Background
Diabetes mellitus (DM) is a leading cause of preventable cardiovascular disease, but the metabolic changes from prediabetes to diabetes have not been fully clarified. This study implemented a metabolomics profiling platform to investigate the variations of metabolites and to elucidate their global profiling from metabolic syndrome to DM. Methods: Male Sprague-Dawley rats (n = 44) were divided into four groups. Three groups were separately fed with a normal diet, a high-fructose diet (HF), or a high-fat (HL) diet while one group was treated with streptozotocin. The HF and HL diet were meant to induce insulin resistance, obesity, and dyslipidemia, which known to induce DM. Results: The most significant metabolic variations in the DM group’s urine samples were the reduced release of citric acid cycle intermediates, the increase in acylcarnitines, and the decrease in urea excretion, all of which indicated energy metabolism abnormalities and mitochondrial dysfunction. Overall, the metabolic analysis revealed tryptophan metabolic pathway variations in the prediabetic phase, even though the mitochondrial function remains unaffected. Conclusion: This study show that widespread methylations and impaired tryptophan metabolism occur in metabolic syndrome and are then followed by a decline in citric acid cycle intermediates, indicating mitochondrial dysfunction in diabetes.
Keywords: Diabetes, Metabolic syndrome, Metabolomics, Methylation, Mitochondrial dysfunction, Prediabetes, Tryptophan
Abbreviations: CAN, acetonitrile; DM, diabetes mellitus; GOT, glutamate oxaloacetate transaminase; GPT, glutamate pyruvate transaminase; LC-MS, liquid chromatography–mass spectrometry; HF, high-fructose; HL, high-fat; HMDB, human metabolome database; PCA, principal component analysis; KEGG, kyoto encyclopedia of genes and genomes; STZ, streptozotocin; TC, total cholesterol; TG, triacylglycerol
1. Introduction
Metabolic syndrome consists of a group of risk factors, and patients with metabolic syndrome are twice as likely as healthy individuals to develop cardiovascular diseases, and five times as likely to develop type 2 diabetes (Eckel et al., 2005, Grundy, 2008). The risk factors that cause metabolic syndrome include insulin resistance, dyslipidemia, obesity, and inflammation (Grundy, 2012). Branched-chain amino acids and, to a lesser extent, the aromatic amino acids phenylalanine and tyrosine were linked to insulin resistance, whereas the gluconeogenic amino acids alanine, glycine, or glutamine, as well as several other amino acids (such as tryptophan, histidine, and arginine), were not. Much remains unknown, however, about the interactions between the various risk factors and the associated metabolic pathways, the variations that occur as the disease progresses (not just amino acids, but also lipid, carbohydrate metabolite, and pathway variations), and the suitability of metabolite profiles as biological indicators (Bloomgarden, 2018). As an emerging platform technology in the post-genomic era, metabolomics is aimed at studying the global profiling and variations of low-molecular-weight metabolites caused by pathophysiological stimuli and genetic variations (Monteiro et al., 2013, Sas et al., 2015). In this study, we sought to explore the early changes that occur in the progression from metabolic syndrome to diabetes mellitus (DM) in rats and to further clarify the underlying mechanisms of that transition through metabolomics and its applicable biomarkers (Huang et al., 2017, Srinivasan et al., 2005).
2. Materials and methods
2.1. Animals and animal study
14-week-old male Sprague-Dawley (SD) rats (n = 44) were obtained from BioLASCO Laboratory Animal Center, Taipei, Taiwan. The animals were kept in an animal house at 22 ± 2 °C and 55 ± 10% relative humidity and maintained under a 12:12-h light–dark cycle. They were randomly divided into four groups. Three groups were fed with normal rodent chow (control group, CT, n = 10), high fructose diet (HF group, n = 10), and high fat diet (HL group, n = 10). The fourth group was treated with streptozotocin (STZ, 65 mg/kg body weight) to induce DM (DM group, n = 14). The diets of the HF and HL groups were intended to induce the insulin resistance, obesity, and dyslipidemia observed in human metabolic syndrome, while the confirmation of DM induction in the DM group was performed as previously described. The experimental protocols were approved by the Institutional Animal Care and Use Committee of Chang Gung University, and basic standards of laboratory animal care were followed.
Starting when they were 16 weeks old, urine and plasma samples were collected from the CT, HF, HL, and DM groups (Fig. 1). The urine samples were filtered through a 0.2 μm filter and stored at −80 °C until they could be analyzed. The blood samples (100 μl) were collected after the animals had fasted overnight. Plasma glucose was determined using a glucose analyzer (Quik-Lab; Ames Divison, Miles Laboratories, Elkhart, IN). Plasma triglyceride and cholesterol were measured using the appropriate enzymatic diagnostic kits. Plasma insulin was measured by an enzyme immunoassay using 10-μl aliquots of plasma with a rat insulin ELISA kit (Mercodia, Uppsala, Sweden).
Fig. 1.
Protocol of animal experiment. CT, rats fed with a control diet; HF, rats fed with a high-fructose diet; HL, rats fed with a high-lipid diet; DM, rats with diabetes induced by a single intraperitoneal injection of streptozotocin (65 mg/kg BW) and fed with a control diet.
2.2. Plasma and urine sample extraction
A fixed volume (400 μl) of acetonitrile (ACN) was added to 100 μl of plasma. The mixture was sonicated and centrifuged at 10,000 g for 25 min. The supernatant was collected into a separate tube, and the pellets were re-extracted once with ACN. The residual pellets were re-extracted with an equivalent volume of 50% aqueous methanol. The aqueous methanolic supernatant and two ACN supernatants were pooled and dried in a nitrogen evaporator and stored at −80 °C. The samples were then re-suspended in 100 μl of 95:5 water/ACN and centrifuged at 14,000 g for 5 min. The clear supernatant was then collected for liquid chromatography–mass spectrometry (LC-MS) analysis. Prior to analysis, urine samples were diluted with distilled water to a creatinine content of 20 μg/ml. The samples were then centrifuged at 14,000 g for 5 min, and the supernatants were collected for LC-MS analysis.
