTO | Effect photoactivatable RNA m6A level reduction | MALAT1, Alp, Bglap, RunX2, and Sp7, Pth1r | Zhao et al. (2020) |
| dCas13b-ALKBH5 | Demethylates m6A of targeted mRNAs to enhance their stability | CYB5A, CTNNB1, EGFR, MYC | Li et al. (2020) |
| Modulating m6A reading | |||
| dCas13b-YTHDF2 | Induces RNA degradation | KRAS, PPIB | Rauch et al. (2018) |
| dCas13b-YTHDF1 | Enhances translation with minimal mRNA destabilization effect | Firefly luciferase | Rauch et al. (2018) |
CRISPR, clustered regularly interspaced short palindromic repeats; m6A, N6-methyladenosine.
Restoring Abnormal Loss of N6-Methyladenosine
Gene transcripts that have lost m6A because of malfunction of the methyltransferases (METTL3 and/or METTL14) in the m6A methylation complex can be repaired using the CRISPR–dCas13 system. This is achievable by fusing dCas13, localized in the nucleus or cytoplasm, with a methyltransferase domain-truncated METTL3 or a modified METTL3:METTL14 complex, respectively. The resultant CRISPR–dCas13 constructs were able to install m6A marks, in a site-specific manner, on hypomethylated mRNAs or mRNAs with amendable m6A levels, including Sox2, Foxm1, and Znf638 in human cells (Wilson et al., 2020). Light-mediated m6A editing has also been put forward as another appealing CRISPR–dCas13 system for engineering the m6A methylome and worth close examination for applicability of the principle in therapeutics. The photoactivatable m6A editing CRISPR–dCas13b tool, which has a coupled component made of the methyltransferase domains of METTL3 and METTL14, was effectively employed in adding m6A to gene transcripts (TPT1, ACTB, TUG1) in human cells (Zhao et al., 2020).
Correcting N6-Methyladenosine Removal Incompetence
In the context of m6A erasure (demethylation), the m6A demethylase FTO and ALKBH5 can be incorporated into the CRISPR–dCas13 system to effect targeted removal of m6A on hypermethylated mRNAs or induce hypomethylation as corrective measures. By utilizing a CRISPR–dCas13b–FTO construct, Mo et al. (2020) were able to make mRNAs more stable via site-directed demethylation (Table 3) (Mo et al., 2020). Similarly, by applying a photoactivatable FTO-coupled CRISPR–dCas13b strategy, m6A marks were effectively and in a targeted manner removed on endogenous gene transcripts (Zhao et al., 2020). Methylated mRNAs that are preferentially demethylated by ALKBH5 can also be targeted by a CRISPR–dCas13b–ALKBH5 construct to remodel their m6A milieu, as applied in reducing the m6A levels associated with transcripts like CYB5A, CTNNB1, EGFR, and MYC, leading to their increased stability and translation (Li et al., 2020).
Rescuing Defective N6-Methyladenosine Reading
It is also possible to specifically target m6A readers to mRNAs of interest using the CRISPR–dCas13b system. It also implies that m6A binding protein dysfunctionality due to mutation can be rectified with a CRISPR–dCas13b construct fused to an engineered functional version of the relevant defective m6A reader. For example, YTHDF1 and/or YTHDF2, the two well-characterized m6A readers, can be fused to CRISPR–dCas13b and guided to specific mRNAs for m6A reading and subsequent fate alteration. Both CRISPR–dCas13b–YTHDF1 and CRISPR–dCas13b–YTHDF2 constructs were able to effect the native functions of YTHDF1 and YTHDF2, leading to translation enhancement and mRNA degradation in cells, respectively (Rauch et al., 2018).
Targeted N6-Methyladenosine Editing in the Diseased Brain as a Promising Treatment Strategy
Based on the intriguing outcomes and specificity of m6A editing application in vitro (Table 3 and Figure 4), we hereby propose the CRISPR–dCas13 system as a highly efficient tool for precise targeting and repair of aberrant m6A-modified mRNAs implicated in the pathophysiology of pertinent neurological disorders. Such a tool can be potentially useful in treating neurological disorders known to have pathologic m6A mRNA methylation, demethylation, or reading as the central underlying pathogenesis mechanism (Table 2). Employing high-resolution single-nucleotide binding techniques will be critical for identifying specific nucleotides bearing the abnormal m6A modification in the diseased brain. This will improve targeting, leading to the desired effect. An example of a strategy for improving the identification and targeting of nucleotides harboring disease-causing m6A marks is by adopting the enhanced crosslinking and immunoprecipitation (eCLIP) technique for robust factor-specific profiling of the m6A methylome in the pathologic brain (Van Nostrand et al., 2016). The in vivo experimental approach for investigating the potency of the CRISPR–dCas13 system for resolving neurological disorders caused by m6A dysregulation would include modeling the disorder in experimental animals and treating them with the rescuing CRISPR–dCas13 construct(s) that would have a target effect in the brain and on the implicated pathogenic mRNA (Figure 4). While the idea of investigating the application of the CRISPR–dCas13 system for rectifying aberrant m6A mRNA methylation (Figure 4) implied in neurological disorders sounds interesting, the approach may be fraught with challenges, especially in preventing off-target effects and in rescuing phenotypes of complex syndromic neurological disorders (e.g., ASD, AD, schizophrenia, MDD). Further investigations that can reveal convergent downstream effectors underlying the pathophysiology of polygenic neurological disorders caused by defective m6A signaling can help streamline an m6A editing-mediated therapeutic strategy.
Conclusion
Methylation of mRNA has emerged as a posttranscriptional regulation of gene expression that modulates protein synthesis in cells. Studies have shown that the m6A mRNA methylation machinery, composed of writers, erasers, and readers (Figure 1A), critically and extensively regulates RNA metabolism (trafficking, stability, processing, and translation efficiency) in cells to impact major biological processes. The brain is a hub of m6A modification, and the enrichment of m6A in the brain is reflective of the essential role it plays in optimal brain morphogenesis and functionality. Hence, the m6A interactome is known to regulate several neurodevelopmental processes in the brain, including neurogenesis, gliogenesis, synapse formation, and neuronal activity. Many of the m6A factors appear to have multiple functional effects during cortical development and in orchestrating several aspects of brain physiology (Table 1). This may make it challenging to effectively disentangle the rather multifactorial downstream causatives or complex phenotypic effects elicited by the dysregulation of the m6A machinery in the brain. As a typical example, whereas the FTO rs9939609 A variant is a risk factor for brain atrophy in old age (Ho et al., 2010) and AD development (Keller et al., 2011), it appears to be neuroprotective against MDD (Samaan et al., 2013). It also implies that m6A signaling is worth considering as a pivotal pathway that can cause novel syndromic neurological disturbances. Indeed, some inherited or acquired defects in the m6A RNA methylome are known causes of syndromes such as ASD, Smith–Magenis syndrome, and FXS. It also goes to reason that genetic variants of factors that make the m6A machinery pose a risk for certain (novel) non-syndromic neurological anomalies of the brain.
The phenomenal neurodevelopmental role played by m6A mRNA methylation and implication for neurological perturbations provoke considerable attention to the emerging involvement of the m6A methylome in normal brain structure and function maintenance. Going forward, more robust and advanced probing techniques are required to finely dissect the mechanistic basis of m6A-mediated neurodevelopment and its involvement in the pathophysiology of pertinent neurological disorders of the brain. Such sophisticated investigations may uncover therapeutic cues that can potentially fend off the neurological disorders caused by defective RNA methylation in the brain or alleviate the associated symptoms. For now, the application of the CRISPR–dCas13 system to edit m6A to restore normality of mRNA state and fate in, say, brain disease conditions is one of the promising approaches for treating abnormal m6A signaling-related neurological disorders.
