Abstract
Xenobiotics are generally known as man-made refractory organic pollutants widely distributed in various environments. For exploring the bioremediation possibility of xenobiotics, two novel xenobiotics-degrading strains affiliated with Azonexaceae were isolated. We report here the phylogenetics, genome, and geo-distribution of a novel and ubiquitous Azonexaceae species that primarily joins in the cometabolic process of some xenobiotics in natural communities. Strains s22 and t15 could be proposed as a novel species within Dechloromonas based on genomic and multi-phylogenetic analysis. Pan-genome analysis showed that the 63 core genes in Dechloromonas include genes for dozens of metabolisms such as nitrogen fixation protein (nifU), nitrogen regulatory protein (glnK), dCTP deaminase, C4-dicarboxylate transporter, and fructose-bisphosphate aldolase. Strains s22 and t15 have the ability to metabolize nitrogen, including nitrogen fixation, NirS-dependent denitrification, and dissimilatory nitrate reduction. Moreover, the novel species possesses the EnvZ-OmpR two-component system for controlling osmotic stress and QseC-QseB system for quorum sensing to rapidly sense environmental changes. It is intriguing that this new species has a series of genes for the biodegradation of some xenobiotics such as azathioprine, 6-Mercaptopurine, trinitrotoluene, chloroalkane, and chloroalkene. Specifically, glutathione S-transferase (GST) and 4-oxalocrotonate tautomerase (praC) in this novel species play important roles in the detoxification metabolism of some xenobiotics like dioxin, trichloroethene, chloroacetyl chloride, benzo[a]pyrene, and aflatoxin B1. Using data from GenBank, DDBJ and EMBL databases, we also demonstrated that members of this novel species were found globally in plants (e.g. rice), guts (e.g. insect), pristine and contaminated regions. Given these data, Dechloromonas sp. strains s22 and t15 take part in the biodegradation of some xenobiotics through key enzymes.
Keywords: Dechloromonas, Xenobiotics biodegradation, Comparative genomics, Novel species, Biogeographic distribution
Dechloromonas; Xenobiotics biodegradation; Comparative genomics; Novel species; Biogeographic distribution.
1. Introduction
Xenobiotics are generally known as man-made refractory organic pollutants [1] and they indeed have harmful health effects on living organisms [2, 3]. Most of them can easily be found in various environments (e.g. soils, sediments, and water bodies) and persist in the environment for months and years [4]. Nonetheless, many biogenic compounds can be biodegraded rapidly [5, 6]. Biodegradation is considered an important way to remove most toxic xenobiotics released into the environment [4] and microorganisms in situ are key players in different xenobiotics-polluted conditions [5, 6, 7]. For this reason, the interest in studying the metabolic potential of biodegrading xenobiotics using microbes for bioremediation purposes has increased, especially in unidentified microorganisms having the xenobiotics-metabolizing capability.
As a member of Azonexaceae, the genus Dechloromonas has been described in the bacterial nomenclature as a chlorate and perchlorate reducer [8, 9, 10, 11]. For instance, D. agitata CKB has been well studied as a model microorganism capable of reducing per(chlorate) [12]. So far, Dechloromonas spp. were studied for their capability to reduce nitrate and nitrite [10, 11, 13, 14, 15]. Members from Dechloromonas are also demonstrated as phosphate accumulating organisms in phosphorus-removing reactors [16, 17, 18]. Additionally, several Dechloromonas strains have been identified as anaerobic Fe (II)- and hydrogen-oxidizing bacteria [13, 14, 19, 20, 21]. They also exhibited a broad range of oxidizing capabilities to simple organic acids, hexoses, reduced humic substances, alcohol, and hydrogen sulphide when nitrate as an electron acceptor was added [22]. Moreover, it is noted that some Dechloromonas isolates were reported to take part in remediating Se- and As-contaminated environments [23, 24].
Many Dechloromonas members have demonstrated exceptional biodegradation potential for xenobiotic compounds. For example, some of them are known for their ability to degrade benzene and polycyclic aromatic hydrocarbons under anaerobic conditions such as Dechloromonas sp. strain JJ [22, 25, 26, 27, 28]. Moreover, they also have the ability to degrade linear alkylbenzene sulfonate and chlorophenol in anoxic environments [29, 30]. Thus, members from Dechloromonas have great potential for biodegrading xenobiotics. However, few studies about Dechloromonas having the ability to biodegrade complex xenobiotics were reported. In the current study, two novel xenobiotics-degrading strains s22 and t15 were isolated from pond water. We investigated their evolution, genome assembly, and biogeography. This study is a supplement to provide a better understanding of the distribution and xenobiotics-degrading potential of members from Dechloromonas.
2. Materials and methods
2.1. Isolation and genomic sequencing
Strains s22 and t15 were isolated from pond water in China. Aliquots (100 μL) of collected pond water were diluted serially 10-fold and an additional 100 μL of the three dilutions (10−4, 10−5, and 10−6) were spread onto 2216E agar plates (pH 7.6–7.8). These 2216E agar plates include the following ingredients (grams per liter): Tryptone, 5.0; Yeast extract, 1.0; Ferric phosphate, 0.005; and Agarose, 15. Of this media, 1.0 L filtered seawater was added as a solvent. All plates were incubated at 25 °C for the next 3 days in order to examine microbial growth. The colonies having different morphological features were streaked individually onto 2216E agar plates and incubated again for another 3 days at 25 °C for growth. This process was repeated three times for purification. Purified bacterial strains were further cultured in marine broth 2216E overnight to cultivate enough bacterial cells for cryopreservation (−80 °C with the addition of 30% glycerol). For DNA extraction of strains s22 and t15, two grams of cells were collected in 2216E solid medium at the exponential growth phase using bamboo stickers of 20 cm, and their genomic DNA was extracted using TIANamp Bacteria DNA Kit (TIANGEN, Beijing, China) following the manufacturer's instructions. NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, United States) was used to measure DNA concentrations and purity. High-quality DNA of strains s22 and t15 was sent to the Beijing Genomics Institute (BGI, Shenzhen, China) for whole-genome sequencing using Illumina HiSeq 2500.
2.2. Genomic assembly and annotation
The quality of raw sequencing data was trimmed and filtered using Trimmomatic at default settings [31]. Quality-filtered and trimmed reads were assigned to de novo assemble using SPAdes assembler with default parameters [32]. In order to check the completeness, contamination, and heterogeneity of assembled genomes of strains s22 and t15, CheckM [33] and Anvi'o software [34] were applied. Other Azonexaceae species genomes were obtained from the NCBI genomes database and related genomic information was listed in the supplementary table S1. Genes prediction on all genomes used in this study was carried out with Prodigal [35] involved in Prokka [36] with default settings to characterize the genomic traits and make FAA protein files compatible for downstream metabolic reconstruction. Furthermore, genome annotations and functional categories were measured based on rapid annotation using the subsystem technology (RAST) server [37], KEGG, and blastp.
