Abstract
Background:
Prostate cancer is the second most common cancer in men in Iran. It can be treated in the early stages of the disease; therefore, early diagnosis can be lifesaving. The aim of this study was to investigate the molecular expression of some oncogenes and predisposing behaviors contributing to the aggressiveness of prostate cancer.
Methods:
In this case-control study, prostate cancer specimens were collected from both patients and healthy volunteers. Several factors such as age, family history, smoking, and stage of the disease, were investigated based on the criteria of this study. Real-time PCR was used to measure the expression of four oncogenes. Statistical analysis of our data was carried out using SPSS software version 22.
Results:
The X2 test showed that there was a difference in the incidence of prostate cancer in different age groups (X2= 9.30; p= 0.026). Although data analysis by the X2 test showed that family history had a significant effect on prostate cancer (X2= 14.43; p= 0.001), smoking did not show a significant effect on the incidence of this disorder (X2= 4.67; p= 0.097). The T2N1M0 stage is the most common form of prostate cancer in patients with family history of prostate cancer and the habit of smoking. Also, the expression of KRAS1P, GLB1L2, SChLAP1 and PACSIN3 oncogenes reduced in prostate cancer samples compared to the control group.
Conclusion:
Overall, functional interpretation of gene expression in the prostate tissue can affect tumor progression. Yet, further practical studies are required to reveal the accurate underlying mechanisms.
Key Words: Age, Family History, Oncogenes, Prostate Cancer, Smoking
Introduction
The study of cancer-related genes, which can be used as markers to predict the stages of a variety of cancers, and especially the aggressiveness of prostate cancer, is of clinical interest to many researchers (1, 2). Research has confirmed that prostate cancer is the second most common cancer and the fifth leading cause of cancer death in men (3, 4). According to the reports of the United States, one in six men suffers from prostate cancer during his lifetime, and more than 200,000 people are diagnosed with the disease each year. However, almost a small number (approximately 20%) of patients are engaged with a high-risk fatal form of prostate cancer (5). Prostate cancer depends on a variety of factors, including age, family history, and genetics; therefore, older men, those with a family history, as well as Africans, Americans, and Asians have a higher chance of developing the cancer (6, 7). Although prostate cancer is usually treatable in early stages, it is life-threatening and poses a therapeutic challenge in its invasive form (7). It is noteworthy that even after the initial treatment of prostate cancer, the cancer will relapse during the first 5 years due to various reasons in 20 to 30% of patients, which is a challenging topic (1, 8). Cancer screening is a vital issue that refers to searching for cancer before the appearance of the symptoms. Although prostate-specific antigen (PSA) detection is used as a routine screening tool for prostate cancer, the gene expression profiling also allows us to identify candidate oncogenes (9, 10). It is important to identify the molecular mechanisms underlying prostate cancer that can help us develop new methods of cancer detection and prevention. In the last two decades, significant efforts have been made to identify genes involved in cancer. Studies have shown that 384 genes that make up about 2% of human genome can be involved in cancer development (10, 11). However, it should be noted that prostate cancer can be predicted by genetic and epigenetic events that occur within the prostate cells. Despite the fact that the potential causes of prostate cancer cannot be attributed to changes in specific inherited genes, somatic mutations and changes in gene expression patterns can lead to prostate cancer (12). Although studies on the expression of prostate cancer related genes are still in their infancy, the impact of the expression of some genes, such as KRAS1P, GLB1L2, SChLAP1 and PACSIN3, on prostate cancer, has attracted the attention of researchers (1, 13). Studies have shown that altered expression of proto-oncogenic KRAS1P leads to the increased KRAS mRNA levels and accelerates cell growth, and this mechanism is involved in many cancers, such as bladder, colorectal, breast, lung, thyroid, and prostate cancers (13, 14). Also, galactosidase beta1-like 2 gene (GLB1L2), can be used as one of the best predictors of the aggressive forms of prostate cancer (1). Subsequently, using different molecular methods, changes in the expression of SChLAP1 (SWI/SNF Complex Antagonist Associated with Prostate Cancer1) independently contributed to the recurrence of prostate cancer, i.e. SPA recurrence was observed in patients with prostate cancer (15). PACSIN3 (Protein Kinase C and Casein Kinase Substrate In Neurons3) is another gene correlated with prostate cancer which is a member of the PACSIN family that regulates intracellular vesicle trafficking through its ability of regulating skeletal rearrangements (1, 16). According to above sentences and the importance of the mentioned genes, the aim of this study was to compare the expression of KRAS1P, GLB1L2, SChLAP1 and PACSIN3 genes in prostate cancer tissues with natural control tissues and to investigate various factors, such as patient age, tumor stage, smoking and family history as predisposing factors for prostate cancer.
