Abstract
Acetyl-CoA carboxylase (ACC) is a crucial enzyme in fatty acid metabolism, which plays a major role in the occurrence and development of certain tumours. Herein, one potential ACC inhibitor (6a) was identified through high-throughput virtual screening (HTVS), and a series of 4-phenoxy-phenyl isoxazoles were synthesised for structure-activity relationship (SAR) studies. Among these compounds, 6g exhibited the most potent ACC inhibitory activity (IC50=99.8 nM), which was comparable to that of CP-640186. Moreover, the antiproliferation assay revealed that compound 6l exhibited the strongest cytotoxicity, with IC50 values of 0.22 µM (A549), 0.26 µM (HepG2), and 0.21 µM (MDA-MB-231), respectively. The preliminary mechanistic studies on 6g and 6l suggested that the compounds decreased the malonyl-CoA levels, arrested the cell cycle at the G0/G1 phase, and induced apoptosis in MDA-MB-231 cells. Overall, these results indicated that the 4-phenoxy-phenyl isoxazoles are potential for further study in cancer therapeutics as ACC inhibitors.
Keywords: Acetyl-CoA carboxylase, antitumor, docking, cell cycle, apoptosis
1. Introduction
Fatty acids are one of the essential biological macromolecules for cell growth and survival, which are widely involved in energy supply, membrane biosynthesis, and signal transduction1,2. In contrast to normal cells that preferentially use circulating lipids to satisfy their requirement for fatty acids, malignant cells primarily undergo high rates of de novo fatty acid synthesis (FASyn) to support their continuous proliferation and division3,4. Therefore, elevated FASyn is accepted as a major characteristic of numerous cancers, including those in the prostate, colon, breast, lung, and liver. The drugs that intervene with the enzymes in FASyn may provide an innovative approach for the fight against cancer5–8.
Acetyl-CoA carboxylase (ACC), a crucial enzyme in FASyn, has two subcellular specific isoforms, namely ACC1 and ACC2, which catalyse the carboxylation of acetyl-CoA to malonyl-CoA and display distinct physiological roles9. ACC1 is a cytosolic enzyme that mainly controls the FASyn process, with its product malonyl-CoA extends the fatty acids chain by two carbon increments under the catalysis of fatty acid synthase10. In contrast, the mitochondrial isoform ACC2 is primarily responsible for fatty acid oxidation (FAOxn) through the inhibition of carnitine palmitoyltransferase I (CPT-1) by localised malonyl-CoA production11. Thus, the functional abnormalities on ACC could provide a viable modality for disturbing the energy metabolism and causing cell damage, which facilitates the utilisation of ACC as an attractive therapeutic target12,13. Currently, cancer therapeutics mainly focuses on the ACC1 isoform due to the over-expression of ACC1 mRNA in most human cancers14–18.
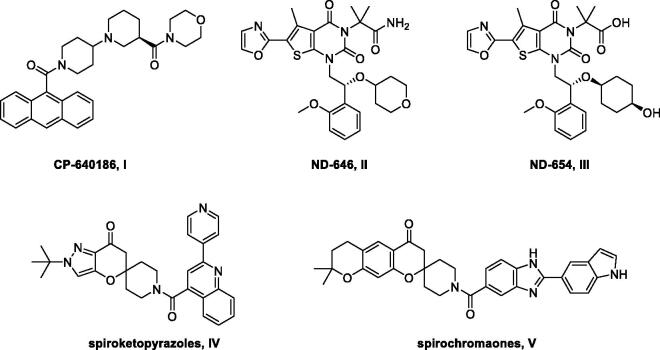
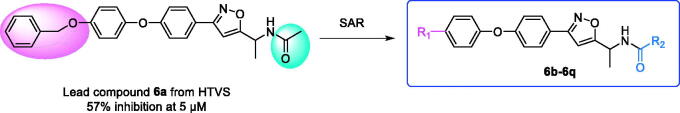
Since the first discovery of the mammalian ACC inhibitor CP-640186 in 2003 (Figure 1), a large amount of ACC inhibitors have been identified, some of which have even been approved in clinical trials for the treatment of glycolipid metabolic diseases, such as type 2 diabetes mellitus (T2DM) and non-alcoholic steatohepatitis (NASH)19–21. However, ACC inhibitors with effective anticancer activities were rarely reported. And most of these inhibitors contain thienopyrimidinone or spirocyclic scaffolds, among which the spiroketopyrazoles (IV) were previously identified by the author to display potent ACC1 inhibitory activity and cytotoxicity22–26. Therefore, an investigation into the generation of novel ACC inhibitors was conducted to further expand our research on antitumor agents and increase the diversity of ACC inhibitors. During the initial stage of this work, compound 6a (Figure 2), an initial high-throughput virtual screening (HTVS) hit with moderate ACC1 inhibitory potency (57% inhibition at 5 µM), was selected as a lead compound. Later, the structure–activity relationship (SAR) study of 4-phenoxy-phenyl isoxazoles was performed with the belief that these new chemical entities would effectively target the ACC enzyme and provide a new approach for cancer treatment.
Figure 1.
CP-640186 and representative ACC inhibitors with anticancer activity.
Figure 2.
The design strategy of 4-phenoxy-phenyl isoxazoles as novel ACC inhibitors.
Herein, starting from the HTVS hit 6a, a series of 4-phenoxy-phenyl isoxazole derivatives were identified. The p-benzyloxy group of 6a was substituted with various alkoxy groups to reduce the structural rigidity and increase the flexibility of target compounds. Furthermore, diverse amide and urea substituents were introduced to improve lipophilicity and increase the number of hydrogen bond donors. All of the compounds were evaluated for biological activity towards ACC1 enzyme and three different cancer cell lines (A549, HepG2, and MDA-MB-231). The preliminary docking and pharmacological studies of the representative compounds were carried out to elucidate the possible action mechanism of 4-phenoxy-phenyl isoxazoles.
2. Results and discussion
2.1. High-throughput virtual screening
To identify potent ACC inhibitors with new structural skeletons, 1,500,000 commercially available compounds from the ChemDiv database were docked into the active site of ACC in silico27,28. However, the co-crystal structure of ACC inhibitor and hACC1 has not been resolved. Therefore, we selected the co-crystal structure of CP-640186 and hACC2 (PDB ID: 3FF6) for HTVS due to the high degree of identity between ACC1 and ACC2 at the nucleotide level (60%) and the amino acid level (75%). Based on the docking results, 500 top-ranked compounds with high total scores were selected for the second round of screening, which mainly focussed on the binding modes and interactions between the candidates and the active site28. Finally, nine potential hits with new skeletons were obtained. Their corresponding properties are summarised in Table 1 (structures are provided in Figure S1). Thereafter, the selected compounds were purchased from the commercial vendors and evaluated for their ACC inhibitory activity using a luminescent ADP detection assay24. As shown in Table 1, compound 6a emerged as a top candidate, exhibiting the strongest inhibition rate at 5 µM. Thus, compound 6a was selected as a lead compound, and its benzyloxy and acetyl groups were modified for SAR studies (Figure 2).
Table 1.
The screening results and biological activity of selected candidates.
| Compounds | Total scorea | Crasha | Polara | hACC1 IRs (%)b |
|---|---|---|---|---|
| C450-0875 | 7.37 | −1.58 | 1.82 | 13.53 |
| C522-2022 | 9.71 | −1.21 | 4.04 | 2.84 |
| D074-0032 | 7.99 | −4.43 | 2.70 | 14.98 |
| D098-1432 (6a) | 8.07 | −1.25 | 2.08 | 57.39 |
| D400-1402 | 6.41 | −1.13 | 0.35 | 3.44 |
| D400-3199 | 6.77 | −1.05 | 0.99 | 6.93 |
| E667-1377 | 8.84 | −1.91 | 4.50 | 12.56 |
| F292-0452 | 7.77 | −0.95 | 2.43 | 6.98 |
| F697-0546 | 9.04 | −0.81 | 2.62 | 20.53 |
| CP-640186 | 7.18 | −1.05 | 2.08 | 86.83 |
aTotal score, crash, and polar values were calculated using Sybyl-X 2.1 software.
bInhibition rates (IRs) of compounds for hACC1 enzyme at 5 μM.
2.2. Chemistry
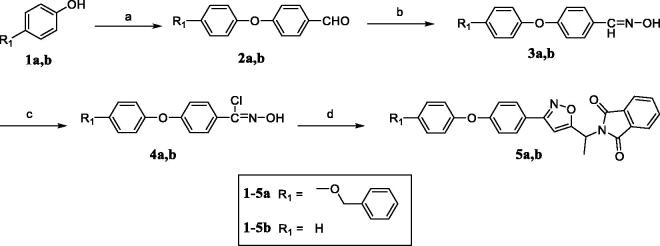
The 4-phenoxy-phenyl isoxazole derivatives (6a–6q) were synthesised according to the pathways described in Schemes 1 and 2. To brief, the commercially available 4-fluorobenzaldehyde was reacted with 1 (phenol or 4-(benzyloxy)phenol) in the presence of K2CO3 as the base at 120 °C to afford intermediate 2. Then, the addition–elimination reactions of 2 and hydroxylamine readily generated the aldoxime 3, which was subsequently chlorinated with N-chlorosuccinimide (NCS) to give the corresponding chloroaldoxime 4. The [3 + 2] dipolar cycloaddition of 4 and 2-(but-3-yn-2-yl)isoindoline-1,3-dione provided the key intermediate 5 in good yields (5a, 86%; 5b, 80%).
