Abstract
Background: Coinfection with bacteria, fungi, and respiratory viruses in SARS-CoV-2 is of particular importance due to the possibility of increased morbidity and mortality. In this meta-analysis, we calculated the prevalence of such coinfections. Methods: Electronic databases were searched from 1 December 2019 to 31 March 2021. Effect sizes of prevalence were pooled with 95% confidence intervals (CIs). To minimize heterogeneity, we performed sub-group analyses. Results: Of the 6189 papers that were identified, 72 articles were included in the systematic review (40 case series and 32 cohort studies) and 68 articles (38 case series and 30 cohort studies) were included in the meta-analysis. Of the 31,953 SARS-CoV-2 patients included in the meta-analysis, the overall pooled proportion who had a laboratory-confirmed bacterial infection was 15.9% (95% CI 13.6–18.2, n = 1940, 49 studies, I2 = 99%, p < 0.00001), while 3.7% (95% CI 2.6–4.8, n = 177, 16 studies, I2 = 93%, p < 0.00001) had fungal infections and 6.6% (95% CI 5.5–7.6, n = 737, 44 studies, I2 = 96%, p < 0.00001) had other respiratory viruses. SARS-CoV-2 patients in the ICU had higher co-infections compared to ICU and non-ICU patients as follows: bacterial (22.2%, 95% CI 16.1–28.4, I2 = 88% versus 14.8%, 95% CI 12.4–17.3, I2 = 99%), and fungal (9.6%, 95% CI 6.8–12.4, I2 = 74% versus 2.7%, 95% CI 0.0–3.8, I2 = 95%); however, there was an identical other respiratory viral co-infection proportion between all SARS-CoV-2 patients [(ICU and non-ICU) and the ICU only] (6.6%, 95% CI 0.0–11.3, I2 = 58% versus 6.6%, 95% CI 5.5–7.7, I2 = 96%). Funnel plots for possible publication bias for the pooled effect sizes of the prevalence of coinfections was asymmetrical on visual inspection, and Egger’s tests confirmed asymmetry (p values < 0.05). Conclusion: Bacterial co-infection is relatively high in hospitalized patients with SARS-CoV-2, with little evidence of S. aureus playing a major role. Knowledge of the prevalence and type of co-infections in SARS-CoV-2 patients may have diagnostic and management implications.
Keywords: SARS-Cov-2, co-infection, coinfection, COVID-19, concurrent, bacterial, fungal, viral, meta-analysis
1. Introduction
Coronavirus disease 2019 (COVID-19) is caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and was first described in Wuhan, China in 2019. Globally, as of 15 April 2021, there have been 137,866,311 confirmed cases of COVID-19, including 2,965,707 deaths, as reported by the World Health Organization [1]. Coinfection with SARS-CoV-2 and other bacterial, fungal, and respiratory viral pathogens [2,3,4], Gram-positive and Gram-negative bacteria [5,6,7], Middle East respiratory syndrome coronavirus (MERS-CoV) [8], and influenza [9,10,11,12,13] has been described. However, the reported frequency is variable. Such coinfections in patients with SARS-CoV-2 may be a cause of increased morbidity and mortality [2,6,7,14,15,16,17,18,19,20,21,22]. Thus, timely diagnosis is important to initiate appropriate therapy and limit the overuse of antimicrobial agents. Previous studies, including case series [2,5,8,11,14,15,16,19,20,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50], cohort studies [3,4,6,7,9,10,12,13,17,18,21,22,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70], and meta-analyses [71,72,73], have shown variable results. In light of recent studies evaluating coinfections in SARS-CoV-2 patients, we aimed to re-evaluate the prevalence of bacterial, fungal, and respiratory viral coinfections in a comprehensive meta-analysis. Moreover, we aimed to identify the risk-factors, characteristics, and consequences associated with SARS-CoV-2 coinfection.
2. Methods
2.1. Design
This is a meta-analysis and was conducted per the Preferred Reporting Items for Systematic Reviews and Meta-Analyses [PRISMA] guidelines [74]. We searched PROQUEST, MEDLINE, EMBASE, PUBMED, CINAHL, WILEY ONLINE LIBRARY, and NATURE for full texts. Search keywords included Coronavirus infection OR SARS coronavirus OR severe acute respiratory syndrome OR COVID OR SARS AND mixed infection OR bacterial pneumonia OR bacteremia OR bacterial infection OR fungal infection OR viral infection OR respiratory infection OR mycosis OR coinfect OR co-infect OR concomitant infect OR concurrent infection OR mixed infect OR coinfection OR co-infection. The search included English language studies from 1 December 2019 to 31 March 2021. Then, articles were kept if the title and abstract contained discussion about bacterial, fungal, and/or respiratory viral co-infection in SARS-CoV-2 patients. In addition, we used manual backward snowballing of the bibliographies of retrieved articles to include additional relevant articles.
2.2. Inclusion and Exclusion Criteria
The included articles were pertinent if these articles included patients with a positive SARS-CoV-2 reverse-transcription polymerase chain reaction (RT-PCR) test of any age and a described co-infection on presentation or developed during the course of the disease or during hospital stay. These cases were retained if bacteria, fungi, and/or viruses were detected in the respiratory tract or blood culture samples and were excluded if they were identified from other samples. We aimed to include randomized controlled trials, cohort studies, and case series, and excluded other types of studies.
2.3. Data Extraction
Three authors (S.A., A.A., and J.A.) reviewed the retrieved studies and chose relevant articles. Data were extracted using key headings as indicated in Table 1. The study designs were classified as well. The extracted information included: authors; study location; study design and setting; publication year; number of SARS-CoV-2 patients tested for co-pathogens; number of coinfected patients; age; proportion of male patients; percentage of patients requiring intensive care unit (ICU) and mechanical ventilation; mortality rates; proportion of patients with bacterial, fungal, and/or respiratory viral coinfections; total organisms identified; antimicrobials prescribed; laboratory techniques for co-pathogen detection; assessment of study risk of bias; and remarks on notable findings.
Table 1.
Summary of the characteristics of the included studies with evidence on SARS-CoV-2 and bacterial, fungal, and/or respiratory viral co-infections (n = 72), 2020–2021.
| Author, Year, Study Location | Study Design, Setting | Number of SARS-CoV-2 Patients Tested for Co-Pathogens, n | Co-Infected Patients, n (%) | Age (Years) | Male, n (%) | Admitted to ICU, n (%) | Mechanical Ventilation, n (%) | Deaths, n (%) | Bacterial Co-Infection, n (%) | Fungal Co-Infection, n (%) | Respiratory Viral Co-Infection, n (%) | Total Organisms, n | Antimicrobials Use, n | Laboratory Techniques for Co-Pathogen Detection | NOS Score | Key Findings |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Alanio et al., 2020 [23], France | Prospective case series, single center | 27 | 7 (25.9) | Median (IQR), 63 (43–79) | 5 (71.4) | 7 (100) | 7 (100) | 4 (75.1) | - | 7 (25.9) | - | 7 Aspergillus fumigatus | 3 Macrolides 2 Antifungals |
Culture from respiratory specimens and GM detection in the BAL and serum | 7 | Death was not related to pulmonary aspergillosis but to bacterial septic shock and organ failure. |
| Allou et al., 2021 [9], France | Prospective cohort, single center | 36 | 5 (13.9) | Median (IQR), 68 (57–82) | 4 (80) | 10 (27.8) | 2 (5.5) | 0 | 2 (5.5) | - | 3 (8.3) | 1 Influenza A virus 1 Branhamella catarrhalis 1 S. pneumoniae 1 H. influenzae 1 Human Coronavirus 229E 1 Rhinovirus 1 MSSA |
Not reported | RT-PCR for naopharyngeal specimens [viruses] AND sputum culture [bacteria and fungi] | 7 | Level of D-dimer was significantly higher in patients with co-infection compared to patients without co-infection (1.36 mg/mL vs. 0.63 mg/mL, p = 0.05). |
| Amin et al., 2021 [14], United States | Retrospective case series, single center | 140 | 79 (56.4) | Mean (SD), 62.3 (16.3) | 55 (69.6) | 29 (36.7) | 26 (32.9) | 38 (48.1) | 79 (56.4) | - | - | 79 M. pneumoniae | All patients received antibiotics coverage against M. pneumoniae, however, agents were not reported | Serum antibody test (IgM) | 6 | Death was significantly higher in patients with M. pneumoniae co-infection compared to patients without M. pneumoniae co-infection (AOR: 2.28, 95% CI: 1.03–5.03). |
| Anton-Vazquez et al., 2021 [24], Spain | Retrospective case series, single center | 917 | 87 (9.5) | Median (IQR), 68 (27–92) | 37 (42.5) | 8 (9.2) | Not reported | 15 (17.2) | 87 (9.5) | - | - | 87 S. pneumoniae | Third Generation Cephalosporins were prescribed in the great majority of cases | Serum antibody test (IgM, IgG) | 6 | Co-infected pneumococcal pneumonia patients compared with COVID-19 patients without pneumococcal testing were mostly female (57% vs. 34%, p < 0.001). No differences in age, length of stay, admission to ICU, or mortality were found between groups. |
| Arentz et al., 2020 [15], United States | Retrospective case series, single center | 21 | 4 (19) | Mean (range), 70 (43–92) | 11 (52) | 21 (100) | 15 (71) | 11 (52.4) | 1 (4.8) | - | 3 (14.3) | 1 Pseudomonas 2 Influenza A virus 1 Parainfluenza 3 virus |
Not reported | Unspecified | 8 | Study included 21 ICU patients who had a high rate of ARDS and a high risk of death. |
| Bardi et al., 2021 [2], United States | Retrospective case series, single center | 140 | 57 (40.7) | Median (IQR), 63 (60–68) | 47 (82) | 57 (100) | 56 (98) | 31 (54) | 51 (36.4) | 6 (4.3) | - | 18 Enterococcus faecium 11 Enterococcus faecalis 16 CoNS 14 P. aeruginosa 9 MRSA1 Klebsiella oxytoca 1 Serratia marcescens 1 Bacteroides spp. 1 Candida glabrata 4 Candida albicans 3 Aspergillus fumigatus 3 Stenotrophomonas maltophilia 2 A. baumannii 2 Enterobacter cloacae 1 Aspergillus terreus 1 Hafnia alvei 1 H. influenzae 1 MSSA 1 K. pneumoniae |
53 Third Generation Cephalosporins 53 Macrolides 47 Other antibiotics |
Respiratory tracheal aspirate and blood cultures | 6 | Co-infection occurred a median of 9 days (IQR 5–11) after admission and was significantly associated with the APACHE II score (p = 0.02). Co-infection was significantly associated with death (OR 2.7, 95% CI 1.2–5.9, p = 0.015) and longer ICU stay (p < 0.001). |
| Barrasa et al., 2020 [16], Spain | Retrospective case series, multi-center | 48 | 6 (12.5) | Median (IQR), 63 (51–75) | 27 (56.2) | 48 (100) | 45 (93.7) | 12 (25) | 5 (10.4) | - | 1 (2.1) | 3 P. aeruginosa 1 Enterococcus faecium 1 H. influenzae 1 MRSA |
17 Fluoroquinolones 22 Third Generation Cephalosporins 10 Macrolides 9 Linezolid 15 Beta-Lactams |
Unspecified | 7 | Procalcitonin plasma above 0.5 mg/L was associated with 16% vs. 19% (p = 0.78) risk of death after 7 days. |
| Bartoletti et al., 2020 [17], Italy | Prospective cohort, multi-center |
108 | 30 (27.7) | Median (IQR), 63 (57–70) | 24 (80) | 108 (100) | 108 (100) | 44 (40.7) | - | 19 (17.6) | - | 15 Aspergillus fumigatus 3 Aspergillus niger 1 Aspergillus flavus |
9 Macrolides 16 Antifungals |
Culture from respiratory specimens and GM detection in the BAL and serum | 7 | Co-infection of aspergillosis occurred after a median of 4 (2–8) days from ICU admission and a median of 14 (11–22) days from SARS-CoV-2 symptom onset. Mortality was higher in ICU patients co-infected with aspergillosis compared to SARS-CoV-2 patients without the fungal co-infection (44% vs. 19%, p = 0.002). |
| Calcagno et al., 2021 [5], Italy | Retrospective case series, single center | 56 | 10 (17.8) | Mean (SD), 63.3 (18) | 6 (60) | Not reported | Not reported | Not reported | 10 (17.8) | - | - | 7 S. aureus 2 H. influenzae 1 E. coli 1 M. catarrhalis 1 Streptococci agalactiae 1 K. pneumoniae 1 Enterobacter cloacae |
Not reported | RT-PCR of respiratory tract specimens (nasopharyngeal, BAL, BA, and sputum) | 7 | Phenomena like viral interference, common receptor usage, different inoculum size, or simply resource competition might explain why dual or multiple concurrent viral respiratory infections are rare. |
| Chen N et al., 2020 [80], China | Retrospective case series, single center | 99 | 5 (5) | Mean (SD), 55.5 (13.1) | 67 (67.7) | 23 (23) | 17 (17) | 11 (11) | 1 (1) | 4 (4) | - | 1 A. baumannii 1 K. pneumoniae 1 Aspergillus flavus 1 Candida glabrata 3 Candida albicans |
70 [cephalosporins, quinolones, carbapenems, tigecycline, and linezolid] 15 Antifungals | RT-PCR via throat swab | 7 | Six (6%) of patients had high procalcitonin levels. |
| Chen T et al., 2020 [25], China | Retrospective case series, single center | 203 | 17 (8.4) | Median (IQR), 54 (20–91) | 108 (53.2) | 34 (16.7) | 39 (19.2) | 26 (12.8) | 2 (0.9) | - | 15 (7.4) | 4 Parainfluenza virus 3 RSV 3 Adenovirus 2 Mycoplasma 2 Influenza A virus 3 Influenza B virus |
Not reported | Unspecified | 7 | Two mortality cases were reported in co-infected patients. |
