Abstract
During nutritional overload and obesity, hepatocyte function is grossly altered, and a subset of hepatocytes begins to accumulate fat droplets, leading to nonalcoholic fatty liver disease (NAFLD). Recent single-cell studies revealed how nonparenchymal cells, such as macrophages, hepatic stellate cells, and endothelial cells, heterogeneously respond to NAFLD. However, it remains to be characterized how hepatocytes, the major constituents of the liver, respond to nutritional overload in NAFLD. Here, using droplet-based, single-cell RNA sequencing (Drop-seq), we characterized how the transcriptomic landscape of individual hepatocytes is altered in response to high-fat diet (HFD) and NAFLD. We showed that the entire hepatocyte population undergoes substantial transcriptome changes upon HFD, although the patterns of alteration were highly heterogeneous, with zonation-dependent and -independent effects. Periportal (zone 1) hepatocytes downregulated many zone 1-specific marker genes, whereas a small number of genes mediating gluconeogenesis were upregulated. Pericentral (zone 3) hepatocytes also downregulated many zone 3-specific genes; however, they upregulated several genes that promote HFD-induced fat droplet formation, consistent with findings that zone 3 hepatocytes accumulate more lipid droplets. Zone 3 hepatocytes also upregulated ketogenic pathways as an adaptive mechanism to HFD. Interestingly, many of the top HFD-induced genes, which encode proteins regulating lipid metabolism, were strongly co-expressed with each other in a subset of hepatocytes, producing a variegated pattern of spatial co-localization that is independent of metabolic zonation. In conclusion, our data set provides a useful resource for understanding hepatocellular alteration during NAFLD at single cell level.
Keywords: hepatocytes, HFD, liver, NAFLD, obesity, scRNA-seq, single cell
INTRODUCTION
The liver is a vital organ that performs essential digestive and metabolic functions within the body, such as glucose and fat metabolism, serum protein production, bile secretion, and chemical detoxification. Most of these functions are mediated by hepatocytes, which constitute the major cell type of the liver, and comprise of 70–85% of the liver’s mass.
With the prevalence of obesity in the modern society, the incidence of nonalcoholic fatty liver disease (NAFLD) is increasing at an alarming rate (1). During obesity, overnutrition and sedentary lifestyle lead to a chronic calorie surplus, resulting in the storage of excessive nutrients in the form of fat. In this condition, the liver also accumulates large fat droplets, which does not typically occur in healthy liver. NAFLD can precipitate further advanced liver diseases such as steatohepatitis (NASH), liver cirrhosis, and hepatocellular carcinoma (HCC; liver cancer) (2).
Recently, pathological NAFLD responses of nonparenchymal cell types, such as inflammatory cells and hepatic stellate cells, were characterized at the single-cell level through scRNA-seq (3). It was reported that, during NAFLD, some macrophages and hepatic stellate cells still retain their normal transcriptome that is almost indistinguishable from those in healthy liver. However, new cell types, which have activated inflammatory signaling (NASH-associated macrophages) or fibrogenic responses (activated hepatic stellate cells), emerged from the normal population and occupied a substantial portion of cells in the diseased liver. Similar observations were made from fibrotic responses of hepatic stellate cells to carbon tetrachloride treatment (4, 5) or human liver cirrhosis (6), indicating the presence of both resting and activated hepatic stellate cell population in fibrotic liver. Another recent study indicated that liver endothelial cells also show similar bipartite response to NASH with responsive and unresponsive populations (7). These findings suggested that at least some nonparenchymal liver cells maintain unaltered transcriptome phenotypes to mediate homeostatic function, whereas other cells alter their transcriptome or migrate from other places to play either adaptive or maladaptive pathological roles during NASH or NAFLD.
Although hepatocytes are often considered functionally homogeneous, studies actually indicate that individual hepatocytes are exposed to different physiological environments, receive different developmental cues, express different sets of genes, and thereby play specialized metabolic functions according to their histological niche (8–11). Recent single-cell RNA-seq (scRNA-seq) studies on normal mouse and human liver samples confirmed the presence of such heterogeneity in mammalian liver (12, 13). Furthermore, histological studies revealed that a subset of hepatocytes in a specific region is more prone to fat accumulation (NAFLD) (14–16), fibrotic disease progression (NASH) (17), liver damage, and hepatocarcinogenesis (HCC) (18–20). Therefore, although transcriptomic analyses of bulk liver mRNAs have revealed that lipogenic, glucogenic, and inflammatory gene transcription levels substantially change upon the development of NAFLD and NASH (21), it is unknown how individual hepatocytes alter their gene expression during liver pathogenesis.
Here, we performed droplet-based, single-cell RNA-sequencing (Drop-seq) (22) on hepatocytes freshly isolated from lean and high-fat diet (HFD)-fed obese mice and characterized their single-cell transcriptome. Our analyses indicate that, unlike nonparenchymal cell types that have both nonresponsive and responsive populations, all hepatocytes altered their transcriptome upon HFD, and each of their single-cell transcriptomes were distinct from the ones isolated from lean mice. However, the patterns of transcriptome alteration were highly heterogeneous across the metabolic zones, and there is also HFD response heterogeneity that is independent of the zonation profile. Some of these interesting single-cell gene expression features were observed at the protein level through immunohistochemistry of liver sections. Collectively, our work reveals how HFD alters the transcriptomic landscape of single hepatocytes across the whole population.
MATERIALS AND METHODS
Data Availability
The scRNA-seq data set generated from this study is available at the Gene Expression Omnibus database (GEO accession no. GSE157281). The data can also be accessed through an interactive online resource (https://lee.lab.medicine.umich.edu/hfd), which has an intuitive graphical user interface for exploring our scRNA-seq data set.
Mice and Diets
Eight-week-old C57BL/6J littermate male mice were separated into two groups and were fed a regular chow diet [low-fat diet (LFD) group; Lab Diet, 5L0D] or high fat diet (HFD group; Bio-Serv, S3282). After 12 wk of dietary modulation, mice whose body weight reached between 48 and 52 g (HFD group) or between 35 and 38 g (LFD group) were euthanized and subjected to single-hepatocyte isolation and Drop-seq. We complied with all relevant ethics regulations for animal testing and research. All experiments were approved by the University of Michigan Institutional Animal Care and Use Committee (PRO00007710 and PRO00009630).
Hepatocyte Isolation
For hepatocyte isolations, the liver was first perfused with calcium-free Hank’s Balanced Salt Solution (HBSS; 14175-095, Gibco) containing 0.2 mg/mL EDTA (51201, AccuGENE) and sodium bicarbonates (7.5%; 25080-094; Gibco) and then sequentially perfused with 0.2% collagenase type II (LS004196; Worthington) in HBSS (14025-092, Gibco) with calcium chloride (2.5 M; C7902-500G; Sigma). The collagenase-treated liver was extracted from the body and further incubated at 37°C for 20 min. Liver cells were diluted in DMEM (11965; Gibco) containing 10% serum and centrifuged at 50 g for 5 min to enrich hepatocytes and passed through a 100-micron nylon cell strainer (10199-659; VWR) multiple times. To remove nonhepatocytes, the gradient precipitation using a 30% percoll solution (17-5445-02; GE Healthcare) was performed, and the resulting hepatocytes were resuspended in 0.5% BSA (A8806; Sigma) in PBS (11965-092; Gibco) for the further analysis of viability and a subsequent Drop-Seq experiment.
Drop-Seq Library Preparation
Drop-seq experiments were performed through a previously described method (22). Hepatocyte preparations were diluted in 2 mL of PBS-BSA to a final concentration of 240,000–300,000 cells. Diluted hepatocytes suspension, barcoded beads (MACOSKO-2011-10; Chemgenes) in lysis buffer (400 mM Tris, pH 7.5, 40 mM EDTA, 12% Ficoll PM-400, 0.4% Sarkosyl, and 100 mM DTT; 100,000 beads/mL), and droplet generation oil (184006; Bio-Rad) were injected into a microfluidics device (FJISUM-QO-180221; FlowJEM) through three separate inlets. The flow rates for the cell and bead suspensions were set as 2,000 µL/h, and the flow rate for the droplet oil was set to 7,500 µL/h. Resulting droplets were sequentially collected in 50-mL falcon tubes, and the total collection time was between 25 and 30 min. Droplets were broken by vigorous shaking to release the beads into the solution, and the beads were collected by centrifugation. Beads were washed multiple times in 6× SSC (diluted from 20× SSC, 15557044; Invitrogen). Excess bead primers were removed by the treatment of Exonuclease I (NEBM0293S; NEB), cDNA synthesis was performed using Template Switch Oligo (TSO), and DNA was amplified using PCR according to the original Drop-seq protocol (22). The resultant PCR product was purified by AMPure XP beads (A63881; Beckman Coulter). The products of the multiple PCR reactions were used for the secondary PCR to construct a full-length cDNA library, which was processed into the sequencing library using the Nextera XT DNA Library Preparation Kit (FC-131-1096; Illumina), with unique barcode sequences for each set. The quality of the libraries was inspected by agarose gel electrophoresis for their average size and concentration before pooling for the sequencing. A total of five sets of cDNA libraries from Drop-seq runs, two from LFD liver and three from HFD liver, were analyzed. The pooled libraries were sequenced using Illumina HiSeq-4000 High-Output at the University of Michigan Sequencing Core and AdmeraHealth, Inc. after an additional quality control process through Agilent BioAnalyzer.