Scheme 1.
Synthetic route of key intermediate 5a and 5b. Reagents and conditions: (a) 4-fluorobenzaldehyde, K2CO3, DMF, 120 °C, 12 h; (b) hydroxylamine hydrochloride, NaOH, H2O/EtOH (3:1), 0 °C to r.t., 2 h; (c) NCS, DMF, r.t., 2 h; (d) 2-(but-3-yn-2-yl)isoindoline-1,3-dione, K2CO3, toluene, reflux, 12 h.
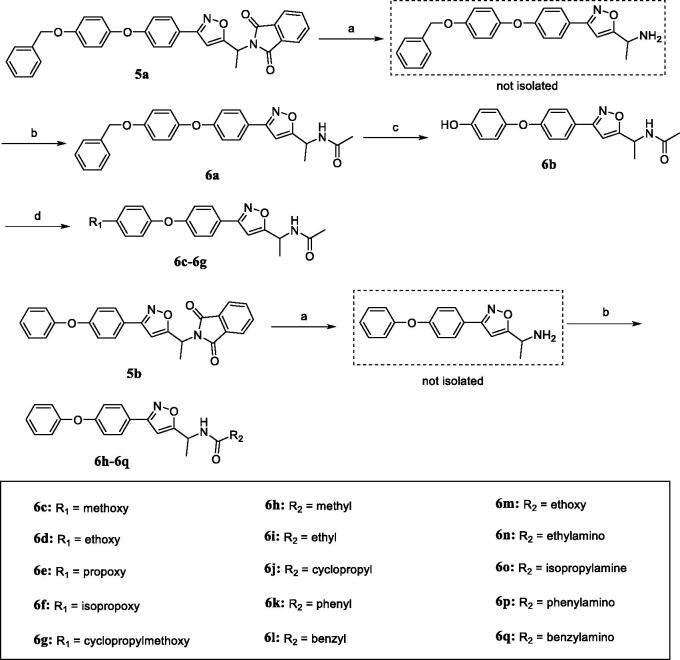
Scheme 2.
Synthetic route of target compounds 6a–6q. Reagents and conditions: (a) hydrazine hydrate, CH2Cl2/EtOH (5:1), reflux, 2 h; (b) appropriate acyl chloride or isocyanate, TEA, CH2Cl2, 0 °C to r.t., 6 h; (c) H2, Pd-C, MeOH, r.t., 3 h; (d) appropriate iodoalkane, Cs2CO3, DMF, reflux, 12 h.
The phthalimide protecting group of 5a was removed by the Ing-Manske reaction, and the resulting primary amine was condensed with acetyl chloride to afford the final compound 6a. Later, compound 6b was generated via Pd/C-catalysed hydrogenolysis. Moreover, the nucleophilic substitution of 6b and appropriate iodoalkane using caesium carbonate as catalyst gave the target compounds 6c–6g. The synthetic routes of the final compounds 6h–6q were similar to that of 6a, namely by the reaction of deprotected 5b with various acyl chlorides or isocyanates.
2.3. Biological evaluation
2.3.1. In vitro ACC inhibitory activity
To explore the SAR of 4-phenoxy-phenyl isoxazoles, the in vitro enzymatic inhibitory effects of 6a–6q were evaluated. The preliminary screening was carried out at two concentrations (5 µM and 0.1 µM), where CP-640186 was selected as the positive control29. As shown in Table 2, the derivatives 6b–6d with short-chain alkoxy groups at R1 exhibited weaker hACC1 inhibitory activity than 6a. In contrast, the propoxy-substituted 6e displayed 78.95 and 31.12% inhibition towards hACC1 at 5 and 0.1 µM, respectively, which indicated that the increase of the chain length is beneficial to the inhibition of ACC. Among the analogues with different substituents at R1 (6a–6h), compound 6g appeared to be the most potential ACC inhibitor, with inhibition rates (IRs) of 95.26 and 51.37% at 5 µM and 0.1 µM, respectively.
Table 2.
The hACC1 inhibitory activity and lipophilicity of compounds 6a–6q.
| Compounds | IRs (%)a |
IC50 (nM)b | cLogPc | |
|---|---|---|---|---|
| 5 μM | 0.1 μM | |||
| 6a | 54.93 | 18.13 | NT | 5.27 |
| 6b | 26.83 | 8.76 | NT | 3.31 |
| 6c | 37.32 | 9.89 | NT | 3.66 |
| 6d | 45.46 | 12.73 | NT | 4.14 |
| 6e | 78.95 | 31.12 | 438.2 ± 16.3 | 4.62 |
| 6f | 52.76 | 16.69 | NT | 4.40 |
| 6g | 95.26 | 51.37 | 99.8 ± 3.7 | 4.74 |
| 6h | 36.37 | 8.78 | NT | 3.57 |
| 6i | 51.83 | 18.51 | NT | 4.07 |
| 6j | 58.36 | 20.79 | NT | 4.13 |
| 6k | 72.47 | 28.11 | 626.3 ± 41.6 | 5.00 |
| 6l | 81.42 | 37.86 | 279.1 ± 13.1 | 5.08 |
| 6m | 16.77 | 3.57 | NT | 4.27 |
| 6n | 44.72 | 13.36 | NT | 4.02 |
| 6o | 49.82 | 17.90 | NT | 4.36 |
| 6p | 65.73 | 21.52 | 1055.0 ± 179.7 | 5.21 |
| 6q | 73.68 | 28.27 | 711.1 ± 52.6 | 5.18 |
| CP-640186 | 87.84 | 48.87 | 108.9 ± 6.2 | 3.59 |
aThe data represent the mean values of two independent experiments.
bIC50 values for ACC are presented as mean ± SD. NT: not test.
bcLogP values were calculated by Molsoft online software.
For 4-phenoxy-phenyl isoxazoles with various amide substituents at R2 (6h–6l), the analogues bearing aromatic ring showed better enzymatic activity, wherein compound 6l exhibited the strongest ACC inhibitory effect (IRs = 81.42 and 37.86%), which could be partly attributed to its bulky phenylacetamide group. Besides, the IRs of compound 6q with the benzylurea group (IRs = 73.68 and 28.27%) were the highest among the ureido-substituted derivatives (6n–6q), although slightly lower than that of the corresponding amide derivative 6l. The inhibitory activity of the carbamate-substituted compound 6m was hardly observed. These results suggested that the introduction of flexible alkoxy groups (such as propoxy and cyclopropylmethoxy, 6e and 6g) at the left-hand hydrophobic terminus, or the incorporation of bulky functional groups (such as phenylacetamide and benzylurea, 6l and 6q) at the right-hand polar end of the molecule was conducive to the inhibition of ACC.
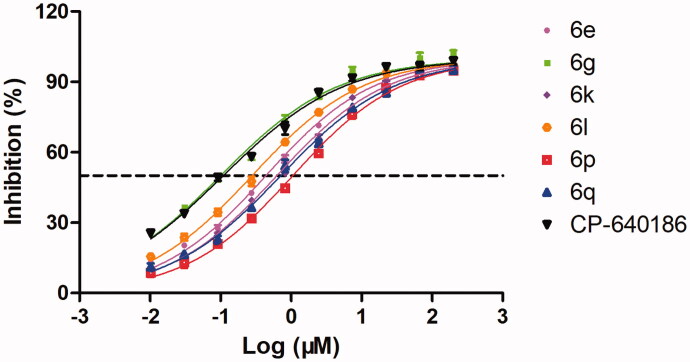
Based on the above results, the IC50 values of the preferred compounds (6e, 6g, 6k, 6l, 6p and 6q) with IRs greater than 60% at 5 µM were calculated and summarised in Table 2. Similar to the preliminary screening results, the cyclopropylmethoxy-substituted derivative 6g exhibited the optimal ACC inhibitory activity among the synthesised compounds. The IC50 value of 6g on the hACC1 enzyme was 99.8 nM, which was lower than that of CP-640186 (IC50=108.9 nM). The dose-response curves of the selected compounds were plotted, as presented in Figure 3.
Figure 3.
The dose–response curves of selected compounds on hACC1 enzyme.