Author Contributions
GS, YX, HN, and TT all contributed to writing and editing the manuscript. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
Funding. This work was supported by TU432/1, TU432/3, TU432/6 DFG grants and Schram-Stiftung to TT. We also acknowledge support by the Open Access Publication Funds of the Ruhr-University Bochum.
References
- Abdelmohsen K., Gorospe M. (2010). Posttranscriptional regulation of cancer traits by HuR. Wiley Interdiscipl. Rev. RNA 1 214–229. 10.1002/wrna.4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abudayyeh O. O., Gootenberg J. S., Konermann S., Joung J., Slaymaker I. M., Cox D. B., et al. (2016). C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353:aaf5573. 10.1126/science.aaf5573 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Agarwala S. D., Blitzblau H. G., Hochwagen A., Fink G. R. (2012). RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet. 8:e1002732. 10.1371/journal.pgen.1002732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alarcón C. R., Goodarzi H., Lee H., Liu X., Tavazoie S., Tavazoie S. F. (2015). HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell 162 1299–1308. 10.1016/j.cell.2015.08.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anders M., Chelysheva I., Goebel I., Trenkner T., Zhou J., Mao Y., et al. (2018). Dynamic m 6 A methylation facilitates mRNA triaging to stress granules. Life Sci. Alliance 1:e201800113. 10.26508/lsa.201800113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ascano M., Jr., Mukherjee N., Bandaru P., Miller J. B., Nusbaum J. D., Corcoran D. L., et al. (2012). FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 492 382–386. 10.1038/nature11737 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagni C., Zukin R. S. (2019). A synaptic perspective of fragile X syndrome and autism spectrum disorders. Neuron 101 1070–1088. 10.1016/j.neuron.2019.02.041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbieri I., Tzelepis K., Pandolfini L., Shi J., Millán-Zambrano G., Robson S. C., et al. (2017). Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552 126–131. 10.1038/nature24678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batista P. J., Molinie B., Wang J., Qu K., Zhang J., Li L., et al. (2014). m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15 707–719. 10.1016/j.stem.2014.09.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell J. L., Wächter K., Mühleck B., Pazaitis N., Köhn M., Lederer M., et al. (2013). Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs): post-transcriptional drivers of cancer progression? Cell. Mol. Life Sci. 70 2657–2675. 10.1007/s00018-012-1186-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benedict C., Jacobsson J. A., Rönnemaa E., Sällman-Almén M., Brooks S., Schultes B., et al. (2011). The fat mass and obesity gene is linked to reduced verbal fluency in overweight and obese elderly men. Neurobiol. Aging 32:1159.e1–5. 10.1016/j.neurobiolaging.2011.02.006 [DOI] [PubMed] [Google Scholar]
- Berson A., Barbash S., Shaltiel G., Goll Y., Hanin G., Greenberg D. S., et al. (2012). Cholinergic-associated loss of hnRNP-A/B in Alzheimer’s disease impairs cortical splicing and cognitive function in mice. EMBO Mol. Med. 4 730–742. 10.1002/emmm.201100995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boccaletto P., Machnicka M. A., Purta E., Piatkowski P., Baginski B., Wirecki T. K., et al. (2018). MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46 D303–D307. 10.1093/nar/gkx1030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bokar J. A., Rath-Shambaugh M. E., Ludwiczak R., Narayan P., Rottman F. (1994). Characterization and partial purification of mRNA N6-adenosine methyltransferase from HeLa cell nuclei. Internal mRNA methylation requires a multisubunit complex. J. Biol. Chem. 269 17697–17704. 10.1016/s0021-9258(17)32497-3 [DOI] [PubMed] [Google Scholar]
- Bokar J. A., Shambaugh M. E., Polayes D., Matera A. G., Rottman F. M. (1997). Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3 1233–1247. [PMC free article] [PubMed] [Google Scholar]
- Boles N. C., Temple S. (2017). Epimetronomics: m6A marks the tempo of corticogenesis. Neuron 96 718–720. 10.1016/j.neuron.2017.11.002 [DOI] [PubMed] [Google Scholar]
- Brown V., Jin P., Ceman S., Darnell J. C., O’Donnell W. T., Tenenbaum S. A., et al. (2001). Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107 477–487. 10.1016/s0092-8674(01)00568-2 [DOI] [PubMed] [Google Scholar]
- Bujnicki J. M., Feder M., Radlinska M., Blumenthal R. M. (2002). Structure prediction and phylogenetic analysis of a functionally diverse family of proteins homologous to the MT-A70 subunit of the human mRNA:m6A methyltransferase. J. Mol. Evol. 55 431–444. 10.1007/s00239-002-2339-8 [DOI] [PubMed] [Google Scholar]
- Burmistrz M., Krakowski K., Krawczyk-Balska A. (2020). RNA-targeting CRISPR-Cas systems and their applications. Int. J. Mol. Sci. 21:1122. 10.3390/ijms21031122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao Y., Zhuang Y., Chen J., Xu W., Shou Y., Huang X., et al. (2020). Dynamic effects of Fto in regulating the proliferation and differentiation of adult neural stem cells of mice. Hum. Mol. Genet. 29 727–735. 10.1093/hmg/ddz274 [DOI] [PubMed] [Google Scholar]
- Chang M., Lv H., Zhang W., Ma C., He X., Zhao S., et al. (2017). Region-specific RNA m(6)A methylation represents a new layer of control in the gene regulatory network in the mouse brain. Open Biol. 7:170166. 10.1098/rsob.170166 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J., Zhang Y. C., Huang C., Shen H., Sun B., Cheng X., et al. (2019). m(6)A regulates neurogenesis and neuronal development by modulating histone methyltransferase Ezh2. Genomics Proteom. Bioinform. 17 154–168. 10.1016/j.gpb.2018.12.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X. C., Yu C. Y., Guo M. J., Zheng X. T., Ali S., Huang H., et al. (2019). Down-regulation of m6A mRNA methylation is involved in dopaminergic neuronal death. Acs Chem. Neurosci. 10 2355–2363. 10.1021/acschemneuro.8b00657 [DOI] [PubMed] [Google Scholar]
- Choe J., Lin S., Zhang W., Liu Q., Wang L., Ramirez-Moya J., et al. (2018). mRNA circularization by METTL3-eIF3h enhances translation and promotes oncogenesis. Nature 561 556–560. 10.1038/s41586-018-0538-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chokkalla A. K., Mehta S. L., Vemuganti R. (2020). Epitranscriptomic regulation by m(6)A RNA methylation in brain development and diseases. J. Cereb. Blood Flow Metab. 40 2331–2349. 10.1177/0271678X20960033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choudhry Z., Sengupta S. M., Grizenko N., Thakur G. A., Fortier M. E., Schmitz N., et al. (2013). Association between obesity-related gene FTO and ADHD. Obesity 21 E738–E744. 10.1002/oby.20444 [DOI] [PubMed] [Google Scholar]
- Cooper-Knock J., Robins H., Niedermoser I., Wyles M., Heath P. R., Higginbottom A., et al. (2017). Targeted genetic screen in amyotrophic lateral sclerosis reveals novel genetic variants with synergistic effect on clinical phenotype. Front. Mol. Neurosci. 10:370. 10.3389/fnmol.2017.00370 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darnell R. B., Ke S., Darnell J. E., Jr. (2018). Pre-mRNA processing includes N(6) methylation of adenosine residues that are retained in mRNA exons and the fallacy of “RNA epigenetics”. RNA 24 262–267. 10.1261/rna.065219.117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Kloet E. R., Joëls M., Holsboer F. (2005). Stress and the brain: from adaptation to disease. Nat. Rev. Neurosci. 6 463–475. 10.1038/nrn1683 [DOI] [PubMed] [Google Scholar]