Drop-Seq Data Processing
We processed raw reads, following the instructions described in the Drop-seq Laboratory Protocol version 3 (27) using DropSeqTools (version 1.13). Reads were aligned to the mm10 mouse genome using STAR (version 2.6.0a) (23), following the default DropSeqTools pipeline. The aligned reads were further processed using a popscle software tool (https://github.com/statgen/popscle) to produce the digital expression matrix. We used a unique molecular identifier (UMI) count 400 as an initial cutoff to filter 44,245 droplets to consider for more stringent filtering. Because hepatocytes are extremely fragile (24, 25), ambient RNAs (soup) released from dead hepatocytes could be easily captured by the majority of droplets that do not have actual single cells. Indeed, preliminary analysis of Drop-seq results revealed several droplet clusters that were suspected of having been formed from soup, not from a single cell. To identify these soup droplets from our data set, a shuffled (Shf) data set was generated by random shuffling of transcriptome information in the original (Org) data set. We assumed that soup droplets in the Org data set would exhibit characteristics similar to the droplets in the Shf data set. To test this, Org and Shf data set were plotted on the t-distributed stochastic neighbor embedding (t-SNE) manifold. Indeed, the results indicate that many of the Org droplets from the Drop-seq experiments have a characteristic similar to droplets of the Shf data set, as they overlap in the t-SNE manifold (Supplemental Fig. S1A; Supplemental Material is available at https://doi.org/10.1101/2020.04.16.045260). From the t-SNE manifold, we identified four small clusters (Supplemental Fig. S1A) that are unique to the Org data set. Among these, one cluster (cluster AA in Supplemental Fig. S1, A and B) contained higher levels of mitochondrial transcripts, whereas another cluster (cluster BB in Supplemental Fig. S1, A and C) contained very low levels of UMI. The other two clusters (clusters CC and DD in Supplemental Fig. S1, A–D) had relatively higher UMI numbers and relatively lower mitochondrial transcript content; therefore, we focused on isolating these two clusters from the data set. Therefore, through a series of multidimensional clustering and subtraction of irrelevant droplets with soup-like profiles (Shf-enriched clusters), higher mitochondrial contents (cutoff: 30%), and lower UMI counts (cutoff: 1,000), we isolated a total of 454 droplets that represent 216 cells from two LFD liver samples and 238 cells from three HFD liver samples Supplemental Fig. S1D).
Cell Clustering and Data Visualization
The digital expression matrix was processed to Seurat version 3 (26), following the “standard processing workflow” in the tutorial. Two-dimensional t-SNE (27) and Uniform Manifold Approximation and Projection (UMAP) (28) manifolds were used to visualize gene expression data across different clusters of single cells. Clustering was performed using the shared nearest neighbor modularity optimization implemented in Seurat’s FindClusters function, using a resolution parameter as 0.2. We observed that batch effects are minimal, and all HFD droplets across three independent batches fell into the cluster corresponding to HFD cells, whereas most of the LFD droplets (97%) across two independent batches fell into the other cluster corresponding to LFD cells.
Imputation of Single-Cell Expression
We performed the imputation of the data using the magic package (29) or the saver package (30). Default parameters were used for the imputation work. The scatterplots and feature plots of imputed data were visualized using customized R scripts with ggplot2.
Construction of Hepatocyte Zonation Profiles
Arg1 and Cyp2e1 are established markers for zone 1 and 3 hepatocytes, respectively (12, 31), and expression levels of these genes were comparable between LFD and HFD livers in our data set. Accordingly, imputed gene expression levels for Arg1 and Cyp2e1 were used for estimating hepatocyte zonation. Zonation score was calculated as the difference between the magic-imputed levels of Arg1 and Cyp2e1 expression. According to the zonation score, hepatocytes were divided into five bins of cells, among which the top three bins were grouped together as zone 1 hepatocytes, and the bottom two bins were grouped as zone 2 and 3 hepatocytes, respectively. The resultant hepatocyte groups appropriately reflect the biological characteristics of zone 1–3 hepatocytes, as supported through independent visualizations using PCA, t-SNE, and UMAP manifolds, as well as gene expression analyses of the other established zone-specific markers (see results and discussion for details).
Pathway Enrichment Analysis
Differentially expressed genes (based on fold enrichment) were identified between LFD and HFD hepatocytes and between zone 1 and zone 3 hepatocytes from the LFD set of hepatocytes, using FindAllMarkers function in Seurat. Networks of GO terms were constructed using ShinyGO version 0.61 (32), using only the top 20 significant terms. The pathway enrichment analysis was also performed using enrichGO and enrichKEGG functions in the clusterProfiler version 3.6 (33).
Immunohistochemistry
For immunohistochemistry, liver tissues were fixed in 10% buffered formalin and embedded in paraffin and subjected to immunohistochemical staining, as previously described (34). In brief, paraffin-embedded liver sections were incubated with primary antibodies obtained from Santa Cruz Biotechnology (Apoa4, sc-374543; Elovl5, sc-374138; Fabp1, sc-271591; Cyp2f2, sc-374540; Cyp1a2, sc-53241) at 1:100, followed by incubation with biotin-conjugated secondary antibodies (BA-9200, 1:200; Vector Laboratories) and horseradish peroxidase (HRP)-conjugated streptavidin (554066, 1:300; BD Biosciences). The HRP activity was visualized with diaminobenzidine staining, and nuclei were visualized by hematoxylin counterstaining. For fluorescence staining of lipid droplets and Cyp2f2, livers were harvested from 4-mo-old mice, which had been either LFD or HFD for 2 mo. Frozen liver sections were fixed with 2% paraformaldehyde, blocked with 1× Western Blocking Reagent (Roche), and incubated with anti-Cyp2f2 primary antibody (sc-374540; Santa Cruz Biotechnology), followed by Alexa 594-conjugated secondary antibody, DAPI, and BODIPY 493/503 (Invitrogen).
RESULTS AND DISCUSSION
Drop-Seq Successfully Captures Single-Hepatocyte Transcriptome Profile
To characterize the effect of HFD on single-cell transcriptome of hepatocytes, we performed Drop-seq in five independent experiments, with freshly isolated hepatocytes from two normal chow (LFD)-fed lean mice and three HFD-fed obese mice. A total of 216 high-quality hepatocytes were identified from the livers of LFD mice, whereas 238 were identified from those of HFD mice (see materials and methods for details). All of these droplets expressed robust levels of Alb (>0.9% of total transcriptome; Supplemental Fig. S2A), an authentic hepatocyte marker encoding albumin protein, confirming that these droplets indeed represent hepatocyte transcriptome.
In contrast, most macrophage markers, such as Emr1, Itgam, and Cd14, as well as many inflammatory cytokines, such as Tnf, Il6, and Ccl2, were undetectable from our single-cell transcriptome data set (Supplemental Fig. S2, B and C), indicating that our Drop-seq preparations did not have contaminating fractions of Kupfer cells, the liver-resident macrophages. Many markers for hepatic stellate cells and fibroblasts, such as Acta2, Col3a1, Pecam1, and Mmp2, were also not detected (Supplemental Fig. S2). Major adipocyte markers, such as Fabp4 and Adipoq, were also undetectable (Supplemental Fig. S2E), indicating that although HFD and fatty liver can render hepatocytes to accumulate lipid droplets (35), they do not alter the tissue identity of hepatocytes to exhibit adipocyte characteristics.
HFD Alters Single-Cell Transcriptome Profile of the Entire Hepatocyte Population in Liver
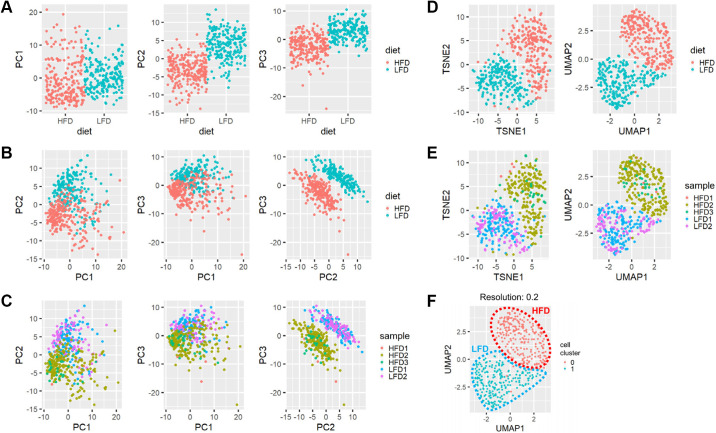
To explore and understand the single-hepatocyte transcriptome data, we first performed the principal component (PC) analysis to determine the signatures of the largest variance in our data set. PC1, which represents the largest variance, did not characterize significant differences between LFD and HFD samples (Fig. 1A, left). However, PC2 and PC3, the orthogonal axes representing the second- and third-largest variance, respectively, were highly effective in separating LFD and HFD transcriptome profiles (Fig. 1A, middle and right). Correspondingly, PC2 and PC3 were sufficient to discriminate LFD and HFD hepatocytes without any additional information (Fig. 1B). The effect of HFD was robust across independent batches of the experiment (Fig. 1C).
Figure 1.
High-fat diet (HFD) alters single-cell transcriptome of the entire hepatocyte population. Eight-week-old C57BL/6J male littermate mice were separated into 2 groups and fed a regular chow diet (LFD group) or high-fat diet (HFD group). Droplet-based, single-cell RNA sequencing (Drop-seq) of hepatocytes was performed after 12 wk of dietary modulation. A–F: principal component analysis (PCA; A–C), t-distributed stochastic neighbor embedding (t-SNE), and Uniform Manifold Approximation and Projection (UMAP; D–F) manifolds colored with diet (A, B, and D), sample (C and E), or the result from multidimensional clustering (F). Individual dots represent single-cell transcriptome. In each manifold, the distance between individual dots represents the difference between the single-cell transcriptome. Approximate boundaries of the area for LFD and HFD samples are indicated as dotted outline (F).
