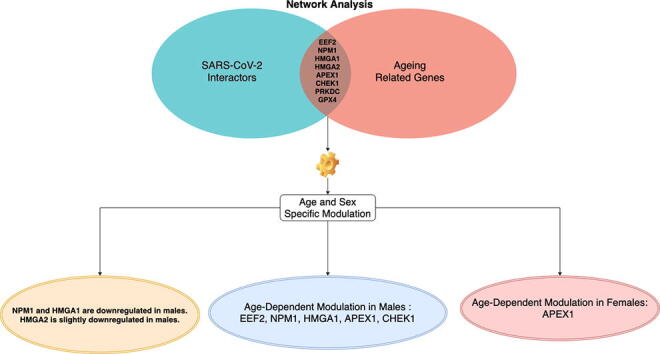
Graphical abstract
Keywords: Data science, SARS-CoV-2, COVID-19, Ageing genes, Interactomes
Abstract
Motivation: Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection (coronavirus disease, 2019; COVID-19) is associated with adverse outcomes in patients. It has been observed that lethality seems to be related to the age of patients. While ageing has been extensively demonstrated to be accompanied by some modifications at the gene expression level, a possible link with COVID-19 manifestation still need to be investigated at the molecular level.
Objectives: This study aims to shed out light on a possible link between the increased COVID-19 lethality and the molecular changes that occur in elderly people.
Methods: We considered public datasets of ageing-related genes and their expression at the tissue level. We selected human proteins interacting with viral ones that are known to be related to the ageing process. Finally, we investigated changes in the expression level of coding genes at the tissue, gender and age level.
Results: We observed a significant intersection between some SARS-CoV-2 interactors and ageing-related genes, suggesting that those genes are particularly affected by COVID-19 infection. Our analysis evidenced that virus infection particularly involves ageing molecular mechanisms centred around proteins EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC, and GPX4. We found that HMGA1 and NPM1 have different expressions in the lung of males, while HMGA1, APEX1, CHEK1, EEF2, and NPM1 present changes in expression in males due to ageing effects.
Conclusion: Our study generated a mechanistic framework to clarify the correlation between COVID-19 incidence in elderly patients and molecular mechanisms of ageing. We also provide testable hypotheses for future investigation and pharmacological solutions tailored to specific age ranges.
1. Introduction
At the end of 2019 in Wuhan (China), medical facilities reported acute pneumonia cases with an unknown origin. Further analysis revealed that a novel coronavirus, named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was responsible for that disease, subsequently called coronavirus disease 2019 (COVID-19) [1], [2]. The clinical manifestations spanned from asymptomatic infection to severe pneumonia and a severe state of inflammation (molecularly characterised by a cytokine storm) leading to a fatal outcome [3], [4], [5], [6], [7], [8].
Starting from China, the virus spread in almost all other countries globally, causing infections and deaths. On 11th March 2020, the World Health Organisation (WHO) declared SARS-CoV-2 as a pandemic. Current data revealed that the impact of COVID-19 presents certain peculiar aspects in different nations that have been deeply investigated [9], [10]. Some authors hypothesised that virus mutations were responsible for these differences [11], [12], [13], [14]. Nevertheless, many independent studies agreed that the mutations might not have a primary role in explaining these differences [15], [16], [17].
Despite the lack of the individuation of the causes, there was a substantial agreement on the fact that the variation of the observed case fatality rate (CFR), i.e. the fraction of confirmed cases leading to fatal outcomes, ranging from 0 to 20% and beyond at country level, needs to be deeply investigated [18], [19], [20]. Among the other differences, we focused on observing that the infection is significantly more lethal in older people [21], [22], [23], [24], [25]. This consideration has also guided the optimisation of vaccination strategy [26].
Some studies have focused on the possible link between increased mortality rate and some characteristics of older people [27], [28]. In addition, these studies suggested the potential effect of the virus as a trigger activating the decompensation of other chronic conditions [29], [30], [31], [32]. Akbar et al., [33], discussed a possible link between the increased chronic inflammatory status occurring during ageing (termed ”inflammaging” [34], [35]), and COVID-19 manifestation that causes the rise of inflammation.
Previous studies have also shown that the understanding of modification of molecular mechanisms related to the ageing process (i.e. modification of gene expression and modulation of regulatory mechanisms) may reveal important insights about ageing [36]. Many studies contributed to identifying such ageing-related diseases despite the lack of having experimental data [37], [38], [39], [35], [40]. Computational predictions have also been made in [36], [41] giving both candidate genes and networks [42], [43].
Consequently, the study of the intersection between SARS-CoV-2 and ageing-related molecular alterations could augment the understanding of COVID-19, thus improving treatment options [44]. Bhattacharyya et al. presented a first analysis based on some preliminary public data reinforcing the rationale that such a possible link exists [45]. The expression of the two human receptors TMPRSS2 and ACE2, which are recognised by the SARS-CoV-2 protein Spike, increases with age in mammals [46], further suggesting a molecular cause for the more severe COVID-19 symptoms with age.
Six functional open reading frames (ORFs) in the SARS-CoV-2 genome encodes for the four main structural proteins, the Spike (S), Envelope (E), Membrane (M), and the Nucleocapsid (N), and ORF1a/ORF1b, which contain information for the replicase–transcriptase complex formed by 16 non-structural proteins (NSP1–NSP16). The SARS-CoV-2 genome also contains 9 accessory factors from sub-genomic ORFs (Orf3a, 3b, 6, 7a, 7b, 8, 9b, 9c and 10) [47]. We investigated the relationships and interactions between these viral components and age-related factors and observed a significant overlap between SARS-CoV-2 and ageing group genes’ interactors, considering possible regulatory mechanisms that may be altered [48], [43], [49].
Starting from these considerations, we hypothesised that SARS-CoV-2 interacting proteins (and genes) might show an overlap with human ageing-related genes higher than chance. Therefore, the infection may affects these mechanisms that can be already impaired in older adults, causing severe outcomes. We downloaded public available interaction data from Guzzi et al. [50] and Gordon et al. [51]. Then we considered the interacting partners that were annotated as ageing genes in MSigDB database [52] ad we also considered the expression at tissue and sex levels extracting data from the GTEx database [53]. We identified a significant fraction of interacting partners of SARS-CoV-2 involved in ageing. These genes are also expressed in the lung, and their expression is modulated by age and sex, (while we also observed that these genes are expressed in adipose tissue as reported in Supplementary Material). The workflow of the experiment is depicted in Fig. 1.
Fig. 1.
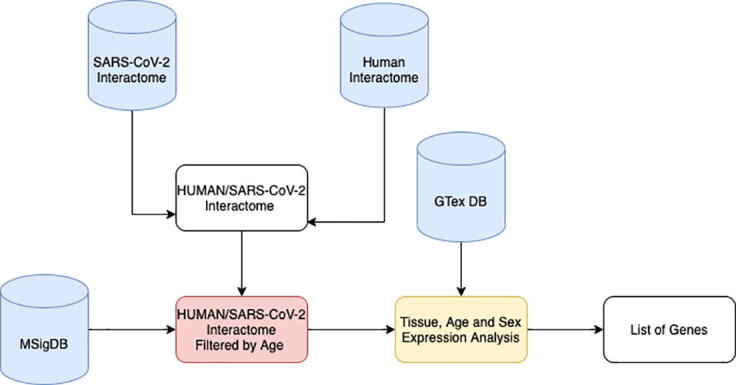
Workflow of the experiment. We downloaded public available interaction data from previous studies. We built the integrated human/SARS-CoV-2 interactome. In parallel, we downloaded the list of genes annotated with ageing keywords as in MSigDB database. Then, for each SARS-CoV-2 protein, we calculated the probability that it contains human interactors annotated with ageing keyword. We obtained a list of SARS-CoV-2 proteins containing a significant number of interactors related to ageing. Then we calculated the intersection of these sets (core interactors) obtaining a list of eight human proteins. For each core interactor, we also considered the expression at tissue level extracting data from GTEx database. We verified that there exist a significant fraction of interacting partners of SARS-CoV-2 that are involved in ageing and that are particularly expressed in lung and in adipose tissue.
