Abstract
Background
The risk for several common cancers is influenced by the transcriptomic landscape of the respective tissue-of-origin. Vitamin D influences in vitro gene expression and cancer cell growth. We sought to determine whether oral vitamin D induces beneficial gene expression effects in human rectal epithelium and identify biomarkers of response.
Methods
Blood and rectal mucosa was sampled from 191 human subjects and mucosa gene expression (HT12) correlated with plasma vitamin D (25-OHD) to identify differentially expressed genes. Fifty subjects were then administered 3200IU/day oral vitamin D3 and matched blood/mucosa resampled after 12 weeks. Transcriptomic changes (HT12/RNAseq) after supplementation were tested against the prioritised genes for gene-set and GO-process enrichment. To identify blood biomarkers of mucosal response, we derived receiver-operator curves and C-statistic (AUC) and tested biomarker reproducibility in an independent Supplementation Trial (BEST-D).
Results
Six hundred twenty-nine genes were associated with 25-OHD level (P < 0.01), highlighting 453 GO-term processes (FDR<0.05). In the whole intervention cohort, vitamin D supplementation enriched the prioritised mucosal gene-set (upregulated gene-set P < 1.0E−07; downregulated gene-set P < 2.6E−05) and corresponding GO terms (P = 2.90E−02), highlighting gene expression patterns consistent with anti-tumour effects. However, only 9 individual participants (18%) showed a significant response (NM gene-set enrichment P < 0.001) to supplementation. Expression changes in HIPK2 and PPP1CC expression served as blood biomarkers of mucosal transcriptomic response (AUC=0.84 [95%CI 0.66–1.00]) and replicated in BEST-D trial subjects (HIPK2 AUC=0.83 [95%CI 0.77–0.89]; PPP1CC AUC=0.91 [95%CI 0.86–0.95]).
Conclusions
Higher plasma 25-OHD correlates with rectal mucosa gene expression patterns consistent with anti-tumour effects, and this beneficial signature is induced by short-term vitamin D supplementation. Heterogenous gene expression responses to vitamin D may limit the ability of randomised trials to identify beneficial effects of supplementation on CRC risk. However, in the current study blood expression changes in HIPK2 and PPP1CC identify those participants with significant anti-tumour transcriptomic responses to supplementation in the rectum. These data provide compelling rationale for a trial of vitamin D and CRC prevention using easily assayed blood gene expression signatures as intermediate biomarkers of response.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12916-021-02044-y.
Keywords: Vitamin D, Colorectal cancer, Gene expression, Biomarker
Background
Vitamin D deficiency is associated with the risk of several common cancers, the strongest evidence supporting a link between vitamin D and colorectal cancer [1, 2]. However, a causal association has yet to be convincingly demonstrated, because the available observational evidence may be participant to several potential confounders. Environmental risk factors associated with CRC are also associated with vitamin D status (i.e. co-causality; e.g. physical activity), while CRC or its treatment may itself lower plasma vitamin D levels (i.e. reverse causation). However, a recent randomised-control trial (RCT) reported an association between supplementation, vitamin D receptor genotype, and risk of colorectal adenoma, supporting the premise that the beneficial effect may be causal [3]. Meanwhile, vitamin D-related genetic variation has been shown to influence the association between 25-OHD level and CRC survival [4–6], with a recent meta-analysis of RCT data strongly supporting a causal effect for vitamin D supplementation on CRC mortality [7, 8].
Differences in gene expression have been reported in CRC and adenoma tissue relative to normal colorectal tissue [9–12], with genes involved in metabolism, transcription, and translation and cellular processes commonly altered [13]. Recent transcriptome wide association studies confirm the importance of gene expression in carcinogenesis [14, 15]. Vitamin D broadly influences gene expression through activation of the ligand-activated transcription factor VDR, which has been shown to influence cancer cell growth in vitro [16]. Therefore, investigation of gene expression in the colorectum in the context of vitamin D status or supplementation may provide fresh insight into mechanisms underlying the relationship between CRC and vitamin D. Recent evidence suggests one mechanism may be that 1,25-dihydroxyvitamin D3 modulates immune and inflammatory pathway genes in large bowel epithelium [17]. However, differential expression in response to high dose 1,25-dihydroxyvitamin D3 may not accurately reflect the relationship between vitamin D status and gene expression at normal or low vitamin D levels, or in response to regular vitamin D3, the most commonly used vitamin D supplement.
We investigated whether circulating vitamin D concentration is associated with differential gene expression in rectal normal mucosa using a 2-Phase approach with validation of putative biomarkers in an independent study dataset. We directly assayed total 25-OHD, which reflects both dietary intake and skin synthesis of vitamin D [18, 19] and investigated its relationship with gene expression in normal mucosa, assessed by microarray. In the Phase 1 correlative study we sought to identify a prioritised list of differentially expressed genes associated with 25-OHD level. In Phase 2, we conducted a study in human volunteers who were supplemented with oral vitamin D to determine whether the corresponding transcriptomic response was induced in vivo. Using blood peripheral blood mononuclear cells (PBMC) transcriptomic analysis, we also identified potential blood biomarkers that indirectly indicate a beneficial response in the host rectal mucosa.
Methods
Study population
Participants recruited to Phase 1 of the Scottish Vitamin D study (see Additional file 1: Protocol) (n = 191) underwent sampling of the blood and normal rectal mucosa by rigid sigmoidoscopic biopsy. Eligibility criteria included age over 16, ability to perform informed consent, absence of bleeding risk (e.g. coagulopathy, anti-coagulants) and absence of acute colorectal/ano-rectal pathology (e.g. peritonitis, diverticulitis, recent colorectal surgery, anal fissure). RNA was extracted for gene expression analysis from matched contemporaneous NM and peripheral blood mononuclear cells (PBMCs). Plasma was collected at the same time for vitamin D analysis, and DNA was extracted from whole blood for genotyping.
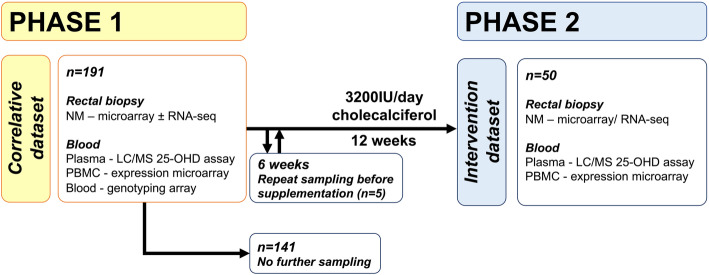
All eligible participants from Phase 1 were invited to proceed to Phase 2 which was an intervention study. Five of the 50 recruited participants to Phase 2 underwent interval sampling before starting supplementation to assess for longitudinal changes in expression before treatment. All Phase 2 participants were then administered vitamin D supplementation and underwent repeat NM, PBMC, and 25-OHD sampling after 12 weeks’ 3200IU/day cholecalciferol (Fultium-D3) supplementation (Fig. 1). No vitamin D supplementation outside of the study protocol was allowed, and concordance with the treatment protocol was assessed through a dose diary and pharmacy log of unused tablets (compliance of 98% of total doses taken achieved). Demographic and clinical data were prospectively collected from patient case notes.
Fig. 1.
Summary of SCOVIDS study protocol. An unselected subset of Phase 1 participants (i.e. selection not based on 25-OHD or baseline gene expression) proceeded to Phase 2 and were given 3200IU cholecalciferol per day. Of these, 5 participants were sampled 6 weeks after initial sampling and without supplementation to provide a control dataset, after which they proceeded to 12 week’s supplementation and final sampling
Sample size considerations
There were no available published data on which to base investigation of the sample size required to determine an association between vitamin D status and global gene expression in normal mucosa. Thus, a formal sample size estimation was not possible.
Blood and mucosa sampling
Participants were sampled in outpatient clinic or during minor surgical procedures. No participant received cleansing oral mechanical bowel preparation. Blood was sampled by standard venepuncture of a peripheral arm vein, with plasma and PBMCs extracted (Additional file 2: Supplementary Methods [20–29]). A separate blood sample was taken to allow extraction of blood leukocyte DNA and genotyping of functionally relevant vitamin D receptor (VDR) polymorphisms (rs1544410, rs10735810, rs7975232, rs11568820 [26–28]) and vitamin D pathway SNPs associated with 25-OHD level (rs2282679, rs10741657, rs2228570, rs6013897 [25]). Normal rectal mucosa (NM) was sampled at the same time via rigid sigmoidoscopic rectal biopsy. NM and PBMC samples were immediately placed in RNAlater and kept immersed for 24–72 h prior to RNA extraction or storage at −80°C.