| Cheng L et al., 2020 [18], Hong Kong | Prospective cohort, single center | 147 | 12 (8.2) | Median (IQR), 49 (30–61) | 9 (75) | 1 (8.3) | Not reported | 0 | 12 (8.2) | - | - | 3 H. influenzae 8 MSSA 1 P. aeruginosa 1 S. pneumoniae |
46 Penicillins & cephalosporins 14 Tetracyclines 3 Fluoroquinolones 3 Macrolides |
RT-PCR of respiratory tract specimens AND sputum and blood cultures | 6 | Co-infected SARS-CoV-2 patients had longer length of hospitalization (median: 20 days vs. 27 days, p = 0.016). |
| Cheng Y et al., 2021 [10], China | Prospective cohort, single center | 213 | 97 (45.5) | Median (IQR), 61 (50–68) | 47 (48.5) | Not reported | 2 (2.1) | 3 (3.1) | - | - | 97 (45.5) | 97 Influenza A virus | Not reported | Serum antibody test (IgM) | 6 | Similar symptoms and clinical outcomes were seen in the SARS-CoV-2 co-infected group compared to the SARS-CoV-2 group without co-infection. Co-infection with Influenza A virus had no effect on disease outcome. |
| Contou et al., 2020 [51], France | Prospective cohort, single center | 92 | 26 (28) | Median (IQR), 61 (55–70) | 73 (79) | 92 (100) | 83 (90) | 45 (49) | 26 (28) | - | - | 10 MSSA 7 H. influenzae 6 S. pneumoniae 5 Enterobacteriaceae 2 P. aeruginosa 1 M. catarrhalis 1 A. baumannii |
14 Third Generation Cephalosporins 14 Beta-Lactam/Beta-Lactamase Inhibitors 6 Beta-Lactams 5 Others antibiotics |
RT-PCR for respiratory specimens [viruses] AND respiratory and blood cultures [bacteria and fungi] | 7 | Resistance by co-pathogens to 3rd generation cephalosporin and to amoxicillin–clavulanate combination was observed in 8% and 21%, respectively. |
| Cuadrado-Payán et al. [11]. 2020, Spain | Retrospective case series, single center | 4 | 4 (100) | Mean (SD), 67 (14.5) | 3 (75) | 3 (75) | 3 (75) | 0 | - | - | 4 (100) | 3 Influenza A virus 2 Influenza B virus |
None | RT-PCR for respiratory specimens | 7 | Clinical courses in co-infected SARS-CoV-2 patients did not differ from those previously reported. |
| De Francesco et al., 2021 [6], Italy | Retrospective cohort, multi-center | 443 | 242 (54.6) | Mean (SD), 71 (19) | 173 (71.4) | Not reported | 16 (6.8) | Not reported | 242 (54.6) | - | - | 242 C. pneumoniae 63 M. pneumoniae |
138 Macrolides | Serum antibody test (IgM, IgG) | 6 | SARS-CoV-2 co-infected patients were more critical than SARS-CoV-2 patients without co-infection (13.2% vs. 5.9%, p = 0.01). Need for ventilatory support was significantly higher in co-infected patients than in only SARS-CoV-2 positive patients (nasal canula: 18.1% vs. 3.6%, p < 0.0001; high flow oxygen support: 45% vs. 23.3%, p < 0.0001; and non-invasive ventilation: 14.7% vs. 4.6%, p = 0.001, respectively). Higher mortality was observed in SARS-CoV-2 patients with M. and/or C. pneumoniae (24.2% vs. 21.8%, p = 0.63). |
| Ding et al., 2020 [19], China | Retrospective case series, single center | 115 | 5 (4.3) | Mean (SD), 50.20 (9.83) | 2 (40) | 0 | 0 | 0 | - | - | 5 (4.3) | 3 Influenza A virus 2 Influenza B virus |
Five patients received antibiotics; however, agents were not reported. | Influenza serology | 7 | SARS-CoV-2 co-infected patients did not show severe disease compared to SARS-CoV-2 without influenza co-infection (similar laboratory results, imaging, and prognosis). Nasal blockade and pharyngeal pain were more in the SARS-CoV-2 con-infected group. |
| Elhazmi et al., 2021 [8], Saudi Arabia | Retrospective case series, multi-center | 67 | 8 (11.9) | Mean (SD), 44.4 (11.8) | 6 (75) | 67 (100) | 7 (87.5) | 3 (37.5) | - | - | 8 (11.9) | 8 MERS-CoV | None | RT-PCR for respiratory specimens | 7 | Seven (87.5%) patients were obese. |
| Garcia-Vidal et al., 2021 [7], Spain | Retrospective cohort, single center | 989 | 31 (3.1) | Median (IQR), 63 (54.5–74) | 18 (58.1) | 8 (25.8) | Not reported | 5 (16.1) | 25 (2.5) | - | 7 (0.7) | 12 S. pneumoniae 7 S. aureus 2 H. influenzae 1 M. catarrhalis 2 P. aeruginosa 1 E. coli 1 K. pneumoniae 1 Enterococcus faecium 1 Proteus mirabilis 1 Citrobacter koseri 6 Influenza A virus 3 Influenza B virus 1 RSV 1 HSV |
26 Macrolides 24 Third Generation Cephalosporins 2 Fifth Generation Cephalosporins |
RT-PCR for respiratory specimens [viruses] AND blood, pleural fluids, sputum cultures [bacteria and fungi] | 7 | Co-infection at COVID-19 diagnosis is uncommon. Worse clinical outcomes were seen in SARS-CoV-2 co-infected patients. |
| Gayam et al., 2020 [52], United States | Retrospective cohort, single center | 350 | 6 (1.7) | Mean (SD), 57 (10.6) | 2 (33.3) | 1 (16.7) | 1 (16.7) | 1 (16.7) | 6 (1.7) | - | - | 6 M. pneumoniae | 6 Third Generation Cephalosporins 3 Macrolides 3 Tetracyclines |
Serum antibody test (IgM, IgG) | 6 | Only one patient (16.7%) required ICU admission and experienced organ failure and death. |
| Hashemi et al., 2021 [12], Iran | Retrospective cohort, multi-center | 105 dead patients | Not reported | Range (0 to >60) | Males were > females | Not reported | Not reported | 105 (100) | - | - | Not reported | 18 Influenza virus (H1N1) 9 Bocavirus 8 RSV 5 Influenza virus (non-H1N1) 4 Parainfluenza virus 3 HMPV 2 Adenovirus |
Not reported | RT-PCR for respiratory specimens | 5 | Most of the co-infected cases were men aged >60 years; and had history of obesity, cancer, hepatitis, and kidney diseases. Prevalence of SARS-CoV-2 and influenza A virus co-infection in dead patients was high. |
| Hazra et al., 2020 [53], United States | Retrospective cohort, single center | 459 | 15 (3.3) | Median, 39 | Not reported | Not reported | Not reported | Not reported | - | - | 15 (3.3) | 2 Adenovirus 1 Coronavirus NL63 2 HMPV 3 Influenza A virus 1 Parainfluenza 2 virus 8 Rhinovirus/Enterovirus |
Not reported | RT-PCR for respiratory specimens | 5 | Co-infected patients were younger than those only infected with SARS-CoV-2 (age: 39 vs. 58 years, p = 0.02). |
| Hughes et al., 2020 [26], United Kingdom | Retrospective case series, multi-center | 836 | 51 (6.1) | Median (IQR), 69 (55–81) | 519 (62) | 3 (5.9) | Not reported | Not reported | 51 (6.1) | 30 (3.6) | - | 8 Enterobacterales 36 CoNS 4 Streptococcus spp. 7 S. aureus 4 Enterococcus spp. 3 Candida albicans 1 P. aeruginosa 12 Pseudomonas spp. 5 Enterobacter spp. 6 Klebsiella spp. 2 Serratia spp. 24 Candida spp. 3 Aspergillus spp. 1 H. influenzae 1 Hafnia spp. 1 Morganella morganii 1 Providencia spp. 2 S. maltophilia |
Not reported | RT-PCR for respiratory specimens [viruses] AND blood, sputum, and BAL cultures [bacteria and fungi] | 6 | Rate of bacterial co-infection in SARS-CoV-2 patients in the early phase of hospital admission was low. |
| Karami et al., 2020 [54], The Netherlands | Retrospective cohort, multi-center | 925 | 12 (1.2) | Median (IQR), 70 (59–77) | 591 (64) | 166 (21.9) | Not reported | 214 (23.3) | 12 (1.2) | - | 2 (0.2) | 7 S. aureus 1 K. oxytoca 1 S. maltophilia 1 Parainfluenzae virus 1 H. influenzae 1 Influenza A virus 1 S. pneumoniae 2 E. coli |
No extractable data | Blood and sputum cultures [bacteria and viruses] | 6 | On presentation to the hospital, bacterial co-infections are rare. |
| Kim et al., 2020 [55], United States | Retrospective cohort, single center | 116 | 23 (19.8) | Median (IQR), 46.9 (14–74) | 12 (52.2) | 0 | 0 | 0 | - | - | 23 (19.8) | 8 Rhinovirus/Enterovirus 6 RSV 5 Coronavirus (non-SARS, non-MERS) 2 HMPV 1 Parainfluenza 1 1 Parainfluenza 3 1 Parainfluenza 4 1 Influenza A virus |
Not reported | RT-PCR via nasopharyngeal swab | 8 | Patients with co-infections did not differ significantly in age (mean, 46.9 years) from those infected with SARS-CoV-2 only (mean, 51.1 years). |
| Koehler et al., 2020 [20], Germany | Retrospective case series, single center | 19 | 5 (26.3) | Mean (SD), 62.6 (8.8) | 3 (60) | 5 (100) | Not reported | 3 (60) | - | 5 (26.3) | 2 (10.5) | 2 HMPV 5 Aspergillus fumigatus |
5 Antifungals | RT-PCR for respiratory specimens [viruses] AND GM detection in the BAL and tracheal aspirates | 6 | Critical cases of SARS-CoV-2 patients were at risk of developing aspergillosis co-infection and had higher mortality. |
| Kreitmann et al., 2020 [56], France | Prospectivecohort, single center | 47 | 13 (27.6) | Median (IQR), 61 (56–74) | 25 (73.5) | 47 (100) | Not reported | 5 (35.8) | 13 (27.6) | - | - | 9 S. aureus 5 H. influenzae 3 S. pneumoniae 1 M. catarrhalis 1 Streptococcus agalactiae |
4 Third Generation Cephalosporins 2 Macrolides 3 Other antibiotics |
RT-PCR for respiratory specimens and/or cultures | 6 | Authors argue for initial empirical antibiotic coverage in SARS-CoV-2 patients. |
| Lehmann et al., 2020 [57], United States | Retrospective cohort, single center | 321 | 12 (3.7) | Mean (SD), 60 (17) | 155 (48) | 17 (5) | Not reported | 22 (7) | 7 (2.2) | - | 5 (1.5) | 2 S. aureusa 1 Proteus mirabilisa 3 Influenza A virus 2 Rhinovirus/Enterovirus 1 Bordetella parapertussis 4 S. pneumoniae |
Antibiotic use was high (222 [69%]); however, agents were not reported. | RT-PCR for respiratory specimens and/or cultures | 7 | Community-acquired co-infection in COVID-19 is infrequent and often viral. Co-infection was more common among ICU patients. |
| Li Y et al., 2021 [27], China | Retrospective case series, single center | 81 | 27 (33.3) | Mean (SD), 76.55 (9.64) | 15 (55.6) | 1 (3.7) | 1 (3.7) | 0 | 27 (33.3) | - | 6 (7.4) | 23 M. Pneumoniae 1 Influenza A virus 2 Influenza B virus 1 RSV 1 Adenovirus 1 Parainfluenza virus 3 M. catarrhalis 1 S. pneumoniae |
No extractable data | Direct immunofluorescence test AND serum antibody test (IgM) | 7 | Almost 1/3 (33.3%) had co-infection. Coinfection did not cause a significant exacerbation in clinical symptoms. |
| Li Z et al., 2020 [28], China | Retrospective case series, multi- center | 32 | 14 (43.7) | Median (IQR), 57 (47–69) | 11 (78.6) | 11 (78.6) | 4 (28.6) | Not reported | 10 (31.2) | 7 (21.9) | 5 (15.6) | 3 Stephanoascus ciferrii 4 Candida albicans 2 Staphylococcus epidermidis 1 Ralstonia mannitolilytica 3 Stenotrophomonas maltophilia 1 Bacteroides fragilis 3 Burkholderia estoste 2 Enterococcus Faecium 1 E. coli 2 Elizabethkingia meningosepticum 1 A. baumannii 1 RSV 1 HMPV 2 HcoV-HKU1 1 Rhinovirus 1 Parainfluenza virus 1 Enterovirus |
Not reported | RT-PCR AND cultres | 6 | SARS-CoV-2 patients with co-infections were admitted more often to ICU (p < 0.05), showed more severe difficulty in breathing (p < 0.05), and experienced more complications such as ARDS and shock (p < 0.05). |
| Lin et al., 2020 [29], China | Retrospective case series, single center | 92 | 6 (6.5) | Majority (≈78%) were in the range (18–65) | 1:1 ratio | Not reported | Not reported | Not reported | - | - | 6 (6.5) | 3 RSV 2 Rhinovirus 2 HMPV 1 Parainfluenza 2 virus 2 HcoV-HKU1 |
Not reported | RT-PCR of respiratory tract specimens (naso- vs. oropharyngeal source not specified) | 7 | Limitation of the sensitivity of method for the different respiratory viruses and low load of virus in specimens might have contributed to negative results. |
| Liu H et al., 2020 [30], China | Retrospective case series, multi-center | 4 | 2 (50) | Range (2 months to 9 years) | 1:1 ratio | 0 | 0 | 0 | 1 (25) | - | 1 (25) | 1 M. pneumoniae 1 RSV |
Not reported | Unspecified | 6 | Pulmonary involvement was more severe, as simultaneous infection of RSV and SARS-CoV-2 in one child was detected. |
| Liu L et al., 2020 [31], China | Retrospective case series, single center | 53 | 31 (58.5) | Median (IQR), 38 (28–47) | 26 (49) | 1 (1.9) | 1 (1.9) | 0 | 25 (47.2) | - | 6 (11.3) | 25 M. pneumoniae 2 Influenza A virus 2 Influenza B virus 2 RSV |
25 Fluoroquinolones | Serum antibody test (IgM, IgG) | 6 | COVID-19 patients co-infected with M. pneumoniae had a higher percentage of monocytes (p < 0.0044) and a lower neutrophils percentage (p < 0.0264). |
| Ma et al., 2020 [32], China | Retrospective case series, single center | 93 | 46 (49.5) | Median (IQR), 67 (54–72) | 51 (54.8) | Not reported | Not reported | 44 (47.3) | - | - | 46 (49.5) | 44 Influenza A virus 2 Influenza B virus 1 Adenovirus 1 Parainfluenza virus |
Not reported | Serum antibody test (IgM) | 6 | Critically ill COVID-19 patients with influenza were more prone to cardiac injury than those without influenza.Critically ill COVID-19 patients with influenza exhibited more severe inflammation and organ injury. |
| Mannheim et al., 2020 [33], United States | Retrospective case series, multi-center | 10 | 4 (40) | Median (IQR), 11 (7–16) | Males were > females | 7 (70) | Not reported | 0 | 2 (20) | - | 3 (30) | 1 M. pneumoniae 1 Adenovirus 1 Rhinovirus/Enterovirus 1 E. coli 1 Rotavirus |
Not reported | RT-PCR for respiratory specimens | 6 | Underlying co-infection might have contributed to severe disease. |
| Massey et al., 2020 [58], United States | Retrospective cohort, multi-center | 1456 | Not reported | Mean (SD), 72.4 (20.9) | Not reported | Not reported | Not reported | Not reported | Not reported | - | Not reported | 937 S. aureus 576 EBV 574 HHV6 328 M. catarrhalis 64 K. pneumoniae 305 HMPV109 Adenovirus |