Similar trends were observed from the nonlinear manifolds generated by t-SNE and UMAP dimension reduction methods (27, 28), which segregated LFD and HFD groups of hepatocytes (Fig. 1D) but were robust against batch effects (Fig. 1E). High-dimensional clustering analysis also clearly differentiated the LFD and HFD groups; all cells from HFD mice fell into the cluster corresponding to the HFD group (group 0 in Fig. 1F), whereas 213 out of 216 cells (98%) from LFD mice fell into the LFD group (group 1 in Fig. 1F). These results indicate that the entire hepatocyte population in mouse liver responded to the HFD challenge by altering their transcriptome profiles.
Metabolic Zonation of Hepatocytes Was Captured in Both HFD and LFD Livers
We were curious about the nature of the PC1 axis, which represents the largest variance of transcriptomic profiles across all hepatocyte populations yet does not strongly represent the diet effect (Fig. 1A). We observed that the hepatocyte marker Alb expression exhibited a substantial negative correlation with PC1 in both HFD and LFD groups (r = −0.54, P < 2.2e-16; Supplemental Fig. S2F). Alb expression is known to be relatively higher in periportal zone 1 hepatocytes and relatively lower in pericentral zone 3 hepatocytes (36); therefore, we suspected that the PC1 axis might represent the metabolic zonation of individual hepatocytes. To further substantiate this conjecture, we examined the expressions of the well-characterized periportal marker Arg1 and the pericentral marker Cyp2e1 (12, 31) to understand the zonation structure of our data set. Both scaled and imputed expression levels of Arg1 showed negative correlation with the PC1 (r = −0.26 and −0.84, respectively, P < 2.5e-8; Supplemental Fig. S3A, top), whereas expression levels of Cyp2e1 showed positive correlation with the PC1 (r = 0.56 and 0.89; P < 2.2e-16; Supplemental Fig. S3A, bottom), indicating that PC1 indeed represents the metabolic zonation structure of hepatocytes.
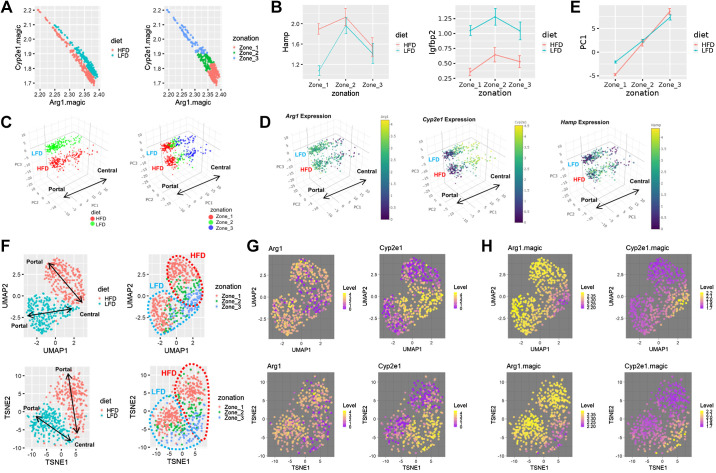
Interestingly, in single cells, imputed levels of Arg1 and Cyp2e1 expression showed a clear negative correlation (r = −0.93; Fig. 2A), consistent with their opposed expression patterns in the liver (12, 31). Based on these levels of Arg1 and Cyp2e1 expression, we partitioned the liver with three zones: zone 1 with periportal characteristics, zone 2 with intermediate characteristics, and zone 3 with pericentral characteristics (Fig. 2A, right).
Figure 2.
Zonation patterns of single-hepatocyte transcriptome is preserved after high-fat diet (HFD). A: inverse correlation between imputed expression levels of Arg1 and Cyp2e1 (magic-imputed expression values). Individual dots represent single-cell transcriptome colored with diet (left) and zone assignment (right). B: analysis of single-cell gene expression in hepatocytes of each zone, expressed as means ± SE (scaled expression values). Data from low-fat diet (LFD) and HFD livers were analyzed separately. C and D: 3-dimensional PCA manifolds depicting the effect of diet (C, left), zonation (C, right), and expression levels of indicated genes (D). Individual dots represent single-cell transcriptome. The size of the dots represents the number of RNA features captured in the droplet. PC1 is composed of genes showing zone-specific expression patterns. PC2 and PC3 are composed of genes showing diet-regulated expression patterns. LFD and HFD area, as well as the directionality of metabolic zonation (from portal to central), are indicated in each manifold. E: analysis of single-cell PC1 values in hepatocytes of each zone, expressed as means ± SE (raw component scores). Data from LFD and HFD livers were analyzed separately. F–H: Uniform Manifold Approximation and Projection (UMAP; top) and t-distributed stochastic neighbor embedding (t-SNE; bottom) manifolds depicting the effect of diet (F, left), zonation (F, right), and scaled (G) and imputed (H) expression levels of indicated genes. LFD and HFD area (F, right), as well as the directionality of metabolic zonation (from portal to central; F, left), are indicated.
Previous studies isolated a long list of zone 1-specific markers, such as Alb, Ass1, Arg1, Cyp2f2, Cps1, Gls2, Pck1, and Sult5a1, and zone 3-specific markers, such as Glul, Oat, Slc1a2, Lect2, Ldhd, Por, Cyp1a2, Cyp2e1, Ahr, and Gstm2, -3 and -6, through various methodologies, including differential isolation, immunohistochemistry, or RNA in situ hybridization analyses (12, 31, 37). All of these genes appear to have corresponding patterns of expression in our data set (Supplemental Fig. S4). Furthermore, Hamp and Igfbp1, genes whose expression is elevated in the intermediate region of the liver (12), showed zone 2-specific expression from our data set (Fig. 2B). These results confirm the validity of our zonation method.
Diet and zonation effects can also be jointly visualized in a three-dimensional PC1/PC2/PC3 space, where PC2 and PC3 axes separate LFD and HFD hepatocytes (Fig. 2C, left), and the PC1 axis visualizes the portal-to-central histological zonation structure (Fig. 2, C, right, and D, and Supplemental Fig. S3B). Indeed, PC1 values were the highest in zone 3 and the lowest in zone 1 hepatocytes, according to our hepatocyte zonation groups (Fig. 2E). Diet and zonation effects were also robustly observed in UMAP and t-SNE (Fig. 2, F–H) manifolds. These results indicate that 1) the structure of metabolic zonation is maintained in HFD liver, 2) HFD produced transcriptome-altering effects on hepatocyte population across entire zonation niches, and 3) zonation effect and diet effect are the two major sources of variation in single-hepatocyte transcriptomes observed from our data set.
HFD Alters the Expression of Genes Controlling Lipid Metabolism
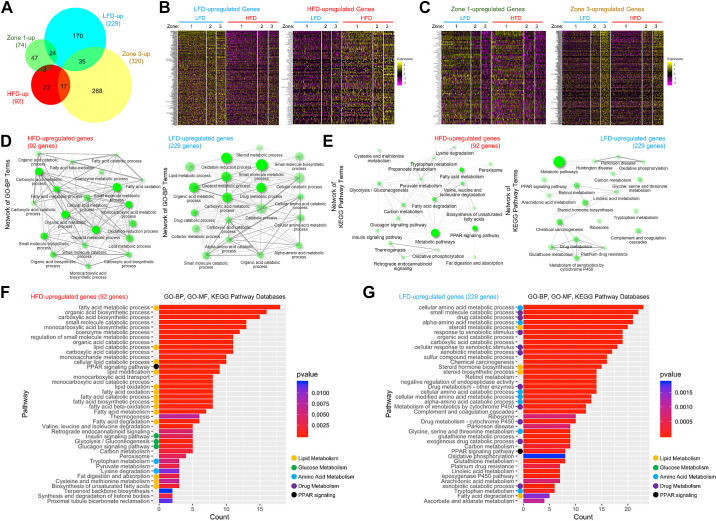
Using the diet and zonation information of individual hepatocytes, we identified a list of genes whose expression patterns are modulated by HFD or dependent on their zonation. Ninety-one genes were significantly upregulated in the HFD group, whereas 226 genes were significantly upregulated in the LFD group (FDR < 0.01; Supplemental Table S1, 1st to 3rd tabs). Partially overlapping with this list (Fig. 3A), 74 genes were found to be specific to zone 1 hepatocytes, whereas 320 genes were specific to zone 3 hepatocytes (FDR < 0.01; Supplemental Table S1, 1st, 4th, and 5th tabs). Heat map analysis of the diet-specific (Fig. 3B) and zone-specific genes (Fig. 3C) confirmed that these gene groups show contrasted gene expression patterns across different populations of hepatocytes.
Figure 3.
Isolation of genes showing diet- and zone-specific expression patterns. A: area-proportional Venn diagram depicting the relationship between diet-regulated genes and zone-specific genes. B and C: heat map analysis depicting gene expression across single-cell population. Single cells were clustered into 6 groups (thick columns) according to diet and zone. Diet-regulated genes (B) and zone-specific genes (C) were analyzed. D and E: network analysis of gene ontology-biological pathway (GO-BP; D) and Kyoto Encyclopedia of Genes and Genomes (KEGG; E) pathway enrichment terms, using ShinyGO (11). Pathways whose enrichment is significant (FDR < 0.05; top 20 terms) were presented as nodes. Two nodes are connected if they share 20% or more genes. Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes. F and G: enrichment analysis of high-fat diet (HFD)-upregulated (left) and downregulated (right) genes, using clusterProfiler (53) with GO-BP, GO-molecular function (GO-MF), and KEGG databases. Color of bars indicates significance (P values), whereas length of bars indicates gene count. Color of circles indicate GO terms related to lipid metabolism (yellow), glucose metabolism (green), amino acid metabolism (blue), drug metabolism (purple), and PPAR pathway (black).
