Abstract
Samples from complete genomes of SARS-CoV-2 isolated during the first wave (December 2019–July 2020) of the global COVID-19 pandemic from 21 countries (Asia, Europe, Middle East and America) around the world, were analyzed using the phylogenetic method with molecular clock dating. Results showed that the first cases of COVID-19 in the human population appeared in the period between July and November 2019 in China. The spread of the virus into other countries of the world began in the autumn of 2019. In mid-February 2020, the virus appeared in all the countries we analyzed. During this time, the global population of SARS-CoV-2 was characterized by low levels of the genetic polymorphism, making it difficult to accurately assess the pathways of infection. The rate of evolution of the coding region of the SARS-CoV-2 genome equal to 7.3 × 10−4 (5.95 × 10−4–8.68 × 10−4) nucleotide substitutions per site per year is comparable to those of other human RNA viruses (Measles morbillivirus, Rubella virus, Enterovirus C). SARS-CoV-2 was separated from its known close relative, the bat coronavirus RaTG13 of the genus Betacoronavirus, approximately 15–43 years ago (the end of the 20th century).
Keywords: SARS-CoV-2, COVID-19, Genomics, Molecular clock, Evolutionary rate, Virus spread
1. Introduction
At the end of 2019, a massive outbreak of acute respiratory infection with complications and a large number of deaths was recorded in Wuhan, People's Republic of China. Analysis of clinical samples from patients in Wuhan, China, from December 23 to 26, 2019, showed that the disease was caused by a new, unknown RNA virus (Wu et al.,2020; Andersen et al., 2020). The first complete genome of the new virus was deposited in the GenBank database on January 5, 2020 (the length of the genome was 29,858, the length of the coding part was 29,265 nucleotides). The virus was genetically close to the subgenus Sarbecovirus of the genus Betacoronavirus, the family Coronaviridae. This group of viruses is known to cause severe acute respiratory syndrome (SARS) in humans and animals. The new virus was named SARS-CoV-2, and the name of the disease it causes was called COVID-19 (Velavan and Meyer, 2020). The genomic analysis of SARS-CoV-2 showed a close genetic relationship with the recently detected SARS-CoV coronavirus, that caused the SARS pandemic of 2002–2003 (Andersen et al., 2020; He et al., 2004) and MERS-CoV, which caused the outbreak of Middle East respiratory syndrome in 2013 in Saudi Arabia (Andersen et al., 2020; van Boheemen et al., 2012). Sequencing of the complete SARS-CoV-2 genome allowed us to understand the biological nature of the COVID-19 disease. Epidemiological data from the beginning of January 2020 showed a serious risk of a new infection for the human population.
In a short period of time, COVID-19 has spread from China to almost every country in the world (Chatterjee et al., 2020; Tabari et al., 2020). Media reports about the first cases of COVID-19 outside of China (other countries in Asia, Europe, America and Australia) became widespread in the second half of January 2020. On 30 January 2020, the World Health Organization (WHO) declared the COVID-19 outbreak as a global health emergency, and on 11 March 2020 it was declared as a worldwide pandemic. The problems of epidemiological studies of COVID-19 lie in the mass spread of asymptomatic and mild forms of the disease (Gao et al., 2020; (Zhao et al., 2020). Therefore, the early stages of the infection spread in the world, or in any country, could be hidden from epidemiologists.
Epidemiological data from the beginning of 2021 showed that globally, since March 2020, there have been two waves (two peaks in incidence) of COVID-19: the spring peak of 2020 and the autumn-winter peak of 2020–2021 (Cacciapaglia et al., 2020). Little epidemiological data was available from December 2019 to February 2020. Detailed analysis of the initial period of COVID-19 spread in the world would allow us to develop a strategy for preventing global pandemics of new viruses in the future.
Genomic phylogeny methods using Bayesian (Larget and Simon, 1999) and other approaches allow researchers to reconstruct the evolutionary history of virus pandemics using the concept of a molecular clock, dating the tree based on the date of strain isolation. It is generally assumed that the molecular clock method can give very approximate results, ranging in millions of years, but viruses mutate at a high rate. Mutations in the genomes of RNA viruses occur millions of times more often than in vertebrate genomes (Pybus and Rambaut, 2009). Therefore, a high accuracy molecular dating of evolutionary events up to several months and even days could be achieved. In fact, the use of Bayesian phylogenetic methods make it possible to identify patterns of virus evolution and distribution that cannot be obtained from epidemiological data (Worobey et al., 2016; Pettersson et al., 2018; Bradshaw et al., 2020) alone.
To date, a large number of complete genomes of various SARS-CoV-2 strains isolated during the global pandemic have been sequenced (Li et al., 2020). The genome-sequencing data are deposited in open GISAID (https://gisaid.org) (Shu and McCauley, 2017) and GenBank databases (https://www.ncbi.nlm.nih.gov/) (Benson et al., 2012).
Our study aimed to reconstruct the SARS-CoV-2 phylogenetic tree at the initial period of the COVID-19 pandemic by the Bayesian phylogenetic method with a molecular clock to identify the date of the first cases in the human population, to determine the origin (country) of the infection, and the time of the appearance and spread of infection in the world during the first wave of COVID-19; to detect the data of separation SARS-CoV-2 with close relatives of the genus Betacoronavirus, and estimate the mutation rate and genetic polymorphism of virus genomes.
2. Materials and methods
80,189 genome records on SARS-CoV-2 from the GISAID database accessed on 8 December 2020 (the first wave of the COVID-19 pandemic) were used in the analysis. From the total data set, sequences were selected according to the following: length greater than 29,000 nucleotides, approximately corresponding to the total length of the virus genome; less than 0.25% of unidentified nucleotides; strains isolated from diseased human patients; known information on strain isolation; and no stop codons within the open reading frame of viral proteins. From the selected genomes the protein coding regions were used for further analysis. The non-coding part of the genomes was not analyzed because of short length (about 592 nucleotides) compared to the length of the coding part of the genome and a large proportion of unknown nucleotides in the data.
