Abstract
Introduction
Several clinical studies have reported the efficacy of favipiravir in reducing viral load and shortening the duration of symptoms. However, the viability of SARS-CoV-2 in the context of favipiravir therapy and the potential for resistance development is unclear.
Methods
We sequenced SARS-CoV-2 in nasopharyngeal specimens collected from patients who participated in a randomized clinical trial of favipiravir at hospitals across Japan between March and May 2020. Paired genomes were sequenced from those who remained RT-PCR-positive 5–8 days into favipiravir therapy. Daily nasopharyngeal specimens from 69 patients who were RT-PCR-positive at randomization were examined for a cytopathic effect (CPE).
Results
Some strains early in the trial belonged to clade 19 B, whereas the majority belonged to clade 20 B. The median time from the disease onset to negative CPE was 9 days. CPE was strongly correlated with the time from disease onset, viral load, age, and male sex. Among 23 patients for whom paired genomes were available, all except one had identical genomes. Two mutations were observed in one patient who received favipiravir, neither in the RdRp gene.
Conclusions
The SARS-CoV-2 genome distribution in this clinical trial conducted in Japan reflected the early influx of strains from China followed by replacement by strains from Europe. CPE was significantly associated with age, male sex, and viral loads but not with favipiravir therapy. There was no evidence of resistance development during favipiravir therapy.
Keywords: SARS-CoV-2, Cytopathic effect, Genome epidemiology, Pharmacotherapy
1. Introduction
Coronavirus disease 2019 (COVID-19), caused by SARS-CoV-2, has claimed nearly three million lives worldwide. Japan has been affected since the relatively early stage of the pandemic, with initial influx of strains from China during the lunar new year followed by strains from Europe over the spring of 2020 [1].
While several immunomodulating agents, including dexamethasone, have shown efficacy in treating moderate to severe COVID-19, antiviral treatment options remain limited. Favipiravir is an inhibitor of viral RNA-dependent RNA polymerase (RdRp) with broad-spectrum activity against RNA viruses that functions as a purine analogue and inhibits RdRp by terminating elongation of the complementary viral RNA strand [2]. It was approved for the treatment of novel or re-emerging influenza infection in Japan in 2014. The barrier to favipiravir resistance is considered to be high for influenza virus. A combination of two mutations in the RdRp gene have been associated with favipiravir resistance in the laboratory setting [3]; however, influenza viruses from 57 patients who participated in clinical trials of favipiravir for influenza did not show differences in favipiravir susceptibility before and 1 or 2 days after starting therapy, suggesting a high resistance threshold [4].
Favipiravir has been examined as a potential treatment option for COVID-19 in several controlled clinical trials. We previously conducted a randomized, open-label trial of 89 patients with asymptomatic to mild COVID-19. The patients were randomized to early favipiravir therapy (starting on day 1 of randomization) and late favipiravir therapy (starting on day 6 of randomization). Viral clearance, as evaluated by RT-PCR, occurred by day 6 in 66.7% and 56.1%, respectively, and the mean time to defervescence was 2.1 and 3.2 days, respectively [5].
Using the nasopharyngeal swab specimens obtained in this clinical trial, the aims of this study were to describe the genome epidemiology in Japan early in the pandemic, to determine the viability of the virus with respect to demographics, time since onset, viral load, and favipiravir therapy, and to identify mutations that occurred during treatment and might affect viral susceptibility to favipiravir.
2. Materials and methods
2.1. The specimens
The nasopharyngeal swab specimens were collected during in a previously described clinical trial [5]. The residual media were stored at −80 °C after extraction of RNA for use in the RT-PCR testing for the trial. A total of 1318 specimens were collected from 89 patients, and 392 specimens were determined as positive for SARS-CoV-2 by RT-PCR targeting the nucleocapsid gene [6]. Among the positive specimens, 89 had Ct values below 34 and were considered amenable to high-throughput sequencing. The RT-PCR-positive specimens with the lowest Ct values collected between day 1 to day 5 were sequenced from each patient. For sequencing of paired genomes, only day 1 specimens were used to generate the first genome. For the subsequent genome from the same patients, an RT-PCR-positive specimen collected between day 5 and day 8 (early therapy group, who were on favipiravir for 4–7 days at the time) or between day 5 and day 6 (late therapy group, who were not yet started on favipiravir at the time) was sequenced from each patient as available.