2. Methods
SARS-CoV-2 Interaction Map. We considered the SARS-CoV-2 protein interaction map provided by Gordon et al., [51], and by Guzzi et al., [50]. Both works provided data about 26 of the 29 SARS-CoV-2 proteins behaviour in human cells by identifying the human proteins that are physically associated with each of the SARS-CoV-2 proteins using affinity-purification mass spectrometry. They found high-confidence protein–protein interactions between SARS-CoV-2 and human proteins; they also provided data about possible interactions with an associated reliability score. We considered both high and low confidence interactions.
Databases. We first defined and labelled genes related to the ageing process as ageing. Then, we considered data provided from the GTEx dataset containing genes positively and negatively correlated with human age [53]. We gathered data from the GenAge dataset that derived human genes by projecting sequence orthologs in model organisms. We also considered the MSigDB gene set collections, which summarised gene information associated with ageing collected from 70 different studies. We selected datasets reporting experiments from Homo sapiens since orthologs’ projection may produce not reliable results for ageing as described in [36].
We used the Search Tool for the Retrieval of Interacting Genes Proteins database (STRING) [54] that is a freely available repository storing both physical and functional association among proteins. Users may search the database through a web interface by specifying a protein identifier or inserting the primary sequence. We queried the database using the identifiers of the nodes of each subnetwork. We used medium confidence as the minimum confidence score for each interaction and all for the sources of interactions. We searched the GTEx Portal [55] using the previously described list of gens. We obtained the expression of those genes in a heat map that shows expression across all GTEx tissues. Gene Ontology analysis was performed by using Gene Ontology web portal [56] while using Reactome Database for identifying related pathways [57].
Bioinformatic and Network Analysis. We selected all known SARS-CoV-2 interacting partners. We used the Gordon dataset [51] to obtain all the partners. Then, for each SARS-COV-2 protein, we retrieved the list of its interactors. We determined the intersection between the list of human interactors and the ageing-related genes for each viral protein. We estimated the probability that this intersection is higher than chance by Fisher’s exact test. In Supplementary Material, we show the sub-networks induced in human interactome by each SARS-COV-2 protein. For each subnetwork, we report the main topological parameters: number of nodes, number of edges, average node degree, average local clustering coefficient, the expected number of edges. For each sub-network, we performed a Gene Ontology enrichment analysis. Network analysis and visualisation were performed in Cytoscape 3.7.0 [58]. We also tested the significance of the difference in the expression of EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC, and GPX4 due to age (we considered six different classes), sex, and tissue. All the p-values of the tests were corrected for multiple testing using Bonferroni correction. We used a Wilcoxon Test for testing difference in the expression among classes (since the expression of genes is not gaussian as reported by a Shapiro test). In addition, the difference among age classes is evaluated using a Kruskal Wallis test.
3. Results
3.1. Network analysis
We selected human interactors for each viral protein. The analysis revealed that only ten viral proteins (M, NSP2, NSP4, NSP6, NSP11, NSP13, Orf3a, Orf7a, Orf8, and Orf9c) have interactors with a significant overlap with respect to ageing-related proteins, as summarised in Table 1 (p-values have been corrected using Bonferroni correction). Then, we considered those that are enriched for ageing in a significant way. Finally, we intersected all these sets, and we obtain a core set of eight proteins: EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC and GPX2 (indicated as core interactors hereafter) as reported in Fig. 2 (see supplementary for the list of interactors for each viral protein, integrated with the topological characteristics of the induced subnetwork in the human interactome).
Table 1.
P-Values of the enrichment. For each protein, we report the significance of the enrichment after correction. A p-value lower than 0.01 means that the interactors are significantly related to ageing (NS stands for not significant).
| Viral Protein | P-Value | Viral Protein | P-Value |
|---|---|---|---|
| Spike | NS | E | NS |
| M | 6.84E−03 | N | NS |
| NSP1 | NS | NSP2 | 1.8E−03 |
| NSP3 | NS | NSP4 | 8.32E−03 |
| NSP5 | NS | NSP6 | 2.6E−03 |
| NSP7 | NS | NSP8 | 3.4E−03 |
| NSP9 | NS | NSP10 | NS |
| NSP11 | 1.8E−04 | NSP12 | NS |
| NSP13 | 2.5E−03 | NSP14 | NS |
| NSP15 | NS | NSP16 | NS |
| Orf3a | 5.06E−03 | Orf3b | NS |
| Orf6 | NS | Orf7a | 1.8E−04 |
| Orf7b | NS | Orf8 | 6.9E−04 |
| Orf9b | NS | Orf9c | 1.50E−02 |
| Orf10 | NS |
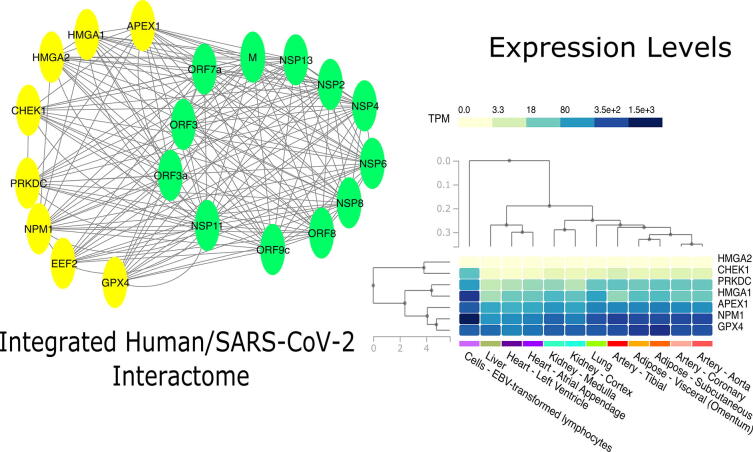
Fig. 2.
Figure shows tissue level analysis of this work. The Network analysis contributed to find a set of human proteins (yellow nodes) related to aging that interact with many SARS-CoV-2 proteins (green nodes). The analysis of the expression of the related genes at tissue level revealed that all these genes are expressed in the lung, as well as in other human tissues. Expression levels are presented as TPMs. (for interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article).
The Gene Ontology analysis revealed that the whole network is enriched with the following terms: (GO:0090402) oncogene-induced cell senescence, (GO:0035986) senescence-associated heterochromatin focus assembly, (GO:2000774) positive regulation of cellular senescence, (GO:2000773) negative regulation of cellular senescence, (GO:2000772) regulation of cellular senescence. The analysis of Reactome DB reveals that the subnetwork is associated with the following pathways: Formation of Senescence Associated Heterochromatin Foci (HSA2559584), Host interactions of HIV factors (HSA162909).