Materials and Methods
Ethical declarations
In this case-control study, as samples were collected from patients with prostate cancer and healthy volunteers, written and informed consent was obtained from participants. The ethics committee of Iran University of Medical Sciences approved the research protocols and the accuracy of the subjects.
Study design and patients
In the present study, sample collection was performed over a period of 8 months (between February and December 2018). Forty patients were treated at Shahid Akbarabadi Hospital in Tehran, and prostate cancer was diagnosed in all patients using pathological biopsy examination. No age limit was considered in sample collecting, and all patients were investigated based on the criteria of this study, including family history, smoking habits, and stage of the disease using TNM staging system (Tumor, Nodule, and Metastasis). In addition, 41 healthy confirmed volunteers were tested as a control group, without any prostate enlargement, history of lower urinary tract symptoms, and family history of prostate cancer.
Sample preparation, RNA extraction, and cDNA synthesis
Initially, isolated prostate tissue samples from the subjects were immediately snap-frozen in liquid nitrogen and stored at -70 °C, and RNA extraction was then performed according to the manufacturer's instructions. At this stage, the Super RNA Extraction Kit for Tissue & Culture Cells (Favorgen Biotech Corp, Taiwan) was used to isolate total RNA from frozen tissues and cell lines. Also, the purity and integrity of extracted RNA of each sample were determined by optical density Ultrospec 2100 (Biochrom, USA) in 260/280 nm ratio. Also, total RNA (A260/280> 1.8) was treated with RNase-free DNase for which 1 μg was used for cDNA synthesis in a 20 μl reaction volume containing a reverse transcriptase enzyme, a primer, dNTPs and an RNase inhibitor (all from Promega, Madison, WI) at 37 °C for 1 h. The synthesized cDNA with optimal quality was diluted in a ratio of 1:10 and stored at -20 °C until use for subsequent experiments.
Primers and probes design
TaqMan probes and primers were used to measure the expression of four oncogenes, including KRAS1P, GLB1L2, SChLAP1 and PACSIN3. Primer Express V.3.0 (Applied Biosystems, USA) software was used to design the probes and primers, according to a study by Zhang and Colleagues in 2008 (17). Table 1 shows the sequences and detailed characteristics of the probes and primers used in this study. Also, 3' and 5' ends of all TaqMan probes were labeled with the quencher dye TAMRA and reporter dye molecule FAM, respectively. In addition, to prevent the probe extension in the PCR process, we phosphorylated the 3′ ends of each probe. Moreover, BLAST analysis was used to examine the specificity of the oligonucleotide sequences of TaqMan probes and primers (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Quantitative real-time PCR (qPCR)
The thermal-cycling conditions in a Rotor-Gene 6000 real-time PCR cycler (Qiagen Corbett, Hilden, Germany) were in two stage as follows: a first denaturation of 1 cycle for 5 min at 95 °C, followed by 40 cycles at 95 °C for 5 (annealing)and at 60 °C for 30s (extension). Each amplification reaction in a volume of 20 ml contained a mixture of 10 μL Probe 2× Taq (Probe qPCR) Master Mix (Takara Bio, Shiga, Japan), 0.4 ml of TaqMan probe, 0.4 ml of primer (each of forward and reverse), one ml of first-strand cDNA and 5.8 ml ddH2O. Also, non-template control (NTC) reactions were performed in all experiments and contained all the materials of the reaction mixture except the template cDNA. Final analysis of the results was performed on the mean values of three Real-time PCR tests on four oncogenes.