For detection of viable virus, the daily specimens collected between day 1 and day 6 from the 69 patients whose day 1 specimen was RT-PCR-positive were subjected to the cytopathic effect (CPE) assay.
2.2. Genome sequencing
SARS-CoV-2 genomic RNA was extracted from the transport medium of nasopharyngeal specimens with Ct values less than 34 using a QIAamp Viral RNA Mini Kit (Qiagen). SARS-CoV-2 genomes were amplified according to the RT-PCR protocol of the ARTIC network (https://artic.network/ncov-2019) or using QIAseq SARS-CoV-2 Primer Panel (Qiagen). Sequencing libraries of the amplicons were prepared using QIAseq FX DNA Library Kit (Qiagen), and the libraries were analyzed by Illumina NextSeq 2000 sequencer using NextSeq 1000/2000 P2 Reagents (300 Cycles) or Illumina MiSeq sequencer using MiSeq Reagent Kits v2 (300 Cycles). Amplicon sequences were mapped to the reference sequence (MN908947.3), and consensus sequences were obtained according to the Utah DoH ARTIC/Illumina Bioinformatic Workflow (https://github.com/CDCgov/SARS-CoV-2_Sequencing/tree/master/protocols/BFX UT_ARTIC_Illumina) with minor modifications to include reads without primers. Specimens with Ct values of 29 or higher were sequenced twice to minimize PCR errors. Consensus sequences obtained from the workflow were confirmed by reading the mapping files (bams) using Integrative Genomics Viewer (IGV) (http://software.broadinstitute.org/software/igv/).
The genomic epidemiology of SARS-CoV-2 identified in this study was analyzed using the Nextclade webpage (https://clades.nextstrain.org/) and the Nextstrain tools (https://nextstrain.github.io/ncov/). Sequencing results were evaluated in the Nextclade webpage (https://clades.nextstrain.org/), and sequence data indicated as good with all QC status under the default settings were defined as high-quality. SARS-CoV-2 genome sequences registered from Japan were also used for this analysis. For sequences registered only with the months of collection and not the dates, the collection dates were randomly assigned within the months to enable analysis through the Nextstrain pipeline.
2.3. Comparative analysis of the SARS-CoV-2 genomes
Sequences extracted from each genome were aligned to the reference genome and inspected. Variants with allele frequencies ≥70% were identified as SNVs (single nucleotide variants). Variant detection was performed using VarScan [7] in addition to manual inspection. An intra-host single nucleotide variation (iSNV) was defined likewise. The web application tool (http://giorgilab.unibo.it/coronannotator/) was used for variant detection and annotation of mutated genes [8]. The amino acid substitutions on SARS-CoV-2 genome were also evaluated on the Nextclade webpage (https://clades.nextstrain.org/).
2.4. CPE assay
The VeroE6/TMPRSS2 cells (Japanese Collection of Research Bioresources Cell Bank, #JCRB1819) were maintained in Dulbecco's modified Eagle's medium (DMEM) supplemented with 5% fetal bovine serum (FBS) and Penicillin-Streptomycin (Sigma-Aldrich). For isolation of SARS-CoV-2, cells were seeded on a 25-cm 2 cell culture flask (FALCON). The next day, the thawed specimen medium (0.5 mL) was centrifuged at low speed, and the supernatant was mixed with 4.5 mL of isolation medium (DMEM supplemented with 2% FBS, penicillin-streptomycin [Sigma-Aldrich], gentamicin [Sigma-Aldrich], and amphotericin B [Sigma-Aldrich]). The maintenance medium in the flask was then removed, the cells were washed once with isolation medium, and the mixture (5 mL) was added to the flask, followed by incubation at 37 °C. The cells were examined daily for 5 days for cytopathic effect (CPE).
2.5. Statistical analysis
The time from randomization to negative CPE or RT-PCR and the time from onset to negative CPE or RT-PCR were depicted by the Kaplan-Meier method. The day of onset was defined as the day when COVID-19-related symptoms appeared or when the positive RT-PCR specimen was collected. The days of negative CPE and RT-PCR were defined as the first negative CPE result following the last positive CPE result and the first negative RT-PCR result following the last positive RT-PCR result, respectively. The effect of favipiravir on time to negative CPE was assessed by the Cox proportional hazard regression model with adjustment for age and time between positive RT-PCR and randomization. In order to quantify the relationship between daily CPE positivity and risk factor variables such as age, sex, and daily viral loads, binomial generalized estimating equations with logit-link adjusted for days from the onset were used. Age, viral loads (log scale), and days from the onset were included in the models using restricted cubic spline with 3 knots.