2.3. Phylogenetic analysis
In order to determine the phylogenetic placement of two novel strains s22 and t15, multiple sequence alignments of 16S rRNA genes were carried out on CLUSTALW using MEGA X software with default parameters [38]. The complete 16S rRNA gene sequences of strains s22 and t15 were extracted from their draft genomes. Phylogenetic tree based on 16S rRNA gene sequences of strains s22 and t15 as well as other type strains belonging to the family Rhodocyclales was constructed using the maximum-likelihood (ML) algorithms. Rhodospirillum rubrum ATCC 11170T was used as the outgroup. Bootstrap values were calculated based on 1000 replications. Moreover, species tree reconstruction based on all lineal homologous genes of total 28 Azonexaceae genomes (s22, t15, and 26 genomes downloaded from GenBank database) was performed using OrthoFinder2 with STAG algorithm [39]. Additionally, single-copy genes from 28 Azonexaceae genomes were selected to construct the evolutionary tree using Anvi'o software [40] with default settings.
2.4. Comparative genomics and metabolic reconstruction
The online tool JSpeciesWS [41] was arranged to calculate average nucleotide identity (ANI) values and aligned nucleotides percentages (ANP) among the genomes of two novel strains (s22 and t15) and type strains from Azonexaceae. The GGD calculator from the DSMZ server [42] was used to measure in silico genome–genome distances (GGDs) among the genomes. Furthermore, CompareM (https://github.com/dparks1134/CompareM) was applied to calculate amino acid identity (AAI) among the genomes. All generated plots were performed with R [43] using the ggplot2 package [44]. Pan-genome analysis was carried out with Roary using the parameters -e, –mafft, -p 8 [45] and the meta-pangenomic workflow of Anvi'o with default settings [34, 40]. In order to visualize all comparative genome information, the program anvi-interactive in Anvi'o software [34, 40] was used. The metabolic reconstructions based on the results of gene prediction were executed using Adobe Illustrator 2019 (Adobe).
2.5. Biogeographic distribution of Dechloromonas strains
In order to better understand the environmental distribution of Dechloromonas sp. strains s22 and t15, the Integrated Microbial Next Generation Sequencing (IMNGS, https://www.imngs.org) server was applied with a minimum DNA size of 200 bp and the set threshold of 97% using complete 16S rRNA gene sequences [46]. A total of 422,877 samples from different niches (e.g. air, soil, sediment, water bodies, and guts) in IMNGS server were assigned to study the abundant and biogeographic characterizations of Dechloromonas sp. strains s22 and t15. Besides, the biogeographic distribution of 16S rRNA gene sequences of other Dechloromonas strains was also investigated.
2.6. Data accessibility
These Whole Genome Shotgun projects of two strains affiliated with Dechloromonas have been deposited at the DDBJ/ENA/GenBank under the accession WYCU00000000 (s22) and WYCV00000000 (t15). The versions described in this paper are version WYCU01000000 (s22) and WYCV01000000 (t15).
3. Results and discussion
3.1. Phylogenetic placement of two novel strains affiliated with Dechloromonas
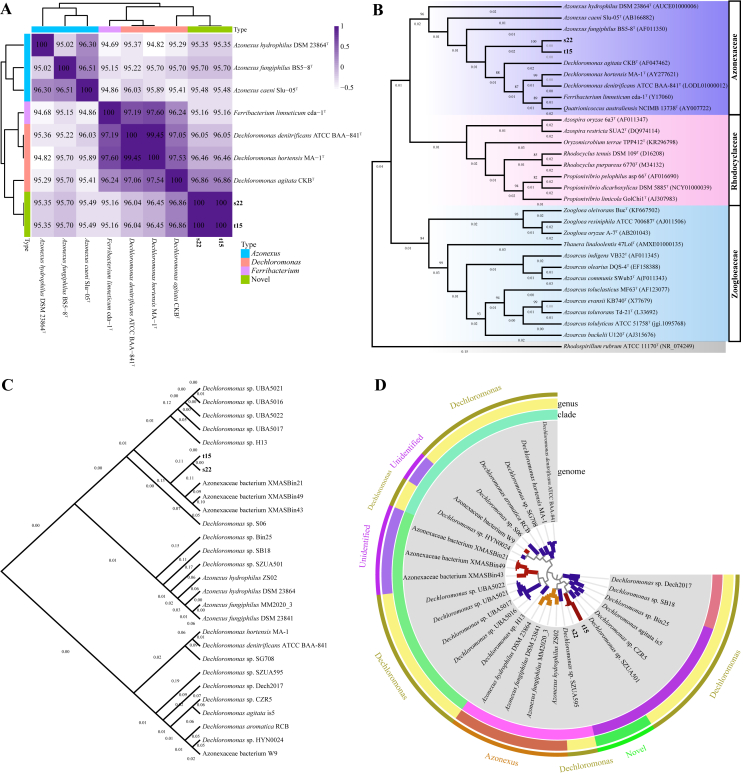
Using the blastn tool, s22 and t15 were closely related to each other (16S rRNA identity: 100%). Moreover, s22 and t15 were closer to members from Dechloromonas (16S rRNA identity: >96.04%) than to the type strains belonging to Azonexus (Figure 1A). It is noted that two strains (s22 and t15) 16S rRNA genes shared the highest sequence similarity of 96.86% with D. agitata CKBT. This identity was lower than the set threshold of 97% for species demarcation, indicating s22 and t15 may be novel species affiliated with Dechloromonas. Additionally, 16S rRNA gene-based phylogeny tree showed that two strains (s22 and t15) were located at the adjacent position of type strain D. agitata CKBT (Figure 1B). Species tree and single-copy genes-based phylogenetic analysis also reflected two strains (s22 and t15) clustered tightly with Dechloromonas strains (Figure 1C, D), which redetermined that s22 and t15 were closer to D. agitata (ANI: 76.67% for both; ANP: 49.74% and 49.75%, respectively; AAI: 81.04% and 81.40%, respectively; GGD: 21.50% for both) than to other Azonexaceae strains (Figure 2). The average ANI, AAI, and GGD values among the Dechloromonas species were ~76%, ~80%, and ~21%, respectively, which are at the lower end of the 62–100% range for interspecies demarcation [47]. On this basis, strains s22 and t15 can be suggested as a new species in the genus Dechloromonas.
