Abstract
Background
Naples prognostic score (NPS) serves as a new prognostic index based on nutritional and inflammatory status in recent years. The aim of the current study was to explore the prognostic effect of NPS and to develop and validate a reliable nomogram based on NPS for individual cancer-specific survival (CSS) prediction in patients with resected ESCC without neoadjuvant therapy.
Methods
The clinical data for 287 (Jan. 2010 to Jun. 2012, Training sets) and 118 (Jan. 2015 to Dec 2015, Validation sets) consecutive resected ESCC cases were retrospectively analyzed. Two NPS models based on the different cut-off values of parameters were compared. Cut-off values in model 1 were derived from previous published studies, while cut-off values in model 2 were obtained in this study based on receiver operating characteristic (ROC) curves. The relationships between NPS and clinical characteristics and CSS were analyzed. The prediction model of nomogram was developed with independent prognostic factors in the training sets and was validated in the validation sets.
Results
The 5-year CSS for NPS 0, 1 and 2 were 61.9%, 34.6% and 13.4% in model 1 and 75.0%, 42.4% and 13.0% in model 2, respectively (P<0.001). Subgroup analyses revealed that NPS was also significantly associated with CSS in both model 1 and model 2 in different TNM stages. Multivariate analyses revealed that NPS was an independent prognostic marker regarding CSS in patients with resected ESCC (P<0.001). A predictive nomogram based on NPS was established and validated. The C-indexes of the nomogram in the training sets and validation sets were 0.68 and 0.72 in model 1 and 0.69 and 0.73 in model 2, respectively. These results confirmed that NPS-based nomogram was a more accurate and effective tool for predicting CSS in patients with resected ESCC.
Conclusion
The current study confirmed that NPS was still a useful independent prognostic score in patients with resected ESCC. The NPS-based nomogram was successfully developed and validated, which may contribute to individual CSS prediction for resected ESCC patients.
Keywords: Naples prognostic score, esophageal squamous cell carcinoma, neutrophil to lymphocyte ratio, lymphocyte to monocyte ratio, cancer-specific survival, prognosis
Introduction
Esophageal cancer (EC) is a common malignant tumor worldwide (1). EC is the 5th of incidence (18.85/100 000) and the 4th of mortality (14.11/100 000) in China (2). The two main types of EC are adenocarcinoma (AC) and squamous cell carcinoma (SCC), of which esophageal SCC (ESCC) accounts for more than 90% in China (3, 4). Although the exact reason of EC is unclear, it is thought to be the result of a combination of various factors, such as diet and lifestyle, demographic factors, environmental and genetic factors. Although the treatment is improved in recent years with the progress of science and technology, the prognosis in patients with EC is still poor. Therefore, the investigation of prognostic factors prior to treatment for patients with EC is more essential.
There is increasing evidence that inflammation and nutrition are associated with tumor prognosis. Studies published in recent years revealed that a range of inflammation-related or nutrition-related indicators, such as c-reactive protein (CRP), controlling nutritional status (CONUT), systemic inflammation score (SIS), platelet to lymphocyte ratio (PLR), neutrophil to lymphocyte ratio (NLR), prognostic nutritional index (PNI), albumin (ALB), Glasgow prognostic score (GPS) and lymphocyte to monocyte ratio (LMR), are associated with tumor prognosis (5–10). However, these inflammation-related and/or nutrition-related indicators mentioned above are to some extent deficient, and the results are still controversial. Therefore, an increasing number of comprehensive prognostic models, including inflammation-related and nutrition-related indicators, are urgently needed.
Naples Prognostic Score (NPS), a novel prognostic index combined with nutritional and inflammatory biomarkers, is recently proposed to predict the survival in patients with resected colorectal cancer (CRC) (11). The study including 562 patients with resected CRC revealed that the NPS, based on a composite score of ALB, LMR, total cholesterol (TC) and NLR, was a useful significant index for overall survival (OS) and disease-free survival (DFS). The result between NPS and OS was also confirmed in another study including 259 patients with metastatic CRC receiving first-line chemotherapy (12). According to the analyses with time-dependent receiver operating characteristic (ROC) curves, moreover, the study revealed that NPS was more sensitive than other conventional prognostic scores for OS prediction in patients with metastatic CRC. Since then, NPS has been further reported in patients with resected endometrial cancer, gastric cancer, early-stage lung cancer, osteosarcoma, and resected pancreatic cancer (13–17).
The prognostic effect of NPS in EC remains unclear. To the best of our knowledge, only one study including 165 ESCC patients with neoadjuvant therapy has concluded the associations between NPS and prognosis (18). However, the recent published study focused on patients with neoadjuvant therapy in small sample. It is well known that neoadjuvant therapy may affect the hematological indicators. Moreover, a reliable nomogram based on NPS for predicting survival in patients with resected ESCC was scarce. Therefore, the aim of this study was to determine the prognostic effect of NPS in patients with resected ESCC without neoadjuvant therapy. In addition, whether or not the NPS provides a better prognostic value than other conventional prognostic scores (GPS, CONUT, SIS and PNI) was also analyzed. Moreover, the prognostic effect of NPS in resected ESCC was verified by using a validation set. Finally, a predictive NPS-based nomogram with other clinical factors was established and validated in resected ESCC patients without neoadjuvant therapy.
Methods
Patient Selection
Between January 2010 and June 2012, 287 consecutive ESCC patients with radical resection in our department (Zhejiang Cancer Hospital) were retrospectively analyzed (Training set). To verify the prognostic significance of NPS and nomogram, a validation set of 118 patients with resected ESCC in our hospital from January 2015 to December 2015 was also analyzed. The present study was consistent with the declaration of Helsinki and was approved by the ethics committee of Zhejiang Cancer Hospital (No.2018-130). Patients according to the following inclusion criteria were recruited in this study: (1) patients were pathologically diagnosed with ESCC, (2) patients in stage TNM I-III with radical resection were conducted, (3) patients received no preoperative treatments, (4) patients were included without any other tumors or distant metastases, and (5) detailed clinical data were obtained within a week before surgery, including preoperative laboratory results.
Treatment and Follow-Up
In the current study, McKeown or Ivor Lewis procedure with two-field lymphadenectomy was the main surgical resection for patients with ESCC (19, 20). McKeown and Ivor Lewis are commonly used procedures of esophagectomy for surgeons because they can make adequate lymph nodes dissection. According to the poor prognostic factors, cancer metastasis or recurrence, in the current study, the adjuvant radiotherapy (45-50.4 Gy) and/or chemotherapy (based on fluoropyrimidine and cisplatin) were conducted after operation. In our hospital, patients were generally followed up every 3 months in the first two years, every 6 months for the next three years, and once a year after five years. The follow-up results were obtained from our medical records. The last follow-up for training sets and validation sets were completed in March 2018 and April 2021, respectively.
Data Analysis
According to the medical records, the main clinical data were collected and analyzed. The laboratory results were obtained within one week before operation, such as lymphocyte count, neutrophil count, monocytes count, ALB, CRP and TC. The NLR and LMR were defined as the ratio of neutrophil count to lymphocyte count and lymphocyte count to monocyte count, respectively. The NPS was composed of the following four serum indicators (ALB, TC, NLR and LMR) according to the previous study (11–18). ESCC has its own characteristics, and patients with ESCC are mostly malnourished, so the above hematological indicators may be different from other cancers. In order to better understand the application of NPS, therefore, two models (model 1 and model 2) were used to verify the prognostic value of NPS in resected ESCC. In model 1, the cut-off points for variables of ALB, TC, NLR and LMR were derived from previous published studies. In model 2, the cut-off values of above parameters in NPS were obtained in the current study based on ROC curves. Then the NPS was calculated into 3 groups (NPS 0, 1 and 2, respectively). The definitions of GPS, PNI, SIS and CONUT were according to the previous studies (7–10). The patients diagnosed with ESCC based on the 7th AJCC/UICC TNM staging system (21).
