Abstract
Snake venoms are complex chemical mixtures of biologically active proteins and non-protein components. Toxins have a wide range of targets and effects to include ion channels and membrane receptors, and platelet aggregation and platelet plug formation. Toxins target these effectors and effects at high affinity and selectivity. From a pharmacological perspective, snake venom compounds are a valuable resource for drug discovery and development. However, a major challenge to drug discovery using snake venoms is isolating and analyzing the bioactive proteins and peptides in these complex mixtures. Getting molecular information from complex mixtures such as snake venoms requires proteomic analyses, generally combined with transcriptomic analyses of venom glands. The present review summarizes current knowledge and highlights important recent advances in venomics with special emphasis on contemporary separation techniques and bioinformatics that have begun to elaborate the complexity of snake venoms. Several analytical techniques such as two-dimensional gel electrophoresis, RP-HPLC, size exclusion chromatography, ion exchange chromatography, MALDI-TOF-MS, and LC-ESI-QTOF-MS have been employed in this regard. The improvement of separation approaches such as multidimensional-HPLC, 2D-electrophoresis coupled to soft-ionization (MALDI and ESI) mass spectrometry has been critical to obtain an accurate picture of the startling complexity of venoms. In the case of bioinformatics, a variety of software tools such as PEAKS also has been used successfully. Such information gleaned from venomics is important to both predicting and resolving the biological activity of the active components of venoms, which in turn is key for the development of new drugs based on these venom components.
Keywords: Animal venoms, Venomic, Drug discovery, Separation methods, Mass spectrometry
1. Introduction
The study of bioactive proteins has long been a key focus of life and health sciences. In 1995, the expression “Proteome” was defined by Marc Wilkins as the entire set of proteins expressed by a genome of an organism, tissue or cell [1]. Consequently, expression “proteomics” is a large-scale comprehensive study of a specific proteome, including information about protein abundance, post-translational modification (PTM), protein–protein interaction, protein function and protein location [2]. In 1980, Celis and his team developed a protein separation method that utilized an exceptionally effective 2-D gel electrophoresis followed by Edman degradation sequencing to identify specific proteins. This along with the wider availability of protein sequence databases opened the door to broad use of proteomics. Since these pioneering efforts, the utility of a proteomic approach now has been demonstrated for many organisms and cell types [3]. Proteomic strategies empower researchers to characterize, compare and distinguish numerous proteins in an organism or multi-species environment. Proteomics is a rapidly developing field, which relies upon rapid development in analytical instrumentation and bioinformatics programming. The main areas in which this field is improving is in understanding protein abundance, half-life, subcellular localization, precise activity, state of modification, structure and interactions with other bioactive molecules. Consequently, proteomics is now widely used in pharmaceutical research allowing scientists and commercial companies to identify novel drugs faster and to test side effects more efficiently. It also has become widely used in clinical research, disease diagnosis, and identification of molecular biomarkers. Biologists use it to investigate basic cell and microbe functions and molecular organizations. It also has become an important tool for the study of animal venoms enabling detailed investigation of venom composition [4–6].
Snake venoms are highly modified saliva containing a complex mixture of biologically active proteins and non-proteins molecules that are most often used to incapacitate, immobilize, and digest prey [7]. Proteins account for about 90–95% of the dry weight of most venoms and play a primary role in the biological effects of venoms. Hence, snake venoms are often considered to be mini-libraries for developing novel drugs [8,9]. Recently, many pharmaceutical companies and experts in the toxinology field have directed their attentions to snake venom components as a natural product resource of materials for biotechnological application [10–13]. Snake venoms have been used in the pharmaceutical industry as a valuable source of therapeutic drugs to include Captopril, Aggrastat, and Eptifibatide [14–16].
Knowledge about the composition of many snake venoms recently has been increased by the progressively rapid improvement of analytical methods to include improvements in mass spectrometry, separation strategies, and bioinformatics [17]. Nonetheless, to manifest the full benefit of proteomics in venomics separation methods must continue to improve. This is particularly true for the initial steps in venom fractionation. The establishment of high quality venom proteomic profiles will (i) reveal the biological diversity of toxin compositional strategies used by snakes to fulfill [18]; (ii) likely will aid prediction of clinical manifestations of envenomation [19,20]; (iii) improve taxonomy [21]; (iv) reveal intriguing ontogenetic changes [22]; (v) provide a useful tool to investigate the possible differences in venom composition associated with distinct geographic origins within the distribution range of species [22]; and (vi) reveal novel, important biologically-active components of venom [23]. The present review summarizes current knowledge and highlights important recent advances in venomics with special emphasis on contemporary separation techniques and bioinformatics that have begun to elaborate the complexity of snake venoms.
2. Venomics approach
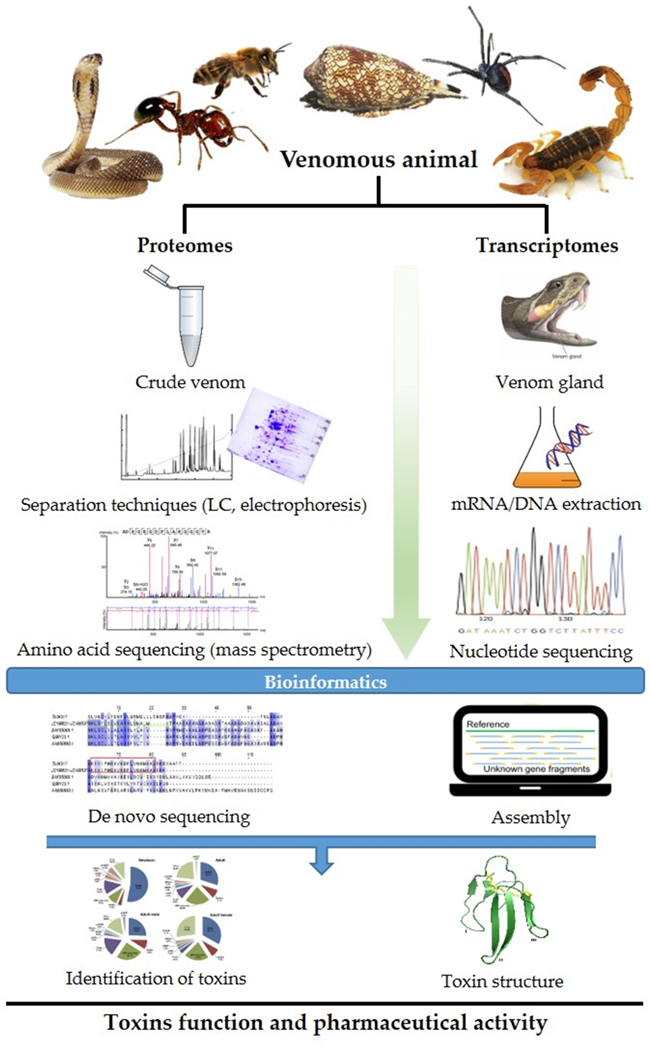
Venomous animals, in particular especially snakes and their venoms have attracted scientific interests mainly for the development of antibody (antivenom) against envenomation caused by snakebites [24–26]. Animal venoms are unique cocktails of often more than 100 different peptides and proteins, making these venoms an appreciable source of millions of peptides and proteins naturally tailored to act on a myriad of exogenous targets, including ion channels, membrane receptors and enzymes [11,27]. The search for new biologically active molecules aimed at discovering novel drugs is usually driven by the use of a specific target through a biological screening test. Proteomics has allowed characterizing and identifying a great number of animal venom bioactive components. Furthermore, ongoing improvements in sequencing have opened new horizons for the understanding of the venomous systems. The use of transcriptomics on venom glands is becoming common in proteomics for identifying compounds of potential interest in pharmaceutical research. The expression “Venomics” was coined to include techniques and methods used to understand venoms and the contents of venom glands [28]. The venomics approach presently encompasses transcriptomics, proteomics and peptidomics to explore the content of animal venoms and venom glands (Fig. 1) [29]. Venomics is an important tool for the study of the phylogenetic relationship between species [21]. Recent developments in animal venomics has made this field an important tool for identifying tomorrow’s drugs from animal venoms.
Fig. 1.
From venoms to drugs. A schematic representation of computational venomic approaches for discovery, identification and development of therapeutics from animal venoms [17].
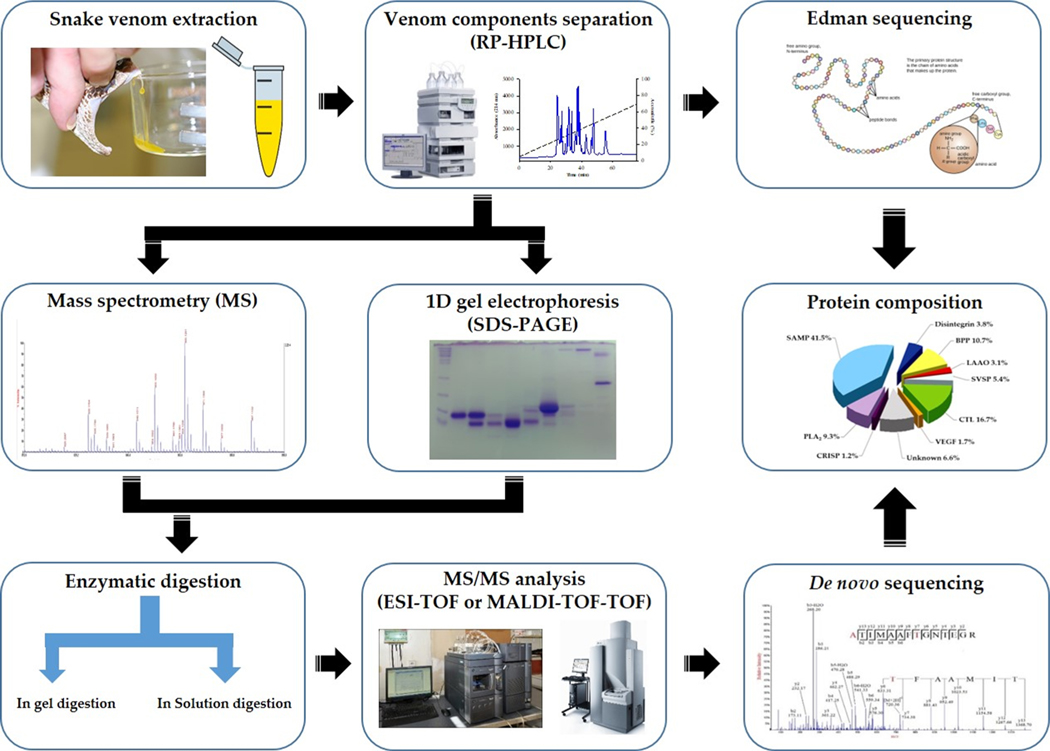
Numerous proteomic approaches have been applied to the investigation of snake venoms [30,31]. The standard snake venom proteomics approach begins with proteolytic digestion, with most commonly trypsin, of venom proteins and peptides that are isolated through single- or multi-dimensional methods, including electrophoretic and chromatographic techniques. The resulting digested peptides are identified using mass spectrometry techniques, such as MALDI or ESI, followed by fragmentation via collision-induced dissociation (CID) energy. Tandem mass spectrometry (MS/MS) data are sequenced using de novo sequencing software. The full sequence of the peptide is identified by database search [32,33]. For instance, Calvete and his colleagues established a snake venomics approach that begins with the fractionation of the snake crude venom by RP-HPLC followed by the initial characterization of each protein fraction by a combination of N-terminal sequencing, SDS-PAGE (or 2DE), and mass spectrometric determination of the molecular masses, and eventually the cysteine (–SH and S–S) content, of the isolated components (Fig. 2) [34–36]. Such investigation gives a good elaboration of the components of the venom and a semi-quantitative estimation of their relative abundance in the venom. Currently, snake venomics studies have been performed on over more than 200 snake species, offering access to an extremely rich source of pharmacologically active proteins and peptides [37].