Plasma vitamin D assay
All plasma samples were measured to a standard, validated, and published protocol by a single laboratory [30]. Total 25-OHD was measured by liquid chromatography tandem mass spectrometry.
Assessment of gene expression
RNA was extracted and purified from NM and blood PBMC using a proprietary RNA extraction kit. Gene expression profiling was undertaken using the Illumina HumanHT-12v4.0 Expression BeadChip Arrays and IScan NO660 scanner, providing coverage of 47,231 transcripts and >31,000 annotated genes. For RNA sequencing, whole-genome transcriptomic patterns were analysed on total RNA from selected normal mucosa samples extracted as described above. RNA was sequenced on the Illumina HiSeq 2500 platform in “rapid mode” with 150bp paired-end reads in a single batch. Transcript indexing and quantification from RNA-seq reads was performed using Salmon v1.1.0 [21].
Statistical analysis
All statistical analysis was undertaken in R [29]. In Phase 1, linear regression modelling was used to test association between 25-OHD level and NM gene expression, adjusting for age, gender, CRC status and anaesthetic status (i.e. sampled under general anaesthetic). VDR haplotype was included in a secondary model for a subset of participants. Genes associated with 25-OHD level at significance level P < 0.01 were termed the ‘candidate gene-set’ and taken forward for testing in the intervention dataset (Phase 2). In Phase 2, differences in 25-OHD level before and after vitamin D supplementation were investigated using paired Wilcoxon rank-sum test and differential gene expression analysis in response to vitamin D supplementation performed using the lmFit and eBayes functions within the ‘limma’ package [31] producing the intervention (Phase 2) dataset. Ranked lists of differentially expressed genes were assessed for functional relevance using the ‘GOrilla', Gene Ontology enRIchment anaLysis and visuaLizAtion tool [32] and replicated using the gseGO function within the ‘clusterProfiler’ package in R [33]. Process ontologies were investigated using gene lists ranked by coefficient (Phase 1) or fold-change (Phase 2).
We tested the Phase 2 dataset (i.e. response to supplementation) for enrichment of the Phase 1 candidate gene-set and top-ranked GO terms. Directional gene-set testing was performed in R, using the gene-setTest function in the ‘limma’ package [34]. We performed technical replication by performing gene-set enrichment testing on differential expression data derived from RNA-seq analysis of the same NM samples from the intervention cohort.
To identify biomarkers of response, we performed participant-level gene-set enrichment testing with a ‘response’ to supplementation defined as enrichment (P < 0.001 given n = 50 subjects) of the candidate gene-set after supplementation. Then, differentially expressed genes in the blood between those with/without rectal NM response were tested for enrichment of the candidate gene-set. Logistic regression testing sought to identify potential blood biomarkers of response and utility of blood biomarkers was calculated using receiver operator curves and C statistic. Finally, we sought to validate putative biomarkers of response in an independent blood gene expression dataset derived from the ‘Biochemical Efficacy and Safety Trial of Vitamin D’ (BEST-D) study [35] (https://www.ebi.ac.uk/arrayexpress/files/E-MTAB-6246/).
Results
Mucosal gene expression signature associated with higher 25-OHD level consistent with anti-tumour effects
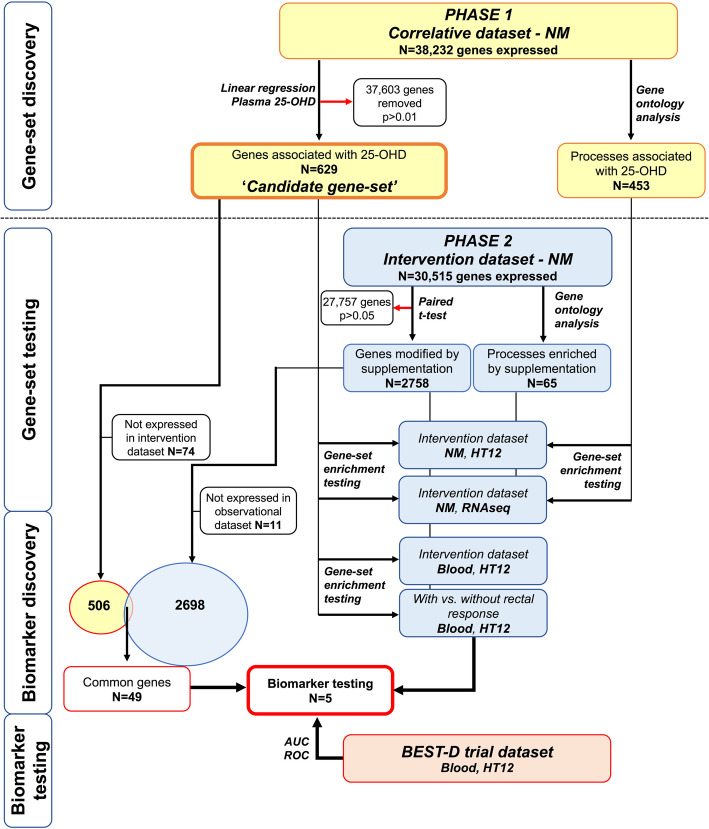
In Phase 1, 191 participants underwent rectal mucosal biopsy and blood sampling (Table 1). 25-OHD was nominally associated with expression of 629 probes (P < 0.01), termed the ‘candidate gene-set’ (‘Gene-set discovery’ Fig. 2).
Table 1.
Baseline characteristics, sampling variables and vitamin D status in included participants
| PHASE 1 Correlative dataset |
PHASE 2 Intervention dataset |
|
|---|---|---|
| N | 191 | 50 |
| Age (median, range) | 63 (24-89) years | 66 (24-88) years |
| Gender (male) | 101 (53%) | 26 (52%) |
| Diagnosis | ||
| Pre-operative; colorectal cancer* | 57 | 1 |
| None (healthy) | 63 | 20 |
| Past medical history of CRC* | 23 | 21 |
| Minor anorectal pathology, no CRC† | 45 | 7 |
| Abdominal tumour (not colorectal) | 3 | 1 |
| Sampled under general anaesthetic | 89 | 3 |
| Median 25-OHD –Baseline | 40 (IQR 37) nmol/l | 36 (IQR 31) nmol/l |
| Median 25-OHD –After supplementation | NA | 89 (IQR 33.5) nmol/l |
Ethnic background for Phase 1 participants were as follows: 183 European, 2 American, 3 African, 3 Asian. * Pre-operative patients had not undergone any neo-adjuvant chemo/radiotherapy at time of sampling. Patients with past history of CRC were previously treated with curative resection +/- adjuvant chemotherapy and no evidence of recurrence at time of recruitment. No significant differences in gene expression were identified according to participant diagnosis. † Full diagnosis list given in Additional file 3: Table S1
Fig. 2.
Analysis flowchart and gene-set selection for biomarker assessment
No individual probe was significantly associated with 25-OHD after adjustment for genome-wide multiple testing (Additional file 3: Table S2), yet the top 3 hits have previous reported association with colorectal tumourigenesis CNN1 [36], COX7A1 [37] and PIP5K1C [38]. Gene ontology analysis demonstrated significant enrichment of 453 processes (Additional file 3: Table S3) with many highly relevant to carcinogenesis e.g. ‘regulation of cell migration’ (FDR=7.55E-08), ‘regulation of programmed cell death’ (FDR=5.38E-03) and ‘regulation of cell differentiation’ (FDR=2.55E-05). Several genes from the candidate gene-set with higher expression associated with higher 25-OHD are included in enriched GO ontology terms relevant to carcinogenesis and have reported tumour suppressor activity (e.g. FOXOs, CAV1, LRP1, Additional file 3: Tables S4, S5 [39–58]). This suggests that the NM gene expression signature, i.e. Phase 1 ‘candidate gene-set’, associated with higher 25-OHD level is consistent with anti-tumour effects.
Oral vitamin D supplementation enriches anti-tumour expression signature in normal rectal mucosa
In Phase 2, 50 participants were administered vitamin D supplementation and underwent repeat sampling after 12-weeks. Post hoc analysis revealed age, gender and baseline 25-OHD to be similar between Phase 1/2 participants (P > 0.05). Supplementation induced an increase in plasma 25-OHD after 12 weeks (median plasma 25-OHD before/after supplementation was 36nmol/l, 89nmol/l; P = 2.5E−09), with a suggestive association between genotype at two vitamin D pathway SNPs (rs2282679, rs2228570) and 25-OHD fold-change after supplementation (Additional file 3: Table S6).