Statistical Analysis
All statistical analyses in the current study were performed by using SPSS 20.0 (SPSS Inc., Chicago, IL, USA) and Medcalc 17.6 (MedCalc Software bvba, Ostend, Belgium). The chi-squared tests or Fisher’s exact test were used to evaluate the correlations grouped by NPS. The ROC curves were used to identify the sensibility and specificity of ALB, TC, NLR and LMR. The ROC curves were also performed to explore the predictive accuracy of NPS, ALB, TC, NLR and LMR. Cut-off values in model 1 for variables of ALB, TC, NLR and LMR were derived from previous published studies. In model 2, the optimal cut-off values for above variables in NPS were selected by ROC curves. The areas under the curve (AUC) for NPS (model 1 and model 2) and its components of ALB, TC, NLR and LMR, as well as other conventional prognostic scores (GPS, SIS, CONUT and PNI) were calculated and compared. The association between CSS and prognostic factors (univariate and multivariate) was analyzed by the Cox regression analyses and Kaplan-Meier methods. Hazard ratios (HRs) with 95% confidence intervals (CIs) were also calculated according to the Cox regression analyses. The predictive nomogram was established based on independent prognostic factors in the training set in multivariate analyses and was validated in the validation set by using R 3.6.0 software (22). Calibrations of the nomogram for survival prediction with 1-, 3-, and 5-year CSS were executed by comparing the training sets and validation sets. P-values less than 0.05 were considered to be statistically significant.
Results
Patient Characteristics
The baseline characteristics of the training sets and validation sets were shown in Table 1 . In the training sets, there were 250 males and 37 females with the mean age of 59.0 ± 7.8 years (range: 36-78 years). There were 84 (29.3%) patients in TNM I stage, 94 (32.8%) patients in TNM II stage and 109 (37.9%) patients in TNM III stage, respectively. Adjuvant treatment was administered to 82 patients (28.6%). In the validation sets, there were 91 males and 27 females with the mean age of 60.2 ± 7.9 years (range: 41-78 years). There were more female patients in validation sets (22.9% vs. 12.9%, P=0.012).
Table 1.
Baseline characteristics of ESCC patients in the training and validation sets.
| Training sets (n=287) | Validation sets (n=118) | P-value | |
|---|---|---|---|
| Age (years) | 59.0 ± 7.8 | 60.2 ± 7.9 | 0.179 |
| Gender | 0.012 | ||
| Female | 37 (12.9%) | 27 (22.9%) | |
| Male | 250 (87.1%) | 91 (77.1%) | |
| Tumor length (cm) | 4.2 ± 1.8 | 4.0 ± 1.8 | 0.215 |
| Tumor location | 0.634 | ||
| Upper | 17 (5.9%) | 10 (8.4%) | |
| Middle | 132 (46.0%) | 54 (45.8%) | |
| Lower | 138 (48.1%) | 54 (45.8%) | |
| Vessel invasion | 0.496 | ||
| Negative | 246 (85.7%) | 98 (83.1%) | |
| Positive | 41 (14.3%) | 20 (16.9%) | |
| Perineural invasion | 0.197 | ||
| Negative | 230 (80.1%) | 101 (75.3%) | |
| Positive | 57 (19.9%) | 17 (24.7%) | |
| Differentiation | 0.098 | ||
| Well | 41 (14.3%) | 19 (16.1%) | |
| Moderate | 191 (66.6%) | 66 (55.9%) | |
| Poor | 55 (19.2%) | 33 (28.0%) | |
| TNM stage | 0.249 | ||
| I | 84 (29.3%) | 25 (21.2%) | |
| II | 94 (32.8%) | 43 (36.4%) | |
| III | 109 (37.9%) | 50 (42.4%) | |
| Adjuvant treatment | 0.241 | ||
| No | 205 (71.4%) | 91 (77.1%) | |
| Yes | 82 (28.6%) | 27 (22.9%) | |
| CRP (mg/L) | 7.0 ± 8.2 | 5.8 ± 8.3 | 0.171 |
| NLR | 3.0 ± 1.25 | 3.1 ± 0.82 | 0.430 |
| LMR | 4.5 ± 1.74 | 4.0 ± 1.51 | 0.008 |
| ALB (mg/dL) | 4.09 ± 0.52 | 4.08 ± 0.71 | 0.824 |
| TC (mg/dL) | 179.7 ± 40.6 | 183.2 ± 42.9 | 0.435 |
| PNI | 48.9 ± 5.76 | 47.7 ± 7.18 | 0.069 |
| GPS | 0.575 | ||
| 0 | 191 (66.6%) | 76 (64.4%) | |
| 1 | 73 (25.4%) | 35 (29.7%) | |
| 2 | 23 (8.0%) | 7 (5.9%) | |
| SIS | 0.020 | ||
| 0 | 96 (33.4%) | 23 (19.5%) | |
| 1 | 109 (38.0%) | 54 (45.8%) | |
| 2 | 82 (28.6%) | 41 (34.7%) | |
| CONUT | 0.054 | ||
| 0 | 136 (47.4%) | 43 (36.4%) | |
| 1 | 139 (48.4%) | 65 (55.1%) | |
| 2 | 12 (4.2) | 10 (8.5%) | |
| NPS (model 1) | 0.056 | ||
| 0 | 63 (22.0%) | 14 (11.9%) | |
| 1 | 127 (44.3%) | 56 (47.5%) | |
| 2 | 97 (33.7%) | 48 (40.7%) | |
| NPS (model 2) | 0.255 | ||
| 0 | 32 (11.1%) | 10 (8.5%) | |
| 1 | 132 (46.0%) | 47 (39.8%) | |
| 2 | 123 (42.9%) | 61 (51.7%) |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; CRP, C-reactive protein; NLR, neutrophil to lymphocyte ratio; LMR, lymphocyte to monocyte ratio; ALB, albumin; TC, total cholesterol; GPS, Glasgow prognostic score; TNM, tumor node metastasis; PNI, prognostic nutritional index; SIS, systemic inflammation score; CONUT, controlling nutritional status. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
Laboratory Results Analysis in the Training Sets
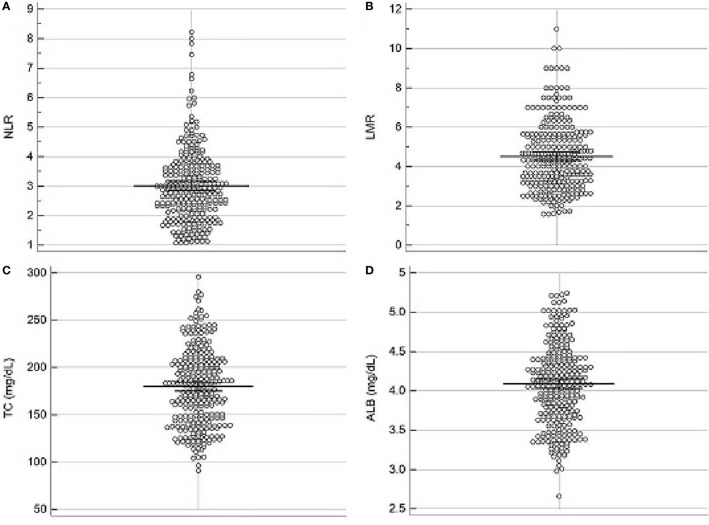
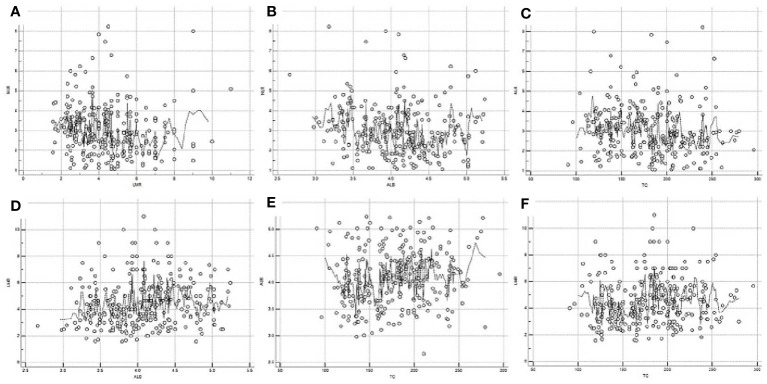
The scatter diagrams regarding NLR, LMR, ALB and TC were shown in Figure 1 . The mean values for NLR, LMR, ALB and TC were 3.0 ± 1.25, 4.5 ± 1.74, 4.1 ± 0.5 mg/dL and 180.0 ± 40.6 mg/dL, respectively. The correlation diagrams regarding NLR, LMR, ALB and TC were shown in Figure 2 . The results revealed that NLR was negatively correlated with LMR (r=-0.12, P=0.041), TC (r=-0.13, P=0.026) and ALB (r=-0.15, P=0.011), and the differences were statistically significant. In addition, positive correlations were found between LMR and ALB (r=0.16, P=0.007), TC and ALB (r=0.12, P=0.038), respectively. However, no correlations were found between TC and LMR (r=0.12, P=0.052).