Figure 1.
Phylogenetic placement of strains s22 and t15 belonging to family Azonexaceae. (A) A heatmap on the basis of 16S rRNA genes similarities. (B) Maximum-likelihood tree based on the 16S rRNA gene of Rhodocyclales. Rhodospirillum rubrum ATCC 11170T was used as the outgroup. (C) Species tree of all lineal homologous genes among 28 Azonexaceae genomes using OrthoFinder2 with STAG algorithm. (D) Evolutionary reconstructions according to single-copy genes from 28 Azonexaceae genomes using Anvi'o software.
Figure 2.
Genomics characteristics of family Azonexaceae. Heatmaps based on (A) average nucleotide identity (ANI), (B) aligned nucleotides percentages, (C) average amino acid identity (AAI), and (D) genome–genome distance (GGD) for all the sequenced genomes from family Azonexaceae.
3.2. Genomic features of two novel strains of Dechloromonas
No plasmid was detected in the genomes of strains s22 and t15. The genomic size of strains s22 and t15 were 3,575,474 bp and 3,590,015 bp, respectively (Table 1). The average G+C content is 58.4%. The final annotation of strains s22 and t15 resulted in 3,183 and 3,173 genes, respectively. 3,127 and 3,173 predicted coding sequences (CDS) were distributed throughout the strains s22 and t15, respectively. A total of 55 RNA sequences, including 2 rRNA genes (16S and 23S) and 53 tRNA genes, were found in strain s22. For strain t15, 67 RNA sequences (4 rRNA genes and 63 tRNA genes) were detected.
Table 1.
General genomic features of strains s22 and t15.
| Feature | Strain s22 | Strain t15 |
|---|---|---|
| Genome size (bp) | 3,575,474 | 3,590,015 |
| GC content (%) | 58.52 | 58.25 |
| Number of CDS | 3,127 | 3,105 |
| Number of Gene | 3,183 | 3,173 |
| Number of rRNA | 2 | 4 |
| Number of tRNA | 53 | 63 |
| Number of Repeat region | 1 | 3 |
3.3. Pan-genome analysis
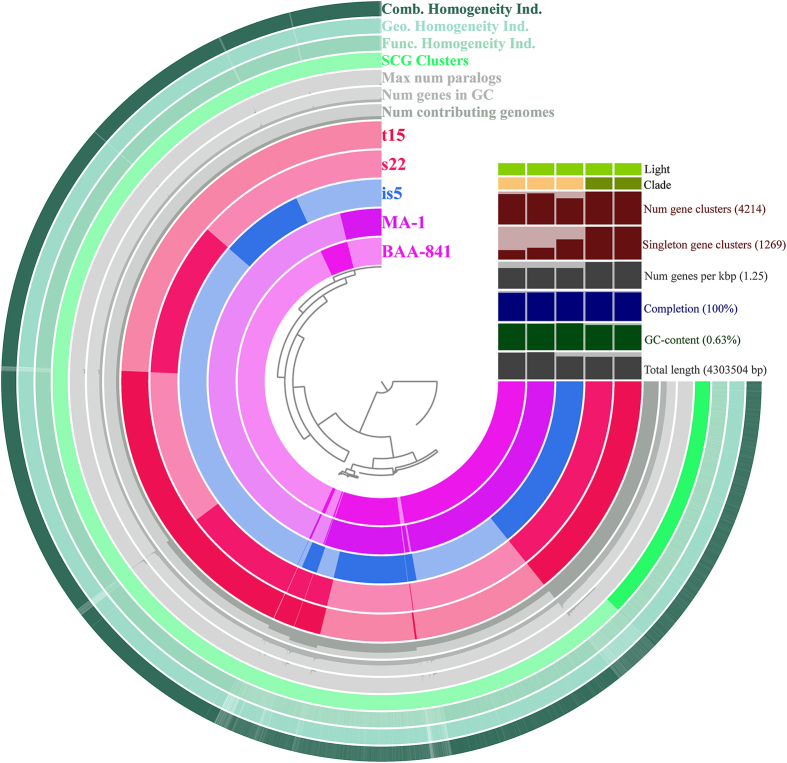
Genomic comparison of the 5 Dechloromonas genomes (s22, t15, is5, MA-1, and BAA-841) was based on pan-genome analysis (Figure 3). Among the 5 Dechloromonas strains analyzed, the 20,670 coding sequences (CDS) of the 5 Dechloromonas genomes clustered into 8,754 protein clusters (PCs). The number of core and pan genes were 63 and 13,607, respectively. The 63 core genes in Dechloromonas include genes for nitrogen fixation protein (nifU), nitrogen regulatory protein (glnK), dCTP deaminase, C4-dicarboxylate transporter, fructose-bisphosphate aldolase, and malate dehydrogenase (mdh). Succinate dehydrogenase/fumarate reductase and thioredoxin 1 were also present in the core genome. With the addition of other Dechloromonas strains, a significant change in the core and pan-genome size occurred (Fig. S1), which suggested a large genomic variation among the species within Dechloromonas [48].
Figure 3.
Communality and uniqueness in the Azonexaceae pangenome as derived from the clustering of 5 Dechloromonas genomes based on 8,754 protein clusters (PCs). Each radial layer represents a genome or a default parameter, and each bar in a layer represents the occurrence of a PC (dark presence, light absence).
3.4. Energy metabolism and environmental information processing
Nitrate as an electron acceptor plays a significant role in the respiration and bioactivity of microbiological cells [49]. The oxidation-reduction of members from Dechloromonas needs the supplement of nitrate [22]. Strains s22 and t15 both contain 25 genes associated with nitrogen metabolism. The metabolic pathways of these genes mainly include nitrogen fixation, NirS-dependent denitrification, and dissimilatory nitrate reduction (Figure 4). Additionally, strains s22 and t15 could uptake and utilize urea due to the presence of urea ABC transporter (UrtABCDE) and urease (UreABCDEFG). The most abundant genes involved in nitrogen utilization are nitronate monooxygenase (npd, 3 copies for each strain) that converts nitroalkane into nitrite. With the presence of ABC transporter CysPUWA, strains s22 and t15 cells could uptake SO42- and S2O32- from extracellular environments. Assimilatory sulfate reduction (sulfate => H2S) pathway and sox system represent the key sulfur metabolic pattern of strains s22 and t15.
Figure 4.