The SARS-CoV-2 genomes from 21 countries around the world (China, South Korea, Thailand, Japan, USA, Mexico, Poland, Germany, Vietnam, France, Spain, Egypt, Israel, Greece, Turkey, Kazakhstan, Italy, Ukraine, Great Britain, Brazil, Russia) were selected for the analysis between December 2019 and July 2020. Countries were chosen based on availability of representative number of SARS-CoV-2 genome data in the databases. These countries were divided into seven regions: (1) China; (2) East Asia (South Korea, Japan); (3) Middle East and North Africa (Egypt, Israel, Turkey); (4) Southeast Asia (Thailand, Vietnam); (5) Europe (Poland, Germany, France, Spain, Greece, Italy, United Kingdom); (6) Post-Soviet countries (Russia, Ukraine, Kazakhstan); and (7) North and Latin America (USA, Mexico, Brazil). In total, three random samples of different sizes of genomes were applied: 248 complete genomes (on average 11 genomes from each country); 509 complete genomes (on average 24 genomes from each country); and 773 complete genomes (on average 36 genomes from each country). The second sample contained approximately twice as many genomes as the first; the third sample contained approximately three times the size of the first. For the analysis, sequences of four virus genomes isolated in China in December 2019, accessed from the GenBank database, were added to each of three generated random samples (252, 513 and 777 SARS-CoV-2 genomes data sets).
For each sample, a separate analysis was performed to determine the sample size effect on the results of phylogenetic reconstructions.
For each of the three differently sized samples, the uncalibrated phylogenetic tree was reconstructed by the maximum likelihood (ML) method in the IQTREE program (Nguyen et al., 2015). The 1 + 2 and 3 codon positions were treated differently in the analysis with the HKY + I + G nucleotide evolutionary model recommended for this phylogenetic reconstruction (recommended for RNA viruses) (Shapiro et al., 2006). A IQTREE preliminary analysis using the ModelFinder algorithm (Kalyaanamoorthy et al., 2017) showed that according to the value of the Bayesian information criterion (BIC), the reconstruction of the tree with differently treated codon positions has a significant advantage over the reconstruction without it. Statistical estimates of the reliability of the tree topology were performed in the IQTREE program using ultrafast bootstrap (1000 replicas) (Minh et al., 2013) and SH-aLRT (Guindon et al., 2010) analysis. The maximum likelihood tree was rooted using data on the strain isolation time by the "residual-mean-squared" method in the TempEst v1.5.3 program (Rambaut et al., 2016).
Phylogenetic analysis of the smallest sample of 252 genomes with molecular clock and calibrating the tree to the time of SARS-CoV-2 strain isolation was performed by the Bayesian phylogenetic method in the BEAST v. 2.6.2 software package (Bouckaert et al., 2014). The 1 + 2 and 3rd codon positions and the HKY + I + G DNA evolution model were tested with the following models: (1) constant population size, strict clock and dated tree; (2) constant population size, strict clock and not dated tree; (3) constant population size, relaxed clock and tip-dating tree - dating of the tree by the time of isolating strains; (4) constant population size, relaxed clock and not dated tree; (5) exponential growth population size, strict clock, tip-dating tree; (6) exponential growth population size, strict clock, not dated tree; (7) Exponential growth population size, relaxed clock, tip-dating tree; (8) exponential growth population size, relaxed clock and not dated tree. The uncorrelated lognormal relaxed clock model was used in the tree reconstruction with relaxed clock.
Testing of the best tree reconstruction model by comparison of the marginal likelihood estimators was provided in BEAST-2 with the “Path sampling” analysis (Baele et al., 2012) using the "Model-selection" package. This analysis was aimed to answer the following questions: (1) does reconstruction of a dated tree based on the virus isolation time have an advantage over reconstruction without time-calibration (in other words, does information about strain isolation contain a time signal for time-calibration of a tree?)?; (2) does tree reconstruction with a relaxed clock have an advantage over reconstruction with a strict clock (in other words, do all the SARS-CoV-2 lines in the human population accumulate mutations at the same rate?)?; (3) did the first wave SARS-CoV-2 pandemics provide exponential increase in the effective population size of the virus (increase in the current number of sick patients?)? This estimation scheme (Bayesian evaluation of temporal signal) of the reliability of molecular clock dating by the time of virus strain isolation using the log marginal likelihood calculated by the “Path sampling" method was held according to the recommendations of Duchene et al. (2020) where a log Bayes factor of at least 5 indicates “very strong” support for a dated model over not dated one.
For the datasets consisting of 513 and 777 genomes, the molecular clock analysis was performed in BEAST v. 2.6.2 using the best-fit reconstruction model of evolution chosen for 252 genomes dataset by the “Path sampling” analysis. This was due to the fact that the “Path sampling” analysis of 252 genomes required a lot of computational resources and time. Therefore, analyzing the datasets consisting of 513 and 777 genomes would be even more labor-intensive. We suppose that if the “Path sampling” analysis finds the applicability of dating of the phylogenetic tree by the time of virus strain isolation for a smaller sample size (252 genomes) then such dating is suitable for a dataset with a large sample size.
For all three datasets, samples from 1000 ultrafast bootstrap IQ-TREE ML trees were combined with topology of Bayesian consensus tree (BEAST). This allowed us to estimate the occurrence of nodes of the consensus of the Bayesian tree among the ultrafast bootstrap ML trees. Thus, the ultrafast bootstrap support was calculated for the Bayesian consensus tree. Comparison of trees and calculation of bootstrap support were carried out with the APE package (Paradis et al., 2004) for R statistical environment.