3.2. Expression analysis
We searched the GTEx database for the expression of core interactors as reported in Fig. 2 expressed as TPM (Transcripts Per Million). We found that all the interactors are expressed in the lung as well as in other human tissues (see supplementary materials for more details). To assess the different outcomes between males and females we focused on lung tissue and we compared the expression of these core interactors in males and females as reported in 3. Since data were not normally distributed (as given by Shapiro Test), we applied a Wilcoxon Test to evaluate significance of the difference in expression between male/female classes.
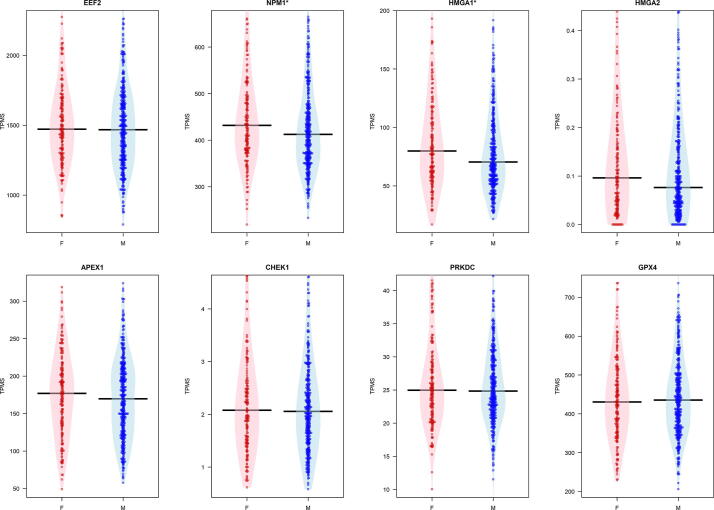
We evidenced a significant difference for NPM1 and HMGA1 which are significantly downregulated in males, without considering age as reported in Fig. 3.
Fig. 3.
Figure reports box plot of the expression of the eight core genes grouped by sex in the lung tissue. The evidences a significant difference tested by using a Wilcoxon Test for NPM1 and HMGA1 genes.
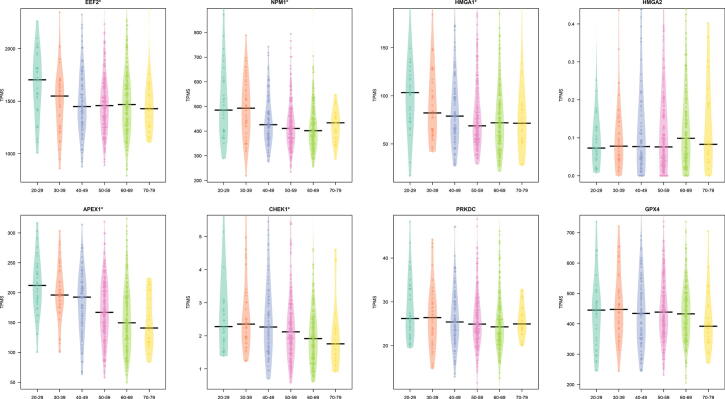
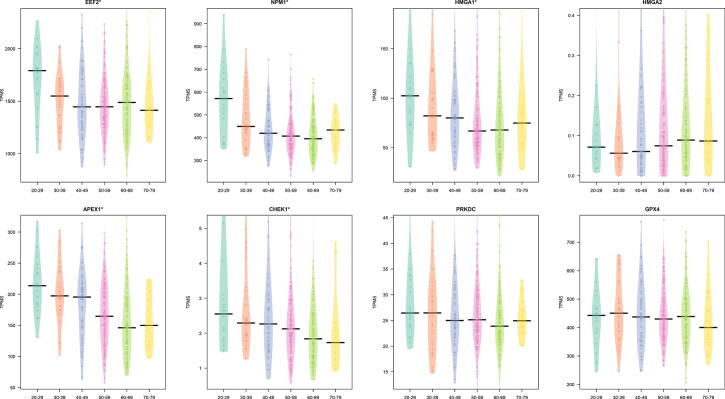
We also explored the trend of the core interactors focusing on lung tissue and six different classes of age (20–29, 30–39, 40–49, 50–59, 60–69, 70–79). We found a significant difference considering age groups for HMGA1, APEX, CHEK1, EEF2, and NPM1 (p 0.05 as evidenced by a Kruskal Wallis test). Fig. 4 reports this trend.
Fig. 4.
Figure reports the difference of the expression of the core genes in lung tissue in different age classes. A on top of the plot means a significant difference ( as evidenced by a Kruskal Wallis test).
4. Discussion
Deaths from COVID-19 occur predominantly among older adults. COVID-19 also appears to be more lethal for men rather than women [23], [9], [10], [24]. This feature has been found in China, as well as in Europe and in the United States of America [59].
Starting from this observation, we investigated the molecular basis of this phenomenon. Next, we recall that ageing is a heterogeneous process that presents differences among individuals. In particular, age-related changes impact many organs producing possible multi-organ failures, even showing many inter-individual differences. Beyond these differences, we tried to explain how the age-related changes at the molecular level can be relevant to COVID-19 pathology.
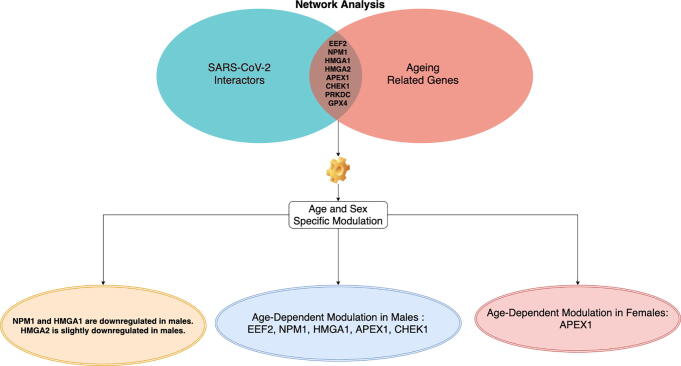
To achieve this goal, we integrated interactomics and expression data related to COVID-19, age and sex. We started from SARS-CoV-2 interactors, and we isolated age-related from those. Then we considered the expression value of these genes, and we further investigated the trend of changes of these genes in age and sex groups. We identified a set of statistically significant interactors for the ageing process: EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC, and GPX4. As reported in Fig. 7, we found some interesting changes of these genes considering tissue, age and sex groups. We also found that NPM1 and HMGA1 are downregulated in males (statistically significant regulation), while HMGA2 is slightly downregulated in males (not significantly) (Fig. 3).
Fig. 7.
Figure summarises main results of the work. Network analysis found that there exist eight proteins related to ageing that are also all targeted by ten SARS-CoV-2 proteins. The analysis of the expression of their genes revealed that there exist difference on the expression of these genes considering both age and sex.
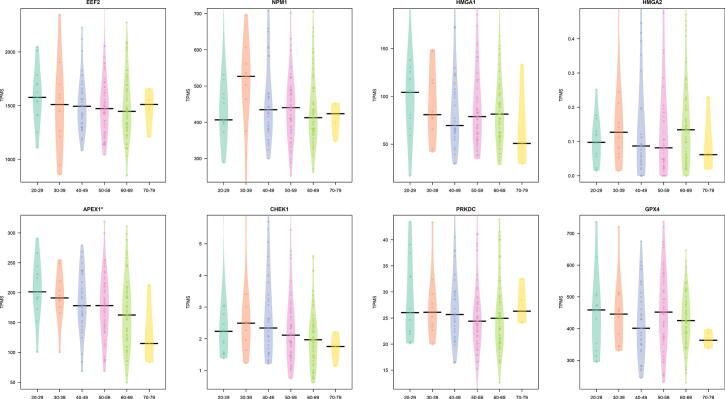
We also found some statistically relevant changes in age for EEF2, NPM1, HMGA1, APEX1, and CHEK1 for males (Fig. 5), and for APEX1 in Females (Fig. 6). With the only exception of HMGA2, all these genes show a decreased expression with ageing in lung tissues.