Statistical analysis
Initially, clinical and pathological data were collected from the subjects via questionnaires and after assessment of clinical information, data were entered into SPSS software version 22 (SPSS incorporate, Chicago) for analysis. Interpretation of the demographic results obtained from all subjects was based on frequency. Four different age groups were defined according to the observed quartiles as following: 1) age≤ 45, 2) 45< age≤ 54, 3) 54< age≤ 63, and 4) age> 63. After approving the normal distribution of all data using the Kolmogorov–Smirnov test, Mann-Whitney test was performed to evaluate between-group differences of the oncogenes. A chi-squared (X2) test was used to examine if age, smoking and family history affected the risk of cancer, and eta (η) correlation ratio was also determined to investigate the association between the oncogenes and the stage of prostate cancer. Moreover, the correlation between family history, smoking and the stage of prostate cancer, as well as the relationship between age and the risk of cancer were analyzed using Carmer's V test. The level of statistical significance was set at p≤ 0.05.
Results
In the present study, the age of patients and healthy individuals were within the range of 25 to 88 (48.70±15.32) and 23 to 89 (53.63±13.35) years old, respectively. The X2 test showed that there was a difference in the incidence of prostate cancer in different age groups (X2= 9.30; p= 0.026). The highest prevalence of prostate cancer was observed in the age group≤ 45 years and the lowest was found within the group 54< age≤ 63 years (figure 1). Among patients’ group, 22 (25.5%) subjects had a family history of prostate cancer (figure 2). The results also showed that 51% and 63% of subjects had smoking habits throughout their lifetime in the control group and the patient group, respectively (Figure 3). Although data analysis by the X2 test showed that family history had a significant effect on prostate cancer (X2= 14.43; p= 0.001), smoking did not show a significant effect with the incidence of prostate cancer (X2= 4.67; p= 0.097).
Fig. 1.
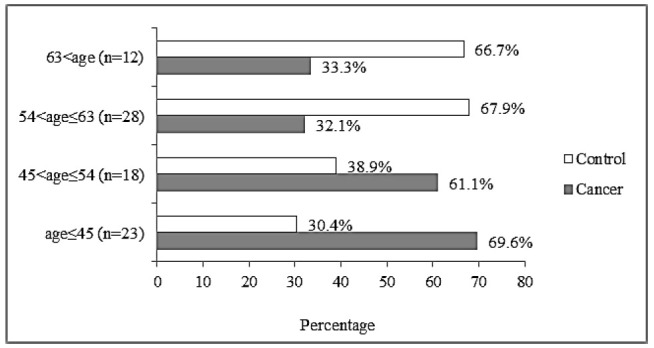
The prevalence of prostate cancer in four different age groups.
Fig. 2.
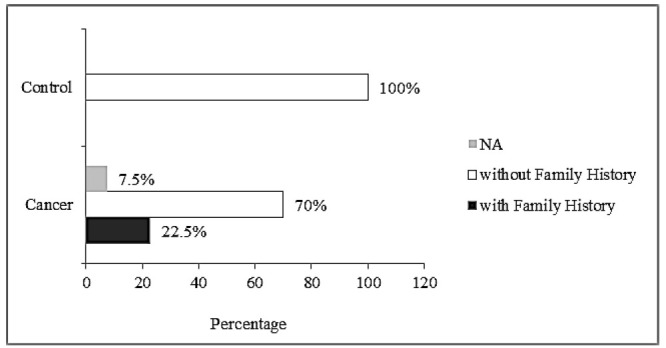
The rate of family history in the two studied groups; NA: not applicable.
Fig. 3.