Fig. 2.
Schematic representation of the ‘snake venomics’ analytical strategy developed by Calvete [34]. This strategy begins with the fractionation of the snake crude venom by RP-HPLC followed by the initial characterization of each protein fraction by a combination of N-terminal sequencing, SDS-PAGE (or 2DE), and mass spectrometric determination of the molecular masses, and eventually the cysteine (–SH and S–S) content, of the isolated components. Reproduced with permission from Ref. [134].
2.1. Separation techniques
Classical protein separation techniques have been successfully employed to isolate and characterize many bioactive components from animal venoms now for over 40 years. Several separation methods are frequently used in combination in proteomic analyses to improve signal-to-noise, proteome coverage, and to reduce interferences between peptides. Thus, a given separation method often allows important supplementary information regarding molecular weights and hydrophobicity and isoelectric point (pI) about each venom component. Such information improves identification of venom components in regard to splice variants and large post-translational modifications. Consequently, pre-fractionation and means to decrease the complexity of snake venoms is still valuable in most analyses. Many fractionation strategies for snake venom compounds are generally available. In snake venom proteomics, both analytical and bioinformatics have been used to describe protein structure and functions. Several analytical techniques such as two-dimensional gel electrophoresis (2-DE), RP-HPLC, size exclusion chromatography (SEC), ion exchange chromatography, MALDI-TOF-MS, and LC-ESI-QTOF-MS have been employed in this regard [38]. In the case of bioinformatics, a variety of software tools also has been used successfully [39].
2.1.1. Two-dimensional gel electrophoresis (2-DE)
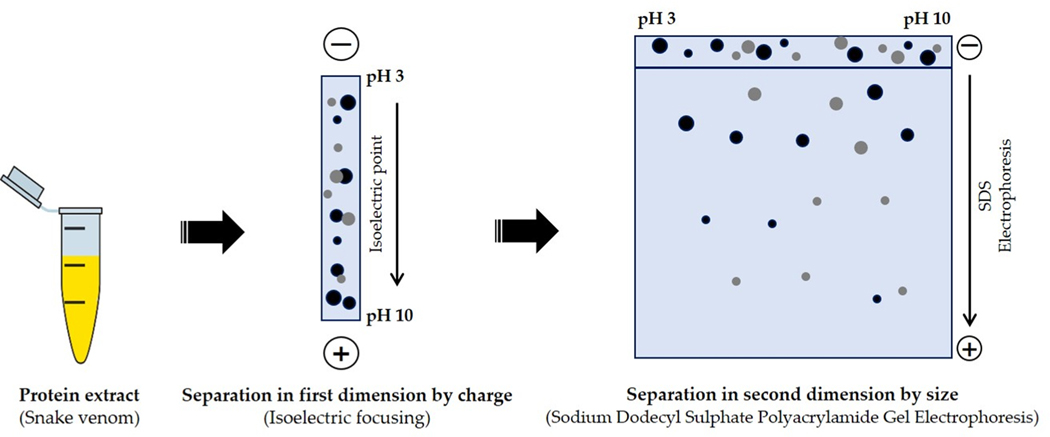
With its appearance thirty years ago, 2-DE has become the primary means for identifying proteins in complex biological solutions [40]. 2-DE separates proteins based on two independent parameters; isoelectric point (pI) values, via isoelectric focusing (IEF) in one dimension and molecular mass by SDS-PAGE in the other. With this approach samples in complex biological solutions can be resolved to a few thousand protein spots (Fig. 3) [41]. The 2-DE strategy has been progressively important methodology in the investigation of snake venom proteomes [42–44]. For instance, venoms of eight snakes to include Ophiophagus hannah, Naja kaouthia, Naja sumatrana, Bungarus fasciatus, Trimeresurus sumatranus, Tropidolaemus wagleri, Enhydrina schistosa and Calloselasma rhodostoma located in Malaysia have been separated based on 2-DE properties [45]. In addition, 2-DE has been used to identify individual compounds like glycoproteins in the venom of Naja kaouthia [46]. However, it is rapidly becoming evident that this approach is far from perfect due to limitations, including (1) limited solubility of hydrophobic proteins; (2) difficulty in focusing highly basic and acidic proteins; (3) poor sensitivity; (4) it being time consuming; and (5) the method is not amenable to automation [47,48]. Regarding these limitations, several research teams have begun pioneering alternative fractionation approaches.
Fig. 3.
2-D electrophoresis workflow chart. Protein/peptides spots result from two separation steps; (i) by charge (Isoelectric focusing) and (ii) by size (SDS-PAGE).
2.1.2. Multidimensional chromatography separation techniques
A large portion of the commonly used fractionation approaches used to separate venom proteins employed complementary and multi-dimensional chromatographic separation procedures in order to isolate compounds in a single phase. Accordingly, RP-HPLC, ion exchange, as well as gel filtration, have been broadly used to fractionate animal venom components [49,50].
2.1.2.1. Ion exchange chromatography.
Ion exchange chromatography was first used in 1930 as powerful methods to purify protein compounds [51]. One of this technique’s benefits is that it can be used a single step processes as well as associated with other chromatographic techniques. Ion exchange chromatography allows the separation of proteins based on their charge due to amino acid composition, where amino acids are ionized as a function of pH. Proteins with a positive charge, at a certain pH below their isoelectric point, can be separated with use of a cation exchange; on the other hand, proteins with negatively charge residues can be separated using a pH value above their isoelectric point via anion exchange [52]. Several scientific publications now support the use of cation exchange chromatography as a first method for the separation of venom components [49,53,54]. For example, cation exchange chromatography, in combination with gel filtration and affinity chromatography, was used for the isolation of bordonein-L (L-amino acid oxidase) from Crotalus durissus terrificus venom [55]. One gram of C. durissus terrificus venom was first suspended in 50 mL of a 0.05 M sodium acetate buffer (pH 5.5) and then applied onto a CM-cellulose-52 column. Cation exchange fractions were eluted at a flow rate of 0.5 mL/min using a concentration gradient from 0 to 1.0 M NaCl. The first chromatographic step fraction that containing bordonein-L was then fractionated further with two other chromatographic steps, to include molecular exclusion and affinity chromatography [55]. However, the resulting fractions contained a high concentration of salts, which could disrupt the properties of the compound and/or impeded functional assays used further analyze venom proteins. Because of this, an additional step for desalting is often required.
2.1.2.2. Size exclusion chromatography (SEC).
Gel filtration chromatography, also called size exclusion chromatography (SEC), is a technique used to separate compounds based on their size [56]. With this approach there is no physical or chemical interaction between the molecules of the sample and the stationary phase [57]. This method has some important properties to include physical chemical stability, operational simplicity, and inertia (the absence of reaction and adsorption properties), which it allows the separation of small molecules with masses under 100 Da as well as very large molecules of several orders larger into the kDa range [58]. Gel filtration chromatography has been used for several years to separate snake venom compounds. In general as a first step, gel filtration chromatography is highly recommended for the purification of PLA2 components of snake venoms. It is particularly useful for separating PLA2 components from smaller peptides such as neurotoxins and cardiotoxins [59]. Batroxase, a new metalloproteinase was isolated from Bothrops atrox venom using a Sephadex G-75 chromatography column [60]. Moreover, gel filtration chromatographic profiles of Cerastes gasperettii, Walterinnesia aegyptia, Bitis arietans, Echis pyramidum, Echis coloratus and Naja arabica were used in the identification and phylogeny of these Arabian snakes [61]. Another study comparing venom elements from four tiger snakes Notechis scutatus scutatus, Notechis ater serventyi, Notechis ater humphreysi and Notechis ater ater using gel filtration resulted in somewhat different elution profiles on a Superose-12 gel filtration column [62]. However, size exclusion chromatography has some disadvantages such as: (1) a limited number of peaks; (2) filtrations must be performed before using the instrument to prevent containment particulates from ruining the columns and interfering with the detectors; (3) large volumes of eluent are usually required for their operation due to the large size of the gel-filtration columns; and (4) it has an inherent low resolution compared to other chromatographic techniques.