Figure 1.
Scatter diagrams regarding NLR (A), LMR (B), TC (C) and ALB (D). The mean values for NLR, LMR, ALB and TC were 3.0 ± 1.25, 4.5 ± 1.74, 4.1 ± 0.5 mg/dL and 180.0 ± 40.6 mg/dL, respectively.
Figure 2.
Correlation diagrams of NLR, LMR, ALB and TC. Negative correlations between NLR and LMR (r=-0.12, P=0.041, A), NLR and ALB (r=-0.15, P=0.011, B), NLR and TC (r=-0.13, P=0.026, C), respectively. Positive correlations between LMR and ALB (r=0.16, P=0.007, D), TC and ALB (r=0.12, P=0.038, E), respectively. No correlations between LMR and TC (r=0.12, P=0.052, F).
NPS Analysis in the Training Sets
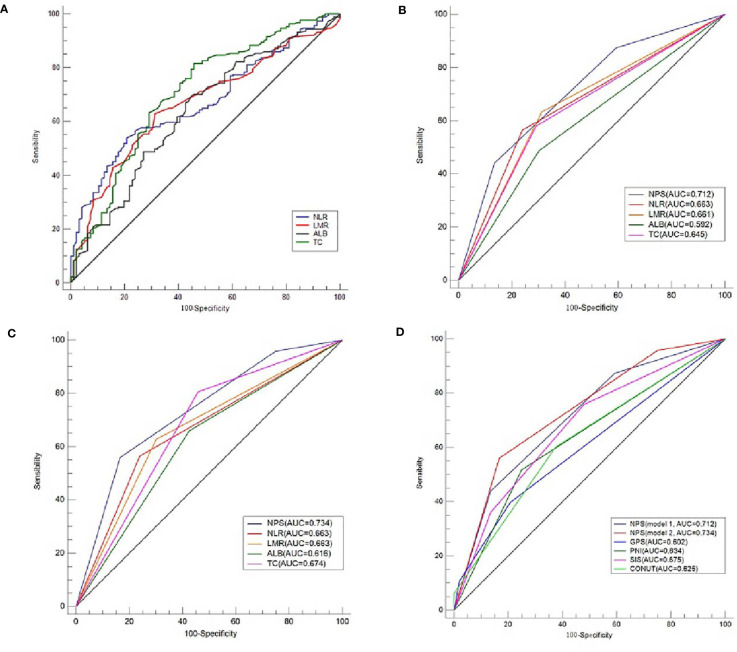
According to the previous published studies, the cut-off points for serum ALB, TC, NLR and LMR were 4.0 mg/dL, 180 mg/dL, 2.96 and 4.44, respectively. According to the ROC curves in the current study, the optimal cut-off points for serum ALB, TC, NLR and LMR were 4.2 mg/dL, 202 mg/dL, 2.97 and 4.40, respectively ( Figure 3A ). In order to better understand the application of NPS, therefore, two models (model 1 and model 2) were used to verify the prognostic value of NPS in resected ESCC. The definition of NPS based on serum ALB, NLR, TC and LMR was shown in Table 2 . The NPS was calculated into 3 groups (NPS 0, 1 and 2, respectively). The sensibilities and specificities of serum ALB, TC, NLR and LMR were identified by ROC curves ( Table 3 ). The baseline characteristics grouped by NPS in both model 1 and model 2 were shown in Table 4 .
Figure 3.
ROC analyses regarding cut-off values and AUC comparison. The optimal cut-off points for serum ALB, TC, NLR and LMR according to the ROC curves (A). The optimal cut-off points for serum ALB, TC, NLR and LMR were 4.2 mg/dL, 202 mg/dL, 2.97 and 4.40, respectively. AUC comparisons between NPS and variables of ALB, TC, NLR and LMR in model 1 (B) and model 2 (C). AUC comparisons between NPS and other conventional prognostic scores of GPS, SIS, CONUT and PNI (D).
Table 2.
Calculation of Naples prognostic score (NPS).
| Variables | Model 1 | Model 2 | Points | NPS group |
|---|---|---|---|---|
| Cut-off value | Cut-off value | |||
| ALB (mg/dL) | ≥ 4.0 | ≥ 4.2 | 0 | NPS 1: 0 point |
| < 4.0 | < 4.2 | 1 | NPS 2: 1 or 2 points | |
| TC (mg/dL) | > 180 | > 202 | 0 | NPS 3: 3 or 4 points |
| ≤ 180 | ≤ 202 | 1 | ||
| NLR | ≤ 2.96 | ≤ 2.97 | 0 | |
| > 2.96 | > 2.97 | 1 | ||
| LMR | > 4.44 | > 4.40 | 0 | |
| ≤ 4.44 | ≤ 4.40 | 1 |
NPS, Naples prognostic score; NLR, neutrophil to lymphocyte ratio; LMR, lymphocyte to monocyte ratio; ALB, albumin; TC, total cholesterol. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
Table 3.
Comparison of sensibility and specificity for the variables of NPS in ESCC.
| Variables | Cut-off value | Sensibility | Specificity | |
|---|---|---|---|---|
| Model 1 | ALB (mg/dL) | 4.0 | 69.8% | 49.7% |
| TC (mg/dL) | 180 | 70.8% | 58.1% | |
| NLR | 2.96 | 56.5% | 76.0% | |
| LMR | 4.44 | 67.7% | 63.4% | |
| Model 2 | ALB (mg/dL) | 4.2 | 57.3% | 67.0% |
| TC (mg/dL) | 202 | 54.2% | 81.7% | |
| NLR | 2.97 | 56.5% | 76.0% | |
| LMR | 4.40 | 68.8% | 62.8% |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; NLR, neutrophil to lymphocyte ratio; LMR, lymphocyte to monocyte ratio; ALB, albumin; TC, total cholesterol. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
Table 4.