2.6. Sequencing data
The sequencing reads generated in this study have been deposited to the GISAID under accession numbers EPI_ISL_862855 to EPI_ISL_862850 and DDBJ DRA under accession number DRA011834.
3. Results
3.1. SARS-CoV-2 clade representation
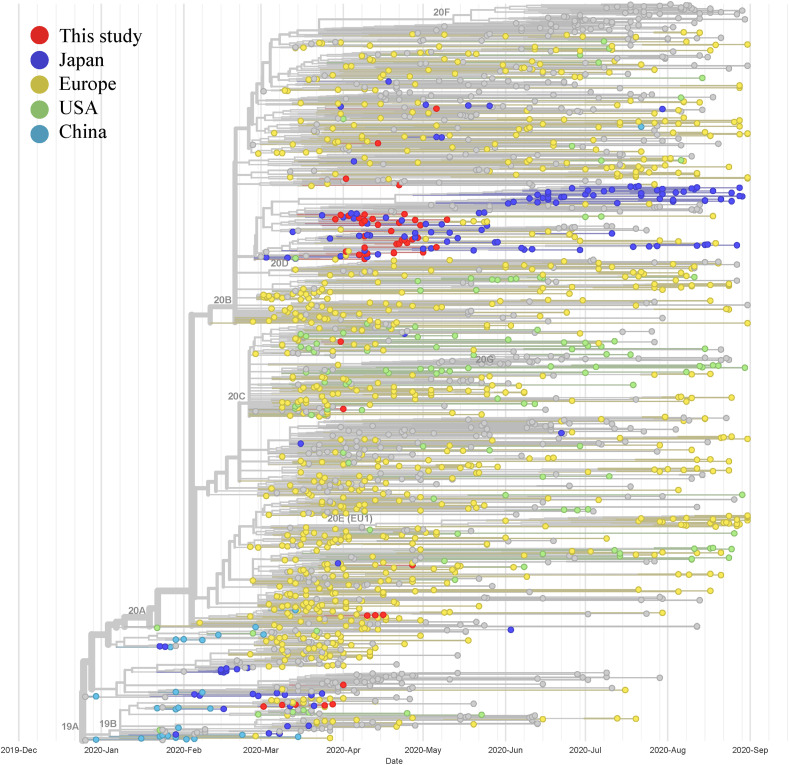
High-quality genome sequences were obtained from 49 of the 89 patients who were enrolled in the clinical trial at hospital sites located across Japan. A previous genome surveillance study suggested that two introductions of SARS-CoV-2 to Japan occurred, the first one between January and February from China and the second one since March from Europe and North America [1]. The clades identified in our trial mirrored this trend. Strains from some of the first enrollments in early March belonged to clades 19A (n = 1) and 19B (n = 5), whereas the rest of the strains belonged to clades 20A (n = 4), 20B (n = 37), and 20C (n = 2), with the majority clustered in clade 20B (Fig. 1 ; Supplemental Fig. 1). Clades 19A and 19B emerged in Wuhan, China, at the onset of the current pandemic, and clade 20A is considered a descendant of clade 19A, with clades 20B and 20C having subsequently emerged as subclades of 20A.
Fig. 1.
Phylogenetic tree of SARS-CoV-2 genomes generated by Nextstrain. Strains from this trial and those worldwide, including those collected between December 2019 and August 2020, are shown. Red: This study, Blue: Japan, Yellow: Europe, Green: the USA, and Pale blue: China.
3.2. Detection of viable virus and its correlates
A total of 421 nasopharyngeal swab specimens collected daily between day 1 and day 6 from 69 patients whose RT-PCR on day 1 was positive for SARS-CoV-2 were subjected to CPE assay. Of the 421 specimens, 323 were positive, whereas the remaining 98 were negative for SARS-CoV-2 by RT-PCR. Of the 421 specimens, 90 were positive for CPE, and all CPE-positive specimens were also positive by RT-PCR. Of the 69 patients, 32 had positive CPE on day 1. For the calculation of time from the onset to negative CPE, 37 patients who had negative CPE on day 1 were considered to have converted to negative on day 1 since pre-enrollment specimens were not available.