2.1.2.3. Reversed phase-high performance liquid chromatography (RP-HPLC).
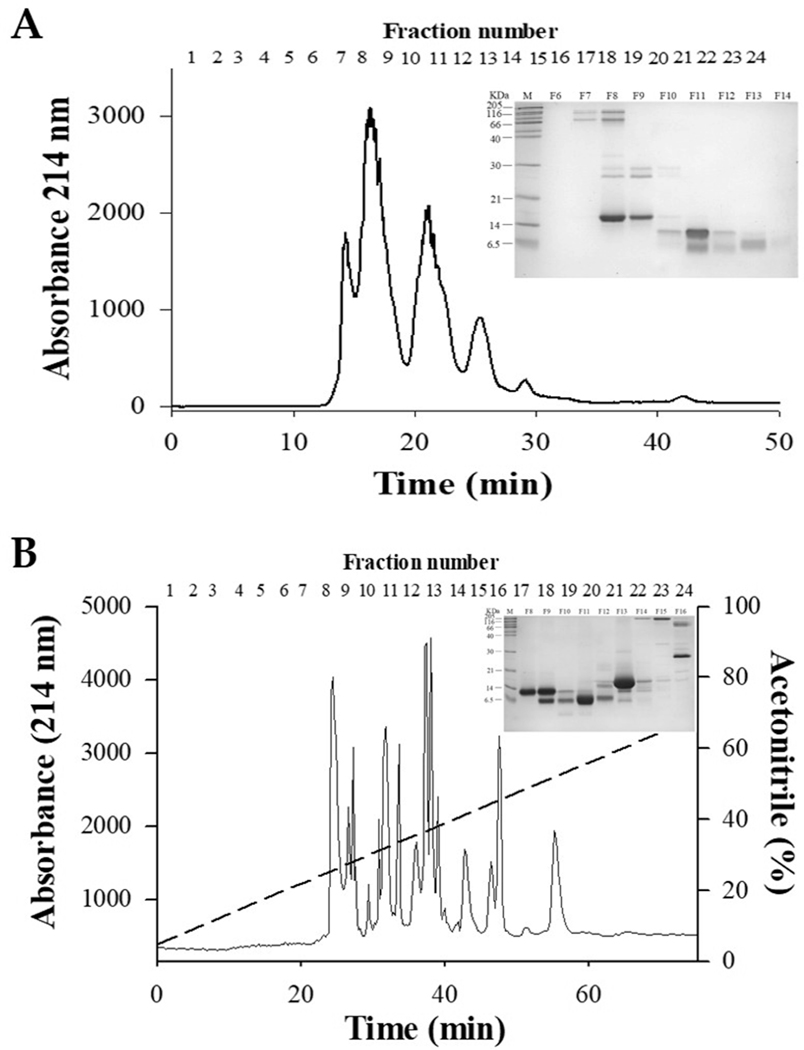
Reversed phase-HPLC has the capacity for high precision. Because of this, it is often used as a more accurate step during the purification process, required for separation of closely related isoforms [63]. The principle of RP-HPLC retention is based on the hydrophobic correlation of the dissolved molecule in the mobile phase with the immobilized hydrophobic ligands coupled to the stationary phase [64]. This technique is very powerful for the analysis of peptides and proteins because of a number of factors including: (1) the use of a mixture of less toxic mobile phases and of slightly lower costs to include water and organic solvents; (2) it uses stationary phases with high stability; (3) it has rapid column equilibrium between stationary and mobile phases; (4) includes flexibility in using gradient elution; (5) has high resolution; and (6) good reproducibility and recovery [63,65]. Complexes of proteins and peptides are usually eluted by the use of an increasing organic solvent concentration gradient. Acetonitrile, methanol and 2-propanol are the most frequently used organic solvents in RP-HPLC. Moreover, a C18-based column and mobile phase are the most often used experimental procedures for RP-HPLC analysis of proteins and peptides. RP-HPLC is extremely flexible when isolating proteins and peptides from numerous synthetic and biological sources to include snake venoms and consequently, is often used in analytical and preparatory applications [65]. The RP-HPLC technique can be used as a single method to separate snake venom compounds or in conjunction with other chromatographic techniques. For example, RP-HPLC has been used as a single chromatographic step to isolate BmTX-I, an Asp49 phospholipase A2, from Bothrops moojeni venom [66]. Moreover, two complementary fractionation techniques have been used to purify a short chain α-neurotoxic protein from the venom of the Mexican coral snake (Micrurus laticollaris). Cation exchange chromatography was usedto fractionate the crude venom and the active fraction was further purified via RP-HPLC on a Vydac C18 column [53]. RP-HPLC provided a good separation of the Walterinnesia aegyptia venom components over a larger scale of fractions than SEC (Fig. 4) [38]. However, this technique does have some drawbacks to include: (1) highly hydrophilic compounds may be missed; (2) columns most appropriate peptides separation often are not perfect for the separation of proteins; and (3) RP-HPLC can cause the irreversible denaturation of protein samples thereby reducing the potential recovery of materials in a biologically active form.
Fig. 4.
Chromatographic profiles of Walterinnesia aegyptia venom. (A) Chromatographic profile of the whole venom was obtained using size exclusion on Sephadex G75. (B) Reversed-phase HPLC chromatogram of Walterinnesia aegyptia venom. An analytic C18 column was used. Retention time is along the x-axis. Venom components were detected at 215 nm and absorbance is indicated on the left axis. The acetonitrile gradient is shown in the HPLC graph and the percentage value corresponds to the right axis for each panel. Fraction numbers are illustrated on the top of the figure. The insert of shows the SDS-PAGE separation of the proteins collected in the fractions. Molecular markers are reported on the side of the gel [38].
2.1.3. OFFGEL electrophoresis
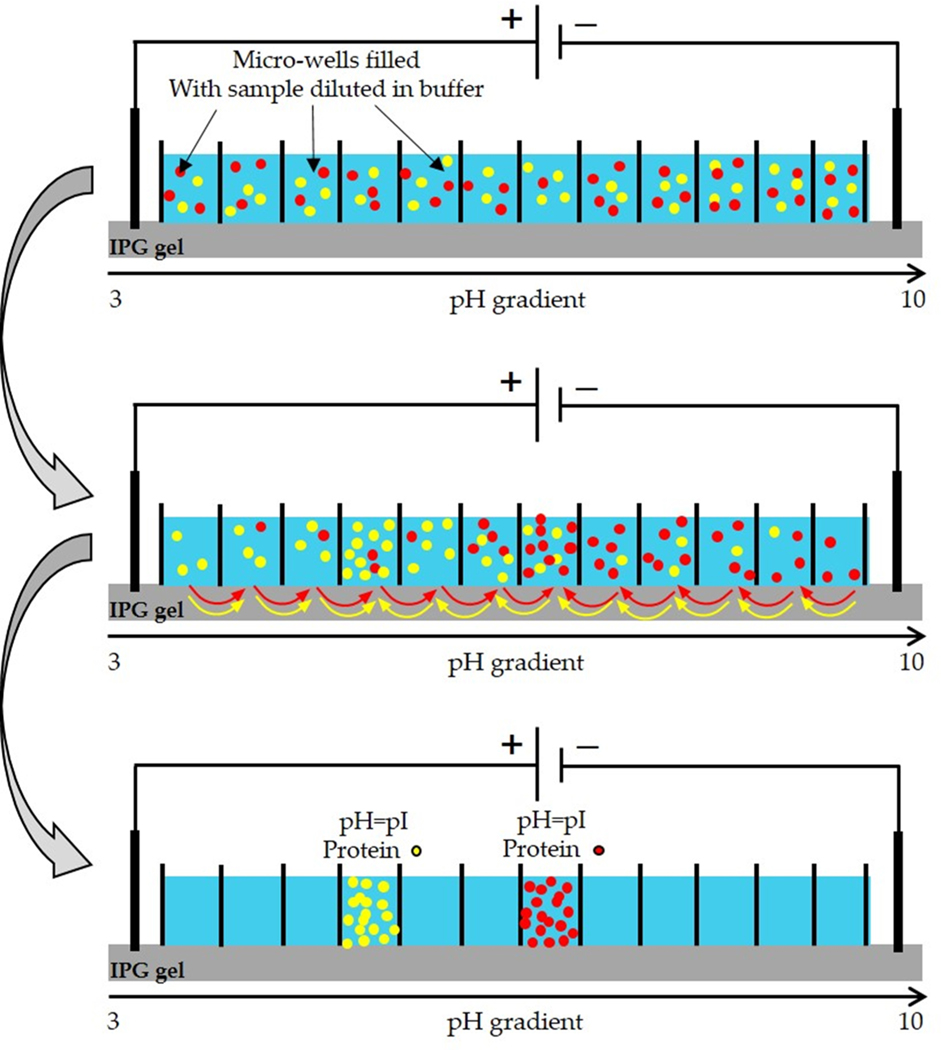
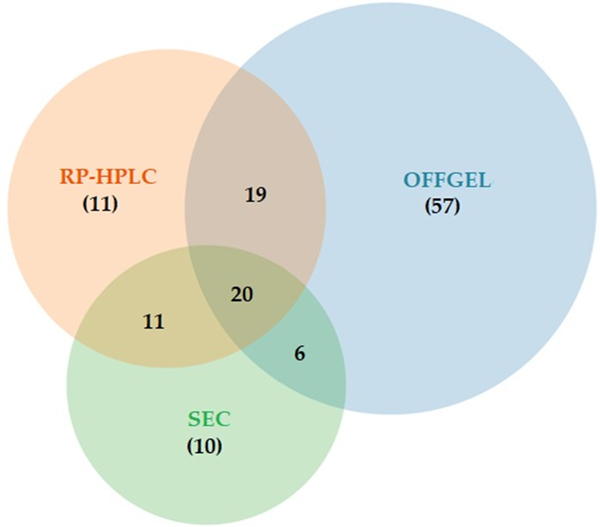
Recently, the OFFGEL fractionator system has been launched by Agilent. This system offers efficient and reproducible separation of biological samples. The Agilent 3100 OFFGEL fractionator utilizes a new isoelectric focus strategy to get premium pI-based fractionation with high reproducibility [67,68]. The pI-dependent separation occurs within the Agilent fractionator in a two-phase system; an upper liquid phase divided into multiple sections and a gel phase used as a pH gradient (IPG) strip. There is no fluid contact between the wells and the charged molecules traveling through the IPG from well to well under an applied electric field until it reaches the well where its net charge equals zero (Fig. 5). The big advantage of OFFGEL in comparison to traditional isoelectric focusing gels, is that the resulting fractions are in liquid phase, making recovery of the compounds much easier prior to further separation techniques or mass spectrometry analysis [69]. In 2006, the first application of OFFGEL electrophoresis for fractionating peptides within the 5–8 pI range from protein digests of Escherichia coli prior to mass detection was reported [67]. Similarly, OFFGEL reagents have been reported to have little effect on the performance of supplementary separation systems. The tryptic peptides from the venom of the box jellyfish Chironex fleckeri, were fractionated using OFFGEL electrophoresis prior to LC-MS/MS analysis [70]. The OFFGEL technique recently also has been used for the first time to separate the venom components of Walterinnesia aegyptia with results comparable to SEC and RP-HPLC (Fig. 6). OFFGEL provided a good separation of the venom components over a larger scale of fractions [38]. Nevertheless, the weakness of OFFGEL electrophoresis is that it requires a long time for completing the fractionation process, usually it takes from a few hours to 2–4 days. Finally, available of different separation techniques are generally complementary among each other and a combination of these different techniques is sometimes required to reach perfect purification of a given individual compound of interest [71,72].
Fig. 5.
Schematic representation of the three steps of OFFGEL electrophoresis principle [135]. First step, micro-wells were filled with the proteins samples diluted in a buffer. Second step, proteins start to migrate throughout the IPG gel to their isoelectric points after applying an electric field. Third step, proteins can be easily recovered in liquid phase after reaching their isoelectric points. Reproduced with permission from Ref. [135].
Fig. 6.
Venn diagram showing the number of ions detected after using SEC, RP-HPLC and OFFGEL separation techniques and those that are common between the three techniques. Reproduced with permission from Ref. [38].