Not reported | RT-PCR for respiratory specimens | 6 | Advanced age and nursing home status were associated with higher co-infection rates in SARS-CoV-2 patients. In SARS-CoV-2 patients, 86.3% had at least one co-infection compared to 75.7% in the negative SARS-CoV-2 group (p < 0.0001). |
| May et al., 2021 [3], United Kingdom | Retrospective cohort, single center | 77 | 39 (50.6) | Not reported | Not reported | 39 (100) | Not reported | Not reported | 28 (36.4) | 11 (14.3) | - | 12 S. aureus 1 Staphylococcus lugdunensis 7 H. influenza 2 S. pneumoniae 10 Klebsiella spp. 3 Serratia marcescens 3 Citrobacter spp. 3 Enterobacter cloacae 3 Proteus mirabilis 2 E. coli 2 P. aeruginosa 1 Hafnia alvei 4 Enterococcus spp. 5 Aspergillus |
Not reported | Unspecified | 5 | There was no significant correlation between hospital mortality and isolation of a pathogen in early or any respiratory sample (p = 0.512 and p = 1.0, respectively). |
| Mo et al., 2020 [81], China | Retrospective cohort, single center | 155 | 12 (7.7) | Median (IQR), 54 (42–66) | 86 (55.5) | 37 (23.9) | 36 (23.2) | 22 (14.2) | 2 (1.3) | - | 13 (8.4) | 3 Parainfluenza virus 3 RSV 3 Adenovirus 2 Mycoplasma 2 Influenza A virus 2 Influenza B virus |
Not reported | Unspecified | 5 | COVID-19 patients were divided into general and refractory groups. |
| Nasir et al., 2020 [34], Pakitstan | Retrospective case series, single center | 23 | 9 (39.1) | Median (IQR), 71 (51–85) | 7 (77.8) | 23 (100) | 2 (22.2) | 4 (17.4) | 9 (39.1) | 5 (21.7) | - | 2 Aspergillus fumigatus 1 Aspergillus niger 6 Aspergillus flavus 2 P. aeruginosa 1 K. pneumoniae 1 MRSA 2 Acinetobacter spp. 1 Clostridium perfringens 2 Stenotrophomonas maltophilia |
7 Macrolides5 Antifungals | Culture from respiratory specimens and GM detection in the BAL, tracheal aspirates and serum | 6 | Invasive aspergillosis is a complication in moderate to severe COVID-19 patients. |
| Nowak et al., 2020 [59], United States | Retrospective cohort, multi-center | 1204 1270 1103 1103 1103 1103 |
1 (0.1) 4 (0.3) 17 (1.5) 8 (0.7) 4 (0.4) 2 (0.2) |
Mean, 60.1 | 16 (44) | Not reported | Not reported | Not reported | - | - | 36 (2.8) | 1 Influenza A virus 4 RSV 17 Other Coronaviridae [7 NL63, 5 HKU1, 4 229E, 1 OC43] 8 Rhinovirus/Enterovirus 4 HMPV 2 Adenovirus |
Not reported | RT-PCR for respiratory specimens | 6 | Study hypothesized that competitive advantage may play a role in the SARS-CoV-2 interaction with other respiratory viruses during co-infection. |
| Oliva et al., 2020 [35], Italy | Retrospective case series, single center | 182 | 7 (3.8) | Median (IQR), 73 (45–79) | 4 (57.1) | 1 (14.3) | Not reported | 0 | 7 (3.8) | - | - | 5 C. pneumoniae 2 M. pneumoniae |
7 Macrolides 1 Teicoplanin 1 Beta-Lactam/Beta-Lactamase Inhibitors 1 Third Generation Cephalosporins |
Serum antibody test (IgM) | 6 | ICU admission and mortality were similar in the SARS-CoV-2 patients co-infected with M. pneumoniae or C. pneumoniae compared to SARS-CoV-2 group without the co-infection (14.2% vs. 13.7% and 0% vs. 14.2%, respectively). |
| Ozaras et al., 2020 [60], Turkey | Retrospective cohort, multi-center | 1103 | 6 (0.54) | Mean (SD), 40.5 (14) | 3 (50) | 0 | 0 | 0 | - | - | 6 (0.5) | 2 Influenza A virus 4 Influenza B virus |
6 Macrolides | Direct immunofluorescence test | 6 | Cases reported in this study were mild to moderate in severity. |
| Peng et al., 2020 [36], China | Retrospective case series, single center | 75 | 42 (56) | Mean (range), 6.06 years (1 month–15 years) | 44 (58.67) | Not reported | Not reported | 0 | 31 (41.3) | - | 8 (10.7) | 28 M. pneumonia 1 M. catarrhalis 1 S. aureus 1 S. pneumoniae 3 Influenza B virus 1 Influenza A virus 2 Adenoviridae 1 CMV 1 RSV |
30 Macrolides Thirty-seven patients received antibiotics; however, agents were not reported. |
Serum antibody test (IgM) | 6 | Co-infection never increased patients’ length of stay or decreased time of SARS-CoV-2 virological clearance. |
| Pongpirul et al., 2020 [37], Thailand | Retrospective case series, multi-center | 11 | 11 (100) | Median (IQR), 61 (28–74) | 6 (54.5) | 0 | 0 | 0 | 5 (45.4) | - | 2 (18.2) | 4 H. influenzae 1 Adenovirus 1 Influenza A virus 1 K. pneumoniae |
5 Third Generation Cephalosporins 2 Beta-Lactam/Beta-Lactamase Inhibitors |
RT-PCR via nasopharyngeal and oropharyngeal swabs and sputum specimens | 8 | Nasopharyngeal and oropharyngeal swabs and sputum specimens were also tested for 33 respiratory pathogens. |
| Ramadan et al., 2020 [21], Egypt | Prospective cohort, multi- center | 260 | 28 (10.8) | Most common age range was between 51 and 70 years (36.2%) | 144 (55.4) | 60 (23) | 8 (13.3) | 24 (40) | 28 (10.8) | 5 (1.9) | - | 5 S. aureus 2 S. pneumoniae 1 E. faecalis 12 K. pneumoniae 7 A. baumannii 4 E. coli 4 P. aeruginosa 2 Enterobacter cloacae 3 Candida albicans 2 Candida glabrata |
28 Macrolides | Respiratory and blood cultures | 7 | Eight (28.6%) patients who had co-infections were moderate cases, while 20 (71.4%) were detected in severe COVID-19 patients. Mortality in 25% of SARS-CoV-2 patients was due to co-infections and increased SARS-CoV-2 severity and complications were observed in co-infected patients. Bacterial co-infection and multidrug resistance among patients with COVID-19 in Upper Egypt is common. |
| Richardson et al., 2020 [38], United States | Retrospective case series, multi-center | 1996 | 42 (2.1) | Not reported | Not reported | Not reported | Not reported | Not reported | Not reported | - | Not reported | 22 Enterovirus/Rhinovirus 7 Coronavirus (non–COVID-19) 4 RSV 3 Parainfluenza 3 2 C. pneumoniae 2 HMPV 1 Influenza A virus 1 M. pneumoniae |
Not reported | Respiratory viral panel | 8 | Most patients were obese (60.7% had a BMI≥30) and old (median (IQR): 63 (52–75)). |
| Rutsaert et al., 2020 [39], Belgium | Retrospective case series, single center | 34 | 6 (17.6) | Median (IQR), 74 (38–86) | 6 (100) | 6 (100) | 6 (100) | 4 (66.7) | - | 6 (17.6) | - | 5 Aspergillus fumigatus 1 Aspergillus flavus |
5 Antifungals | Culture from respiratory specimens and GM detection in the BAL and serum | 6 | Patients were old and had deteriorating outcomes due to many medical conditions and risk factors. |
| Schirmer et al., 2021 [13], United States | Retrospective cohort, multi-center | 3757 | 56 (1.5) | Median (IQR), 68 (56–74) | 55 (98) | 10 (26) | Not reported | 10 (18) | 1 (0.03) | - | 55 (1.5) | 2 Adenovirus 1 C. pneumoniae 13 Coronaviruses (HKU1, NL63, 229E, & OC43) 3 HMPV 2 Parainfluenza virus 4 12 Influenza A virus 3 Influenza B virus 4 RSV 19 Rhinovirus/Enterovirus |
Not reported | Molecular and/or viral culture respiratory assays [multiplex respiratory pathogen panels] | 6 | Individuals with COVID-19 co-infection had higher odds of being male. |
| Sepulveda et al., 2020 [61], United States | Retrospective cohort, multi- center | 4185 | 159 (3.8) | Not reported | Not reported | Not reported | Not reported | Not reported | 156 (3.7) | 3 (0.07) | - | 39 Staphylococcus epidermidis 28 Staphylococcus hominis 8 E. coli 8 Staphylococcus haemolyticus 8 CoNS 5 Corynebacterium 5 Enterobacter cloacae complex 5 Micrococcus luteus 5 Staphylococcus warneri 1 Actinomyces turicensis 1 Aerococcus urinae 1 Candida glabrata 1 Comamonas estosterone 1 Dolosigranulum pigrum 1 Eneterobacter 1 Enterococcus faecium, Vancomycin-Resistant 1 Globicatella sanguinis 1 Granulicatella adiacens 1 Kocuria marina 1 Moraxella osloensis 1 Rothia aeria 1 S. aureus 1 Staphylococcus auricularis 1 Staphylococcus lugdunensis 1 Streptococcus intermedius 1 Streptococcus sanguinis 2 Enterococcus faecalis 2 E. coli 2 Fusobacterium spp. 2 Lactobacillus 2 Streptococci, Viridans Group 2 Streptococcus anginosus 2 Streptococcus spp. 6 K. pneumoniae 6 MSSA 11 Staphylococcus capitis 10 Methicillin Susceptible- CoNS 9 Bacillus non-anthracis 7 Methicillin Resistant-CoNS 4 MRSA 3 Candida albicans |
Not reported | Blood cultures | 6 | Rate of bacteremia was significantly lower among COVID-19 patients (3.8%) than among COVID-19-negative patients (8.0%) (p < 0.001). More than 98% of all positive cultures were detected within 4 days of incubation. The most common causes of true bacteremia among COVID-19 patients were E. coli (16.7%), S. aureus (13.3%), K. pneumoniae (10.0%), and Enterobacter cloacae complex (8.3%). |
| Singh et al., 2021 [62], United States | Retrospective cohort, multi-center | 4259 | 1,558 (36.59) | Mean (SD), 45.21 (20.43) | 692 (44.4) | Not reported | Not reported | Not reported | 517 (12.1) | - | 53 (1.2) | 53 H. influenzae 75 S. aureus 1 Bordetella pertussis 1 C. pneumoniae 11 K. pneumoniae 1 M. pneumoniae 49 S. pneumoniae 2 Adenovirus 1 Coronavirus 1 Herpes virus 5 12 EBV 1 RSV 3 Rhinovirus 1 HSV 1 HMPV 1 PIV 1 Influenza virus |
Not reported | RT-PCR for respiratory specimens | 6 | Co-infections were significantly higher in the older age group (60+ years). |
| Song et al., 2020 [63], China | Retrospective cohort, single center | 89 | 18 (20.2) | Median (IQR), 35.5 (15–76) | Not reported | 2 (11.1) | Not reported | Not reported | 18 (20.2) | - | - | 6 K. pneumoniae 5 E. coli 4 M. catarrhalis 4 H. influenzae 2 A. baumannii 2 S. aureus 1 P. aeruginosa 1 Streptococcus Group A |
Not reported | RT-PCR for respiratory specimens | 6 | Authors did not detect co-infection of SARS-CoV-2 with other viruses. |
| Sun et al., 2020 [40], China | Retrospective case series, single center | 36 | ≈23 (62.86) | Mean (range), 6.43 months (2–12 months) | 22 (61.11) | 1 (2.78) | 1 (2.78) | 1 (2.78) | 1 (2.8) | - | 1 (2.8) | 1 M. pneumonia 1 Influenza A virus |
15 Second Generation Cephalosporins15 Macrolides | Unspecified | 6 | Co-infections were common in infants with COVID-19, which were different from adults with COVID-19; however, authors never provided details of all co-pathogens. |
| Tagarro et al., 2021 [41], Spain | Retrospective case series, multi-center | 41 | 2 (4.8) | Mean (range), 1 (0–15) | Females were > males | 4 (9.7) | 1 (2) | 0 | - | - | 2 (4.9) | 2 Influenza B virus | Not reported | Unspecified | 7 | Most patients who tested positive for SARS-CoV-2 had no comorbidities (67%). |
| Tang et al., 2021 [64], China | Retrospective cohort, single center | 78 | 11 (14.1) | Mean (SD), 42.7 (14.9) | 41 (52.6) | 2 (18.2) | 2 (18.2) | 0 | 6 (7.7) | - | 6 (7.7) | 5 M. pneumoniae 4 RSV 2 C. pneumoniae 1 Influenza B virus 1 Adenoviruses 1 Legionella pneumophila |
48 Fluoroquinolones 5 Beta-Lactam/Beta-Lactamase Inhibitors 3 Linezolid 1 Vancomycin 3 Carbapenems |
Serum antibody test (IgM) | 6 | SARS-CoV-2 patients with co-infections had significantly higher levels of procalcitonin compared to SARS-CoV-2 patients with no co-infections (p = 0.002). |
| Thelen et al., 2021 [65], The Netherlands | Retrospective cohort, multi-center | 678 | 61 (9) | Median (IQR), 70 (58–78) | 443 (65.1) | 6 (0.9) | Not reported | Not reported | 61 (9) | - | - | 2 E. coli 1 K. pneumoniae 1 P. aeruginosa 2 S. pneumoniae 1 Other Streptococcus spp. 1 S. aureus 55 CoNS 1 Corynebacterium spp. |
Not reported | RT-PCR for respiratory specimens AND blood cultures | 6 | Prevalence of co-infection in SARS-CoV-2 patients was very low compared to influenza patient group. |
| Van Arkel et al., 2020 [42], The Netherlands | Retrospective case series, single center | 31 | 6 (19.3) | Median (IQR), 62.5 (43–83) | 6 (100) | 6 (100) | 6 (100) | 4 (66.7) | - | - | 5 Aspergillus fumigatus | 6 Antifungals | Culture from respiratory specimens and GM detection in the BAL, tracheal aspirates, and serum. | 6 | Pulmonary aspergillosis co-infections occurred after a median of 11.5 days (8–42) after COVID-19 symptom onset and at a median of 5 days (3–28) after ICU admission. | |
| Wang L et al., 2021 [22], United Kingdom | Retrospective cohort, multi- center | 1396 | 37 (2.7) | Median (IQR), 76 (64–82) | 28 (75.7) | 11 (29.7) | Not reported | 10 (27) | 37 (2.7) | 4 (0.3) | - | 12 E. coli 2 K. pneumoniae 2 Klebsiella variicola 4 Proteus mirabilis 2 P. aeruginosa 1 MRSA 7 MSSA 1 Staphylococcus epidermidis 1 Candida albicans 2 Group A Streptococcus 1 H. influenzae 3 Candida spp. 2 Enterococcus faecalis 3 S. pneumoniae 1 Serratia spp. 1 Klebsiella oxytoca 1 Streptococcus anginosus 1 Bacteroides ovatus 1 Granulicatella adiacens 1 S. aureus |
Not reported | Unspecified | 7 | ICU admission and mortality were not different in SARS-CoV-2 patients with co-infections compared to SARS-CoV-2 patients without co-infections [215 (15.8%) vs. 11 (29.7%), p = 0.075] and [410 (30.2%) vs. 10 (27.0%), p = 0.68], respectively. Bacterial co-infection was infrequent in hospitalized COVID-19 patients within 48 hours of admission. |