The phylogenetic tree was reconstructed by the date of virus isolation in BEAST v. 2.6.2. to identify the time period when SARS-CoV-2 split from its closest relatives in the genus Betacoronavirus. The coding regions of virus genomes of closely related species were added to three SARS-CoV-2 datasets (252, 513 and 777 genomes), each of which was analyzed separately to determine the sample size effect. Based on published data (Hul et al., 2021), four strains were selected as viruses that appeared at about the same time period as SARS-CoV-2: bat coronavirus RaTG13 (genome-wide sequence, NCBI accession number MN996532), isolated in 2013 (Andersen et al., 2020); bat coronavirus RmYN02 (two fragments of the complete genome, NCBI accession numbers MW201981 and MW201982) isolated in 2019 (Zhou et al., 2020); two strains of the bat coronavirus RShSTT182 and bat coronavirus RShSTT200 (genome-wide sequences, GISAID accession numbers EPI_ISL_852604 and EPI_ISL_852605) isolated in 2010 (Hul et al., 2021). For the comparative analysis, calculations in BEAST were carried out using evolutionary models with constant population size and exponential growth population size with uncorrelated lognormal relaxed clock.
3. Data availability
All supplementary materials such as the genomes with GISAID and GenBank numbers used in the study, the xml files for the BEAST v. 2.6.2 program, and original reconstructed phylogenetic trees are available at https://doi.org/10.6084/m9.figshare.14830899.
4. Results
Analysis of 252 SARS-CoV-2 genomes in the IQTREE program showed 547 polymorphic sites in the coding region, of which 112 sites were parsimony-informative. There were 277 polymorphic sites in the 1 + 2 codon position dataset, of which 52 were parsimony-informative. The 3rd codon positions had 365 polymorphic sites, of which 60 were parsimony-informative. The mutation rate in the 3rd codon positions was 30.05% higher than that in the 1 + 2 codon positions. The data set from 513 SARS-CoV-2 genomes in the coding region contained 1203 polymorphic sites, of which 219 sites were parsimony-informative (675 polymorphic sites in the 1 + 2 codon position dataset, of which 122 were parsimony-informative and 528 polymorphic sites, of which 97 were parsimony-informative). The data set from 777 SARS-CoV-2 genomes in the coding region contained 1667 polymorphic sites, of which 325 sites were parsimony-informative (938 polymorphic sites in the 1 + 2 codon position dataset, of which 179 were parsimony-informative and 729 polymorphic sites, of which 146 were parsimony-informative).
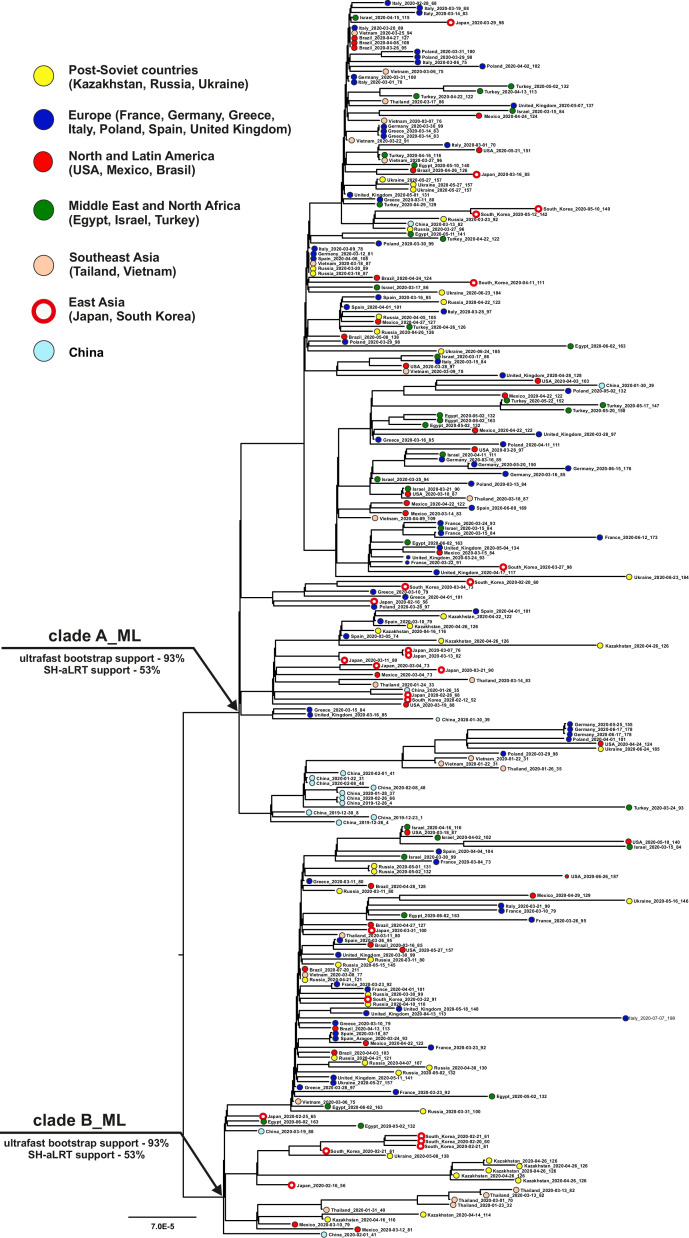
The rooted maximum likelihood tree for 252 SARS-CoV-2 genomes dataset is shown in Fig. 1 . The tree is divided into two large clades A_ML and B_ML in its basal part. For both clades (Fig. 1), the ultrafast bootstrap support values were <95%, and the SH-aLRT support values were <60%. These values (ultrafast bootstrap < 95%, SH-aLRT support < 80%) (Minh et al., 2013; Guindon et al., 2010) do not support reliable division of SARS-CoV-2 into independent phylogenetic lines in the period from the beginning of the pandemic to July 2020. The chosen method of rooting the tree, using information about the time of strain isolation, shows that SARS-CoV-2 genomes isolated in China in December 2019 were not the root of the tree and fall into the internal clade A_ML. This indicates that the virus began to circulate in the human population long before December 2019. None of the groups of genomes from any country forms its own monophyletic clade on the tree. Moreover, the genomes from 18 of the 21 countries occur in both the A_ML and B_ML clades, and the root nodes of the subtrees from the genomes in these 18 countries coincide with the root node of the whole tree. It suggests the penetration of SARS-CoV-2 into each country occurred independently from multiple sources. Similar conclusions can be made from the analysis of datasets consisting of 513 and 777 virus genomes. The topology of the rooted phylogenetic trees (see IQ-TREE_513_tree.tree and IQ-TREE_777_tree.tree files from supplementary materials) for these datasets was similar to the topology of the tree in Fig. 1, dividing the entire sequence set into two clusters, each with low ultrafast bootstrap and SH-aLRT supports. The virus genomes isolated in any of the countries did not form monophyletic clusters, supporting the conclusion about the entry of the virus into each of the countries from several independent sources.