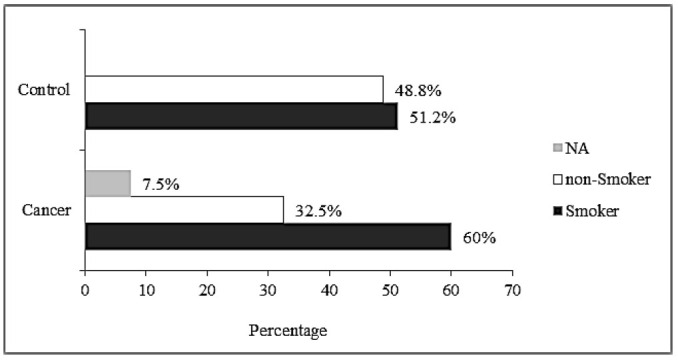
Rate of smokers in patients and the control group; NA: not applicable.
In this study, patients with prostate cancer were placed in different stages, as seen in Table 1. The T2N1M0 stage is the most common form of prostate cancer in patients with the highest rate of family history and smoking habit. In this study, analyzes exhibited that there was no significant relationship between different stages of the disease and smoking. In the group of patients, only 8 (20%) subjects had a family history and there was no significant relationship between family history and stage of the disease.
Table 1.
The frequency of different TNM staging system of prostate cancer and the corresponding relationship with family history and smoking.
| TNM staging system (n= 40) | Frequency (%) | Smoking habit | Family history | ||||
|---|---|---|---|---|---|---|---|
| Yes (%) | No (%) | NA (%) | Yes (%) | No (%) | NA (% | ||
| T1N0M0 | 1 (2.5) | 1 (2.5) | - | - | - | 1 (2.5) | - |
| T1N1M1 | 1 (2.5) | 1 (2.5) | - | - | 1 (2.5) | - | - |
| T2N1M0 | 19 (47.5) | 11 (27.5) | 7 (17.5) | 1 (2.5) | 3 (7.5) | 15 (37.5) | 1 (2.5) |
| T2N1M1 | 4 (10) | - | 4 (10) | - | - | 4 (10) | - |
| T2N2M1 | 7 (17.5) | 4 (10) | 3 (7.5) | - | 2 (5) | 4 (10) | 1 (2.5) |
| T3N1M1 | 2 (5) | - | 2 (5) | - | 2 (5) | - | - |
| T4N1M1 | 1 (2.5) | 1 (2.5) | - | - | - | 1 (2.5) | - |
| NA | 5 (12.5) | 2 (5) | 1 (2.5) | 2 (5) | 1 (2.5) | 3 (7.5) | 1 (2.5) |
T0: In these cases, there is no evidence of tumor in the prostate tissue.
T1: The tumor cannot be detected during a digital rectal exam (DRE) and cannot be seen using imaging tests.
T2: The tumor is large enough to be detected during DRE.
T3: The tumor has grown through the prostate on 1 side and into the tissue just outside the prostate.
T4: The tumor is fixed, or it is growing into adjacent structures other than the seminal vesicles.
N0: Prostate cancer has not yet extended to the regional lymph nodes.
N1: In these cases, the cancer has extended to the pelvic lymph nodes.
M0: Prostate cancer has not metastasized.
M1: Metastasis to distant tissues has occurred.
* TNM staging system was not applicable on five samples.
The results of qPCR and CT examination of four oncogenes in subjects showed that the expression of the studied oncogenes had reduced in prostate cancer samples compared to the control group (Figure 3). Statistical analysis showed that there was a moderate relationship between KRAS1P expression and the stage of the prostate cancer (Eta= 0.50), as well as a weak relationship between stage of the disease and the expression of GLB1L2 (Eta= 0.32), SChLAP1 (Eta= 0.40) and PACSIN3 (Eta= 0.19).
Fig. 4.