Comparison of AUC areas between NPS and other markers in ESCC.
| AUC | 95% CI | P-value | ||
|---|---|---|---|---|
| Model 1 | NPS | 0.712 | 0.656-0.763 | Reference |
| NLR | 0.663 | 0.605-0.717 | 0.0603 | |
| LMR | 0.661 | 0.603-0.715 | 0.0631 | |
| ALB | 0.592 | 0.533-0.650 | 0.0001 | |
| TC | 0.645 | 0.586-0.700 | 0.0077 | |
| GPS | 0.602 | 0.543-0.659 | 0.0005 | |
| PNI | 0.634 | 0.576-0.690 | 0.0038 | |
| SIS | 0.675 | 0.618-0.729 | 0.0799 | |
| CONUT | 0.625 | 0.567-0.682 | 0.0020 | |
| Model 2 | NPS | 0.734 | 0.679-0.784 | Reference |
| NLR | 0.663 | 0.605-0.717 | 0.0055 | |
| LMR | 0.663 | 0.605-0.718 | 0.0135 | |
| ALB | 0.616 | 0.557-0.673 | 0.0001 | |
| TC | 0.674 | 0.616-0.728 | 0.0367 | |
| GPS | 0.602 | 0.543-0.659 | 0.0001 | |
| PNI | 0.634 | 0.576-0.690 | 0.0003 | |
| SIS | 0.675 | 0.618-0.729 | 0.0220 | |
| CONUT | 0.625 | 0.567-0.682 | 0.0003 | |
| Model 1 vs. Model 2 | NPS (model 1) | 0.712 | 0.656-0.763 | Reference |
| NPS (model 2) | 0.734 | 0.679-0.784 | 0.1689 |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; CRP, C-reactive protein; NLR, neutrophil to lymphocyte ratio; LMR, lymphocyte to monocyte ratio; ALB, albumin; TC, total cholesterol; GPS, Glasgow prognostic score; TNM, tumor node metastasis; PNI, prognostic nutritional index; SIS, systemic inflammation score; CONUT, controlling nutritional status. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
ROC Analysis Regarding AUC Comparison in the Training Sets
The ROC curves regarding categorical variables for NPS and its components of ALB, TC, NLR and LMR, as well as other conventional prognostic scores (GPS, SIS, CONUT and PNI) were shown in Figures 3B–D . Compared with its components (ALB, TC, NLR and LMR) and other conventional prognostic scores (GPS, SIS, CONUT and PNI), NPS had the largest AUC (both in model 1 and model 2) according to the ROC curves ( Table 5 ). Although the AUC of NPS in model 2 (0.734) was greater than that of NPS in model 1 (0.712), there was no statistical difference between model 1 and model 2.
Table 5.
Comparison of baseline characteristics of ESCC patients based on NPS in training sets.
| NPS Model 1 | NPS Model 2 | |||||||
|---|---|---|---|---|---|---|---|---|
| 0(n=97) | 1(n=127) | 2(n=97) | P-value | 0(n=32) | 1(n=132) | 2(n=123) | P-value | |
| Age (years) | 0.494 | 0.779 | ||||||
| ≤ 60 | 33(52.4) | 77(60.6) | 59(60.8) | 17(53.1) | 79(59.8) | 73(59.3) | ||
| > 60 | 30(47.6) | 50(39.4) | 38(39.2) | 15(46.9) | 53(40.2) | 50(40.7) | ||
| Gender | 0.078 | 0.193 | ||||||
| Female | 12(19.0) | 18(14.2) | 7(12.9) | 6(18.8) | 20(15.2) | 11(8.9) | ||
| Male | 51(81.0) | 109(85.8) | 90(87.1) | 26(81.2) | 112(84.8) | 112(91.1) | ||
| Tumor length (cm) | <0.001 | 0.007 | ||||||
| ≤ 3.0 | 30(47.6) | 40(31.5) | 15(15.5) | 14(43.8) | 46(34.8) | 25(20.3) | ||
| > 3.0 | 33(52.4) | 87(68.5) | 82(84.5) | 18(56.2) | 86(65.2) | 98(79.7) | ||
| Tumor location | 0.491 | 0.314 | ||||||
| Upper | 1(1.6) | 10(7.9) | 6(6.2) | 0(0.0) | 11(8.3) | 6(4.9) | ||
| Middle | 31(49.2) | 59(46.5) | 42(43.3) | 17(53.1) | 62(47.0) | 53(43.1) | ||
| Lower | 31(49.2) | 58(45.7) | 49(50.5) | 15(46.9) | 59(44.7) | 64(52.0) | ||
| Vessel invasion | 0.562 | 0.966 | ||||||
| Negative | 53(84.1) | 112(88.2) | 81(83.5) | 27(84.4) | 113(85.6) | 106(86.2) | ||
| Positive | 10(15.9) | 15(11.8) | 16(16.5) | 5(15.6) | 19(14.4) | 17(13.8) | ||
| Perineural invasion | 0.024 | 0.288 | ||||||
| Negative | 58(92.1) | 99(78.0) | 73(75.3) | 29(90.6) | 104(78.8) | 97(78.9) | ||
| Positive | 5(7.9) | 28(22.0) | 24(24.7) | 3(9.4) | 28(21.2) | 26(21.1) | ||
| Differentiation | 0.465 | 0.546 | ||||||
| Well | 9(14.3) | 18(14.2) | 14(14.4) | 4(12.5) | 17(12.9) | 20(16.3) | ||
| Moderate | 42(66.7) | 90(70.9) | 59(60.8) | 23(71.9) | 93(70.5) | 75(61.0) | ||
| Poor | 12(19.0) | 19(14.9) | 24(24.8) | 5(15.6) | 22(16.6) | 28(22.8) | ||
| TNM stage | <0.001 | 0.002 | ||||||
| I | 29(46.0) | 40(31.5) | 15(15.5) | 12(37.5) | 49(37.1) | 23(18.7) | ||
| II | 23(36.5) | 40(31.5) | 31(32.0) | 13(40.6) | 42(31.8) | 39(31.7) | ||
| III | 11(17.5) | 47(37.0) | 51(52.5) | 7(21.9) | 41(31.1) | 61(49.6) | ||
| Adjuvant treatment | 0.921 | 0.408 | ||||||
| No | 46(73.0) | 91(71.7) | 68(70.1) | 20(62.5) | 98(74.2) | 87(70.7) | ||
| Yes | 17(27.0) | 36(28.3) | 29(29.9) | 12(37.5) | 34(25.8) | 36(29.3) | ||
| CRP (mg/L) | <0.001 | <0.001 | ||||||
| ≤ 10.0 | 58(92.1) | 107(84.3) | 50(51.5) | 27(84.4) | 115(87.1) | 73(59.3) | ||
| > 10.0 | 5(7.9) | 20(15.7) | 47(48.5) | 5(15.6) | 17(12.9) | 50(40.7) | ||
| GPS | <0.001 | <0.001 | ||||||
| 0 | 58(92.1) | 95(74.8) | 38(39.2) | 27(84.4) | 104(78.8) | 60(48.8) | ||
| 1 | 5(7.9) | 30(23.6) | 38(39.2) | 5(15.6) | 27(20.5) | 41(33.3) | ||
| 2 | 0(0.0) | 2(1.6) | 21(21.6) | 0(0.0) | 1(0.8) | 22(17.9) | ||
| PNI | <0.001 | <0.001 | ||||||
| ≤ 47.5 | 1(1.6) | 44(34.6) | 78(80.4) | 0(0.0) | 31(23.5) | 92(74.8) | ||
| > 47.5 | 62(98.4) | 83(65.4) | 19(19.6) | 32(100) | 101(76.5) | 31(25.2) | ||
| SIS | <0.001 | <0.001 | ||||||
| 0 | 63(100) | 33(26.0) | 0(0.0) | 32(100) | 57(43.2) | 7(5.7) | ||
| 1 | 0(0.0) | 77(60.6) | 32(33.0) | 0(0.0) | 63(47.7) | 46(37.4) | ||