Fig. 5.
Difference in the expression in lung tissue by age classes in males. Expression is reported as TPM.A on top reveals a modulation in groups.
Fig. 6.
Difference in the expression in lung tissue by age classes in females. Expression is reported as TPM. A on top reveals a modulation in groups.
As investigated in [60], ageing is characterised by the decline of the immune function. Older adults are not immuno-deficient, but the immune system’s response is often not sufficient to be effective against antigens. This effect is particularly evident when they are subject to novel antigens. For example, it is known that both responses to influenza and vaccination are not efficient in the elderly [61], [62]. Moreover, the elderly accumulate inflammatory mediators in tissues (inflammageing process), which may occur by the accumulation of DNA lesions that, in turn, triggers the increased production of inflammatory mediators [63]. In parallel, the link between COVID-19 and the suppression of the immune system has been observed in [64]. Authors found that many proteins related to the immune response were modulated, causing the possible suppression of such a system.
HMGA1 and HMGA2 genes encode four proteins (HMGA1a, HMGA1b, HMGA1c, and HMGA2) belonging to the High-mobility group A (HMGA) protein family [65]. All the proteins bind AT-rich regions in DNA and modulate gene expression by acting as transcription factors. Literature reports that HMGA1 has critical roles in tumorigenesis and the progression of various cancers. However, the role of HMGA1 in COVID-19 has not been explored in the past. We now provide a hypothesis framework for future research in the functional interplay between ageing and SARS-CoV-2 infection. HMGA1 is significantly downregulated both in males and the elderly, and these differences may be associated with poor outcomes observed in these classes. It has been shown that HMGA1 induces inflammatory pathways in many cancers, enhance the expression of genes related to neural stmness and pathways involved cell cycle progression. HMGA1 dysregulation causes aberration in cellular development and hematopoiesis [66]. Furthermore, the involvement of HMGA1 in the transcriptional regulation of genes essential in both the inflammatory response and atherosclerosis has been established [67].
Our results suggest that low HMGA1 levels may be a risk factor in COVID-19 patients, given the possibility that interactions between SARS-CoV-2 and HMGA1 may impair/trigger inflammatory pathways. Furthermore, it has been demonstrated that low HMGA1 levels in basal stem/progenitor cells of the human airway epithelium are associated with suppression of the expression of genes critical to normal differentiation and up-regulation of genes linked to abnormal differentiation relevant to smoking and chronic obstructive pulmonary disease [68], which have been demonstrated to be risk factors associated with COVID-19 mortality [69].
Similarly to HMGA1, the Nucleophosmin (NPM1) is also downregulated in males. NPM1 is related to DNA and cell cycle control such as ribosome biogenesis, protein chaperoning, centrosome duplication, histone assembly, and cell proliferation [70], [71]. Previous studies investigated the age incidence of acute myeloid leukaemia with mutated nucleophosmin (NPM1) [72], [73], while there are no studies related to these mutations and other diseases. In [74] the impact of NPM1 modification in older patients has been investigated for AML, suggesting a worse prognosis for older patients due to NPM1 changes. The interaction between NPM1 and the nucleocapsid protein of the previous SARS-CoV is known to affect the viral particle assembly [75], [76], [77]. The role of NPM1 and Histone H2AX targeted by other viral proteins has also been reported in other viruses such as Epstein-Barr and KSHV as a common strategy to manipulate translation and to promote virus latency [78], [79]. A case of SARS-CoV-2 associated sudden death in an NPM-mutated AML 50-year-old male patient was reported in [80]. Together with our findings, this suggests that further studies on interactions between SARS-CoV-2 and NPM1 are required. Moreover, for older men, the scenario is furtherly complicated by the downregulation of EEF2, APEX1 and CHEK1.
The dysregulation of EEF2 may cause the accumulation of DNA damage [81]. The role of EEF2 in severe cases of COVID-19 has also been elucidated in [64], and the possible association of downregulation of EEF2 with COVID-19 severity is also suggested by our study. Moreover, this protein is targeted together with the Eukaryotic translation initiation factor 2 subunit 1 (EIF2S1) by Orf3a, Orf8, NSP2, NSP6, NSP11, NSP13, indicating a possible role of the virus to promote viral translation over cellular translation [82]. In [83] the synergistic downregulation of both APEX1 and NPM1 has been clearly observed in oligodendrocyte cells in relation to ageing APEX1 plays a protective role in the cellular response to oxidative stress [84], and has a major role in DNA repair and in redox regulation of transcription factors [73]. CHEK1 is targeted together with CDK1 by many SARS-CoV-2 interactors (NSP2, NSP4, NSP11, NSP13) and with CDKN2A (Orf3, NSP13), suggesting an additive effect on the disruption of pathways of apoptosis mediated by TP53 [85] yet dis-regulated by both senescence and ageing. [86].
Differently, for females we found only the age-dependent modulation of APEX1. Thus, this may suggest that females may have less risk factors than males.
In parallel, in supplementary material we report that core interactors are also significantly overexpressed in adipose tissue, therefore suggesting a second factor of co-morbidity. Changes in adipose tissue promote a chronic state of low-grade systemic inflammation on a phenotypic level, thus increasing the risk of age-associated diseases [35], [87]. Here, we report that core interactors are expressed in adipose tissue, suggesting a possible role that should be further investigated. We hypothesise that the molecular relationship between SARS-CoV-2 and aging is intrinsical: on one side, SARS-CoV-2 induces a major change to the host cell’s transcriptome/proteome, with hundreds of transcripts/proteins affected [51], [88]; on the other side, this effect is larger in older transcriptomes [89]. Secondly, ageing modulates the expression of proteins necessary for the viral cycle of SARS-CoV-2 [46], including those included in the interactome described in this study.
5. Conclusion
We applied a bioinformatic analysis to perform a qualitative study of mechanisms of infection by SARS-CoV-2 in older people.
Several studies have shown in the past the modifications of genes and proteins that occur in older adults. Other studies have partially elucidated the mechanism of infections and the dysregulated pathways in COVID-19 patients.
We detected a statistically significant overlap between SARS-CoV-2 interacting proteins and those related to ageing, suggesting a potentially different response in older people. Our analysis showed that virus infection mainly affects ageing molecular mechanisms centred around proteins EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC, and GPX4. We also found that some of these genes are differentially expressed in lung tissues of the elderly, suggesting an increased susceptibility of the elderly to COVID-19 inflammatory-related manifestations. Finally, we found that there is a significant difference in the expression considering both age and sex.
While causality is often hard to derive in high-throughput datasets such as the proteomics/transcriptomics data on which our study is based [90], we believe that the capability of SARS-CoV-2 to interact with proteins increasing in abundance with ageing may justify part of the increased severity of COVID-19 in older individuals.
These results will provide a first step for understanding the molecular basis of the mechanism of infection and will shed light on infection progression. The limitation of this study is that the dataset is correlative, and thus it should be confirmed by in vivo experiments.