Vary shared metabolic pathways of strains s22 and t15 illustrating predicted ABC transporters, two-component system, nitrogen metabolism, sulfur metabolism, and xenobiotics biodegradation and metabolism. Genes arranged in glutathione S-transferase also take part in degrading benzo[a]pyrene, bromobenzene, naphthalene, 1-nitronaphthalene, aflatoxin B1, and 1,2-Dibromoethane.
Despite the lack of genes encoding for NtrY, strains s22 and t15 encode the NarX-NarL two-component system and the GlnL-GlnG system for the regulation of nitrate metabolism. Moreover, strains s22 and t15 possess the EnvZ-OmpR two-component system for controlling osmotic stress and QseC-QseB system for quorum sensing such as the regulation of motility [50, 51, 52]. The identified two-component regulatory systems can help these organisms to rapidly sense environmental changes [52]. Furthermore, strains s22 and t15 have the PhoR-PhoB two-component system for the detection of inorganic phosphate limitation, which may be useful to sense nutrient enrichment and depletion [53]. Lastly, the RegB-RegA two-component system detected in strains s22 and t15 can be used for electron transfer and aerobic respiration in addition to regulating nitrogen assimilation.
3.5. Xenobiotics biodegradation and metabolism
Dechloromonas sp. strains s22 and t15 possess many genes involved in xenobiotics biodegradation and metabolism that may play an important role in the detoxification of xenobiotics (Table S2). Strains s22 and t15 have a complete biodegradation pathway of azathioprine and 6-Mercaptopurine, which are oral immunosuppressants and have many pathogenic risks [54, 55]. Additionally, it is intriguing that this novel species has a series of genes that catalyzes the first step of xenobiotics biodegradation. For instance, with the presence of genes encoding UbiX, phenol can be catalyzed into 4-Hydroxybenzoate. Trinitrotoluene can be degraded into 2-Hydroxylamino-2,6-dinitrotoluene or 4-Hydroxylamino-2,6-dinitrotoluene by genes encoding NfnB and NemA. The two novel strains also take part in the downstream metabolism of degrading xenobiotics. In the late stage of benzoyl-CoA degradation, 3-Hydroxy-pimeloyl-CoA can continue to catalyze. The results of the genome analysis showed that Ferribacterium limneticum strain RCB harbored more genes required for xenobiotics degradation than other Azonexaceae strains. Even so, for chloroalkane and chloroalkene degradation, strains s22 and t15 can biodegrade chloroacetaldehyde and trans/cis-3-Chloroallyl aldehyde due to genes arranged in aldehyde dehydrogenase (ALDH). Genomic analysis also showed that D. denitrificans ATCC BAA-841, D. hortensis MA-1, and D. sp. CZR5 had this gene to take part in biodegrading chloroalkane and chloroalkene (Table S2). Moreover, strains s22 and t15 take part in the last two steps of the process of caprolactam degradation before entering the benzoate degradation. It is noted that a key enzyme named 4-oxalocrotonate tautomerase (praC) also joins in xenobiotics degradation such as dioxin, xylene, and some aromatic compounds. Moreover, glutathione S-transferase (GST) which is a major Phase II enzyme plays a significant role in the detoxification metabolism of exogenous substances [56]. Genes arranged in GST take part in degrading trichloroethene, chloroacetyl chloride, benzo[a]pyrene, bromobenzene, naphthalene, 1-nitronaphthalene, aflatoxin B1, and 1,2-Dibromoethane (Figure 4 and Fig. S2). These multifunctional enzymes may play an essential role as the cometabolic potential of some xenobiotics in microbial communities. It is important to note that Dechloromonas spp. have the ability to degrade xenobiotics and related metabolisms should be characterized deeply in the future. To some extent, the cometabolism within microbial communities can determine the fate of xenobiotic compounds in nature [57]. In addition, potential functional genes involved in the isolates should be explored due to the incomplete annotation of xenobiotics-degrading metabolic pathways. In the future, more works should focus on the discovery of novel xenobiotics-degrading enzymes and microorganisms.
It is reported that Dechloromonas spp. are facultative anaerobic nitrate-reducing bacteria and are known as aromatic compound-degrading bacteria [25]. They are frequently present in aerobic/anaerobic WWTPs [58, 59, 60, 61], where they contribute to the degradation of organic compounds and removal of N and P under aerobic and anaerobic conditions. For strains s22 and t15, apart from reducing nitrate, degrading some aromatic compounds and accumulating polyphosphate, they could be used to remove some poisonous medicines such as azathioprine and 6-Mercaptopurine to reduce pathogenic risk, as well as some small molecular xenobiotics in the wastewater treatment. To sum up, strains s22 and t15 possess a broad range of metabolisms integral to the wastewater treatment ecosystem. The rationale behind the use of this novel species to treat wastewater lies in their ability to (i) biodegrade some xenobiotics through key enzymes like GST and praC; (ii) use the EnvZ-OmpR two-component system for controlling osmotic stress and QseC-QseB system for quorum sensing to rapidly sense environmental changes; (iii) acclimatize and survive in plants, guts, pristine and contaminated regions.