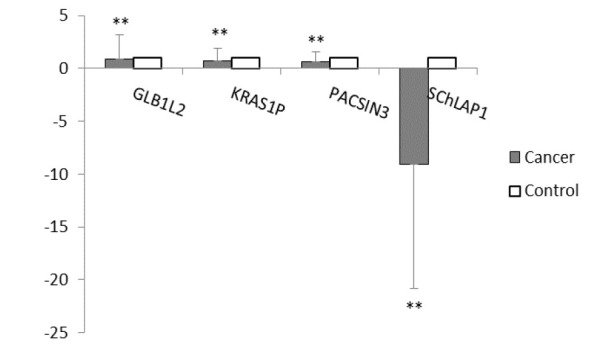
Comparison of the studied oncogenes expression based on fold change in patients and the control group.
Discussion
In 2018, the prevalence of prostate cancer in Iran was low and 9.11 per 100,000 cases were challenged with this disease; however, epidemiological studies show that prostate cancer is the sixth most common cancer among the Iranian population and the second most common cancer among Iranian men (18, 19). Prostate cancer as a chronic disease has led to widespread public health concerns as geographical and racial differences have made it more prevalent in Western countries than in Asian countries (20). We showed that different factors such as family history and age can affect the prostate cancer, and a significant relationship was observed between these factors and the disease. However, in our study, there was no significant relationship between smoking and prostate cancer. Although many etiological studies have been conducted in different parts of the world, the modifiable risk factors for prostate cancer are still unknown, especially in Iran. Although smoking has been shown to be involved in the development of tumors and complications, the results of studies on smoking and its effect on prostate cancer are contradictory (20). A cross-sectional study in Italy found no significant association between smoking and the disease in people with prostate cancer but in another study by Tang et al., a significant association was reported between smoking and prostate cancer (21, 22). Studies on the relationship between age and prostate cancer have shown that there is a significant relationship between age and the incidence of this cancer, however, the average age of patients with prostate cancer varies from study to study. Different factors, such as changes in diet and lifestyle, enhanced life expectancy, more access to screening and diagnosis of prostate cancer, occupational risk factors, the improvement in cancer registry completeness, and more exposure to the environment, can affect patients at different ages (20, 23, 24). In this study, only 20% of patients had a family history of prostate cancer. Although family history has been shown to be a major predictor of prostate cancer risk, a small proportion of prostate cancer cases can be attributed to the inheritance of cancer-prone genes (25). However, several studies (including ours) have reported a link between family history and prostate cancer in patients (9, 26).
The high mortality rate of prostate cancer in men and the lack of effective treatment options in the advanced forms of this cancer have doubled the importance of early detection of the disease (27). Hence, a lot of research has been done to discover specific and sensitive markers for accurate and timely identification of the cases. Changes in gene expression can pave the way for the diagnosis of this disease (28, 29). Therefore, many studies should be done to determine the gene expression pattern in different stages of prostate cancer to this end. Accordingly, in this study, changes in the expression of oncogenes as new molecular markers for prognostic factors of prostate cancer were investigated. Altered expression of KRAS1P, GLB1L2, SChLAP1 and PACSIN3 oncogenes in patients with prostate cancer was observed in our study compared to the control group. In one study, Rose et al. reported decreased expression of the GLB1L2 gene in cancer. In this study, it was shown that reduced GLB1L2 expression decreased the expression of a tumor-suppressing protein called ITIH5 (Inter-Alpha-Trypsin Inhibitor Heavy Chain Family Member 5), which is involved in tissue cancer; however, its molecular mechanisms of action are still unclear (300). The association of GLB1L2 with prostate cancer has also been identified in a study by Jhun et al., who showed the reduced expression of this gene in samples isolated from prostate cancer patients (1). Today, another gene that has been considered in the detection of cancers (prostate cancer, hepatocellular, neuroblastoma, and retinoblastoma carcinomas) is the KRAS1P gene, whose expression has been studied by several research groups (including ours) in cancers (31). Another study found an inverse relationship of expression between miR-143 and KRAS protein in prostate cancer. The results showed that suppression of KRAS and inactivation of the MAPK pathway can be associated with prostate cancer (32). In addition, altered expression of SChLAP1 has been implicated in cancer initiation. Mehra et al. reported that changes in the expression of SChLAP1 gene could be a promising marker in predicting prostate cancer in American men and in identifying at-risk patients (15). Another study also found that the expression of SChLAP1 changes with prostate cancer progression, and according to our study, SChLAP1 could be considered as a biomarker for the diagnosis of prostate cancer (33). As shown, PACSIN3 is one of the genes that is down-regulated in gastric cancer and serves as a good marker for carcinogenesis and subsequent progression (34). Similar to the results of the present study, another study also showed that the decreased expression of PACSIN3 gene was associated with prostate cancer outcomes in a radical prostatectomy group (1). The results of the above studies suggest KRAS1P, GLB1L2, SChLAP1 and PACSIN3 oncogenes as potential markers for the diagnosis of prostate cancer. These genes can be used to detect prostate cancer in combination with other biomarkers.