Fig. 1.
The undated Maximal Likelihood tree (ML) reconstructed in the IQTREE program. The tree was rooted based on time of the virus isolation with the “residual-mean-squared” method in the TempEst program. The tips names contain information about the country of isolation of the strain, the time of isolation of the strain and the number of days that have passed since the isolation of the first genome.
The Markov chain Monte Carlo (MCMC) calculations for 252 SARS-CoV-2 genomes dataset in BEAST were carried out in triplicates with 0.6 × 109 generations, saving the results at every 30,000. Only this scheme of MCMC analysis allowed us to achieve a stable value of ESS statistics, 200 units, for all parameters of the reconstructions. For the “Path sampling” method calculations, analysis was run in triplicate with 300 steps and the chain length of 2,000,000 generations (MCMC) each as recommended in the tutorial (https://github.com/BEAST2-Dev/beast-docs/releases/download/v1.0/BFD-tutorial-2017.zip). The results of the "Path sampling" analysis (Table 1 ) showed that the reconstructed evolutionary tree (dated to the time of strain isolation with an exponential increase in the effective population size of the virus and a relaxed molecular clock in all three replicates) had a higher log marginal likelihood value. This model for a tree reconstruction was shown to be the best fit for the analysis. The "Path sampling" analysis proved that: (1) information on the virus strain isolation time has a signal for time-calibration of a phylogenetic tree; (2) the mutation rates in various SARS-CoV-2 lines differed from each other; (3) during the first wave of the COVID-19 pandemic until July 2020, there was an increase in the effective population size of the virus (and a simultaneous increase in the number of sick patients). The "Path sampling" analysis results indicates that the evolutionary model with exponential growth population size, relaxed clock, tip-dating tree is suitable for the 513 and 777 genomes data sets. The analysis based on this model required 1.2 × 109 generations (saving the results at every 30,000 generations) for the dataset of 513 genomes and 1.9 × 109 generations (saving the results at every 30,000 generations) for the dataset of 777 genomes to achieve a stable value of ESS statistics 200 units for all parameters of the reconstructions.
Table 1.
Path sampling analysis results for testing phylogenetic tree reconstruction models.
| Evolutionary model | Run 1 log marginal likelihood value | Run 2 log marginal likelihood value | Run 3 log marginal likelihood value |
| Сonstant population size, strict clock, dated tree | −46094.203 | −46090.993 | −46091.278 |
| Сonstant population size, strict clock, not dated tree | −46184.348 | −−46182.243 | −46176.5 |
| Сonstant population size, relaxed clock, dated tree | −46086.055 | −46088.769 | −46087.755 |
| Сonstant population size, relaxed clock, not dated tree | −46181.297 | −46179.925 | 46181.091 |
| Exponential growth population size, strict clock, dated tree | −46086.246 | −46083.521 | −46084.368 |
| Exponential growth population size, strict clock, not dated tree | −46177.031 | −46177.089 | −46177.638 |
| Exponential growth population size, relaxed clock, dated tree | −46080.018 | −46080.919 | −46080.425 |
| Exponential growth population size, relaxed clock, not dated tree | −46177.998 | −46178.772 | −46177.865 |
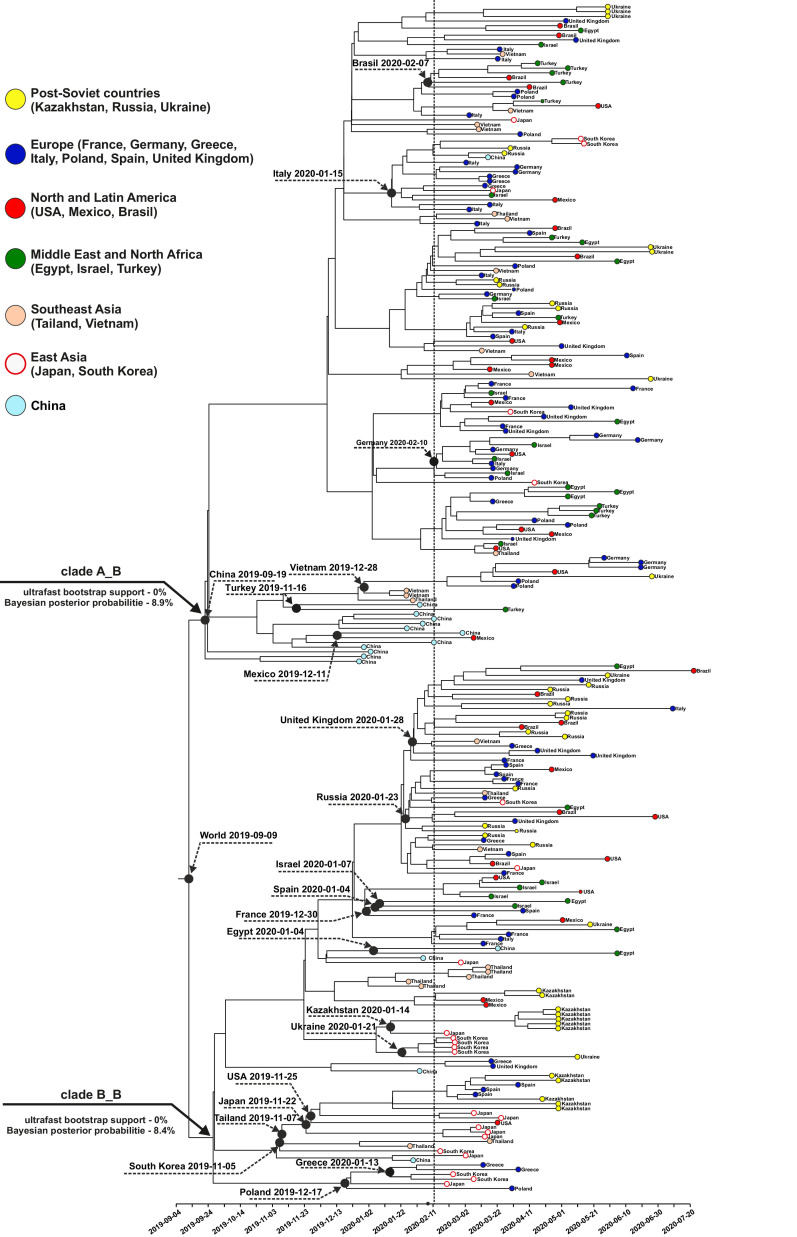
There are two distinguished clades A_B and B_B in the basal part on the Bayesian phylogenetic tree (Fig. 2 ), reconstructed according to the best fit DNA model chosen for the dataset of 252 SARS-CoV-2 genomes. These two clades are similar to those on the maximum likelihood tree. Statistical support for these clades, calculated with ultrafast bootstrap analysis, and Bayesian posterior probabilities are close to zero. Groups of genomes from the 252 SARS-CoV-2 genomes datasets from different countries of the world on both the Bayesian and maximum likelihood trees do not form monophyletic clades. The genomes isolated in most countries of the world (18 out of 21) simultaneously belong to both clades A_B and B_B. The root nodes of the subtrees of these countries coincided with the root of the entire tree. It should be noted that the composition of the A_B and B_B clades on the Bayesian tree differs from that on the maximum likelihood tree. This suggests that the clustering order of genomic sequences in the basal part