3.6. Global distribution and abundance of Dechloromonas bacteria
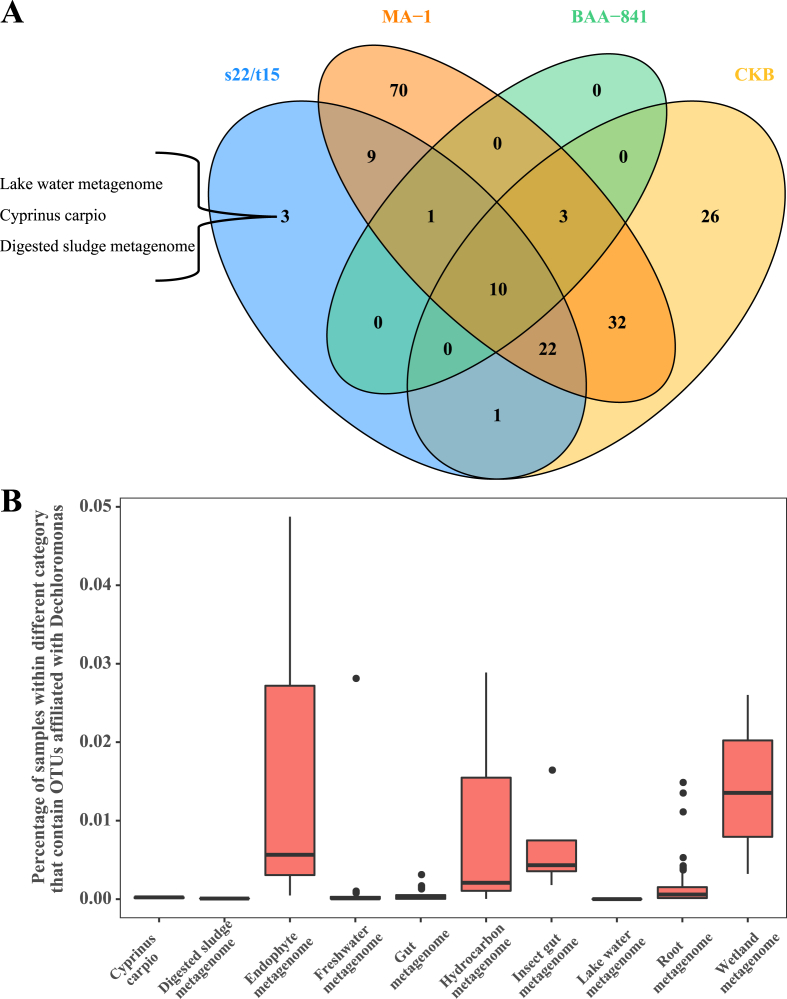
Generally, it was consistent with our results that members of the genus Dechloromonas are ubiquitous and have been isolated and identified from almost all screened environments (e.g. lake and river sediments, paper and pulp mill waste, groundwater, soil as well as invertebrate guts) as shown in Supplementary Table S3 and Fig. S3 [19, 62 ,63]. Investigation of biogeographical characterization of strains s22 and t15 based on IMNGS analysis showed that the target 16S rRNA genes were determined (at >97% similarity) in 760 environmental samples (Table S3). It is intriguing that bacteria closely related with strains s22 and t15 could exist in three distinct geographical samples (Figure 5A), including lake water metagenome from Australia (e.g. SRR2516495), Cyprinus carpio gut metagenomes from China (e.g. ERR238326), and digested sludge metagenome from China (e.g. SRR3068598). The target 16S rRNA genes of this new species were abundant in plants (e.g. rice), guts (e.g. insect), water bodies (e.g. lake), and hydrocarbon-contaminated regions (Figure 5B). The presence and relative abundances of Dechloromonas sp. strains s22 and t15 provide evidence that this new species is more abundant in rice endotrophic metagenomes (e.g. 0.0487% in SRR1197948) than other Dechloromonas species (Table S3). Moreover, there was a report that Dechloromonas could be dominant in the rice rhizospheric microbial community with the supplement of nitrate [64]. Thus, it can be inferred that s22 and t15 as facultative anaerobes take part in the metabolic process in plant organisms, animal guts, pristine and contaminated environments, and most especially, they are associated with detoxification metabolism of xenobiotics.
Figure 5.
Biogeographic distribution and abundance profiles of Dechloromonas strains in kinds of samples. (A) Venn diagram of different categories that contain OTUs affiliated with Dechloromonas. (B) Abundance of OTUs belonging to strains s22 and t15 in 10 representative samples.
4. Conclusion
In the present study, strains s22 and t15 were proposed as novel species within Dechloromonas. They played important roles in the biodegradation of some xenobiotics through key enzymes like GST. They possess the EnvZ-OmpR two-component system for controlling osmotic stress and QseC-QseB system for quorum sensing to rapidly sense environmental changes. Pan-genome analysis showed that there are 63 core genes in Dechloromonas and revealed the potential survival strategies of strains within Dechloromonas dwelling in eutrophic and polluted environments. Moreover, biogeographic characterization underlined members of this novel species are found globally in plants, guts, pristine and contaminated regions. The foregoing findings provide a deeper understanding of the biogeographic distribution and the xenobiotics-degrading potential of members from Dechloromonas.
Declarations
Author contribution statement
Shuangfei Zhang: Conceived and designed the experiments; Performed the experiments; Analyzed and interpreted the data; Wrote the paper.
Charles Amanze, Xueduan Liu, Yili Liang: Conceived and designed the experiments; Analyzed and interpreted the data; Wrote the paper.
Chongran Sun, Kai Zou, Shaodong Fu, Yan Deng: Analyzed and interpreted the data; Contributed reagents, materials, analysis tools or data; Wrote the paper.
Funding statement
This work was supported by the following grants: Chinese National Science and Technology Support Program (2013BAC09B00), Chinese National Natural Science Foundation of China (31570113 and 302001124), and the Fundamental Research Funds for the Central Universities of Central South University (2020zzts207).
Data availability statement
These Whole Genome Shotgun projects of two strains affiliated with Dechloromonas have been deposited at the DDBJ/ENA/GenBank under the accession WYCU00000000 (s22) and WYCV00000000 (t15). The versions described in this paper are version WYCU01000000 (s22) and WYCV01000000 (t15).
Competing interest statement
The authors declare no conflict of interest.
Additional information
No additional information is available for this paper.