As prostate cancer is an increasing challenge among Iranian men, it is vital to propose new methods to diagnose patients more quickly and accurately. In the current study, we measured the expression of KRAS1P, GLB1L2, SChLAP1 and PACSIN3 oncogenes in clinical prostate samples to confirm the relationship between the expression level of these genes and some predisposing factors. Our results verified the data from the previous research regarding the down-regulation of KRAS1P, GLB1L2, SChLAP1 and PACSIN3 oncogenes in prostate cancer. On the whole, these functional interpretations of gene expression in the prostate tissue can affect the progression of tumor. Yet, further practical studies are required to reveal the accurate underlying mechanisms.
Acknowledgements
We are grateful to Islamic Azad University for general support. The authors declare no conflict of interest.
References
- 1.Jhun MA, Geybels MS, Wright JL, Kolb S, April C, Bibikova M, et al. Gene expression signature of Gleason score is associated with prostate cancer outcomes in a radical prostatectomy cohort. . Oncotarget. . 2017;8(26):43035, 43047. doi: 10.18632/oncotarget.17428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Jonsson M, Ragnum HB, Julin CH, Yeramian A, Clancy T, Frikstad K-AM, et al. Hypoxia-independent gene expression signature associated with radiosensitisation of prostate cancer cell lines by histone deacetylase inhibition. . Br J Cancer. 2016;115(8):929, 939. doi: 10.1038/bjc.2016.278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kazantseva M, Mehta S, Eiholzer RA, Gimenez G, Bowie S, Campbell H, et al. The Δ133p53β isoform promotes an immunosuppressive environment leading to aggressive prostate cancer. . Cell Death Dis. . 2019;10(9):631. doi: 10.1038/s41419-019-1861-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Caraceni A, Zecca E, Formaglio F, Ricchini F. Bone Metastases from Prostate Cancer: From Symptom Control to Pain Palliation. Bone Metastases from Prostate Cancer. 2017:251–270. [Google Scholar]
- 5.Prensner JR, Iyer MK, Sahu A, Asangani IA, Cao Q, Patel L, et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. . Nat Genet. 2013;45(11):1392–8. doi: 10.1038/ng.2771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ito K. Prostate cancer in Asian men. . Nature Reviews Urology. 2014;11(4):197–212. doi: 10.1038/nrurol.2014.42. [DOI] [PubMed] [Google Scholar]
- 7.Joniau S, Briganti A, Gontero P, Gandaglia G, Tosco L, Fieuws S, et al. Stratification of high-risk prostate cancer into prognostic categories: a European multi-institutional study. . Eur Urol. . 2015;67(1):157–164. doi: 10.1016/j.eururo.2014.01.020. [DOI] [PubMed] [Google Scholar]
- 8.Han M, Partin AW, Zahurak M, Piantadosi S, Epstein JI, Walsh PC. Biochemical (prostate specific antigen) recurrence probability following radical prostatectomy for clinically localized prostate cancer. . J Urol. . 2003;169(2):517–23. doi: 10.1097/01.ju.0000045749.90353.c7. [DOI] [PubMed] [Google Scholar]
- 9.Hosseinzadeh O, Hekmat Z, Nekoufar S, Ahmad M, Mohammadzadeh N, Monfaredan A. Evaluate the gene expression of TPT1, EDN3, and ANO7 in prostate cancer tissues and their relation with age, tumor stage and family history. Meta Gene. 2020;24:100671. [Google Scholar]