2.2. Mass spectrometry
Since its invention in the early 20th century, mass spectrometry (MS) rapidly has become an essential tool for biology and biochemistry researchers with respect to elaboration of cellular function. It is now the most used discovery-based biosynthesis tool in the field of protein science. MS is well defined as an analytical approach, which generates spectra of the masses of atoms, or molecules that comprise a sample. These spectra are used to establish: (1) the sample elements or isotopes signatures; (2) the mass of sample molecules; and (3) illuminate the chemical structures of molecules in the sample to include peptides and other chemical compounds. The principle of MS employs ionized chemical compounds to produce charged molecules or molecule fragments in combination to measuring their mass to charge ratios [73]. Since 1988, MALDI and electrospray have emerged as two powerful ionization techniques to ionize proteins, peptides, and other large non-volatile molecules of biological importance[74]. For proteomic applications, these two ionization approaches are coupled with a diversity of mass analyzers including numerous types of ion traps and time-of-flight (TOF) systems. It is important for systems that use a tandem mass spectrometer (MS/MS) approach to first determine the precise masses of sample molecules and then to use a subsequent step to determine masses [75]. MS/MS spectra can then be used to identify proteins by searching databases or de novo interpretation to identify the sequence and/or post-translation modifications of sample molecules. The diverse application of mass spectrometry provides a chance to gain information about simple molecular masses to the sequences of sample peptides. For instance, on-line liquid chromatography (LC) analysis alongside ESI-MS or MALDI-MS analysis of crude venoms gives a molecular mass fingerprint of venom components.
2.2.1. Matrix-assisted laser desorption/ionization (MALDI)
Matrix-assisted laser desorption/ionization (MALDI) is a soft ionization approach used in mass spectrometry analysis. MALDI-MS was first introduced in 1988 by Hillenkamp and Karas [76]. It is particularly well adapted for the analysis of complex natural mixtures to include bio-fluids and natural extracts such as venoms [74,77]. In MALDI analysis, the analyte is first co-crystallized with a major molar excess of an organic matrix compound that strongly absorbes at the laser wavelength used, and then laser irradiation of this analyte–matrix mixture leads to the vaporization of the matrix along with analyte that is in it. Depending on the type of sample, 2,5-dihydroxybenzoic acid (DHB), α-cyano-4-hydroxycinnamic acid (CHCA), and 3,5-dimethoxy-4-hydroxycinnamic acid (sinapinic acid or SA) are the most frequently utilized matrices. Some matrices can complement each other with respect to obtaining complete protein sequence coverage. For instance, peptide mapping with CHCA can resolve low-mass peptide ions (2500 Da), while SA may provide better coverage for higher-mass peptides (greater than2500 Da) [78,79]. MALDI-MS is frequently used in complement with TOF mass analyzers. MALDI-TOF-MS was used to obtain mass patterns of the major peptides (< 8 kDa) found in pooled venoms from Bothrops jararaca, Bothrops insularis, Bothrops alternatus, Bothrops jararacussu, Bothrops neuwiedi, Crotalus viridis, Crotalus adamanteus, and Crotalus durissus terrificus [80]. Currently, investigators are further developing this technique to obtain more structural information from peptides/proteins using a combination of chemical and enzymatic modifications of the analyte prior to analysis by MALDI-TOF-MS. MALDI-TOF-MS analysis of gel tryptic-digested peptides of the venom of four Malaysian snake species to include Naja kaouthia, Ophiophagus hannah, Bungarus fasciatus, and Calloselasma rhodostoma has been performed [42]. Moreover, the full sequence of eight single charged ions of m/z 873–1277 (M + H)+ were identified in venom of two elapid snakes, Naja mossambica mossambica and Notechis scutatus, using MALDI-TOF/TOF mass spectrometry [81]. However, MALDI is not compatible with HPLC. Furthermore, the salts and detergents commonly used in biomolecular research can interfere with either crystallization or ionization.
2.2.2. Electrospray ionization (ESI)
Over the last decade, electrospray ionization mass spectrometry (ESI-MS) has developed into a rapid and precise method for determining masses of proteins and peptides and can be used to validate protein sequences [82]. In linear mode detection, ESI-MS is more sensitive and accurate in mass determination than MALDI-TOF-MS. Furthermore, along with the HPLC approach for molecular fractionation of samples prior to mass spectrometric analysis, LC-ESI-MS has turned into an extremely effective approach capable of analyzing both small and large molecules of different polarities in a complex biological sample [83]. Currently, ESI-MS is not only employed as a balance to weigh protein molecules but additionally to get a much deeper understanding of 3D protein structures, non-covalent interactions, post-translation modifications, and amino acid sequences [84]. In general, ESI process produces multiple charged ions. The development of ESI-MS has rapidally enabled the elaboration of animal venom proteomics. The bioactive proteins/peptides profile of seven Bothrops venom species to include B. jararaca, B. jararacussu, B. alternatus, B. insularis, B. moojeni, B. erythromelas and B. leucurus have been studied using direct infusion nano-electrospray ionization mass spectrometry (nano-ESI-MS) [85]. ESI-MS has also provided new means for the study of the functions and structures of Chinese Agkistrodon blomhoffii snake venom PLA2 [86]. Moreover, collision induced dissociation (CID) using an ESI-MS/MS spectra has been used to identify and describe a bradykinin-potentiating peptide (BPP) from Amazon Bothrops atrox venom with m/z 1384.7386 (M + H)+ [87].
2.3. Peptide sequencing
In addition to separation and mass spectrometry techniques, bioinformatics has become the third significant development that has advanced proteomics. Bioinformatics allows scientist to collect, store, process, and visualize the vast amount of data produced by proteomics studies [17]. There are two basic approaches to peptide sequencing. MS/MS data are typically either MALDI-TOF/TOF-MS/MS or LC-ESI-QTOF-MS/MS. The most popular approach is to search databases of known genomes, which looks for the peptide that produced the mass spectrum in a protein database. Mascot and Sequest are the most used approaches for protein identification [88,89]. These methods are effective but often give false positives or incorrect identifications. For unknown genomes, de novo sequencing utilizes computational approaches to deduce the sequences or partial sequences of peptides directly from the experimental MS/MS spectra [90]. To aid the assignment of sequences a number of chemical techniques have been developed to favor the formation of more stable ‘y’ or ‘b’ ions [91,92]. Currently, several software algorithms packages have been produced to determine protein sequences from MS/MS data [93]. De novo sequencing is embodied in programs such as PEAKS, which can deduce sequences directly from MS/MS spectra data. It computes the best possible sequence among all possible amino acid combinations [94]. Sequence similarity searches have also been employed through BLAST to identify proteins via their known homologues in other species. A recent study used PEAKS to create a de novo sequence for a new PLA2 from Bothriopsis taeniata venom [95].
The older standard of N-terminal sequencing (Edman degradation) also remains an important tool for peptide sequencing. It remains useful for partial or full amino acid sequencing of animal venom toxins. Edman degradation can be complemented with MS/MS analysis to identify the isobaric amino acids isoleucine/leucine and for N-terminal sequencing. A potential N-glycosylation site in a novel C-lectin protein from snake Trimeresurus stejnegeri venom was sequenced by Edman degradation [96]. Another example, a fibrinogenolytic metalloproteinase, named moojenin, was isolated from Bothrops moojeni venom and sequenced by Edman sequencing [97].
Since identification of the first animal venom peptides using the traditional methods, the analytical description of animal venoms has been continuously enriched by the new approaches and technologies of venomics. The implementation of next generation sequencing and high-resolution mass spectrometry into proteomic workflows have significantly advanced the analysis of animal venoms. More recently, a protein profiling strategy called bottom up or shotgun proteomics has been developed to identify proteins, characterize their amino acid sequences and post-translational modifications. This strategy involves proteolytic digestion of a complex mixture of proteins prior to chromatographic separation coupled with mass spectrometry [98]. Nevertheless, the proteolytic digestion step rises up the number of ions existing in a sample and disconnect structural relation within highly homologous isoforms and ultimately happening post-translational modified proteins. To avoid these issues and to overcome the details by the digestion step, top-down proteomics was introduced in a study of Indonesian King cobra Ophiophagus hannah venom [99]. As a result of using top-down mass spectrometry, a number of proteins from different toxin families to include 11 three-finger toxins (7–7.9 kDa), a Kunitz-type inhibitor (6.3 kDa), ohanin (11.9 kDa), a novel phospholipase A2 molecule (13.8 kDa), and the cysteine-rich secretory protein (CRISP) ophanin (25 kDa) have been identified from the Indonesian king cobra venom. The first venom study using a bottom-up proteomic approach was conducted on the venom of the Western diamondback rattlesnake Crotalus atrox [100]. Currently, a combination of bottom-up and top-down proteomics, middle-down proteomics, has received a lot of attention because it avoids redundant peptide sequences and other drawbacks. Middle-down proteomics has been used to identify several compounds, including metalloproteases, metalloprotease inhibitor, PLA2, disintegrins, C-type lectin, and Kunitz-type protease inhibitor from the venom of Vipera anatolica [101].
Recently, a new method entitled at-line nanofractionation methodology, which is based on the principles of bioassay-guided fractionation. Briefly, this method combines liquid chromatography analysis of a crude snake venom with parallel mass spectrometry detection and high-resolution nanofractionation onto 384-well plates for bio-assaying, enabled by the presence of a post-column flow split. Subsequently, bioactivity chromatograms can be constructed by plotting the readout of each well against the time of the corresponding fraction. A recent study showed that this approach enables them to separate snake venom into fractions, and identify the proteins present in those fractions that cause coagulopathic effects in the bioassay [102–104].
3. Application of snake venom proteomics
To date, snake venomic studies have been on more than 200 snake species and these numbers continue to grow (Table 1) [105–115]. Identification of snake venom components using proteomic approaches has had several benefits for different fields to include basic research, development of new research tools, drug discovery and clinical diagnosis. These studies offer the potential to discover an extremely rich source of pharmacologically active proteins and peptides. Calciseptine is a specific blocker of the L-type calcium channel but has no effect on N-type and T-type channels [116]. Calciseptine is a 60-amino acid peptide isolated from the venom of the black mamba, Dendroaspis polylepis polylepis, using three purification steps: gel filtration, ion exchange and RP-HPLC. The peptide was fully sequenced using Edman degradation. With recent advances in snake venomics, the combination of proteomic and transcriptomic were used to to identify several toxins within the venom of the medically important Australian snake Pseudonaja textilis [117]. Prothrombinase coagulation factors, neurotoxic textilotoxin phospholipase A2 (PLA2), three-finger toxins (3FTx), and the Kunitz-type protease inhibitor textilinin, venom metalloproteinase, C-type lectins, cysteine rich secretory proteins, calreticulin, and dipeptidase from venom glands all have been sequenced via de novo transcriptome analysis [117].
Table 1.