2.3.2. In vitro cytotoxic activity
The cytotoxicity of 6a–6q on three different cancer cell lines with a high expression of ACC1 (A549, HepG2, and MDA-MB-231) was determined using the MTT assay24. The initial screening was performed at a single concentration of 10 µM, where doxorubicin (DOX) was served as a reference drug. As illustrated in Figure S2, the 4-phenoxyphenylisoxazoles showed the best antiproliferative activity on MDA-MB-231 cells, followed by A549 cells, and the weakest on HepG2 cells. Besides, most of the target compounds displayed significant inhibitory effects on cell proliferation. Among them, compounds 6a, 6g, and 6l exhibited inhibitory rates greater than 80% against the three cancer cell lines, which were comparable to that of DOX.
Based on the good initial screening results, the IC50 values and calculated logarithm of the octanol-water partition coefficient (cLogp) of 4-phenoxy-phenyl isoxazoles were determined for further SAR studies30. As shown in Table 3, when R2 was fixed as a methyl group, the derivatives 6a–6h exhibited varying degrees of antiproliferative activity. Among them, the compounds 6e and 6g with good ACC inhibitory activities exhibited satisfactory antitumor effects, in which 6g displayed the best inhibition on A549, HepG2, and MDA-MB-231 cells, with IC50 values of 1.10, 1.73, and 1.50 µM, respectively. Besides, compound 6a with moderate ACC activity and high lipophilicity showed considerable antiproliferative activity on the three cancer cell lines. However, the cytotoxicity of the derivatives (6b–6d, 6h) with relatively weak enzymatic activity and low lipophilicity decreased remarkably (IC50 >10 µM).
Table 3.
In vitro anti-proliferative activity of compounds 6a–6q.
| Compounds | IC50 (μM)a |
Cell viability (%) of HUVEC at 10 μMb | ||
|---|---|---|---|---|
| A549 | HepG2 | MDA-MB-231 | ||
| 6a | 1.40 ± 0.22 | 1.91 ± 0.66 | 1.57 ± 0.09 | 45.79 ± 1.16 |
| 6b | >10 | >10 | >10 | 88.14 ± 3.83 |
| 6c | >10 | >10 | >10 | 82.39 ± 4.85 |
| 6d | >10 | >10 | >10 | 89.06 ± 6.09 |
| 6e | 2.66 ± 0.47 | 3.48 ± 0.74 | 1.53 ± 0.38 | 83.68 ± 4.40 |
| 6f | >10 | >10 | 6.77 ± 0.74 | 34.96 ± 1.66 |
| 6g | 1.10 ± 0.12 | 1.73 ± 0.55 | 1.50 ± 0.58 | 87.33 ± 6.19 |
| 6h | >10 | >10 | >10 | 91.53 ± 2.58 |
| 6i | 6.02 ± 0.63 | 4.21 ± 0.53 | 2.97 ± 1.49 | 70.33 ± 0.25 |
| 6j | 2.66 ± 0.47 | 3.48 ± 0.74 | 1.13 ± 0.38 | 59.46 ± 0.95 |
| 6k | 2.61 ± 0.34 | 3.23 ± 0.89 | 0.57 ± 0.18 | 80.00 ± 2.47 |
| 6l | 0.22 ± 0.07 | 0.26 ± 0.02 | 0.21 ± 0.01 | 89.54 ± 3.21 |
| 6m | >10 | >10 | >10 | 79.12 ± 4.76 |
| 6n | 5.78 ± 0.86 | 9.04 ± 1.43 | 8.63 ± 1.14 | 83.23 ± 2.33 |
| 6o | 3.12 ± 0.19 | 2.31 ± 0.44 | 1.90 ± 0.52 | 77.65 ± 5.67 |
| 6p | 5.30 ± 0.46 | 1.80 ± 0.62 | 0.84 ± 0.08 | 85.60 ± 7.35 |
| 6q | 5.73 ± 0.11 | 2.47 ± 0.50 | 0.89 ± 0.26 | 84.92 ± 3.98 |
| DOX | 0.52 ± 0.07 | 0.67 ± 0.13 | 0.42 ± 0.09 | 18.95 ± 4.02 |
aIC50 values for A549, HepG2, and MDA-MB-231 cells are presented as mean ± SD (n = 3).
bCell viability was evaluated after incubation of compounds at a concentration of 10 μM.
On the other hand, when R1 was fixed as a hydrogen atom (6h–6q), most of the synthesised compounds exhibited potent in vitro antiproliferative activity. Of the five amide derivatives (6h–6l), compound 6l showed the best potency in cancer treatment, with IC50 values of 0.22, 0.26, and 0.21 µM for A549, HepG2, and MDA-MB-231 cells, respectively, which were apparently superior to DOX. The outstanding cell activity of 6l was supposed to be attributed to its optimal balance between the enzymatic activity and lipophilicity. Besides, the ureido-substituted compounds (6n–6q) exhibited remarkable performances towards the inhibition of MDA-MB-231 cells. The IC50 values of 6p (0.84 µM) and 6q (0.89 µM) against MDA-MB-231 cells were both less than 1 µM. In contrast, the carbamate-substituted 6m only provided a faint antitumor effect, suggesting that the carbamate group was detrimental to bioactivity.
To investigate the selectivity of the synthesised compounds towards cancer cells, the non-cancerous HUVEC cell line was selected for testing24. As shown in Table 2, most of the compounds exhibited weak toxic to HUVEC. Moreover, the compounds with strong ACC inhibitory and antitumor activities, namely 6e, 6g, 6k, 6l, 6p and 6q, showed less than 20% IRs towards HUVEC, which indicated the cancer cell selectivity of ACC inhibitors. Overall, the SAR studies revealed that the ACC inhibitory activity and lipophilicity of the target compounds play an important role in their antiproliferative activity.
2.3.3. Effect on intracellular malonyl-CoA levels
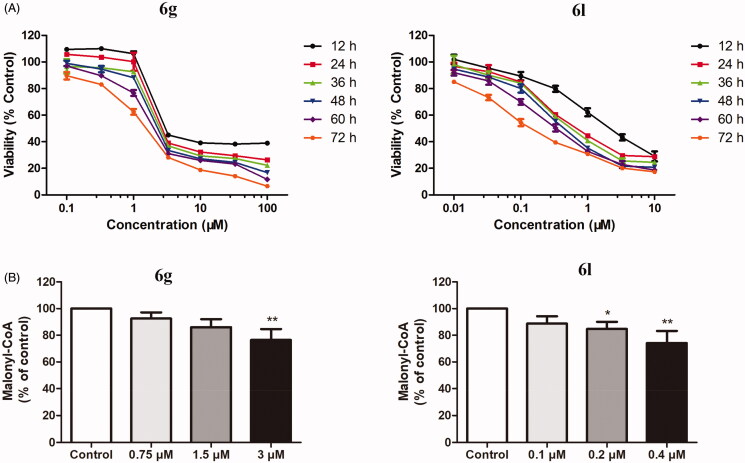
Due to the most potent ACC1 inhibitory activity of 6g and strongest antitumor activity of 6l, these compounds were selected for further biological studies. Moreover, MDA-MB-231 was chosen as the model cell line due to the best antiproliferative effect of 4-phenoxy-phenyl isoxazoles on these cells. As illustrated in Figure 4(A), compounds 6g and 6l inhibited the proliferation of MDA-MB-231 cells in a concentration- and time-dependent manner. Later, the effects of 6g and 6l on the levels of malonyl-CoA in MDA-MB-231 cells were investigated. The results suggested that the treatment of the cells with 6g and 6l significantly decreased the intracellular malonyl-CoA levels, with an approximate reduction of 23.59 and 25.85% compared to the control group at the maximum test concentration, respectively (Figure 4(B)).
Figure 4.
(A) Concentration- and time-dependent cytotoxicity of compounds 6g and 6l. The MDA-MB-231 cells were treated with drugs at increasing concentrations for 12, 24, 36, 48, 60, and 72 h, respectively. (B) Effects of 6g and 6l on the levels of malonyl-CoA in MDA-MB-231 cells. MDA-MB-231 cells were treated with diverse concentrations of drugs for 48 h, with the intracellular malonyl-CoA quantified. Data were expressed as mean ± SD (n = 3). *p < 0.05, **p < 0.01 versus control group.