of the phylogenetic tree is unstable and depends on the choice of the clustering method. Besides low statistical support of nodes on the tree, the variability of the clustering which depends on the clustering method, confirms the conclusion that from the beginning of the pandemic to July 2020, genomic data do not allow us to find stable phylogenetic lineages of SARS-CoV-2. The dated Bayesian phylogenetic trees, based on 513 and 777 genomes datasets (see BEAST_Exp_RC_513_dated_tree.tree and BEAST_Exp_RC_777_dated_tree.tree files from supplementary materials), in the basal part do not have any resolved clustering and clades with high support values of topology. The order of clustering resembles a ladder structure; on the tree, phylogenetic SARS-CoV-2 lines from different countries are evenly distributed across different clusters. In the Bayesian phylogenetic tree based on 513 genomes dataset, the root is a clade containing two genomes: first, the SARS-CoV-2 genome isolated in December 2019 in China, and second, genome isolated in January 2020 in Thailand. On the Bayesian phylogenetic tree based on 777 genomes, the root is a single genome clade isolated in January 2020 in China.
Fig. 2.
The dated phylogenetic tree of SARS-CoV-2 was reconstructed based on complete coding genome regions of viruses isolated from December 2019 to July 2020. Different world regions are marked with colored circles. Dating of the appearance of the first phylogenetic SARS-CoV-2 lineages in each of 21 studied countries is shown at the nodes.
A comparison of the topologies of the Bayesian phylogenetic trees based on 252, 513 and 777 genomes shows that, due to low topology support values, changes in the dataset (using other virus genomes for analysis) result in chaotic changes in the clustering order of the analyzed SARS-CoV-2 genomes taken for analysis from the first wave of the COVID-19 pandemic.
Analysis of the time-calibrated Bayesian phylogenetic tree based on 252 SARS-CoV-2 genomes showed (Fig. 2, Table 2 ) that the common ancestor of COVID-19 in the human population appeared in the period from 21 July 2019 to 27 October 2019. Reconstructions based on 513 and 777 genomes (Table 3 ) indicate a similar time period (Table 3). Notably, an increase in the sample size does not lead to a narrowing of the confidence interval (an increase in the accuracy of analysis) of the estimated lifetime of the common ancestor of all SARS-CoV-2 phylogenetic lines circulating during the first wave of the pandemic. On the contrary, there is a slight enlargement of confidence interval boundaries when applying the largest dataset of 777 genomes. On the dated Bayesian phylogenetic tree for each country (Fig. 2), there is a clade (or single genome) – a phylogenetic lineage that separated from the ancestral node together with genomes from other countries at the earliest point in time. With some approximation, we can suggest that the time of separation of this phylogenetic lineage from the nearest ancestor is the time of the appearance of the SARS-CoV-2 virus in a certain country. Examples of such phylogenetic lineages are given on the Bayesian phylogenetic tree based on 252 SARS-CoV-2 genomes (Fig. 2). The results of the time-calibrated divergence of the earliest phylogenetic lineages for all datasets (252, 513, and 777 SARS-CoV-2 genomes), and confidence intervals, are shown in Fig. 3 and Table 2. The result of analysis of all three datasets shows that the earliest phylogenetic lineage of SARS-CoV-2 (Fig. 3, Table 3) originated from China (summer-autumn 2019). In the autumn of 2019, local phylogenetic lineages of the virus appeared in Thailand, the USA, Mexico, Japan, and Vietnam. For some datasets, the confidence intervals of the appearance of phylogenetic lineages of the virus in these and other countries overlap with the time period calculated for China (Fig. 3, Table 2). Results based on all datasets (252, 513 and 777 SARS-CoV-2 genomes) showed (Fig. 3, Table 2) that in the autumn of 2019 the SARS-CoV-2 virus had already been introduced to Asia, America, the Middle East and Europe. The global spread of SARS-CoV-2 had already begun in the autumn of 2019. By mid-February 2020, the virus had spread across almost all countries of the world. Significantly, the results of calculated time for the appearance of the first SARS-CoV-2 lineages into each of the countries are unstable and highly dependent on the choice of genomes used in the analysis.
Table 2.