Most recent snake species whose venoms have been examined by proteomic and transcriptomic approaches.
| Snake species | Proteomic | Transcriptomic |
|---|---|---|
| Ahaetulla prasina and Borikenophis portoricensis | [114] | |
| Boiga irregularis | [105] | |
| Vipera ammodytes ammodytes | [106] | [106] |
| Bothrops Atrox | [107] | |
| Micrurus corallinus, M. lemniscatus carvalhoi, M. lemniscatus lemniscatus, M. paraensis, M. spixii spixii and M. surinamensis | [108] | [108] |
| Pseudonaja Textilis | [109] | |
| Dendroaspis angusticeps, D. polylepis, D. jamesoni jamesoni, D. j. kaimosae and D. viridis | [110] | [110] |
| Crotalus simus, C. tzbacan and C. culminates | [111] | [111] |
| Gloydius intermedius | [112] | [112] |
| Bothrops jararaca | [113] | [113] |
| Mixcoatlus melanurus | [115] | |
In addition, snake venom proteomes may help in understanding the biology and ecology of snakes and provide a valuable tool to identify possible variances in snake venom composition resulting from ontogenetic variability and differing geographic distribution of species. Prior proteomic and transcriptomic studies showed a significant intraspecific variation in venom composition of Mojave Rattlesnakes (Crotalus scutulatus) according to the geographic distribution of these snakes [118]. Mojave Rattlesnakes are widely distributed in the deserts of North America and south-central Mexico. Furthermore, the venom of individual Crotalus scutulatus snakes generally exhibit two venom phenotypes, highly neurotoxic (type A) and hemotoxic (type B) [119]. The type A neurotoxic venom was present in cold and heavy rainfall areas indicating a relation between the specific enzymatic activity of venom and climate [120].
One of the important contributions made by the venomic strategy is the ability to combine emerging information about the composition of snake venoms with development of antivenoms. This combined approach is named antivenomics. Antivenomics is a proteomics-based approach first established in 2008 to examine the immunological profile of antivenoms [121]. Death resulting from snakebites poses a serious health problem in the rural communities of developing countries. Globally, more than 5 million snakebite envenomings happen every year, resulting in about 400,000 amputations and more than 100,000 deaths [122]. At the end of the nineteenth century, snake antivenoms were first developed by Calmette using horses as a host for production [123]. To evaluate the preclinical characterization of antivenoms, the antivenomic strategy employs different immunological procedures, such as western blot analyses, in vitro and in vivo venom toxicity neutralization assays, and ELISA. First generation antivenomics is based on the immuno-depletion of antivenom-binding toxins upon incubating the whole venom with antivenom. This is then followed by the addition of a secondary antibody to precipitate the antigen–antibody complexes from the reaction mixture. Venom components that remain in the supernatant are called non-immune depleted proteins, which failed to raise antibodies to the antivenom. RP-HPLC separation of the non-precipitated fraction, supernatant from the immunoprecipitation, can then be compared with the HPLC profile of the whole venom in order to assess the degree of immune-depletion [121]. Such as first generation antivenomics approach was applied to test the immunoreactivity of crotalic antivenom against the subspecies cascavella and collilineatus of the Brazilian tropical rattlesnake Crotalus durissus [124]. Second generation antivenomics is a solid-phase interaction that arises from the combination of immune-affinity chromatography and proteomic analysis of the venom components in the non-bound fractions [125]. It was developed to overcome the drawbacks of the immune-depletion in-solution methods. The whole venom is incubated with antivenom antibodies immobilized onto beads of an immune-affinity column. The immune-captured or antivenom-bound venom fractions is eluted from the unbound venom fraction by a change in pH. The HPLC profiles of both samples are compared to that of the control venom sample. The degree of immune-recognition for venom components is then quantified by integrating the chromatographic peak areas. Recently, second generation antivenomics has become the preferred approach to characterize of antivenoms [126,127]. In combination with this, several types of immunoassays, including immunoblotting and enzyme-immunoassays HPLC/ELISA, have been used with the venomic approach to characterize antivenom toward specific venom toxins [128–130]. Thus, along with the venomic appraoch, antivenomics has evolved as an important tool in the preclinical elaboration of antivenoms.
4. Concluding remarks
Advanced venom fractionation techniques, advances in mass spectrometry and NMR spectroscopy, miniaturization of functional assays, and the ability to directly analyze toxin transcripts from venom-gland cDNA libraries together have driven the recent rapid advancement in venomics [131–133]. Along with this continuing refinement in technologies that enable high-throughput screening and structural characterization of toxins promise to rapidly, improve drug discovery based on animal venoms.
Acknowledgments
Sources of funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Abbreviations:
- PLA2
phospholipase A2
- 3FTx
three-finger toxins
- DHB
2,5-dihydroxybenzoic acid
- CHCA
α-cyano-4-hydroxycinnamic acid
- SA
3,5-dimethoxy-4-hydroxycinnamic acid
- BPP
bradykinin-potentiating peptide
Footnotes
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- [1].Wilkins MR, Sanchez JC, Gooley AA, Appel RD, Humphery-Smith I, Hochstrasser DF, Williams KL, Progress with proteome projects: why all proteins expressed by a genome should be identified and how to do it, Biotechnology & genetic engineering reviews 13 (1996) 19–50. [DOI] [PubMed] [Google Scholar]
- [2].Jensen ON, Interpreting the protein language using proteomics, Nat Rev Mol Cell Biol 7 (2006) 391–403. [DOI] [PubMed] [Google Scholar]
- [3].Bauw G, Van Damme J, Puype M, Vandekerckhove J, Gesser B, Ratz GP, Lauridsen JB, Celis JE, Protein-electroblotting and -microsequencing strategies in generating protein data bases from two-dimensional gels, Proceedings of the National Academy of Sciences of the United States of America, 86 (1989) 7701–7705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].Diz AP, MartÍNez-FernÁNdez M, RolÁN-Alvarez E, Proteomics in evolutionary ecology: linking the genotype with the phenotype, Molecular Ecology 21 (2012) 1060–1080. [DOI] [PubMed] [Google Scholar]
- [5].Chandramouli K, Qian P-Y, Proteomics: challenges, techniques and possibilities to overcome biological sample complexity, Hum Genomics Proteomics 2009 (2009) 239204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Aslam B, Basit M, Nisar MA, Khurshid M, Rasool MH, Proteomics: Technologies and Their Applications, Journal of Chromatographic Science 55 (2017) 182–196. [DOI] [PubMed] [Google Scholar]
- [7].Koh DC, Armugam A, Jeyaseelan K, Snake venom components and their applications in biomedicine, Cell Mol Life Sci 63 (2006) 3030–3041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Harvey AL, Toxins and drug discovery, Toxicon 92 (2014) 193–200. [DOI] [PubMed] [Google Scholar]
- [9].Abd El-Aziz TM, Al Khoury S, Jaquillard L, Triquigneaux M, Martinez G, Bourgoin-Voillard S, Sève M, Arnoult C, Beroud R, De Waard M, Actiflagelin, a new sperm activator isolated from Walterinnesia aegyptia venom using phenotypic screening, Journal of Venomous Animals and Toxins including Tropical Diseases 24 (2018) 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Da Silva SL, Rowan EG, Albericio F, Stabeli RG, Calderon LA, Soares AM, Animal toxins and their advantages in biotechnology and pharmacology, Biomed Res Int 2014 (2014) 951561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [11].Palatty PL, Saldanha E, The medicinal use of venoms and toxins, Journal of the Indian Medical Association 111 (2013) 51–53. [PubMed] [Google Scholar]
- [12].Lewis RJ, Garcia ML, Therapeutic potential of venom peptides, Nat Rev Drug Discov 2 (2003) 790–802. [DOI] [PubMed] [Google Scholar]
- [13].Mohamed Abd El-Aziz T, Garcia Soares A, Stockand JD, Snake Venoms in Drug Discovery: Valuable Therapeutic Tools for Life Saving, Toxins (Basel) 11 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Marrakchi N, Guermazi S, El Ayeb M, The anti-platelet aggregation drugs: new approaches based on snake venom polypeptides, Arch Inst Pasteur Tunis 74 (1997) 9–15. [PubMed] [Google Scholar]
- [15].Bazan-Socha S, Marcinkiewicz C, Musial J, Snake venom disintegrins–source of knowledge and development of the new drugs, Pol Arch Med Wewn 110 (2003) 1345–1351. [PubMed] [Google Scholar]
- [16].Kemp E, Steinbruchel D, Starklint H, Larsen S, Henriksen I, Dieperink H, Renal xenograft rejection: prolonging effect of captopril, ACE-inhibitors, prostacyclin, and cobra venom factor, Transplant Proc 19 (1987) 4471–4474. [PubMed] [Google Scholar]
- [17].Kaas Q, Craik DJ, Bioinformatics-Aided Venomics, Toxins (Basel) 7 (2015) 2159–2187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Fernández J, Lomonte B, Sanz L, Angulo Y, Gutiérrez JM, Calvete JJ, Snake Venomics of Bothriechis nigroviridis Reveals Extreme Variability among Palm Pitviper Venoms: Different Evolutionary Solutions for the Same Trophic Purpose, Journal of Proteome Research 9 (2010) 4234–4241. [DOI] [PubMed] [Google Scholar]