2.3. Liquid chromatography/time-of-flight mass spectrometry (LC/TOF-MS) analysis
Liquid chromatographic separation was achieved on a 100 mm × 2.1 mm Acquity 1.7-μm C18 column (Waters Corp, Milford, MA) by using an HPLC system (1200 rapid resolution system; Agilent Technologies, Santa Clara, CA). The column was maintained at 40 °C with a flow rate of 0.25 ml/min. Samples were eluted from the LC column with linear gradients of solvent A (2 mM ammonium formate in water) and solvent B (100% ACN): 0–1 min: 0% B; 1–9.6 min: 0–98% B; 9.6–15 min: 98% B; and 15–18 min 0% B.
Mass spectrometry was performed on an Agilent Q-TOFMS (6510 Q-TOF MS; Agilent Technologies, Santa Clara, CA) operated in electrospray positive-ion (ESI+) and electrospray negative-ion (ESI−) modes. The scan range was from 50 to 1000 m/z. The capillary and skimmer voltages were set at 4000 and 65 V, respectively; the liquid nebulizer was set to 30 psig, and the nitrogen drying gas was set at a flow rate of 10 l/min; and the drying gas temperature was maintained at 350 °C. To maintain a constant mass accuracy, purine ([M+H]+ = 121.050873), hexakis (1H,1H,3H-tetrafluoropropoxy) phosphazine ([M+H]+ = 922.009798), or purine ([ M – H ] − = 119.036320) and formate adduct ([M–H] − = 966.000725) were used as internal reference ions. Data were collected in the profile mode by using data acquisition software (Agilent MassHunter Workstation).
2.4. Data processing
Information regarding the metabolites of interest was extracted from the raw data by using the molecular feature extraction algorithm (MassHunter, Agilent). In addition to time-aligned ion features, the algorithm provided monoisotopic neutral mass, retention time, and ion abundances. GeneSpring-MS was used to analyze and visualize the pattern of MassHunter data matrices. Principal component analysis (PCA) was used for clustering analysis and correlation analysis (Fig. 2) (Dai et al., 2014). The relative concentrations of metabolites were compared by ANOVA with Tukey HSD correction. The same software was also used for multivariate data analysis and representation. Accurate masses of features that were significantly different between the control and test groups were searched using the METLIN, Human Metabolome Database (HMDB), and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. The compound prediction was performed by the Metabolite Database and Molecular Formula Generation software (Agilent Technologies).
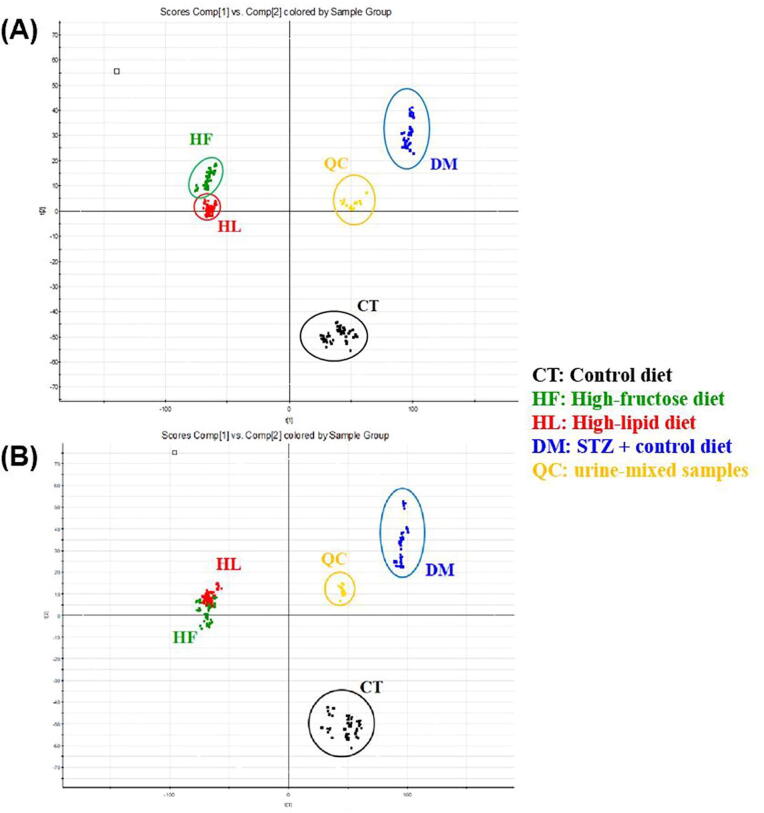
Fig. 2.
The PCA plot of urine samples. (A) ESI positive mode (B) ESI negative mode. CT, rats fed with a control diet; HF, rats fed with a high-fructose diet; HL, rats fed with a high-lipid diet; DM, rats with diabetes induced by a single intraperitoneal injection of streptozotocin (65 mg/kg BW) and fed with a control diet; QC, urine-mixed samples (used as a quality control).
2.5. Metabolite identification
For structural identification, we used the identical chromatographic conditions that were employed in the profiling experiment for the metabolite standards. Mass spectrometry using an Agilent 6510 Q-TOF MS was performed under the following conditions: gas temperature, 350 °C; flow rate of drying gas, 10 l/min; nebulizer pressure, 30 psig; capillary voltage, 4000 V; fragmentor voltage, 175 V; skimmer voltage, 65 V. MS and MS/MS spectra were both collected at 1.0 spectrum per second, with a medium isolation window of ~4 m/z. For MS/MS, the collision energy was set from 5 to 35 V. Several metabolites were further confirmed by an ion mobility mass spectrometer (SYNAPT HDMS, Waters) under similar chromatographic conditions.
2.6. Statistical analysis
All the data are expressed as means ± SE. One-way ANOVA was conducted for multiple-group comparisons, and a Tukey post-hoc test was used to evaluate the significance of differences between group means. In all of the analyses, p < 0.05 was considered significant.