- Dermentzaki G., Lotti F. (2020). New insights on the role of N (6)-methyladenosine RNA methylation in the physiology and pathology of the nervous system. Front. Mol. Biosci. 7:555372. 10.3389/fmolb.2020.555372 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Desrosiers R., Friderici K., Rottman F. (1974). Identification of methylated nucleosides in messenger RNA from novikoff hepatoma cells. Proc. Natl. Acad. Sci. U.S.A. 71 3971–3975. 10.1073/pnas.71.10.3971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dictenberg J. B., Swanger S. A., Antar L. N., Singer R. H., Bassell G. J. (2008). A direct role for FMRP in activity-dependent dendritic mRNA transport links filopodial-spine morphogenesis to fragile X syndrome. Dev. Cell 14 926–939. 10.1016/j.devcel.2008.04.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dominissini D., Moshitch-Moshkovitz S., Schwartz S., Salmon-Divon M., Ungar L., Osenberg S., et al. (2012). Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485 201–206. 10.1038/nature11112 [DOI] [PubMed] [Google Scholar]
- Donega V., Marcy G., Lo Giudice Q., Zweifel S., Angonin D., Fiorelli R., et al. (2018). Transcriptional dysregulation in postnatal glutamatergic progenitors contributes to closure of the cortical neurogenic period. Cell Rep. 22 2567–2574. 10.1016/j.celrep.2018.02.030 [DOI] [PubMed] [Google Scholar]
- Du H., Zhao Y., He J., Zhang Y., Xi H., Liu M., et al. (2016). YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex. Nat. Commun. 7:12626. 10.1038/ncomms12626 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du T. F., Li G. X., Yang J. L., Ma K. L. (2020). RNA demethylase Alkbh5 is widely expressed in neurons and decreased during brain development. Brain Res. Bull. 163 150–159. 10.1016/j.brainresbull.2020.07.018 [DOI] [PubMed] [Google Scholar]
- Du T. F., Rao S. Q., Wu L., Ye N., Liu Z. Y., Hu H. L., et al. (2015). An association study of the m6A genes with major depressive disorder in Chinese Han population. J. Affect. Disord. 183 279–286. 10.1016/j.jad.2015.05.025 [DOI] [PubMed] [Google Scholar]
- Edens B. M., Vissers C., Su J., Arumugam S., Xu Z., Shi H., et al. (2019). FMRP modulates neural differentiation through m(6)A-dependent mRNA nuclear export. Cell Rep. 28 845.e5–854.e5. 10.1016/j.celrep.2019.06.072 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edupuganti R. R., Geiger S., Lindeboom R. G. H., Shi H., Hsu P. J., Lu Z., et al. (2017). N(6)-methyladenosine (m(6)A) recruits and repels proteins to regulate mRNA homeostasis. Nat. Struct. Mol. Biol. 24 870–878. 10.1038/nsmb.3462 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engel M., Chen A. (2018). The emerging role of mRNA methylation in normal and pathological behavior. Genes Brain Behav. 17:e12428. 10.1111/gbb.12428 [DOI] [PubMed] [Google Scholar]
- Engel M., Eggert C., Kaplick P. M., Eder M., Roh S., Tietze L., et al. (2018). The role of m(6)A/m-RNA methylation in stress response regulation. Neuron 99 389.e9–403.e9. 10.1016/j.neuron.2018.07.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan X. C., Steitz J. A. (1998). Overexpression of HuR, a nuclear-cytoplasmic shuttling protein, increases the in vivo stability of ARE-containing mRNAs. EMBO J. 17 3448–3460. 10.1093/emboj/17.12.3448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fava M., Kendler K. S. (2000). Major depressive disorder. Neuron 28 335–341. 10.1016/s0896-6273(00)00112-4 [DOI] [PubMed] [Google Scholar]
- Feng Z., Li Q., Meng R., Yi B., Xu Q. (2018). METTL3 regulates alternative splicing of MyD88 upon the lipopolysaccharide-induced inflammatory response in human dental pulp cells. J. Cell Mol. Med. 22 2558–2568. 10.1111/jcmm.13491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernandez-Funez P., Nino-Rosales M. L., de Gouyon B., She W.-C., Luchak J. M., Martinez P., et al. (2000). Identification of genes that modify ataxin-1-induced neurodegeneration. Nature 408 101–106. 10.1038/35040584 [DOI] [PubMed] [Google Scholar]
- Flint J., Kendler K. S. (2014). The genetics of major depression. Neuron 81 484–503. 10.1016/j.neuron.2014.01.027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu Y., Dominissini D., Rechavi G., He C. (2014). Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15 293–306. 10.1038/nrg3724 [DOI] [PubMed] [Google Scholar]
- Fustin J. M., Doi M., Yamaguchi Y., Hida H., Nishimura S., Yoshida M., et al. (2013). RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155 793–806. 10.1016/j.cell.2013.10.026 [DOI] [PubMed] [Google Scholar]
- Gao X., Wan J., Liu B., Ma M., Shen B., Qian S.-B. (2015). Quantitative profiling of initiating ribosomes in vivo. Nat. Methods 12 147–153. 10.1038/nmeth.3208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geula S., Moshitch-Moshkovitz S., Dominissini D., Mansour A. A., Kol N., Salmon-Divon M., et al. (2015). m 6 A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347 1002–1006. 10.1126/science.1261417 [DOI] [PubMed] [Google Scholar]
- Gu T. T., Zhao T., Kohli U., Hewes R. S. (2017). The large and small SPEN family proteins stimulate axon outgrowth during neurosecretory cell remodeling in Drosophila. Dev. Biol. 431 226–238. 10.1016/j.ydbio.2017.09.013 [DOI] [PubMed] [Google Scholar]
- Hagerman R. J., Berry-Kravis E., Hazlett H. C., Bailey D. B., Jr., Moine H., Kooy R. F., et al. (2017). Fragile X syndrome. Nat. Rev. Dis. Primers 3:17065. 10.1038/nrdp.2017.65 [DOI] [PubMed] [Google Scholar]
- Han M., Liu Z., Xu Y., Liu X., Wang D., Li F., et al. (2020). Abnormality of m6A mRNA methylation is involved in alzheimer’s disease. Front. Neurosci. 14:98. 10.3389/fnins.2020.00098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harfmann A. M., Nayler O., Schwaiger F. W., Obermeier A., Stamm S. (1999). The interaction and colocalization of Sam68 with the splicing-associated factor YT521-B in nuclear dots is regulated by the Src family kinase p59(fyn). Mol. Biol. Cell 10 3909–3926. 10.1091/mbc.10.11.3909 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harvey R., Dezi V., Pizzinga M., Willis A. E. (2017). Post-transcriptional control of gene expression following stress: the role of RNA-binding proteins. Biochem. Soc. Trans. 45 1007–1014. 10.1042/bst20160364 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes M. T. (2019). Parkinson’s disease and Parkinsonism. Am. J. Med. 132 802–807. 10.1016/j.amjmed.2019.03.001 [DOI] [PubMed] [Google Scholar]
- Hazra D., Chapat C., Graille M. (2019). m6A mRNA destiny: chained to the rhYTHm by the YTH-containing proteins. Genes 10:49. 10.3390/genes10010049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heck A. M., Russo J., Wilusz J., Nishimura E. O., Wilusz C. J. (2020). YTHDF2 destabilizes m(6)A-modified neural-specific RNAs to restrain differentiation in induced pluripotent stem cells. RNA 26 739–755. 10.1261/rna.073502.119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hess M. E., Hess S., Meyer K. D., Verhagen L. A., Koch L., Brönneke H. S., et al. (2013). The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat. Neurosci. 16 1042–1048. 10.1038/nn.3449 [DOI] [PubMed] [Google Scholar]