| Wang R et al., 2020 [43], China | Retrospective case series, single center | 118 | 35 (29.7) | Mean (SD), 38.76 (13.79) | (56.8) | 19 (16.1) | 4 (3.4) | 0 | 35 (29.7) | - | 1 (0.8) | 40 M. pneumoniae 1 Adenovirus 1 Influenza B virus 1 Influenza A virus |
Seventy-nine patients received antibiotics; however, agents were not reported. | Serum Antibody test (IgM) | 6 | Old age, chronic underlying diseases, and smoking history may be risk factors that worsen SARS-CoV-2 disease. |
| Wang Y et al., 2020 [44], China | Retrospective case series, single center | 55 | 4 (7.3) | Median (IQR), 49 (2–69) | 22 (40) | 0 | 0 | 0 | 3 (12.7) | - | 1 (1.8) | 1 EBV 3 M. pneumoniae |
Not reported | Serologically | 7 | All patients included in this study had laboratory-confirmed positive results for SARS-CoV-2 and were asymptomatic. |
| Wang Z et al., 2020 [45], China | Retrospective case series, single center | 29 sputum 28 blood |
5 (17.2) 4 (14.3) |
Majority (51%) were in the range (30–49) | Females were > males | Not reported | Not reported | 5 (7.5) | 5 (≈17.2) | 2 (6.9) | 2 (7.1) | 2 Candida albicans 2 Enterobacter cloacae 1 A. baumannii 2 Chlamydia 1 RSV 1 Adenovirus |
39 Fluoroquinolones 8 Antifungals |
Serum Antibody test (IgM, IgG) | 7 | Source of patients’ samples tested for co-pathogens were sputum and blood. |
| Wee et al., 2020 [66], Singapore | Prospective cohort, single center | 431 | 6 (1.4) | Mean (SD), 29.2 (1.7) | 6 (100) | 0 | 0 | 0 | 0 | - | 6 (1.4) | 3 Rhinovirus 2 Parainfluenza 1 Other coronavirus (229E/NL63/OC43) |
Not reported | RT-PCR for respiratory specimens | 6 | Co-infections in patients with SARS-CoV-2 shown no increase in morbidity or mortality. All cases of COVID-19 co-infections were young, healthy, and had no medical comorbidities. |
| Wu C et al., 2020 [67], China | Retrospective cohort, single center | 173 | 1 (0.6) | Majority (80.1%) had a median age <65 | Males were > females | 53 (26.4) | 67 (33.3) | 44 (21.9) | - | - | 1 (0.6) | 1 Influenza A virus | Not reported | RT-PCR for respiratory specimens [viruses] AND sputum culture [bacteria and fungi] | 8 | Most (n = 173 [86.1%]) patients were tested for 9 additional respiratory pathogens. Bacteria and fungi cultures were collected from 148 (73.6%) patients. |
| Wu Q et al., 2020 [46], China | Retrospective case series, multi-center | 34 | 19 (55.9) | Range (≤3 month to >10 years) | Males were > females | 0 | 1 (2.9) | 0 | 16 (47) | - | 10 (29.4) | 16 M. pneumoniae 2 RSV 2 EBV 3 CMV 1 Influenza A virus 1 Influenza B virus |
15 Macrolides | Unspecified | 7 | Nearly one-half of the infected children had co-infection with other common respiratory pathogens. |
| Xia et al., 2020 [47], China | Retrospective case series, single center | 20 | 8 (40) | Range (<1 month to >6 years) | Males were > females | 0 | 0 | 0 | 4 (20) | - | 5 (25) | 1 CMV 2 Influenza B virus 1 Influenza A virus 4 Mycoplasma 1 RSV |
Not reported | Unspecified | 5 | Procalcitonin increased in most of the cases (80%). |
| Yang et al., 2020 [48], China | Retrospective case series, single center | 52 | 7 (13.5) | Majority (73%) were in the range (50–79) | Males were > females | 52 (100) | 37 (71) | 32 (61.5) | 4 (7.7) | 3 (5.8) | - | 2 K. pneumoniae 1 Aspergillus flavus 1 Aspergillus fumigatus 1 P. aeruginosa 1 Serratia marcescens 1 Candida albicans |
Forty-nine patients received antibiotics; however, agents were not reported. | Respiratory and blood cultures | 8 | Those isolated pathogens caused hospital-acquired infections. |
| Yue et al., 2020 [68], China | Retrospective cohort, single center | 307 | 176 (57.3) | Mean (SD), 60.3 (16.5) | 75 (42.6) | Not reported | Not reported | Not reported | - | - | 176 (57.3) | 153 Influenza A virus 23 Influenza B virus |
None | Serum antibody test (IgM) | 6 | Patients co-infected with SARS-CoV-2 and Influenza B virus developed poor outcomes (30.4% vs. 5.9%). |
| Zha et al., 2020 [82], China | Retrospective case series, single center | 874 | 22 (2.5) | Median (IQR), 56.5 (52.5–66.5) | 11 (50) | Not reported | Not reported | 1 (4.5) | 22 (2.5) | - | - | 22 M. pneumoniae | 18 Fluoroquinolones 11 Cephalosporins 3 Beta-Lactam/Beta-Lactamase Inhibitors |
RT-PCR for respiratory specimens OR serum antibody test (IgM) | 6 | Length of cough was longer in the M. pneumoniae co-infection group (20 vs. 16.25, p = 0.043), while the length of hospital stay was slightly longer (16 vs. 14, p = 0.145). |
| Zhang et al., 2020 [50], China | Retrospective case series, single center | 140 | 7 (5) | Majority (70%) were > 50 | 1:1 ratio | Not reported | Not reported | Not reported | 5 (3.6) | - | 2 (1.4) | 5 M. pneumonia 1 RSV 1 EBV |
Not reported | Serum antibody test (IgM, IgG) | 5 | No clinical and radiological signs of co-infection caused by these pathogens were identified. Increased procalcitonin (p = 0.004) was more commonly observed in severe patients. |
| Zhao et al., 2020 [69], China | Prospective cohort, multi-center | 19 | 2 (10.5) | Median (IQR), 48 (27–56) | Males were > females | 0 | 0 | 0 | 1 (5.3) | - | 1 (5.3) | 1 Coxsackie virus 1 Mycoplasma |
None | RT-PCR for respiratory specimens AND serum antibody test (IgM) | 6 | Sample size was very small. |
| Zheng F et al., 2020 [49], China | Retrospective case series, multi-center | 25 | 6 (24) | Range (1 month to ≥6 years) | Males were > females | 2 (8) | 2 (8) | 0 | 4 (16) | - | 2 (8) | 2 Influenza B virus 3 M. pneumonia 1 Klebsiella aerogenes |
1 Beta-Lactam/Beta-Lactamase Inhibitors 1 Carbapenems 1 Linezolid |
Unspecified | 5 | Highest incidence of infection occurred in children aged <3 years. |
| Zheng X et al., 2020 [70], China | Retrospective cohort, single center | 1001 | 4 (0.4) | Mean (SD), 35 (19.6) | 1:1 | 0 | 0 | 0 | - | - | 4 (0.4) | 3 Influenza A virus 3 Influenza B virus |
Three patients received antibiotics; however, agents were not reported. | RT-PCR for respiratory specimens | 7 | Patients with both SARS-CoV-2 and influenza virus infection showed similar clinical characteristics to those patients with SARS-CoV-2 infection only. Co-infection of SARS-CoV-2 and influenza viruses was low. |
| Zhu et al., 2020 [4], China | Retrospective cohort, single center | 257 | 243 (94.5) | Median (IQR), 51 (2−99) | 138 (53.7) | 3 (1.2) | 0 | 0 | 236 (91.8) | 60 (23.3) | 81 (31.5) | 153 S. pneumoniae 143 K. pneumoniae 103 H. influenza 60 Aspergillus 52 EBV 24 E. coli 21 S. aureus 12 Rhinovirus 12 P. aeruginosa 11 M. catarrhalis 10 Adenovirus 8 HSV 7 A. baumannii 6 C. pneumoniae 6 Mucor 5 Influenza B 4 M. pneumonia 3 Bordetella pertussis 2 Candida 3 CMV 2 Influenza A virus 1 Bocavirus 1 HMPV 1 Cryptococcus |
Not reported | RT-PCR for respiratory specimens | 7 | Highest and lowest rates of co-infections were found in patients aged 15–44 and below 15, respectively. Most co-infections occurred within 1–4 days of onset of COVID-19 disease. Proportion of viral, fungal and bacterial co-infections were the highest in severe COVID-19 cases. |
Abbreviations: BA, bronchoaspirate; BAL, bronchoalveolar lavage; GM, galactomannan; IgG, immunoglobulin G; IgM, immunoglobulin M; RT-PCR, reverse transcription polymerase chain reaction; COVID-19, coronavirus disease 2019; ICU, intensive care unit; MERS-CoV, Middle East respiratory syndrome coronavirus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; NOS, Newcastle–Ottawa scale; C. pneumoniae, Chlamydia pneumoniae; M. pneumoniae, Mycoplasma pneumoniae; RSV, Respiratory syncytial virus; H. influenzae, Haemophilus influenzae; K. pneumoniae, Klebsiella pneumoniae; EBV, Epstein–Barr virus; P. aeruginosa, Pseudomonas aeruginosa; HcoV-HKU1, human coronavirus HKU1; S. pneumoniae, Streptococcus pneumoniae; M. catarrhalis, Moraxella catarrhalis; ARDS, acute respiratory distress syndrome; MRSA, methicillin-resistant Staphylococcus aureus; CMV, cytomegalovirus; S. aureus, Staphylococcus aureus; HSV, herpes simplex virus; A. baumannii, Acinetobacter baumannii; MSSA, methicillin-susceptible Staphylococcus aureus; CoNS, coagulase-negative staphylococci; AOR, adjusted odds ratio; CI, confidence interval; HHV6, human herpes virus 6; E. coli, Escherichia coli; spp., species; HMPV, human metapneumovirus.
2.4. Quality Assessment
The Newcastle–Ottawa Scale [NOS] was the primary tool for examining the quality of included studies, as described previously [75]. The tool provides maximum scores of 4 for selection, 2 for comparability, and 3 for exposure/outcome. High-quality studies have a score of >7, and moderate-quality studies have a score of 5–7. Quality assessment was performed independently by four authors (A.M.A., S.A.A., G.Y.A., and A.R.) and a consensus was used to resolve any disagreement.
2.5. Data Analysis
We examined primarily the proportion of confirmed acute bacterial, fungal and/or respiratory viral infections in patients with SARS-CoV-2. This proportion was further classified based on initial presentation or during the course of the illness. Taking a conservative approach, a random effects with the DerSimoniane–Laird model was used [76], which produces wider confidence intervals [CIs] than a fixed effect model. Results were illustrated using forest plots. The Cochran’s chi-square (χ2) and the I2 statistic provided the tools of examining statistical heterogeneity [77]. An I2 value of >50% suggested significant heterogeneity [78]. Examining the source of heterogeneity, a subgroup analysis was conducted based on ICU and non-ICU admission or only ICU admission. Funnel plots and Egger’s correlation test estimate publication bias and p value < 0.05 indicates statistical significance [79]. R version 4.1.0 with the packages metafor and meta was used for all statistical analyses.
3. Results
3.1. Characteristics and Quality of Included Studies
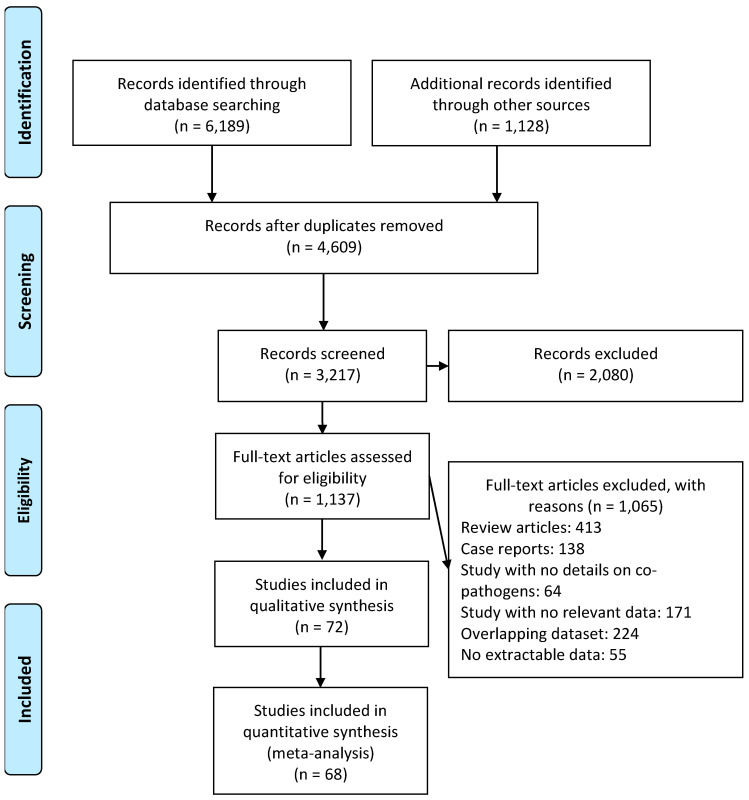
Of the initial 7317 retrieved publications, there were 4609 duplicate articles, and 2080 articles were found to be irrelevant based on their titles and abstracts and were excluded. An additional 1065 articles were excluded after review, meaning that we included 72 articles in the systematic review [2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,80,81,82], while 68 articles were included in the meta-analysis [2,3,4,5,6,7,8,9,10,11,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,39,40,41,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,59,60,61,62,63,64,65,66,67,68,69,70,80,81,82] (Figure 1).
Figure 1.
Flow diagram of literature search and data extraction from studies included in the systematic review and meta-analysis.
The included studies had a total of 31,953 SARS-CoV-2 infected patients as detailed in Table 1. Of those patients, 25,302 (79.2%) were from 32 cohort studies and 20.8% were from 40 case series. The geographical distribution of these studies was as follows: Asia (n = 36), Europe (n = 22), and North America (n = 14). The majority of the studies were single center and only 24 studies were multi-center. Laboratory techniques for co-pathogen detection within studies included 19 that used respiratory samples and RT-PCR tests [4,5,8,11,12,13,29,33,37,38,53,55,58,59,62,63,66,70,80], 17 that used serologic tests (antibodies) [6,10,14,19,24,31,32,35,36,43,44,45,50,52,60,64,68], 15 that used RT-PCR tests with respiratory and/or blood cultures [7,9,17,18,23,26,28,34,39,42,51,56,57,65,67], 12 that did not specify their testing methods [3,15,16,22,25,30,40,41,46,47,49,81], five that only used respiratory and/or blood cultures [2,21,48,54,61], and three that tested both serology and RT-PCR [27,69,82] (Table 1). Seven studies examined patients for influenza A and B only [10,11,19,41,60,68,70]; while five studies evaluated patients for the presence of Chlamydia or Mycoplasma [6,24,35,52,82]; and four studies only evaluated for the presence of fungi [17,23,39,42]. The proportion of patients receiving antibiotic agents was reported in 34 studies [2,6,7,14,16,17,18,19,20,21,23,24,31,34,35,36,37,39,40,42,43,45,46,48,49,51,52,56,57,60,64,70,80,82]. The most commonly used antimicrobials were macrolides (n = 355), 2nd/3rd/5th generation cephalosporins (n = 157), fluoroquinolones, (n = 150), antifungals (n = 62), beta-lactams/beta-lactam inhibitors (n = 26), beta-lactams (n = 21), tetracyclines (n = 17), linezolid (n = 13), carbapenems (n = 4), and glycopeptides (n = 2). The median NOS score was 6 with a range from 5 to 8. The NOS quality was moderate for 66 studies, and high quality for 6 studies. The majority (60/72, 83.3%) of the studies included only adult patients. The proportion of male patients had a median of 55.9% [interquartile range (IQR) 48.9–71.9%]. The majority (n = 58) of the studies included any hospitalized patient, and 14 studies included only critically ill. Sixteen, thirteen, and four studies exclusively reported on respiratory viral, bacterial, and fungal co-infections, respectively; and the remaining 39 studies reported on bacterial, fungal, and respiratory viral co-infections; Table 1.