Appendix A. Supplementary data
The following is the supplementary data related to this article:
References
- 1.Sahin A.D., Sacan M.T. Understanding the toxic potencies of xenobiotics inducing TCDD/TCDF-like effects. SAR QSAR Environ. Res. 2018;29(2):117–131. doi: 10.1080/1062936X.2017.1414075. [DOI] [PubMed] [Google Scholar]
- 2.Faggio C., Tsarpali V., Dailianis S. Mussel digestive gland as a model tissue for assessing xenobiotics: an overview. Sci. Total Environ. 2018;636:220–229. doi: 10.1016/j.scitotenv.2018.04.264. [DOI] [PubMed] [Google Scholar]
- 3.Messina C.M., Faggio C., Laudicella V.A., Sanfilippo M., Trischitta F., Santulli A. Effect of sodium dodecyl sulfate (SDS) on stress response in the Mediterranean mussel (Mytilus Galloprovincialis): regulatory volume decrease (Rvd) and modulation of biochemical markers related to oxidative stress. Aquat. Toxicol. 2014;157:94–100. doi: 10.1016/j.aquatox.2014.10.001. [DOI] [PubMed] [Google Scholar]
- 4.Singh R. Biodegradation of xenobiotics-a way for environmental detoxification. Int. J. Dev. Res. 2017;7(1):14082–14087. [Google Scholar]
- 5.Aranda E. Promising approaches towards biotransformation of polycyclic aromatic hydrocarbons with Ascomycota fungi. Curr. Opin. Biotechnol. 2016;38:1–8. doi: 10.1016/j.copbio.2015.12.002. [DOI] [PubMed] [Google Scholar]
- 6.Marco-Urrea E., García-Romera I., Aranda E. Potential of non-ligninolytic fungi in bioremediation of chlorinated and polycyclic aromatic hydrocarbons. N. Biotech. 2015;32(6):620–628. doi: 10.1016/j.nbt.2015.01.005. [DOI] [PubMed] [Google Scholar]
- 7.Zhang C.L., Bennett G.N. Biodegradation of xenobiotics by anaerobic bacteria. Appl. Microbiol. Biotechnol. 2005;67(5):600–618. doi: 10.1007/s00253-004-1864-3. [DOI] [PubMed] [Google Scholar]
- 8.Achenbach L.A., Michaelidou U., Bruce R.A., Fryman J., Coates J.D. Dechloromonas agitata gen. nov., sp nov and Dechlorosoma suillum gen. nov., sp nov., two novel environmentally dominant (per)chlorate-reducing bacteria and their phylogenetic position. Int. J. Syst. Evol. Microbiol. 2001;51:527–533. doi: 10.1099/00207713-51-2-527. [DOI] [PubMed] [Google Scholar]
- 9.Logan B.E., Zhang H., Mulvaney P., Milner M.G., Head I.M., Unz R.F. Kinetics of perchlorate- and chlorate-respiring bacteria. Appl. Environ. Microbiol. 2001;67(6):2499–2506. doi: 10.1128/AEM.67.6.2499-2506.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wang Y.L., Zhao J.W., Wang D.B., Liu Y.W., Wang Q.L., Ni B.J., Chen F., Yang Q., Li X.M., Zeng G.M., Yuan Z.G. Free nitrous acid promotes hydrogen production from dark fermentation of waste activated sludge. Water Res. 2018;145:113–124. doi: 10.1016/j.watres.2018.08.011. [DOI] [PubMed] [Google Scholar]
- 11.Xu Z.S., Chai X.L. Effect of weight ratios of PHBV/PLA polymer blends on nitrate removal efficiency and microbial community during solid-phase denitrification. Int. Biodeterior. Biodegrad. 2017;116:175–183. [Google Scholar]
- 12.Liebensteiner M.G., Oosterkamp M.J., Stams A.J.M. Microbial respiration with chlorine oxyanions: diversity and physiological and biochemical properties of chlorate- and perchlorate-reducing microorganisms. In: Lambertz M., Perry S.F., editors. Special Issue: Respiratory Science. Blackwell Science Publ; Oxford: 2016. pp. 59–72. [DOI] [PubMed] [Google Scholar]
- 13.Coby A.J., Picardal F., Shelobolina E., Xu H.F., Roden E.E. Repeated anaerobic microbial redox cycling of iron. Appl. Environ. Microbiol. 2011;77(17):6036–6042. doi: 10.1128/AEM.00276-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gibney B.P., Nusslein K. Arsenic sequestration by nitrate respiring microbial communities in urban lake sediments. Chemosphere. 2007;70(2):329–336. doi: 10.1016/j.chemosphere.2007.05.094. [DOI] [PubMed] [Google Scholar]
- 15.Ginige M.P., Keller J., Blackall L.L. Investigation of an acetate-fed denitrifying microbial community by stable isotope probing, full-cycle rRNA analysis, and fluorescent in situ hybridization-microautoradiography. Appl. Environ. Microbiol. 2005;71(12):8683–8691. doi: 10.1128/AEM.71.12.8683-8691.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Liu Y., Zhang T., Fang H.H.P. Microbial community analysis and performance of a phosphate-removing activated sludge. Bioresour. Technol. 2005;96(11):1205–1214. doi: 10.1016/j.biortech.2004.11.003. [DOI] [PubMed] [Google Scholar]
- 17.Yuan C., Wang B., Peng Y., Li X., Zhang Q., Hu T. Enhanced nutrient removal of simultaneous partial nitrification, denitrification and phosphorus removal (SPNDPR) in a single-stage anaerobic/micro-aerobic sequencing batch reactor for treating real sewage with low carbon/nitrogen. Chemosphere. 2020;257:127097. doi: 10.1016/j.chemosphere.2020.127097. [DOI] [PubMed] [Google Scholar]
- 18.Zhang X.X., Wang C.C., Wu P., Yin W., Xu L.Z. New insights on biological nutrient removal by coupling biofilm-based CANON and denitrifying phosphorus removal (CANDPR) process: long-term stability assessment and microbial community evolution. Sci. Total Environ. 2020;730:11. doi: 10.1016/j.scitotenv.2020.138952. [DOI] [PubMed] [Google Scholar]
- 19.Chakraborty A., Picardal F. Neutrophilic, nitrate-dependent, Fe(II) oxidation by a Dechloromonas species. World J. Microbiol. Biotechnol. 2013;29(4):617–623. doi: 10.1007/s11274-012-1217-9. [DOI] [PubMed] [Google Scholar]
- 20.Weber K.A., Urrutia M.M., Churchill P.F., Kukkadapu R.K., Roden E.E. Anaerobic redox cycling of iron by freshwater sediment microorganisms. Environ. Microbiol. 2006;8(1):100–113. doi: 10.1111/j.1462-2920.2005.00873.x. [DOI] [PubMed] [Google Scholar]
- 21.Zhang H.S., Bruns M.A., Logan B.E. Perchlorate reduction by a novel chemolithoautotrophic, hydrogen-oxidizing bacterium. Environ. Microbiol. 2002;4(10):570–576. doi: 10.1046/j.1462-2920.2002.00338.x. [DOI] [PubMed] [Google Scholar]
- 22.Singh D.N., Kumar A., Sarbhai M.P., Tripathi A.K. Cultivation-independent analysis of archaeal and bacterial communities of the formation water in an Indian coal bed to enhance biotransformation of coal into methane. Appl. Microbiol. Biotechnol. 2012;93(3):1337–1350. doi: 10.1007/s00253-011-3778-1. [DOI] [PubMed] [Google Scholar]