3. Results
3.1. Effects of high fructose and high fat diet in SD rats
The HF and HL groups were fed high-fructose feed and high-fat feed over a period of 36 days. The water intake of the HF and HL groups and 24-hour urine output of the HF group was significantly lower than that of the CT group. The HF and HL groups recorded significantly higher liver weights than the CT group. With respect to the weight of adipose tissue, the HL group had a significantly higher weight than the CT group.
As for the ratio of liver weight to body weight, the HF group had a significantly higher ratio than the CT and HL group, while HL group’s adipose weight to body weight was higher than the other two groups (Table 1). Both the HF and HL group had higher total cholesterol (TC) and triacylglycerol (TG) concentration than CT group, with HF group’s TG concentration having much higher levels than that of the HL group (Table 1).
Table 1.
Physiological and biochemical parameters of experimental animals.
| CT (n = 10) | HF (n = 10) | HL (n = 10) | DM (n = 14) | |
|---|---|---|---|---|
| Food intake (g/d) | 24 ± 5 | 24 ± 4 | 23 ± 3 | 52 ± 14a,b,c |
| Water intake (g/d) | 38 ± 8 | 28 ± 5a | 29 ± 8a | 252 ± 102a,b,c |
| Body weight (BW) (g) | ||||
| Initial | 460 ± 43 | 446 ± 52 | 451 ± 36 | 456 ± 42 |
| Final | 492 ± 22 | 514 ± 59 | 581 ± 42a,b | 339 ± 41a,b,c |
| Liver (g) | 14.93 ± 1.91 | 18.47 ± 4.23a | 17.53 ± 1.66a | 15.89 ± 2.74 |
| Fat pad (g) | 5.95 ± 1.39 | 7.76 ± 2.28 | 11.09 ± 3.05a,b | 0.93 ± 0.91a,b,c |
| Liver/body weight (100 g/g) | 3.04 ± 0.31 | 3.57 ± 0.30a | 3.02 ± 0.44b | 4.71 ± 0.24a,b,c |
| Fat pad/body weight (100 g/g) | 1.22 ± 0.31 | 1.49 ± 0.30 | 1.90 ± 0.44a,b | 0.20 ± 0.24a,b,c |
| Urine | ||||
| Volume (mL/day) | 20 ± 7 | 13 ± 6a | 16 ± 16 | 212 ± 55a,b,c |
| Creatinine (µg/mL) | 885 ± 283 | 1143 ± 311 | 1456 ± 796 | 81 ± 17a,b,c |
| Total creatinine (mg) | 15.9 ± 2.3 | 13.6 ± 2.4 | 14.1 ± 3.0 | 16.9 ± 3.9b |
| Serum | ||||
| Glucose (mg/dL) | 246 ± 37 | 269 ± 40 | 256 ± 22 | 540 ± 79a,b,c |
| Total cholesterol (mg/dL) | 71 ± 10 | 91 ± 20a | 95 ± 21a | 120 ± 44a,b,c |
| Triacylglycerol (mg/dL) | 88 ± 27 | 222 ± 92a | 136 ± 32a,b | 497 ± 373a,b,c |
| Glutamate oxaloacetate transaminase (U/L) | 104 ± 42 | 79 ± 14a | 84 ± 17 | 480 ± 228a,b,c |
| Glutamate pyruvate transaminase (U/L) | 33 ± 16 | 16 ± 8a | 11 ± 4a,b | 213 ± 91a,b,c |
*The 14-week-old male SD rats (n = 44) were randomly divided into four groups (CT, DM, HF, and HL groups). CT, rats fed with a control diet. DM, rats injected with streptozotocin (65 mg/kg BW) and fed with the control diet. HF, rats fed with a high-fructose diet. HL, rats fed with a high-lipid diet. **Data were presented as mean ± SD. a, p < 0.05 compared with CT group; b, p < 0.05 compared with HF group; c, p < 0.05 compared with HL group by one-way ANOVA with Tukey’s post-hoc test.
With respect to liver function, the CT group's glutamate oxaloacetate transaminase (GOT) concentration was significantly higher than that of the HF group. Contrary to our expectation, both the HF and HL groups had lower concentrations of glutamate pyruvate transaminase (GPT) concentration than the CT group. However, the HF group still had a significantly higher concentration than the HL group (Table 1).
3.2. Physiological and biochemical parameters of DM group
When the rats were 14 weeks old, the mean weight of the DM group was significantly lower than that of the CT group even though their daily food intake was higher. In addition, the DM group’s 24-hour urine output was considerably higher than the CT group’s and urinary creatinine concentration was lower. (Table 1)
The DM group had lower weight of adipose tissue and ratio of fat weight to body weight than the CT group. Moreover, the ratio of liver weight to body weight for the DM group was higher than the CT group (Table 1). For both TC and TG concentrations, the DM group recorded significantly higher levels compared to the CT group.
3.3. Identification of target metabolites
Using the S-plot, points that were comparatively differentiated were selected from both the top and bottom sections and compared with the HMDB to identify metabolites matched by the mass spectrometry with a mass value error of no more than 5 ppm. Cross-comparison was then carried out using MarkerLynx XS to identify any molecular formulas (Cai et al., 2009). Results from the comparison of the HF and CT groups revealed 34 metabolites that may be positively correlated to the CT group and 58 metabolites that may be negatively correlated to the CT group.
The comparison of the HL and CT groups showed 42 metabolites that may be positively correlated to the CT group and 55 metabolites that may be negatively correlated to the CT group. Results from the comparison of the DM and CT groups revealed 47 metabolites that may be positively correlated to the CT group and 39 metabolites that may be negatively correlated to the CT group (Table 2). All of the metabolites that were found were compared with databases (HMDB and KEGG) to identify the metabolic pathways that underwent variations during disease progression.
Table 2.