- 10.Santarius T, Shipley J, Brewer D, Stratton MR, Cooper CS. A census of amplified and overexpressed human cancer genes. . Nature Reviews Cancer. . 2010;10(1):59–64. doi: 10.1038/nrc2771. [DOI] [PubMed] [Google Scholar]
- 11.Quinn DI, Henshall SM, Sutherland RL. Molecular markers of prostate cancer outcome. . European journal of cancer. 2005;41(6):858–887. doi: 10.1016/j.ejca.2004.12.035. [DOI] [PubMed] [Google Scholar]
- 12.Fredriksson NJ, Ny L, Nilsson JA, Larsson E. Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. . Nat Genet. . 2014;46(12):1258–63. doi: 10.1038/ng.3141. [DOI] [PubMed] [Google Scholar]
- 13.Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP, et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. . Nature. 2010;465(7301):1033–1038. doi: 10.1038/nature09144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Silan F, Gultekin Y, Atik S, Kilinc D, Alan C, Yildiz F, et al. Combined point mutations in codon 12 and 13 of KRAS oncogene in prostate carcinomas. Mol Biol Rep. . 2012;39(2):1595–9. doi: 10.1007/s11033-011-0898-8. [DOI] [PubMed] [Google Scholar]
- 15.Mehra R, Udager AM, Ahearn TU, Cao X, Feng FY, Loda M, et al. Overexpression of the long non-coding RNA SChLAP1 independently predicts lethal prostate cancer. . Eur Urol. 2016;70(4):549–552. doi: 10.1016/j.eururo.2015.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Roach W, Plomann M. PACSIN3 overexpression increases adipocyte glucose transport through GLUT1. . Biochem Biophys Res Commun. 2007;355(3):745–50. doi: 10.1016/j.bbrc.2007.02.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang L, Wang C-Y, Yang R, Shi J, Fu R, Chen L, et al. Real-time quantitative RT-PCR assay of prostate-specific antigen and prostate-specific membrane antigen in peripheral blood for detection of prostate cancer micrometastasis. . Urologic Oncology: Seminars and Original Investigations. . 2008:634–640. doi: 10.1016/j.urolonc.2007.07.016. [DOI] [PubMed] [Google Scholar]
- 18.Hassanipour S, Fathalipour M, Salehiniya H. The incidence of prostate cancer in Iran: a systematic review and meta-analysis. Prostate Int. 2018;6(2):41–45. doi: 10.1016/j.prnil.2017.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mousavi SM, Gouya MM, Ramazani R, Davanlou M, Hajsadeghi N, Seddighi Z, editors. Cancer incidence and mortality in Iran. . Annals of oncology. . 2009;20(3):556–563. doi: 10.1093/annonc/mdn642. [DOI] [PubMed] [Google Scholar]
- 20.Sadeghi-Gandomani H, Yousefi M, Rahimi S, Yousefi S, Karimi-Rozveh A, Hosseini S, et al. The incidence, risk factors, and knowledge about the prostate cancer through worldwide and Iran. World Cancer Research Journal. . 2017;4(4):e972. [Google Scholar]
- 21.Tang B, Han CT, Gan HL, Zhang GM, Zhang CZ, Yang WY, et al. Smoking increased the risk of prostate cancer with grade group≥ 4 and intraductal carcinoma in a prospective biopsy cohort. . Prostate. 2017;77(9):984–989. doi: 10.1002/pros.23354. [DOI] [PubMed] [Google Scholar]
- 22.De Nunzio C, Tema G, Lombardo R, Trucchi A, Bellangino M, Esperto F, et al. Cigarette smoking is not associated with prostate cancer diagnosis and aggressiveness: a cross sectional Italian study. . Minerva Urol Nefrol. . 2018;70(6):598–605. doi: 10.23736/S0393-2249.18.03182-X. [DOI] [PubMed] [Google Scholar]