No individual gene from the candidate gene-set showed significant differential expression after adjustment for multiple testing (Additional file 3: Table S7). However, testing of the Phase 1 candidate gene-set showed significant enrichment after supplementation (upregulated gene-set P < 1.0E−07; downregulated gene-set 2.8E−05, see ‘Gene-set testing’ Fig. 2, Table 2, Additional file 3: Figure S1), confirmed in RNA-seq data. Gene-set testing of a candidate gene-set from Phase 1 adjusted for VDR haplotypes, also showed significant enrichment (P < 0.0001).
Table 2.
Gene-set testing for enrichment of the candidate gene-set from Phase 1 after supplementation in Phase 2 and the BEST-D study
| PHASE 1 Correlative dataset |
SCOVIDS Phase 2 | BEST-D trial | |||
|---|---|---|---|---|---|
| Intervention dataset, NM All participants |
Intervention dataset, blood, HT12 | Blood, HT12 | |||
| HT12 | RNA-seq | All participants | With vs. without rectal response | All participants | |
| Candidate gene-set: positive association with 25-OHD level |
P < 1.0E−07 n = 349 |
2.05E−07 n = 242 |
P = 3.89E−13 n = 239 |
P = 3.65E−12 n = 291 |
P = 4.72E−06 n = 191 |
| Candidate gene-set: negative association with 25-OHD level |
P = 2.8E−05 n = 206 |
6.87E−09 n = 155 |
P = 9.60E−05 n = 185 |
P = 3.50E−17 n = 217 |
P = 0.02 n = 130 |
NM normal mucosa. P value given for directional gene-set enrichment test of whether Phase 1 candidate gene-set showed greater change after supplementation when compared to randomly chosen genes. n = number of genes tested. Rectal response is defined by candidate gene-set enrichment in NM P < 0.001. To explore the role of VDR genotype in modifying the association between 25-OHD and gene expression, VDR haplotypes were derived for the four genotyped VDR polymorphisms, rs1544410, rs10735810, rs7975232, rs11568820, using BEAGLE software (version 3.3.2) with standard settings [59] and used as additional covariates in the linear regression model in a subset of 125 participants, resulting in a smaller candidate gene-set which remained significantly enriched after supplementation (P < 0.0001)
Enrichment of the candidate gene-set was not associated with 25-OHD response to supplementation, with those with the lowest 25-OHD FCs still showing gene-set enrichment (Additional file 3: Figure. S1). Meanwhile, there was no enrichment of the candidate gene-set in interval NM samples taken before commencement of supplementation indicating it is a treatment effect (median interval 8 weeks, Additional file 3: Figure S1).
Of the 629 candidate genes associated with circulating vitamin D in the Phase 1 gene-set, 55 had nominally significant expression change after supplementation (P < 0.05). Concordance in direction of effect between the coefficient of association with plasma 25-OHD level and expression change after supplementation was observed in 49 of these (R0.93, P < 2.2E−16, Additional file 3: Table S7), with these genes taken forward for biomarker discovery (see ‘Biomarker discovery’ section Fig. 2).
GO term enrichment indicates modulation of anti-tumour effects in normal mucosa by supplementation
Functional annotation of the intervention dataset gene list identified 65 significantly enriched pathways after supplementation, with many terms relevant to carcinogenesis including ‘regulation of programmed cell death (FDR=9.66E-03) and ‘regulation of cell migration’ (FDR=7.83E−03) (Additional file 3: Table S8). Taken together, genes in the top 50 GO terms from Phase 1 were significantly enriched after supplementation (P = 2.90E−02). Common processes across both the Phase 1 and Phase 2 datasets included terms relevant to carcinogenesis including ‘regulation of programmed cell death’, ‘regulation of cell migration’ (Additional file 3: Figure S2), demonstrating that biologically relevant patterns of gene expression changes associated with higher 25-OHD level and consistent with anti-tumour effects could be imparted by oral supplementation. Results from GOrilla analysis were replicated using both DAVID and GSEA in R, which confirmed enrichment of common processes depicted in Additional file 3: Figure S2, and direction of effect, with terms consistent with ‘anti-tumour’ effects activated in relation to both higher 25-OHD and after vitamin D supplementation.
Blood expression biomarkers identify participants with gene expression response to supplementation
We identified 9 individual participants (18%) with a significant response (i.e. candidate gene-set enrichment in NM P < 0.001) to supplementation. Of the top 50 ranked GO terms from Phase 1, 43 were enriched in these 9 participants after supplementation indicating a biologically relevant NM gene expression response to supplementation, which was absent in the remaining participants (Additional file 3: Table S9). NM gene expression response was associated with an increased allele risk score of the four functionally relevant VDR SNPs (p = 0.006), but not with VDR gene expression, increased 25-OHD fold-change or baseline 25-OHD when adjusting for multiple testing (Additional file 3: Table S10).
Changes in PBMC gene expression after supplementation reflected those in the rectum, with the Phase 1 candidate gene-set significantly enriched in the blood after supplementation (Table 2). Moderate correlation between blood and rectum fold-change in the 49 genes taken forward for biomarker discovery was seen (R=0.64, P = 5.63E−06). When we compared PBMC gene expression after supplementation between those participants with and without a rectal mucosal gene expression response, the differentially expressed genes in PBMCs were enriched for the candidate gene-set, indicating potential blood biomarkers of mucosal response (Table 2, see ‘Biomarker discovery’ section Fig. 2). Five genes identified from the Phase 1 which were both differentially expressed in NM after supplementation and also differentially expressed in blood between participants with and without a rectal response to supplementation (SMEK2, HIPK2, PPP1C, DDR1 and SNX21), indicating biomarker potential (Table 3). When the genes were combined, a blood expression signature based on the best derived cut-off showed strong utility in predicting NM response (AUC=0.99, 95%CI 0.97–1.00, Additional file 3: Table S11, Figure S3).
Table 3.
Genes prioritised from Phase 1 correlative dataset modified by supplementation in NM with evidence of potential biomarker utility in PHASE 2 and BEST-D trial
| Gene | Phase 1 NM |
Phase 2 NM |
Phase 2 Blood |
Phase 2 With vs. without response, Blood |
BEST D trial Blood |
BEST-D trial With vs. without response, Blood |
||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Coeff. | P value | FC | P value | FC | P value | FC | P value | FC | P value | FC | P value | |
| SMEK2 | −0.002 | 0.007 | 0.93 | 0.020 | 0.97 | 0.272 | 0.74 | 0.007 | - | - | - | - |
| HIPK2 | 0.004 | 0.006 | 1.17 | 0.016 | 1.11 | 0.068 | 1.46 | 0.014 | 1.07 | 0.002 | 1.36 | 8.47E−12 |
| PPP1CC | −0.002 | 0.007 | 0.93 | 0.020 | 0.91 | 0.006 | 0.82 | 0.047 | 0.96 | 0.14 | 0.61 | 1.98E−17 |
| SNX21 | 0.004 | 0.003 | 1.07 | 0.047 | 0.99 | 0.750 | 1.26 | 0.039 | 1.01 | 0.27 | 1.04 | 0.02 |
| DDR1 | 0.007 | 0.008 | 1.14 | 0.0008 | 1.03 | 0.572 | 0.72 | 0.040 | 1.02 | 0.03 | 1.04 | 0.08 |
NM normal mucosa, FC fold-change. Results from Phase 1 and Phase 2 analysis given. For Phase 1, coefficient given for association with 25-OHD level. For Phase 2, fold-change and P value given for gene expression response to supplementation in NM and in the blood. In final column, blood expression fold-change difference between participants with and without NM response to the candidate gene-set is given
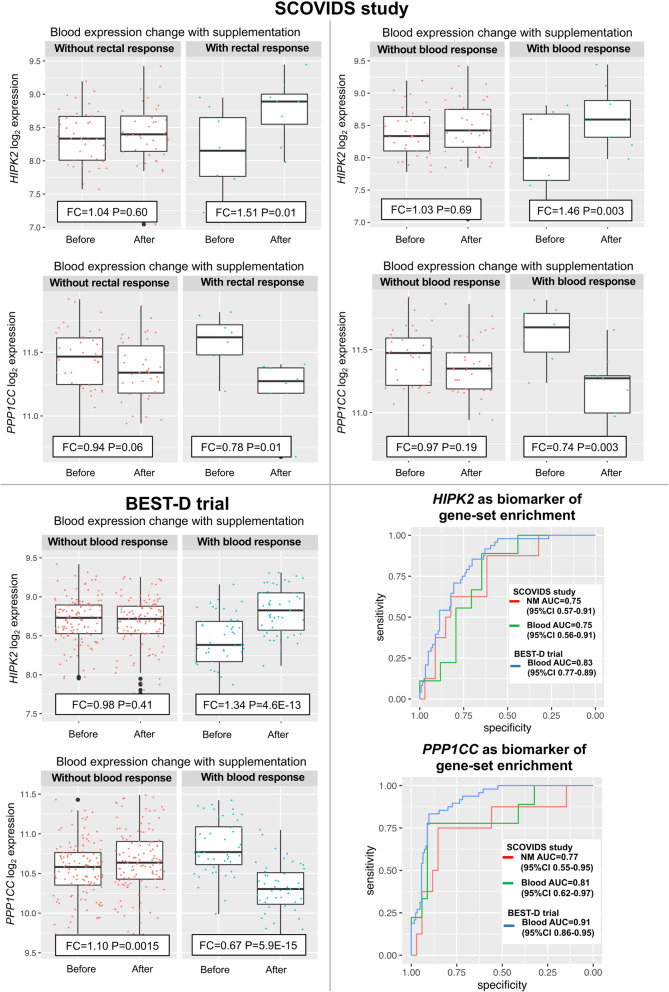
We then explored the value of these same genes as blood biomarkers of the blood response to supplementation. The HIPK2 and PPP1CC genes (Table 3, Fig. 3) demonstrated the best predictive utility in identifying participants with response in both rectum (AUC=0.84, 95%CI 0.66–1.00) and blood (AUC=0.87, 95%CI 0.71–1.00, Additional file 3: Table S11).