6. Key Points
-
•
A network-based analysis identified some molecular mechanisms that could play a role in the SARS-CoV-2 molecular aetiology and ultimately affect COVID-19 outcome.
-
•
Our analysis evidenced that virus infection particularly affects ageing molecular mechanisms centred around proteins EEF2, NPM1, HMGA1, HMGA2, APEX1, CHEK1, PRKDC, and GPX4.
-
•
We found an age-dependent modulation of EEF2, NPM1, HMGA1, APEX1 and CHEK1 in lung tissue of males.
-
•
We found an age-dependent modulation of APEX1 in females.
-
•
Our study generated a mechanistic framework aiming at clarifying the correlation between COVID-19 incidence in elderly patients and molecular mechanisms of ageing considering differences by age and sex.
Author contribution
F.M.G and P.H.G. conceived the main idea of this manuscript. D.M. performed the experimental analysis. F.M.G., D.M., and P.H.G. participated in the experimental phase and the discussion of the results. P.V participated in the design and implementation of data analysis and integration. E.P. participated in the writing of Discussion Section and also validated the clinical aspects of this work. All authors read and approved the manuscript.
CRediT authorship contribution statement
Daniele Mercatelli: Data curation, Visualization, Writing - original draft. Elisabetta Pedace: Writing - original draft, Conceptualization. Pierangelo Veltri: Supervision, Writing - review & editing. Federico M. Giorgi: Conceptualization, Methodology, Writing - original draft. Pietro Hiram Guzzi: Conceptualization, Methodology, Writing - original draft.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgement
Authors thank C.V. Cannistraci for useful suggestions and discussion during the preparation of this work.
Footnotes
Supplementary data associated with this article can be found, in the online version, at https://doi.org/10.1016/j.csbj.2021.07.002.
Supplementary data
The following are the Supplementary data to this article:
References
- 1.Burki Talha. Outbreak of coronavirus disease 2019. Lancet Infect Dis. 2020;20(3):292–293. doi: 10.1016/S1473-3099(20)30076-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Das Jayanta Kumar, Roy Swarup, Guzzi Pietro Hiram. Analyzing host-viral interactome of sars-cov-2 for identifying vulnerable host proteins during covid-19 pathogenesis. Infect Genet Evol. 2021:104921. doi: 10.1016/j.meegid.2021.104921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Huang Chaolin, Wang Yeming, Li Xingwang, Ren Lili, Zhao Jianping, Yi Hu, Zhang Li, Fan Guohui, Jiuyang Xu., Xiaoying Gu. Clinical features of patients infected with 2019 novel coronavirus in wuhan, china. Lancet. 2020;395(10223):497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang Dawei, Bo Hu., Chang Hu., Zhu Fangfang, Liu Xing, Zhang Jing, Wang Binbin, Xiang Hui, Cheng Zhenshun, Xiong Yong. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus–infected pneumonia in wuhan, china. Jama. 2020;323(11):1061–1069. doi: 10.1001/jama.2020.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Petrilli Christopher M, Jones Simon A, Yang Jie, Rajagopalan Harish, O’Donnell Luke, Chernyak Yelena, Tobin Katie A, Cerfolio Robert J, Francois Fritz, Horwitz Leora I. Factors associated with hospital admission and critical illness among 5279 people with coronavirus disease 2019 in new york city: prospective cohort study. BMJ 2020;369 [DOI] [PMC free article] [PubMed]
- 6.Biamonte Flavia, Botta Cirino, Mazzitelli Maria, Rotundo Salvatore, Trecarichi Enrico Maria, Foti Daniela, Torti Carlo, Viglietto Giuseppe, Torella Daniele, Costanzo Francesco. Combined lymphocyte/monocyte count, d-dimer and iron status predict covid-19 course and outcome in a long-term care facility. Researchsquare. 2020 doi: 10.1186/s12967-021-02744-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Meftahi Gholam Hossein, Jangravi Zohreh, Sahraei Hedayat, Bahari Zahra. The possible pathophysiology mechanism of cytokine storm in elderly adults with covid-19 infection: the contribution of ”inflame-aging”. Inflamm Res. 2020:1–15. doi: 10.1007/s00011-020-01372-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Guzzi Pietro Hiram, Tradigo Giuseppe, Veltri Pierangelo. Spatio-temporal resource mapping for intensive care units at regional level for covid-19 emergency in Italy. Int J Environ Res Public Health. 2020;17(10):3344. doi: 10.3390/ijerph17103344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dong Ensheng, Hongru Du., Gardner Lauren. An interactive web-based dashboard to track covid-19 in real time. Lancet Infect Dis. 2020;20(5):533–534. doi: 10.1016/S1473-3099(20)30120-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Guzzi Pietro H., Tradigo Giuseppe, Veltri Pierangelo. Regional resource assessment during the covid-19 pandemic in Italy: modeling study. JMIR Med Inf. 2021;9(3):e18933. doi: 10.2196/18933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ramírez Juan David, Munoz Marina, Hernández Carolina, Flórez Carolina, Gomez Sergio, Rico Angelica, Pardo Lisseth, Barros Esther C., Paniz-Mondolfi Alberto E. Genetic diversity among sars-cov2 strains in south america may impact performance of molecular detection. Pathogens. 2020;9(7):580. doi: 10.3390/pathogens9070580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Korber Bette, Fischer Will M., Gnanakaran Sandrasegaram, Yoon Hyejin, Theiler James, Abfalterer Werner, Hengartner Nick, Giorgi Elena E., Bhattacharya Tanmoy, Foley Brian. Tracking changes in sars-cov-2 spike: evidence that d614g increases infectivity of the covid-19 virus. Cell. 2020;182(4):812–827. doi: 10.1016/j.cell.2020.06.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gagliardi Ida, Patella Gemma, Michael Ashour, Serra Raffaele, Provenzano Michele, Andreucci Michele. Covid-19 and the kidney: From epidemiology to clinical practice. J Clin Med. 2020;9(8):2506. doi: 10.3390/jcm9082506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cannataro Mario, Harrison Andrew. Bioinformatics helping to mitigate the impact of covid-19-editorial; 2021 [DOI] [PMC free article] [PubMed]
- 15.Mercatelli Daniele, Holding Andrew N., Giorgi Federico M. Web tools to fight pandemics: the covid-19 experience. Briefings Bioinf. 2020 doi: 10.1093/bib/bbaa261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ortuso Francesco, Mercatelli Daniele, Guzzi Pietro Hiram, Giorgi Federico Manuel. Structural genetics of circulating variants affecting the sars-cov-2 spike/human ace2 complex. J Biomol Struct Dyna. 2021:1–11. doi: 10.1080/07391102.2021.1886175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Galicia Johnah C., Guzzi Pietro H., Giorgi Federico M., Khan Asma A. Predicting the response of the dental pulp to sars-cov2 infection: a transcriptome-wide effect cross-analysis. Genes Immunity. 2020;21(5):360–363. doi: 10.1038/s41435-020-00112-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sorci Gabriele, Faivre Bruno, Morand Serge. Explaining among-country variation in covid-19 case fatality rate. Scientific Rep. 2020;10(1):1–11. doi: 10.1038/s41598-020-75848-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Koff Wayne C., Williams Michelle A. Covid-19 and immunity in aging populations–a new research agenda. New Engl J Med. 2020 doi: 10.1056/NEJMp2006761. [DOI] [PubMed] [Google Scholar]