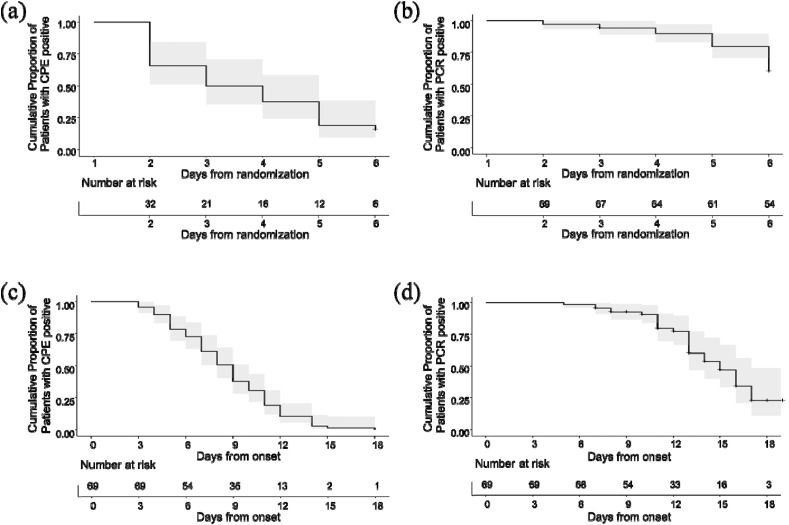
Kaplan-Meier product-limit estimates for the cumulative probability of being free of negative CPE or RT-PCR negativity over time from the time of randomization and the time of disease onset are plotted in Fig. 2 . The median time to the negative CPE was 2.5 days from the time of randomization (95% confidence interval [CI], 2–4), and 9 days from the onset of the disease (95% CI, 8–10). The median time to negative RT-PCR was 15 days from the time of onset of the disease (95% CI, 13–17). Thus, the median time from the onset to the negativity was shorter by 7 days for CPE than RT-PCR (95% CI, 4–8; p < 0.001 by bootstrap method).
Fig. 2.
Kaplan-Meier product-limit estimates for the cumulative probability of being free of negative CPE or RT-PCR negativity over time from the time of randomization and the time of disease onset. The gray shaded area corresponds to pointwise 95% confidence intervals. (a) Cumulative proportion of patients with CPE positive against days from randomization. (b) Cumulative proportion of patients with RT-PCR positive against days from randomization. (c) Cumulative proportion of patients with CPE positive against days from onset. (c) Cumulative proportion of patients with RT-PCR positive against days from onset.
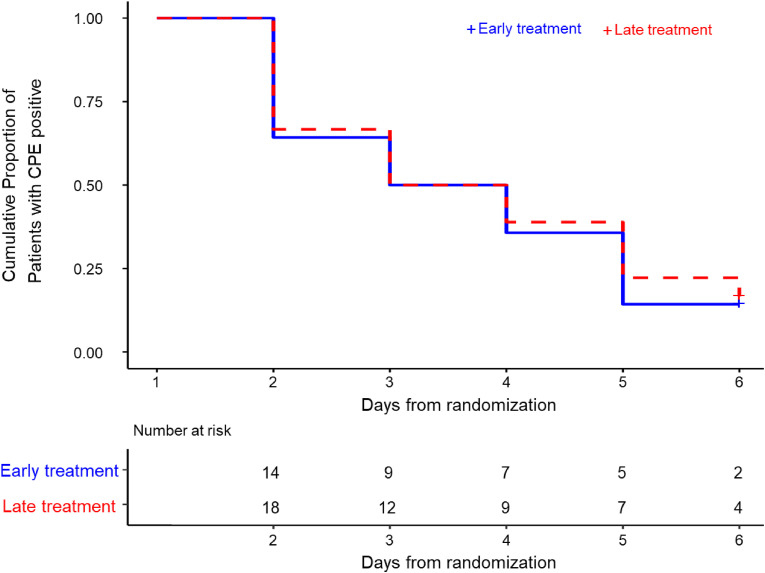
The effect of favipiravir on CPE negativity was examined using data from 32 patients with positive CPE on day 1. Kaplan-Meier product-limit estimates for the cumulative probability of being free of negative CPE over time from the time of randomization stratified by therapy group are depicted in Fig. 3 . There was no significant difference in the time to negative CPE between the early therapy group (i.e., received favipiravir between day 1 and day 6) and the late therapy group (i.e., did not receive favipiravir until day 6) (adjusted hazard ratio [aHR], 1.09; 95% CI, 0.49–2.42; p = 0.837). We previously showed that the proportion of patients who achieved RT-PCR negativity by day 6 were nominally higher in the early therapy group than the late therapy group (66.7% and 56.1%, respectively), and there was a trend towards earlier RT-PCR negativity in the early therapy group (aHR, 1.42; 95% CI, 0.76–2.62) [5]. Overall, the CPE assay did not demonstrate additional virological efficacy of early favipiravir therapy compared with late favipiravir therapy beyond what has been observed in terms of time to RT-PCR negativity under this clinical trial setting [5].