Fig. 3.
Blood HIPK2 and PPP1CC expression before and after supplementation in SCOVIDS and BEST-D trial with ROC of biomarker utility. Response defined as participant level gene-set enrichment to Phase 1 candidate gene-set from our SCOVIDS study after HIPK2, PPP1CC, SMEK2, DDR1 and SNX21 excluded
HIPK2 and PPP1CC are independent biomarkers of expression response to supplementation in the BEST-D trial expression dataset
In the BEST-D trial, 48/172 (28%) participants showed significant enrichment of our Phase 1 candidate gene-set. HIPK2 expression increased with supplementation in the BEST-D trial (FC=1.07, p = 0.002), with the increase greatest in those with a blood response to supplementation as defined by our candidate gene-set (FC=1.34; P = 4.6E−13, Fig. 3) and a 1.36 fold-difference in expression change after supplementation between those with/without a response to our candidate gene-set (P = 8.48E−12, Table 3). In the current intervention study (SCOVIDS), we observed an average HIPK2 FC=1.11 in the blood, with 21 (47%) participants showing a FC>1.19, the optimum threshold determined by AUC calculations. In the BEST-D study, 63 (37%) participants taking oral vitamin D supplementation showed a FC>1.19 response, with this signature more prevalent in those taking 4000IU per day (42%), suggesting a dose-response. HIPK2 expression change showed utility in identifying those with a response to our candidate gene-set, AUC=0.83 (95%CI 0.77–0.89, Fig. 3) and when the threshold from our study was used, HIPK2 FC>1.19 AUC=0.74 (95%CI 0.66–0.81, Additional file 3: Table S11).
PPP1CC expression showed non-significant decrease after supplementation in the BEST-D cohort overall, but when the participants were stratified by blood response, PPP1CC was seen to increase in those without blood response (P = 0.0015, Fig. 3) and markedly decreased in those with a blood response (FC=0.67; P = 5.9E−15). There was a 0.61 fold-difference in PPP1CC expression change between those with and without a response to our candidate gene-set (P = 1.98E−17, Table 3). A total of 47 (27%) participants had PPP1CC FC<0.76 after supplementation, closely reflecting the 11 (26%) participants in the current intervention study. Crucially, PPP1CC expression change after supplementation showed utility as a biomarker of blood response to our candidate gene-set, AUC=0.91 (95%CI=0.86−0.95, Fig. 3) and when using the threshold from the SCOVIDs study, PPP1CC FC<0.76 AUC=0.83 (95%CI 0.76−0.89, Additional file 3: Table S11).
Discussion
This study reveals demonstrable differences in gene expression patterns in normal rectal mucosa correlated with plasma 25-OHD level. These differences are consistent with beneficial effects on processes relevant to colorectal carcinogenesis. Furthermore, we show that oral supplementation with vitamin D induces changes in the prioritised gene list. This indicates that the beneficial expression “signature” is not static, but rather can be modified by oral vitamin D supplementation, at least within the timescale tested here. Although we were not able to directly test cancer endpoints, there is considerable published evidence supporting the premise that enrichment of this favourable gene-set imparts anti-tumour effects.
Homeodomain-interacting protein kinase 2 (HIPK2) is a known tumour suppressor gene [60] and the Protein Phosphatase 1 Catalytic Subunit Gamma gene (PPP1CC), a published molecular marker of CRC [61]. We found expression changes in these genes in blood to have predictive value in reflecting rectal mucosa response to supplementation. Hence, these may have utility as blood biomarkers of a beneficial epithelial response to supplementation. In addition, the effect on gene expression in blood PBMCs appears robust, since we replicated the effect in a large, independent, expression dataset, namely the BEST-D trial in which subjects were administered oral vitamin D supplementation (2000/4000IU 12 months).
We devised a 2-Phase in vivo approach, firstly to identify differentially expressed genes in the rectal epithelium associated with plasma 25-OHD and determine the GO terms and processes linked to that prioritised gene list. However, our ultimate aim was to establish whether these transcriptomic responses could be recapitulated by oral vitamin D supplementation, thereby demonstrating a modifiable transcriptomic landscape. Many of the top-ranked genes associated with higher 25-OHD level have links with CRC, for instance CNN1 [36], COX7A1 [37], PEG3 [62], PIP5K1C [38], TAGLN [63] and DAAM2 [64]. Furthermore, we highlight a number of genes within processes relevant to tumourigenesis which are associated with 25-OHD level and influenced by supplementation. The directions of effect of these genes were consistent with tumour suppressor activity. Enrichment of pathways involved in cell migration and cell death validate published in vitro data which demonstrate vitamin D-induced growth arrest and apoptosis of CRC cell lines, modulation of the Wnt signalling pathway, DNA repair and immunomodulation [16]. Published clinical data also corroborate our current findings, for example Protiva et al., reported upregulation of genes involved in cell adhesion in response to 1,25(OH)2D3 [17], while Bostick, reported increased cell differentiation and apoptosis in the normal human colorectal epithelium [65]. Taken together, these data suggest possible mechanisms underlying the widely reported link between vitamin D deficiency and increased CRC risk [1, 2]. It also might explain the recently reported beneficial impact of supplementation on CRC survival outcomes [8].
Despite compelling published observational and pre-clinical data, the link between vitamin D and risk of cancer and several other traits remains controversial. Indeed, several large intervention trials have shown no benefit on cancer endpoints (VITAL Trial [66], Vitamin D Assessment (ViDA) study [67, 68] and Baron et al. [69]). However, participants in these trials were predominantly sufficient for vitamin D at the trial outset, thereby potentially blunting beneficial effects [70]. We have previously rehearsed potential reasons why previous study designs might have failed to detect real effects [70]. To counter potential confounding effects, we conducted a Mendelian randomisation study but this also did not demonstrate a beneficial effect of circulating vitamin D on CRC risk. However, available genetic instrumental variables are weak and explain only a small portion of variance of 25OHD levels [71, 72].
In this study, 18% of participants receiving vitamin D supplementation exhibited a response in the colorectal epithelium (the putative target tissue). Unmeasured variables may account for the marked inter-individual variation in response including ethnic or genetic background, dietary, lifestyle, pharmacological (e.g. concurrent medication) or pathological effects (e.g. unknown viral infection during course of supplementation). Variation in the activity of the vitamin D enzymes or carriers (e.g. CYP24A1, GC and CYP27B1 undetected in the current dataset) may impact responses in the mucosa relative to 25-OHD change, and we noted differential CYP2R1 change in the rectum after supplementation between those with/without a response to our candidate gene-set (FC 0.80 vs. 1.04, P = 0.007). Irrespective of the cause of inter-individual variation in response, if our hypothesis holds that expression changes translate to cancer endpoints, this low response rate would adversely impact on statistical power of trials conducted to date which have tested the effect of vitamin D supplementation on clinical endpoints. Such trials routinely perform subgroup analyses based on change in circulating 25-OHD level, yet the current study reveals poor correlation between plasma level and mucosal gene expression changes, suggesting non-linear responses to vitamin D may introduce further heterogeneity to clinical endpoints. Future work, including GWAS and machine learning approaches will aim to define whether ‘response’ can be determined at baseline is required.