| 2 | 0(0.0) | 17(13.4) | 65(67.0) | 0(0.0) | 12(9.1) | 70(56.9) | ||
| CONUT | <0.001 | <0.001 | ||||||
| 0 | 59(93.7) | 60(47.2) | 17(17.5) | 31(96.9) | 79(59.8) | 26(21.1) | ||
| 1 | 4(6.3) | 67(52.8) | 68(70.1) | 1(3.1) | 53(40.2) | 85(69.1) | ||
| 2 | 0(0.0) | 0(0.0) | 12(12.4) | 0(0.0) | 0(0.0) | 12(9.8) | ||
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; CRP, C-reactive protein; GPS, Glasgow prognostic score; TNM, tumor node metastasis; PNI, prognostic nutritional index; SIS, systemic inflammation score; CONUT, controlling nutritional status. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
CSS Analysis in the Training Sets
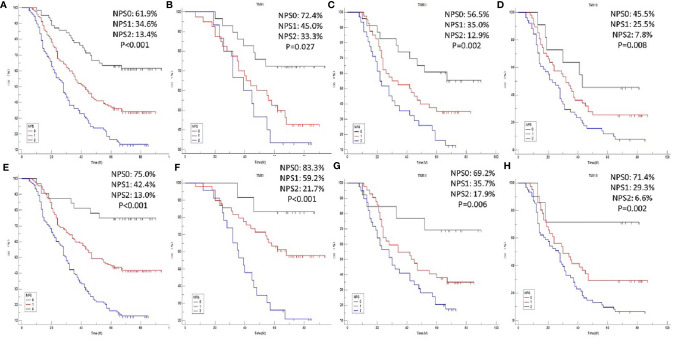
The 5-year CSS for NPS 0, 1 and 2 were 61.9%, 34.6% and 13.4% in model 1 ( Figure 4A ) and 75.0%, 42.4% and 13.0% in model 2 ( Figure 4E ), respectively (P<0.001). To better understand the prognostic significance of NPS in different TNM stages, subgroup analyses in model 1 ( Figures 4B–D ) and model 2 ( Figures 4F–H ) were also performed. In both model 1 and model 2, significant correlations between NPS and CSS were shown in different TNM stages.
Figure 4.
CSS analyses. Kaplan-Meier for CSS grouped by NPS in model 1 (A) and model 2 (E). CSS analyses for NPS in subgroup analyses based on TNM stage in model 1 [TNM I: P=0.027, (B) TNM II: P=0.002, (C) TNM III: P=0.008, (D)] and model 2 [TNM I: P<0.001, (F) TNM II: P=0.006, (G) TNM III: P=0.002, (H)], respectively.
Cox Analyses With Univariate and Multivariate Analyses in the Training Sets
Univariate analyses were used to explore the significantly clinical variables associated with CSS ( Table 6 ). Significant prognostic variables then were recruited in multivariate analyses. The results revealed that NPS, TNM and CRP were independent prognostic markers regarding CSS. Compared with NPS 0, patients in NPS 1 or 2 had worse 5-year CSS in model 1 (NPS 1 vs. NPS 0: HR=1.978, 95% CI: 1.250-3.130, P=0.004; NPS 2 vs. NPS 0: HR=2.903, 95% CI: 1.803-4.675, P<0.001) and model 2 (NPS 1 vs. NPS 0: HR=3.072, 95% CI: 1.480-6.380, P=0.003; NPS 2 vs. NPS 0: HR=5.239, 95% CI: 2.536-10.825, P<0.001), respectively ( Table 7 ).
Table 6.
Univariate Cox analyses of CSS in training sets.
| HR (95% CI) | P-value | |
|---|---|---|
| Age (years) | 0.925 | |
| ≤ 60 | 1.000 | |
| > 60 | 1.014 (0.760-1.353) | |
| Gender | 0.704 | |
| Female | 1.000 | |
| Male | 1.088 (0.704-1.682) | |
| Tumor length (cm) | 0.135 | |
| ≤ 3.0 | 1.000 | |
| > 3.0 | 1.272 (0.928-1.743) | |
| Tumor location | 0.764 | |
| Upper | 1.000 | |
| Middle | 1.277 (0.664-2.456) | 0.464 |
| Lower | 1.252 (0.652-2.406) | 0.499 |
| Vessel invasion | 0.026 | |
| Negative | 1.000 | |
| Positive | 1.540 (1.053-2.252) | |
| Perineural invasion | 0.007 | |
| Negative | 1.000 | |
| Positive | 1.585 (1.134-2.216) | |
| Differentiation | 0.407 | |
| Well | 1.000 | |
| Moderate | 1.156 (0.753-1.774) | 0.508 |
| Poor | 1.394 (0.842-2.310) | 0.197 |
| TNM stage | <0.001 | |
| I | 1.000 | |
| II | 1.830 (1.230-2.721) | 0.003 |
| III | 2.874 (1.973-4.186) | <0.001 |
| Adjuvant treatment | 0.480 | |
| No | 1.000 | |
| Yes | 1.119 (0.818-1.531) | |
| CRP (mg/L) | <0.001 | |
| ≤ 10.0 | 1.000 | |
| > 10.0 | 2.126 (1.559-2.901) | |
| NLR (model 1) | <0.001 | |
| ≤ 2.96 | 1.000 | |
| > 2.96 | 2.235 (1.676-2.982) | |
| NLR (model 2) | <0.001 | |
| ≤ 2.97 | 1.000 | |
| > 2.97 | 2.235 (1.676-2.982) | |
| LMR (model 1) | <0.001 | |
| > 4.44 | 1.000 | |
| ≤ 4.44 | 2.174 (1.616-2.923) | |
| LMR (model 2) | <0.001 | |
| > 4.40 | 1.000 | |
| ≤ 4.40 | 2.198 (1.636-2.953) | |
| ALB (mg/dL, model 1) | 0.001 | |
| ≥ 4.0 | 1.000 | |
| < 4.0 | 1.615 (1.215-2.147) | |
| ALB (mg/dL, model 2) | <0.001 | |
| ≥ 4.2 | 1.000 | |
| < 4.2 | 1.806 (1.337-2.438) | |
| TC (mg/dL, model 1) | <0.001 | |
| > 180 | 1.000 | |
| ≤ 180 | 1.881 (1.409-2.511) | |
| TC (mg/dL, model 1) | ||
| > 202 | 1.000 | <0.001 |
| ≤ 202 | 2.586 (1.802-3.711) | |
| NPS (model 1) | <0.001 | |
| 0 | 1.000 | |
| 1 | 2.230 (1.415-3.515) | 0.001 |
| 2 | 4.006 (2.537-6.323) | <0.001 |
| NPS (model 2) | <0.001 | |
| 0 | 1.000 | |
| 1 | 2.937 (1.417-6.090) | 0.004 |
| 2 | 6.313 (3.069-12.990) | <0.001 |
| GPS | <0.001 | |
| 0 | 1.000 | |
| 1 | 1.897 (1.374-2.619) | <0.001 |
| 2 | 3.283 (2.047-5.267) | <0.001 |
| PNI | <0.001 | |
| ≤ 47.5 | 1.000 | |
| > 47.5 | 0.527 (0.396-0.701) | |
| SIS | <0.001 | |
| 0 | 1.000 | |
| 1 | 1.800 (1.248-2.598) | 0.002 |
| 2 | 2.790 (1.916-4.064) | <0.001 |
| CONUT | <0.001 | |
| 0 | 1.000 | |
| 1 | 1.474 (1.095-1.985) | 0.010 |
| 2 | 4.127 (2.226-7.651) | <0.001 |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; CRP, C-reactive protein; NLR, neutrophil to lymphocyte ratio; LMR, lymphocyte to monocyte ratio; ALB, albumin; TC, total cholesterol; GPS, Glasgow prognostic score; PNI, prognostic nutritional index; SIS, systemic inflammation score; CONUT, controlling nutritional status; TNM, tumor node metastasis; CI, confidence interval; HR, hazard ratio; CSS, cancer-specific survival. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
Table 7.