Proposed disturbances of urine metabolome and metabolic pathways.
| Pathway | Metabolites tentatively identified | HF | HL | DM |
|---|---|---|---|---|
| Tryptophan metabolism | Tryptophan | — | — | ↑ |
| Indoxyl | ↑ | ↑ | ↓ | |
| Indoxyl sulfate | ↑ | — | ↓ | |
| Tryptamine | — | — | ↑ | |
| N-Methyltryptamine | ↓ | ↓ | — | |
| Indoleacetaldehyde | ↓ | ↓ | — | |
| Tryptophanol | ↓ | ↓ | — | |
| 4,6-Dihydroxyquinoline | ↓ | ↓ | — | |
| Serotonin | — | — | ↓ | |
| N-Acetylserotonin | ↓ | ↓ | — | |
| 6-Hydroxymelatonin | ↓ | ↓ | — | |
| 5-Hydroxyindoleacetate | ↑ | — | — | |
| Indolelactic acid/ 5-Methoxyindoleacetate | ↓ | ↓ | — | |
| Kynurenine | ↓ | ↓ | — | |
| Kynurenate | — | — | — | |
| Formylanthranilate | ↑ | ↑ | ↓ | |
| Anthranilate | ↑ | ↑ | — | |
| 3-Hydroxyanthranilate | ↓ | ↓ | — | |
| Nicotinate and nicotinamide metabolism | Nicotinamide | — | — | ↓ |
| N1-Methylnicotinamide | ↓ | ↓ | ↓ | |
| Nicotinate | ↓ | ↓ | ↑ | |
| Nicotinurate | ↓ | ↓ | ↑ | |
| Trigonelline | ↓ | ↓ | ↑ | |
| Histidine metabolism | Oxoglutaric acid | — | — | ↓ |
| Methylhistidine | ↑ | — | — | |
| Methylimidazole acetaldehyde | ↓ | ↓ | — | |
| N-Acetylhistamine | ↓ | ↓ | — | |
| Glycine, serine, and threonine metabolism | Sarcosine | — | — | ↑ |
| Glycine | ↑ | ↑ | — | |
| Glyoxylic acid | ↑ | ↑ | — | |
| Arginine and proline metabolism | Sarcosine | — | — | ↑ |
| Creatinine | ↑ | ↑ | ↓ | |
| Urea | — | — | ↓ | |
| Glyoxylic acid | ↑ | ↑ | — | |
| L-Glutamine | — | ↑ | — | |
| Taurine and hypotaurine metabolism | L-Alanine | — | — | ↑ |
| Taurine | — | ↑ | ↓ | |
| 5-L-Glutamyl-taurine | ↓ | ↓ | ↑ | |
| Oxoglutaric acid | — | — | ↓ | |
| Tyrosine metabolism | Phenylacetylglycine | ↑ | ↑ | — |
| 3-Hydroxyphenylacetic acid | ↓ | ↓ | — | |
| Vanylglycol | ↓ | ↓ | — | |
| 4-Hydroxyphenylacetaldehyde | ↓ | ↓ | — | |
| Tyrosol | ↓ | ↓ | — | |
| Lysine degradation | D-1-Piperideine-2-carboxylic acid | — | — | ↑ |
| N6-Acetyl-L-lysine | — | ↑ | ↑ | |
| Valine, leucine, and isoleucine biosynthesis (BCAAs) & degradation | L-Leucine / L-Isoleucine | — | — | ↑ |
| Phenylalanine metabolism | N-Acetyl-L-phenylalanine | — | — | ↑ |
| 2-Methylhippuric acid | ↑ | ↑ | — | |
| 3-Hydroxyphenylacetic acid | ↓ | ↓ | — | |
| Phenylacetic acid | ↓ | ↓ | — | |
| Citric acid cycle | Oxoglutaric acid | — | — | ↓ |
| cis-Aconitic acid | — | — | ↓ | |
| Isocitric acid | — | — | ↓ | |
| Citric acid | — | — | ↓ | |
| Pyrimidine metabolism | Methylcytosine | ↑ | — | ↓ |
| Urea | — | — | ↓ | |
| Thymidine | ↑ | — | — | |
| L-Glutamine | — | ↑ | — | |
| Purine metabolism | Glyoxylic acid | ↑ | ↑ | — |
| Adenosine 2′,3′-cyclic phosphate | ↑ | ↑ | — | |
| L-Glutamine | — | ↑ | — | |
| Aminobenzoate metabolism | 3-Hydroxyanthranilate | ↓ | ↓ | — |
| Diethylphosphate | — | ↑ | — | |
| 4-Aminophenol | ↓ | ↓ | — | |
| Mandelic acid | ↓ | ↓ | — | |
| Aminobenzoic acid | ↓ | ↓ | — |
3.4. Quantifying target metabolites in tryptophan metabolic pathways
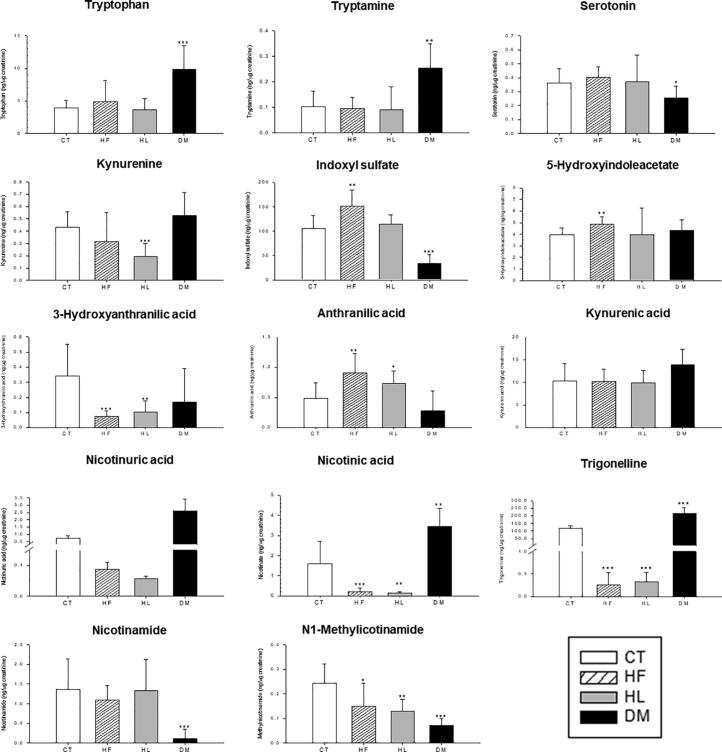
Of the above metabolic pathway variations, tryptophan metabolism was shown to have the highest levels of variation. With respect to serotonin, melatonin, and 5-hydroxyindoleacetate, the 5-hydroxyindoleacetate concentration levels in the HF group were significantly higher than those in the CT group. However, for the second pathway, no significant changes were detected among the HF, HL, and CT groups. The third pathway consists of metabolites related to and generated by intestinal micro-organisms, such as indoxyl and indoxyl sulfate; the results showed that the HF group's indoxyl sulfate concentration was significantly higher than the CT group’s. For the fourth pathway, the changes in the HF and HL groups exhibited a high level of consistency. For example, the HL group's kynurenine concentration (0.195 ± 0.107 ng/μg creatinine) was significantly lower than the CT group’s (0.433 ± 0.127 ng/μg creatinine) (p < 0.05). Meanwhile, the HF group's anthranilate concentration (0.910 ± 0.312 ng/μg creatinine) was also significantly higher than the CT group’s (0.487 ± 0.251 ng/μg creatinine) (p < 0.05). In contrast, the 3-hydroxyanthranilate, N1-methylnicotinamide, nicotinate, nicotinurate, and trigonelline concentration levels of the HF and HL groups were both significantly lower than those of the CT group.