2.3.4. Cell-cycle assay
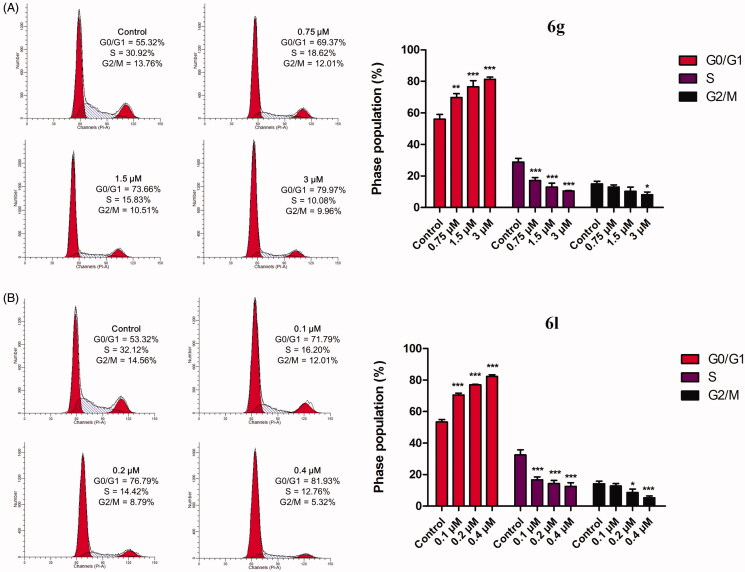
The cell-cycle arrest creates a stopping point at which the cells no longer participate in the process of duplication and division that represents an important approach for cancer treatment31,32. To investigate the cell-cycle progression, a flow cytometric analysis was performed on the MDA-MB-231 cells treated with various concentrations (1/2 IC50, IC50, and 2 × IC50) of 6g and 6l for 48 h. As shown in Figure 5, compound 6g led to an obvious increase in the cells at the G0/G1 phase from 55.32 to 79.97%, accompanied by a decrease in the cells at the S and G2/M phases from 30.93 and 13.76% to 10.08 and 9.96%, respectively. Moreover, the treatment of the MDA-MB-231 cells with increasing concentrations of 6l (0.1, 0.2, and 0.4 µM) increased the proportion of cells at the G0/G1 phase from 53.32% to 81.93%, while the proportion of cells at the S and G2/M phases decreased. These results confirmed that 6g and 6l induced cell-cycle arrest at the G0/G1 phase in a dose-dependent manner, which may be a possible mechanism for their antitumor activity.
Figure 5.
The effect of compounds 6g (A) and 6l (B) on cell-cycle distribution. MDA-MB-231 cells were treated with 6g and 6l at the indicated concentrations for 48 h, and stained with propidium iodide for flow cytometry. Data are expressed as mean ± SD (n = 3). *p < 0.05, **p < 0.01, and ***p < 0.001 versus control group.
2.3.5. Cell apoptosis assay
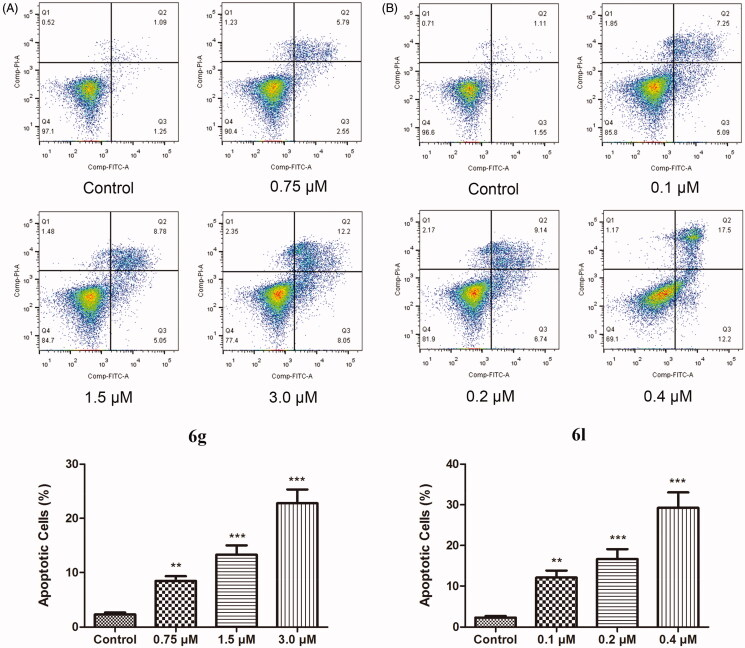
Apoptosis is a form of programmed cell death, which plays an important role in many cancer treatment strategies33,34. To explore the relationship between the antiproliferative effects of 4-phenoxy-phenyl isoxazoles and cell apoptosis, Annexin V and propidium iodide (PI) double staining was performed on the vehicle- and drug-treated MDA-MB-231 cells. Upon incubation with compound 6g (0.75, 1.5, and 3 µM) for 48 h, the total percentage of the early (bottom right quadrant) and late (upper right quadrant) apoptotic cells was increased from 2.34 to 8.34, 13.83, and 20.25%, respectively, as depicted in Figure 6. In addition, compound 6l induced early (from 1.55 to 12.20%) and late apoptosis (from 1.11 to 17.50%) in a dose-dependent manner. These data suggested that the antitumor activity of compounds 6g and 6l might be involved in the induction of apoptosis.
Figure 6.
The effect of compounds 6g (A) and 6l (B) on cell apoptosis. MDA-MB-231 cells were incubated with 6g and 6l at the indicated concentrations for 48 h, and stained with Annexin V-FITC/PI for flow cytometry. Data are expressed as mean ± SD (n = 3). **p < 0.01 and ***p < 0.001 versus control group.
2.4. Molecular docking
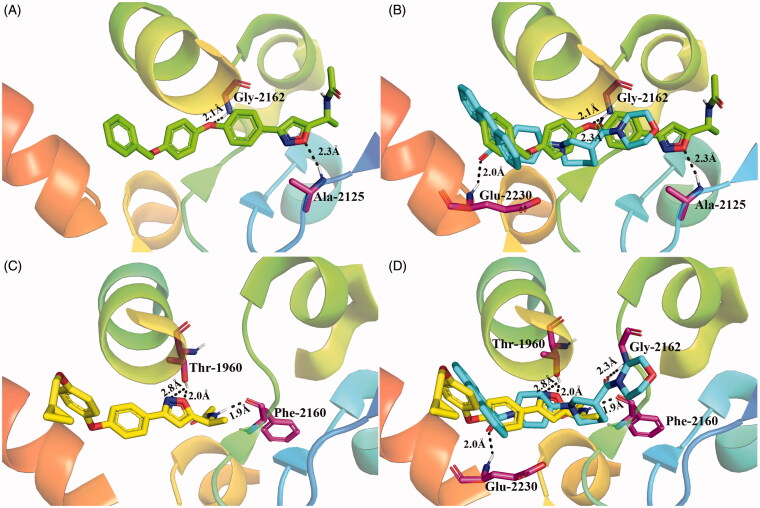
To gain insight into the specific interactions between the target compounds and ACC enzyme, molecular docking studies of the representative compounds (6a and 6g) were conducted using the co-crystal structure of ACC and CP-640186 (PDB ID: 3FF6) as the template27. As illustrated in Figure 7, compounds 6a and 6g were located at the dimer interface of the carboxyltransferase (CT) domain of ACC and exhibited different binding modes. Compared with CP-640186, 6a formed one conserved hydrogen bond with Gly-2162 (2.1 Å), and one additional hydrogen bond between the oxygen of oxazole and Ala-2125 (2.3 Å) was observed (Figure 7(A,B)). However, there was no hydrogen bond between 6a and Glu-2230, which partly explained its moderate ACC inhibitory activity. The substitution of a cyclopropylmethoxy group at R1 (6g) led to the movement of the binding site and formed three hydrogen bonds between 6g and the CT domain (Figure 7(C)). To our surprise, 6g and CP-640186 did not have the same hydrogen bond interaction with ACC (Figure 7(D)). In detail, the Thr-1960 residue formed two hydrogen bonds with the nitrogen (2.8 Å) and oxygen atoms (2.0 Å) of oxazole, and the Phe-2160 residue formed one hydrogen bond with the nitrogen of amide (1.9 Å). The stronger interaction of 6g with ACC than that of 6a might account for its better activity in enzyme inhibition.
Figure 7.
Molecular docking studies for compounds 6a and 6g (PDB ID: 3FF6). (A) Docking model of compound 6a (green) with the active site of ACC. (B) Overlapping conformation of 6a (green) with CP-640186 (cyan). (C) Docking model of compound 6g (yellow) with the active site of ACC. (D) Overlapping conformation of 6g (yellow) with CP-640186 (cyan). All hydrogen bonds are shown with black dashed lines, and related amino acids are highlighted with magenta sticks.
3. Conclusions
In this study, a series of structurally novel 4-phenoxy-phenyl isoxazoles were synthesised and evaluated for their antiproliferative activities towards a panel of human cancer cell lines as ACC inhibitors. The lead compound 6a was discovered from HTVS, and further SAR studies resulted in the identification of several derivatives with potent biological activity. Among them, compound 6g, which formed three hydrogen bonds with the CT domain of ACC, exhibited the most potent hACC1 inhibitory activity (IC50=99.8 nM). Compound 6l displayed the strongest antiproliferative activity, with IC50 values of 0.22, 0.26, and 0.21 µM against A549, HepG2, and MDA-MB-231 cells, respectively, which were superior to that of DOX. The best antitumor activity of compound 6l might be attributed to its optimal balance between the enzymatic activity and lipophilicity. Furthermore, treatment of the MDA-MB-231 cells with 6g and 6l significantly decreased the intracellular malonyl-CoA levels. The flow cytometry analysis confirmed that 6g and 6l arrested cell cycle at the G0/G1 phase and induced apoptosis in a dose-dependent manner. Overall, these results indicated that the 4-phenoxy-phenyl isoxazoles exhibited potent anticancer activity as novel ACC inhibitors, which demonstrated the causal relationship between ACC inhibition and loss of cell viability.