Multivariate analyses regarding CSS in patients with ESCC.
| HR (95% CI) | P-value | ||
|---|---|---|---|
| Model 1 | CRP (mg/L, > 10.0 vs. ≤ 10.0) | 1.532 (1.102-2.128) | 0.011 |
| NPS (model 1) | |||
| 1 vs. 0 | 1.978 (1.250-3.130) | 0.004 | |
| 2 vs. 0 | 2.903 (1.803-4.675) | <0.001 | |
| TNM stage | |||
| II vs. I | 1.620 (1.087-2.416) | 0.018 | |
| III vs. I | 2.196 (1.494-3.227) | <0.001 | |
| Model 2 | CRP (mg/L, > 10.0 vs. ≤ 10.0) | 1.618 (1.171-2.235) | 0.004 |
| TNM stage | |||
| II vs. I | 1.643 (1.101-2.451) | 0.015 | |
| III vs. I | 2.269 (1.544-3.334) | <0.001 | |
| NPS (model 2) | |||
| 1 vs. 0 | 3.072 (1.480-6.380) | 0.003 | |
| 2 vs. 0 | 5.239 (2.536-10.825) | <0.001 |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; CRP, C-reactive protein; TNM, tumor node metastasis; CI, confidence interval; HR, hazard ratio; CSS, cancer-specific survival. Model 1: The cut-off values according to the previous published studies. Model 2: The cut-off values according to the ROC curves in the current study.
Nomogram Development and Validation
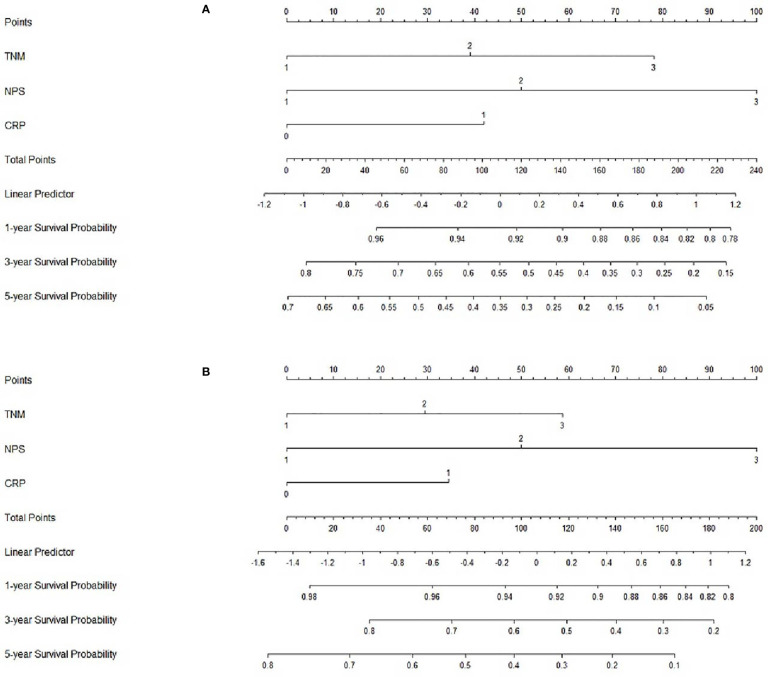
Based on the analyses of prognostic factors in multivariate analyses, three variables (NPS, TNM and CRP) were selected to develop a nomogram for predicting 1-, 3- and 5-year CSS in resected ESCC patients. The predictive nomogram based on NPS in model 1 and model 2 was established in Figure 5 . The C-indexes of the nomograms in the training sets and validation sets were 0.68 and 0.72 in model 1 and 0.69 and 0.73 in model 2, respectively. Regarding the individual 1-, 3- and 5-year CSS prediction, the calibration curves presented an acceptable agreement between the training sets and validation sets ( Figure 6 ). The results in the present study confirmed the NPS-based nomogram as a more accurate and effective tool for survival prediction with 1-, 3- and 5-year CSS in patients with resected ESCC.
Figure 5.
Nomogram analyses. Nomogram including NPS, TNM and CRP for predicting the 1-, 3- and 5-year CSS in patients with resected ESCC in model 1 (A) and model 2 (B).
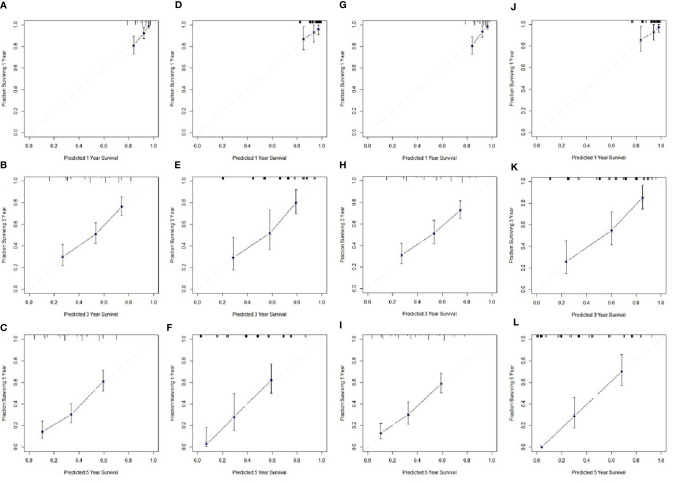
Figure 6.
Calibration curves of the nomogram. Calibration curves presented an acceptable agreement between the training sets and validation sets. Calibration curves for CSS for 1-year, 3-year and 5-year survival nomogram calibration curves in training set (A–C) and validation sets (D–F) in model 1. Calibration curves for CSS for 1-year, 3-year and 5-year survival nomogram calibration curves in training set (G–I) and validation sets (J–L) in model 2.
Discussion
The present study confirmed the prognostic effect of the NPS, and its prognostic effect was significantly greater than other conventional prognostic scores. Compared with patients in NPS 0 group, the present study also revealed that patients in group NPS 1-2 had worse CSS. Multivariate analyses revealed that NPS, TNM stage and CRP were independent prognostic markers regarding CSS. Moreover, we firstly established and validated a new prognostic nomogram based on NPS and other independent prognostic factors. The results revealed that the NPS-based nomogram was a more accurate and effective tool for survival prediction with 1-, 3- and 5-year CSS in patients with resected ESCC.