- [19].Angulo Y, Escolano J, Lomonte B, Gutiérrez JM, Sanz L, Calvete JJ, Snake Venomics of Central American Pitvipers: Clues for Rationalizing the Distinct Envenomation Profiles of Atropoides nummifer and Atropoides picadoi, Journal of Proteome Research 7 (2008) 708–719. [DOI] [PubMed] [Google Scholar]
- [20].Abd El-Aziz TM, Shoulkamy MI, Hegazy AM, Stockand JD, Mahmoud A, Mashaly AMA, Comparative study of the in vivo toxicity and pathophysiology of envenomation by three medically important Egyptian snake venoms, Archives of Toxicology 94 (2020) 335–344. [DOI] [PubMed] [Google Scholar]
- [21].Lomonte B, Rey-Suárez P, Tsai W-C, Angulo Y, Sasa M, Gutiérrez JM, Calvete JJ, Snake venomics of the pit vipers Porthidium nasutum, Porthidium ophryomegas, and Cerrophidion godmani from Costa Rica: Toxicological and taxonomical insights, Journal of Proteomics 75 (2012) 1675–1689. [DOI] [PubMed] [Google Scholar]
- [22].Alape-Giron A, Sanz L, Escolano J, Flores-Diaz M, Madrigal M, Sasa M, Calvete JJ, Snake venomics of the lancehead pitviper Bothrops asper: geographic, individual, and ontogenetic variations, J Proteome Res 7 (2008) 3556–3571. [DOI] [PubMed] [Google Scholar]
- [23].Ching ATC, Paes Leme AF, Zelanis A, Rocha MMT, Furtado MDFD, Silva DA, Trugilho MRO, da Rocha SLG, Perales J, Ho PL, Serrano SMT, Junqueira-de-Azevedo ILM, Venomics Profiling of Thamnodynastes strigatus Unveils Matrix Metalloproteinases and Other Novel Proteins Recruited to the Toxin Arsenal of Rear-Fanged Snakes, Journal of Proteome Research 11 (2012) 1152–1162. [DOI] [PubMed] [Google Scholar]
- [24].Zhang Y, Why do we study animal toxins? Dongwuxue Yanjiu 36 (2015) 183–222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Utkin YN, Animal venom studies: Current benefits and future developments, World J Biol Chem 6 (2015) 28–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].El-Aziz TMA, Ravelet C, Molgo J, Fiore E, Pale S, Amar M, Al-Khoury S, Dejeu J, Fadl M, Ronjat M, Taiwe GS, Servent D, Peyrin E, De Waard M, Efficient functional neutralization of lethal peptide toxins in vivo by oligonucleotides, Scientific reports (2017) 7202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Lee CY, Snake Venoms, Springer, Berlin Heidelberg, 2012. [Google Scholar]
- [28].Menez A, Stocklin R, Mebs D, ‘Venomics’ or : The venomous systems genome project, Toxicon 47 (2006) 255–259. [DOI] [PubMed] [Google Scholar]
- [29].de Graaf DC, Aerts M, Danneels E, Devreese B, Bee, wasp and ant venomics pave the way for a component-resolved diagnosis of sting allergy, J Proteomics 72 (2009) 145–154. [DOI] [PubMed] [Google Scholar]
- [30].Calvete JJ, Proteomic tools against the neglected pathology of snake bite envenoming, Expert review of proteomics 8 (2011) 739–758. [DOI] [PubMed] [Google Scholar]
- [31].Fox JW, Ma L, Nelson K, Sherman NE, Serrano SMT, Comparison of indirect and direct approaches using ion-trap and Fourier transform ion cyclotron resonance mass spectrometry for exploring viperid venom proteomes, Toxicon 47 (2006) 700–714. [DOI] [PubMed] [Google Scholar]
- [32].Mitchell Wells J, McLuckey SA, Collision-Induced Dissociation (CID) of Peptides and Proteins, Methods in Enzymology, Academic Press (2005) 148–185. [DOI] [PubMed] [Google Scholar]
- [33].Cottrell JS, Protein identification using MS/MS data, Journal of Proteomics 74 (2011) 1842–1851. [DOI] [PubMed] [Google Scholar]
- [34].Calvete JJ, Juarez P, Sanz L, Snake venomics, Strategy and applications, J Mass Spectrom 42 (2007) 1405–1414. [DOI] [PubMed] [Google Scholar]
- [35].Calvete JJ, Sanz L, Angulo Y, Lomonte B, Gutiérrez JM, Venoms, venomics, antivenomics, FEBS Letters 583 (2009) 1736–1743. [DOI] [PubMed] [Google Scholar]
- [36].Gutiérrez JM, Lomonte B, León G, Alape-Girón A, Flores-Díaz M, Sanz L, Angulo Y, Calvete JJ, Snake venomics and antivenomics: Proteomic tools in the design and control of antivenoms for the treatment of snakebite envenoming, Journal of Proteomics 72 (2009) 165–182. [DOI] [PubMed] [Google Scholar]
- [37].Calvete JJ, Snake venomics: From the inventory of toxins to biology, Toxicon 75 (2013) 44–62. [DOI] [PubMed] [Google Scholar]
- [38].Abd El Aziz TM, Bourgoin-Voillard S, Combemale S, Beroud R, Fadl M, Seve M, De Waard M, Fractionation and proteomic analysis of the Walterinnesia aegyptia snake venom using OFFGEL and MALDI-TOF-MS techniques, Electrophoresis 36 (2015) 2594–2605. [DOI] [PubMed] [Google Scholar]
- [39].Tan PT, Khan AM, Brusic V, Bioinformatics for venom and toxin sciences, Brief Bioinform 4 (2003) 53–62. [DOI] [PubMed] [Google Scholar]
- [40].O’Farrell PH, High resolution two-dimensional electrophoresis of proteins, J Biol Chem 250 (1975) 4007–4021. [PMC free article] [PubMed] [Google Scholar]
- [41].Carrette O, Burkhard PR, Sanchez JC, Hochstrasser DF, State-of-the-art two-dimensional gel electrophoresis: a key tool of proteomics research, Nature protocols 1 (2006) 812–823. [DOI] [PubMed] [Google Scholar]
- [42].Vejayan J, Khoon TL, Ibrahim H, Comparative analysis of the venom proteome of four important Malaysian snake species, Journal of Venomous Animals and Toxins including Tropical Diseases 20 (2014) 1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Menezes MC, Furtado MF, Travaglia-Cardoso SR, Camargo AC, Serrano SM, Sex-based individual variation of snake venom proteome among eighteen Bothrops jararaca siblings, Toxicon 47 (2006) 304–312. [DOI] [PubMed] [Google Scholar]
- [44].Serrano SM, Shannon JD, Wang D, Camargo AC, Fox JW, A multifaceted analysis of viperid snake venoms by two-dimensional gel electrophoresis: an approach to understanding venom proteomics, Proteomics 5 (2005) 501–510. [DOI] [PubMed] [Google Scholar]
- [45].Vejayan J, Shin Yee L, Ponnudurai G, Ambu S, Ibrahim I, Protein profile analysis of Malaysian snake venoms by two-dimensional gel electrophoresis, Journal of Venomous Animals and Toxins including Tropical Diseases 16 (2010) 623–630. [Google Scholar]
- [46].Nawarak J, Phutrakul S, Chen ST, Analysis of lectin-bound glycoproteins in snake venom from the Elapidae and Viperidae families, J Proteome Res 3 (2004) 383–392. [DOI] [PubMed] [Google Scholar]
- [47].Yanes O, Avilés FX, Wenzel R, Nazabal A, Zenobi R, Calvete JJ, Proteomic Profiling of a Snake Venom Using High Mass Detection MALDI-TOF Mass Spectrometry, Journal of the American Society for Mass Spectrometry 18 (2007) 600–606. [DOI] [PubMed] [Google Scholar]
- [48].Rabilloud T, Two-dimensional gel electrophoresis in proteomics: old, old fashioned, but it still climbs up the mountains, Proteomics 2 (2002) 3–10. [PubMed] [Google Scholar]
- [49].Bordon KC, Perino MG, Giglio JR, Arantes EC, Isolation, enzymatic characterization and antiedematogenic activity of the first reported rattlesnake hyaluronidase from Crotalus durissus terrificus venom, Biochimie 94 (2012) 2740–2748. [DOI] [PubMed] [Google Scholar]
- [50].Siigur J, Vija H, Samel M, Tõnismägi K, Trummal K, Aaspõllu A, Subbi J, Rönnholm G, Kalkkinen N, Siigur E, Separation and analysis of peptides and proteins from Vipera lebetina snake venom, Procedia Chemistry 2 (2010) 109–115. [Google Scholar]
- [51].Wixom RL, Chapter 1 The beginnings of chromatography – The pioneers (1900–1960), in: Gehrke RLWCW, Bayer E. (Eds.), Journal of Chromatography Library, Elsevier, Ny, 2001, pp. 1–38. [Google Scholar]
- [52].Liu X, Pohl CA, Comparison of reversed-phase/cation-exchange/anion-exchange trimodal stationary phases and their use in active pharmaceutical ingredient and counterion determinations, Journal of chromatography. A 1232 (2012) 190–195. [DOI] [PubMed] [Google Scholar]
- [53].Carbajal-Saucedo A, Lopez-Vera E, Benard-Valle M, Smith EN, Zamudio F, de Roodt AR, Olvera-Rodriguez A, Isolation, characterization, cloning and expression of an alpha-neurotoxin from the venom of the Mexican coral snake Micrurus laticollaris (Squamata: Elapidae), Toxicon 66 (2013) 64–74. [DOI] [PubMed] [Google Scholar]
- [54].Weinstein SA, Smith LA, Preliminary fractionation of tiger rattlesnake (Crotalus tigris) venom, Toxicon 28 (1990) 1447–1455. [DOI] [PubMed] [Google Scholar]
- [55].Bordon KC, Wiezel GA, Cabral H, Arantes EC, Bordonein-L, a new L-amino acid oxidase from Crotalus durissus terrificus snake venom: isolation, preliminary characterization and enzyme stability, J Venom Anim Toxins Incl Trop Dis 21 (2015) 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [56].Hagel L, Gel-filtration chromatography, Current protocols in molecular biology / edited by Frederick M Ausubel: et al. , Chapter 10 (2001) Unit 10.19. [DOI] [PubMed] [Google Scholar]
- [57].Cutler P, Size-exclusion chromatography, Methods Mol Biol 244 (2004) 239–252. [DOI] [PubMed] [Google Scholar]
- [58].Hagel L, Gel filtration: size exclusion chromatography, Methods of biochemical analysis 54 (2011) 51–91. [DOI] [PubMed] [Google Scholar]
- [59].Rochat H, Martin-Eauclaire MF, Animal Toxins: Facts and Protocols, Birkhäuser Basel (2013). [Google Scholar]
- [60].Cintra ACO, De Toni LGB, Sartim MA, Franco JJ, Caetano RC, Murakami MT, Sampaio SV, Batroxase, a new metalloproteinase from B. atrox snake venom with strong fibrinolytic activity, Toxicon 60 (2012) 70–82. [DOI] [PubMed] [Google Scholar]
- [61].Al Asmari A, Manthiri RA, Khan HA, Identification and phylogeny of Arabian snakes: Comparison of venom chromatographic profiles versus 16S rRNA gene sequences, Saudi Journal of Biological Sciences 21 (2014) 436–441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [62].John TR, Kaiser II, Comparison of venom constituents from four tiger snake (Notechis) subspecies, Toxicon 28 (1990) 1117–1122. [DOI] [PubMed] [Google Scholar]