4. Experimental
4.1. Chemistry
All reagents and solvents were purchased from commercial sources without further purification. The reaction progress was monitored by TLC on Silica Gel 60 F254 plates, and the target compounds were purified by column chromatography with silica gel (200–300 mesh size). Melting points were measured in open capillary tubes using YRT-3 melting point apparatus and were uncorrected. 1H NMR and 13 C NMR spectra were recorded on JNM-ECZR 400 MHz spectrometer with tetramethyl silane (TMS) as an internal standard. High-resolution mass spectra (HRMS) were obtained from a (UHR-TOF) maXis 4 G instrument. Analytical HPLC was run on the Agilent 1260 HPLC instrument, equipped with Agilent SB-C18 column (Agilent Technologies, Palo Alto, CA) and UV detection at 250 nm. Eluent system was: 70% MeOH in H2O; flow rate = 0.2 ml/min.
4.1.1. General procedure for the preparation of key intermediate 5a–b
To a solution of phenol or 4-benzyloxy phenol (1, 27.5 mmol) in DMF (25 ml), K2CO3 (50 mmol) and 4-fluorobenzaldehyde (27.5 mmol) was added. After stirring at 120 °C for 12 h, the mixture was poured into water (100 ml) and extracted with ethyl acetate (3 × 25 ml). The combined organics were dried over anhydrous Na2SO4 and concentrated to give 2. Then, hydroxylamine hydrochloride (20 mmol) and 50% NaOH aqueous solution (40 mmol) was slowly added to a solution of 2 (20 mmol) in H2O/ethanol (3:1, 20 ml) at 0 °C. After stirring at room temperature for 2 h, the mixture was acidified to pH 6 by concentrated HCl and extracted with ethyl acetate (3 × 5 ml). The organic phases were dried over anhydrous Na2SO4 and concentrated in vacuo to afford 3. To a solution of oxime 3 (20 mmol) in DMF (25 ml), N-chlorosuccinimide (22 mmol) was added slowly at ambient temperature, the solution was stirred for 2 h before being poured into ice water (100 ml). The resulting precipitate was collected by filtration and dried to provide 4. Finally, 2-(But-3-yn-2-yl)isoindoline-1,3-dione (10 mmol), 4 (11 mmol), and K2CO3 (30 mmol) was dissolved in toluene (50 ml) and heated at reflux for 12 h. After cooling to room temperature, the mixture was filtered, concentrated, and purified by a silica gel flash column to give the key intermediate 5.
4.1.1.1. 2-(1-(3-(4-(4-(benzyloxy)phenoxy)phenyl)isoxazol-5-yl)ethyl)isoindoline-1,3-dione (5a)
White solid, yield 70%, m.p. 160.5–162.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.87–7.85 (m, 2H), 7.77–7.67 (m, 4H), 7.49–7.31 (m, 5H), 7.06–6.92 (m, 6H), 6.55 (d, J = 0.6 Hz, 1H), 5.70 (q, J = 7.3 Hz, 1H), 5.06 (s, 2H), 1.93 (d, J = 7.3 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 170.6, 167.3, 161.9, 160.1, 155.4, 149.4, 136.8, 134.3, 131.7, 128.6, 128.3, 128.0, 127.5, 123.5, 122.9, 121.2, 117.5, 116.0, 100.3, 70.5, 42.3, 16.8. HRMS m/z calcd for C32H24N2O5 [M + Na]+ 539.1577, found 539.1577.
4.1.1.2. 2-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)isoindoline-1,3-dione (5b)
White solid, yield 72%, m.p. 133.5–134.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.87–7.84 (m, 2H), 7.78–7.72 (m, 4H), 7.40–7.33 (m, 2H), 7.15 (t, J = 7.4 Hz, 1H), 7.08–7.01 (m, 4H), 6.57 (s, 1H), 5.70 (q, J = 7.0 Hz, 1H), 1.94 (d, J = 7.3 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 170.7, 167.3, 161.9, 159.0, 156.3, 134.3, 131.7, 129.9, 128.4, 123.9, 123.6, 119.5, 118.6, 100.3, 42.3, 16.8. HRMS m/z calcd for C26H24N2O4 [M + Na]+ 433.1159, found 433.1167.
4.1.2. General procedure for the preparation of target compounds 6a–6q
(A) Hydrazine monohydrate (8 mmol) was added dropwise to a solution of 5 (1.2 mmol) in methylene chloride/ethanol (5:1, 12 ml) and refluxed for 2 h. The mixture was extracted with methylene chloride (3 × 10 ml), dried over anhydrous Na2SO4, and concentrated in vacuo to give corresponding amines. The amine was dissolved in methylene chloride (8 ml) under an ice-bath. Then, triethylamine (3 mmol) and appropriate acyl chloride or isocyanate (1 mmol) were added sequentially at 0 °C, with the mixture stirred for 6 h. After removal of the solvent, the residue was purified by column chromatography to yield target compounds 6a and 6h–6q.
(B) To a solution of 6a (10 mmol) in anhydrous methanol (60 ml), Pd-C catalyst (10%, 1.5 mmol) was added, and the mixture was hydrogenated for 3 h at atmospheric pressure. Upon completion of the reaction, the catalyst was removed by filtration and the solvent was evaporated to give 6b.
(C) A mixture of 6b (1 mmol), Cs2CO3 (2 mmol), and corresponding iodoalkane (1.2 mmol) in DMF (10 ml) was stirred at reflux for 12 h. After cooling to room temperature, the mixture was poured into ice water (50 ml) and extracted with methylene dichloride (3 × 10 ml). The combined organic phases were dried over anhydrous Na2SO4, filtered, and concentrated under reduced pressure. The residue was purified by column chromatography to afford target compounds 6c–6g.
4.1.2.1. N-(1-(3-(4-(4-(Benzyloxy)phenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6a)
White solid, yield 70%, m.p. 152.5–153.5 °C. 1H NMR (400 MHz, DMSO-d6): δ 7.74–7.65 (m, 2H), 7.49–7.31 (m, 5H), 7.05–6.94 (m, 6H), 6.39 (s, 1H), 5.94 (d, J = 8.2 Hz, 1H), 5.44–5.33 (m, 1H), 5.06 (s, 2H), 2.03 (s, 3H), 1.57 (d, J = 7.1 Hz, 3H). 13 C NMR (100 MHz, DMSO-d6): δ 173.0, 169.5, 162.0, 160.3, 155.6, 149.6, 137.0, 128.7, 128.4, 128.2, 127.6, 123.0, 121.4, 117.6, 116.1, 99.1, 70.6, 42.3, 23.3, 19.6. HRMS m/z calcd for C26H24N2O4 [M + Na]+ 451.1628, found 451.1630. Purity: 99.3%.
4.1.2.2. N-(1-(3-(4-(4-Hydroxyphenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6b)
White solid, yield 65%, m.p. 103–105 °C. 1H NMR (400 MHz, CDCl3): δ 7.70 (dd, J = 13.5, 8.3 Hz, 2H), 7.12–6.79 (m, 6H), 6.40 (s, 1H), 5.90 (d, J = 16.1 Hz, 1H), 5.46–5.31 (m, 1H), 2.05 (s, 3H), 1.59 (d, J = 5.9 Hz, 3H). 13 C NMR (100 MHz, CDCl3): 13 C NMR (101 MHz,) δ 172.7, 170.1, 162.1, 160.6, 153.2, 148.7, 128.4, 121.6, 117.9, 117.4, 116.6, 99.3, 42.4, 23.3, 19.5. HRMS m/z calcd for C19H18N2O4 [M + Na]+ 361.1159, found 361.1162. Purity: 99.2%.
4.1.2.3. N-(1-(3-(4-(4-Methoxyphenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6c)
White solid, yield 70%, m.p. 127.5–128.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.81–7.60 (m, 2H), 7.03–6.91 (m, 6H), 6.40 (s, 1H), 5.97 (d, J = 6.2 Hz, 1H), 5.50–5.28 (m, 1H), 3.82 (s, 3H), 2.03 (s, 3H), 1.58 (d, J = 6.4 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 169.4, 162.0, 160.4, 156.4, 149.3, 128.4, 121.4, 118.0, 117.5, 115.1, 99.1, 55.8, 42.2, 23.3, 19.6. HRMS m/z calcd for C20H20N2O4 [M + Na]+ 375.1315, found 375.1321. Purity: 98.4%.
4.1.2.4. N-(1-(3-(4-(4-Ethoxyphenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6d)
White solid, yield 72%, m.p. 106.5–107.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.83–7.58 (m, 2H), 7.01–6.89 (m, 6H), 6.40 (s, 1H), 5.98 (d, J = 5.8 Hz, 1H), 5.50–5.25 (m, 1H), 4.07 (q, J = 12.0 Hz, 2H), 2.03 (s, 3H), 1.58 (d, J = 5.9 Hz, 3H), 1.43 (t, J = 6.3 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 169.5, 162.1, 160.4, 155.8, 149.1, 128.4, 121.4, 118.0, 117.5, 115.7, 99.1, 64.0, 42.2, 23.3, 19.6, 15.0. HRMS m/z calcd for C21H22N2O4 [M + Na]+ 389.1472, found 389.1477. Purity: 97.4%.