- 20.Marcon Gabriella, Tettamanti Mauro, Capacci Giorgia, Fontanel Giulia, Spanò Marco, Nobili Alessandro, Forloni Gianluigi, Franceschi Claudio. Covid-19 mortality in lombardy: the vulnerability of the oldest old and the resilience of male centenarians. Aging (Albany NY) 2020;12(15):15186. doi: 10.18632/aging.103872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mallapaty Smriti. The coronavirus is most deadly if you are older and male-new data reveal the risks. Nature. 2020:16–17. doi: 10.1038/d41586-020-02483-2. [DOI] [PubMed] [Google Scholar]
- 22.Li Tao, Lu Lei, Tao Yu, Zhang Weishuo, Wang Liuming, Bao Jing, Liu Bao, Duan Jun. Clinical characteristics of 312 hospitalized older patients with covid-19 in wuhan, china. Arch Gerontol Geriatrics. 2020;91:104185. doi: 10.1016/j.archger.2020.104185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cannistraci Carlo Vittorio, Valsecchi Maria Grazia, Capua Ilaria. Age-sex population adjusted analysis of disease severity in epidemics as a tool to devise public health policies for covid-19. Scientific Rep. 2021;11(1):1–8. doi: 10.1038/s41598-021-89615-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Milano Marianna, Cannataro Mario. Statistical and network-based analysis of italian covid-19 data: communities detection and temporal evolution. Int J Environ Res Public Health. 2020;17(12):4182. doi: 10.3390/ijerph17124182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kang Seung-Ji, Jung Sook In. Age-related morbidity and mortality among patients with covid-19. Infect Chemother. 2020;52(2):154. doi: 10.3947/ic.2020.52.2.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Matrajt Laura, Eaton Julia, Leung Tiffany, Brown Elizabeth R. Vaccine optimization for covid-19: who to vaccinate first? medRxiv. 2020 doi: 10.1126/sciadv.abf1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pietrobon Anna Julia, Teixeira Franciane Mouradian Emidio, Sato Maria Notomi. Immunosenescence and inflammaging: Risk factors of severe covid-19 in older people. Front Immunol. 2020;11 doi: 10.3389/fimmu.2020.579220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hazeldine Jon, Lord Janet M. Immunesenescence: a predisposing risk factor for the development of covid-19? Front Immunol. 2020;11:2381. doi: 10.3389/fimmu.2020.573662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Napoli Claudio, Tritto Isabella, Mansueto Gelsomina, Coscioni Enrico, Ambrosio Giuseppe. Immunosenescence exacerbates the covid-19. Arch Gerontol Geriatrics. 2020 doi: 10.1016/j.archger.2020.104174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Grasselli Giacomo, Pesenti Antonio, Cecconi Maurizio. Critical care utilization for the covid-19 outbreak in lombardy, italy: early experience and forecast during an emergency response. Jama. 2020;323(16):1545–1546. doi: 10.1001/jama.2020.4031. [DOI] [PubMed] [Google Scholar]
- 31.Docherty Annemarie B., Harrison Ewen M., Green Christopher A., Hardwick Hayley E., Pius Riinu, Norman Lisa, Holden Karl A., Read Jonathan M., Dondelinger Frank, Carson Gail. Features of 20 133 uk patients in hospital with covid-19 using the isaric who clinical characterisation protocol: prospective observational cohort study. BMJ. 2020;369 doi: 10.1136/bmj.m1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cianflone Eleonora, Torella Michele, Biamonte Flavia, De Angelis Antonella, Urbanek Konrad, Costanzo Francesco S., Rota Marcello, Ellison-Hughes Georgina M., Torella Daniele. Targeting cardiac stem cell senescence to treat cardiac aging and disease. Cells. 2020;9(6):1558. doi: 10.3390/cells9061558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Akbar Arne N., Gilroy Derek W. Aging immunity may exacerbate covid-19. Science. 2020;369(6501):256–257. doi: 10.1126/science.abb0762. [DOI] [PubMed] [Google Scholar]
- 34.Franceschi C., Garagnani P., Parini P., Giuliani C., Santoro A. Inflammaging: a new immune-metabolic viewpoint for age-related diseases. Nat Rev Endocrinol. 2018;14(10):576–590. doi: 10.1038/s41574-018-0059-4. [DOI] [PubMed] [Google Scholar]
- 35.Santoro A., Guidarelli G., Ostan R., Giampieri E., Fabbri C., Bertarelli C., Nicoletti C., Kadi F., de Groot L.C.P.G.M., Feskens E., Berendsen A., Brzozowska A., Januszko O., Kozlowska K., Fairweather-Tait S., Jennings A., Meunier N., Caumon E., Napoli A., Mercatelli D., Battista G., Capri M., Franceschi C., Bazzocchi A. Gender-specific association of body composition with inflammatory and adipose-related markers in healthy elderly Europeans from the NU-AGE study. Eur Radiol. 2019;29(9):4968–4979. doi: 10.1007/s00330-018-5973-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Faisal Fazle E., Milenković Tijana. Dynamic networks reveal key players in aging. Bioinformatics. 2014;30(12):1721–1729. doi: 10.1093/bioinformatics/btu089. [DOI] [PubMed] [Google Scholar]
- 37.Bolignano Davide, Mattace-Raso Francesco, Sijbrands Eric JG, Zoccali Carmine. The aging kidney revisited: a systematic review. Ageing Res Rev. 2014;14:65–80. doi: 10.1016/j.arr.2014.02.003. [DOI] [PubMed] [Google Scholar]
- 38.Kerepesi Csaba, Daróczy Bálint, Sturm Ádám, Vellai Tibor, Benczúr András. Prediction and characterization of human ageing-related proteins by using machine learning. Scientific Rep. 2018;8(1):4094. doi: 10.1038/s41598-018-22240-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Abenavoli L., Luigiano C., Guzzi P.H., Milic N., Morace C., Stelitano L., Consolo P., Miraglia S., Fagoonee S., Virgilio C. Serum adipokine levels in overweight patients and their relationship with non-alcoholic fatty liver disease. Panminerva Medica. 2014;56(2):189–193. [PubMed] [Google Scholar]
- 40.Guzzi Pietro H., Cannataro Mario. μ-cs: an extension of the tm4 platform to manage affymetrix binary data. BMC Bioinf. 2010;11(1):315. doi: 10.1186/1471-2105-11-315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tacutu Robi, Craig Thomas, Budovsky Arie, Wuttke Daniel, Lehmann Gilad, Taranukha Dmitri, Costa Joana, Fraifeld Vadim E., De Magalhaes Joao Pedro. Human ageing genomic resources: integrated databases and tools for the biology and genetics of ageing. Nucl Acids Res. 2012;41(D1):D1027–D1033. doi: 10.1093/nar/gks1155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Guzzi Pietro Hiram, Milenković Tijana. Survey of local and global biological network alignment: the need to reconcile the two sides of the same coin. Briefings Bioinf. 2018;19(3):472–481. doi: 10.1093/bib/bbw132. [DOI] [PubMed] [Google Scholar]