- [63].Aguilar MI, Hearn MT, High-resolution reversed-phase high-performance liquid chromatography of peptides and proteins, Methods Enzymol 270 (1996) 3–26. [DOI] [PubMed] [Google Scholar]
- [64].Mant CT, Hodges RS, [1] Analysis of peptides by high-performance liquid chromatography, Methods in Enzymology, Academic Press1996, pp. 3–50. [DOI] [PubMed] [Google Scholar]
- [65].Aguilar M-I, Reversed-Phase High-Performance Liquid Chromatography, in: Aguilar M−I(Ed.), HPLC of Peptides and Proteins: Methods and Protocols, Springer, New York, Totowa, NJ, 2004, pp. 9–22. [DOI] [PubMed] [Google Scholar]
- [66].Calgarotto AK, Damico DC, Ponce-Soto LA, Baldasso PA, Da Silva SL, Souza GH, Eberlin MN, Marangoni S, Biological and biochemical characterization of new basic phospholipase A(2) BmTX-I isolated from Bothrops moojeni snake venom, Toxicon 51 (2008) 1509–1519. [DOI] [PubMed] [Google Scholar]
- [67].Horth P, Miller CA, Preckel T, Wenz C, Efficient fractionation and improved protein identification by peptide OFFGEL electrophoresis, Mol Cell Proteomics 5 (2006) 1968–1974. [DOI] [PubMed] [Google Scholar]
- [68].Chenau J, Michelland S, Sidibe J, Seve M, Peptides OFFGEL electrophoresis: a suitable pre-analytical step for complex eukaryotic samples fractionation compatible with quantitative iTRAQ labeling, Proteome Sci 6 (2008) 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [69].Heller M, Michel PE, Morier P, Crettaz D, Wenz C, Tissot JD, Reymond F, Rossier JS, Two-stage Off-Gel isoelectric focusing: protein followed by peptide fractionation and application to proteome analysis of human plasma, Electrophoresis 26 (2005) 1174–1188. [DOI] [PubMed] [Google Scholar]
- [70].Brinkman DL, Aziz A, Loukas A, Potriquet J, Seymour J, Mulvenna J, Venom Proteome of the Box Jellyfish Chironex fleckeri, PLoS ONE 7 (2012) e47866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [71].Bian LJ, Wu P, Yang XY, Two-step chromatographic method for separation and purification of nerve growth factor from venom of Chinese cobra, J Chromatogr B Analyt Technol Biomed Life Sci 805 (2004) 119–125. [DOI] [PubMed] [Google Scholar]
- [72].Kishino S, Miyazaki K, Separation methods for glycoprotein analysis and preparation, Journal of chromatography, B, Biomedical sciences and applications 699 (1997) 371–381. [DOI] [PubMed] [Google Scholar]
- [73].Sparkman OD, Mass Spectrometry Desk Reference, Global View Pub. (2000). [Google Scholar]
- [74].Chapman JR, Mass spectrometry of proteins and peptides, Humana Press, Totowa, NJ, 2000. [Google Scholar]
- [75].Hunt DF, Yates JR, Shabanowitz J, Winston S, Hauer CR, Protein sequencing by tandem mass spectrometry, Proceedings of the National Academy of Sciences of the United States of America, 83 (1986) 6233–6237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [76].Karas M, Hillenkamp F, Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons, Analytical chemistry 60 (1988) 2299–2301. [DOI] [PubMed] [Google Scholar]
- [77].Tubbs KA, Nelson RW, Krone JR, Bieber AL, MASS SPECTRAL STUDIES OF SNAKE VENOMS AND SOME OF THEIR TOXINS, Journal of Toxicology: Toxin Reviews 19 (2000) 1–22. [Google Scholar]
- [78].Fitzgerald MC, Parr GR, Smith LM, Basic matrixes for the matrix-assisted laser desorption/ionization mass spectrometry of proteins and oligonucleotides, Analytical chemistry 65 (1993) 3204–3211. [DOI] [PubMed] [Google Scholar]
- [79].Beavis RC, Chaudhary T, Chait BT, α-Cyano-4-hydroxycinnamic acid as a matrix for matrixassisted laser desorption mass spectromtry, Organic Mass Spectrometry 27 (1992) 156–158. [Google Scholar]
- [80].Wermelinger LS, Dutra DL, Oliveira-Carvalho AL, Soares MR, Bloch C Jr., R.B. Zingali, Fast analysis of low molecular mass compounds present in snake venom: identification of ten new pyroglutamate-containing peptides, Rapid Commun Mass Spectrom 19 (2005) 1703–1708. [DOI] [PubMed] [Google Scholar]
- [81].Munawar A, Trusch M, Georgieva D, Hildebrand D, Kwiatkowski M, Behnken H, Harder S, Arni R, Spencer P, Schlüter H, Betzel C, Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis, Toxins 6 (2014) 850–868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [82].Ashton DS, Beddell CR, Green BN, Oliver RW, Rapid validation of molecular structures of biological samples by electrospray-mass spectrometry, FEBS Lett 342 (1994) 1–6. [DOI] [PubMed] [Google Scholar]
- [83].Ho CS, Lam CWK, Chan MHM, Cheung RCK, Law LK, Lit LCW, Ng KF, Suen MWM, Tai HL, Electrospray Ionisation Mass Spectrometry: Principles and Clinical Applications, The Clinical Biochemist Reviews 24 (2003) 3–12. [PMC free article] [PubMed] [Google Scholar]
- [84].Banerjee S, Mazumdar S, Electrospray Ionization Mass Spectrometry: A Technique to Access the Information beyond the Molecular Weight of the Analyte, International Journal of Analytical Chemistry 2012 (2012) 40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [85].Souza GH, Catharino RR, Ifa DR, Eberlin MN, Hyslop S, Peptide fingerprinting of snake venoms by direct infusion nano-electrospray ionization mass spectrometry: potential use in venom identification and taxonomy, J Mass Spectrom 43 (2008) 594–599. [DOI] [PubMed] [Google Scholar]
- [86].Liu S, Zhang C, Xu YF, Yang F, Sun MZ, Electrospray ionization mass spectrometry as a critical tool for revealing new properties of snake venom phospholipase A2, Rapid Commun Mass Spectrom 23 (2009) 1158–1166. [DOI] [PubMed] [Google Scholar]
- [87].Coutinho-Neto A, Caldeira CAS, Souza GHMF, Zaqueo KD, Kayano AM, Silva RS, Zuliani JP, Soares AM, Stábeli RG, Calderon LA, ESI-MS/MS Identification of a Bradykinin-Potentiating Peptide from Amazon Bothrops atrox Snake Venom Using a Hybrid Qq-oaTOF Mass Spectrometer, Toxins 5 (2013) 327–335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [88].Perkins DN, Pappin DJ, Creasy DM, Cottrell JS, Probability-based protein identification by searching sequence databases using mass spectrometry data, Electrophoresis 20 (1999) 3551–3567. [DOI] [PubMed] [Google Scholar]
- [89].Eng JK, McCormack AL, Yates JR, An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database, Journal of the American Society for Mass Spectrometry 5 (1994) 976–989. [DOI] [PubMed] [Google Scholar]
- [90].Seidler J, Zinn N, Boehm ME, Lehmann WD, De novo sequencing of peptides by MS/MS, PROTEOMICS 10 (2010) 634–649. [DOI] [PubMed] [Google Scholar]
- [91].Keough T, Lacey MP, Youngquist RS, Derivatization procedures to facilitate de novo sequencing of lysine-terminated tryptic peptides using postsource decay matrix-assisted laser desorption/ionization mass spectrometry, Rapid Commun Mass Spectrom 14 (2000) 2348–2356. [DOI] [PubMed] [Google Scholar]
- [92].Munchbach M, Quadroni M, Miotto G, James P, Quantitation and facilitated de novo sequencing of proteins by isotopic N-terminal labeling of peptides with a fragmentation-directing moiety, Analytical chemistry 72 (2000) 4047–4057. [DOI] [PubMed] [Google Scholar]
- [93].Allmer J, Algorithms for the de novo sequencing of peptides from tandem mass spectra, Expert review of proteomics 8 (2011) 645–657. [DOI] [PubMed] [Google Scholar]
- [94].Ma B, Zhang K, Hendrie C, Liang C, Li M, Doherty-Kirby A, Lajoie G, PEAKS: powerful software for peptide de novo sequencing by tandem mass spectrometry, Rapid Commun Mass Spectrom 17 (2003) 2337–2342. [DOI] [PubMed] [Google Scholar]
- [95].Carregari VC, Dai J, Verano-Braga T, Rocha T, Ponce-Soto LA, Marangoni S, Roepstorff P, Revealing the functional structure of a new PLA2 K49 from Bothriopsis taeniata snake venom employing automatic “de novo” sequencing using CID/HCD/ETD MS/MS analyses, J Proteomics 131 (2016) 131–139. [DOI] [PubMed] [Google Scholar]
- [96].Zeng R, Xu Q, Shao XX, Wang KY, Xia QC, Characterization and analysis of a novel glycoprotein from snake venom using liquid chromatography-electrospray mass spectrometry and Edman degradation, Eur J Biochem 266 (1999) 352–358. [DOI] [PubMed] [Google Scholar]
- [97].de Morais NCG, Neves Mamede CC, Fonseca KC, de Queiroz MR, Gomes-Filho SA, Santos-Filho NA, Bordon KDCF, Beletti ME, Sampaio SV, Arantes EC, de Oliveira F, Isolation and characterization of moojenin, an acid-active, anticoagulant metalloproteinase from Bothrops moojeni venom, Toxicon 60 (2012) 1251–1258. [DOI] [PubMed] [Google Scholar]
- [98].Calvete JJ, Next-generation snake venomics: protein-locus resolution through venom proteome decomplexation, Expert review of proteomics 11 (2014) 315–329. [DOI] [PubMed] [Google Scholar]
- [99].Petras D, Heiss P, Sussmuth RD, Calvete JJ, Venom Proteomics of Indonesian King Cobra, Ophiophagus hannah: Integrating Top-Down and Bottom-Up Approaches, J Proteome Res 14 (2015) 2539–2556. [DOI] [PubMed] [Google Scholar]
- [100].Bandeira N, Clauser KR, Pevzner PA, Shotgun Protein Sequencing: Assembly of Peptide Tandem Mass Spectra from Mixtures of Modified Proteins, Molecular & Cellular Proteomics 6 (2007) 1123–1134. [DOI] [PubMed] [Google Scholar]
- [101].Gocmen B, Heiss P, Petras D, Nalbantsoy A, Sussmuth RD, Mass spectrometry guided venom profiling and bioactivity screening of the Anatolian Meadow Viper, Vipera anatolica, Toxicon 107 (2015) 163–174. [DOI] [PubMed] [Google Scholar]
- [102].Still KBM, Nandlal RSS, Slagboom J, Somsen GW, Casewell NR, Kool J, Multipurpose HTS Coagulation Analysis: Assay Development and Assessment of Coagulopathic Snake Venoms, Toxins 9 (2017) 382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [103].Mladic M, Zietek BM, Iyer JK, Hermarij P, Niessen WMA, Somsen GW, Kini RM, Kool J, At-line nanofractionation with parallel mass spectrometry and bioactivity assessment for the rapid screening of thrombin and factor Xa inhibitors in snake venoms, Toxicon 110 (2016) 79–89. [DOI] [PubMed] [Google Scholar]