4.1.2.5. N-(1-(3-(4-(4-Propoxyphenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6e)
White solid, yield 68%, m.p. 118.5–119.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.71 (t, J = 8.2 Hz, 2H), 6.99 (t, J = 7.7 Hz, 4H), 6.91 (d, J = 8.8 Hz, 2H), 6.40 (s, 1H), 5.95 (d, J = 9.3 Hz, 1H), 5.47–5.28 (m, 1H), 3.92 (t, J = 4.5 Hz, 2H), 2.04 (s, 3H), 1.88–1.75 (m, 2H), 1.58 (d, J = 5.9 Hz, 3H), 1.06 (t, J = 6.2 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 169.5, 162.1, 160.5, 156.0, 149.1, 128.4, 121.4, 117.9, 117.5, 115.7, 99.2, 70.1, 42.3, 23.3, 22.7, 19.6, 10.6. HRMS m/z calcd for C22H24N2O4 [M + Na]+ 403.1628, found 403.1633. Purity: 99.1%.
4.1.2.6. N-(1-(3-(4-(4-Isopropoxyphenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6f)
White solid, yield 65%, m.p. 124–126 °C. 1H NMR (400 MHz, CDCl3): δ 7.71 (t, J = 8.9 Hz, 2H), 7.07–6.81 (m, 6H), 6.40 (s, 1H), 5.95 (d, J = 6.8 Hz, 1H), 5.49–5.25 (m, 1H), 4.51 (s, 1H), 2.04 (s, 3H), 1.58 (d, J = 5.4 Hz, 3H), 1.37 (dd, J = 15.2, 5.6 Hz, 6H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 169.5, 162.1, 160.4, 154.7, 149.1, 128.4, 121.4, 118.1, 117.6, 117.3, 99.1, 70.6, 42.2, 23.3, 22.2, 19.6. HRMS m/z calcd for C22H24N2O4 [M + Na]+ 403.1628, found 403.1633. Purity: 98.8%.
4.1.2.7. N-(1-(3-(4-(4-(Cyclopropylmethoxy)phenoxy)phenyl)isoxazol-5-yl)ethyl)acetamide (6g)
White solid, yield 50%, m.p. 131.5–132.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.71 (t, J = 9.0 Hz, 2H), 7.14–6.81 (m, 6H), 6.40 (s, 1H), 5.89 (d, J = 8.4 Hz, 1H), 5.55–5.24 (m, 1H), 3.84 (dd, J = 33.2, 5.9 Hz, 2H), 2.04 (s, 3H), 1.58 (d, J = 6.7 Hz, 3H), 1.40–1.19 (m, 1H), 0.73–0.57 (m, 2H), 0.45–0.30 (m, 2H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 169.4, 162.0, 160.4, 155.9, 149.2, 128.4, 121.4, 118.0, 117.5, 115.84, 99.1, 73.4, 42.3, 23.3, 19.6, 10.4, 3.3. HRMS m/z calcd for C23H24N2O4 [M + Na]+ 415.1628, found 415.1634. Purity: 99.3%.
4.1.2.8. N-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)acetamide (6h)
White solid, yield 85%, m.p. 118.5–119.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.76–7.68 (m, 2H), 7.40–7.32 (m, 2H), 7.19–7.11 (m, 1H), 7.07–7.00 (m, 4H), 6.40 (s, 1H), 6.01 (d, J = 71.1 Hz, 1H), 5.45–5.31 (m, 1H), 2.03 (s, 3H), 1.57 (d, J = 7.1 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 173.0, 169.4, 161.9, 159.2, 156.3, 129.9, 128.4, 124.0, 123.5, 119.6, 118.6, 99.1, 42.2, 23.2, 19.5. HRMS m/z calcd for C19H18N2O3 [M + Na]+ 345.1210, found 345.1225. Purity: 96.9%.
4.1.2.9. N-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)propionamide (6i)
White solid, yield 83%, m.p. 129.5–130.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.78–7.68 (m, 2H), 7.40–7.36 (m, 2H), 7.21–7.12 (m, 1H), 7.09–7.00 (m, 4H), 6.41 (s, 1H), 5.87 (d, J = 7.8 Hz, 1H), 5.46–5.35 (m, 1H), 2.26 (q, J = 7.6 Hz, 2H), 1.59 (d, J = 7.1 Hz, 3H), 1.18 (t, J = 7.6 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 173.1, 173.1, 162.0, 159.3, 156.4, 130.0, 128.5, 124.1, 123.7, 119.6, 118.7, 99.1, 42.1, 29.6, 19.6, 9.7. HRMS m/z calcd for C20H20N2O3 [M + Na]+ 359.1366, found 359.1369. Purity: 98.2%.
4.1.2.10. N-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)cyclopropanecarboxamide (6j)
White solid, yield 75%, m.p. 168.5–169.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.79–7.70 (m, 2H), 7.44–7.33 (m, 2H), 7.18–7.14 (m, 1H), 7.10–7.01 (m, 4H), 6.41 (s, 1H), 6.09 (d, J = 8.3 Hz, 1H), 5.45–5.38 (m, 1H), 1.60 (d, J = 7.1 Hz, 3H), 1.43–1.32 (m, 1H), 1.06–0.94 (m, 2H), 0.81–0.77 (m, 2H). 13 C NMR (100 MHz, CDCl3): δ 173.3, 162.0, 159.2, 156.4, 130.0, 128.5, 124.1, 123.7, 119.6, 118.7, 99.1, 42.3, 19.7, 14.8, 7.7. HRMS m/z calcd for C21H20N2O3 [M + Na]+ 371.1366, found 371.1367. Purity: 100%.
4.1.2.11. N-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)benzamide (6k)
White solid, yield 72%, m.p. 155.5–156.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.80 (dd, J = 5.3, 3.3 Hz, 2H), 7.76–7.71 (m, 2H), 7.56–7.50 (m, 1H), 7.48–7.42 (m, 2H), 7.40–7.34 (m, 2H), 7.19–7.12 (m, 1H), 7.07–7.03 (m, 4H), 6.52 (d, J = 7.7 Hz, 1H), 6.47 (s, 1H), 5.64–5.57 (m, 1H), 1.70 (d, J = 7.1 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 173.0, 166.8, 162.1, 159.3, 156.4, 133.8, 132.0, 130.0, 128.8, 128.5, 127.2, 124.1, 123.6, 119.6, 118.7, 99.3, 42.8, 19.7. HRMS m/z calcd for C24H20N2O3 [M + Na]+ 407.1366, found 407.1368. Purity: 99.0%.
4.1.2.12. N-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)-2-phenylacetamide (6l)
White solid, yield 70%, m.p. 164.5–165.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.70 (d, J = 8.6 Hz, 2H), 7.41–7.26 (m, 7H), 7.16 (t, J = 7.3 Hz, 1H), 7.05 (dd, J = 11.0, 4.4 Hz, 4H), 6.27 (s, 1H), 5.78 (d, J = 8.2 Hz, 1H), 5.41–5.33 (m, 1H), 3.62 (s, 2H), 1.50 (d, J = 7.0 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 172.9, 170.4, 161.9, 159.3, 156.4, 134.5, 130.0, 129.5, 129.2, 128.5, 127.7, 124.1, 123.6, 119.7, 118.7, 99.0, 43.8, 42.4, 19.4. HRMS m/z calcd for C25H22N2O3 [M + Na]+ 421.1523, found 421.1527. Purity: 96.7%.
4.1.2.13. Ethyl (1-(3-(4-phenoxyphenyl)isoxazol-5-yl)ethyl)carbamate (6m)
White solid, yield 70%, m.p. 106.5–107.5 °C. 1H NMR (400 MHz, CDCl3): δ 7.74 (d, J = 7.5 Hz, 2H), 7.38 (dd, J = 9.2, 4.2 Hz, 2H), 7.29–7.23 (m, 1H), 7.16 (dd, J = 7.7, 6.8 Hz, 1H), 7.10–6.99 (m, 4H), 6.56–6.34 (m, 1H), 5.27–4.83 (m, 1H), 4.15 (q, J = 8.6 Hz, 2H), 1.59 (d, J = 3.0 Hz, 3H), 1.26 (t, J = 6.5 Hz, 3H). 13 C NMR (100 MHz, CDCl3): δ 173.5, 161.9, 159.2, 156.4, 155.7, 130.0, 128.5, 124.1, 123.7, 119.6, 118.7, 98.9, 61.4, 44.2, 19.9, 14.6. HRMS m/z calcd for C25H22N2O3 [M + Na]+ 421.1523, found 421.1527. HRMS m/z calcd for C20H20N2O4 [M + Na]+ 375.1315, found 375.1320. Purity: 99.4%.