Time of appearance of the first lineage of the SARS-CoV-2 in each of the studied countries.
| Country | Run with genomes sample size | Appearance of the earliest lineage | Upper value 95% confidence interval | Lower value 95% confidence interval |
| The whole world | 252 | 2019-09-09 | 2019-07-21 | 2019-10-27 |
| 513 | 2019-09-19 | 2019-08-04 | 2019-10-29 | |
| 777 | 2019-09-05 | 2019-07-03 | 2019-10-26 | |
| China | 252 | 2019-09-19 | 2019-08-16 | 2019-10-25 |
| 513 | 2019-10-18 | 2019-09-13 | 2019-11-10 | |
| 777 | 2019-09-05 | 2019-07-03 | 2019-10-26 | |
| Thailand | 252 | 2019-11-07 | 2019-10-12 | 2019-12-02 |
| 513 | 2019-10-22 | 2019-09-14 | 2019-11-22 | |
| 777 | 2019-09-08 | 2019-08-26 | 2019-09-10 | |
| USA | 252 | 2019-11-25 | 2019-10-23 | 2019-12-23 |
| 513 | 2019-10-24 | 2019-09-27 | 2019-11-13 | |
| 777 | 2019-10-13 | 2019-09-18 | 2019-11-03 | |
| Mexico | 252 | 2019-12-11 | 2019-11-11 | 2020-01-06 |
| 513 | 2019-10-25 | 2019-09-28 | 2019-11-17 | |
| 777 | 2019-10-23 | 2019-09-25 | 2019-11-11 | |
| Japan | 252 | 2019-11-22 | 2019-10-23 | 2019-12-20 |
| 513 | 2019-10-27 | 2019-09-27 | 2019-11-16 | |
| 777 | 2019-11-19 | 2019-10-24 | 2019-12-09 | |
| Vietnam | 252 | 2019-12-28 | 2019-12-09 | 2020-01-13 |
| 513 | 2019-12-23 | 2019-11-30 | 2020-01-31 | |
| 777 | 2019-10-07 | 2019-10-04 | 2019-10-26 | |
| Turkey | 252 | 2019-11-16 | 2019-10-11 | 2020-01-03 |
| 513 | 2019-12-25 | 2019-11-29 | 2020-01-17 | |
| 777 | 2019-11-22 | 2019-10-22 | 2019-12-18 | |
| Germany | 252 | 2020-02-10 | 2020-01-23 | 2020-02-16 |
| 513 | 2019-10-24 | 2019-10-01 | 2019-11-19 | |
| 777 | 2019-11-29 | 2019-11-20 | 2019-12-02 | |
| South Korea | 252 | 2019-11-05 | 2019-10-06 | 2019-12-03 |
| 513 | 2019-12-24 | 2019-11-27 | 2020-01-18 | |
| 777 | 2020-01-04 | 2019-12-11 | 2020-01-25 | |
| Poland | 252 | 2019-12-17 | 2019-11-09 | 2020-01-20 |
| 513 | 2019-12-24 | 2019-12-22 | 2019-12-25 | |
| 777 | 2019-11-29 | 2019-11-20 | 2019-12-02 | |
| Italy | 252 | 2020-01-15 | 2020-01-08 | 2020-01-28 |
| 513 | 2019-12-24 | 2019-12-22 | 2019-12-25 | |
| 777 | 2019-11-23 | 2019-11-19 | 2019-12-08 | |
| United Kingdom | 252 | 2020-01-28 | 2020-01-21 | 2020-02-13 |
| 513 | 2020-01-17 | 2019-12-27 | 2020-02-03 | |
| 777 | 2019-11-15 | 2019-10-23 | 2019-12-07 | |
| Egypt | 252 | 2020-01-04 | 2019-12-15 | 2020-01-17 |
| 513 | 2019-12-30 | 2019-12-12 | 2020-01-13 | |
| 777 | 2020-01-14 | 2019-12-28 | 2020-01-26 | |
| Russia | 252 | 2020-01-23 | 2019-12-20 | 2020-01-29 |
| 513 | 2019-12-18 | 2019-12-16 | 2019-12-23 | |
| 777 | 2020-01-19 | 2020-01-14 | 2020-01-21 | |
| Greece | 252 | 2020-01-13 | 2019-12-17 | 2020-02-04 |
| 513 | 2019-12-26 | 2019-12-05 | 2020-01-15 | |
| 777 | 2020-01-25 | 2020-01-24 | 2020-01-26 | |
| Spain | 252 | 2020-01-04 | 2019-12-10 | 2020-01-24 |
| 513 | 2020-01-18 | 2019-12-28 | 2020-02-03 | |
| 777 | 2020-01-25 | 2020-01-17 | 2020-02-04 | |
| France | 252 | 2019-12-30 | 2019-12-11 | 2020-01-16 |
| 513 | 2020-01-26 | 2020-01-10 | 2020-02-07 | |
| 777 | 2020-01-28 | 2020-01-18 | 2020-02-04 | |
| Brazil | 252 | 2020-02-07 | 2020-02-03 | 2020-02-13 |
| 513 | 2019-12-18 | 2019-12-16 | 2019-12-23 | |
| 777 | 2020-01-30 | 2020-01-18 | 2020-02-04 | |
| Israel | 252 | 2020-01-07 | 2019-12-15 | 2020-01-27 |
| 513 | 2020-01-31 | 2020-01-12 | 2020-02-12 | |
| 777 | 2020-01-29 | 2020-01-24 | 2020-02-06 | |
| Kazakhstan | 252 | 2020-01-14 | 2019-12-20 | 2020-02-07 |
| 513 | 2020-01-18 | 2019-12-28 | 2020-02-03 | |
| 777 | 2020-02-05 | 2020-01-20 | 2020-02-20 | |
| Ukraine | 252 | 2020-01-21 | 2020-01-02 | 2020-02-09 |
| 513 | 2020-01-25 | 2020-01-07 | 2020-02-10 | |
| 777 | 2020-01-27 | 2020-01-11 | 2020-02-11 |
Table 3.