- Hinman M. N., Lou H. (2008). Diverse molecular functions of Hu proteins. Cell. Mol. Life Sci. 65 3168–3181. 10.1007/s00018-008-8252-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho A. J., Stein J. L., Hua X., Lee S., Hibar D. P., Leow A. D., et al. (2010). A commonly carried allele of the obesity-related FTO gene is associated with reduced brain volume in the healthy elderly. Proc. Natl. Acad. Sci. U.S.A. 107 8404–8409. 10.1073/pnas.0910878107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horiuchi K., Kawamura T., Iwanari H., Ohashi R., Naito M., Kodama T., et al. (2013). Identification of Wilms’ tumor 1-associating protein complex and its role in alternative splicing and the cell cycle. J. Biol. Chem. 288 33292–33302. 10.1074/jbc.M113.500397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H., Camats-Perna J., Medeiros R., Anggono V., Widagdo J. (2020). Altered expression of the m6A methyltransferase METTL3 in Alzheimer’s disease. eNeuro 7:ENEURO.0125-20.2020. 10.1523/eneuro.0125-20.2020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H., Weng H., Sun W., Qin X., Shi H., Wu H., et al. (2018). Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20 285–295. 10.1038/s41556-018-0045-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H., Weng H., Zhou K., Wu T., Zhao B. S., Sun M., et al. (2019). Histone H3 trimethylation at lysine 36 guides m6A RNA modification co-transcriptionally. Nature 567 414–419. 10.1038/s41586-019-1016-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang J., Yin P. (2018). Structural insights into N(6)-methyladenosine (m(6)A) modification in the transcriptome. Genomics Proteom. Bioinform. 16 85–98. 10.1016/j.gpb.2018.03.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang R. R., Zhang Y., Bai Y., Han B., Ju M. Z., Chen B. L., et al. (2020). N-6-methyladenosine modification of fatty acid amide hydrolase messenger RNA in circular RNA STAG1-regulated astrocyte dysfunction and depressive-like behaviors. Biol. Psychiat. 88 392–404. 10.1016/j.biopsych.2020.02.018 [DOI] [PubMed] [Google Scholar]
- Huang T., Gao Q., Feng T., Zheng Y., Guo J., Zeng W. (2018). FTO knockout causes chromosome instability and G2/M arrest in mouse GC-1 cells. Front. Genet. 9:732. 10.3389/fgene.2018.00732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai Y., Matsuo N., Ogawa S., Tohyama M., Takagi T. (1998). Cloning of a gene, YT521, for a novel RNA splicing-related protein induced by hypoxia/reoxygenation. Brain Res. Mol. Brain Res. 53 33–40. 10.1016/s0169-328x(97)00262-3 [DOI] [PubMed] [Google Scholar]
- Iyer L. M., Zhang D., Aravind L. (2016). Adenine methylation in eukaryotes: apprehending the complex evolutionary history and functional potential of an epigenetic modification. BioEssays 38 27–40. 10.1002/bies.201500104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia G., Fu Y., Zhao X., Dai Q., Zheng G., Yang Y., et al. (2011). N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7 885–887. 10.1038/nchembio.687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia G., Yang C.-G., Yang S., Jian X., Yi C., Zhou Z., et al. (2008). Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Lett. 582 3313–3319. 10.1016/j.febslet.2008.08.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung Y., Goldman D. (2018). Role of RNA modifications in brain and behavior. Genes Brain Behav. 17:e12444. 10.1111/gbb.12444 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalia L. V., Lang A. E. (2015). Parkinson’s disease. Lancet 386 896–912. 10.1016/S0140-6736(14)61393-3 [DOI] [PubMed] [Google Scholar]
- Kang H.-J., Jeong S.-J., Kim K.-N., Baek I.-J., Chang M., Kang C.-M., et al. (2014). A novel protein, Pho92, has a conserved YTH domain and regulates phosphate metabolism by decreasing the mRNA stability of PHO4 in Saccharomyces cerevisiae. Biochem. J. 457 391–400. 10.1042/bj20130862 [DOI] [PubMed] [Google Scholar]
- Kaufmann W. E., Kidd S. A., Andrews H. F., Budimirovic D. B., Esler A., Haas-Givler B., et al. (2017). Autism spectrum disorder in fragile X syndrome: cooccurring conditions and current treatment. Pediatrics 139 S194–S206. 10.1542/peds.2016-1159F [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ke S., Alemu E. A., Mertens C., Gantman E. C., Fak J. J., Mele A., et al. (2015). A majority of m6A residues are in the last exons, allowing the potential for 3’ UTR regulation. Genes Dev. 29 2037–2053. 10.1101/gad.269415.115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keller L., Xu W., Wang H. X., Winblad B., Fratiglioni L., Graff C. (2011). The obesity related gene, FTO, interacts with APOE, and is associated with Alzheimer’s disease risk: a prospective cohort study. J. Alzheimer’s Dis. 23 461–469. 10.3233/jad-2010-101068 [DOI] [PubMed] [Google Scholar]
- Kiernan M. C., Vucic S., Cheah B. C., Turner M. R., Eisen A., Hardiman O., et al. (2011). Amyotrophic lateral sclerosis. Lancet 377 942–955. 10.1016/S0140-6736(10)61156-7 [DOI] [PubMed] [Google Scholar]
- Kim H. J., Kim N. C., Wang Y. D., Scarborough E. A., Moore J., Diaz Z., et al. (2013). Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495 467–473. 10.1038/nature11922 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kisliouk T., Rosenberg T., Ben-Nun O., Ruzal M., Meiri N. (2020). Early-Life m(6)A RNA demethylation by fat mass and obesity-associated protein (FTO) influences resilience or vulnerability to heat stress later in life. eNeuro 7 ENEURO.0549-19.2020. 10.1523/ENEURO.0549-19.2020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knuckles P., Lence T., Haussmann I. U., Jacob D., Kreim N., Carl S. H., et al. (2018). Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev. 32 415–429. 10.1101/gad.309146.117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kontur C., Jeong M., Cifuentes D., Giraldez A. J. (2020). Ythdf m(6)A readers function redundantly during Zebrafish development. Cell Rep. 33:108598. 10.1016/j.celrep.2020.108598 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koranda J. L., Dore L., Shi H. L., Patel M. J., Vaasjo L. O., Rao M. N., et al. (2018). Mettl14 is essential for epitranscriptomic regulation of striatal function and learning. Neuron 99 283–292. 10.1016/j.neuron.2018.06.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretschmer J., Rao H., Hackert P., Sloan K. E., Höbartner C., Bohnsack M. T. (2018). The m6A reader protein YTHDC2 interacts with the small ribosomal subunit and the 5’–3’ exoribonuclease XRN1. RNA 24 1339–1350. 10.1261/rna.064238.117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogan N. J., Cagney G., Yu H., Zhong G., Guo X., Ignatchenko A., et al. (2006). Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440 637–643. 10.1038/nature04670 [DOI] [PubMed] [Google Scholar]
- Lasman L., Krupalnik V., Viukov S., Mor N., Aguilera-Castrejon A., Schneir D., et al. (2020). Context-dependent functional compensation between Ythdf m(6)A reader proteins. Genes Dev. 34 1373–1391. 10.1101/gad.340695.120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leighton L. J., Ke K., Zajaczkowski E. L., Edmunds J., Spitale R. C., Bredy T. W. (2018). Experience-dependent neural plasticity, learning, and memory in the era of epitranscriptomics. Genes Brain Behav. 17:e12426. 10.1111/gbb.12426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lesbirel S., Viphakone N., Parker M., Parker J., Heath C., Sudbery I., et al. (2018). The m(6)A-methylase complex recruits TREX and regulates mRNA export. Sci. Rep. 8:13827. 10.1038/s41598-018-32310-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lesbirel S., Wilson S. A. (2019). The m(6)A-methylase complex and mRNA export. Biochim. Biophys. Acta Gene Regul. Mech. 1862 319–328. 10.1016/j.bbagrm.2018.09.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li A., Chen Y.-S., Ping X.-L., Yang X., Xiao W., Yang Y., et al. (2017). Cytoplasmic m6A reader YTHDF3 promotes mRNA translation. Cell Res. 27:444. 10.1038/cr.2017.10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li F., Zhao D., Wu J., Shi Y. (2014). Structure of the YTH domain of human YTHDF2 in complex with an m6A mononucleotide reveals an aromatic cage for m6A recognition. Cell Res. 24:1490. 10.1038/cr.2014.153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H., Ren Y., Mao K., Hua F., Yang Y., Wei N., et al. (2018). FTO is involved in Alzheimer’s disease by targeting TSC1-mTOR-Tau signaling. Biochem. Biophys. Res. Commun. 498 234–239. 10.1016/j.bbrc.2018.02.201 [DOI] [PubMed] [Google Scholar]