4.1.2.14. 1-Ethyl-3-(1-(3-(4-phenoxyphenyl)isoxazol-5-yl)ethyl)urea (6n)
White solid, yield 80%, m.p. 158.5–159.5 °C. 1H NMR (400 MHz, DMSO-d6): δ 7.91–7.80 (m, 2H), 7.50–7.37 (m, 2H), 7.27–7.17 (m, 1H), 7.15–7.04 (m, 4H), 6.78 (s, 1H), 6.48 (d, J = 8.3 Hz, 1H), 5.87 (t, J = 5.6 Hz, 1H), 5.03–4.96 (m, 1H), 3.03 (q, J = 3.6 Hz, 2H), 1.42 (d, J = 7.1 Hz, 3H), 1.00 (t, J = 7.2 Hz, 3H). 13 C NMR (100 MHz, DMSO-d6): δ 176.5, 161.6, 159.0, 157.5, 156.3, 130.8, 129.0, 124.7, 124.1, 120.0, 119.0, 99.0, 42.8, 34.7, 20.5, 16.2. HRMS m/z calcd for C20H21N3O3 [M + Na]+ 374.1475, found 374.1480. Purity: 98.6%.
4.1.2.15. 1-Isopropyl-3-(1-(3-(4-phenoxyphenyl)isoxazol-5-yl)ethyl)urea (6o)
White solid, yield 75%, m.p. 197.5–198.5 °C. 1H NMR (400 MHz, DMSO-d6): δ 7.91–7.79 (m, 2H), 7.48–7.39 (m, 2H), 7.25–7.18 (m, 1H), 7.16–7.03 (m, 4H), 6.77 (s, 1H), 6.34 (d, J = 8.4 Hz, 1H), 5.75 (d, J = 7.7 Hz, 1H), 5.08–4.91 (m, 1H), 3.72–3.64 (m, 1H), 1.42 (d, J = 7.1 Hz, 3H), 1.04 (dd, J = 6.5, 1.8 Hz, 6H). 13 C NMR (100 MHz, DMSO-d6): δ 176.4, 161.6, 159.0, 156.9, 156.3, 130.8, 129.0, 124.7, 124.1, 120.0, 119.0, 99.0, 42.7, 41.5, 23.7, 23.7, 20.6. HRMS m/z calcd for C21H23N3O3 [M + Na]+ 388.1632, found 388.1638. Purity: 100%.
4.1.2.16. 1-(1-(3-(4-Phenoxyphenyl)isoxazol-5-yl)ethyl)-3-phenylurea (6p)
White solid, yield 73%, m.p. 195.5–196.5 °C. 1H NMR (400 MHz, DMSO-d6): δ 8.48 (s, 1H), 7.94–7.81 (m, 2H), 7.50–7.35 (m, 4H), 7.22 (dd, J = 16.1, 7.8 Hz, 3H), 7.14–7.05 (m, 4H), 6.96–6.86 (m, 2H), 6.80 (d, J = 8.1 Hz, 1H), 5.12–5.07 (m, 1H), 1.51 (d, J = 7.0 Hz, 3H). 13 C NMR (100 MHz, DMSO-d6): δ 176.4, 161.6, 159.0, 156.9, 156.3, 130.8, 129.0, 124.7, 124.1, 120.0, 119.0, 99.0, 42.7, 41.5, 23.7, 23.7, 20.6. HRMS m/z calcd for C24H21N3O3 [M + Na]+ 422.1475, found 422.1480. Purity: 96.7%.
4.1.2.17. 1-Benzyl-3-(1-(3-(4-phenoxyphenyl)isoxazol-5-yl)ethyl)urea (6q)
White solid, yield 76%, m.p. 161.5–162.5 °C. 1H NMR (400 MHz, DMSO-d6): δ 7.93–7.78 (m, 2H), 7.54–7.39 (m, 2H), 7.35–7.19 (m, 6H), 7.14–7.05 (m, 4H), 6.78 (s, 1H), 6.64 (d, J = 8.3 Hz, 1H), 6.42 (t, J = 6.0 Hz, 1H), 5.07–4.99 (m, 1H), 4.32–4.14 (m, 2H), 1.45 (d, J = 7.1 Hz, 3H). 13 C NMR (100 MHz, DMSO-d6): δ 176.4, 161.6, 159.0, 157.7, 156.3, 141.2, 130.8, 129.0, 128.8, 127.6, 127.2, 124.7, 124.1, 120.0, 119.0, 99.1, 43.5, 43.0, 20.5. HRMS m/z calcd for C25H23N3O3 [M + Na]+ 436.1632, found 436.1636. Purity: 98.9%.
4.2. HTVS
The ChemDiv database was provided by Topscience Co., and the HTVS was performed using the Surflex docking module in Sybyl-X 2.128. The crystal complex of ACC and CP-640186 (PDB ID: 3FF6) was downloaded from the Protein Data Bank. To improve the efficiency of screening, the maximum quantity of conformations was reduced from 20 to 10, and the maximum quantity of rotatable bonds was decreased from 100 to 50, with other parameters kept as usual. Compounds with high total-scores (top 500) were extracted to observe their binding modes and interactions with the CT domain. Finally, nine candidates with new scaffolds were obtained. The cLogp values and drug-likeness scores were calculated by Molsoft software (http://www.molsoft.com/mprop/).
4.3. ACC inhibitory assay
The hACC1 inhibitory activity of target compounds was evaluated using the ADP-GloTM kinase assay according to our previous report24. In brief, 4.5 µL of working solution containing recombinant ACC1 (BPS Biosciences, 50200) and 0.5 µL of diluted compound solution were added to a 384-well Optiplate plate (Perkin Elmer, 6007290, PerkinElmer, Waltham, MA, USA), and incubated at room temperature for 15 min. Then, 5 µL of substrate mixture was added to the 384-well plate to start the reaction. After incubating for 60 min at room temperature, 10 µL of ADP-Glo reagents were added to the 384-well plate, and incubated for another 40 min to deplete the remaining ATP. Finally, 20 µL of kinase detection reagents were added to the plate and incubated for 40 min to convert ADP to ATP. The luminescent signal of ATP was determined using an Envision multifunction reader (Perkin Elmer, 2104–0010, PerkinElmer, Waltham, MA, USA). The IC50 values were calculated using GraphPad Prism 5 software (GraphPad Software, La Jolla, CA, USA).
4.4. Molecular docking
The docking study was carried out with the Sybyl-X 2.1 software, and the crystal complex of ACC and CP-640186 (PDB ID: 3FF6) was downloaded from the Protein Data Bank. The whole waters and superfluous ligands were removed, and hydrogen atoms were added to the original protein structure. For the docked ligands, they were added with non-polar hydrogens and further energy was minimised with the Tripos force field. Then, the optimised ligands were docked to the prepared protein using a Surflex-Dock protocol. The optimal binding pose of the ligand was determined by the total score and its interaction with ACC. The best docking conformations of 6a and 6g, as well as their corresponding overlapping conformations with CP-640186 was visualised by PyMOL.
4.5. MTT assay
The cytotoxicity of target compounds was assessed by MTT assay. Briefly, cells were seeded in 96-well plates at a density of 2500–3000 per well and incubated for 24 h in a CO2 incubator. Then, the cells were treated with candidate compounds at seven different concentrations for 72 h. At the end of treatment, the MTT reagent (5 mg/mL) was added to determine the cell viability. After 4 h of incubation, the resulting medium was removed, and 100 µL of DMSO was added to dissolve the formed formazan. Finally, the absorbance at 550 nm was measured by a microplate reader (Varioskan LUX, Promega, Madison, WI), with the IC50 values calculated using GraphPad Prism 5 software (GraphPad Software, La Jolla, CA, USA).
4.6. Measurement of malonyl-CoA levels
MDA-MB-231 cells were seeded on 6-well plates and treated with various concentrations of 6g and 6l for 48 h. Cells were collected, washed with PBS, and lysed with 1% Triton X-100. The lysates were centrifuged at 4 °C for 15 min. The malonyl-CoA levels were determined using Elisa assay kits (Ruixin Biotech, Beijing, China).
4.7. Cell-cycle assay
MDA-MB-231 cells were seeded on 6-well plates (3 × 104 cells/well) and incubated with indicated concentrations of 6g (0.75, 1.5, and 3 µM) and 6l (0.1, 0.2, and 0.4 µM) for 48 h in a 5% CO2 incubator. Then, the cells were collected by trypsinization and centrifugation, followed by fixing in 70% pre-cooled ethanol overnight. Finally, the cells were stained with PI/RNase A according to the manual and measured by flow cytometer (Facs Canto II, Becton Dickinson, San Jose, CA’, USA)35.