- 23.Suhadolnik M.L.S., Salgado A.P.C., Scholte L.L.S., Bleicher L., Costa P.S., Reis M.P., Dias M.F., Avila M.P., Barbosa F.A.R., Chartone-Souza E., Nascimento A.M.A. Novel arsenic-transforming bacteria and the diversity of their arsenic-related genes and enzymes arising from arsenic-polluted freshwater sediment. Sci. Rep. 2017;7:17. doi: 10.1038/s41598-017-11548-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhang Y.Q., Frankenberger W.T. Supplementing Bacillus sp. RS1 with Dechloromonas sp. HZ for enhancing selenate reduction in agricultural drainage water. Sci. Total Environ. 2007;372(2-3):397–405. doi: 10.1016/j.scitotenv.2006.10.027. [DOI] [PubMed] [Google Scholar]
- 25.Coates J.D., Chakraborty R., Lack J.G., O'Connor S.M., Cole K.A., Bender K.S., Achenbach L.A. Anaerobic benzene oxidation coupled to nitrate reduction in pure culture by two strains of Dechloromonas. Nature. 2001;411(6841):1039–1043. doi: 10.1038/35082545. [DOI] [PubMed] [Google Scholar]
- 26.Xu M., He Z., Zhang Q., Liu J., Guo J., Sun G., Zhou J. Responses of aromatic-degrading microbial communities to elevated nitrate in sediments. Environ. Sci. Technol. 2015;49(20):12422–12431. doi: 10.1021/acs.est.5b03442. [DOI] [PubMed] [Google Scholar]
- 27.Yan Z.S., He Y.H., Cai H.Y., Van Nostrand J.D., He Z.L., Zhou J.Z., Krumholz L.R., Jiang H.L. Interconnection of key microbial functional genes for enhanced benzo[a]pyrene biodegradation in sediments by microbial electrochemistry. Environ. Sci. Technol. 2017;51(15):8519–8529. doi: 10.1021/acs.est.7b00209. [DOI] [PubMed] [Google Scholar]
- 28.Yu X., Sun J., Li G., Huang Y., Li Y., Xia D., Jiang F. Integration of •SO4--based AOP mediated by reusable iron particles and a sulfidogenic process to degrade and detoxify Orange II. Water Res. 2020;174:115622. doi: 10.1016/j.watres.2020.115622. [DOI] [PubMed] [Google Scholar]
- 29.Carosia M.F., Okada D.Y., Sakamoto I.K., Silva E.L., Varesche M.B.A. Microbial characterization and degradation of linear alkylbenzene sulfonate in an anaerobic reactor treating wastewater containing soap powder. Bioresour. Technol. 2014;167:316–323. doi: 10.1016/j.biortech.2014.06.002. [DOI] [PubMed] [Google Scholar]
- 30.Yu Y., Ndayisenga F., Yu Z., Zhao M., Lay C.-H., Zhou D. Co-substrate strategy for improved power production and chlorophenol degradation in a microbial fuel cell. Int. J. Hydrogen Energy. 2019;44(36):20312–20322. [Google Scholar]
- 31.Bolger A.M., Lohse M., Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114–2120. doi: 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bankevich A., Nurk S., Antipov D., Gurevich A.A., Dvorkin M., Kulikov A.S., Lesin V.M., Nikolenko S.I., Pham S., Prjibelski A.D., Pyshkin A.V., Sirotkin A.V., Vyahhi N., Tesler G., Alekseyev M.A., Pevzner P.A. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012;19(5):455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Parks D.H., Imelfort M., Skennerton C.T., Hugenholtz P., Tyson G.W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25(7):1043–1055. doi: 10.1101/gr.186072.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Delmont T.O., Eren A.M. Linking pangenomes and metagenomes: the Prochlorococcus metapangenome. PeerJ. 2018;6 doi: 10.7717/peerj.4320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hyatt D., Chen G.L., LoCascio P.F., Land M.L., Larimer F.W., Hauser L.J. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf. 2010;11:11. doi: 10.1186/1471-2105-11-119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068–2069. doi: 10.1093/bioinformatics/btu153. [DOI] [PubMed] [Google Scholar]
- 37.Aziz R.K., Bartels D., Best A.A., DeJongh M., Disz T., Edwards R.A., Formsma K., Gerdes S., Glass E.M., Kubal M., Meyer F., Olsen G.J., Olson R., Osterman A.L., Overbeek R.A., McNeil L.K., Paarmann D., Paczian T., Parrello B., Pusch G.D., Reich C., Stevens R., Vassieva O., Vonstein V., Wilke A., Zagnitko O. The RAST server: rapid annotations using subsystems technology. BMC Genom. 2008;9:15. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kumar S., Stecher G., Li M., Knyaz C., Tamura K., MEGA X. Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018;35(6):1547–1549. doi: 10.1093/molbev/msy096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Emms D.M., Kelly S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 2019;20(1):238. doi: 10.1186/s13059-019-1832-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Eren A.M., Esen Ö.C., Quince C., Vineis J.H., Morrison H.G., Sogin M.L., Delmont T.O. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ. 2015;3:e1319. doi: 10.7717/peerj.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Richter M., Rossello-Mora R., Glockner F.O., Peplies J., JSpeciesWS A web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics. 2016;32(6):929–931. doi: 10.1093/bioinformatics/btv681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Meier-Kolthoff J.P., Klenk H.P., Goker M. Taxonomic use of DNA G plus C content and DNA-DNA hybridization in the genomic age. Int. J. Syst. Evol. Microbiol. 2014;64:352–356. doi: 10.1099/ijs.0.056994-0. [DOI] [PubMed] [Google Scholar]
- 43.Team R.C. 2016. A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. 2017, Vienna, Austria.https://www.R-project.org/ [Google Scholar]
- 44.Wickham H. Springer; 2016. ggplot2: Elegant Graphics for Data Analysis. [Google Scholar]
- 45.Page A.J., Cummins C.A., Hunt M., Wong V.K., Reuter S., Holden M.T.G., Fookes M., Falush D., Keane J.A., Parkhill J. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics. 2015;31(22):3691–3693. doi: 10.1093/bioinformatics/btv421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lagkouvardos I., Joseph D., Kapfhammer M., Giritli S., Horn M., Haller D., Clavel T. IMNGS: a comprehensive open resource of processed 16S rRNA microbial profiles for ecology and diversity studies. Sci. Rep. 2016;6(1):33721. doi: 10.1038/srep33721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kim M., Oh H.S., Park S.C., Chun J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014;64:346–351. doi: 10.1099/ijs.0.059774-0. [DOI] [PubMed] [Google Scholar]