- 43.Nassa Giovanni, Tarallo Roberta, Guzzi Pietro H., Ferraro Lorenzo, Cirillo Francesca, Ravo Maria, Nola Ernesto, Baumann Marc, Nyman Tuula A., Cannataro Mario. Comparative analysis of nuclear estrogen receptor alpha and beta interactomes in breast cancer cells. Mol BioSyst. 2011;7(3):667–676. doi: 10.1039/c0mb00145g. [DOI] [PubMed] [Google Scholar]
- 44.Blagosklonny Mikhail V. From causes of aging to death from covid-19. Aging (Albany NY) 2020;12(11):10004. doi: 10.18632/aging.103493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Bhattacharyya Upasana, Thelma B.K. Age-related gene expression alterations by sars-cov-2 infection contribute to poor prognosis in elderly. J Genet. 2020;99(1):1–9. doi: 10.1007/s12041-020-01233-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bilinska Katarzyna, Jakubowska Patrycja, Von Bartheld Christopher S., Butowt Rafal. Expression of the sars-cov-2 entry proteins, ace2 and tmprss2, in cells of the olfactory epithelium: identification of cell types and trends with age. ACS Chem Neurosci. 2020;11(11):1555–1562. doi: 10.1021/acschemneuro.0c00210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Das Jayanta Kumar, Tradigo Giuseppe, Veltri Pierangelo, Guzzi Pietro H., Roy Swarup. Data science in unveiling covid-19 pathogenesis and diagnosis: evolutionary origin to drug repurposing. Briefings Bioinf. 2021;22(2):855–872. doi: 10.1093/bib/bbaa420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Cho Young-Rae, Mina Marco, Lu Yanxin, Kwon Nayoung, Guzzi Pietro H. M-finder: uncovering functionally associated proteins from interactome data integrated with go annotations. Proteome Sci. 2013;11(S1):S3. doi: 10.1186/1477-5956-11-S1-S3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Bernabeu-Wittel M., Ternero-Vega J.E., Díaz-Jiménez P., Conde-Guzmán C., Nieto-Martín M.D., Moreno-Gavino L., Delgado-Cuesta J., Rincón-Gómez M., Giménez-Miranda L., Navarro-Amuedo M.D. Death risk stratification in elderly patients with covid-19. a comparative cohort study in nursing homes outbreaks. Arch Gerontol Geriatrics. 2020;91:104240. doi: 10.1016/j.archger.2020.104240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Guzzi Pietro H., Mercatelli Daniele, Ceraolo Carmine, Giorgi Federico M. Master regulator analysis of the sars-cov-2/human interactome. J Clin Med. 2020;9(4):982. doi: 10.3390/jcm9040982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gordon David E., Jang Gwendolyn M., Bouhaddou Mehdi, Xu Jiewei, Obernier Kirsten, White Kris M., O’Meara Matthew J., Rezelj Veronica V., Guo Jeffrey Z., Swaney Danielle L. A sars-cov-2 protein interaction map reveals targets for drug repurposing. Nature. 2020:1–13. doi: 10.1038/s41586-020-2286-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Liberzon Arthur, Subramanian Aravind, Pinchback Reid, Thorvaldsdóttir Helga, Tamayo Pablo, Mesirov Jill P. Molecular signatures database (msigdb) 3.0. Bioinformatics. 2011;27(12):1739–1740. doi: 10.1093/bioinformatics/btr260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jia Kaiwen, Cui Chunmei, Gao Yuanxu, Zhou Yuan, Cui Qinghua. An analysis of aging-related genes derived from the genotype-tissue expression project (gtex) Cell Death Discovery. 2018;4(1):1–14. doi: 10.1038/s41420-018-0093-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Katerina C Nastou, David Lyon, Rebecca Kirsch, Sampo Pyysalo, Nadezhda T Doncheva, Marc Legeay, Tao Fang, Peer Bork, et al. The string database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nu- cleic Acids Research, page gkaa1074, 2020
- 55.Lonsdale John, Thomas Jeffrey, Salvatore Mike, Phillips Rebecca, Lo Edmund, Shad Saboor, Hasz Richard, Walters Gary, Garcia Fernando, Young Nancy. The genotype-tissue expression (gtex) project. Nat Genet. 2013;45(6):580–585. doi: 10.1038/ng.2653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gene Ontology Consortium. The gene ontology (go) database and informatics resource. Nucl Acids Res 2004;32(suppl_1):D258–D261 [DOI] [PMC free article] [PubMed]
- 57.Fabregat Antonio, Sidiropoulos Konstantinos, Garapati Phani, Gillespie Marc, Hausmann Kerstin, Haw Robin, Jassal Bijay, Jupe Steven, Korninger Florian, McKay Sheldon. The reactome pathway knowledgebase. Nucl Acids Res. 2016;44(D1):D481–D487. doi: 10.1093/nar/gkv1351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Shannon Paul, Markiel Andrew, Ozier Owen, Baliga Nitin S., Wang Jonathan T., Ramage Daniel, Amin Nada, Schwikowski Benno, Ideker Trey. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Cristina Polidori M., Sies Helmut, Ferrucci Luigi, Benzing Thomas. Covid-19 mortality as a fingerprint of biological age. Ageing Res Rev. 2021:101308. doi: 10.1016/j.arr.2021.101308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Encarnacion M.R., Beata B.M., Kenneth D. Cause, consequences, and reversal of immune system aging. J Clin Invest. 2013;123:65–958. doi: 10.1172/JCI64096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Thompson William W., Shay David K., Weintraub Eric, Brammer Lynnette, Cox Nancy, Anderson Larry J. Mortality associated with influenza and respiratory syncytial virus in the united states. JAMA. 2003;289(2):179–186. doi: 10.1001/jama.289.2.179. [DOI] [PubMed] [Google Scholar]
- 62.Reichert Thomas A., Simonsen Lone, Sharma Ashutosh, Pardo Scott A., Fedson David S., Miller Mark A. Influenza and the winter increase in mortality in the United States, 1959–1999. Am J Epidemiol. 2004;160(5):492–502. doi: 10.1093/aje/kwh227. [DOI] [PubMed] [Google Scholar]
- 63.Franceschi Claudio, Campisi Judith. Chronic inflammation (inflammaging) and its potential contribution to age-associated diseases. J Gerontol Ser A. 2014;69(Suppl_1):S4–S9. doi: 10.1093/gerona/glu057. [DOI] [PubMed] [Google Scholar]
- 64.Tian Wenmin, Zhang Nan, Jin Ronghua, Feng Yingmei, Wang Siyuan, Gao Shuaixin, Gao Ruqin, Guizhen Wu., Tian Di, Tan Wenjie. Immune suppression in the early stage of covid-19 disease. Nat Commun. 2020;11(1):1–8. doi: 10.1038/s41467-020-19706-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cleynen Isabelle, Van de Ven Wim J.M. The hmga proteins: a myriad of functions. Int J Oncol. 2008;32(2):289–305. [PubMed] [Google Scholar]
- 66.Schuldenfrei A., Belton A., Kowalski J., Talbot C.C., Jr, Di Cello F., Poh W., Tsai H.L., Shah S.N., Huso T.H., Huso D.L., Resar L.M. HMGA1 drives stem cell, inflammatory pathway, and cell cycle progression genes during lymphoid tumorigenesis. BMC Genomics. 2011 Nov;4(12):549. doi: 10.1186/1471-2164-12-549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Carvajal Irvith M., Baron Rebecca M., Perrella Mark A. High-mobility group-i/y proteins: potential role in the pathophysiology of critical illnesses. Crit Care Med. 2002;30(1):S36–S42. [PubMed] [Google Scholar]