The NPS, combined with serum TC, ALB, NLR and LMR, was initially reported by Galizia et al. in 2017 in patients undergoing surgery with CRC (11). The study including 562 patients with resected CRC revealed that the NPS was an independent significant predictor of OS and DFS. The NPS was also confirmed in another study including 259 patients with metastatic CRC receiving first-line systemic chemotherapy (12). Moreover, the results revealed that NPS was more sensitive than other conventional prognostic scores for OS prediction in patients with metastatic CRC according to the time-dependent ROC analysis. Since then, NPS has been further reported in patients with various cancers (13–17). We summarized published articles regarding the association between NPS and prognosis in cancers ( Table 8 ).
Table 8.
Comparison of NPS in published studies.
| Study | Year | Country | Tumor types | Sample | Survival |
|---|---|---|---|---|---|
| Galizia | (11) | Italy | colorectal cancer | 562 | OS,DFS |
| Miyamoto | (12) | Japan | colorectal cancer | 259 | OS |
| Galizia | (16) | Italy | gastric cancer | 415 | OS,DFS |
| Li | (13) | China | endometrial cancer | 1038 | OS,PFS |
| Yang | (15) | China | osteosarcoma | 133 | OS,PFS |
| Nakagawa | (17) | Japan | pancreatic cancer | 196 | OS |
| Li | (14) | China | lung Cancer | 457 | OS,DFS |
| Kano | (18) | Japan | ESCC | 165 | RFS, OS |
| Current | 2021 | China | ESCC | 287 | CSS |
ESCC, esophageal squamous cell carcinoma; NPS, Naples prognostic score; OS, overall survival; DFS, disease-free survival; PFS, progression-free survival; RFS, recurrence-free survival; CSS, cancer-specific survival.
It is well known that the components of NPS (TC, ALB, NLR and LMR) are common clinical biomarkers in daily clinical practice. The prognostic effect of NPS in EC remains unclear. Recently, Kano et al. (18) analyzed the associations between NPS and prognosis in locally advanced ESCC with neoadjuvant therapy followed by surgery. There were some differences between the Kano’s study and the current study. Firstly, the Kano’s study focused on patients with neoadjuvant therapy while the patients in the present study were recruited without any neoadjuvant therapy. Secondly, the samples in the current study were larger than that of the Kano’s study, and the results in the current study were established in the training sets and validated in the validation sets, respectively. Thirdly, the Kano’s study did not add other conventional prognostic scores in univariate and multivariate analyses. Although the NPS had the largest AUC according to ROC curves, the prognostic significance of NPS as an independent prognostic factor in univariate and multivariate analyses should be regard with caution. Fourthly, ESCC has its own characteristics, and patients with ESCC are mostly malnourished, so the above hematological indicators may be different from other cancers. In the current study, therefore, the cut-off values of parameters in NPS based on previous studies and current study were compared. Last but not least, a predictive nomogram based on NPS and other clinical factors was established and validated in the current study for survival prediction in resected ESCC patients without neoadjuvant therapy. Therefore, we performed this study to explore the prognostic effect of NPS and develop a nomogram based on NPS for survival prediction with 1-, 3- and 5-year CSS in resected ESCC patients without neoadjuvant therapy. Results from our research provided new insights regarding prognostic significance of NPS in the field of resected ESCC.
There is increasing evidence that inflammation and nutrition are associated with tumor prognosis. Studies published in recent years reported that a range of inflammation-related and/or nutrition-related indicators, such as CONUT, SIS, GPS and PNI, are associated with tumor prognosis (7–10). Compared with SIS, CONUT and PNI, NPS was considered to be the highest scoring system for grouping patients with the same OS and DFS survival rate (11). Yang et al. (15) also compared the predictive results among different independent prognostic factors through the time-dependent ROC analyses. The results demonstrated that NPS obtained the highest AUC. The above conventional prognostic scores were still controversial in patients with EC. In the current study, compared with its components (ALB, TC, NLR and LMR) and other conventional prognostic scores (GPS, SIS, CONUT and PNI), NPS had the largest AUC (both in model 1 and model 2) according to the ROC curves. GPS, SIS, CONUT and PNI were also included in our study for Cox multivariate analyses. Compared with NPS, however, the results revealed that these conventional prognostic scores were not independent significant prognostic factors.
Recently, more and more studies have revealed that nomogram based on nutritional and inflammatory status can better predict prognosis of various cancers (23–25). The nomogram can incorporate multiple factors into the prediction and consider the weight of each variable, making predictive nomogram more accurate and practical. Nomogram can also develop risk stratification and help clinicians to perform suitable treatments and survival predictions. In the current study, all three variables included in the nomogram could be obtained easily, which facilitates the application of this nomogram in clinical practice. Therefore, clinicians could use this nomogram to predict 1-, 3- and 5- year CSS rates of resected ESCC patients.
Previous published study by Galizia et al. (11) reported that NPS may have important clinical implications. They believed that improvement of inflammation and malnutrition can improve patient prognosis and prevent postoperative complications. The prognostic effect of NPS was also suitable in the present study in the clinical practice in resected ESCC patients without neoadjuvant therapy. Compared with patients in NPS 0 group, the present study revealed that patients in group NPS 1-2 had worse CSS. If patients with status of NPS 2-3, therefore, it is suggested to improve the status of inflammation and/or malnutrition before radical resection, or to conduct adjuvant therapy after surgery.
Some limitations should be mentioned in this study. Firstly, this was a retrospective study in a single-center. Secondly, the levels of serum markers such as ALB, NLR, LMR and TC may be influenced by various conditions, the applications of NPS based on these variables should be regarded with caution. Thirdly, patients treated without any preoperative therapy in the present study, which may have influenced results. Finally, the training sets and validation sets were from the same center, which may affect the generalizability of the findings in this study. Despite these limitations, the NPS-based nomogram in the present study was still an accurate and effective tool to perform survival prediction (CSS) in resected ESCC patients.
Conclusion
In summary, the present study determined the relationships between NPS and CSS in resected ESCC patients without neoadjuvant therapy. The results indicated that NPS is still a useful prognostic score in resected ESCC patients. A new prognostic predictive model based on NPS was successfully developed and validated, which may contribute to 1-, 3- and 5-year survival prediction for resected ESCC patients.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/ Supplementary Material .
Ethics Statement
The present study was approved by the ethics committee of Zhejiang Cancer hospital and was consistent with the declaration of Helsinki. The patients/participants provided their written informed consent to participate in this study.