Gene ontology analysis of HFD-upregulated and -downregulated (LFD-upregulated) genes showed that, consistent with previous bulk gene expression studies (38–40), genes controlling lipid and fatty acid metabolism are upregulated in HFD, whereas genes controlling amino acid and drug catabolism are downregulated (FDR < 0.05 for all presented pathways; Fig. 3, D–G). However, genes mediating inflammation and fibrosis were not included here (Fig. 3, D–G and Supplemental Table S1), since our data set was exclusive to hepatocytes and did not include hepatic stellate cells or inflammatory cells (Supplemental Fig. S1). Pathway enrichment analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database identified the metabolic pathways as the top enriched pathway for both HFD-upregulated and -downregulated gene lists (FDR = 1.6e-10 and 2.2e-29, respectively; Fig. 3E), consistent with the central role of the liver in metabolism. Interestingly, among various biological pathways, the PPAR pathway was represented in both HFD-upregulated and HFD-downregulated gene lists (Fig. 3, E–G), consistent with the former studies indicating that the pathway is among the major pathways altering hepatocellular transcriptome during HFD (41, 42).
HFD Alters Expression Patterns of a Subset of Zone-Specific Genes
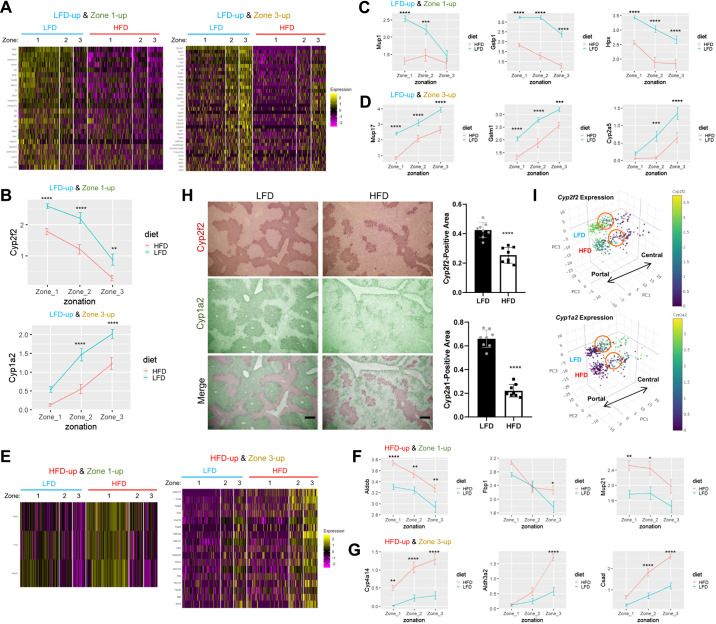
Next, we focused on the genes that exhibit both diet- and zone-specific expression patterns (Fig. 3A and Supplemental Table S1, 1st tab). We found that many markers for zone 1 hepatocytes, such as Cyp2f2, Mup1, Gstp1, and Hpx, were strongly downregulated upon HFD feeding (Fig. 4, A–D). Only a very small number of zone 1-specific genes, such as Aldob, Fbp1, and Mup21, were upregulated (Fig. 4, E and F). Many zone 3 hepatocyte markers, such as Cyp1a2, Mup17, Gstm1, and Cyp2a5, were also downregulated (Fig. 4, A–D). However, several zone 3-specific genes, including Cyp4a14, Aldh3a2, and Csad, were substantially upregulated in response to HFD (Fig. 4, E and G). Therefore, the HFD effect on zone-specific gene expression could be variable across individual genes.
Figure 4.
Isolation of genes that are substantially influenced by both diet and zonation. A and E: heat map analysis depicting gene expression across single-cell population. Cells were clustered into 6 groups according to diet and zone. High-fat diet (HFD)-downregulated (A) and -upregulated (E) genes that show periportal (zone 1-high; left in each panel) or pericentral (zone 3-high; right in each panel) patterns of expression were analyzed. B–D, F, and G: analysis of single-cell gene expression in hepatocytes of each zone, expressed as means ± SE (scaled expression values). Data from low-fat diet (LFD) and HFD livers were analyzed separately. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 in Sidak’s multiple-comparison test. H: Cyp2f2 and Cyp1a2 protein expression was visualized through immunohistochemistry from serial sections of LFD and HFD mouse liver (left). Cyp2f2 and Cyp1a2 staining signals were artificially colored with red (1st row) and green (2nd row), respectively, to produce merged images (3rd row) of the serial liver sections. Cyp2f2- and Cyp1a2-positive areas were quantified (right). Scale bars, 200 µm. I: 3-dimensional PCA manifold depicting the single-cell expression levels of indicated genes. Individual dots represent single-cell transcriptome. The size of the dots represents the number of RNA features captured in the droplet. LFD and HFD area, as well as the directionality of metabolic zonation (from portal to central), are indicated in each manifold. Orange circles indicate the approximate position of zone 2 hepatocytes.
We assessed whether mRNA expression changes observed from our Drop-seq analysis could lead to alterations of the protein level by examining Cyp2f2 and Cyp1a2 genes, which are among the genes that show the strongest zonation patterns in our data set and previous data sets (12). In our data set, the expression of these genes in their corresponding metabolic zones was strongly reduced after HFD (Fig. 4B). These observations were reproduced through immunohistochemical staining of Cyp2f2 and Cyp1a2 proteins in liver sections; the areas expressing these two proteins were dramatically shrunken (Fig. 4H). Correspondingly, although the regions expressing Cyp2f2 and Cyp1a2 substantially overlapped in LFD liver, they hardly overlapped in HFD, creating the gap area where none of these proteins were expressed (Fig. 4H). Similar patterns were also observed in Drop-seq data, where many zone 2 hepatocytes reduced expression of both Cyp2f2 and Cyp1a2 upon HFD (cells in orange circles of Fig. 4I). These data exemplify the relevance of our Drop-seq data set for understanding single-cell gene expression of hepatocytes in LFD and HFD mouse liver.
Zonation-Independent Heterogeneity in Single Hepatocyte Responses to HFD
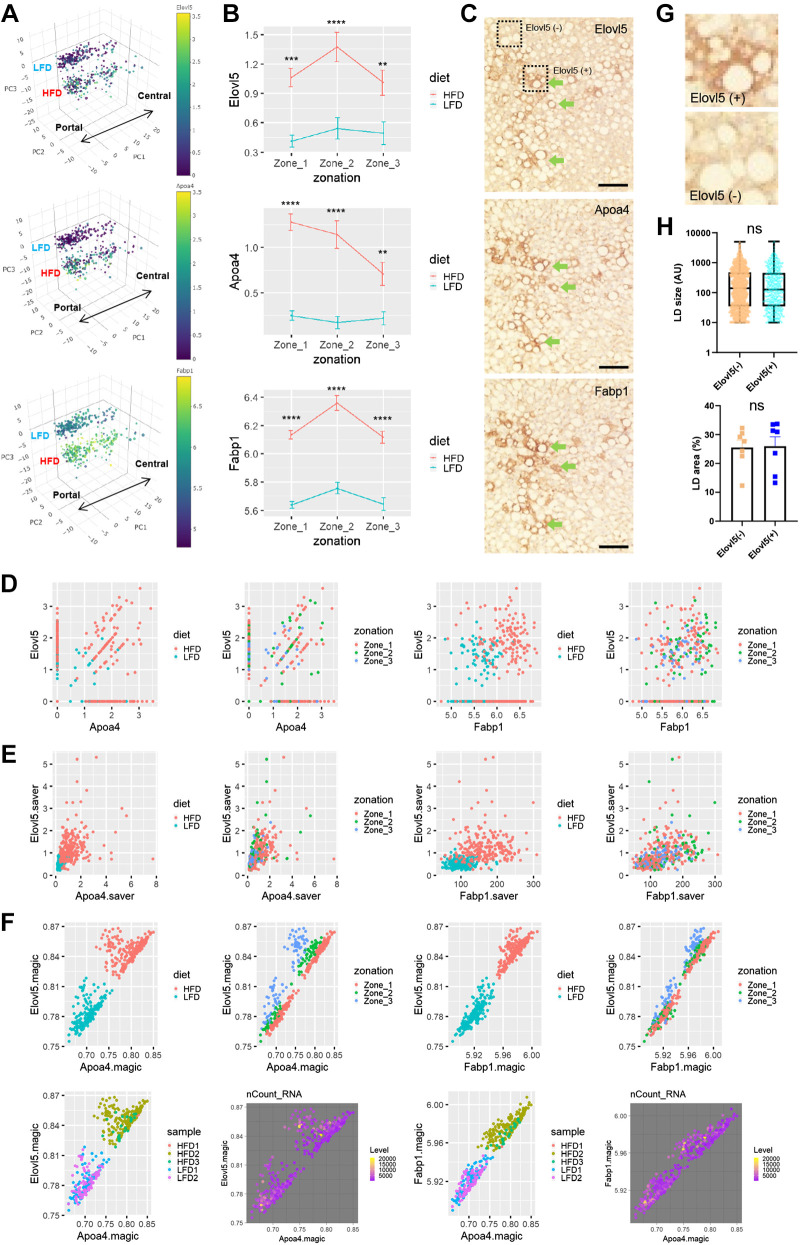
Elovl5, Apoa4, and Fabp1 are among the genes that show the strongest upregulation of gene expression after HFD (Supplemental Table S1, 3rd tab). Although the HFD induction of these genes was very robust in both the Drop-seq data set (Fig. 5, A and B) and immunohistochemical staining of liver sections (Supplemental Fig. S5), they did not show strong zone-specific expression patterns (Fig. 5, A and B, and Supplemental Fig. S5). Interestingly, in liver immunohistochemistry, Elovl5, Apoa4, and Fabp1 proteins exhibited variegated expression patterns across the hepatocytes (Supplemental Fig. S5A), indicating that their induction after HFD is highly heterogeneous between different hepatocytes independent of metabolic zonation.