- Li J., Chen Z., Chen F., Xie G., Ling Y., Peng Y., et al. (2020). Targeted mRNA demethylation using an engineered dCas13b-ALKBH5 fusion protein. Nucleic Acids Res. 48 5684–5694. 10.1093/nar/gkaa269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J., Yang X., Qi Z., Sang Y., Liu Y., Xu B., et al. (2019). The role of mRNA m(6)A methylation in the nervous system. Cell Biosci. 9:66. 10.1186/s13578-019-0330-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L., Zang L., Zhang F., Chen J., Shen H., Shu L., et al. (2017). Fat mass and obesity-associated (FTO) protein regulates adult neurogenesis. Hum. Mol. Genet. 26 2398–2411. 10.1093/hmg/ddx128 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M., Zhao X., Wang W., Shi H., Pan Q., Lu Z., et al. (2018). Ythdf2-mediated m(6)A mRNA clearance modulates neural development in mice. Genome Biol. 19:69. 10.1186/s13059-018-1436-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin S., Choe J., Du P., Triboulet R., Gregory R. I. (2016). The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Mol. Cell 62 335–345. 10.1016/j.molcel.2016.03.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder B., Grozhik A. V., Olarerin-George A. O., Meydan C., Mason C. E., Jaffrey S. R. (2015). Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12 767–772. 10.1038/nmeth.3453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J., Yue Y., Han D., Wang X., Fu Y., Zhang L., et al. (2014). A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10 93–95. 10.1038/nchembio.1432 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu K., Ding Y., Ye W., Liu Y., Yang J., Liu J., et al. (2016). Structural and functional characterization of the proteins responsible for N(6)-methyladenosine modification and recognition. Curr. Protein Pept. Sci. 17 306–318. 10.2174/1389203716666150901113553 [DOI] [PubMed] [Google Scholar]
- Liu N., Dai Q., Zheng G., He C., Parisien M., Pan T. (2015). N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518 560–564. 10.1038/nature14234 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu N., Zhou K. I., Parisien M., Dai Q., Diatchenko L., Pan T. (2017). N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res. 45 6051–6063. 10.1093/nar/gkx141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Z., Hu X., Cai J., Liu B., Peng X., Wegner M., et al. (2007). Induction of oligodendrocyte differentiation by Olig2 and Sox10: evidence for reciprocal interactions and dosage-dependent mechanisms. Dev. Biol. 302 683–693. 10.1016/j.ydbio.2006.10.007 [DOI] [PubMed] [Google Scholar]
- Livneh I., Moshitch-Moshkovitz S., Amariglio N., Rechavi G., Dominissini D. (2020). The m(6)A epitranscriptome: transcriptome plasticity in brain development and function. Nat. Rev. Neurosci. 21 36–51. 10.1038/s41583-019-0244-z [DOI] [PubMed] [Google Scholar]
- Ma C., Chang M., Lv H., Zhang Z. W., Zhang W., He X., et al. (2018). RNA m(6)A methylation participates in regulation of postnatal development of the mouse cerebellum. Genome Biol. 19:68. 10.1186/s13059-018-1435-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masuda S., Das R., Cheng H., Hurt E., Dorman N., Reed R. (2005). Recruitment of the human TREX complex to mRNA during splicing. Genes Dev. 19 1512–1517. 10.1101/gad.1302205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mauer J., Luo X., Blanjoie A., Jiao X., Grozhik A. V., Patil D. P., et al. (2017). Reversible methylation of m(6)A(m) in the 5’ cap controls mRNA stability. Nature 541 371–375. 10.1038/nature21022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McEwen B. S., Bowles N. P., Gray J. D., Hill M. N., Hunter R. G., Karatsoreos I. N., et al. (2015). Mechanisms of stress in the brain. Nat. Neurosci. 18 1353–1363. 10.1038/nn.4086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merkurjev D., Hong W. T., Iida K., Oomoto I., Goldie B. J., Yamaguti H., et al. (2018). Synaptic N(6)-methyladenosine (m(6)A) epitranscriptome reveals functional partitioning of localized transcripts. Nat. Neurosci. 21 1004–1014. 10.1038/s41593-018-0173-6 [DOI] [PubMed] [Google Scholar]
- Meyer K. D., Jaffrey S. R. (2017). Rethinking m(6)A readers, writers, and erasers. Annu. Rev. Cell Dev. Biol. 33 319–342. 10.1146/annurev-cellbio-100616-060758 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer K. D., Patil D. P., Zhou J., Zinoviev A., Skabkin M. A., Elemento O., et al. (2015). 5’ UTR m(6)A promotes cap-independent translation. Cell 163 999–1010. 10.1016/j.cell.2015.10.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer K. D., Saletore Y., Zumbo P., Elemento O., Mason C. E., Jaffrey S. R. (2012). Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149 1635–1646. 10.1016/j.cell.2012.05.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milaneschi Y., Lamers F., Mbarek H., Hottenga J. J., Boomsma D. I., Penninx B. W. (2014). The effect of FTO rs9939609 on major depression differs across MDD subtypes. Mol. Psychiatry 19 960–962. 10.1038/mp.2014.4 [DOI] [PubMed] [Google Scholar]
- Mitropoulos K., Merkouri Papadima E., Xiromerisiou G., Balasopoulou A., Charalampidou K., Galani V., et al. (2017). Genomic variants in the FTO gene are associated with sporadic amyotrophic lateral sclerosis in Greek patients. Hum. Genomics 11:30. 10.1186/s40246-017-0126-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mo J., Chen Z., Qin S., Li S., Liu C., Zhang L., et al. (2020). TRADES: targeted RNA demethylation by SunTag system. Adv. Sci. 7:2001402. 10.1002/advs.202001402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mo X. B., Lei S. F., Qian Q. Y., Guo Y. F., Zhang Y. H., Zhang H. (2019). Integrative analysis revealed potential causal genetic and epigenetic factors for multiple sclerosis. J. Neurol. 266 2699–2709. 10.1007/s00415-019-09476-w [DOI] [PubMed] [Google Scholar]
- Nainar S., Marshall P. R., Tyler C. R., Spitale R. C., Bredy T. W. (2016). Evolving insights into RNA modifications and their functional diversity in the brain. Nat. Neurosci. 19 1292–1298. 10.1038/nn.4378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narayan P., Rottman F. (1988). An in vitro system for accurate methylation of internal adenosine residues in messenger RNA. Science 242 1159–1162. 10.1126/science.3187541 [DOI] [PubMed] [Google Scholar]