Estimates of the evolution rates in the coding part of the genomes of the SARS-CoV-2.
| Substitution type | Run with genomes sample size | Evolution rate - nucleotide substitutions per site per year (95% confidence interval) | Average number of substitutions per month for the coding part of the genome (95% confidence interval) |
| 1 + 2 codon positions | 252 | 6.02 × 10−4 4.98 × 10−4– 7.16 × 10−4 |
0.98 0.81–1.16 |
| 513 | 6.66 × 10−4 5.51 × 10−4–7.92 × 10−4 |
1.08 0.89–1.27 |
|
| 777 | 5.97 × 10−4 4.83 × 10−4–7.18 × 10−4 |
0.97 0.78–1.163 |
|
| 3 codon position | 252 | 8.59 × 10−4 6.92 × 10−4–10.2 × 10−4 |
0.7 0.56–0.83 |
| 513 | 9.53 × 10−4 7.68 × 10−4––11.32 × 10−4 |
0.77 0.62–0.92 |
|
| 777 | 8.58 × 10−4 6.91 × 10−4 - 10.4 × 10−4 |
0.699 0.559 - 0,85 |
|
| 1 + 2 + 3 codon positions | 252 | 7.31 × 10−4 5.95 × 10−4–8.68 × 10−4 |
1.68 1.37–1.99 |
| 513 | 8.11 × 10−4 6.60 × 10−4–9.63 × 10−4 |
1.86 1.52–2.07 |
|
| 777 | 7.26 × 10−4 5.87 × 10−4–8.79 × 10−4 |
1.67 1.35–2.02 |
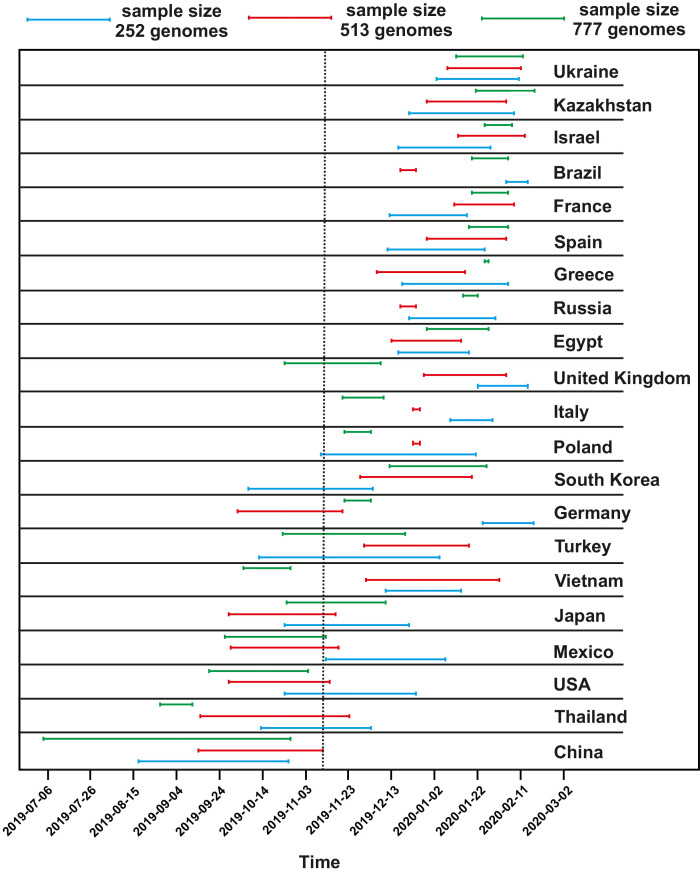
Fig. 3.
The distribution of confidence intervals on the time scale shows the time periods of the appearance of SARS-CoV-2 first phylogenetic lineages in studied countries. Results are based on the datasets of 252, 513 and 777 SARS-CoV-2 genomes. The black dotted vertical bar separates the countries’ confidence intervals, which intersect with the confidence interval calculated for China at the initial period of the COVID-19 pandemic.
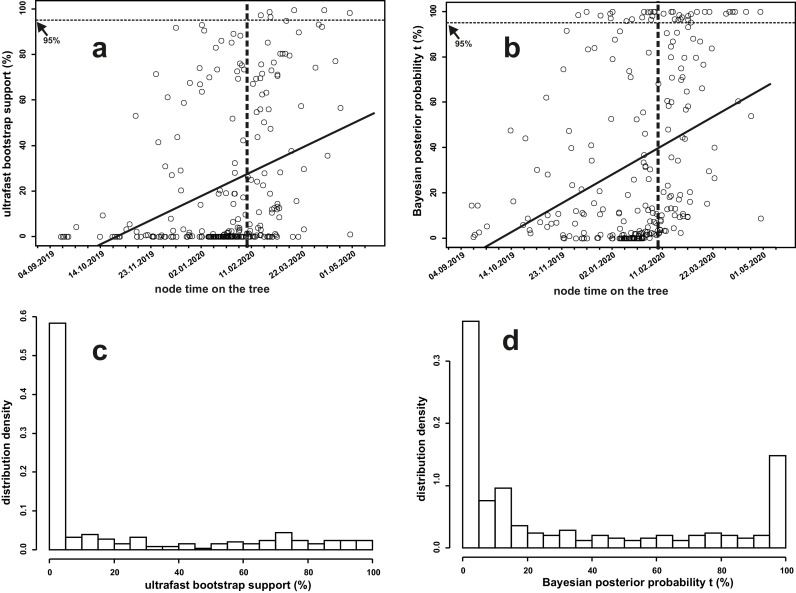
Genomic data can be used to track a path of the virus from one country to another based on the topology of a phylogenetic tree and time estimates inferred from the date of virus isolation. To track the virus pathway, high support of the tree topology is required. For the dataset of 252 SARS-CoV-2 genomes, Fig. 4 shows the correlation between the bootstrap/probability value on the Bayesian phylogenetic tree and the time estimator at the node (Fig. 4a - ultrafast bootstrap support, Fig. 4b - Bayesian posterior probability). Analysis of this correlation shows that the percentage of ultrafast bootstrap support and Bayesian posterior probability values ≥ 95% increases from the root of the tree to the end of branches. Only slightly more than 10% of the nodes in the tree exceed the 95% confidence threshold (Fig. 4a,b). For the ultrafast bootstrap supported tree, all these nodes appear after 13 February 2020, when the SARS-CoV-2 had already spread around the world. From the analysis of Bayesian posterior probabilities, less than half of the nodes (16 out of 37) with support values greater than 95%wwwww occurred before 13 February 2020. This distribution of support values at time-calibrated nodes on the tree demonstrate that SARS-CoV-2 genome data do not provide a sufficient signal to track the spread and path of the virus without accurate epidemiological data on the movement of virus carriers between countries. There are similar distributions of support values on the Bayesian phylogenetic trees based on datasets of 513 and 777 virus genomes (see BEAST_Exp_RC_513_dated_tree.tree, BEAST_Exp_RC_777_dated_tree.tree, BEAST_Exp_RC_dated_513_tree_with_ultrafast_bootstrap_supports.tree and BEAST_Exp_RC_dated_777_tree_with_ultrafast_bootstrap_supports.tree files from supplementary materials). An increase in sample size does not improve the support values of a topology of the reconstructed phylogenetic trees. Average mutation rates calculated on the basis of the SARS-CoV-2 coding region (datasets of 252, 513, and 777 genomes) are shown in Table 3 For all data sets, average values of mutation rates and confidence interval measurements differ less than 11%. The maximal variability of values (the largest confidence interval) was observed in the analysis of the dataset consisting of 777 SARS-CoV-2 genomes (Table 3).