Fig. 3.
Kaplan-Meier product-limit estimates for the cumulative probability of being free of negative CPE over time from the time of randomization stratified by treatment group.
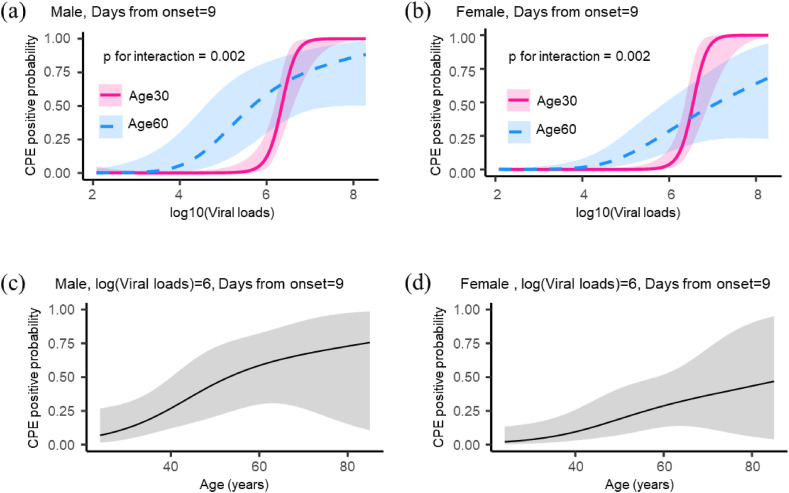
We then investigated factors that may be associated with CPE positivity. The probability of having CPE positivity was quantified by the function of age, sex, and viral loads using daily data from 69 patients. Age, viral loads, and their interaction were significantly associated with CPE positivity (p < 0.001, p = 0.007, respectively, and p = 0.002 for the interaction, Fig. 4 ). The apparent lack of this correlation in older patients may reflect difficulties in accurately identifying time of onset in this population. Male sex was also significantly associated with CPE positivity (odds ratio 3.51; 95% CI, 1.22–10.15; p = 0.020).
Fig. 4.
The CPE positive probabilities quantified by the function of age, sex, viral loads. (a) CPE positive probability against log10 (Viral loads) for patients with age 30 or 60, male sex, and 9 days from onset. (b) CPE positive probability against log10 (Viral loads) for patients with age 30 or 60, female sex, and 9 days from onset. (c) CPE positive probability against age for patients with log10 (Viral loads) = 6, male sex, and 9 days from onset. (d) CPE positive probability against age for patients with log10 (Viral loads) = 6, female sex, and 9 days from onset. For patients with age 30 at 9 days from onset, viral loads around 106 copies/mL corresponded to CPE positive probability nearly 0, and viral loads around 107 copies/mL corresponded to CPE positive probability nearly 1. For patients with age 60 at 9 days from onset, viral loads around 104 copies/mL corresponded to CPE positive probability of nearly 0, and viral loads around 108 copies/mL corresponded to CPE positive probability of 50% or more. The association between CPE positive probability and viral loads was more marked in younger patients.
3.3. Impact of favipiravir therapy on viral sequences
Genomes could be sequenced from day 1 specimen and day 5–8 specimens for 11 patients in the early therapy group and 12 patients in the delayed therapy group (Table 1 ). None of the 12 paired genomes in the delayed therapy group contained SNVs between the genomes from day 1 and day 5 or 6, during which period the patients were not on favipiravir. In the early therapy group in which patients received favipiravir from day 1, one paired genome from day 1 and day 5 contained 3 SNVs, including 2 non-synonymous mutations. These mutations affected the amino acid sequences of NSP4 and NSP16 genes but not the RdRp gene (Table 2 ). None of the remaining paired genomes in the early therapy group contained SNVs.
Table 1.