Crucially, we have identified blood biomarkers that reliably identify participants who respond to vitamin D supplementation by inducing gene expression changes in the target tissue. The value of these biomarkers is replicated in a larger independent expression dataset. Further work is required to assess the utility of respective blood protein assays (e.g. ELISA) and reproducibility of these blood biomarkers in identifying mucosal response across a larger cohort. Nevertheless, these exciting and novel findings provide rationale for a trial of vitamin D and CRC prevention using easily assayed blood gene expression signatures as intermediate biomarkers of response.
Whilst the 2-Phase design of an intervention study informed by our correlative dataset has many positive attributes, the study has a number of limitations. First, the low median level of 25-OHD and narrow positively skewed distribution of 25-OHD in the Phase 1 cohort may have masked some true associations between vitamin D level and gene expression. Failure to identify individual gene significance after adjustment for genome-wide multiple testing may also indicate inadequate sample size, physiological autoregulation maintaining constant gene expression despite differences in circulating 25-OHD, differences between plasma and rectal mucosa concentrations of 25-OHD or 1,25-OHD [73] or heterogeneity in cell type in sampled mucosal tissue.
This intervention study is larger than many published studies of gene expression and vitamin D supplementation [17, 74–77], yet may still have limited power to achieve individual gene significance. Phase 2 participants were recruited as a subset of the Phase 1 cohort, which may influence gene-set enrichment test results. However, we did not select those who received supplementation based on 25-OHD level or baseline gene expression, which could have led to overfitting of the data, but instead took an unselected group. If anything, this approach could blunt the observed effect of supplementation, as mucosal response to supplementation may be capped in those with specific 25-OHD or favorable patterns of gene expression at baseline. Despite this, we observed significant enrichment of the candidate gene-set derived in Phase 1 in those receiving supplementation. Unmeasured variation in environmental exposures (e.g. diet or UVB exposure) may have influenced responses, which should be accurately detailed in future studies of vitamin D supplementation. Sampling of rectal mucosa but not colonic mucosa avoided the use of cleansing bowel laxatives which may influence gene expression [78], yet limits the generalisability of our findings to more proximal colonic mucosa. Sampling after 12 weeks of supplementation may not adequately capture early or later gene expression changes yet more frequent or delayed sampling would provide additional practical and ethical challenges. Finally, we recognise responses to vitamin D supplementation and the biomarker utility of HIPK2 and PPP1CC require further mechanistic study (e.g. protein expression) to validate and expand these findings towards a fuller understanding of both biological mechanisms and biomarker potential.
Conclusions
In conclusion, we report for the first time patterns of gene expression and functional pathways in the normal rectal mucosa that are associated with circulating plasma vitamin D level. Oral vitamin D supplementation induces transcriptomic changes consistent with beneficial anti-tumour effects. Blood leukocyte expression of HIPK2 and PPP1CC predicted well those participants with the greatest expression response following supplementation. Whilst further replication in a separate cohort is desirable, these data provide compelling rationale for a trial of vitamin D and CRC prevention using easily assayed blood gene expression signatures as intermediate biomarkers of response.
Supplementary Information
Additional file 1. Full protocol for SCOVIDS study.
Additional file 2. Supplementary methods to be read as an adjunct to the main methods section in the manuscript.
Additional file 3: Table S1. Full list of diagnoses in recruited participants. Table S2. Top 50 ranked genes in PHASE 1. Table S3. Top 50 ranked GO terms enriched in PHASE 1. Table S4. Candidate genes positively associated with 25-OHD level in the GO term ‘regulation of cell migration’ and published evidence of tumour suppressor/ biomarker activity. Table S5. Candidate genes positively associated with 25-OHD level in the GO term ‘regulation of programmed cell death’ and published evidence of tumour suppressor/ biomarker activity. Table S6. Association between relevant genetic variants and 25OHD fold-change after supplementation. Table S7. Direction and magnitude of effect in candidate genes with expression change after supplementation. Table S8. Top ranked Enriched GO terms after vitamin D supplementation. Table S9. Enrichment of candidate GO terms after supplementation in RNAseq data. Table S10. Characteristics of participants with rectal mucosa response to supplementation. Table S11. AUC values for blood biomarkers of rectal and blood response in the SCOVIDS and BEST-D studies. Figure S1. Gene-set enrichment plot for enrichment of candidate genes associated with 25-OHD level in those undergoing vitamin D supplementation. Figure S2. GO terms enriched in PHASE 1 and PHASE 2. Figure S3. Receiver-operator curve for putative blood biomarkers and rectal mucosal response to supplementation.
Acknowledgements
We acknowledge statistical advice from Dr Catalina Vallejos, Biomedical Data Science research group, University of Edinburgh. We acknowledge the excellent technical support from Stuart Reid. We are grateful to Donna Markie, and all those who continue to contribute to recruitment, data collection and data curation for the Scottish Vitamin D Study. We acknowledge the expert support on sample preparation from the Genetics Core of the Edinburgh Wellcome Trust Clinical Research Facility in addition to the nursing and study facilities provided by the Clinical Research Facility.
Abbreviations
- 25-OHD
25-Hydroxyvitamin D
- AUC
Area under curve
- BEST-D
Biochemical Efficacy and Safety Trial of Vitamin D trial
- CRC
Colorectal cancer
- DNA
Deoxyribonucleic acid
- FC
Fold-change
- GO
Gene ontology
- NM
Normal mucosa
- PBMC
Peripheral blood mononuclear cells
- RCT
Randomised control trial
- RNA
Ribonucleic acid
- SCOVIDS
Scottish Vitamin D study
- SNP
Single-nucleotide polymorphism
- VDR
Vitamin D receptor gene
Authors’ contributions
Conceptualization, PGVS, MGD; Methodology, PGVS, LYO, GG, ET, FVND, SMF, MGD; Investigation, PGVS, LYO, JPB, GG, MT, MW, VS, KD; Writing-original draft PVS, SMF, MGD; Writing – review & editing, JPB, MT, SMF, FVND, MGD; Funding acquisition, MGD.; Resources, MGD; Supervision, SMF and MGD. SMF and MGD shared co-senior authorship. The authors read and approved the final manuscript.
Funding
The work reported in this manuscript was supported by a Cancer Research UK Programme Grant (C348/A18927) and a Project Leader Grant to MGD (MRC Human Genetics Unit Centre Grant - U127527198). MGD is an MRC Investigator. PVS was supported by a NES SCREDS clinical lectureship, MRC Clinical Research Training Fellowship (MR/M004007/1), a Research Fellowship from the Harold Bridges bequest and by the Melville Trust for the Care and Cure of Cancer. JPB is supported by an ECAT-linked CRUK ECRC Clinical training award (C157/A23218). FVND is supported by a CSO Senior Clinical Fellowship. LYO was supported by a CRUK Research Training Fellowship (C10195/A12996). The work was also supported by the infrastructure and staffing of the Edinburgh CRUK Cancer Research Centre.
Availability of data and materials
Transcriptomic profiling available at https://www.ncbi.nlm.nih.gov/geo/ GEO ID GSE157982 [79]. Full phenotypic data available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
All participants provided informed written consent, and research was approved by the local research ethics committees (13/SS/0248) and National Health Service management (2014/0058).