- 23.Putra IBO, Hamid AR, Mochtar CA, Umbas R. Relationship of age, prostate-specific antigen, and prostate volume in Indonesian men with benign prostatic hyperplasia. . Prostate Int. . 2016;4(2):43–8. doi: 10.1016/j.prnil.2016.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kurian CJ, Leader AE, Thong MS, Keith SW, Zeigler-Johnson CM. Examining relationships between age at diagnosis and health-related quality of life outcomes in prostate cancer survivors. BMC Public Health. . 2018;18(1):1060. doi: 10.1186/s12889-018-5976-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Taheri M, Habibi M, Noroozi R, Rakhshan A, Sarrafzadeh S, Sayad A, et al. HOTAIR genetic variants are associated with prostate cancer and benign prostate hyperplasia in an Iranian population. Gene. 2017;613:20–24. doi: 10.1016/j.gene.2017.02.031. [DOI] [PubMed] [Google Scholar]
- 26.Barber L, Gerke T, Markt SC, Peisch SF, Wilson KM, Ahearn T, et al. Family history of breast or prostate cancer and prostate cancer risk. . Clin Cancer Res. . 2018;24(23):5910–5917. doi: 10.1158/1078-0432.CCR-18-0370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. . CA Cancer J Clin. . 2011;61(2):69–90. doi: 10.3322/caac.20107. [DOI] [PubMed] [Google Scholar]
- 28.Nodouzi V, Nowroozi M, Hashemi M, Javadi G, Mahdian R. Concurrent down-regulation of PTEN and NKX3. 1 expression in Iranian patients with prostate cancer. . Int Braz J Urol. 2015;41(5):898, 905. doi: 10.1590/S1677-5538.IBJU.2014.0036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Luo JH, Yu YP, Zhargy Y, Cieply K, Lin F, Deflavia P, Dhir R, et al. Gene expression analysis of prostate cancers. . Mol Carcinog. 2002;33(1):25, 35. doi: 10.1002/mc.10018. [DOI] [PubMed] [Google Scholar]
- 30.Rose M, Kloten V, Noetzel E, Gola L, Ehling J, Heide T, et al. ITIH5 mediates epigenetic reprogramming of breast cancer cells. . Mol Cancer. . 2017;16(1):44. doi: 10.1186/s12943-017-0610-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Su X, Xing J, Wang Z, Chen L, Cui M, Jiang B. microRNAs and ceRNAs: RNA networks in pathogenesis of cancer. Chin J Cancer Res. . 2013;25(2):235, 239. doi: 10.3978/j.issn.1000-9604.2013.03.08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Xu B, Niu X, Zhang X, Tao J, Wu D, Wang Z, et al. miR-143 decreases prostate cancer cells proliferation and migration and enhances their sensitivity to docetaxel through suppression of KRAS. Mol Cell Biochem. 2011;350(1-2):207, 13. doi: 10.1007/s11010-010-0700-6. [DOI] [PubMed] [Google Scholar]
- 33.Chua ML, Lo W, Pintilie M, Murgic J, Lalonde E, Bhandari V, et al. A prostate cancer “nimbosus”: genomic instability and SChLAP1 dysregulation underpin aggression of intraductal and cribriform subpathologies. . Eur Urol. . 2017;72(5):665, 674. doi: 10.1016/j.eururo.2017.04.034. [DOI] [PubMed] [Google Scholar]
- 34.Gou Wf, Shen Df, Yang Xf, Zhao S, Liu Yp, Sun Hz, et al. ING5 suppresses proliferation, apoptosis, migration and invasion, and induces autophagy and differentiation of gastric cancer cells: a good marker for carcinogenesis and subsequent progression. Oncotarget. . 2015;6(23):19552, 79. doi: 10.18632/oncotarget.3735. [DOI] [PMC free article] [PubMed] [Google Scholar]