- 68.Zhang Haijun, Yang Jing, Walters Matthew S., Staudt Michelle R., Strulovici-Barel Yael, Salit Jacqueline, Mezey Jason G., Leopold Philip L., Crystal Ronald G. Mandatory role of hmga1 in human airway epithelial normal differentiation and post-injury regeneration. Oncotarget. 2018;9(18):14324. doi: 10.18632/oncotarget.24511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Reyes Felix M., Hache-Marliere Manuel, Karamanis Dimitris, Berto Cesar G., Estrada Rodolfo, Langston Matthew, Ntaios George, Gulani Perminder, Shah Chirag D., Palaiodimos Leonidas. Assessment of the association of copd and asthma with in-hospital mortality in patients with covid-19. A systematic review, meta-analysis, and meta-regression analysis. J Clin Med. 2021;10(10):2087. doi: 10.3390/jcm10102087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.López David J., Blas Ander de, Hurtado Mikel, García-Alija Mikel, Mentxaka Jon, Arada Igor de la, Urbaneja María A., no Marián Alonso-Mari, nuelos Sonia Ba. Nucleophosmin interaction with ape1: Insights into dna repair regulation. DNA Repair. 2020;88:102809. doi: 10.1016/j.dnarep.2020.102809. [DOI] [PubMed] [Google Scholar]
- 71.Storci Gianluca, Bacalini Maria Giulia, Bonifazi Francesca, Garagnani Paolo, Carolis Sabrina De, Salvioli Stefano, Olivieri Fabiola, Bonafè Massimiliano. Ribosomal dna instability: an evolutionary conserved fuel for inflammaging. Ageing Res Rev. 2020;58:101018. doi: 10.1016/j.arr.2020.101018. [DOI] [PubMed] [Google Scholar]
- 72.Thiede C., Creutzig E., Reinhardt D., Ehninger G., Creutzig U. Different types of npm1 mutations in children and adults: evidence for an effect of patient age on the prevalence of the tctg-tandem duplication in npm1-exon 12. Leukemia. 2007;21(2):366–367. doi: 10.1038/sj.leu.2404519. [DOI] [PubMed] [Google Scholar]
- 73.Johansson Helena, Simonsson Stina. Core transcription factors, oct4, sox2 and nanog, individually form complexes with nucleophosmin (npm1) to control embryonic stem (es) cell fate determination. Aging (Albany NY) 2010;2(11):815. doi: 10.18632/aging.100222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Lachowiez Curtis A., Loghavi Sanam, Kadia Tapan M., Daver Naval, Borthakur Gautam, Pemmaraju Naveen, Naqvi Kiran, Alvarado Yesid, Yilmaz Musa, Short Nicholas. Outcomes of older patients with npm1-mutated aml: current treatments and the promise of venetoclax-based regimens. Blood Adv. 2020;4(7):1311–1320. doi: 10.1182/bloodadvances.2019001267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zeng Yingchun, Ye Linbai, Zhu Shengli, Zheng Hong, Zhao Peng, Cai Weijia, Su Liya, She Yinglong, Wu Zhenghui. The nucleocapsid protein of sars-associated coronavirus inhibits b23 phosphorylation. Biochem Biophys Res Commun. 2008;369(2):287–291. doi: 10.1016/j.bbrc.2008.01.096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Sun Lei, Li Pan, Ju Xiaohui, Rao Jian, Huang Wenze, Zhang Shaojun, Xiong Tuanlin, Xu Kui, Zhou Xiaolin, Ren Lili, Ding Qiang, Wang Jianwei, Zhang Qiangfeng Cliff. In vivo structural characterization of the whole sars-cov-2 rna genome identifies host cell target proteins vulnerable to re-purposed drugs. bioRxiv. 2020 doi: 10.1016/j.cell.2021.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Levi Marcel. Covid-19 coagulopathy vs disseminated intravascular coagulation. Blood Adv. 2020;4(12):2850. doi: 10.1182/bloodadvances.2020002197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Kang Myung-Soo, Kieff Elliott. Epstein–barr virus latent genes. Experimental Mol Med. 2015;47(1):e131. doi: 10.1038/emm.2014.84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Uppal Timsy, Banerjee Sagarika, Sun Zhiguo, Verma Subhash C., Robertson Erle S. Kshv lana–the master regulator of kshv latency. Viruses. 2014;6(12):4961–4998. doi: 10.3390/v6124961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Giammarco Sabrina, Chiusolo Patrizia, Sica Simona, Rossi Monica, Minnella Gessica, Zini Gina. Sudden death of a sars-cov-2 patient with npm1+ acute myeloid leukemia mimicking acute promyelocytic leukemia. Int J Lab Hematol. 2021 doi: 10.1111/ijlh.13446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Beckelman Brenna C., Yang Wenzhong, Kasica Nicole P., Zimmermann Helena R., Xueyan Zhou C., Keene Dirk, Ryazanov Alexey G., Ma Tao. Genetic reduction of eef2 kinase alleviates pathophysiology in alzheimer’s disease model mice. J Clin Invest. 2019;129(2):820–833. doi: 10.1172/JCI122954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Jaafar Zane A, Kieft Jeffrey S. Viral rna structure-based strategies to manipulate translation. Nat Rev Microbiol. 2019;17(2):110–123. doi: 10.1038/s41579-018-0117-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.de la Fuente Alerie G., Queiroz Rayner M.L., Ghosh Tanay, McMurran Christopher E., Cubillos Juan F., Bergles Dwight E., Fitzgerald Denise C., Jones Clare A., Lilley Kathryn S., Glover Colin P. Changes in the oligodendrocyte progenitor cell proteome with ageing. Mol Cell Proteomics. 2020 doi: 10.1074/mcp.RA120.002102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Shan Jin-Lu, He Hai-Tao, Li Meng-Xia, Zhu Jian-Wu, Cheng Yi, Nan Hu., Wang Ge, Wang Dong, Yang Xue-Qin, He Yong. Ape1 promotes antioxidant capacity by regulating nrf-2 function through a redox-dependent mechanism. Free Radical Biol Med. 2015;78:11–22. doi: 10.1016/j.freeradbiomed.2014.10.007. [DOI] [PubMed] [Google Scholar]
- 85.Xiong Yong, Liu Yuan, Cao Liu, Wang Dehe, Guo Ming, Jiang Ao, Guo Dong, Hu Wenjia, Yang Jiayi, Tang Zhidong. Transcriptomic characteristics of bronchoalveolar lavage fluid and peripheral blood mononuclear cells in covid-19 patients. Emerg Microbes Infect. 2020;9(1):761–770. doi: 10.1080/22221751.2020.1747363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Rufini A., Tucci P., Celardo I., Melino G. Senescence and aging: the critical roles of p53. Oncogene. 2013;32(43):5129–5143. doi: 10.1038/onc.2012.640. [DOI] [PubMed] [Google Scholar]
- 87.Mancuso Peter, Bouchard Benjamin. The impact of aging on adipose function and adipokine synthesis. Front Endocrinol. 2019;10:137. doi: 10.3389/fendo.2019.00137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Blanco-Melo Daniel, Nilsson-Payant Benjamin E., Liu Wen-Chun, Uhl Skyler, Hoagland Daisy, Møller Rasmus, Jordan Tristan X., Oishi Kohei, Panis Maryline, Sachs David. Imbalanced host response to sars-cov-2 drives development of covid-19. Cell. 2020;181(5):1036–1045. doi: 10.1016/j.cell.2020.04.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Chow Ryan D., Majety Medha, Chen Sidi. The aging transcriptome and cellular landscape of the human lung in relation to sars-cov-2. Nat Commun. 2021;12(1):1–13. doi: 10.1038/s41467-020-20323-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Pearl Judea. Cambridge University Press; 2009. Causality. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.