4.8. Cell apoptosis assay
MDA-MB-231 cells were plated on 6-well plates (3 × 104 cells/well) and incubated with various concentrations of 6g (0.75, 1.5, and 3 µM) and 6l (0.1, 0.2, and 0.4 µM) for 48 h. Then, cells were harvested by trypsinization and centrifugation, followed by staining with annexin-V-FITC in binding buffer for 15 min protected from light. Subsequently, the cells were stained with PI, and the proportion of apoptotic cells was measured by flow cytometer (Facs Canto II, Becton Dickinson, San Jose, CA, USA)36.
Supplementary Material
Funding Statement
This study was supported by National Natural Science Foundation of China (81671744), Natural Science Foundation of Jiangsu Province (BK20171184), and Science and Technology Planning Project of Xuzhou, China (KC20104).
Disclosure statement
No potential conflict of interest was reported by the author(s).
References
- 1.Braig S. Chemical genetics in tumor lipogenesis. Biotechnol Adv 2018;36:1724–9. [DOI] [PubMed] [Google Scholar]
- 2.Rohrig F, Schulze A.. The multifaceted roles of fatty acid synthesis in cancer. Nat Rev Cancer 2016;16:732–49. [DOI] [PubMed] [Google Scholar]
- 3.Hanahan D, Weinberg RA.. Hallmarks of cancer: the next generation. Cell 2011;144:646–74. [DOI] [PubMed] [Google Scholar]
- 4.Ngoi NYL, Eu JQ, Hirpara J, et al. Targeting cell metabolism as cancer therapy. Antioxid Redox Signal 2020;32:285–308. [DOI] [PubMed] [Google Scholar]
- 5.Tripathi SC, Fahrmann JF, Vykoukal JV, et al. Targeting metabolic vulnerabilities of cancer: small molecule inhibitors in clinic. Cancer Rep 2019;2:e1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wang W, Bai L, Li W, et al. The lipid metabolic landscape of cancers and new therapeutic perspectives. Front Oncol 2020;10:605154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Corn KC, Windham MA, Rafat M.. Lipids in the tumor microenvironment: from cancer progression to treatment. Prog Lipid Res 2020;80:101055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Montesdeoca N, Lopez M, Ariza X, et al. Inhibitors of lipogenic enzymes as a potential therapy against cancer. Faseb J 2020;34:11355–81. [DOI] [PubMed] [Google Scholar]
- 9.Chen LY, Duan YQ, Wei HQ, et al. Acetyl-CoA carboxylase (ACC) as a therapeutic target for metabolic syndrome and recent developments in ACC1/2 inhibitors. Expert Opin Investig Drug 2019;28:917–30. [DOI] [PubMed] [Google Scholar]
- 10.Tong L. Acetyl-coenzyme A carboxylase: crucial metabolic enzyme and attractive target for drug discovery. Cell Mol Life Sci 2005;62:1784–803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tong L, Harwood HJ Jr.. Acetyl-coenzyme A carboxylases: versatile targets for drug discovery. J Cell Biochem 2006;99:1476–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wang C, Ma J, Zhang N, et al. The acetyl-CoA carboxylase enzyme: a target for cancer therapy? Expert Rev Anticancer 2015;15:667–76. [DOI] [PubMed] [Google Scholar]
- 13.Wu X, Huang T.. Recent development in acetyl-CoA carboxylase inhibitors and their potential as novel drugs. Future Med Chem 2020;12:533–61. [DOI] [PubMed] [Google Scholar]
- 14.Li EQ, Zhao W, Zhang C, et al. Synthesis and anti-cancer activity of ND-646 and its derivatives as acetyl-CoA carboxylase 1 inhibitors. Eur J Pharm Sci 2019;137:105010. [DOI] [PubMed] [Google Scholar]
- 15.Saab J, Santos-Zabala ML, Loda M, et al. Fatty acid synthase and acetyl-CoA carboxylase are expressed in nodal metastatic melanoma but not in benign intracapsular nodal nevi. Am J Dermatopathol 2018;40:259–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Yahagi N, Shimano H, Hasegawa K, et al. Co-ordinate activation of lipogenic enzymes in hepatocellular carcinoma. Eur J Cancer 2005;41:1316–22. [DOI] [PubMed] [Google Scholar]
- 17.Chajes V, Cambot M, Moreau K, et al. Acetyl-CoA carboxylase alpha is essential to breast cancer cell survival. Cancer Res 2006;66:5287–94. [DOI] [PubMed] [Google Scholar]
- 18.Petrova E, Scholz A, Paul J, et al. Acetyl-CoA carboxylase inhibitors attenuate wnt and hedgehog signaling and suppress pancreatic tumor growth. Oncotarget 2017;8:48660–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Griffith DA, Kung DW, Esler WP, et al. Decreasing the rate of metabolic ketone reduction in the discovery of a clinical acetyl-CoA carboxylase inhibitor for the treatment of diabetes. J Med Chem 2014;57:10512–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kim CW, Addy C, Kusunoki J, et al. Acetyl CoA carboxylase inhibition reduces hepatic steatosis but elevates plasma triglycerides in mice and humans: a bedside to bench investigation. Cell Metab 2017;26:394–406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lawitz EJ, Poordad F, Coste A, et al. Acetyl-CoA carboxylase (acc) inhibitor GS-0976 leads to suppression of hepatic de novo lipogenesis and significant improvements in MRI-PDFF, MRE, and markers of fibrosis after 12 weeks of therapy in patients with NASH. J Hepatol 2017;66:S34. [Google Scholar]
- 22.Wei Q, Mei L, Chen P, et al. Design, synthesis and biological evaluation of novel chroman derivatives as non-selective acetyl-CoA carboxylase inhibitors. Bioorg Chem 2020;101:103943. [DOI] [PubMed] [Google Scholar]
- 23.Wei Q, Mei L, Yang Y, et al. Design, synthesis and biological evaluation of novel spiro-pentacylamides as acetyl-CoA carboxylase inhibitors. Bioorg Med Chem 2018;26:3866–74. [DOI] [PubMed] [Google Scholar]
- 24.Huang T, Wu X, Yan S, et al. Synthesis and in vitro evaluation of novel spiroketopyrazoles as acetyl-CoA carboxylase inhibitors and potential antitumor agents. Eur J Med Chem 2021;212:113036. [DOI] [PubMed] [Google Scholar]
- 25.Lally JSV, Ghoshal S, DePeralta DK, et al. Inhibition of acetyl-CoA carboxylase by phosphorylation or the inhibitor ND-654 suppresses lipogenesis and hepatocellular carcinoma. Cell Metab 2019;29:174–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Svensson RU, Parker SJ, Eichner LJ, et al. Inhibition of acetyl-CoA carboxylase suppresses fatty acid synthesis and tumor growth of non-small-cell lung cancer in preclinical models. Nat Med 2016;22:1108–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Madauss KP, Burkhart WA, Consler TG, et al. The human ACC2 CT-domain C-terminus is required for full functionality and has a novel twist. Acta Crystallogr D 2009;65:449–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Huang T, Sun J, Zhou S, et al. Identification of direct activator of adenosine monophosphate-activated protein kinase (AMPK) by structure-based virtual screening and molecular docking approach. Int J Mol Sci 2017;18:1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Harwood HJ, Petras SF, Shelly LD, et al. Isozyme-nonselective n-substituted bipiperidylcarboxamide acetyl-CoA carboxylase inhibitors reduce tissue malonyl-CoA concentrations, inhibit fatty acid synthesis, and increase fatty acid oxidation in cultured cells and in experimental animals. J Biol Chem 2003;278:37099–111. [DOI] [PubMed] [Google Scholar]
- 30.Huang TH, Wu X, Liu TY, et al. Synthesis and anticancer activity evaluation of novel oxacalix[2]arene[2]pyrimidine derivatives. Med Chem Res 2019;28:580–90. [Google Scholar]
- 31.Icard P, Fournel L, Wu Z, et al. Interconnection between metabolism and cell cycle in cancer. Trends Biochem Sci 2019;44:490–501. [DOI] [PubMed] [Google Scholar]
- 32.Leal-Esteban LC, Fajas L.. Cell cycle regulators in cancer cell metabolism. BBA Mol Basis Dis 2020;1866:165715. [DOI] [PubMed] [Google Scholar]
- 33.Carneiro BA, El-Deiry WS.. Targeting apoptosis in cancer therapy. Nat Rev Clin Oncol 2020;17:395–417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.D'Arcy MS. Cell death: a review of the major forms of apoptosis, necrosis and autophagy. Cell Biol Int 2019;43:582–92. [DOI] [PubMed] [Google Scholar]
- 35.Wang C, Li Y, Liu Z, et al. Design, synthesis and biological evaluation of 1-Aryl-5-(4-arylpiperazine-1-carbonyl)-1H-tetrazols as novel microtubule destabilizers. J Enzyme Inhib Med Chem 2021;36:549–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Han X, Yu YL, Ma D, et al. Synthesis, telomerase inhibitory and anticancer activity of new 2-phenyl-4H-chromone derivatives containing 1,3,4-oxadiazole moiety. J Enzyme Inhib Med Chem 2021;36:344–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.