Ct values and iSNVs between 1st and 2nd specimens among paired sequenced patients.
| Patient group | Patient number | Ct value of day 1 specimen | Ct value of 2nd specimen (day) | The number of iSNVs | The number of amino acid substitutions |
|---|---|---|---|---|---|
| Early therapy group | 9530–09 | 21.43 | 25.04 [5] | 0 | 0 |
| 9530–11 | 28.75 | 26.68 [5] | 3 | 2 | |
| 9532–01 | 24.74 | 22.09 [8] | 0 | 0 | |
| 9538–06 | 21.42 | 30.8 [6] | 0 | 0 | |
| 9539–02 | 28.13 | 28.66 [6] | 0 | 0 | |
| 9545–04 | 17.98 | 29.33 [6] | 0 | 0 | |
| 9550–09 | 22.68 | 29.16 [8] | 0 | 0 | |
| 9551–01 | 26.1 | 30.91 [6] | 0 | 0 | |
| 9722–03 | 23.33 | 31.43 [6] | 0 | 0 | |
| 9722–06 | 19.58 | 29.56 [5] | 0 | 0 | |
| 9736–01 | 28.74 | 28.15 [5] | 0 | 0 | |
| Delayed therapy group | 9530–01 | 25.98 | 30.77 [6] | 0 | 0 |
| 9530–02 | 20.13 | 28.87 [6] | 0 | 0 | |
| 9534–03 | 28.37 | 30.48 [6] | 0 | 0 | |
| 9545–01 | 27.33 | 21.52 [5] | 0 | 0 | |
| 9550–06 | 24.2 | 30.54 [6] | 0 | 0 | |
| 9556–01 | 16.15 | 24.9 [6] | 0 | 0 | |
| 9722–04 | 25.15 | 23.27 [5] | 0 | 0 | |
| 9722–05 | 19.39 | 28.7 [6] | 0 | 0 | |
| 9530–20 | 21.76 | 29.68 [6] | 0 | 0 | |
| 9530–22 | 21.1 | 25.45 [6] | 0 | 0 | |
| 9530–24 | 19.18 | 24.92 [6] | 0 | 0 | |
| 9530–26 | 18.78 | 19.7 [6] | 0 | 0 |
Table 2.
SNVs found in the genomes from patient 9530–11.
| position | 9051 | 15120 | 20429 |
|---|---|---|---|
| ORF | NSP4 | NSP12 (RdRp) | NSP16 |
| day 1 (Ct 28.75) | C | C | C |
| day 5 (Ct 26.68) | T (P2929L) | T | T (P2321L) |
4. Discussion
Leveraging the extensive collection of nasopharyngeal swabs and clinical data available through a previously described multicenter randomized clinical trial of favipiravir for the treatment of COVID-19 [5], we set out to examine the genome epidemiology of SARS-CoV-2 in Japan at the beginning of the pandemic when the trial took place, determine factors associated with prolonged shedding of viable virus, and examine whether favipiravir treatment was associated with mutations that may be associated with resistance.
In this collection of SARS-CoV-2 genomes obtained from patients at hospitals across Japan, strains from early March mainly belonged to clades 19A and 19B that had emerged in Wuhan, and the later strains belonged to clades 20A, 20B and 20C that were likely introduced from Europe and North America over the spring of 2020, a trend that is consistent with a previous nationwide genome analysis [1].
Based on the CPE assay of over 400 nasopharyngeal specimens obtained from 69 patients whose day 1 specimens were positive for SARS-CoV-2 by RT-PCR, the median time to negative CPE was 9 days from the onset of the disease, 7 days shorter than that to negative RT-PCR. Viable virus is generally considered to be present up to approximately 8 days from disease onset among immunocompetent individuals with COVID-19 [9,10]. Our data suggest that shedding of viable virus may occur for a longer period but that the widely adopted practice of 10-day isolation from symptom onset is still adequate to prevent transmission in most cases, even when RT-PCR remains positive for longer.
Male sex, age, viral loads, and interaction of the latter two were significantly associated with CPE positivity in this clinical trial cohort. On the other hand, receipt of favipiravir was not associated with CPE positivity, suggesting that its administration did not translate to suppression of viral growth in this setting. Female sex and young age have been associated with more robust T cell activation during COVID-19, which may contribute to the better clinical outcome in these groups [11]. Association of CPE positivity with male sex and age observed in our study may represent the virological correlate of this immunological observation.