- Narayanan R., Pham L., Kerimoglu C., Watanabe T., Hernandez R. C., Sokpor G., et al. (2018). Chromatin remodeling BAF155 subunit regulates the genesis of basal progenitors in developing cortex. Iscience 4 109–126. 10.1016/j.isci.2018.05.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen H., Kerimoglu C., Pirouz M., Pham L., Kiszka K. A., Sokpor G., et al. (2018). Epigenetic regulation by BAF complexes limits neural stem cell proliferation by suppressing wnt signaling in late embryonic development. Stem Cell Rep. 10 1734–1750. 10.1016/j.stemcr.2018.04.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen H., Sokpor G., Pham L., Rosenbusch J., Stoykova A., Staiger J. F., et al. (2016). Epigenetic regulation by BAF (mSWI/SNF) chromatin remodeling complexes is indispensable for embryonic development. Cell Cycle 15 1317–1324. 10.1080/15384101.2016.1160984 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noack F., Calegari F. (2018). Epitranscriptomics: a new regulatory mechanism of brain development and function. Front. Neurosci. 12:85. 10.3389/fnins.2018.00085 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oldmeadow C., Mossman D., Evans T. J., Holliday E. G., Tooney P. A., Cairns M. J., et al. (2014). Combined analysis of exon splicing and genome wide polymorphism data predict schizophrenia risk loci. J. Psychiatr. Res. 52 44–49. 10.1016/j.jpsychires.2014.01.011 [DOI] [PubMed] [Google Scholar]
- Otte C., Gold S. M., Penninx B. W., Pariante C. M., Etkin A., Fava M., et al. (2016). Major depressive disorder. Nat. Rev. Dis. Primers 2:16065. 10.1038/nrdp.2016.65 [DOI] [PubMed] [Google Scholar]
- Park O. H., Ha H., Lee Y., Boo S. H., Kwon D. H., Song H. K., et al. (2019). Endoribonucleolytic cleavage of m6A-containing RNAs by RNase P/MRP complex. Mol. Cell 74 494.e8–507.e8. 10.1016/j.molcel.2019.02.034 [DOI] [PubMed] [Google Scholar]
- Patil D. P., Chen C. K., Pickering B. F., Chow A., Jackson C., Guttman M., et al. (2016). m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537 369–373. 10.1038/nature19342 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pendleton K. E., Chen B., Liu K., Hunter O. V., Xie Y., Tu B. P., et al. (2017). The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell 169 824.e14–835.e14. 10.1016/j.cell.2017.05.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng S. S., Chen C. Y., Xu N., Shyu A. B. (1998). RNA stabilization by the AU-rich element binding protein, HuR, an ELAV protein. EMBO J. 17 3461–3470. 10.1093/emboj/17.12.3461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perry R. P., Kelley D. E., Friderici K., Rottman F. (1975a). The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5’ terminus. Cell 4 387–394. 10.1016/0092-8674(75)90159-2 [DOI] [PubMed] [Google Scholar]
- Perry R. P., Kelley D. E., Friderici K. H., Rottman F. M. (1975b). Methylated constituents of heterogeneous nuclear RNA: presence in blocked 5’ terminal structures. Cell 6 13–19. 10.1016/0092-8674(75)90068-9 [DOI] [PubMed] [Google Scholar]
- Ping X. L., Sun B. F., Wang L., Xiao W., Yang X., Wang W. J., et al. (2014). Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24 177–189. 10.1038/cr.2014.3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rauch S., He C., Dickinson B. C. (2018). Targeted m(6)A reader proteins to study epitranscriptomic regulation of single RNAs. J. Am. Chem. Soc. 140 11974–11981. 10.1021/jacs.8b05012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy K. S. (2005). Cytogenetic abnormalities and fragile-X syndrome in autism spectrum disorder. BMC Med. Genet. 6:3. 10.1186/1471-2350-6-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitz C., Tosto G., Mayeux R., Luchsinger J. A. (2012). Genetic variants in the Fat and Obesity Associated (FTO) gene and risk of Alzheimer’s disease. PLoS One 7:e50354. 10.1371/journal.pone.0050354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ricard G., Molina J., Chrast J., Gu W., Gheldof N., Pradervand S., et al. (2010). Phenotypic consequences of copy number variation: insights from Smith-Magenis and Potocki-Lupski syndrome mouse models. PLoS Biol. 8:e1000543. 10.1371/journal.pbio.1000543 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richard E. M., Polla D. L., Assir M. Z., Contreras M., Shahzad M., Khan A. A., et al. (2019). Bi-allelic variants in METTL5 cause autosomal-recessive intellectual disability and microcephaly. Am. J. Hum. Genet. 105 869–878. 10.1016/j.ajhg.2019.09.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ries R. J., Zaccara S., Klein P., Olarerin-George A., Namkoong S., Pickering B. F., et al. (2019). m6A enhances the phase separation potential of mRNA. Nature 571 424–428. 10.1038/s41586-019-1374-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rockwell A. L., Hongay C. F. (2019). The m(6)A dynamics of profilin in neurogenesis. Front. Genet. 10:987. 10.3389/fgene.2019.00987 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roundtree I. A., Evans M. E., Pan T., He C. (2017a). Dynamic RNA modifications in gene expression regulation. Cell 169 1187–1200. 10.1016/j.cell.2017.05.045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roundtree I. A., Luo G.-Z., Zhang Z., Wang X., Zhou T., Cui Y., et al. (2017b). YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. eLife 6:e31311. 10.7554/eLife.31311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowland L. P., Shneider N. A. (2001). Amyotrophic lateral sclerosis. N. Engl. J. Med. 344 1688–1700. 10.1056/NEJM200105313442207 [DOI] [PubMed] [Google Scholar]
- Rùžièka K., Zhang M., Campilho A., Bodi Z., Kashif M., Saleh M., et al. (2017). Identification of factors required for m(6) A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. N. Phytol. 215 157–172. 10.1111/nph.14586 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samaan Z., Anand S. S., Zhang X., Desai D., Rivera M., Pare G., et al. (2013). The protective effect of the obesity-associated rs9939609 A variant in fat mass- and obesity-associated gene on depression. Mol. Psychiatry 18 1281–1286. 10.1038/mp.2012.160 [DOI] [PubMed] [Google Scholar]
- Samuels T. J., Jarvelin A. I., Ish-Horowicz D., Davis I. (2020). Imp/IGF2BP levels modulate individual neural stem cell growth and division through myc mRNA stability. eLife 9:e51529. 10.7554/eLife.51529 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz S., Mumbach M. R., Jovanovic M., Wang T., Maciag K., Bushkin G. G., et al. (2014). Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep. 8 284–296. 10.1016/j.celrep.2014.05.048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shafik A. M., Zhang F., Guo Z., Dai Q., Pajdzik K., Li Y., et al. (2021). N6-methyladenosine dynamics in neurodevelopment and aging, and its potential role in Alzheimer’s disease. Genome Biol. 22:17. 10.1186/s13059-020-02249-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah A., Rashid F., Awan H. M., Hu S., Wang X., Chen L. (2017). The DEAD-Box RNA helicase DDX3 interacts with m(6)A RNA demethylase ALKBH5. Stem Cell Int. 2017:8596135. 10.1155/2017/8596135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H., Wang X., Lu Z., Zhao B. S., Ma H., Hsu P. J., et al. (2017). YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 27:315. 10.1038/cr.2017.15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H., Zhang X., Weng Y. L., Lu Z., Liu Y., Lu Z., et al. (2018). m(6)A facilitates hippocampus-dependent learning and memory through YTHDF1. Nature 563 249–253. 10.1038/s41586-018-0666-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shima H., Matsumoto M., Ishigami Y., Ebina M., Muto A., Sato Y., et al. (2017). S-Adenosylmethionine synthesis is regulated by selective N6-adenosine methylation and mRNA degradation involving METTL16 and YTHDC1. Cell Rep. 21 3354–3363. 10.1016/j.celrep.2017.11.092 [DOI] [PubMed] [Google Scholar]