As for tryptophan, it was significantly higher in the DM group than the CT group (10.054 ± 3.609 versus 3.975 ± 1.227 ng/μg creatinine) (p < 0.05). Moreover, for the first pathway, the DM group's serotonin concentration (0.255 ± 0.085 ng/μg creatinine) was significantly lower than the CT group’s (0.362 ± 0.103 ng/μg creatinine) (p < 0.05). However, for the second pathway, the DM group's tryptamine concentration (0.254 ± 0.095 ng/μg creatinine) was significantly higher than the CT group’s (0.103 ± 0.061 ng/μg creatinine) (p < 0.05). As for the third pathway (related to intestinal microbes), the DM group's indoxyl sulfate concentration (31.528 ± 19.468 ng/μg creatinine) was significantly lower than the CT group’s (105.947 ± 28.724 ng/μg creatinine) (p < 0.05). Lastly, in the fourth pathway, the DM group's nicotinamide and N1-methylnicotinamide concentration levels (0.122 ± 0.228, 0.073 ± 0026 ng/μg creatinine) were significantly lower than the CT group’s (1.357 ± 0.782, 0.243 ± 0.079 ng/μg creatinine), while the DM group's nicotinate, nicotinurate, and trigonelline levels were significantly higher than the CT group’s. The correlations between the urine metabolites of the tryptophan metabolism pathways in all the groups are shown in Table S1.
4. Discussion
In this study, in order to induce metabolic syndrome, the HF group were given high-fructose feed to increase the development of two risks factors, insulin resistance and dyslipidemia. A rise in the liver/body weight ratios, signs of fatty liver disease, an increase in blood TG values, and a slight increase in TC values could be observed in the HF group, but obesity was not observed. In this liver, fructose is phosphorylated by fructokinase to fructose-1-phosphate, which could bypass the glycogenolytic metabolic pathway determined by phosphofructokinase-1 and finally break down into dihydroxyacetone phosphate and glyceraldehyde. Glyceraldehyde will be phosphorylated into glyceraldehyde 3-phosphate, which produces acyl CoA. Simultaneously, dihydroxyacetone phosphate further generates glycerol-3-phosphate, which synthesizes with acyl CoA to form TG, thus leading to a rise in TG concentration levels in the blood, and subsequently to a rise in free fatty acid concentration levels due to the influence of lipase (Gugliucci, 2017, Alwahsh and Gebhardt, 2017, Ter Horst et al., 2016).
The HL group displayed clear signs of obesity, fatty liver, high TC and high TG (although to a lesser extent than the HF group) (Buettner et al., 2006, Marques et al., 2016, Buettner et al., 2007). Consuming high-fat feed leads to fat accretion, as well as an increase in the concentration of free fatty acids in the blood. Moreover, fat will accumulate in the muscles and liver, resulting in the generation of ectopic fat, which in turn causes the surrounding tissues to exhibit increased insulin resistance (Liu et al., 2016, Gheibi et al., 2017).
The DM group rats were found to have the lightest weight, in spite of their high food and water intake. Since STZ causes β cell necrosis and disrupts insulin synthesis, it will induce high blood sugar levels in rats (Islam and Loots du, 2009, Zou et al., 2017, Gharibi et al., 2017). The DM rats exhibited high blood sugar, TG, and cholesterol levels, as well as significantly higher concentrations of serum GOT and GTP compared to the CT group- an indicator of impaired liver function.
Urine samples contain small molecule metabolites (excreted through terminal metabolic pathways), which can be studied as biological indicators (Emwas et al., 2016, Emwas et al., 2015). The urine samples from the HF and HL groups, being the groups in which metabolic syndrome was simulated, showed partial overlapping and significant clustering of these metabolites compared to the CT and DM groups, which indicated there were some common metabolites causing the variances.
The present study found that the trends in the variations of the metabolites in the HF and HL groups were not identical but were consistent (Fig. 3). The tryptophan metabolic pathways are such an example and they can be categorized into several parts. One of these routes consists of tryptophan-based physiological amines such as serotonin, melatonin, and 5-hydroxyindoleacetate (Oh et al., 2016, Palego et al., 2016, Bender, 1983). In the HF group, urinary 5-hydroxyindoleacetate concentration levels increased significantly. Relatedly, recent studies have indicated that 5-hydroxyindoleacetate rises in the plasma of those with metabolic syndrome (Fukui et al., 2012). With regard to another pathway, the release of urinary indoxyl was observed to have risen in both the HF and HL groups, while indoxyl sulfate rose significantly in the HF group only. Since indoxyl sulfate is a product of intestinal microbial metabolism, it is speculated that high-fructose and high-fat diets will result in comparatively abnormal gut microbiomes. Previous studies have shown that indoxyl sulfate negatively affects renal function (Barrios et al., 2015, Meijers and Evenepoel, 2011, Nazzal et al., 2017). Moreover, methylhippuric acid and phenylacetylglycine both rose in the HF and HL groups, results which support the involvement of intestinal microbes in metabolic variations (Martin et al., 2012, Tjellstrom et al., 2005, Martin et al., 2011). The upward trend of phenylacetylglycine in the HF and HL groups may have been due to a rise in the phenylacetylglycine precursors produced by intestinal microbes, proving the link between phenylacetylglycine and obesity (Pelantova et al., 2016). With regard to the tryptophan metabolic pathways in the DM group, the serotonin concentration in the groups decreased significantly. Serotonin is a type of neurotransmitter that is primarily used to manage one’s mood, appetite, and sleep behavior (Voigt and Fink, 2015). Similarly, diabetes is often accompanied with depression and sleep disorders, as there is a fall in serotonin concentration.