Development of resistance is a potential concern when antiviral therapy is implemented. While the number of patients is limited, it is reassuring that SARS-CoV-2 genomes from pre-treatment and on-treatment specimens of 11 patients who received favipiravir showed no evidence of selection for favipiravir-dependent mutations.
In conclusion, in this virological study that complements a randomized clinical trial of favipiravir for COVID-19 conducted in Japan between March and May 2020, SARS-CoV-2 initially belonged to clades 19A and 19B, which were then replaced mostly by clade 20B, reflecting the influx of virus first from China and subsequently from Europe and the U.S. that occurred before the travel ban was implemented at the end of March 2020. Recovery of the viable virus was associated with age, male sex and viral load, but not with favipiravir therapy.
Financial support
This study was by Japan Agency for Medical Research and Development (AMED) under Grant Numbers JP19fk0108150 and JP20fk0108150.
Author contribution
Study design: A.S., Y.D., T.M. Data collection: M.S., S.K., T. Id., C.K.L., T.M. Data analysis: M.S., T. Im., A.S. Manuscript preparation: T. Im., A.S., C.K.L., Y.D., T.M.
Declaration of competing interest
Y.D. has received speaking fees from FujiFilm Toyama Chemical, which donated favipirvir to the underlying clinical trial but was otherwise not involved in the study. All other authors declare no competing interest.
Acknowledgments
We thank Drs. M. Takahashi and Y. Niwa (International Center for Cell and Gene Therapy, Fujita Health University) for technical support.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jiac.2021.06.010.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Sekizuka T., Itokawa K., Hashino M., Kawano-Sugaya T., Tanaka R., Yatsu K., et al. A genome epidemiological study of SARS-CoV-2 introduction into Japan. mSphere. 2020;5:e00786–20. doi: 10.1128/mSphere.00786-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Shiraki K., Daikoku T. Favipiravir, an anti-influenza drug against life-threatening RNA virus infections. Pharmacol Ther. 2020;209:107512. doi: 10.1016/j.pharmthera.2020.107512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Goldhill D.H., Te Velthuis A.J.W., Fletcher R.A., Langat P., Zambon M., Lackenby A., et al. The mechanism of resistance to favipiravir in influenza. Proc Natl Acad Sci U S A. 2018;115:11613–11618. doi: 10.1073/pnas.1811345115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Takashita E., Ejima M., Ogawa R., Fujisaki S., Neumann G., Furuta Y., et al. Antiviral susceptibility of influenza viruses isolated from patients pre- and post-administration of favipiravir. Antivir Res. 2016;132:170–177. doi: 10.1016/j.antiviral.2016.06.007. [DOI] [PubMed] [Google Scholar]
- 5.Doi Y., Hibino M., Hase R., Yamamoto M., Kasamatsu Y., Hirose M., et al. A prospective, randomized, open-label trial of early versus late favipiravir therapy in hospitalized patients with COVID-19. Antimicrob Agents Chemother. 2020;64:e01897–20. doi: 10.1128/AAC.01897-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shirato K., Nao N., Katano H., Takayama I., Saito S., Kato F., et al. Development of genetic diagnostic methods for detection for novel coronavirus 2019(nCoV-2019) in Japan. Jpn J Infect Dis. 2020;73:304–307. doi: 10.7883/yoken.JJID.2020.061. [DOI] [PubMed] [Google Scholar]
- 7.Koboldt D.C., Zhang Q., Larson D.E., Shen D., McLellan M.D., Lin L., et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568–576. doi: 10.1101/gr.129684.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mercatelli D., Triboli L., Fornasari E., Ray F., Giorgi F.M. Coronapp: a web application to annotate and monitor SARS-CoV-2 mutations. J Med Virol. 2021;93:3238–3245. doi: 10.1002/jmv.26678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.La Scola B., Le Bideau M., Andreani J., Hoang V.T., Grimaldier C., Colson P., et al. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur J Clin Microbiol Infect Dis. 2020;39:1059–1061. doi: 10.1007/s10096-020-03913-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wolfel R., Corman V.M., Guggemos W., Seilmaier M., Zange S., Muller M.A., et al. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581:465–469. doi: 10.1038/s41586-020-2196-x. [DOI] [PubMed] [Google Scholar]
- 11.Takahashi T., Ellingson M.K., Wong P., Israelow B., Lucas C., Klein J., et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature. 2020;588:315–320. doi: 10.1038/s41586-020-2700-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.