- Skene J. H. P., Jacobson R. D., Snipes G. J., Mcguire C. B., Norden J. J., Freeman J. A. (1986). A protein-induced during nerve growth (Gap-43) is a major component of growth-cone membranes. Science 233 783–786. 10.1126/science.3738509 [DOI] [PubMed] [Google Scholar]
- Śledź P., Jinek M. (2016). Structural insights into the molecular mechanism of the m(6)A writer complex. eLife 5:e18434. 10.7554/eLife.18434 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song H., Feng X., Zhang H., Luo Y., Huang J., Lin M., et al. (2019). METTL3 and ALKBH5 oppositely regulate m6A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy 15 1419–1437. 10.1080/15548627.2019.1586246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soria Lopez J. A., González H. M., Léger G. C. (2019). Alzheimer’s disease. Handbook Clin. Neurol. 167 231–255. 10.1016/b978-0-12-804766-8.00013-3 [DOI] [PubMed] [Google Scholar]
- Spychala A., Ruther U. (2019). FTO affects hippocampal function by regulation of BDNF processing. PLoS One 14:e0211937. 10.1371/journal.pone.0211937 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun L., Ma L., Zhang H., Cao Y., Wang C., Hou N., et al. (2019). Fto deficiency reduces anxiety- and depression-like behaviors in mice via alterations in gut microbiota. Theranostics 9 721–733. 10.7150/thno.31562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang C., Klukovich R., Peng H., Wang Z., Yu T., Zhang Y., et al. (2018). ALKBH5-dependent m6A demethylation controls splicing and stability of long 3’-UTR mRNAs in male germ cells. Proc. Natl. Acad. Sci. U.S.A. 115 E325–E333. 10.1073/pnas.1717794115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulmke P. A., Xie Y., Sokpor G., Pham L., Shomroni O., Berulava T., et al. (2021). Post-transcription regulation by the exosome complex is required for cell survival and forebrain development by repressing P53 signaling. Development 148:dev188276. 10.1242/dev.188276 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Nostrand E. L., Pratt G. A., Shishkin A. A., Gelboin-Burkhart C., Fang M. Y., Sundararaman B., et al. (2016). Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13 508–514. 10.1038/nmeth.3810 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velders F. P., De Wit J. E., Jansen P. W., Jaddoe V. W., Hofman A., Verhulst F. C., et al. (2012). FTO at rs9939609, food responsiveness, emotional control and symptoms of ADHD in preschool children. PLoS One 7:e49131. 10.1371/journal.pone.0049131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verkerk A. J., Pieretti M., Sutcliffe J. S., Fu Y. H., Kuhl D. P., Pizzuti A., et al. (1991). Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65 905–914. 10.1016/0092-8674(91)90397-h [DOI] [PubMed] [Google Scholar]
- Walters B. J., Mercaldo V., Gillon C. J., Yip M., Neve R. L., Boyce F. M., et al. (2017). The role of The RNA demethylase FTO (Fat Mass and Obesity-Associated) and mRNA methylation in hippocampal memory formation. Neuropsychopharmacology 42 1502–1510. 10.1038/npp.2017.31 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang C. X., Cui G. S., Liu X., Xu K., Wang M., Zhang X. X., et al. (2018). METTL3-mediated m6A modification is required for cerebellar development. PLoS Biol. 16:e2004880. 10.1371/journal.pbio.2004880 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Wang L., Zou X., Duan S., Li Z., Deng Z., et al. (2019). Advances in CRISPR-Cas systems for RNA targeting, tracking and editing. Biotechnol. Adv. 37 708–729. 10.1016/j.biotechadv.2019.03.016 [DOI] [PubMed] [Google Scholar]
- Wang P., Doxtader K. A., Nam Y. (2016). Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol. Cell 63 306–317. 10.1016/j.molcel.2016.05.041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X., Feng J., Xue Y., Guan Z., Zhang D., Liu Z., et al. (2016). Structural basis of N6-adenosine methylation by the METTL3–METTL14 complex. Nature 534:575. 10.1038/nature18298 [DOI] [PubMed] [Google Scholar]
- Wang X., Lu Z., Gomez A., Hon G. C., Yue Y., Han D., et al. (2014). N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505 117–120. 10.1038/nature12730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X., Zhao B. S., Roundtree I. A., Lu Z., Han D., Ma H., et al. (2015). N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161 1388–1399. 10.1016/j.cell.2015.05.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Li Y., Toth J. I., Petroski M. D., Zhang Z., Zhao J. C. (2014). N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat. Cell Biol. 16 191–198. 10.1038/ncb2902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Li Y., Yue M. H., Wang J., Kumar S., Wechsler-Reya R. J., et al. (2018). N-6-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nat. Neurosci. 21 195–206. 10.1038/s41593-017-0057-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Zhao J. C. (2016). Update: mechanisms underlying N(6)-methyladenosine modification of eukaryotic mRNA. Trends Genet. 32 763–773. 10.1016/j.tig.2016.09.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei H., Peng J., Lu J., Song T., Xie X., Yang Y., et al. (2019). Zfp217 mediates m6A mRNA methylation to orchestrate transcriptional and post-transcriptional regulation to promote adipogenic differentiation. Nucleic Acids Res. 47 6130–6144. 10.1093/nar/gkz312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei J., Liu F., Lu Z., Fei Q., Ai Y., He P. C., et al. (2018). Differential m6A, m6Am, and m1A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol. Cell. 71 973.e5–985.e5. 10.1016/j.molcel.2018.08.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weller J., Budson A. (2018). Current understanding of Alzheimer’s disease diagnosis and treatment. F1000Research 7:F1000 Faculty Rev–1161. 10.12688/f1000research.14506.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen J., Lv R., Ma H., Shen H., He C., Wang J., et al. (2018). Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Mol. Cell. 69 1028.e6–1038.e6. 10.1016/j.molcel.2018.02.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weng Y. L., Wang X., An R., Cassin J., Vissers C., Liu Y., et al. (2018). Epitranscriptomic m(6)A regulation of axon regeneration in the adult mammalian nervous system. Neuron 97 313.e6–325.e6. 10.1016/j.neuron.2017.12.036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Widagdo J., Anggono V. (2018). The m6A-epitranscriptomic signature in neurobiology: from neurodevelopment to brain plasticity. J. Neurochem. 147 137–152. 10.1111/jnc.14481 [DOI] [PubMed] [Google Scholar]
- Widagdo J., Zhao Q. Y., Kempen M. J., Tan M. C., Ratnu V. S., Wei W., et al. (2016). Experience-dependent accumulation of N6-methyladenosine in the prefrontal cortex is associated with memory processes in mice. J. Neurosci. 36 6771–6777. 10.1523/jneurosci.4053-15.2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson C., Chen P. J., Miao Z., Liu D. R. (2020). Programmable m(6)A modification of cellular RNAs with a Cas13-directed methyltransferase. Nat. Biotechnol. 38 1431–1440. 10.1038/s41587-020-0572-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu B., Su S., Patil D. P., Liu H., Gan J., Jaffrey S. R., et al. (2018). Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1. Nat. Commun. 9:420. 10.1038/s41467-017-02770-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu R., Li A., Sun B., Sun J.-G., Zhang J., Zhang T., et al. (2019). A novel m(6)A reader Prrc2a controls oligodendroglial specification and myelination. Cell Res. 29 23–41. 10.1038/s41422-018-0113-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao W., Adhikari S., Dahal U., Chen Y.-S., Hao Y.-J., Sun B.-F., et al. (2016). Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61 507–519. 10.1016/j.molcel.2016.01.012 [DOI] [PubMed] [Google Scholar]