Figure 5.
Spatial co-expression pattern of high-fat diet (HFD)-induced genes regulating lipid metabolism. A: 3-dimensional PCA manifold depicting the single-cell expression levels of indicated genes. Individual dots represent single-cell transcriptome. The size of the dots represents the number of RNA features captured in the droplet. Low-fat diet (LFD) and HFD area, as well as the directionality of metabolic zonation (from portal to central), are indicated in each manifold. B: analysis of single-cell gene expression in hepatocytes of each zone, expressed as means ± SE (scaled expression values). Data from LFD and HFD livers were analyzed separately. **P < 0.01, ***P < 0.001, and ****P < 0.0001 in Sidak’s multiple-comparison test. C, G, and H: Elovl5, Apoa4, and Fabp1 protein expression was visualized through immunohistochemistry from serial sections of HFD mouse liver (C). Green arrows indicate areas of positive staining that are congruently observed across all three staining images. Elovl5-high (+) and -low (−) areas (dotted boxes) are magnified in G. Lipid droplet (LD) size (n ≥ 479; H, top) and area (n = 7; H, bottom) in Elovl5-high and -low areas were quantified. Data are expressed as a box plot [arbitrary unit (AU); top] or means ± SE (%area; bottom) with individual data points. Student’s t tests failed to detect a significant difference between the 2 groups (NS). Scale bars, 100 µm. D–F: correlation between expression levels of Elovl5, Apoa4, and Fabp1 genes from scaled (D), saver-imputed (E), and magic-imputed (F) droplet-based, single-cell RNA sequencing (Drop-seq) data set. Individual dots represent single-cell expression levels colored by diet, zone, sample information, and level of total RNA counts (nCount_RNA).
Given the spatially restricted patterns of Elovl5, Apoa4, and Fabp1 protein expression in liver sections (Supplemental Fig. S5A), we became curious about whether the patterns between these genes are correlated with each other. To assess this, we stained each of these proteins in a serial section of the same histological block. Interestingly, it was found that the regions of high Elovl5, Apoa4, and Fabp1 expression were substantially overlapping with each other, indicating that protein products of these genes are expressed in a positive correlation with each other (Fig. 5C).
We then examined whether the positive correlation between Elovl5, Apoa4, and Fabp1 expression could be observed from the Drop-seq data set. A query of the most significantly correlated gene for Elovl5 expression resulted in Apoa4, Cyp4a14, and Fabp1 as the top three genes, among which both Apoa4 and Fabp1 are included. Correlation scatterplot between Elovl5 and these two genes showed the trend of positive correlation in scaled data (r = 0.22 and 0.21, respectively, P < 1.5e-6; Fig. 5D); however, due to the sparsity of the specific mRNA observation and subsequent technical noise, the observed correlation may not be as strong as the true correlation. After applying two independent imputation methods correcting for the technical noise effect, saver (Fig. 5E) (30) and magic (Fig. 5F) (29), we were able to detect more robust correlation between gene expression profiles of Elovl5 and Apoa4 [r = 0.90 (magic) and 0.50 (saver)] and between Elovl5 and Fabp1 [r = 0.97 (magic) and 0.41 (saver)] (Fig. 5, E and F). Importantly, these patterns of correlation were independent of the zonation (zonation panels in Fig. 5, D–F), batches (sample panels in Fig. 5F), or mRNA reads (nCount_RNA/level panels in Fig. 5F). Therefore, these results suggest the presence of zonation-independent heterogeneity in hepatocyte responses to HFD.
Elovl5-High and -Low Hepatocytes Accumulate Similar Levels of Fat Droplets
In HFD mice, hepatocytes expressing high levels of Elovl5, Apoa4, and Fabp1 were not morphologically different from other hepatocytes in terms of lipid droplet accumulation (Fig. 5, C and G). Quantification of the lipid droplet size did not reveal any obvious differences in lipid droplet size (Fig. 5H, top) or area (Fig. 5H, bottom) between Elovl5-high and -low hepatocyte populations. Therefore, the levels of HFD-induced Elovl5, Apoa4, and Fabp1 do not seem to substantially alter the steady-state level of fat accumulation in the hepatocytes.
Considering that Fabp1, Elovl5, and Apoa4 are all involved in fatty acid metabolism, it could be inferred that hepatocytes expressing high levels of these genes might be more active in lipid processes. Because the histological analysis indicates that the expression of these genes does not substantially alter the intracellular amounts of fat droplets (Fig. 5, G and H), the biological relevance of this heterogeneous gene expression pattern is unclear in the context of HFD feeding. It is possible that active processes in lipid metabolism, mediated by these genes, alter the flux of lipid metabolites without affecting the steady-state fat levels. It is also possible that heterogeneity in expression of these genes is temporarily generated; therefore, over time, other hepatocytes might also express high levels of these proteins, producing similar metabolic profiles.
HFD Induces Stronger Fat Accumulation in Zone 3 Hepatocytes
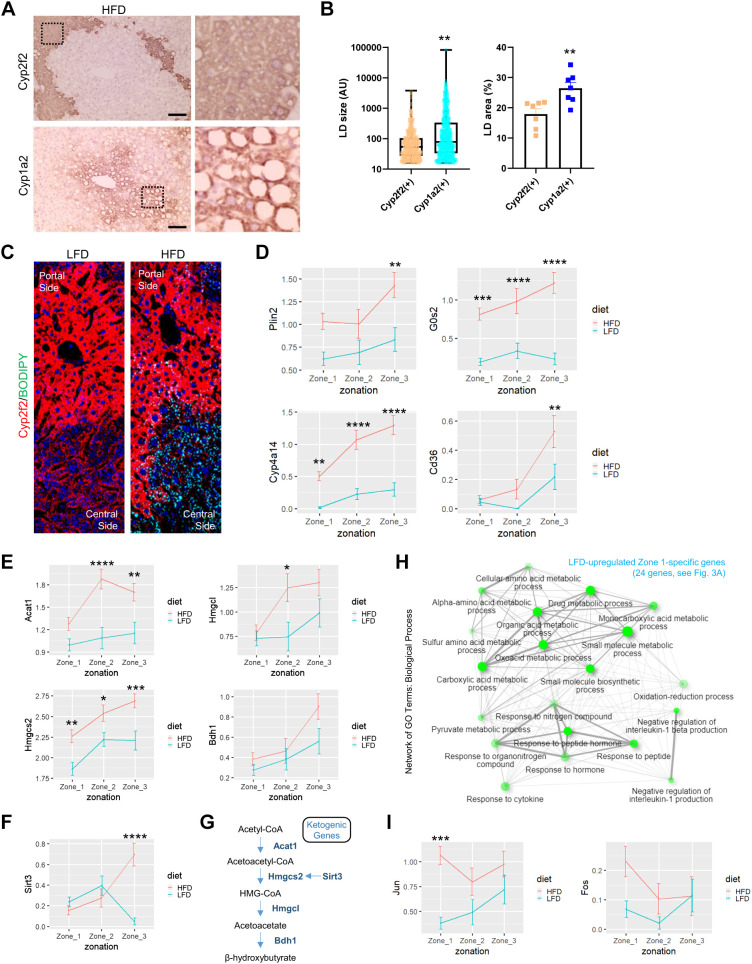
It was well documented that obesity and hepatosteatosis disparately affect individual hepatocytes across their histological zonation. Some hepatocytes, especially the ones in zone 3, which are deprived of nutrients and oxygen, are more prone to accumulate lipid droplets, whereas the ones in zone 1, a nutrient- and oxygen-rich environment, are less susceptible to steatotic progression (15, 16). Consistent with these former studies, we observed from the histology results that Cyp1a2-positive zone 3 hepatocytes contain more and bigger lipid droplets when compared to Cyp2f2-positive zone 1 hepatocytes (Fig. 6, A and B). The observation of zone 3-specific fat accumulation was reproduced when we directly stained lipid droplets in freshly frozen tissue sections from LFD and HFD livers (Fig. 6C), again supporting that HFD-induced fat accumulation is more pronounced in zone 3.
Figure 6.
High-fat diet (HFD) induces zone 3 hepatocytes to express genes promoting lipid droplet accumulation. A and B: serial sections of HFD mouse liver were stained with zone 1 marker Cyp2f2 (A, top) and zone 2 marker Cyp1a2 (A, bottom). Boxed areas in A were magnified at right. Lipid droplet (LD) size (n ≥ 535; B, left) and area (n = 7; B, right) in each compartment were quantified. Data are expressed as box plot [arbitrary units (AU); left] or means ± SEM (%area; right) with individual data points. Student’s t tests were used to examine significant differences between the 2 groups (**P < 0.01 ). Scale bars, 100 µm. C: Fresh frozen sections from low-fat diet (LFD) and HFD mouse liver were immunostained to visualize Cyp2f2 (red), LDs (green, stained by BODIPY 493/503), and DNA (blue, by DAPI). D–F and I: analysis of single-cell gene expression in hepatocytes of each zone, expressed as means ± SE (scaled expression values). Data from LFD and HFD livers were analyzed separately. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 in Sidak’s multiple-comparison test. G: Ketogenic genes induced by HFD in zone 3 hepatocytes are presented in a pathway diagram. H: network analysis of gene ontology-biological pathway (GO-BP) enrichment terms in the HFD-downregulated zone 1-specific gene list, using ShinyGO (11). Pathways whose enrichment is significant (FDR < 0.05) are presented as nodes. Two nodes are connected if they share 20% or more genes. Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes.