Fig. 3.
Metabolic disturbance of tryptophan-nicotinamide metabolism pathway. The identity of the metabolites was validated by tandem MS/MS. The level of each metabolite is normalized by creatinine (ng/μg creatinine). CT, rats fed with a control diet; HF, rats fed with a high-fructose diet; HL, rats fed with a high-lipid diet; DM, rats with diabetes induced by a single intraperitoneal injection of streptozotocin (65 mg/kg BW) and fed with a control diet *p < 0.05, **p < 0.01, ***p < 0.001 compared with CT group by one-way ANOVA with Tukey’s post-hoc test.
In another pathway, the DM group's indoxyl sulfate urine level decreased significantly, which may have further increased its serum levels and then caused a deterioration in renal function (Atoh et al., 2009, Chung et al., 2019). The other pathway relates to nicotinamide metabolism, which primarily produces coenzymes like NAD+ and NADP+ that play a role in glycogenolysis, glucoamylation, and the citric acid cycle (Huang et al., 2013). The results showed that the terminal metabolites, nicotinate, nicotinurate, and trigonelline, increased significantly in the DM group, which implied that the intermediate products of this metabolic pathway cannot be fully undertaken by pathways like glycogenolysis, glucoamylation, and the citric acid cycle. This may lead to deterioration of energy metabolism in these rats.
Another variation in the DM group worthy of investigating was the variation in the citric acid cycle. The oxoglutaric acid, cis-aconitic acid, isocitric acid, and citric acid levels in the urine samples of the group all showed a significant decline. These metabolites are the intermediate products of the citric acid cycle, indicating that there was a drop in metabolism and mitochondrial function in the group. Previous studies have shown that hyperglycemia can cause mitochondrial dysfunction, inhibit the activities of pyruvate kinase, glucokinase, and phosphofructokinase, and further promote the formation of gluconeogenesis (Blake and Trounce, 2014, Bhatti et al., 2017, Seidman et al., 1967). The results of our study support the conclusion that a reduction in mitochondrial function occurred in the DM group given the observation that carnitine decreased while acetylcarnitine increased in the urine samples of the group, indicating impairment of the citric acid cycle. In brief, the products of β-oxidation during fatty acid oxidation cannot be undertaken by the citric acid cycle, so they leave the cells and enter the blood in the form of acylcarnitines, before finally being excreted out the urine. Moreover, there is a reduction in the excretion level of urea, the end product of the urea cycle, which partially implies mitochondrial perturbation (Bonham et al., 1999). Taken together, the comprehensive evidence leads to the conclusion that a decline in mitochondrial utility is the most prominent dysfunction in diabetes. Meanwhile, although the HF and HL rats were progressing toward diabetes, the analysis of their metabolites did not indicate mitochondrial dysfunction.
5. Conclusions
Through the metabolomics approach, we clarified a range of observations from mild metabolic defects in metabolic syndrome to variations in multiple metabolic pathways in diabetes, indicating that the prediabetic phase is accompanied with tryptophan metabolic pathway-based disparities and widespread methylation, even as the mitochondrial function remains unaffected. However, eventually, through lipid metabolism variations, the perturbation of mitochondria with impairment of β-oxidation and the citric acid cycle can be detected in the diabetic phase.
CRediT authorship contribution statement
Chun-Feng Huang: Conceptualization, Writing - original draft, Writing - review & editing. Ann Chen: Methodology, Formal analysis, Investigation, Project administration. Siao-Yun Lin: . Mei-Ling Cheng: Methodology, Validation, Investigation, Writing - review & editing. Ming-Shi Shiao: Conceptualization, Writing - review & editing. Tso-Yen Mao: Writing - original draft, Writing - review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Acknowledgments
The authors would like to thank Dr. Jan-Kan Chen for his feedback regarding the interpretation of the study.
Funding
This work was supported by grants from Chang Gung University and National Yang Ming Chiao Tung University.
Footnotes
Peer review under responsibility of King Saud University.