- Xie Y. B., Castro-Hernandez R., Sokpor G., Pham L., Narayanan R., Rosenbusch J., et al. (2019). RBM15 modulates the function of chromatin remodeling factor BAF155 through RNA methylation in developing cortex. Mol. Neurobiol. 56 7305–7320. 10.1007/s12035-019-1595-1 [DOI] [PubMed] [Google Scholar]
- Xu C., Liu K., Ahmed H., Loppnau P., Schapira M., Min J. (2015). Structural basis for the discriminative recognition of N6-methyladenosine RNA by the human YT521-B homology domain family of proteins. J. Biol. Chem. 290 24902–24913. 10.1074/jbc.M115.680389 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu C., Wang X., Liu K., Roundtree I. A., Tempel W., Li Y., et al. (2014). Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat. Chem. Biol. 10:927. 10.1038/nchembio.1654 [DOI] [PubMed] [Google Scholar]
- Xu H., Dzhashiashvili Y., Shah A., Kunjamma R. B., Weng Y. L., Elbaz B., et al. (2020). m(6)A mRNA methylation is essential for oligodendrocyte maturation and CNS myelination. Neuron 105 293.e5–309.e5. 10.1016/j.neuron.2019.12.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang F., Jin H., Que B., Chao Y., Zhang H., Ying X., et al. (2019). Dynamic m6A mRNA methylation reveals the role of METTL3-m6A-CDCP1 signaling axis in chemical carcinogenesis. Oncogene 38 4755–4772. 10.1038/s41388-019-0755-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao B., Christian K. M., He C., Jin P., Ming G. L., Song H. (2016). Epigenetic mechanisms in neurogenesis. Nat. Rev. Neurosci. 17 537–549. 10.1038/nrn.2016.70 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoon K. J., Ringeling F. R., Vissers C., Jacob F., Pokrass M., Jimenez-Cyrus D., et al. (2017). Temporal control of mammalian cortical neurogenesis by m(6)A methylation. Cell 171 877.e17–889.e17. 10.1016/j.cell.2017.09.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoon K. J., Vissers C., Ming G. L., Song H. (2018). Epigenetics and epitranscriptomics in temporal patterning of cortical neural progenitor competence. J. Cell Biol. 217 1901–1914. 10.1083/jcb.201802117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu J., Chen M., Huang H., Zhu J., Song H., Zhu J., et al. (2018). Dynamic m6A modification regulates local translation of mRNA in axons. Nucleic Acids Res. 46 1412–1423. 10.1093/nar/gkx1182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yue Y., Liu J., Cui X., Cao J., Luo G., Zhang Z., et al. (2018). VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 4:10. 10.1038/s41421-018-0019-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaccara S., Jaffrey S. R. (2020). A unified model for the function of YTHDF proteins in regulating m(6)A-modified mRNA. Cell 181 1582.e18–1595.e18. 10.1016/j.cell.2020.05.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zarnack K., König J., Tajnik M., Martincorena I., Eustermann S., Stévant I., et al. (2013). Direct competition between hnRNP C and U2AF65 protects the transcriptome from the exonization of Alu elements. Cell 152 453–466. 10.1016/j.cell.2012.12.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang F., Kang Y., Wang M., Li Y., Xu T., Yang W., et al. (2018). Fragile X mental retardation protein modulates the stability of its m6A-marked messenger RNA targets. Hum. Mol. Genet. 27 3936–3950. 10.1093/hmg/ddy292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L., Du K., Wang J., Nie Y., Lee T., Sun T. (2020). Unique and specific m(6)A RNA methylation in mouse embryonic and postnatal cerebral cortices. Genes 11:1139. 10.3390/genes11101139 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X., Wei L.-H., Wang Y., Xiao Y., Liu J., Zhang W., et al. (2019). Structural insights into FTO’s catalytic mechanism for the demethylation of multiple RNA substrates. Proc. Natl. Acad. Sci. U.S.A. 116 2919–2924. 10.1073/pnas.1820574116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Wang X., Zhang X., Wang J., Ma Y., Zhang L., et al. (2019). RNA-binding protein YTHDF3 suppresses interferon-dependent antiviral responses by promoting FOXO3 translation. Proc. Natl. Acad. Sci. U.S.A. 116 976–981. 10.1073/pnas.1812536116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Z., Theler D., Kaminska K. H., Hiller M., de la Grange P., Pudimat R., et al. (2010). The YTH domain is a novel RNA binding domain. J. Biol. Chem. 285 14701–14710. 10.1074/jbc.M110.104711 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Z. Y., Wang M., Xie D. F., Huang Z. H., Zhang L. S., Yang Y., et al. (2018). METTL3-mediated N-6-methyladenosine mRNA modification enhances long-term memory consolidation. Cell Res. 28 1050–1061. 10.1038/s41422-018-0092-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao B. S., Roundtree I. A., He C. (2017). Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18 31–42. 10.1038/nrm.2016.132 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J., Li B., Ma J., Jin W., Ma X. (2020). Photoactivatable RNA N(6) -methyladenosine editing with CRISPR-Cas13. Small 16:e1907301. 10.1002/smll.201907301 [DOI] [PubMed] [Google Scholar]
- Zhao X., Yang Y., Sun B.-F., Shi Y., Yang X., Xiao W., et al. (2014). FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res. 24 1403–1419. 10.1038/cr.2014.151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng G., Dahl J. A., Niu Y., Fedorcsak P., Huang C.-M., Li C. J., et al. (2013). ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49 18–29. 10.1016/j.molcel.2012.10.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong S., Li H., Bodi Z., Button J., Vespa L., Herzog M., et al. (2008). MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell 20 1278–1288. 10.1105/tpc.108.058883 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou H., Wang B., Sun H., Xu X., Wang Y. (2018). Epigenetic regulations in neural stem cells and neurological diseases. Stem cells Int. 2018:6087143. 10.1155/2018/6087143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J., Wan J., Gao X., Zhang X., Jaffrey S. R., Qian S. B. (2015). Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature 526 591–594. 10.1038/nature15377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J., Wan J., Shu X. E., Mao Y., Liu X.-M., Yuan X., et al. (2018). N(6)-methyladenosine guides mRNA alternative translation during integrated stress response. Mol. Cell 69 636.e7–647.e7. 10.1016/j.molcel.2018.01.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu C., Yi C. (2014). Switching demethylation activities between AlkB family RNA/DNA demethylases through exchange of active-site residues. Angew. Chem. Int. Ed. 53 3659–3662. 10.1002/anie.201310050 [DOI] [PubMed] [Google Scholar]
- Zhuang M., Li X., Zhu J., Zhang J., Niu F., Liang F., et al. (2019). The m6A reader YTHDF1 regulates axon guidance through translational control of Robo3.1 expression. Nucleic Acids Res. 47 4765–4777. 10.1093/nar/gkz157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou S., Toh J. D. W., Wong K. H. Q., Gao Y.-G., Hong W., Woon E. C. Y. (2016). N(6)-Methyladenosine: a conformational marker that regulates the substrate specificity of human demethylases FTO and ALKBH5. Sci. Rep. 6:25677. 10.1038/srep25677 [DOI] [PMC free article] [PubMed] [Google Scholar]