3.2. Meta-Analysis of Bacterial, Fungal, and Respiratory Viral Co-Infections in Patients with SARS-CoV-2
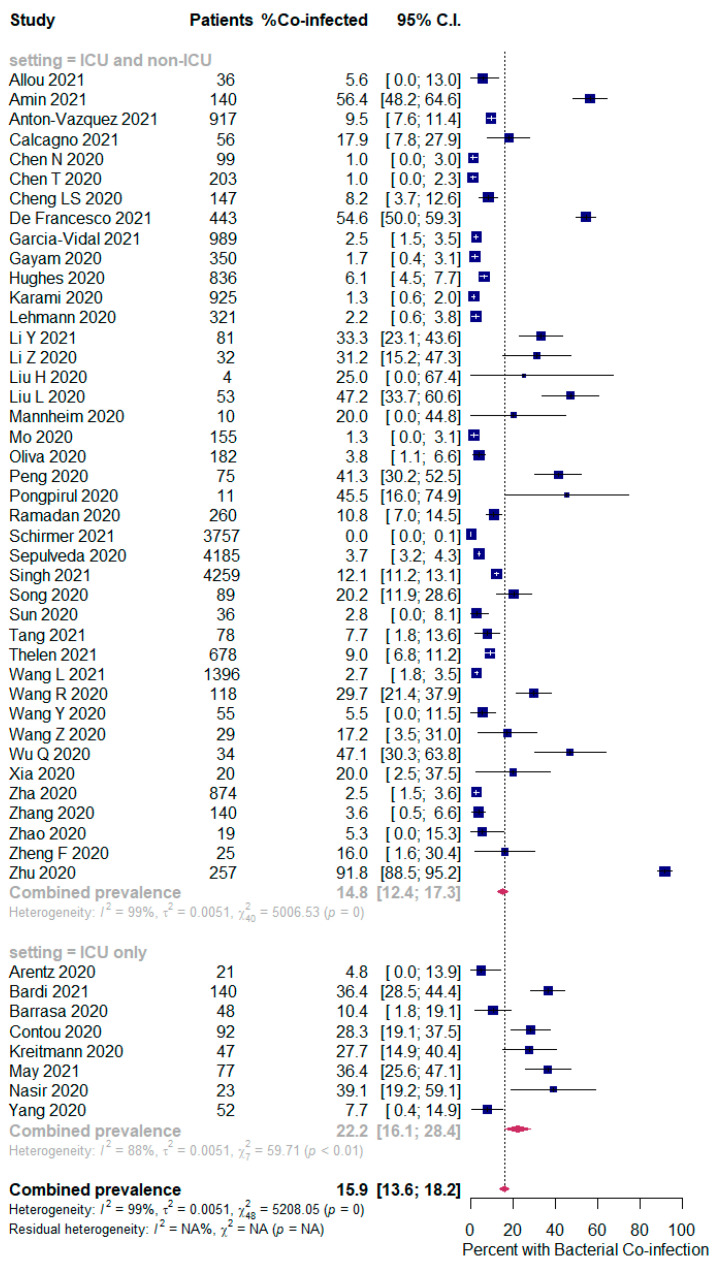
The overall pooled proportions of SARS-CoV-2 patients who had laboratory-confirmed bacterial, fungal, and respiratory viral coinfections were 15.9% (95% CI 13.6 to 18.2, n = 1940, 49 studies, I2 99%, p < 0.00001), 3.7% (95% CI 2.6 to 4.8, n = 177, 16 studies, I2 93%, p < 0.00001), and 6.6% (95% CI 5.5 to 7.6, n = 737, 44 studies, I2 96%, p < 0.00001), respectively; (Figure 2,Figure 3,Figure 4).
Figure 2.
Forest plot of proportion of SARS-CoV-2 patients with bacterial co-infections (all patients in the upper panel and only ICU patients in the lower panel).
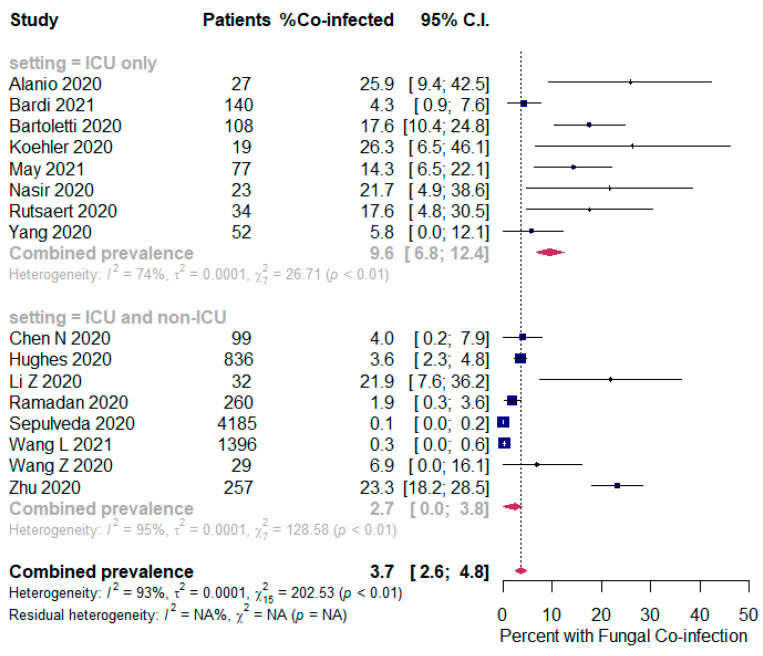
Figure 3.
Forest plot of proportion of SARS-CoV-2 patients with fungal co-infections (all patients in the upper panel and only ICU patients in the lower panel).
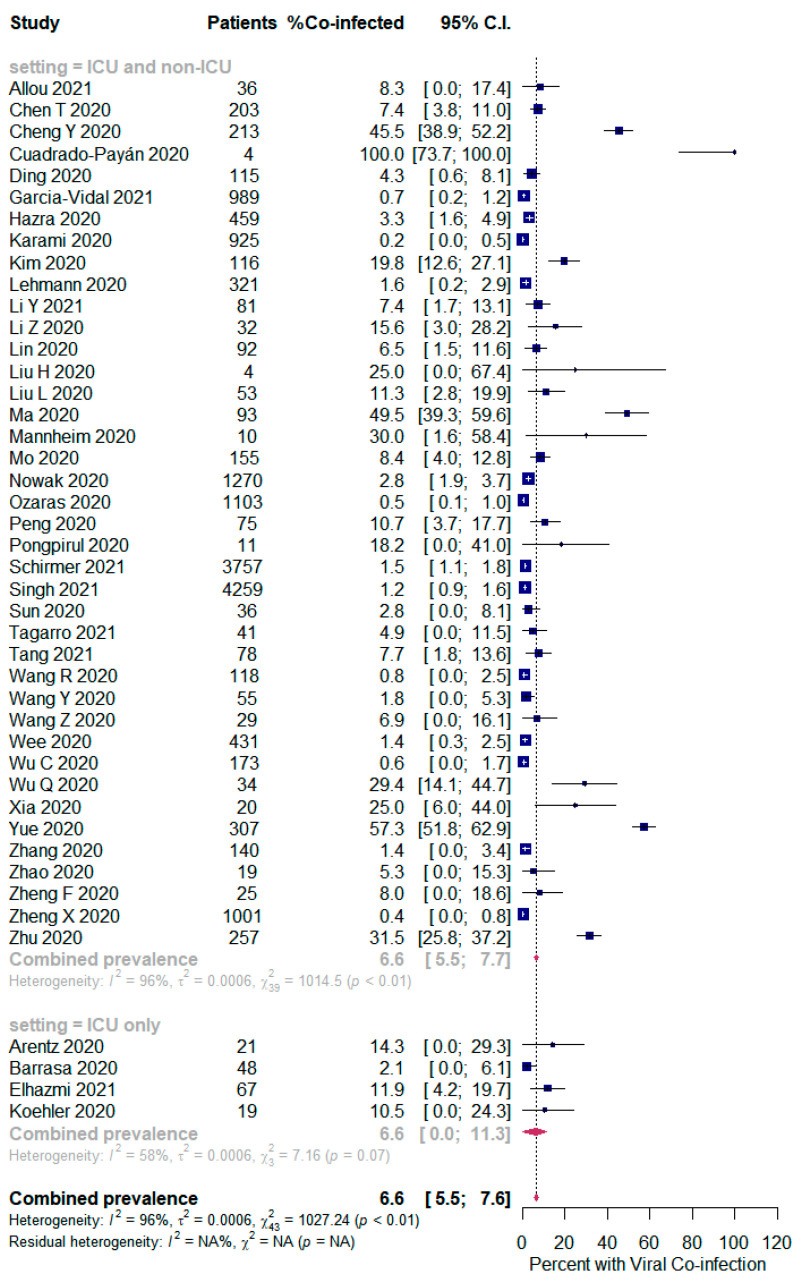
Figure 4.
Forest plot of proportion of SARS-CoV-2 patients with respiratory viral co-infections (all patients in the upper panel and only ICU patients in the lower panel).
In bacterial coinfected SARS-CoV-2 patients, subgroup analysis showed some difference in the rates between all patients (ICU and non-ICU group); and the ICU only group (14.8% (95% CI 12.4 to 17.3, n = 1802, 41 studies, I2 = 99%); and 22.2% (95% CI 16.1 to 28.4, n = 137, 8 studies, I2 = 88%), respectively); Figure 2. In the fungal co-infected SARS-CoV-2 patients, subgroup analysis showed a significant difference in the rates between all patients (ICU and non-ICU); and ICU only patients [2.7% (95% CI 0.0 to 3.8, n = 155, 8 studies, I2 = 95%); and 9.6% (95% CI 6.8 to 12.4, n = 62, 8 studies, I2 = 74%), respectively]; Figure 3.
However, in the respiratory viral co-infected SARS-CoV-2 patients, subgroup analysis showed an identical proportion between all patients (ICU and non-ICU) and the ICU only patients [6.6% (95% CI 5.5 to 7.7, n = 723, 40 studies, I2 = 96%); and 6.6% (95% CI 0.0 to 11.3, n = 14, 4 studies, I2 = 58%), respectively]; Figure 4.
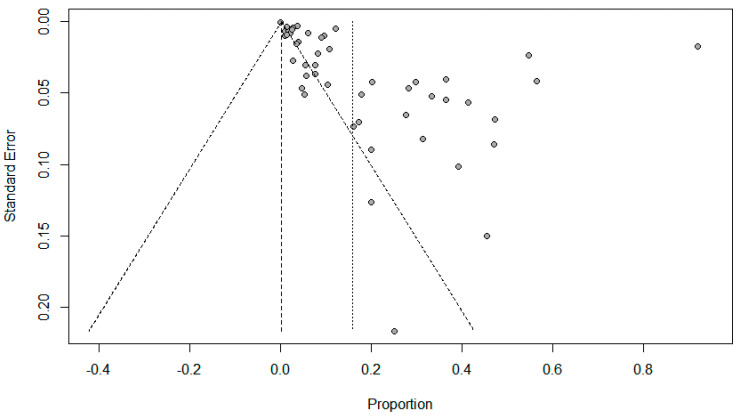
Funnel plots for possible publication bias for the pooled effect size to determine the prevalence of coinfections in SARS-Cov-2 patients appeared asymmetrical on visual inspection, and Egger’s tests confirmed asymmetry with p values < 0.05; Figure 5, Figure 6 and Figure 7.
Figure 5.
Funnel plots evaluating publication bias for the pooled effect size determining the prevalence of bacterial co-infections in SARS-Cov-2 patients.
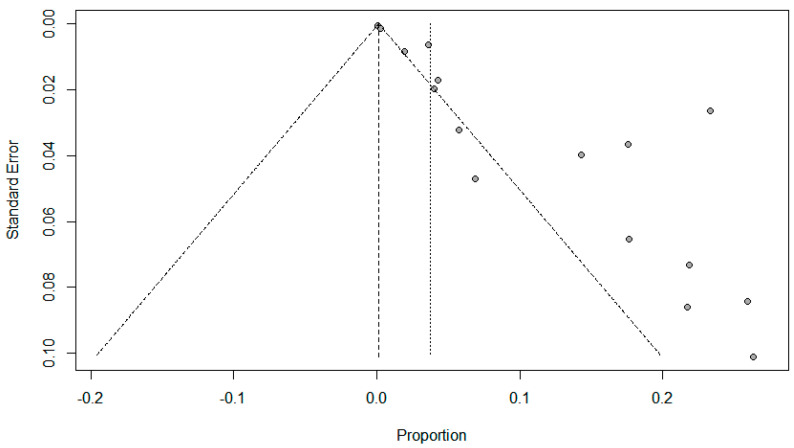
Figure 6.
Funnel plots evaluating publication bias for the pooled effect size to determine the prevalence of fungal co-infections in SARS-Cov-2 patients.
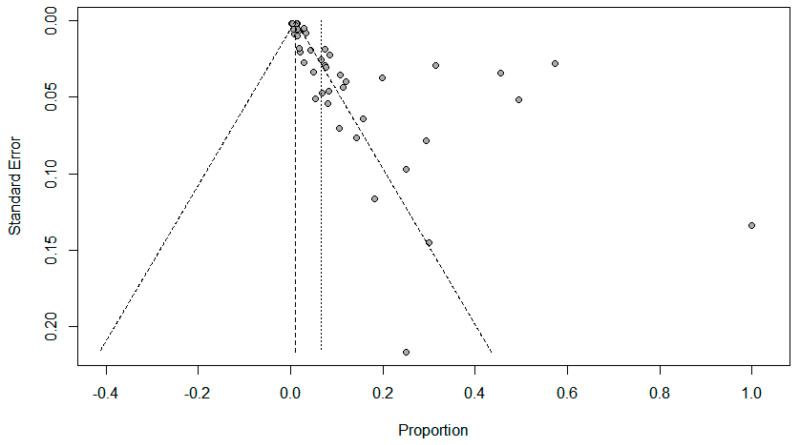
Figure 7.
Funnel plots to evaluate publication bias for the pooled effect size to determine the prevalence of other respiratory viral co-infections in SARS-Cov-2 patients.
3.3. Bacterial, Fungal and Respiratory Viral Co-Pathogens
Specific bacterial co-pathogens were reported in 49/72 (68%) studies, which is about 57.3% of the reported co-infections. The most common bacteria were S. aureus (n = 1095), M. catarrhalis (n = 352), M. pneumoniae (n = 338), S. pneumoniae (n = 316), C. pneumoniae (n = 261), K. pneumoniae (n = 259), and H. influenzae (n = 197) (Table 2).
Table 2.
Proportion of all identified SARS-CoV-2 bacterial co-infections (N = 3468).
| Bacterial Pathogen Type | Identified Number (%) | Bacterial Pathogen Type | Identified Number (%) |
|---|---|---|---|
| S. aureus | 1,095 (31.6) | Corynebacterium spp. | 6 (0.2) |
| M. catarrhalis | 352 (10.1) | Bordetella pertussis | 5 (0.1) |
| M. pneumoniae | 338 (9.7) | Micrococcus luteus | 5 (0.1) |
| S. pneumoniae | 316 (9.1) | Citrobacter koseri | 4 (0.1) |
| C. pneumoniae | 261 (7.5) | Hafnia alvei | 3 (0.1) |
| K. pneumoniae | 259 (7.5) | S. maltophilia | 3 (0.1) |
| H. influenzae | 197 (5.7) | Streptococcus anginosus | 3 (0.1) |
| CoNS | 115 (3.3) | Streptococcus Group A | 3 (0.1) |
| E. coli | 65 (1.9) | Burkholderia cepacia | 3 (0.1) |
| P. aeruginosa | 48 (1.4) | Bacteroides spp. | 3 (0.1) |
| Staphylococcus epidermidis | 42 (1.2) | Stephanoascus ciferrii | 3 (0.1) |
| MSSA | 31 (0.9) | Elizabethkingia meningosepticum | 2 (0.1) |
| Other Enterococcus spp. | 31 (0.9) | Granulicatella adiacens | 2 (0.1) |
| Staphylococcus hominis | 28 (0.8) | Lactobacillus | 2 (0.1) |
| A. baumannii | 24 (0.7) | Streptococci agalactiae | 2 (0.1) |
| Enterococcus faecium | 23 (0.7) | Fusobacterium spp. | 2 (0.1) |
| MRSA | 18 (0.5) | Aerococcus urinae | 1 (0.03) |
| Enterococcus faecalis | 17 (0.5) | Streptococcus intermedius | 1 (0.03) |
| Other Klebsiella spp. | 15 (0.4) | Streptococcus sanguinis | 1 (0.03) |
| Enterobacter cloacae | 15 (0.4) | Actinomyces turicensis | 1 (0.03) |
| Pseudomonas spp. | 13 (0.4) | Providencia spp. | 1 (0.03) |
| Streptococcus pneumoniae | 12 (0.3) | Ralstonia mannitolilytica | 1 (0.03) |
| Staphylococcus capitis | 11 (0.3) | Rothia aeria | 1 (0.03) |
| Methicillin Susceptible- CoNS | 10 (0.3) | Legionella pneumophila | 1 (0.03) |
| Other Streptococcus spp. | 9 (0.3) | Clostridium perfringens | 1 (0.03) |
| Proteus mirabilis | 9 (0.3) | Comamonas testosteroni | 1 (0.03) |
| Bacillus non-anthracis | 9 (0.3) | Dolosigranulum pigrum | 1 (0.03) |
| Other Staphylococcus spp. | 8 (0.2) | Globicatella sanguinis | 1 (0.03) |
| Serratia marcescens | 8 (0.2) | Kocuria marina | 1 (0.03) |
| Staphylococcus haemolyticus | 8 (0.2) | Morganella morganii | 1 (0.03) |
| Stenotrophomonas maltophilia | 8 (0.2) | Moraxella osloensis | 1 (0.03) |
| Methicillin Resistant- CoNS | 7 (0.2) |
Abbreviations: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; C. pneumoniae, Chlamydia pneumoniae; M. pneumoniae, Mycoplasma pneumoniae; H. influenzae, Haemophilus influenzae; K. pneumoniae, Klebsiella pneumoniae; P. aeruginosa, Pseudomonas aeruginosa; S. pneumoniae, Streptococcus pneumoniae; M. catarrhalis, Moraxella catarrhalis; MRSA, methicillin-resistant Staphylococcus aureus; S. aureus, Staphylococcus aureus; A. baumannii, Acinetobacter baumannii; MSSA, methicillin-susceptible Staphylococcus aureus; CoNS, coagulase-negative staphylococci; E. coli, Escherichia coli; spp., species.