References
- Alwahsh S.M., Gebhardt R. Dietary fructose as a risk factor for non-alcoholic fatty liver disease (NAFLD) Arch. Toxicol. 2017;91(4):1545–1563. doi: 10.1007/s00204-016-1892-7. [DOI] [PubMed] [Google Scholar]
- Atoh K., Itoh H., Haneda M. Serum indoxyl sulfate levels in patients with diabetic nephropathy: relation to renal function. Diab. Res. Clin. Pract. 2009;83(2):220–226. doi: 10.1016/j.diabres.2008.09.053. [DOI] [PubMed] [Google Scholar]
- Barrios C., Beaumont M., Pallister T., Villar J., Goodrich J.K., Clark A., Pascual J., Ley R.E., Spector T.D., Bell J.T., Menni C. Gut-microbiota-metabolite axis in early renal function decline. PLoS ONE. 2015;10, (8):e0134311. doi: 10.1371/journal.pone.0134311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bender D.A. Biochemistry of tryptophan in health and disease. Mol. Aspects Med. 1983;6(2):101–197. doi: 10.1016/0098-2997(83)90005-5. [DOI] [PubMed] [Google Scholar]
- Bhatti J.S., Bhatti G.K., Reddy P.H. Mitochondrial dysfunction and oxidative stress in metabolic disorders – a step towards mitochondria based therapeutic strategies. Biochim. Biophys. Acta, Mol. Basis Dis. 2017;1863(5):1066–1077. doi: 10.1016/j.bbadis.2016.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blake R., Trounce I.A. Mitochondrial dysfunction and complications associated with diabetes. BBA. 2014;1840(4):1404–1412. doi: 10.1016/j.bbagen.2013.11.007. [DOI] [PubMed] [Google Scholar]
- Bloomgarden Z. Diabetes and branched-chain amino acids: what is the link? J. Diabetes. 2018;10(5):350–352. doi: 10.1111/1753-0407.12645. [DOI] [PubMed] [Google Scholar]
- Bonham J.R., Guthrie P., Downing M., Allen J.C., Tanner M.S., Sharrard M., Rittey C., Land J.M., Fensom A., O'Neill D., Duley J.A., Fairbanks L.D. The allopurinol load test lacks specificity for primary urea cycle defects but may indicate unrecognized mitochondrial disease. J. Inherit. Metab. Dis. 1999;22(2):174–184. doi: 10.1023/a:1005406205548. [DOI] [PubMed] [Google Scholar]
- Buettner R., Parhofer K.G., Woenckhaus M., Wrede C.E., Kunz-Schughart L.A., Scholmerich J., Bollheimer L.C. Defining high-fat-diet rat models: metabolic and molecular effects of different fat types. J. Mol. Endocrinol. 2006;36(3):485–501. doi: 10.1677/jme.1.01909. [DOI] [PubMed] [Google Scholar]
- Buettner R., Scholmerich J., Bollheimer L.C. High-fat diets: modeling the metabolic disorders of human obesity in rodents. Obesity (Silver Spring) 2007;15(4):798–808. doi: 10.1038/oby.2007.608. [DOI] [PubMed] [Google Scholar]
- Cai X., Zou L., Dong J., Zhao L., Wang Y., Xu Q., Xue X., Zhang X., Liang X. Analysis of highly polar metabolites in human plasma by ultra-performance hydrophilic interaction liquid chromatography coupled with quadrupole-time of flight mass spectrometry. Anal. Chim. Acta. 2009;650(1):10–15. doi: 10.1016/j.aca.2009.01.054. [DOI] [PubMed] [Google Scholar]
- Chung S., Barnes J.L., Astroth K.S. Gastrointestinal microbiota in patients with chronic kidney disease: a systematic review. Adv. Nutr. 2019;10(5):888–901. doi: 10.1093/advances/nmz028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai W., Yin P., Zeng Z., Kong H., Tong H., Xu Z., Lu X., Lehmann R., Xu G. Nontargeted modification-specific metabolomics study based on liquid chromatography-high-resolution mass spectrometry. Anal. Chem. 2014;86(18):9146–9153. doi: 10.1021/ac502045j. [DOI] [PubMed] [Google Scholar]
- Eckel R.H., Grundy S.M., Zimmet P.Z. The metabolic syndrome. Lancet. 2005;365(9468):1415–1428. doi: 10.1016/S0140-6736(05)66378-7. [DOI] [PubMed] [Google Scholar]
- Emwas A.H., Luchinat C., Turano P., Tenori L., Roy R., Salek R.M., Ryan D., Merzaban J.S., Kaddurah-Daouk R., Zeri A.C., Nagana Gowda G.A., Raftery D., Wang Y., Brennan L., Wishart D.S. Standardizing the experimental conditions for using urine in NMR-based metabolomic studies with a particular focus on diagnostic studies: a review. Metabolomics. 2015;11(4):872–894. doi: 10.1007/s11306-014-0746-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emwas A.H., Roy R., McKay R.T., Ryan D., Brennan L., Tenori L., Luchinat C., Gao X., Zeri A.C., Gowda G.A., Raftery D., Steinbeck C., Salek R.M., Wishart D.S. Recommendations and standardization of biomarker quantification using NMR-based metabolomics with particular focus on urinary analysis. J. Proteome Res. 2016;15(2):360–373. doi: 10.1021/acs.jproteome.5b00885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukui M., Tanaka M., Toda H., Asano M., Yamazaki M., Hasegawa G., Imai S., Nakamura N. High plasma 5-hydroxyindole-3-acetic acid concentrations in subjects with metabolic syndrome. Diab. Care. 2012;35(1):163–167. doi: 10.2337/dc11-1619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gharibi F., Soltani N., Maleki M., Talebi A., Nasiri M., Shirdavani S., Nematbakhsh M. The protective effect of L-arginine in cisplatin-induced nephrotoxicity in streptozotocin-induced diabetic rats. Adv Biomed Res. 2017;6:100. doi: 10.4103/2277-9175.212928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gheibi S., Kashfi K., Ghasemi A. A practical guide for induction of type-2 diabetes in rat: incorporating a high-fat diet and streptozotocin. Biomed. Pharmacother. 2017;95:605–613. doi: 10.1016/j.biopha.2017.08.098. [DOI] [PubMed] [Google Scholar]
- Grundy S.M. Metabolic syndrome pandemic. Arterioscler. Thromb. Vasc. Biol. 2008;28(4):629–636. doi: 10.1161/ATVBAHA.107.151092. [DOI] [PubMed] [Google Scholar]
- Grundy S.M. Pre-diabetes, metabolic syndrome, and cardiovascular risk. J. Am. Coll. Cardiol. 2012;59(7):635–643. doi: 10.1016/j.jacc.2011.08.080. [DOI] [PubMed] [Google Scholar]