Zone 3 Hepatocytes Robustly Express Genes Mediating Fat Accumulation during HFD
We then tried to identify the features of single hepatocyte transcriptome that may explain the preferential accumulation of lipid droplets in zone 3 hepatocytes. For this, we surveyed the function of all genes that show either HFD- or zone 3-specific expression patterns (Supplemental Table S1, 3rd and 5th tabs) through literature search. We found that there are at least four genes, Plin2, G0s2, Cyp4a14, and Cd36, that are known to play a mechanistic role in fat accumulation (43–49) and at the same time are strongly induced by HFD in zone 3 hepatocytes (Fig. 6D).
Plin2 is a protein directly associated with hepatic lipid droplets (44). Plin2 surrounds the lipid droplet and assists the storage of neutral lipids within the lipid droplets. Consistent with increased lipid droplet accumulation in zone 3 hepatocytes, Plin2 expression is more strongly upregulated in zone 3 during HFD (Fig. 6D). The Plin2 induction could be critical for zone 3-specific accumulation of lipid droplets because hepatic deletion of Plin2 is known to attenuate hepatic fat accumulation (44, 45).
G0s2, whose product is a well-established inhibitor of lipase activity in hepatocytes (47), was also strongly upregulated upon HFD, specifically in zone 3 hepatocytes (Fig. 6D). Considering that G0s2 is important for the accumulation of triglycerides in hepatocytes by inhibiting lipase activities, it is likely that the pericentral expression of G0s2 is responsible for lipid droplet accumulation in zone 3. Indeed, in a recent study, G0s2 knockout mice and liver-specific knockdown mice did not show hepatosteatosis upon HFD, whereas G0s2 overexpression sufficed to induce hepatosteatosis (49).
Cyp4a14 is another gene that is induced upon HFD and critical for generating HFD-induced hepatosteatosis (48). HFD-induced Cyp4a14 expression is also much more pronounced in zone 3 compared with the other zones (Fig. 6D).
Cyp4a14 was suggested to promote hepatosteatosis partly thorough inducing Cd36/FAT, whose products play a role in importing fatty acids into hepatocytes (48). Cd36/FAT was also highly induced in zone 3 hepatocytes of HFD-fed mouse liver (Fig. 6D). Notably, prior studies showed that hepatic Cd36 overexpression was sufficient to provoke hepatosteatosis even without HFD challenges (43), whereas liver-specific Cd36 disruption was sufficient to attenuate fatty liver in HFD mice (46).
Collectively, these observations, combined with former genetic studies performed on these genes (43–49), suggest that zone 3-specific upregulation of Plin2, G0s2, Cyp4a14, and Cd36 plays an important role for producing zone 3-specific steatosis phenotype in response to HFD challenges.
Zone 3 Hepatocytes Upregulate Genes Mediating Ketogenic Pathway
In addition to the genes responsible for producing lipid droplet accumulation, we also observed that HFD-induced expression of ketogenic genes, such as Acat1, Hmgcs2, Hmgcl, and Bdh1, were relatively higher in zone 3 hepatocytes (Fig. 6E). Sirt3, whose product deacetylates and activates Hmgcs2, was also more strongly expressed in zone 3 hepatocytes of HFD liver (Fig. 6F). These results suggest that the HFD-induced ketogenesis pathway (Fig. 6G) is preferentially activated in zone 3 hepatocytes of mouse liver. Activation of ketogenesis in zone 3 hepatocytes might be critical for distributing the energy to peripheral tissues and generating metabolic adaptation to HFD-induced hypernutrition (50).
Zone 1 Hepatocytes Also Transcriptionally Respond to HFD
In contrast to zone 3 hepatocytes, zone 1 hepatocytes strongly downregulated many zone 1-specific transcripts that mediate various metabolic processes, including drug and amino acid catabolism and redox metabolism (Fig. 6H). Reduction of these functions may be critical for HFD adaptation by accommodating an increased need for lipid metabolism. Although many of the zone 1-specific genes were downregulated (Fig. 4A), Aldob and Fbp1, two genes that are involved in gluconeogenesis, were strongly upregulated in zone 1 hepatocytes after HFD (Fig. 4, E and F). This is consistent with the previous findings indicating that gluconeogenesis activity is the most active in zone 1 hepatocytes (16). This zone 1-specific regulation of Aldob and Fbp1 might be contributing to decreased glucose tolerance during HFD-induced obesity (51). In addition, stress-induced AP1 transcription factors Jun and Fos were also specifically upregulated in zone 1 hepatocytes upon HFD stimulation (Fig. 6I). These results indicate that, although zone 1 hepatocytes are relatively resistant to fat droplet accumulation, they also respond transcriptionally to HFD challenges and contribute to physiological HFD responses in a substantial way.
PPAR Pathway is Implicated in HFD Modulation of Single-Hepatocyte Transcriptome
It is interesting to note that many of the HFD-induced genes reviewed above are targets of the PPAR family of transcription factors; genes with a variegated co-expression pattern (Elovl5, Apoa4, and Fabp1), as well as genes that show zone 3-specific patterns and mediate fat droplet accumulation (G0s2, Plin2, Cyp4a14, and Cd36) and ketogenesis upregulation (Acat1, Hmgcs2, Hmgcl, and Bdh1), are all targets or PPARα (Fig. 7) (42, 52, 53). As observed above (Fig. 3, E–G), the PPAR pathway is the only transcription factor-targeted group that is enriched in both HFD-upregulated and HFD-downregulated gene lists. Importantly, PPAR is known to be activated upon stimulation with fatty acids, as they are direct ligands for transcriptional activation of PPAR (52, 53). Former bulk analysis of fatty liver transcriptome also revealed that various targets of PPAR, as represented in our data set, are strongly upregulated upon HFD challenges (38–41). Notably, many of these genes did not show such diet-dependent modulations in Ppara-deleted knockout mutant strains (41, 42). Therefore, many transcriptome features observed from our data set could be at least partly mediated by PPAR activation by excessive fatty acids from dietary sources.
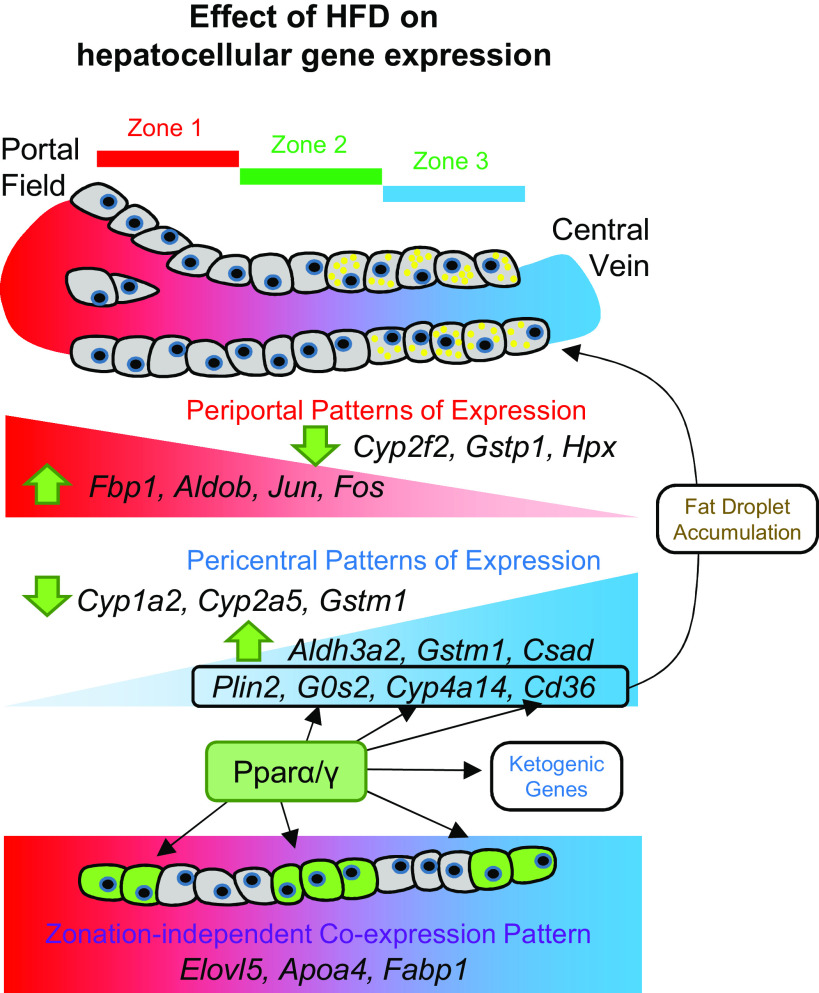
Figure 7.
Holistic understanding of heterogeneous hepatocyte responses to high-fat diet (HFD). The schematic diagram depicts the heterogeneous effect of HFD on single hepatocellular gene expression. Our data set indicates that the entire hepatocyte population undergoes substantial transcriptome changes upon HFD and that the patterns of alteration were highly heterogeneous across the hepatocyte population with zonation-dependent and -independent effects.
Limitations of the Current Study
Our Drop-seq data set contains a large number of droplets containing ambient RNA (Supplemental Fig. S1; see materials and methods for details), indicating that a considerable number of single hepatocytes were damaged during isolation and microfluidics. It is possible that the single-hepatocyte data presented in the current study is biased toward the hepatocyte population that is resistant to physical damage. In addition, although our method of partitioning single-hepatocyte transcriptome profiles into three zones is robust and consistent with previous studies, it is possible that this is an oversimplification of the complex histological architecture of the liver. These issues could be potentially addressed in future studies by utilizing microfluidics-free methods for sorting single cells (12, 54) or spatial profiling of liver transcriptome through tissue sections (55).