- [104].Slagboom J, Mladić M, Xie C, Kazandjian TD, Vonk F, Somsen GW, Casewell NR, Kool J, High throughput screening and identification of coagulopathic snake venom proteins and peptides using nanofractionation and proteomics approaches, PLoS neglected tropical diseases 14 (2020) e0007802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [105].Pla D, Petras D, Saviola AJ, Modahl CM, Sanz L, Pérez A, Juárez E, Frietze S, Dorrestein PC, Mackessy SP, Calvete JJ, Transcriptomics-guided bottom-up and top-down venomics of neonate and adult specimens of the arboreal rear-fanged Brown Treesnake, Boiga irregularis, from Guam, Journal of proteomics 174 (2018) 71–84. [DOI] [PubMed] [Google Scholar]
- [106].Leonardi A, Sajevic T, Pungerčar J, Križaj I, Comprehensive Study of the Proteome and Transcriptome of the Venom of the Most Venomous European Viper: Discovery of a New Subclass of Ancestral Snake Venom Metalloproteinase Precursor-Derived Proteins, J Proteome Res 18 (2019) 2287–2309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [107].Sousa LF, Portes-Junior JA, Nicolau CA, Bernardoni JL, Nishiyama MY Jr., Amazonas DR, Freitas-de-Sousa LA, Mourão RH, Chalkidis HM, Valente RH, Moura-da-Silva AM, Functional proteomic analyses of Bothrops atrox venom reveals phenotypes associated with habitat variation in the Amazon, Journal of proteomics 159 (2017) 32–46. [DOI] [PubMed] [Google Scholar]
- [108].Aird SD, da Silva NJ, Qiu L, Villar-Briones A, Saddi VA, Pires de Campos Telles M, Grau ML, Mikheyev AS, Coralsnake Venomics: Analyses of Venom Gland Transcriptomes and Proteomes of Six Brazilian Taxa, Toxins (Basel) 9 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [109].Viala VL, Hildebrand D, Trusch M, Fucase TM, Sciani JM, Pimenta DC, Arni RK, Schlüter H, Betzel C, Mirtschin P, Dunstan N, Spencer PJ, Venomics of the Australian eastern brown snake (Pseudonaja textilis): Detection of new venom proteins and splicing variants, Toxicon 107 (2015) 252–265. [DOI] [PubMed] [Google Scholar]
- [110].Ainsworth S, Petras D, Engmark M, Süssmuth RD, Whiteley G, Albulescu L-O, Kazandjian TD, Wagstaff SC, Rowley P, Wüster W, Dorrestein PC, Arias AS, Gutiérrez JM, Harrison RA, Casewell NR, Calvete JJ, The medical threat of mamba envenoming in sub-Saharan Africa revealed by genus-wide analysis of venom composition, toxicity and antivenomics profiling of available antivenoms, Journal of proteomics 172 (2018) 173–189. [DOI] [PubMed] [Google Scholar]
- [111].Durban J, Sanz L, Trevisan-Silva D, Neri-Castro E, Alagón A, Calvete JJ, Integrated Venomics and Venom Gland Transcriptome Analysis of Juvenile and Adult Mexican Rattlesnakes Crotalus simus, C. tzabcan, and C. culminatus Revealed miRNA-modulated Ontogenetic Shifts, Journal of Proteome Research 16 (2017) 3370–3390. [DOI] [PubMed] [Google Scholar]
- [112].Yang ZM, Yang YE, Chen Y, Cao J, Zhang C, Liu LL, Wang ZZ, Wang XM, Wang YM, Tsai IH, Transcriptome and proteome of the highly neurotoxic venom of Gloydius intermedius, Toxicon 107 (2015) 175–186. [DOI] [PubMed] [Google Scholar]
- [113].Gonçalves-Machado L, Pla D, Sanz L, Jorge RJB, Leitão-De-Araújo M, Alves MLM, Alvares DJ, De Miranda J, Nowatzki J, de Morais-Zani K, Fernandes W, Tanaka-Azevedo AM, Fernández J, Zingali RB, Gutiérrez JM, Corrêa-Netto C, Calvete JJ, Combined venomics, venom gland transcriptomics, bioactivities, and antivenomics of two Bothrops jararaca populations from geographic isolated regions within the Brazilian Atlantic rainforest, Journal of proteomics 135 (2016) 73–89. [DOI] [PubMed] [Google Scholar]
- [114].Modahl CM, Frietze S, Mackessy SP, Transcriptome-facilitated proteomic characterization of rear-fanged snake venoms reveal abundant metalloproteinases with enhanced activity, Journal of proteomics 187 (2018) 223–234. [DOI] [PubMed] [Google Scholar]
- [115].Neri-Castro E, Sanz L, Olvera-Rodríguez A, Bénard-Valle M, Alagón A, Calvete JJ, Venomics and biochemical analysis of the black-tailed horned pitviper, Mixcoatlus melanurus, and characterization of Melanurutoxin, a novel crotoxin homolog, Journal of proteomics, Doi: 10.1016/j.jprot.2020.103865(2020) 103865. [DOI] [PubMed] [Google Scholar]
- [116].de Weille JR, Schweitz H, Maes P, Tartar A, Lazdunski M, Calciseptine, a peptide isolated from black mamba venom, is a specific blocker of the L-type calcium channel, Proceedings of the National Academy of Sciences of the United States of America, 88 (1991) 2437–2440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [117].Viala VL, Hildebrand D, Trusch M, Fucase TM, Sciani JM, Pimenta DC, Arni RK, Schlüter H, Betzel C, Mirtschin P, Dunstan N, Spencer PJ, Venomics of the Australian eastern brown snake (Pseudonaja textilis): Detection of new venom proteins and splicing variants, Toxicon 107 (2015) 252–265. [DOI] [PubMed] [Google Scholar]
- [118].Gibbs HL, Sanz L, Sovic MG, Calvete JJ, Phylogeny-based comparative analysis of venom proteome variation in a clade of rattlesnakes (Sistrurus sp.), PloS one 8 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [119].Glenn JL, Straight RC, Wolfe MC, Hardy DL, Geographical variation in Crotalus scutulatus scutulatus (Mojave rattlesnake) venom properties, Toxicon 21 (1983) 119–130. [DOI] [PubMed] [Google Scholar]
- [120].Massey DJ, Calvete JJ, Sánchez EE, Sanz L, Richards K, Curtis R, Boesen K, Venom variability and envenoming severity outcomes of the Crotalus scutulatus scutulatus (Mojave rattlesnake) from Southern Arizona, Journal of proteomics 75 (2012) 2576–2587. [DOI] [PubMed] [Google Scholar]
- [121].Lomonte B, Escolano J, Fernández J, Sanz L, Angulo Y, Gutiérrez JM, Calvete JJ, Snake Venomics and Antivenomics of the Arboreal Neotropical Pitvipers Bothriechis lateralis and Bothriechis schlegelii, Journal of Proteome Research 7 (2008) 2445–2457. [DOI] [PubMed] [Google Scholar]
- [122].Arnold C, Vipers, mambas and taipans: the escalating health crisis over snakebites, Nature 537 (2016) 26–28. [DOI] [PubMed] [Google Scholar]
- [123].Williams D, Gutierrez JM, Harrison R, Warrell DA, White J, Winkel KD, Gopalakrishnakone P, The Global Snake Bite Initiative: an antidote for snake bite, Lancet (London, England) 375 (2010) 89–91. [DOI] [PubMed] [Google Scholar]
- [124].Boldrini-França J, Corrêa-Netto C, Silva MMS, Rodrigues RS, De La Torre P, Pérez A, Soares AM, Zingali RB, Nogueira RA, Rodrigues VM, Sanz L, Calvete JJ, Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management, Journal of proteomics 73 (2010) 1758–1776. [DOI] [PubMed] [Google Scholar]
- [125].Pla D, Gutiérrez JM, Calvete JJ, Second generation snake antivenomics: Comparing immunoaffinity and immunodepletion protocols, Toxicon 60 (2012) 688–699. [DOI] [PubMed] [Google Scholar]
- [126].Calvete JJ, Arias AS, Rodríguez Y, Quesada-Bernat S, Sánchez LV, Chippaux JP, Pla D, Gutiérrez JM, Preclinical evaluation of three polyspecific antivenoms against the venom of Echis ocellatus: Neutralization of toxic activities and antivenomics, Toxicon 119 (2016) 280–288. [DOI] [PubMed] [Google Scholar]
- [127].Madrigal M, Pla D, Sanz L, Barboza E, Arroyo-Portilla C, Corrêa-Netto C, Gutiérrez JM, Alape-Girón A, Flores-Díaz M, Calvete JJ, Cross-reactivity, antivenomics, and neutralization of toxic activities of Lachesis venoms by polyspecific and monospecific antivenoms, PLOS Neglected Tropical Diseases 11 (2017) e0005793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [128].Ciscotto PHC, Rates B, Silva DAF, Richardson M, Silva LP, Andrade H, Donato MF, Cotta GA, Maria WS, Rodrigues RJ, Sanchez E, De Lima ME, Pimenta AMC, Venomic analysis and evaluation of antivenom cross-reactivity of South American Micrurus species, Journal of proteomics 74 (2011) 1810–1825. [DOI] [PubMed] [Google Scholar]
- [129].Gutiérrez JM, Tsai W-C, Pla D, Solano G, Lomonte B, Sanz L, Angulo Y, Calvete JJ, Preclinical assessment of a polyspecific antivenom against the venoms of Cerrophidion sasai, Porthidium nasutum and Porthidium ophryomegas: Insights from combined antivenomics and neutralization assays, Toxicon 64 (2013) 60–69. [DOI] [PubMed] [Google Scholar]
- [130].Lauridsen LP, Laustsen AH, Lomonte B, Gutiérrez JM, Exploring the venom of the forest cobra snake: Toxicovenomics and antivenom profiling of Naja melanoleuca, Journal of proteomics 150 (2017) 98–108. [DOI] [PubMed] [Google Scholar]
- [131].Vetter I, Davis JL, Rash LD, Anangi R, Mobli M, Alewood PF, Lewis RJ, King GF, Venomics: a new paradigm for natural products-based drug discovery, Amino acids 40 (2011) 15–28. [DOI] [PubMed] [Google Scholar]
- [132].Escoubas P, Quinton L, Nicholson GM, Venomics: unravelling the complexity of animal venoms with mass spectrometry, J Mass Spectrom 43 (2008) 279–295. [DOI] [PubMed] [Google Scholar]
- [133].Wagstaff SC, Sanz L, Juarez P, Harrison RA, Calvete JJ, Combined snake venomics and venom gland transcriptomic analysis of the ocellated carpet viper, Echis ocellatus, J Proteomics 71 (2009) 609–623. [DOI] [PubMed] [Google Scholar]
- [134].Lomonte B, Fernández J, Sanz L, Angulo Y, Sasa M, Gutiérrez JM, Calvete JJ, Venomous snakes of Costa Rica: Biological and medical implications of their venom proteomic profiles analyzed through the strategy of snake venomics, Journal of proteomics 105 (2014) 323–339. [DOI] [PubMed] [Google Scholar]
- [135].Moreda-Piñeiro A, García-Otero N, Bermejo-Barrera P, A review on preparative and semi-preparative offgel electrophoresis for multidimensional protein/peptide assessment, Analytica Chimica Acta 836 (2014) 1–17. [DOI] [PubMed] [Google Scholar]