Consent for publication
Not required.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Theodoratou E, Tzoulaki I, Zgaga L, Ioannidis JP. Vitamin D and multiple health outcomes: umbrella review of systematic reviews and meta-analyses of observational studies and randomised trials. BMJ. 2014;348(apr01 2):g2035. doi: 10.1136/bmj.g2035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Autier P, Boniol M, Pizot C, Mullie P. Vitamin D status and ill health: a systematic review. Lancet Diabetes Endocrinol. 2014;2(1):76–89. doi: 10.1016/S2213-8587(13)70165-7. [DOI] [PubMed] [Google Scholar]
- 3.Barry EL, Peacock JL, Rees JR, Bostick RM, Robertson DJ, Bresalier RS, et al. Vitamin D receptor genotype, vitamin D3 supplementation, and risk of colorectal adenomas: a randomized clinical trial. JAMA Oncol. 2017;3(5):628-35. [DOI] [PMC free article] [PubMed]
- 4.Hermann J, Eder P, Banasiewicz T, Matysiak K, Lykowska-Szuber L. Current management of anal fistulas in Crohn's disease. Prz Gastroenterol. 2015;10(2):83–88. doi: 10.5114/pg.2015.49684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vaughan-Shaw PG, O'Sullivan F, Farrington SM, Theodoratou E, Campbell H, Dunlop MG, Zgaga L. The impact of vitamin D pathway genetic variation and circulating 25-hydroxyvitamin D on cancer outcome: systematic review and meta-analysis. Br J Cancer. 2017;116(8):1092–1110. doi: 10.1038/bjc.2017.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Vaughan-Shaw PG, Zgaga L, Ooi LY, Theodoratou E, Timofeeva M, Svinti V, Walker M, O'Sullivan F, Ewing A, Johnston S, et al. Low plasma vitamin D is associated with adverse colorectal cancer survival after surgical resection, independent of systemic inflammatory response. Gut. 2020;69(1):103–111. doi: 10.1136/gutjnl-2018-317922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Keum N, Lee DH, Greenwood DC, Manson JE, Giovannucci E. Vitamin D supplementation and total cancer incidence and mortality: a meta-analysis of randomized controlled trials. Ann Oncol. 2019;30(5):733–743. doi: 10.1093/annonc/mdz059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Vaughan-Shaw PG, Buijs LF, Blackmur JP, Theodoratou E, Zgaga L, Din FVN, Farrington SM, Dunlop MG. The effect of vitamin D supplementation on survival in patients with colorectal cancer: systematic review and meta-analysis of randomised controlled trials. Br J Cancer. 2020;123(11):1705–1712. doi: 10.1038/s41416-020-01060-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3(6):415–428. doi: 10.1038/nrg816. [DOI] [PubMed] [Google Scholar]
- 10.Bertucci F, Salas S, Eysteries S, Nasser V, Finetti P, Ginestier C, Charafe-Jauffret E, Loriod B, Bachelart L, Montfort J, Victorero G, Viret F, Ollendorff V, Fert V, Giovaninni M, Delpero JR, Nguyen C, Viens P, Monges G, Birnbaum D, Houlgatte R. Gene expression profiling of colon cancer by DNA microarrays and correlation with histoclinical parameters. Oncogene. 2004;23(7):1377–1391. doi: 10.1038/sj.onc.1207262. [DOI] [PubMed] [Google Scholar]
- 11.Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D, Levine AJ. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci U S A. 1999;96(12):6745–6750. doi: 10.1073/pnas.96.12.6745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chan SK, Griffith OL, Tai IT, Jones SJM. Meta-analysis of colorectal cancer gene expression profiling studies identifies consistently reported candidate biomarkers. Cancer Epidemiol Biomark Prev. 2008;17(3):543–552. doi: 10.1158/1055-9965.EPI-07-2615. [DOI] [PubMed] [Google Scholar]
- 13.Birkenkamp-Demtroder K, Christensen LL, Olesen SH, Frederiksen CM, Laiho P, Aaltonen LA, Laurberg S, Sorensen FB, Hagemann R, Orntoft TF. Gene expression in colorectal cancer. Cancer Res. 2002;62(15):4352–4363. [PubMed] [Google Scholar]
- 14.Dong X, Su YR, Barfield R, Bien SA, He Q, Harrison TA, Huyghe JR, Keku TO, Lindor NM, Schafmayer C, et al. A general framework for functionally informed set-based analysis: application to a large-scale colorectal cancer study. PLoS Genet. 2020;16(8):e1008947. doi: 10.1371/journal.pgen.1008947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bosse Y, Li Z, Xia J, Manem V, Carreras-Torres R, Gabriel A, Gaudreault N, Albanes D, Aldrich MC, Andrew A, et al. Transcriptome-wide association study reveals candidate causal genes for lung cancer. Int J Cancer. 2020;146(7):1862–1878. doi: 10.1002/ijc.32771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fleet JC, DeSmet M, Johnson R, Li Y. Vitamin D and cancer: a review of molecular mechanisms. Biochem J. 2012;441(1):61–76. doi: 10.1042/BJ20110744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Protiva P, Pendyala S, Nelson C, Augenlicht LH, Lipkin M, Holt PR. Calcium and 1,25-dihydroxyvitamin D3 modulate genes of immune and inflammatory pathways in the human colon: a human crossover trial. Am J Clin Nutr. 2016;103(5):1224–1231. doi: 10.3945/ajcn.114.105304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ross AC, Manson JE, Abrams SA, Aloia JF, Brannon PM, Clinton SK, Durazo-Arvizu RA, Gallagher JC, Gallo RL, Jones G, et al. The 2011 report on dietary reference intakes for calcium and vitamin d from the institute of medicine: what clinicians need to know editorial comment. Obstet Gynecol Surv. 2011;66(6):356–357. doi: 10.1097/OGX.0b013e31822c197a. [DOI] [Google Scholar]
- 19.Seamans KM, Cashman KD. Existing and potentially novel functional markers of vitamin D status: a systematic review. Am J Clin Nutr. 2009;89(6):1997s–2008s. doi: 10.3945/ajcn.2009.27230D. [DOI] [PubMed] [Google Scholar]
- 20.Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8(1):118–127. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- 21.Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14(4):417–419. doi: 10.1038/nmeth.4197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Soneson C, Love MI, Robinson MD. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 2015;4:1521. doi: 10.12688/f1000research.7563.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Durinck S, Spellman PT, Birney E, Huber W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat Protoc. 2009;4(8):1184–1191. doi: 10.1038/nprot.2009.97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Durinck S, Moreau Y, Kasprzyk A, Davis S, De Moor B, Brazma A, Huber W. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics. 2005;21(16):3439–3440. doi: 10.1093/bioinformatics/bti525. [DOI] [PubMed] [Google Scholar]
- 25.Wang TJ, Zhang F, Richards JB, Kestenbaum B, van Meurs JB, Berry D, Kiel DP, Streeten EA, Ohlsson C, Koller DL, Peltonen L, Cooper JD, O'Reilly PF, Houston DK, Glazer NL, Vandenput L, Peacock M, Shi J, Rivadeneira F, McCarthy MI, Anneli P, de Boer IH, Mangino M, Kato B, Smyth DJ, Booth SL, Jacques PF, Burke GL, Goodarzi M, Cheung CL, Wolf M, Rice K, Goltzman D, Hidiroglou N, Ladouceur M, Wareham NJ, Hocking LJ, Hart D, Arden NK, Cooper C, Malik S, Fraser WD, Hartikainen AL, Zhai G, Macdonald HM, Forouhi NG, Loos RJF, Reid DM, Hakim A, Dennison E, Liu Y, Power C, Stevens HE, Jaana L, Vasan RS, Soranzo N, Bojunga J, Psaty BM, Lorentzon M, Foroud T, Harris TB, Hofman A, Jansson JO, Cauley JA, Uitterlinden AG, Gibson Q, Järvelin MR, Karasik D, Siscovick DS, Econs MJ, Kritchevsky SB, Florez JC, Todd JA, Dupuis J, Hyppönen E, Spector TD. Common genetic determinants of vitamin D insufficiency: a genome-wide association study. Lancet. 2010;376(9736):180–188. doi: 10.1016/S0140-6736(10)60588-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Uitterlinden AG, Fang Y, Van Meurs JB, Pols HA, Van Leeuwen JP. Genetics and biology of vitamin D receptor polymorphisms. Gene. 2004;338(2):143–156. doi: 10.1016/j.gene.2004.05.014. [DOI] [PubMed] [Google Scholar]
- 27.Staal A, van Wijnen AJ, Birkenhager JC, Pols HA, Prahl J, DeLuca H, Gaub MP, Lian JB, Stein GS, van Leeuwen JP, et al. Distinct conformations of vitamin D receptor/retinoid X receptor-alpha heterodimers are specified by dinucleotide differences in the vitamin D-responsive elements of the osteocalcin and osteopontin genes. Mol Endocrinol. 1996;10(11):1444–1456. doi: 10.1210/mend.10.11.8923469. [DOI] [PubMed] [Google Scholar]