Fungal co-pathogens were reported in 16/72 (22.2%) studies, which is equal to only 3.2% of the reported co-infections. The most common fungal organisms were Aspergillus spp. (n = 68), Aspergillus fumigatus (n = 43), Other Candida spp. (n = 29), Candida albicans (n = 25) and Aspergillus flavus (n = 10) (Table 3).
Table 3.
Proportion of all identified SARS-CoV-2 fungal co-infections (N = 192).
| Fungal Pathogen Type | Identified Number (%) |
|---|---|
| Aspergillus spp. | 68 (35.4) |
| Aspergillus fumigatus | 43 (22.4) |
| Other Candida spp. | 29 (15.1) |
| Candida albicans | 25 (13) |
| Aspergillus flavus | 10 (5.2) |
| Mucor | 6 (3.1) |
| Candida glabrata | 5 (2.6) |
| Aspergillus niger | 4 (2.1) |
| Aspergillus terreus | 1 (0.5) |
| Cryptococcus | 1 (0.5) |
Abbreviations: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; spp., species.
Respiratory viral co-pathogens were reported in 44/72 (61.1%) studies, representing about 39.5% of the reported co-infections. The most common respiratory viruses were EBV (n = 644), HHV6 (n = 574), Influenza A virus (n = 355), HMPV (n = 328), and Adenovirus (n = 144) (Table 4).
Table 4.
Proportion of all identified SARS-CoV-2 respiratory viral co-infections (N = 2392).
| Respiratory Viral Pathogen Type | Identified Number (%) |
|---|---|
| EBV | 644 (26.9) |
| HHV6 | 574 (24) |
| Influenza A virus | 355 (14.8) |
| HMPV | 328 (13.7) |
| Adenovirus | 144 (6) |
| Influenza B virus | 68 (2.8) |
| Rhinovirus/Enterovirus | 68 (2.8) |
| RSV | 52 (2.2) |
| Parainfluenza [1, 2, 3 and 4] virus | 29 (1.2) |
| HcoV-OC43 | 11 (0.5) |
| Rhinovirus | 22 (0.9) |
| Influenza virus (H1N1) | 18 (0.8) |
| HcoV-HKU1 | 16 (0.7) |
| HcoV-NL63 | 13 (0.5) |
| Bocavirus | 10 (0.4) |
| HSV | 10 (0.4) |
| HcoV-229E | 9 (0.4) |
| CMV | 8 (0.3) |
| MERS-CoV | 8 (0.3) |
| Enterovirus | 1 (0.04) |
| Rotavirus | 1 (0.04) |
| Coxsackie virus | 1 (0.04) |
| Human Coronavirus 229E | 1 (0.04) |
| Herpes virus 5 | 1 (0.04) |
Abbreviations: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; RSV, respiratory syncytial virus; EBV, Epstein–Barr virus; HcoV-HKU1, human coronavirus HKU1; CMV, cytomegalovirus; HSV, herpes simplex virus; HHV6, human herpes virus 6; HMPV, human metapneumovirus.
4. Discussion
In this large systematic review and meta-analysis, we included 31,953 patients with laboratory-confirmed SARS-CoV-2 from 72 observational studies in order to estimate the prevalence of coinfections with bacterial, fungal, and respiratory viral pathogens. This study showed the following microbial coinfection prevalences: bacterial (15.9%, 95% CI 13.6–18.2); fungal (3.7%, 95% CI 2.6–4.8); and respiratory viral (6.6%, 95% CI 5.5–7.6) coinfections. Bacterial and fungal coinfections were more common in ICU patients ((22.2%%, 95% CI 16.1–28.4) and (9.6%, 95% CI 6.8–12.4), respectively) than mixed ICU and non-ICU patients, as expected. However, respiratory viral co-infection rate in SARS-CoV-2 patients was identical in both groups (6.6%, 95% CI 0.0–11.3). Nevertheless, the included studies in this meta-analysis are case series and cohort studies and we did not identify any randomized controlled trials addressing this issue. In addition, the included studies comprised only admitted patients, which may skew the findings and should not be generalized to all SARS-COV-2 patients. Non-admitted COVID-19 patients were not represented in these studies and thus the exact prevalence of coinfections could not be calculated for all SARS-CoV-2 infected patients [83,84,85]. The findings in this meta-analysis showed different results from previous systematic meta-analyses that evaluated coinfections among COVID-19 patients [71,72,73]. We reported a higher prevalence of coinfections in hospitalized SARS-CoV-2 patients. The current meta-analysis is more comprehensive and included a total of 71 studies [2,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,80] and one abstract [3], including a total of 31,953 patients. The inclusion of 18 recently published studies [2,3,5,6,7,8,9,10,12,13,14,22,24,27,41,62,64,65] contributed to the refinement of the estimate of the pooled prevalence of pathogens contributing to coinfections in SARS-CoV-2 patients.
In this meta-analysis, bacterial coinfection was more prevalent than fungal and other respiratory viruses. This finding may reflect high rates of antimicrobial use for admitted patients with SARS-CoV-2 infection to treat documented or presumed bacterial co-infections. Thus, it is important to study the occurrence, type, and intended antimicrobial agent use in SARS-COV-2 patients in order to develop additional strategies for the optimal use of antimicrobial agents in this population. As expected, bacterial, fungal, and other respiratory viral co-infections in SARS-CoV-2 patients were more frequent in ICUs compared with non-ICU locations [2,20,28,57], a finding which has previously been described in systematic reviews [71,72] and may reflect the epicenter role of ICUs in both infections and antimicrobial resistance. One of the reasons for the increase in infection rate in ICUs could be due to the simultaneous infection of the virus and bacterium. Viruses can facilitate the attachment and colonization of the bacteria in the respiratory tract, which is certainly no exception for SARS-CoV-2 [86]. Nevertheless, other factors such as ICU type, used equipment rate, admission or discharge criteria, high workload or nurse ratio, etc. can also affect the quality of care and the rate of ICU-acquired, healthcare-associated infections [87,88]. With observed strains currently being placed on healthcare systems during the upstroke of the SARS-CoV-2 pandemic, guidelines must focus on the maintenance of good knowledge and compliance of infection prevention and control [89], antimicrobial stewardship [90], and robust surveillance for healthcare-associated infections and antimicrobial resistance [91,92].
The most common method used to detect co-infections in the studies included in this review was RT-PCR tests for respiratory samples. The choice of diagnostic test for pathogens depends in part upon test availability and how soon the results are needed. If available, molecular assays (RT-PCR or, alternatively, a rapid molecular assay) are preferred over antigen detection tests (e.g., direct and indirect immunofluorescence assays) because molecular tests are the most sensitive [93]. Nevertheless, positive RT-PCR tests might indicate recently resolved infection or colonization [94,95]. In addition, many studies evaluated serological (antibodies) tests with this method detecting co-infections in SARS-CoV-2 patients. Application of serologic laboratory technique for co-pathogens detection across all studies was likely to reveal an even higher overall co-infection proportion than found in our study. Consecutively, it is possible that positive serology indicated recent and not acute infection in included patients [96]. Serologic testing is useful primarily for research purposes and antibody-based tests might produce false negative results during the window period. It is worthwhile to mention that administration of broad-spectrum antimicrobials to a large percentage of the patients included in this review might relatively have lowered the sensitivity of microbial culture methods, which could have resulted in underestimation of the true numbers of co-infections.
Specific co-infecting pathogens in SARS-CoV-2 patients were identified in this study from the 72 included studies. In line with the previous systematic reviews and meta-analyses [71,72], M. pneumoniae, K. pneumoniae, and H. influenzae were among the predominant co-pathogens. However, in this meta-analysis, S. aureus was the most common bacterial pathogens co-infecting SARS-CoV-2 patients. However, this finding needs to be carefully interpreted, as 85.6% of all S. aureus co-pathogens in our review were reported by one study [58]. S. aureus infections are a known complication of other viral pandemics, such as the Spanish flu and the H1N1 influenza pandemic [97,98]. S. aureus is known to act synergistically in SARS-CoV-2 patients, increasing mortality and severity of disease [38,99]. The proposed mechanisms of viral-induced S. aureus co-infections include viral modification of airway structures and increased adherence of the organism to respiratory mucosa, as well as initiation of immune-suppressive responses [22,100,101]. Further investigations are necessary to confirm an association between SARS-CoV-2 infection and susceptibility to S. aureus coinfections.
It was noted that male patients with SARS-CoV-2 were more likely to have coinfections than female [13]. However, patients with pneumococcal pneumoniae and SARS-CoV-2 were mostly females [24]. Older age appears to be the major risk factor associated with coinfections with bacteria and respiratory viruses [12,38,43,58,62] and fungi [39]. This might be attributed mainly to the differences in the inclusion criteria and the population age groups included in the studies, or it could be explained by the gender-based biological differences in the host immune response to COVID-19 infection [102]. The age-dependent defects in T-cell and B-cell function and the excess production of type 2 cytokines could lead to a deficiency in control of viral replication and more prolonged proinflammatory responses, potentially leading to poorer outcomes [103]. Yet, SARS-CoV-2 patients of any age may develop such coinfections and experience severe disease, especially in those with comorbidities, even in young people [4,53], children [27,49], and infants [40].
A few underlying comorbidities were associated with increased risk of coinfections, and these included obesity [8,12,38], cancer, hepatitis, and kidney disease [12,43]. Laboratory abnormalities that have been described in SARS-CoV-2 patients with bacterial and respiratory viral coinfections were high procalcitonin [47,50,64,80], d-dimer [9], and monocytes [31]; and low neutrophils [31]. Some conclusions could be drawn from available data as to whether patients who have a concurrent bacterial, fungal, and/or respiratory viral infection have a worse prognosis than those in whom SARS-CoV-2 is the only detected pathogen. Mortality in SARS-CoV-2 patients was increased due to bacterial [2,6,14,21], fungal [2,17,20,21], or respiratory viral [20] co-infections compared to SARS-CoV-2 patients with no co-infections. Few studies observed no increase in mortality in COVID-19 patients compared to those who did not have bacterial [3,22,24,35,66], fungal [3,22], or other respiratory viral [66] coinfections. Clinical presentation, laboratory results, radiological findings, and outcome are likely to differ between SARS-CoV-2 positive patients with and without co-infections. Bacterial coinfection increased SARS-CoV-2 patients’ hospital length of stay [18,50], need for ventilatory support [6,28], ARDS [28], shock [28], multi-organ injury [23,32], and caused more severe COVID-19 disease [2,21,28,33,34,53,68]. Two studies reported conflicting results on the role of bacterial [24,36] or respiratory viral [36] coinfection in relation to increasing length of hospital stay or ICU admission [22,24,35]. It was shown that the patterns of SARS-CoV-2 symptoms and clinical outcomes were not different in the bacterial [27] and respiratory viral [10,11,27,66,70] co-infected patients. The severity and time of SARS-CoV-2 disease clearance were not different in patients with respiratory viral co-infections [19,36].
The data on the timing of the occurrence of co-infection was variable. The occurrence of co-infections has a median time of 4–11.5 days (IQR 2–42) of ICU admission [2,17,42]. Bacterial co-infection was infrequent within 2–4 days of hospital admission [22,26]. Nonetheless, considering the high number and severity of bacterial co-infections previously reported in patients with SARS-CoV-2, initiation of antibiotic therapy for all hospitalized patients with COVID-19 is recommended [7]. The approach of administering empiric antibiotic therapy solely to patients who were admitted for SARS-CoV-2 and who presented with a chest X-ray suggestive of bacterial infection, have a need for direct ICU admission, or are severely immunocompromised should be reconsidered. When bacterial co-infection in SARS-CoV-2 patients is suspected, an antibiotic approach with optimal S. aureus coverage, such as ceftaroline, ceftriaxone, or cefazolin plus levofloxacin, is recommended in areas with methicillin-sensitive S. aureus prevalence [104].
Limitations
The main limitation of this meta-analysis is that included studies were observational with no randomized controlled trials; and there was no standardized microbiologic testing at specified intervals. In interpreting funnel plots, the different possible reasons for funnel plot asymmetry should be distinguished. Possible sources of asymmetry in funnel plots might be the wide differences between the included populations in the different studies, publication bias and selective outcome and/or analysis reporting, poor methodological design and inadequate analysis, or asymmetry might have occurred by chance. Furthermore, the analysis was limited to the English literature and thus may miss other studies published in other languages.
5. Conclusions
Bacterial co-infection is relatively high in hospitalized patients with SARS-CoV-2, with little evidence of S. aureus having a major role. Empiric antibiotic therapy should be considered in SARS-CoV-2 patients who present with a chest X-ray suggestive of bacterial infection, the need for direct ICU admission, or a severely immunocompromised condition. Knowledge of the prevalence and type of co-infections in SARS-CoV-2 patients may have diagnostic and management implications.
Acknowledgments
We would like to thank Hani N. Mufti for precious guidance and support to create the forest and funnel plots using RStudio. We would also like to thank the reviewers for very helpful and valuable comments and suggestions for improving the paper.
Author Contributions
S.A., A.A.M., A.A.R., Z.A.A. and J.A.A.-T. contributed equally to this study. S.A., A.A.M., Z.A.A., and A.A.R. were the core team leading the systematic review. S.A., A.A.M., and J.A.A.-T. identified and selected the studies. A.M.A. (Abeer M. Alshawi), S.A.A., G.Y.A., and A.A.R. did the quality assessment of the studies. S.A., A.A.M., M.S.A., and A.A.A. collected the data. S.A., A.H.B.S., A.M.A. (Abdullah M. Alotaibi), and A.A.-O. analyzed the data. S.A., A.A.M., Z.A.A., A.A.R., and J.A.A.-T. drafted the manuscript. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data are available upon request. Please contact author for data requests.
Conflicts of Interest
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.World Health Organization WHO Coronavirus (COVID-19) Dashboard. [(accessed on 5 April 2021)];2021 Available online: https://covid19.who.int.