- Gugliucci A. Formation of fructose-mediated advanced glycation end products and their roles in metabolic and inflammatory diseases. Adv Nutr. 2017;8(1):54–62. doi: 10.3945/an.116.013912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang C.F., Cheng M.L., Fan C.M., Hong C.Y., Shiao M.S. Nicotinuric acid: a potential marker of metabolic syndrome through a metabolomics-based approach. Diabetes Care. 2013;36(6):1729–1731. doi: 10.2337/dc12-1067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang L., Yang L., Luo L., Wu P., Yan S. Osteocalcin improves metabolic profiles, body composition and arterial stiffening in an induced diabetic rat model. Exp. Clin. Endocrinol. Diab. 2017;125(4):234–240. doi: 10.1055/s-0042-122138. [DOI] [PubMed] [Google Scholar]
- Islam M.S., Loots du T. Experimental rodent models of type 2 diabetes: a review. Methods Find. Exp. Clin. Pharmacol. 2009;31(4):249–261. doi: 10.1358/mf.2009.31.4.1362513. [DOI] [PubMed] [Google Scholar]
- Liu J., Han L., Zhu L., Yu Y. Free fatty acids, not triglycerides, are associated with non-alcoholic liver injury progression in high fat diet induced obese rats. Lipids Health Dis. 2016;15:27. doi: 10.1186/s12944-016-0194-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marques C., Meireles M., Norberto S., Leite J., Freitas J., Pestana D., Faria A., Calhau C. High-fat diet-induced obesity Rat model: a comparison between Wistar and Sprague-Dawley Rat. Adipocyte. 2016;5(1):11–21. doi: 10.1080/21623945.2015.1061723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin F.P., Collino S., Rezzi S. 1H NMR-based metabonomic applications to decipher gut microbial metabolic influence on mammalian health. Magn. Reson. Chem. 2011;49(Suppl 1):S47–S54. doi: 10.1002/mrc.2810. [DOI] [PubMed] [Google Scholar]
- Martin F.P., Collino S., Rezzi S., Kochhar S. Metabolomic applications to decipher gut microbial metabolic influence in health and disease. Front. Physiol. 2012;3:113. doi: 10.3389/fphys.2012.00113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meijers B.K., Evenepoel P. The gut-kidney axis: indoxyl sulfate, p-cresyl sulfate and CKD progression. Nephrol. Dial. Transplant. 2011;26(3):759–761. doi: 10.1093/ndt/gfq818. [DOI] [PubMed] [Google Scholar]
- Monteiro M.S., Carvalho M., Bastos M.L., Guedes de Pinho P. Metabolomics analysis for biomarker discovery: advances and challenges. Curr. Med. Chem. 2013;20(2):257–271. doi: 10.2174/092986713804806621. [DOI] [PubMed] [Google Scholar]
- Nazzal L., Roberts J., Singh P., Jhawar S., Matalon A., Gao Z., Holzman R., Liebes L., Blaser M.J., Lowenstein J. Microbiome perturbation by oral vancomycin reduces plasma concentration of two gut-derived uremic solutes, indoxyl sulfate and p-cresyl sulfate, in end-stage renal disease. Nephrol. Dial. Transplant. 2017;32(11):1809–1817. doi: 10.1093/ndt/gfx029. [DOI] [PubMed] [Google Scholar]
- Oh C.M., Park S., Kim H. Serotonin as a new therapeutic target for diabetes mellitus and obesity. Diabetes Metab J. 2016;40(2):89–98. doi: 10.4093/dmj.2016.40.2.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palego L., Betti L., Rossi A., Giannaccini G. Tryptophan biochemistry: structural, nutritional, metabolic, and medical aspects in humans. J Amino Acids. 2016;2016:8952520. doi: 10.1155/2016/8952520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelantova H., Bartova S., Anyz J., Holubova M., Zelezna B., Maletinska L., Novak D., Lacinova Z., Sulc M., Haluzik M., Kuzma M. Metabolomic profiling of urinary changes in mice with monosodium glutamate-induced obesity. Anal. Bioanal. Chem. 2016;408(2):567–578. doi: 10.1007/s00216-015-9133-0. [DOI] [PubMed] [Google Scholar]
- Sas K.M., Karnovsky A., Michailidis G., Pennathur S. Metabolomics and diabetes: analytical and computational approaches. Diabetes. 2015;64(3):718–732. doi: 10.2337/db14-0509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seidman I., Horland A.A., Teebor G.W. Hepatic glycolytic and gluconeogenic enzymes of the obese-hyperglycemic mouse. BBA. 1967;146(2):600–603. doi: 10.1016/0005-2744(67)90249-5. [DOI] [PubMed] [Google Scholar]
- Srinivasan K., Viswanad B., Asrat L., Kaul C.L., Ramarao P. Combination of high-fat diet-fed and low-dose streptozotocin-treated rat: a model for type 2 diabetes and pharmacological screening. Pharmacol. Res. 2005;52(4):313–320. doi: 10.1016/j.phrs.2005.05.004. [DOI] [PubMed] [Google Scholar]
- Ter Horst K.W., Schene M.R., Holman R., Romijn J.A., Serlie M.J. Effect of fructose consumption on insulin sensitivity in nondiabetic subjects: a systematic review and meta-analysis of diet-intervention trials. Am. J. Clin. Nutr. 2016;104(6):1562–1576. doi: 10.3945/ajcn.116.137786. [DOI] [PubMed] [Google Scholar]
- Tjellstrom B., Stenhammar L., Hogberg L., Falth-Magnusson K., Magnusson K.E., Midtvedt T., Sundqvist T., Norin E. Gut microflora associated characteristics in children with celiac disease. Am. J. Gastroenterol. 2005;100(12):2784–2788. doi: 10.1111/j.1572-0241.2005.00313.x. [DOI] [PubMed] [Google Scholar]
- Voigt J.P., Fink H. Serotonin controlling feeding and satiety. Behav. Brain Res. 2015;277:14–31. doi: 10.1016/j.bbr.2014.08.065. [DOI] [PubMed] [Google Scholar]
- Zou W., Yuan J., Tang Z.J., Wei H.J., Zhu W.W., Zhang P., Gu H.F., Wang C.Y., Tang X.Q. Hydrogen sulfide ameliorates cognitive dysfunction in streptozotocin-induced diabetic rats: involving suppression in hippocampal endoplasmic reticulum stress. Oncotarget. 2017;8(38):64203–64216. doi: 10.18632/oncotarget.19448. [DOI] [PMC free article] [PubMed] [Google Scholar]