- 28.Yamamoto H, Miyamoto K, Li B, Taketani Y, Kitano M, Inoue Y, Morita K, Pike JW, Takeda E. The caudal-related homeodomain protein Cdx-2 regulates vitamin D receptor gene expression in the small intestine. J Bone Miner Res. 1999;14(2):240–247. doi: 10.1359/jbmr.1999.14.2.240. [DOI] [PubMed] [Google Scholar]
- 29.R Development Core Team . R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2013. [Google Scholar]
- 30.Knox S, Harris J, Calton L, Wallace AM. A simple automated solid-phase extraction procedure for measurement of 25-hydroxyvitamin D3 and D2 by liquid chromatography-tandem mass spectrometry. Ann Clin Biochem. 2009;46(Pt 3):226–230. doi: 10.1258/acb.2009.008206. [DOI] [PubMed] [Google Scholar]
- 31.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009;10(1):48. doi: 10.1186/1471-2105-10-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–287. doi: 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Berlanga-Taylor AJ, Plant K, Dahl A, Lau E, Hill M, Sims D, Heger A, Emberson J, Armitage J, Clarke R, Knight JC. Genomic response to vitamin D supplementation in the setting of a randomized, placebo-controlled trial. EBioMedicine. 2018;31:133–142. doi: 10.1016/j.ebiom.2018.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yanagisawa Y, Takeoka M, Ehara T, Itano N, Miyagawa S, Taniguchi S. Reduction of Calponin h1 expression in human colon cancer blood vessels. Eur J Surg Oncol. 2008;34(5):531–537. doi: 10.1016/j.ejso.2007.05.010. [DOI] [PubMed] [Google Scholar]
- 37.Sandberg TP, Oosting J, van Pelt GW, Mesker WE, Tollenaar R, Morreau H. Molecular profiling of colorectal tumors stratified by the histological tumor-stroma ratio - Increased expression of galectin-1 in tumors with high stromal content. Oncotarget. 2018;9(59):31502–31515. doi: 10.18632/oncotarget.25845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Xue J, Ge X, Zhao W, Xue L, Dai C, Lin F, Peng W. PIPKIgamma regulates CCL2 expression in colorectal cancer by activating AKT-STAT3 signaling. J Immunol Res. 2019;2019:3690561. doi: 10.1155/2019/3690561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lo PK, Lee JS, Liang X, Han L, Mori T, Fackler MJ, Sadik H, Argani P, Pandita TK, Sukumar S. Epigenetic inactivation of the potential tumor suppressor gene FOXF1 in breast cancer. Cancer Res. 2010;70(14):6047–6058. doi: 10.1158/0008-5472.CAN-10-1576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kops GJ, Medema RH, Glassford J, Essers MA, Dijkers PF, Coffer PJ, Lam EW, Burgering BM. Control of cell cycle exit and entry by protein kinase B-regulated forkhead transcription factors. Mol Cell Biol. 2002;22(7):2025–2036. doi: 10.1128/MCB.22.7.2025-2036.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sung JC, McCarthy S, Turner J, Li CG, Yeatman TJ. The NBL1 tumor suppressor gene is downregulated in colon cancer by promoter methylation. J Am Coll Surg. 2006;199(3):S92. doi: 10.1016/j.jamcollsurg.2004.05.200. [DOI] [Google Scholar]
- 42.Romano G, Santi L, Bianco MR, Giuffre MR, Pettinato M, Bugarin C, Garanzini C, Savarese L, Leoni S, Cerrito MG, et al. The TGF-beta pathway is activated by 5-fluorouracil treatment in drug resistant colorectal carcinoma cells. Oncotarget. 2016;7(16):22077–22091. doi: 10.18632/oncotarget.7895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Visuttijai K, Pettersson J, Mehrbani Azar Y, van den Bout I, Orndal C, Marcickiewicz J, Nilsson S, Hornquist M, Olsson B, Ejeskar K, et al. Lowered expression of tumor suppressor candidate MYO1C stimulates cell proliferation, suppresses cell adhesion and activates AKT. PLoS One. 2016;11(10):e0164063. doi: 10.1371/journal.pone.0164063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ha TK, Her NG, Lee MG, Ryu BK, Lee JH, Han J, Jeong SI, Kang MJ, Kim NH, Kim HJ, Chi SG. Caveolin-1 increases aerobic glycolysis in colorectal cancers by stimulating HMGA1-mediated GLUT3 transcription. Cancer Res. 2012;72(16):4097–4109. doi: 10.1158/0008-5472.CAN-12-0448. [DOI] [PubMed] [Google Scholar]
- 45.Badiola I, Olaso E, Crende O, Friedman SL, Vidal-Vanaclocha F. Discoidin domain receptor 2 deficiency predisposes hepatic tissue to colon carcinoma metastasis. Gut. 2012;61(10):1465–1472. doi: 10.1136/gutjnl-2011-300810. [DOI] [PubMed] [Google Scholar]
- 46.Boulagnon-Rombi C, Schneider C, Leandri C, Jeanne A, Grybek V, Bressenot AM, Barbe C, Marquet B, Nasri S, Coquelet C, Fichel C, Bouland N, Bonnomet A, Kianmanesh R, Lebre AS, Bouché O, Diebold MD, Bellon G, Dedieu S. LRP1 expression in colon cancer predicts clinical outcome. Oncotarget. 2018;9(10):8849–8869. doi: 10.18632/oncotarget.24225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wu JB, Sarmiento AL, Fiset PO, Lazaris A, Metrakos P, Petrillo S, Gao ZH. Histologic features and genomic alterations of primary colorectal adenocarcinoma predict growth patterns of liver metastasis. World J Gastroenterol. 2019;25(26):3408–3425. doi: 10.3748/wjg.v25.i26.3408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Dasgupta N, Kumar Thakur B, Chakraborty A, Das S. Butyrate-induced in vitro colonocyte differentiation network model identifies ITGB1, SYK, CDKN2A, CHAF1A, and LRP1 as the prognostic markers for colorectal cancer recurrence. Nutr Cancer. 2019;71(2):257–271. doi: 10.1080/01635581.2018.1540715. [DOI] [PubMed] [Google Scholar]
- 49.Jiang H, Dong L, Gong F, Gu Y, Zhang H, Fan D, Sun Z. Inflammatory genes are novel prognostic biomarkers for colorectal cancer. Int J Mol Med. 2018;42(1):368–380. doi: 10.3892/ijmm.2018.3631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Luque-Garcia JL, Martinez-Torrecuadrada JL, Epifano C, Canamero M, Babel I, Casal JI. Differential protein expression on the cell surface of colorectal cancer cells associated to tumor metastasis. Proteomics. 2010;10(5):940–952. doi: 10.1002/pmic.200900441. [DOI] [PubMed] [Google Scholar]
- 51.Nakajima H, Koizumi K, Tanaka T, Ishigaki Y, Yoshitake Y, Yonekura H, Sakuma T, Fukushima T, Umehara H, Ueno S, et al. Loss of HITS (FAM107B) expression in cancers of multiple organs: tissue microarray analysis. Int J Oncol. 2012;41(4):1347–1357. doi: 10.3892/ijo.2012.1550. [DOI] [PubMed] [Google Scholar]
- 52.Slattery ML, Lundgreen A, Wolff RK. MAP kinase genes and colon and rectal cancer. Carcinogenesis. 2012;33(12):2398–2408. doi: 10.1093/carcin/bgs305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ghadimi BM, Grade M, Difilippantonio MJ, Varma S, Simon R, Montagna C, Fuzesi L, Langer C, Becker H, Liersch T, et al. Effectiveness of gene expression profiling for response prediction of rectal adenocarcinomas to preoperative chemoradiotherapy. J Clin Oncol. 2005;23(9):1826–1838. doi: 10.1200/JCO.2005.00.406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Slattery ML, Lundgreen A, Bondurant KL, Wolff RK. Tumor necrosis factor-related genes and colon and rectal cancer. Int J Mol Epidemiol Genet. 2011;2(4):328–338. [PMC free article] [PubMed] [Google Scholar]
- 55.Meulmeester E, Jochemsen AG. p53: a guide to apoptosis. Curr Cancer Drug Targets. 2008;8(2):87–97. doi: 10.2174/156800908783769337. [DOI] [PubMed] [Google Scholar]
- 56.Ju YT, Kwag SJ, Park HJ, Jung EJ, Jeong CY, Jeong SH, Lee YJ, Choi SK, Kang KR, Hah YS, Hong SC. Decreased expression of heat shock protein 20 in colorectal cancer and its implication in tumorigenesis. J Cell Biochem. 2015;116(2):277–286. doi: 10.1002/jcb.24966. [DOI] [PubMed] [Google Scholar]
- 57.Kim J, Takeuchi H, Lam ST, Turner RR, Wang HJ, Kuo C, Foshag L, Bilchik AJ, Hoon DS. Chemokine receptor CXCR4 expression in colorectal cancer patients increases the risk for recurrence and for poor survival. J Clin Oncol. 2005;23(12):2744–2753. doi: 10.1200/JCO.2005.07.078. [DOI] [PubMed] [Google Scholar]
- 58.Zhou J, Liu M, Zhai Y, Xie W. The antiapoptotic role of pregnane X receptor in human colon cancer cells. Mol Endocrinol. 2008;22(4):868–880. doi: 10.1210/me.2007-0197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Browning SR, Browning BL. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet. 2007;81(5):1084–1097. doi: 10.1086/521987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Puca R, Nardinocchi L, Givol D, D'Orazi G. Regulation of p53 activity by HIPK2: molecular mechanisms and therapeutical implications in human cancer cells. Oncogene. 2010;29(31):4378–4387. doi: 10.1038/onc.2010.183. [DOI] [PubMed] [Google Scholar]