- 2.Bardi T., Pintado V., Gomez-Rojo M., Escudero-Sanchez R., Lopez A.A., Diez-Remesal Y., Castro N.M., Ruiz-Garbajosa P., Pestaña D. Nosocomial infections associated to COVID-19 in the intensive care unit: Clinical characteristics and outcome. Eur. J. Clin. Microbiol. Infect. Dis. 2021;40:495–502. doi: 10.1007/s10096-020-04142-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.May A., Swetenham N., Pandey M., Taylor V., Hughes H., Underwood J. P197 Bacterial and fungal respiratory co-infection among patients admitted to ICU with COVID-19: A retrospective cohort study in a UK hospital. BMJ. 2021;76:A196–A197. [Google Scholar]
- 4.Zhu X., Ge Y., Wu T., Zhao K., Chen Y., Wu B., Zhu F., Zhu B., Cui L. Co-infection with respiratory pathogens among COVID-2019 cases. Virus Res. 2020;285:198005. doi: 10.1016/j.virusres.2020.198005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Calcagno A., Ghisetti V., Burdino E., Trunfio M., Allice T., Boglione L., Bonora S., Di Perri G. Co-infection with other respiratory pathogens in COVID-19 patients. Clin. Microbiol. Infect. 2021;27:297–298. doi: 10.1016/j.cmi.2020.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.De Francesco M.A., Poiesi C., Gargiulo F., Bonfanti C., Pollara P., Fiorentini S., Caccuri F., Carta V., Mangeri L., Pellizzeri S. Co-infection of Chlamydia pneumoniae and Mycoplasma pneumoniae with SARS-CoV-2 is associated with more severe features. J. Infect. 2021;8 doi: 10.1016/j.jinf.2021.01.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Garcia-Vidal C., Sanjuan G., Moreno-García E., Puerta-Alcalde P., Garcia-Pouton N., Chumbita M., Fernandez-Pittol M., Pitart C., Inciarte A., Bodro M. Incidence of co-infections and superinfections in hospitalized patients with COVID-19: A retrospective cohort study. Clin. Microbiol. Infect. 2021;27:83–88. doi: 10.1016/j.cmi.2020.07.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Elhazmi A., Al-Tawfiq J.A., Sallam H., Al-Omari A., Alhumaid S., Mady A., Al Mutair A. Severe respiratory syndrome Coronavirus 2 (SARS-CoV-2) and middle east respiratory syndrome Coronavirus (MERS-CoV) coinfection: A unique case series. Travel Med. Infect. Dis. 2021;41:102026. doi: 10.1016/j.tmaid.2021.102026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Allou N., Larsen K., Dubernet A., Traversier N., Masse L., Foch E., Bruneau L., Maillot A., André M., Lagrange-Xelot M. Co-infection in patients with hypoxemic pneumonia due to COVID-19 in Reunion Island. Medicine. 2021;100:e24524. doi: 10.1097/MD.0000000000024524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cheng Y., Ma J., Wang H., Wang X., Hu Z., Li H., Zhang H., Liu X. Co-infection of influenza A virus and SARS-CoV-2: A retrospective cohort study. J. Med. Virol. 2021;93 doi: 10.1002/jmv.26817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cuadrado-Payán E., Montagud-Marrahi E., Torres-Elorza M., Bodro M., Blasco M., Poch E., Soriano A., Piñeiro G.J. SARS-CoV-2 and influenza virus co-infection. Lancet. 2020;395:e84. doi: 10.1016/S0140-6736(20)31052-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hashemi S.A., Safamanesh S., Ghasemzadeh-Moghaddam H., Ghafouri M., Azimian A. High prevalence of SARS-CoV-2 and influenza A virus (H1N1) coinfection in dead patients in northeastern Iran. J. Med. Virol. 2021;93:1008–1012. doi: 10.1002/jmv.26364. [DOI] [PubMed] [Google Scholar]
- 13.Schirmer P., Lucero-Obusan C., Sharma A., Sohoni P., Oda G., Holodniy M. Respiratory co-infections with COVID-19 in the Veterans Health Administration, 2020. Diagn. Microbiol. Infect. Dis. 2021;100:115312. doi: 10.1016/j.diagmicrobio.2021.115312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Amin D., McKitish K., Shah P.S. Association of mortality and recent Mycoplasma pneumoniae infection in COVID-19 patients. J. Med. Virol. 2021;93:1180–1183. doi: 10.1002/jmv.26467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Arentz M., Yim E., Klaff L., Lokhandwala S., Riedo F.X., Chong M., Lee M. Characteristics and outcomes of 21 critically ill patients with COVID-19 in Washington state. JAMA. 2020;323:1612–1614. doi: 10.1001/jama.2020.4326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Barrasa H., Rello J., Tejada S., Martín A., Balziskueta G., Vinuesa C., Fernández-Miret B., Villagra A., Vallejo A., San Sebastián A. SARS-CoV-2 in Spanish intensive care units: Early experience with 15-day survival in Vitoria. Anaesth. Crit. Care Pain Med. 2020;39:553–561. doi: 10.1016/j.accpm.2020.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bartoletti M., Pascale R., Cricca M., Rinaldi M., Maccaro A., Bussini L., Fornaro G., Tonetti T., Pizzilli G., Francalanci E. Epidemiology of invasive pulmonary aspergillosis among COVID-19 intubated patients: A prospective study. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa1065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cheng L.S., Chau S.K., Tso E.Y., Tsang S.W., Li I.Y., Wong B.K., Fung K.S. Bacterial co-infections and antibiotic prescribing practice in adults with COVID-19: Experience from a single hospital cluster. Ther. Adv. Infect. Dis. 2020;7 doi: 10.1177/2049936120978095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ding Q., Lu P., Fan Y., Xia Y., Liu M. The clinical characteristics of pneumonia patients coinfected with 2019 novel coronavirus and influenza virus in Wuhan, China. J. Med. Virol. 2020;92:1549–1555. doi: 10.1002/jmv.25781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Koehler P., Cornely O.A., Böttiger B.W., Dusse F., Eichenauer D.A., Fuchs F., Hallek M., Jung N., Klein F., Persigehl T. COVID-19 associated pulmonary aspergillosis. Mycoses. 2020;63:528–534. doi: 10.1111/myc.13096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ramadan H.K.-A., Mahmoud M.A., Aburahma M.Z., Elkhawaga A.A., El-Mokhtar M.A., Sayed I.M., Hosni A., Hassany S.M., Medhat M.A. Predictors of severity and co-infection resistance profile in COVID-19 patients: First report from upper Egypt. Infect. Drug Res. 2020;13:3409. doi: 10.2147/IDR.S272605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wang L., Amin A.K., Khanna P., Aali A., McGregor A., Bassett P., Gopal Rao G. An observational cohort study of bacterial co-infection and implications for empirical antibiotic therapy in patients presenting with COVID-19 to hospitals in north west London. J. Antimicrob. Chemother. 2021;76:796–803. doi: 10.1093/jac/dkaa475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Alanio A., Dellière S., Fodil S., Bretagne S., Mégarbane B. Prevalence of putative invasive pulmonary aspergillosis in critically ill patients with COVID-19. Lancet Respir. Med. 2020;8:e48–e49. doi: 10.1016/S2213-2600(20)30237-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Anton-Vazquez V., Clivillé R. Streptococcus pneumoniae coinfection in hospitalised patients with COVID-19. Eur. J. Clin. Microbiol. Infect. Dis. 2021;40:1353–1355. doi: 10.1007/s10096-021-04166-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chen T., Dai Z., Mo P., Li X., Ma Z., Song S., Chen X., Luo M., Liang K., Gao S. Clinical characteristics and outcomes of older patients with coronavirus disease 2019 (COVID-19) in Wuhan, China: A single-centered, retrospective study. J. Gerontol. Ser. A. 2020;75:1788–1795. doi: 10.1093/gerona/glaa089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hughes S., Troise O., Donaldson H., Mughal N., Moore L.S. Bacterial and fungal coinfection among hospitalized patients with COVID-19: A retrospective cohort study in a UK secondary-care setting. Clin. Microbiol. Infect. 2020;26:1395–1399. doi: 10.1016/j.cmi.2020.06.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Li Y., Wang H., Wang F., Lu X., Du H., Xu J., Han F., Zhang L., Zhang M. Co-infections of SARS-CoV-2 with multiple common respiratory pathogens in infected children: A retrospective study. Medicine. 2021;100 doi: 10.1007/s11427-020-1668-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Z., Chen Z., Chen L.D., Zhan Y.Q., Li S.Q., Cheng J., Zhu A., Chen L.Y., Zhong N.S., Li S.Y. Coinfection with SARS-CoV-2 and other respiratory pathogens in patients with COVID-19 in Guangzhou, China. J. Med. Virol. 2020;92:2381–2383. doi: 10.1002/jmv.26073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lin D., Liu L., Zhang M., Hu Y., Yang Q., Guo J., Guo Y., Dai Y., Xu Y., Cai Y. Co-infections of SARS-CoV-2 with multiple common respiratory pathogens in infected patients. Sci. China Life Sci. 2020;63:606–609. doi: 10.1007/s11427-020-1668-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Liu H., Liu F., Li J., Zhang T., Wang D., Lan W. Clinical and CT imaging features of the COVID-19 pneumonia: Focus on pregnant women and children. J. Infect. 2020;80:e7–e13. doi: 10.1016/j.jinf.2020.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Liu L., Lei X., Xiao X., Yang J., Li J., Ji M., Du W., Tan H., Zhu J., Li B. Epidemiological and clinical characteristics of patients with coronavirus disease-2019 in Shiyan City, China. Front. Cell. Infect. Microbiol. 2020;10 doi: 10.3389/fcimb.2020.00284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ma S., Lai X., Chen Z., Tu S., Qin K. Clinical characteristics of critically ill patients co-infected with SARS-CoV-2 and the influenza virus in Wuhan, China. Int. J. Infect. Dis. 2020;96:683–687. doi: 10.1016/j.ijid.2020.05.068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mannheim J., Gretsch S., Layden J.E., Fricchione M.J. Characteristics of hospitalized pediatric coronavirus disease 2019 cases in Chicago, Illinois, March–April 2020. J. Pediatr. Infect. Dis. Soc. 2020;9:519–522. doi: 10.1093/jpids/piaa070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nasir N., Farooqi J., Mahmood S.F., Jabeen K. COVID-19-associated pulmonary aspergillosis (CAPA) in patients admitted with severe COVID-19 pneumonia: An observational study from Pakistan. Mycoses. 2020;63:766–770. doi: 10.1111/myc.13135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Oliva A., Siccardi G., Migliarini A., Cancelli F., Carnevalini M., D’Andria M., Attilia I., Danese V.C., Cecchetti V., Romiti R. Co-infection of SARS-CoV-2 with Chlamydia or Mycoplasma pneumoniae: A case series and review of the literature. Infection. 2020;48:871–877. doi: 10.1007/s15010-020-01483-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Peng H., Gao P., Xu Q., Liu M., Peng J., Wang Y., Xu H. Coronavirus disease 2019 in children: Characteristics, antimicrobial treatment, and outcomes. J. Med. Virol. 2020;128:104425. doi: 10.1016/j.jcv.2020.104425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pongpirul W.A., Mott J.A., Woodring J.V., Uyeki T.M., MacArthur J.R., Vachiraphan A., Suwanvattana P., Uttayamakul S., Chunsuttiwat S., Chotpitayasunondh T. Clinical characteristics of patients hospitalized with coronavirus disease, Thailand. Emerg. Infect. Dis. 2020;26:1580–1585. doi: 10.3201/eid2607.200598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Richardson S., Hirsch J.S., Narasimhan M., Crawford J.M., McGinn T., Davidson K.W., Barnaby D.P., Becker L.B., Chelico J.D., Cohen S.L. Presenting characteristics, comorbidities, and outcomes among 5700 patients hospitalized with COVID-19 in the New York City area. JAMA. 2020;323:2052–2059. doi: 10.1001/jama.2020.6775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rutsaert L., Steinfort N., Van Hunsel T., Bomans P., Naesens R., Mertes H., Dits H., Van Regenmortel N. COVID-19-associated invasive pulmonary aspergillosis. Ann. Intensiv. Care. 2020;10:71. doi: 10.1186/s13613-020-00686-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sun D., Chen X., Li H., Lu X.-X., Xiao H., Zhang F.-R., Liu Z.-S. SARS-CoV-2 infection in infants under 1 year of age in Wuhan City, China. World J. Pediatr. 2020;16:260–266. doi: 10.1007/s12519-020-00368-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tagarro A., Epalza C., Santos M., Sanz-Santaeufemia F.J., Otheo E., Moraleda C., Calvo C. Screening and severity of coronavirus disease 2019 (COVID-19) in children in Madrid, Spain. JAMA Pediatr. 2021;175:316–317. doi: 10.1001/jamapediatrics.2020.1346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Van Arkel A.L., Rijpstra T.A., Belderbos H.N., Van Wijngaarden P., Verweij P.E., Bentvelsen R.G. COVID-19–associated pulmonary aspergillosis. Am. J. Respir. Crit. Care Med. 2020;202:132–135. doi: 10.1164/rccm.202004-1038LE. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wang R., Pan M., Zhang X., Han M., Fan X., Zhao F., Miao M., Xu J., Guan M., Deng X. Epidemiological and clinical features of 125 hospitalized patients with COVID-19 in Fuyang, Anhui, China. Int. J. Infect. Dis. 2020;95:421–428. doi: 10.1016/j.ijid.2020.03.070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wang Y., Liu Y., Liu L., Wang X., Luo N., Li L. Clinical outcomes in 55 patients with severe acute respiratory syndrome coronavirus 2 who were asymptomatic at hospital admission in Shenzhen, China. J. Infect. Dis. 2020;221:1770–1774. doi: 10.1093/infdis/jiaa119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wang Z., Yang B., Li Q., Wen L., Zhang R. Clinical features of 69 cases with coronavirus disease 2019 in Wuhan, China. Clin. Infect. Dis. 2020;71:769–777. doi: 10.1093/cid/ciaa272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wu Q., Xing Y., Shi L., Li W., Gao Y., Pan S., Wang Y., Wang W., Xing Q. Coinfection and other clinical characteristics of COVID-19 in children. Pediatrics. 2020;146 doi: 10.1542/peds.2020-0961. [DOI] [PubMed] [Google Scholar]
- 47.Xia W., Shao J., Guo Y., Peng X., Li Z., Hu D. Clinical and CT features in pediatric patients with COVID-19 infection: Different points from adults. Pediatr. Pulmonol. 2020;55:1169–1174. doi: 10.1002/ppul.24718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yang X., Yu Y., Xu J., Shu H., Liu H., Wu Y., Zhang L., Yu Z., Fang M., Yu T. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: A single-centered, retrospective, observational study. Lancet Respir. Med. 2020;8:475–481. doi: 10.1016/S2213-2600(20)30079-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zheng F., Liao C., Fan Q.-H., Chen H., Zhao X., Xie Z., Li X., Chen C., Lu X., Liu Z. Clinical characteristics of children with coronavirus disease 2019 in Hubei, China. Curr. Med. Sci. 2020:275–280. doi: 10.1007/s11596-020-2172-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zhang J., Dong X., Cao Y., Yuan Y., Yang Y., Yan Y., Akdis C.A., Gao Y. Clinical characteristics of 140 patients infected with SARS-CoV-2 in Wuhan, China. Allergy. 2020;75:1730–1741. doi: 10.1111/all.14238. [DOI] [PubMed] [Google Scholar]
- 51.Contou D., Claudinon A., Pajot O., Micaëlo M., Flandre P.L., Dubert M., Cally R., Logre E., Fraissé M., Mentec H. Bacterial and viral co-infections in patients with severe SARS-CoV-2 pneumonia admitted to a French ICU. Ann. Intensive Care. 2020;10:119. doi: 10.1186/s13613-020-00736-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Gayam V., Konala V.M., Naramala S., Garlapati P.R., Merghani M.A., Regmi N., Balla M., Adapa S. Presenting characteristics, comorbidities, and outcomes of patients coinfected with COVID-19 and Mycoplasma pneumoniae in the USA. J. Med. Virol. 2020;92:2181–2187. doi: 10.1002/jmv.26026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hazra A., Collison M., Pisano J., Kumar M., Oehler C., Ridgway J.P. Coinfections with SARS-CoV-2 and other respiratory pathogens. Infect. Control Hosp. Epidemiol. 2020;41:1228–1229. doi: 10.1017/ice.2020.322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Karami Z., Knoop B.T., Dofferhoff A.S., Blaauw M.J., Janssen N.A., van Apeldoorn M., Kerckhoffs A.P., van de Maat J.S., Hoogerwerf J.J., Ten Oever J. Few bacterial co-infections but frequent empiric antibiotic use in the early phase of hospitalized patients with COVID-19: Results from a multicentre retrospective cohort study in The Netherlands. Infect. Dis. 2020 doi: 10.1080/23744235.2020.1839672. [DOI] [PubMed] [Google Scholar]