Author Contributions
J-FF, J-MZ, and Q-XC contributed to the study design. J-FF, J-MZ, SC, and Q-XC were responsible for interpretation of the results. J-MZ, CS, and J-FF contributed to statistical analysis. J-FF, J-MZ, and Q-XC were prepared for the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This study was supported by grants from Zhejiang Medical and Health Science and Technology Project (2018KY290, 2019RC129). This study was also supported by Zhejiang TCM Science and Technology Project (2021ZB034).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.652537/full#supplementary-material
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
- 1. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2015. CA: Cancer J Clin (2015) 65(1):5–29. 10.3322/caac.21254 [DOI] [PubMed] [Google Scholar]
- 2. Chen W, Sun K, Zheng R, Zeng H, Zhang S, Xia C, et al. Cancer Incidence and Mortality in China, 2014. Chin J Cancer Res (2018) 30(1):1–12. 10.21147/j.issn.1000-9604.2018.01.01 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Lin Y, Totsuka Y, He Y, Kikuchi S, Qiao Y, Ueda J, et al. Epidemiology of Esophageal Cancer in Japan and China. J Epidemiol (2013) 23(3):233–42. 10.2188/jea.JE20120162 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer Statistics in China, 2015. CA: Cancer J Clin (2016) 66(2):115–32. 10.3322/caac.21338 [DOI] [PubMed] [Google Scholar]
- 5. Feng JF, Sheng C, Zhao Q, Chen P. Prognostic Value of Mean Platelet Volume/Platelet Count Ratio in Patients With Resectable Esophageal Squamous Cell Carcinoma: A Retrospective Study. PeerJ (2019) 7:e7246. 10.7717/peerj.7246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Sun Y, Zhang L. The Clinical Use of Pretreatment NLR, PLR, and LMR in Patients With Esophageal Squamous Cell Carcinoma: Evidence From a Meta-Analysis. Cancer Manag Res (2018) 10:6167–79. 10.2147/CMAR.S171035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Castillo-Martínez L, Castro-Eguiluz D, Copca-Mendoza ET, Pérez-Camargo DA, Reyes-Torres CA, Ávila EA, et al. Nutritional Assessment Tools for the Identification of Malnutrition and Nutritional Risk Associated With Cancer Treatment. Rev Invest Clin (2018) 70(3):121–5. 10.24875/RIC.18002524 [DOI] [PubMed] [Google Scholar]
- 8. Yılmaz A, Tekin SB, Bilici M, Yılmaz H. The Significance of Controlling Nutritional Status (CONUT) Score as a Novel Prognostic Parameter in Small Cell Lung Cancer. Lung (2020) 198(4):695–704. 10.1007/s00408-020-00361-2 [DOI] [PubMed] [Google Scholar]
- 9. McMillan DC. The Systemic Inflammation-Based Glasgow Prognostic Score: A Decade of Experience in Patients With Cancer. Cancer Treat Rev (2013) 39(5):534–40. 10.1016/j.ctrv.2012.08.003 [DOI] [PubMed] [Google Scholar]
- 10. Hu Y, Shen J, Liu R, Feng Z, Zhang C, Ling L, et al. Prognostic Value of Pretreatment Prognostic Nutritional Index in Non-Small Cell Lung Cancer: A Systematic Review and Meta-Analysis. Int J Biol Markers (2018) 33(4):372–8. 10.1177/1724600818799876 [DOI] [PubMed] [Google Scholar]
- 11. Galizia G, Lieto E, Auricchio A, Cardella F, Mabilia A, Podzemny V, et al. Naples Prognostic Score, Based on Nutritional and Inflammatory Status, is an Independent Predictor of Long-Term Outcome in Patients Undergoing Surgery for Colorectal Cancer. Dis Colon Rectum (2017) 60(12):1273–84. 10.1097/DCR.0000000000000961 [DOI] [PubMed] [Google Scholar]
- 12. Miyamoto Y, Hiyoshi Y, Daitoku N, Okadome K, Sakamoto Y, Yamashita K, et al. Naples Prognostic Score is a Useful Prognostic Marker in Patients With Metastatic Colorectal Cancer. Dis Colon Rectum (2019) 62(12):1485–93. 10.1097/DCR.0000000000001484 [DOI] [PubMed] [Google Scholar]
- 13. Li Q, Cong R, Wang Y, Kong F, Ma J, Wu Q, et al. Naples Prognostic Score is an Independent Prognostic Factor in Patients With Operable Endometrial Cancer: Results From a Retrospective Cohort Study. Gynecol Oncol (2020) 160(1):91–8. 10.1016/j.ygyno.2020.10.013 [DOI] [PubMed] [Google Scholar]
- 14. Li S, Wang H, Yang Z, Zhao L, Lv W, Du H, et al. Naples Prognostic Score as a Novel Prognostic Prediction Tool in Video-Assisted Thoracoscopic Surgery for Early-Stage Lung Cancer: A Propensity Score Matching Study. Surg Endosc (2020). 10.1007/s00464-020-07851-7 [DOI] [PubMed] [Google Scholar]
- 15. Yang Q, Chen T, Yao Z, Zhang X. Prognostic Value of Pre-Treatment Naples Prognostic Score (NPS) in Patients With Osteosarcoma. World J Surg Oncol (2020) 18(1):24. 10.1186/s12957-020-1789-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Galizia G, Auricchio A, de Vita F, Cardella F, Mabilia A, Basile N, et al. Inflammatory and Nutritional Status is a Predictor of Long-Term Outcome in Patients Undergoing Surgery for Gastric Cancer. Validation of the Naples Prognostic Score. Ann Ital Chir (2019) 90:404–16. [PubMed] [Google Scholar]
- 17. Nakagawa N, Yamada S, Sonohara F, Takami H, Hayashi M, Kanda M, et al. Clinical Implications of Naples Prognostic Score in Patients With Resected Pancreatic Cancer. Ann Surg Oncol (2020) 27(3):887–95. 10.1245/s10434-019-08047-7 [DOI] [PubMed] [Google Scholar]
- 18. Kano K, Yamada T, Yamamoto K, Komori K, Watanabe H, Takahashi K, et al. The Impact of Pretherapeutic Naples Prognostic Score on Survival in Patients With Locally Advanced Esophageal Cancer. Ann Surg Oncol (2021). 10.1245/s10434-020-09549-5 [DOI] [PubMed] [Google Scholar]
- 19. Yang YS, Shang QX, Yuan Y, Wu XY, Hu WP, Chen LQ. Comparison of Long-Term Quality of Life in Patients With Esophageal Cancer After Ivor-Lewis, Mckeown, or Sweet Esophagectomy. J Gastrointest Surg (2019) 23(2):225–31. 10.1007/s11605-018-3999-z [DOI] [PubMed] [Google Scholar]
- 20. Helminen O, Mrena J, Sihvo E. Can We Increase the Resection Rate by Minimally Invasive Approach? Experience From 100 Minimally Invasive Esophagectomies. J Oncol (2019) 2019:3809383. 10.1155/2019/3809383 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Rice TW, Rusch VW, Ishwaran H, Blackstone EH. Worldwide Esophageal Cancer Collaboration. Cancer of the Esophagus and Esophagogastric Junction: Data-Driven Staging for the Seventh Edition of the American Joint Committee on Cancer/International Union Against Cancer Staging Manuals. Cancer (2010) 116(16):3763–73. 10.1002/cncr.25146 [DOI] [PubMed] [Google Scholar]
- 22. Iasonos A, Schrag D, Raj GV, Panageas KS. How to Build and Interpret a Nomogram for Cancer Prognosis. J Clin Oncol (2008) 26(8):1364–70. 10.1200/JCO.2007.12.9791 [DOI] [PubMed] [Google Scholar]
- 23. Li HX, Chang H, Xu BQ, Tao YL, Gao J, Chen C, et al. An Inflammatory Biomarker Based Nomogram to Predict Prognosis of Patients With Nasopharyngeal Carcinoma: An Analysis of a Prospective Study. Cancer Med (2017) 6(1):310–9. 10.1002/cam4.947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Zeng X, Liu G, Pan Y, Li Y. Development and Validation of Immune Inflammation-Based Index for Predicting the Clinical Outcome in Patients With Nasopharyngeal Carcinoma. J Cell Mol Med (2020) 24(15):8326–49. 10.1111/jcmm.15097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Wang Y, Sun K, Shen J, Li B, Kuang M, Cao Q, et al. Novel Prognostic Nomograms Based on Inflammation-Related Markers for Patients With Hepatocellular Carcinoma Underwent Hepatectomy. Cancer Res Treat (2019) 51(4):1464–78. 10.4143/crt.2018.657 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/ Supplementary Material .