- 61.Kou Y, Zhang S, Chen X, Hu S. Gene expression profile analysis of colorectal cancer to investigate potential mechanisms using bioinformatics. Onco Targets Ther. 2015;8:745–752. doi: 10.2147/OTT.S78974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhou T, Lin W, Zhu Q, Renaud H, Liu X, Li R, Tang C, Ma C, Rao T, Tan Z, Guo Y. The role of PEG3 in the occurrence and prognosis of colon cancer. Onco Targets Ther. 2019;12:6001–6012. doi: 10.2147/OTT.S208060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zhou HM, Fang YY, Weinberger PM, Ding LL, Cowell JK, Hudson FZ, Ren M, Lee JR, Chen QK, Su H, Dynan WS, Lin Y. Transgelin increases metastatic potential of colorectal cancer cells in vivo and alters expression of genes involved in cell motility. BMC Cancer. 2016;16(1):55. doi: 10.1186/s12885-016-2105-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Galamb O, Kalmar A, Peterfia B, Csabai I, Bodor A, Ribli D, Krenacs T, Patai AV, Wichmann B, Bartak BK, et al. Aberrant DNA methylation of WNT pathway genes in the development and progression of CIMP-negative colorectal cancer. Epigenetics. 2016;11(8):588–602. doi: 10.1080/15592294.2016.1190894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bostick RM. Effects of supplemental vitamin D and calcium on normal colon tissue and circulating biomarkers of risk for colorectal neoplasms. J Steroid Biochem Mol Biol. 2015;148:86–95. doi: 10.1016/j.jsbmb.2015.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Manson JE, Cook NR, Lee IM, Christen W, Bassuk SS, Mora S, et al. Vitamin D supplements and prevention of cancer and cardiovascular disease. N Engl J Med. 2018. [DOI] [PMC free article] [PubMed]
- 67.Scragg R, Khaw KT, Toop L, Sluyter J, Lawes CMM, Waayer D, Giovannucci E, Camargo CA., Jr Monthly high-dose vitamin D supplementation and cancer risk: a post hoc analysis of the vitamin D assessment randomized clinical trial. JAMA Oncol. 2018;4(11):e182178. doi: 10.1001/jamaoncol.2018.2178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Chandler PD, Chen WY, Ajala ON, Hazra A, Cook N, Bubes V, Lee IM, Giovannucci EL, Willett W, Buring JE, Manson JAE, VITAL Research Group Effect of vitamin D3 supplements on development of advanced cancer: a secondary analysis of the VITAL randomized clinical trial. JAMA Netw Open. 2020;3(11):e2025850. doi: 10.1001/jamanetworkopen.2020.25850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Baron JA, Barry EL, Mott LA, Rees JR, Sandler RS, Snover DC, Bostick RM, Ivanova A, Cole BF, Ahnen DJ, Beck GJ, Bresalier RS, Burke CA, Church TR, Cruz-Correa M, Figueiredo JC, Goodman M, Kim AS, Robertson DJ, Rothstein R, Shaukat A, Seabrook ME, Summers RW. A trial of calcium and vitamin D for the prevention of colorectal adenomas. N Engl J Med. 2015;373(16):1519–1530. doi: 10.1056/NEJMoa1500409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Vaughan-Shaw PG, Zgaga L, Theodoratou E, Blackmur JP, Dunlop MG. Whether vitamin D supplementation protects against colorectal cancer risk remains an open question. Eur J Cancer. 2019;115:1–3. doi: 10.1016/j.ejca.2019.03.024. [DOI] [PubMed] [Google Scholar]
- 71.He Y, Timofeeva M, Farrington SM, Vaughan-Shaw P, Svinti V, Walker M, Zgaga L, Meng X, Li X, Spiliopoulou A, et al. Exploring causality in the association between circulating 25-hydroxyvitamin D and colorectal cancer risk: a large Mendelian randomisation study. BMC Med. 2018;16(1):142. doi: 10.1186/s12916-018-1119-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Cornish AJ, Law PJ, Timofeeva M, Palin K, Farrington SM, Palles C, Jenkins MA, Casey G, Brenner H, Chang-Claude J, Hoffmeister M, Kirac I, Maughan T, Brezina S, Gsur A, Cheadle JP, Aaltonen LA, Tomlinson I, Dunlop MG, Houlston RS. Modifiable pathways for colorectal cancer: a mendelian randomisation analysis. Lancet Gastroenterol Hepatol. 2020;5(1):55–62. doi: 10.1016/S2468-1253(19)30294-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Wagner D, Dias AG, Schnabl K, Van der Kwast T, Vieth R. Determination of 1,25-dihydroxyvitamin D concentrations in human colon tissues and matched serum samples. Anticancer Res. 2012;32(1):259–263. [PubMed] [Google Scholar]
- 74.Ryynanen J, Neme A, Tuomainen TP, Virtanen JK, Voutilainen S, Nurmi T, de Mello VD, Uusitupa M, Carlberg C. Changes in vitamin D target gene expression in adipose tissue monitor the vitamin D response of human individuals. Mol Nutr Food Res. 2014;58(10):2036–2045. doi: 10.1002/mnfr.201400291. [DOI] [PubMed] [Google Scholar]
- 75.Gerke AK, Pezzulo AA, Tang F, Cavanaugh JE, Bair TB, Phillips E, Powers LS, Monick MM. Effects of vitamin D supplementation on alveolar macrophage gene expression: preliminary results of a randomized, controlled trial. Multidiscip Respir Med. 2014;9(1):18. doi: 10.1186/2049-6958-9-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Hossein-nezhad A, Spira A, Holick MF. Influence of vitamin D status and vitamin D3 supplementation on genome wide expression of white blood cells: a randomized double-blind clinical trial. PLoS One. 2013;8(3):e58725. doi: 10.1371/journal.pone.0058725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Saksa N, Neme A, Ryynanen J, Uusitupa M, de Mello VD, Voutilainen S, Nurmi T, Virtanen JK, Tuomainen TP, Carlberg C. Dissecting high from low responders in a vitamin D3 intervention study. J Steroid Biochem Mol Biol. 2015;148:275–282. doi: 10.1016/j.jsbmb.2014.11.012. [DOI] [PubMed] [Google Scholar]
- 78.Buckmire M, Parquet G, Seeburger JL, Fukuchi SG, Rolandelli RH. Effect of bowel preparation and a fiber-free liquid diet on expression of transforming growth factor and procollagen in colonic tissue preoperatively and postoperatively. Dis Colon Rectum. 1998;41(10):1273–1280. doi: 10.1007/BF02258228. [DOI] [PubMed] [Google Scholar]
- 79.Vaughan Shaw PG, Grimes G, Dunlop MG: Scottish Colorectal Cancer Vitamin D Intervention Study (SCoViDS). GEO. 2021 https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?&acc=GSE157982.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1. Full protocol for SCOVIDS study.
Additional file 2. Supplementary methods to be read as an adjunct to the main methods section in the manuscript.
Additional file 3: Table S1. Full list of diagnoses in recruited participants. Table S2. Top 50 ranked genes in PHASE 1. Table S3. Top 50 ranked GO terms enriched in PHASE 1. Table S4. Candidate genes positively associated with 25-OHD level in the GO term ‘regulation of cell migration’ and published evidence of tumour suppressor/ biomarker activity. Table S5. Candidate genes positively associated with 25-OHD level in the GO term ‘regulation of programmed cell death’ and published evidence of tumour suppressor/ biomarker activity. Table S6. Association between relevant genetic variants and 25OHD fold-change after supplementation. Table S7. Direction and magnitude of effect in candidate genes with expression change after supplementation. Table S8. Top ranked Enriched GO terms after vitamin D supplementation. Table S9. Enrichment of candidate GO terms after supplementation in RNAseq data. Table S10. Characteristics of participants with rectal mucosa response to supplementation. Table S11. AUC values for blood biomarkers of rectal and blood response in the SCOVIDS and BEST-D studies. Figure S1. Gene-set enrichment plot for enrichment of candidate genes associated with 25-OHD level in those undergoing vitamin D supplementation. Figure S2. GO terms enriched in PHASE 1 and PHASE 2. Figure S3. Receiver-operator curve for putative blood biomarkers and rectal mucosal response to supplementation.
Data Availability Statement
Transcriptomic profiling available at https://www.ncbi.nlm.nih.gov/geo/ GEO ID GSE157982 [79]. Full phenotypic data available from the corresponding author on reasonable request.