- 55.Kim D., Quinn J., Pinsky B., Shah N.H., Brown I. Rates of co-infection between SARS-CoV-2 and other respiratory pathogens. JAMA. 2020;323:2085–2086. doi: 10.1001/jama.2020.6266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kreitmann L., Monard C., Dauwalder O., Simon M., Argaud L. Early bacterial co-infection in ARDS related to COVID-19. Intensive Care Med. 2020;46:1787–1789. doi: 10.1007/s00134-020-06165-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lehmann C.J., Pho M.T., Pitrak D., Ridgway J.P., Pettit N.N. Community acquired co-infection in COVID-19: A retrospective observational experience. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Massey B.W., Jayathilake K., Meltzer H.Y. Respiratory microbial co-infection with SARS-CoV-2. Front. Microbiol. 2020;11 doi: 10.3389/fmicb.2020.02079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Nowak M.D., Sordillo E.M., Gitman M.R., Paniz Mondolfi A.E. Coinfection in SARS-CoV-2 infected patients: Where are influenza virus and rhinovirus/enterovirus? J. Med. Virol. 2020;92:1699–1700. doi: 10.1002/jmv.25953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ozaras R., Cirpin R., Duran A., Duman H., Arslan O., Bakcan Y., Kaya M., Mutlu H., Isayeva L., Kebanlı F. Influenza and COVID-19 coinfection: Report of six cases and review of the literature. J. Med. Virol. 2020;92:2657–2665. doi: 10.1002/jmv.26125. [DOI] [PubMed] [Google Scholar]
- 61.Sepulveda J., Westblade L.F., Whittier S., Satlin M.J., Greendyke W.G., Aaron J.G., Zucker J., Dietz D., Sobieszczyk M., Choi J.J. Bacteremia and blood culture utilization during COVID-19 surge in New York City. J. Med. Virol. 2020;58 doi: 10.1128/JCM.00875-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Singh V., Upadhyay P., Reddy J., Granger J. SARS-CoV-2 respiratory co-infections: Incidence of viral and bacterial co-pathogens. Int. J. Infect. Dis. 2021 doi: 10.1016/j.ijid.2021.02.087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Song W., Jia X., Zhang X., Ling Y., Yi Z. Co-infection in COVID-19, a cohort study. J. Infect. 2020 doi: 10.1016/j.jinf.2020.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Tang M.-L., Li Y.-Q., Chen X., Lin H., Jiang Z.-C., Gu D.-L., Chen X., Tang C.-X., Xie Z.-Q. Co-infection with common respiratory pathogens and SARS-CoV-2 in patients with COVID-19 pneumonia and laboratory biochemistry findings: A retrospective cross-sectional study of 78 patients from a single center in China. Int. Med. J. Exp. Clin. Res. 2021;27:e929783-1. doi: 10.12659/MSM.929783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Thelen J.M., Buenen A.N., van Apeldoorn M., Wertheim H.F., Hermans M.H., Wever P.C. Community-acquired bacteraemia in COVID-19 in comparison to influenza A and influenza B: A retrospective cohort study. BMC Infect. Dis. 2021;21:199. doi: 10.1186/s12879-021-05902-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Wee L.E., Ko K.K.K., Ho W.Q., Kwek G.T.C., Tan T.T., Wijaya L. Community-acquired viral respiratory infections amongst hospitalized inpatients during a COVID-19 outbreak in Singapore: Co-infection and clinical outcomes. J. Clin. Virol. 2020;128:104436. doi: 10.1016/j.jcv.2020.104436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wu C., Chen X., Cai Y., Zhou X., Xu S., Huang H., Zhang L., Zhou X., Du C., Zhang Y. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern. Med. 2020;180:934–943. doi: 10.1001/jamainternmed.2020.0994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Yue H., Zhang M., Xing L., Wang K., Rao X., Liu H., Tian J., Zhou P., Deng Y., Shang J. The epidemiology and clinical characteristics of co-infection of SARS-CoV-2 and influenza viruses in patients during COVID-19 outbreak. J. Med. Virol. 2020;92:2870–2873. doi: 10.1002/jmv.26163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Zhao D., Yao F., Wang L., Zheng L., Gao Y., Ye J., Guo F., Zhao H., Gao R. A comparative study on the clinical features of coronavirus 2019 (COVID-19) pneumonia with other pneumonias. Clin. Infect. Dis. 2020;71:756–761. doi: 10.1093/cid/ciaa247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Zheng X., Wang H., Su Z., Li W., Yang D., Deng F., Chen J. Co-infection of SARS-CoV-2 and Influenza virus in Early Stage of the COVID-19 Epidemic in Wuhan, China. J. Infect. 2020 doi: 10.1016/j.jinf.2020.05.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Lansbury L., Lim B., Baskaran V., Lim W.S. Co-infections in people with COVID-19: A systematic review and meta-analysis. J. Infect. 2020;81:266–275. doi: 10.1016/j.jinf.2020.05.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Langford B.J., So M., Raybardhan S., Leung V., Westwood D., MacFadden D.R., Soucy J.-P.R., Daneman N. Bacterial co-infection and secondary infection in patients with COVID-19: A living rapid review and meta-analysis. Clin. Microbiol. Infect. 2020 doi: 10.1016/j.cmi.2020.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Davis B., Rothrock A.N., Swetland S., Andris H., Davis P., Rothrock S.G. Viral and atypical respiratory co-infections in COVID-19: A systematic review and meta-analysis. J. Am. Coll. Emerg. Phys. Open. 2020;1:533–548. doi: 10.1002/emp2.12128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Moher D., Liberati A., Tetzlaff J., Altman D.G., Group P. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009;6:e1000097. doi: 10.1371/journal.pmed.1000097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Wells G., Shea B., O’Connell D., Peterson J., Welch V., Losos M., Tugwell P. The Newcastle-Ottawa Scale (NOS) for Assessing the Quality of Nonrandomized Studies in Meta-Analyses. Ottawa Health Research Institute; Ottawa, ON, Canada: 2015. [Google Scholar]
- 76.DerSimonian R., Kacker R. Random-effects model for meta-analysis of clinical trials: An update. Contemp. Clin. Trials. 2007;28:105–114. doi: 10.1016/j.cct.2006.04.004. [DOI] [PubMed] [Google Scholar]
- 77.Higgins J.P., Thompson S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 2002;21:1539–1558. doi: 10.1002/sim.1186. [DOI] [PubMed] [Google Scholar]
- 78.Higgins J.P., Thompson S.G., Deeks J.J., Altman D.G. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–560. doi: 10.1136/bmj.327.7414.557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Egger M., Smith G.D., Schneider M., Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315:629–634. doi: 10.1136/bmj.315.7109.629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Chen N., Zhou M., Dong X., Qu J., Gong F., Han Y., Qiu Y., Wang J., Liu Y., Wei Y. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet. 2020;395:507–513. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Mo P., Xing Y., Xiao Y., Deng L., Zhao Q., Wang H., Xiong Y., Cheng Z., Gao S., Liang K. Clinical characteristics of refractory COVID-19 pneumonia in Wuhan, China. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Zha L., Shen J., Tefsen B., Wang Y., Lu W., Xu Q. Clinical features and outcomes of adult COVID-19 patients co-infected with Mycoplasma pneumoniae. J. Infect. 2020 doi: 10.1016/j.jinf.2020.07.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Al Mutair A., Alhumaid S., Alhuqbani W.N., Zaidi A.R.Z., Alkoraisi S., Al-Subaie M.F., AlHindi A.M., Abogosh A.K., Alrasheed A.K., Alsharafi A.A. Clinical, epidemiological, and laboratory characteristics of mild-to-moderate COVID-19 patients in Saudi Arabia: An observational cohort study. Eur. J. Med. Res. 2020;25 doi: 10.1186/s40001-020-00462-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Alhumaid S., Al Mutair A., Al Alawi Z., Al Salman K., Al Dossary N., Omar A., Alismail M., Al Ghazal A.M., Jubarah M.B., Al Shaikh H. Clinical features and prognostic factors of intensive and non-intensive 1014 COVID-19 patients: An experience cohort from Alahsa, Saudi Arabia. Eur. J. Med. Res. 2021;26:47. doi: 10.1186/s40001-021-00517-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Al-Omari A., Alhuqbani W.N., Zaidi A.R.Z., Al-Subaie M.F., AlHindi A.M., Abogosh A.K., Alrasheed A.K., Alsharafi A.A., Alhuqbani M.N., Salih S. Clinical characteristics of non-intensive care unit COVID-19 patients in Saudi Arabia: A descriptive cross-sectional study. J. Infect. Public Health. 2020;13:1639–1644. doi: 10.1016/j.jiph.2020.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sharifipour E., Shams S., Esmkhani M., Khodadadi J., Fotouhi-Ardakani R., Koohpaei A., Doosti Z., Golzari S.E. Evaluation of bacterial co-infections of the respiratory tract in COVID-19 patients admitted to ICU. BMC Infect. Dis. 2020;20:646. doi: 10.1186/s12879-020-05374-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Lee A., Cheung Y.S.L., Joynt G.M., Leung C.C.H., Wong W.-T., Gomersall C.D. Are high nurse workload/staffing ratios associated with decreased survival in critically ill patients? A cohort study. Ann. Intensive Care. 2017;7:46. doi: 10.1186/s13613-017-0269-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Dasgupta S., Das S., Chawan N.S., Hazra A. Nosocomial infections in the intensive care unit: Incidence, risk factors, outcome and associated pathogens in a public tertiary teaching hospital of eastern India. Indian J. Crit. Care Med. 2015;19:14. doi: 10.4103/0972-5229.148633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Alhumaid S., Al Mutair A., Al Alawi Z., Alsuliman M., Ahmed G.Y., Rabaan A.A., Al-Tawfiq J.A., Al-Omari A. Knowledge of infection prevention and control among healthcare workers and factors influencing compliance: A systematic review. Antimicrob. Resist. Infect. Control. 2021;10:1–32. doi: 10.1186/s13756-021-00957-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Al-Omari A., Al Mutair A., Alhumaid S., Salih S., Alanazi A., Albarsan H., Abourayan M., Al Subaie M. The impact of antimicrobial stewardship program implementation at four tertiary private hospitals: Results of a five-years pre-post analysis. Antimicrob. Resist. Infect. Control. 2020;9:95. doi: 10.1186/s13756-020-00751-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Al Mutair A., Alhumaid S., Al Alawi Z., Zaidi A.R.Z., Alzahrani A.J., Al-Tawfiq J., Al-Shammari H., Rabaan A., Khojah O., Al-Omari A. Five-year resistance trends in pathogens causing healthcare-associated infections at a multi-hospital healthcare system in Saudi Arabia, 2015–2019. J. Glob. Antimicrob. Resist. 2021 doi: 10.1016/j.jgar.2021.03.009. [DOI] [PubMed] [Google Scholar]
- 92.Alhumaid S., Al Mutair A., Al Alawi Z., Alzahrani A.J., Tobaiqy M., Alresasi A.M., Bu-Shehab I., Al-Hadary I., Alhmeed N., Alismail M., et al. Antimicrobial susceptibility of gram-positive and gram-negative bacteria: A 5-year retrospective analysis at a multi-hospital healthcare system in Saudi Arabia. Ann. Clin. Microbiol. Antimicrob. 2021;20:43. doi: 10.1186/s12941-021-00450-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Uyeki T.M., Bernstein H.H., Bradley J.S., Englund J.A., File T.M., Jr., Fry A.M., Gravenstein S., Hayden F.G., Harper S.A., Hirshon J.M. Clinical practice guidelines by the Infectious Diseases Society of America: 2018 update on diagnosis, treatment, chemoprophylaxis, and institutional outbreak management of seasonal influenza. Clin. Infect. Dis. 2019;68:e1–e47. doi: 10.1093/cid/ciy866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Unnewehr M., Friederichs H., Bartsch P., Schaaf B. High diagnostic value of a new real-time Pneumocystis PCR from bronchoalveolar lavage in a real-life clinical setting. Respiration. 2016;92:144–149. doi: 10.1159/000448626. [DOI] [PubMed] [Google Scholar]
- 95.Byington C.L., Ampofo K., Stockmann C., Adler F.R., Herbener A., Miller T., Sheng X., Blaschke A.J., Crisp R., Pavia A.T. Community surveillance of respiratory viruses among families in the Utah better identification of germs-longitudinal viral epidemiology (BIG-LoVE) study. Clin. Infect. Dis. 2015;61:1217–1224. doi: 10.1093/cid/civ486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Patel R., Babady E., Theel E.S., Storch G.A., Pinsky B.A., St George K., Smith T.C., Bertuzzi S. Report from the American Society for Microbiology COVID-19 international summit, 23 March 2020: Value of diagnostic testing for SARS–CoV-2/COVID-19. Am. Soc. Microbiol. 2020 doi: 10.1128/mBio.00722-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Morens D.M., Taubenberger J.K., Fauci A.S. Predominant role of bacterial pneumonia as a cause of death in pandemic influenza: Implications for pandemic influenza preparedness. J. Infect. Dis. 2008;198:962–970. doi: 10.1086/591708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Leung C.-H., Tseng H.-K., Wang W.-S., Chiang H.-T., Wu A.Y.-J., Liu C.-P. Clinical characteristics of children and adults hospitalized for influenza virus infection. J. Microbiol. Immunol. Infect. 2014;47:518–525. doi: 10.1016/j.jmii.2013.06.002. [DOI] [PubMed] [Google Scholar]
- 99.Cusumano J.A., Dupper A.C., Malik Y., Gavioli E.M., Banga J., Berbel Caban A., Nadkarni D., Obla A., Vasa C.V., Mazo D., editors. Open Forum Infectious Diseases. Oxford University Press; Oxford, UK: 2020. Staphylococcus aureus bacteremia in patients infected with COVID-19: A case series. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Navarini A.A., Recher M., Lang K.S., Georgiev P., Meury S., Bergthaler A., Flatz L., Bille J., Landmann R., Odermatt B. Increased susceptibility to bacterial superinfection as a consequence of innate antiviral responses. Proc. Nat. Acad. Sci. USA. 2006;103:15535–15539. doi: 10.1073/pnas.0607325103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Didierlaurent A., Goulding J., Patel S., Snelgrove R., Low L., Bebien M., Lawrence T., van Rijt L.S., Lambrecht B.N., Sirard J.-C. Sustained desensitization to bacterial Toll-like receptor ligands after resolutionof respiratory influenza infection. J. Exp. Med. 2008;205:323–329. doi: 10.1084/jem.20070891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Gadi N., Wu S.C., Spihlman A.P., Moulton V.R. What’s sex got to do with COVID-19? Gender-based differences in the host immune response to coronaviruses. Front. Immunol. 2020;11:2147. doi: 10.3389/fimmu.2020.02147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Rabaan A.A., Al-Ahmed S.H., Garout M.A., Al-Qaaneh A.M., Sule A.A., Tirupathi R., Mutair A.A., Alhumaid S., Hasan A., Dhawan M. Diverse immunological factors influencing pathogenesis in patients with COVID-19: A review on viral dissemination, immunotherapeutic options to counter cytokine storm and inflammatory responses. Pathogens. 2021;10:565. doi: 10.3390/pathogens10050565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Kamfose M.M., Muriithi F.G., Knight T., Lasserson D., Hayward G. Intravenous ceftriaxone versus multiple dosing regimes of intravenous anti-Staphylococcal antibiotics for methicillin-susceptible Staphylococcus aureus (MSSA): A systematic review. Antibiotics. 2020;9:39. doi: 10.3390/antibiotics9020039. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data are available upon request. Please contact author for data requests.