Summary
Recent single-cell transcriptome studies revealed that hepatocyte gene expression and function are highly heterogeneous across their metabolic zonation, revealing global division of metabolic labor of the liver (12, 13). Building on these previous findings, our study provides the first snapshot of how the single-hepatocyte transcriptome landscape is altered in response to HFD and subsequent development of NAFLD. Through this data set, we were able to find that HFD makes an impact on the transcriptome of the entire hepatocyte population. We also found that HFD responses of hepatocytes can be heterogeneous with zonation-dependent and -independent effects. Our observations detailed above systematically characterize HFD-induced changes in hepatocellular transcriptome and their relationship to NAFLD pathogenesis. Furthermore, we made our data set available in an interactive web tool (https://lee.lab.medicine.umich.edu/hfd), where individual investigators can reproduce our analyses and test their hypothesis using our publicly available data set. We believe that this resource will be greatly useful for future NAFLD studies.
GRANTS
This work was supported by the National Institutes of Health (NIH; R01-DK-102850 and R01-DK-114131 to J.H.L., U-01-HL-137182 to H.M.K., T32AG000114 to C.S.C., and P30-AG-024824, P30-DK-034933, P30-DK-089503, and P30-CA-046592), the Chan Zuckerberg Initiative (to H.M.K.), the MCubed initiative (to J.H.L. and H.M.K.), and the AASLD Pilot Research Award (to J.H.L. and H.M.K.).
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the authors.
AUTHOR CONTRIBUTIONS
H.K. and J.L. conceived and designed research; S.P. and C.C. performed experiments; S.P., J.X., H.K., and J.L. analyzed data; S.P., H.K., and J.L. interpreted results of experiments; S.P., H.K. and J.L. prepared figures; S.P., H.K., and J.L. drafted manuscript; H.K. and J.L. edited and revised manuscript; S.P., C.C., J.X., H.K., and J.L. approved final version of manuscript.
ACKNOWLEDGMENTS
We thank Drs. Jiandie Lin, Yatrik Shah, and Lei Yin labs for their help with hepatocyte isolation and scRNA-seq optimization. We thank Dr. Myungjin Kim for advice, Dr. Steve Norris for editing, Bondong Gu and Boyoung Kim for assistance, and Santa Cruz Biotechnology for antibodies.
REFERENCES
- 1.Lazo M, Hernaez R, Eberhardt MS, Bonekamp S, Kamel I, Guallar E, Koteish A, Brancati FL, Clark JM. Prevalence of nonalcoholic fatty liver disease in the United States: the Third National Health and Nutrition Examination Survey, 1988–1994. Am J Epidemiol 178: 38–45, 2013. doi: 10.1093/aje/kws448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Michelotti GA, Machado MV, Diehl AM. NAFLD, NASH and liver cancer. Nat Rev Gastroenterol Hepatol 10: 656–665, 2013. doi: 10.1038/nrgastro.2013.183. [DOI] [PubMed] [Google Scholar]
- 3.Xiong X, Kuang H, Ansari S, Liu T, Gong J, Wang S, Zhao X-Y, Ji Y, Li C, Guo L, Zhou L, Chen Z, Leon-Mimila P, Chung MT, Kurabayashi K, Opp J, Campos-Pérez F, Villamil-Ramírez H, Canizales-Quinteros S, Lyons R, Lumeng CN, Zhou B, Qi L, Huertas-Vazquez A, Lusis AJ, Xu XZS, Li S, Yu Y, Li JZ, Lin JD. Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol Cell 75: 644–660.e5, 2019. doi: 10.1016/j.molcel.2019.07.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dobie R, Wilson-Kanamori JR, Henderson BEP, Smith JR, Matchett KP, Portman JR, Wallenborg K, Picelli S, Zagorska A, Pendem SV, Hudson TE, Wu MM, Budas GR, Breckenridge DG, Harrison EM, Mole DJ, Wigmore SJ, Ramachandran P, Ponting CP, Teichmann SA, Marioni JC, Henderson NC. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep 29: 1832–1847, 2019. e1838doi: 10.1016/j.celrep.2019.10.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Krenkel O, Hundertmark J, Ritz TP, Weiskirchen R, Tacke F. Single cell RNA sequencing identifies subsets of hepatic stellate cells and myofibroblasts in liver fibrosis. Cells 8: 503, 2019. doi: 10.3390/cells8050503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ramachandran P, Dobie R, Wilson-Kanamori JR, Dora EF, Henderson BEP, Luu NT, Portman JR, Matchett KP, Brice M, Marwick JA, Taylor RS, Efremova M, Vento-Tormo R, Carragher NO, Kendall TJ, Fallowfield JA, Harrison EM, Mole DJ, Wigmore SJ, Newsome PN, Weston CJ, Iredale JP, Tacke F, Pollard JW, Ponting CP, Marioni JC, Teichmann SA, Henderson NC. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575: 512–518, 2019. doi: 10.1038/s41586-019-1631-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ægidius HM, Veidal SS, Feigh M, Hallenborg P, Puglia M, Pers TH, Vrang N, Jelsing J, Kornum BR, Blagoev B, Rigbolt KTG. Multi-omics characterization of a diet-induced obese model of non-alcoholic steatohepatitis. Sci Rep 10: 1148, 2020. doi: 10.1038/s41598-020-58059-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Birchmeier W. Orchestrating Wnt signalling for metabolic liver zonation. Nat Cell Biol 18: 463–465, 2016. doi: 10.1038/ncb3349. [DOI] [PubMed] [Google Scholar]
- 9.Jungermann K, Katz N. Functional hepatocellular heterogeneity. Hepatology 2: 385S–395S, 1982. doi: 10.1002/hep.1840020316. [DOI] [PubMed] [Google Scholar]
- 10.Jungermann K, Katz N. Functional specialization of different hepatocyte populations. Physiol Rev 69: 708–764, 1989. doi: 10.1152/physrev.1989.69.3.708. [DOI] [PubMed] [Google Scholar]
- 11.Katz NR. Metabolic heterogeneity of hepatocytes across the liver acinus. J Nutr 122: 843–849, 1992. doi: 10.1093/jn/122.suppl_3.843. [DOI] [PubMed] [Google Scholar]
- 12.Bahar Halpern K, Shenhav R, Matcovitch-Natan O, Toth B, Lemze D, Golan M, Massasa EE, Baydatch S, Landen S, Moor AE, Brandis A, Giladi A, Stokar-Avihail A, David E, Amit I, Itzkovitz S. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542: 352–356, 2017. doi: 10.1038/nature21065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.MacParland SA, Liu JC, Ma XZ, Innes BT, Bartczak AM, Gage BK, , et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun 9: 4383, 2018. doi: 10.1038/s41467-018-06318-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chalasani N, Wilson L, Kleiner DE, Cummings OW, Brunt EM, Unalp A; NASH Clinical Research Network. Relationship of steatosis grade and zonal location to histological features of steatohepatitis in adult patients with non-alcoholic fatty liver disease. J Hepatol 48: 829–834, 2008. doi: 10.1016/j.jhep.2008.01.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hall Z, Bond NJ, Ashmore T, Sanders F, Ament Z, Wang X, Murray AJ, Bellafante E, Virtue S, Vidal-Puig A, Allison M, Davies SE, Koulman A, Vacca M, Griffin JL. Lipid zonation and phospholipid remodeling in nonalcoholic fatty liver disease. Hepatology 65: 1165–1180, 2017. doi: 10.1002/hep.28953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hijmans BS, Grefhorst A, Oosterveer MH, Groen AK. Zonation of glucose and fatty acid metabolism in the liver: mechanism and metabolic consequences. Biochimie 96: 121–129, 2014. doi: 10.1016/j.biochi.2013.06.007. [DOI] [PubMed] [Google Scholar]
- 17.Font-Burgada J, Shalapour S, Ramaswamy S, Hsueh B, Rossell D, Umemura A, Taniguchi K, Nakagawa H, Valasek MA, Ye L, Kopp JL, Sander M, Carter H, Deisseroth K, Verma IM, Karin M. Hybrid periportal hepatocytes regenerate the injured liver without giving rise to cancer. Cell 162: 766–779, 2015. doi: 10.1016/j.cell.2015.07.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Stanger BZ. Probing hepatocyte heterogeneity. Cell Res 25: 1181–1182, 2015. doi: 10.1038/cr.2015.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Takahashi Y, Fukusato T. Histopathology of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. World J Gastroenterol 20: 15539–15548, 2014. doi: 10.3748/wjg.v20.i42.15539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang B, Zhao L, Fish M, Logan CY, Nusse R. Self-renewing diploid Axin2(+) cells fuel homeostatic renewal of the liver. Nature 524: 180–185, 2015. doi: 10.1038/nature14863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Berlanga A, Guiu-Jurado E, Porras JA, Auguet T. Molecular pathways in non-alcoholic fatty liver disease. Clin Exp Gastroenterol 7: 221–239, 2014. doi: 10.2147/CEG.S62831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, Tirosh I, Bialas AR, Kamitaki N, Martersteck EM, Trombetta JJ, Weitz DA, Sanes JR, Shalek AK, Regev A, McCarroll SA. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161: 1202–1214, 2015. doi: 10.1016/j.cell.2015.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29: 15–21, 2013. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Smets FN, Chen Y, Wang LJ, Soriano HE. Loss of cell anchorage triggers apoptosis (anoikis) in primary mouse hepatocytes. Mol Genet Metab 75: 344–352, 2002. doi: 10.1016/S1096-7192(02)00004-5. [DOI] [PubMed] [Google Scholar]