- 48.Panthee S., Paudel A., Blom J., Hamamoto H., Sekimizu K. Complete genome sequence of Weissella hellenica 0916-4-2 and its comparative genomic analysis. Front. Microbiol. 2019;10:13. doi: 10.3389/fmicb.2019.01619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ogilvie B.G., Rutter M., Nedwell D.B. Selection by temperature of nitrate-reducing bacteria from estuarine sediments: species composition and competition for nitrate. FEMS Microbiol. Ecol. 1997;23(1):11–22. [Google Scholar]
- 50.Audia J.P. Amer Soc Microbiology; 1752 N Street Nw, Washington, Dc 20036-2904 USA: 2012. Rickettsial Physiology and Metabolism in the Face of Reductive Evolution. [Google Scholar]
- 51.Sharma V.K., Casey T.A. Escherichia coli O157:H7 lacking the qseBC-encoded quorum-sensing system outcompetes the parental strain in colonization of cattle intestines. Appl. Environ. Microbiol. 2014;80(6):1882–1892. doi: 10.1128/AEM.03198-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Smith D.P., Thrash J.C., Nicora C.D., Lipton M.S., Burnum-Johnson K.E., Carini P., Smith R.D., Giovannoni S.J. Proteomic and transcriptomic analyses of ‘Candidatus Pelagibacter ubique’ describe the first PII-independent response to nitrogen limitation in a free-living Alphaproteobacterium. mBio. 2013;4(6) doi: 10.1128/mBio.00133-12. e00133-00112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Klinges J.G., Rosales S.M., McMinds R., Shaver E.C., Shantz A.A., Peters E.C., Eitel M., Worheide G., Sharp K.H., Burkepile D.E., Silliman B.R., Thurber R.L.V. Phylogenetic, genomic, and biogeographic characterization of a novel and ubiquitous marine invertebrate-associated Rickettsiales parasite, Candidatus Aquarickettsia rohweri, gen. nov., sp. nov. ISME J. 2019;13(12):2938–2953. doi: 10.1038/s41396-019-0482-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Belizna C., Meroni P.L., Shoenfeld Y., Devreese K., Alijotas-Reig J., Esteve-Valverde E., Chighizola C., Pregnolato F., Cohen H., Fassot C., Mattera P.M., Peretti P., Levy A., Bernard L., Saiet M., Lagarce L., Briet M., Riviere M., Pellier I., Gascoin G., Rakotonjanahary J., Borghi M.O., Stojanovich L., Djokovic A., Stanisavljevic N., Bromley R., Elefant-Amoura E., Bahi Buisson N., Pindi Sala T., Kelchtermans H., Makatsariya A., Bidsatze V., Khizroeva J., Latino J.O., Udry S., Henrion D., Loufrani L., Guihot A.L., Muchardt C., Hasan M., Ungeheuer M.N., Voswinkel J., Damian L., Pabinger I., Gebhart J., Lopez Pedrera R., Cohen Tervaert J.W., Tincani A., Andreoli L. In utero exposure to Azathioprine in autoimmune disease. Where do we stand? Autoimmun. Rev. 2020;19(9):102525. doi: 10.1016/j.autrev.2020.102525. [DOI] [PubMed] [Google Scholar]
- 55.Kotlyar D.S., Lewis J.D., Beaugerie L., Tierney A., Brensinger C.M., Gisbert J.P., Loftus E.V., Peyrin-Biroulet L., Blonski W.C., Van Domselaar M., Chaparro M., Sandilya S., Bewtra M., Beigel F., Biancone L., Lichtenstein G.R. Risk of lymphoma in patients with inflammatory bowel disease treated with azathioprine and 6-Mercaptopurine: a meta-analysis. Clin. Gastroenterol. Hepatol. 2015;13(5) doi: 10.1016/j.cgh.2014.05.015. 847-U541. [DOI] [PubMed] [Google Scholar]
- 56.Zhou Z.Z., Wang B., Zeng S.M., Gong Z., Jing F., Zhang J.S. Glutathione S-transferase (GST) genes from marine copepods Acartia tonsa: cDNA cloning and mRNA expression in response to 1,2-dimethylnaphthalene. Aquat. Toxicol. 2020;224:8. doi: 10.1016/j.aquatox.2020.105480. [DOI] [PubMed] [Google Scholar]
- 57.Rieger P.G., Meier H.M., Gerle M., Vogt U., Groth T., Knackmuss H.J. Xenobiotics in the environment: present and future strategies to obviate the problem of biological persistence. J. Biotechnol. 2002;94(1):101–123. doi: 10.1016/s0168-1656(01)00422-9. [DOI] [PubMed] [Google Scholar]
- 58.Hu M., Wang X., Wen X., Xia Y. Microbial community structures in different wastewater treatment plants as revealed by 454-pyrosequencing analysis. Bioresour. Technol. 2012;117:72–79. doi: 10.1016/j.biortech.2012.04.061. [DOI] [PubMed] [Google Scholar]
- 59.Saunders A.M., Albertsen M., Vollertsen J., Nielsen P.H. The activated sludge ecosystem contains a core community of abundant organisms. ISME J. 2016;10(1):11–20. doi: 10.1038/ismej.2015.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tu Y., Schuler A.J. Low acetate concentrations favor polyphosphate-accumulating organisms over glycogen-accumulating organisms in enhanced biological phosphorus removal from wastewater. Environ. Sci. Technol. 2013;47(8):3816–3824. doi: 10.1021/es304846s. [DOI] [PubMed] [Google Scholar]
- 61.Zhang T., Shao M.F., Ye L. 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J. 2012;6(6):1137–1147. doi: 10.1038/ismej.2011.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Coates J.D., Achenbach L.A. Microbial perchlorate reduction: rocket-fuelled metabolism. Nat. Rev. Microbiol. 2004;2(7):569–580. doi: 10.1038/nrmicro926. [DOI] [PubMed] [Google Scholar]
- 63.Coates J.D., Michaelidou U., Bruce R.A., O'Connor S.M., Crespi J.N., Achenbach L.A. Ubiquity and diversity of dissimilatory (per)chlorate-reducing bacteria. Appl. Environ. Microbiol. 1999;65(12):5234–5241. doi: 10.1128/aem.65.12.5234-5241.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Scheid D., Stubner S., Conrad R. Identification of rice root associated nitrate, sulfate and ferric iron reducing bacteria during root decomposition. FEMS Microbiol. Ecol. 2004;50(2):101–110. doi: 10.1016/j.femsec.2004.06.001. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
These Whole Genome Shotgun projects of two strains affiliated with Dechloromonas have been deposited at the DDBJ/ENA/GenBank under the accession WYCU00000000 (s22) and WYCV00000000 (t15). The versions described in this paper are version WYCU01000000 (s22) and WYCV01000000 (t15).
These Whole Genome Shotgun projects of two strains affiliated with Dechloromonas have been deposited at the DDBJ/ENA/GenBank under the accession WYCU00000000 (s22) and WYCV00000000 (t15). The versions described in this paper are version WYCU01000000 (s22) and WYCV01000000 (t15).