- 25.Sufiandi S, Obara H, Enosawa S, Hsu HC, Matsuno N, Mizunuma H. Improvement of infusion process in cell transplantation: effect of shear stress on hepatocyte viability under horizontal and vertical syringe orientation. Cell Med 7: 59–66, 2015. doi: 10.3727/215517914X685150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM 3rd, Hao Y, Stoeckius M, Smibert P, Satija R. Comprehensive integration of single-cell data. Cell 177: 1888–1902.e21, 2019. doi: 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.van der Maaten L, Hinton G. Visualizing data using t-SNE. JMLR 9: 2579–2605, 2008. [Google Scholar]
- 28.Becht E, McInnes L, Healy J, Dutertre CA, Kwok IWH, Ng LG, Ginhoux F, Newell EW. Dimensionality reduction for visualizing single-cell data using UMAP. Nat Biotechnol 37: 38–44, 2019. doi: 10.1038/nbt.4314. [DOI] [PubMed] [Google Scholar]
- 29.van Dijk D, Sharma R, Nainys J, Yim K, Kathail P, Carr AJ, Burdziak C, Moon KR, Chaffer CL, Pattabiraman D, Bierie B, Mazutis L, Wolf G, Krishnaswamy S, Pe’er D. Recovering gene interactions from single-cell data using data diffusion. Cell 174: 716–729.e27, 2018. doi: 10.1016/j.cell.2018.05.061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Huang M, Wang J, Torre E, Dueck H, Shaffer S, Bonasio R, Murray JI, Raj A, Li M, Zhang NR. SAVER: gene expression recovery for single-cell RNA sequencing. Nat Methods 15: 539–542, 2018. doi: 10.1038/s41592-018-0033-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ghafoory S, Breitkopf-Heinlein K, Li Q, Scholl C, Dooley S, Wolfl S. Zonation of nitrogen and glucose metabolism gene expression upon acute liver damage in mouse. PLoS One 8: e78262, 2013. doi: 10.1371/journal.pone.0078262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ge SX, Jung D, Yao R. ShinyGO: a graphical enrichment tool for animals and plants. Bioinformatics 36: 2628–2629, 2020. doi: 10.1093/bioinformatics/btz931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16: 284–287, 2012. doi: 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Cho CS, Park HW, Ho A, Semple IA, Kim B, Jang I, Park H, Reilly S, Saltiel AR, Lee JH. Lipotoxicity induces hepatic protein inclusions through TANK binding kinase 1-mediated p62/sequestosome 1 phosphorylation. Hepatology 68: 1331–1346, 2018. doi: 10.1002/hep.29742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pan X, Wang P, Luo J, Wang Z, Song Y, Ye J, Hou X. Adipogenic changes of hepatocytes in a high-fat diet-induced fatty liver mice model and non-alcoholic fatty liver disease patients. Endocrine 48: 834–847, 2015. doi: 10.1007/s12020-014-0384-x. [DOI] [PubMed] [Google Scholar]
- 36.Evarts RP, Nagy P, Marsden E, Thorgeirsson SS. In situ hybridization studies on expression of albumin and alpha-fetoprotein during the early stage of neoplastic transformation in rat liver. Cancer Res 47: 5469–5475, 1987. [PubMed] [Google Scholar]
- 37.Torre C, Perret C, Colnot S. Transcription dynamics in a physiological process: beta-catenin signaling directs liver metabolic zonation. Int J Biochem Cell Biol 43: 271–278, 2011. doi: 10.1016/j.biocel.2009.11.004. [DOI] [PubMed] [Google Scholar]
- 38.Kim S, Sohn I, Ahn JI, Lee KH, Lee YS, Lee YS. Hepatic gene expression profiles in a long-term high-fat diet-induced obesity mouse model. Gene 340: 99–109, 2004. doi: 10.1016/j.gene.2004.06.015. [DOI] [PubMed] [Google Scholar]
- 39.Siersbæk M, Varticovski L, Yang S, Baek S, Nielsen R, Mandrup S, Hager GL, Chung JH, Grøntved L. High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss. Sci Rep 7: 40220, 2017. doi: 10.1038/srep40220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Soltis AR, Kennedy NJ, Xin X, Zhou F, Ficarro SB, Yap YS, Matthews BJ, Lauffenburger DA, White FM, Marto JA, Davis RJ, Fraenkel E. Hepatic dysfunction caused by consumption of a high-fat diet. Cell Rep 21: 3317–3328, 2017. doi: 10.1016/j.celrep.2017.11.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Patsouris D, Reddy JK, Muller M, Kersten S. Peroxisome proliferator-activated receptor alpha mediates the effects of high-fat diet on hepatic gene expression. Endocrinology 147: 1508–1516, 2006. doi: 10.1210/en.2005-1132. [DOI] [PubMed] [Google Scholar]
- 42.Wang Y, Botolin D, Xu J, Christian B, Mitchell E, Jayaprakasam B, Nair MG, Peters JM, Busik JV, Olson LK, Jump DB. Regulation of hepatic fatty acid elongase and desaturase expression in diabetes and obesity. J Lipid Res 47: 2028–2041, 2006. doi: 10.1194/jlr.M600177-JLR200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Koonen DP, Jacobs RL, Febbraio M, Young ME, Soltys CL, Ong H, Vance DE, Dyck JR. Increased hepatic CD36 expression contributes to dyslipidemia associated with diet-induced obesity. Diabetes 56: 2863–2871, 2007. doi: 10.2337/db07-0907. [DOI] [PubMed] [Google Scholar]
- 44.Libby AE, Bales E, Orlicky DJ, McManaman JL. Perilipin-2 deletion impairs hepatic lipid accumulation by interfering with sterol regulatory element-binding protein (SREBP) activation and altering the hepatic lipidome. J Biol Chem 291: 24231–24246, 2016. doi: 10.1074/jbc.M116.759795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Najt CP, Senthivinayagam S, Aljazi MB, Fader KA, Olenic SD, Brock JR, Lydic TA, Jones AD, Atshaves BP. Liver-specific loss of Perilipin 2 alleviates diet-induced hepatic steatosis, inflammation, and fibrosis. Am J Physiol Gastrointest Liver Physiol 310: G726–G738, 2016. doi: 10.1152/ajpgi.00436.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wilson CG, Tran JL, Erion DM, Vera NB, Febbraio M, Weiss EJ. Hepatocyte-specific disruption of CD36 attenuates fatty liver and improves insulin sensitivity in HFD-fed mice. Endocrinology 157: 570–585, 2016. doi: 10.1210/en.2015-1866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhang X, Heckmann BL, Campbell LE, Liu J. G0S2: A small giant controller of lipolysis and adipose-liver fatty acid flux. Biochim Biophys Acta Mol Cell Biol Lipids 1862: 1146–1154, 2017. doi: 10.1016/j.bbalip.2017.06.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhang X, Li S, Zhou Y, Su W, Ruan X, Wang B, Zheng F, Warner M, Gustafsson JA, Guan Y. Ablation of cytochrome P450 omega-hydroxylase 4A14 gene attenuates hepatic steatosis and fibrosis. Proc Natl Acad Sci USA 114: 3181–3185, 2017. doi: 10.1073/pnas.1700172114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang X, Xie X, Heckmann BL, Saarinen AM, Czyzyk TA, Liu J. Targeted disruption of G0/G1 switch gene 2 enhances adipose lipolysis, alters hepatic energy balance, and alleviates high-fat diet-induced liver steatosis. Diabetes 63: 934–946, 2014. doi: 10.2337/db13-1422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Schleicher J, Tokarski C, Marbach E, Matz-Soja M, Zellmer S, Gebhardt R, Schuster S. Zonation of hepatic fatty acid metabolism - The diversity of its regulation and the benefit of modeling. Biochim Biophys Acta 1851: 641–656, 2015. doi: 10.1016/j.bbalip.2015.02.004. [DOI] [PubMed] [Google Scholar]
- 51.Winzell MS, Ahren B. The high-fat diet-fed mouse: a model for studying mechanisms and treatment of impaired glucose tolerance and type 2 diabetes. Diabetes 53: S215–S219, 2004. doi: 10.2337/diabetes.53.suppl_3.S215. [DOI] [PubMed] [Google Scholar]
- 52.Kersten S. Integrated physiology and systems biology of PPARalpha. Mol Metab 3: 354–371, 2014. doi: 10.1016/j.molmet.2014.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.McMullen PD, Bhattacharya S, Woods CG, Sun B, Yarborough K, Ross SM, Miller ME, McBride MT, LeCluyse EL, Clewell RA, Andersen ME. A map of the PPARalpha transcription regulatory network for primary human hepatocytes. Chem Biol Interact 209: 14–24, 2014. doi: 10.1016/j.cbi.2013.11.006. [DOI] [PubMed] [Google Scholar]
- 54.Gierahn TM, Wadsworth MH 2nd, Hughes TK, Bryson BD, Butler A, Satija R, Fortune S, Love JC, Shalek AK. Seq-Well: portable, low-cost RNA sequencing of single cells at high throughput. Nat Methods 14: 395–398, 2017. doi: 10.1038/nmeth.4179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Saviano A, Henderson NC, Baumert TF. Single-cell genomics and spatial transcriptomics: Discovery of novel cell states and cellular interactions in liver physiology and disease biology. J Hepatol 73: 1219–1230, 2020. doi: 10.1016/j.jhep.2020.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The scRNA-seq data set generated from this study is available at the Gene Expression Omnibus database (GEO accession no. GSE157281). The data can also be accessed through an interactive online resource (https://lee.lab.medicine.umich.edu/hfd), which has an intuitive graphical user interface for exploring our scRNA-seq data set.