Fig. 4.
Assessment of the relationship between the supports of the topology of nodes and the time of node appearance on the dated Bayesian phylogenetic tree (a - ultrafast bootstrap support, b - Bayesian posterior probability) and histograms of the distribution of the supports of the tree topology (c - ultrafast bootstrap support, d - Bayesian posterior probability).
An enlargement of the sample size more than threefold (from 252 to 777 genomes) does not increase the accuracy of the analysis. On average, in the coding part of the genome, one SARS-CoV-2 lineage circulating in the human population can accumulate 1.35 – 2.02 nucleotide substitutions per month. Mutations in the 1 + 2 codon positions, as a rule, lead to amino acid changes. Therefore, SARS-CoV-2 lineages could accumulate in average of 0.78–1.163 amino acid substitutions per month. The actual number of nucleotide and amino acid substitutions accumulated by the SARS-CoV-2 phylogenetic lineage will be a random variable distributed according to Poisson's law. With the estimated average mutation rates, the probability that the virus genome will not accumulate substitutions for a month of circulation in the human population is 0.156 for nucleotide and 0.34 for amino acid substitutions. The spread of SARS-CoV-2 from its origin around the world has occurred during two and a half months. During this time, only a few, if any, substitutions were accumulated in the SARS-CoV-2 genomes circulated in the human population. Therefore, the estimated rates of accumulation of substitutions confirm the conclusion that the pathways of the spread and movement of the virus between countries cannot be traced on the basis of complete genomes alone.
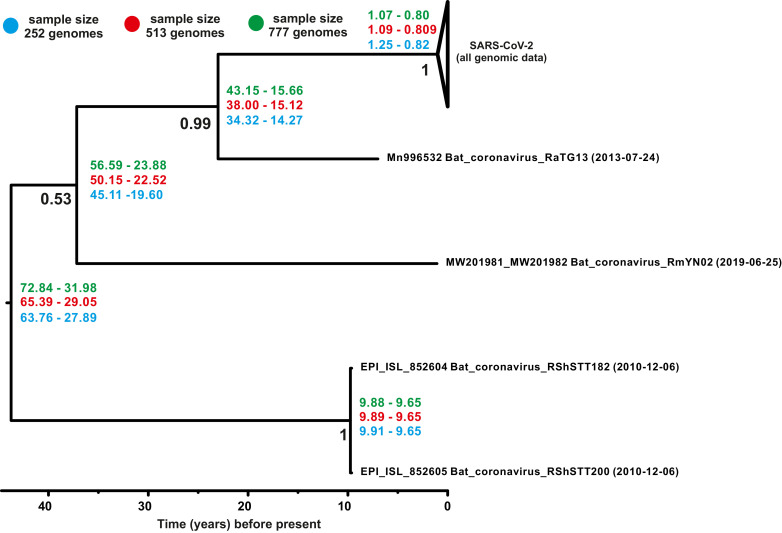
A dendrogram of the Bayesian time-calibrated phylogenetic tree including the genomes of SARS-CoV-2 and the most evolutionary close members of Betacoronavirus is shown in Fig. 5 . The clustering order of SARS-CoV-2 and Betacoronavirus strains does not depend on the sample size (datasets of 252, 513 and 777 virus genomes), nor the choice of the constant population size, nor exponential growth population size evolutionary model; the topology of the tree has always been consistent with that shown on Fig. 5. The use of the exponential growth population size model is preferable in this case since most of the genomes characterize a pandemic with an exponentially increasing effective population size of the virus. Estimated data indicate the split of SARS-CoV-2 and its closest relative, bat coronavirus RaTG13, from their common ancestor from 15 to 43 years ago (late 20th–early 21st century). The split of all analyzed closely related viruses happened 31–72 years ago (the second half of 20 century). The use of evolutionary model with constant population size narrows down confidence intervals (see BEAST_Betacoronavirus_tree_const.pdf files from supplementary materials) and reduce a nodes’ age (the split of SARS-CoV-2, Bat coronavirus RaTG13 and its closest relative occurred from 13 to 23 years ago, and for all dataset of closely related viruses – from 24 to 47 years ago). Reconstructions using two evolutionary models (constant population size and exponential growth population size) show intersection of confidence intervals of dating of nodes on the tree. Notably, in the analysis that included closely related strains (Fig. 5), the left border of the confidence interval for dating the divergence of all SARS-CoV-2 strains from their common ancestor (the date of existence of the closest common ancestor of the SARS-CoV-2 strains circulating in the first wave of the pandemic) moved back by two-three months (to the first half of 2019). As in the analysis without closely related strains, an increase in the sample size of SARS-CoV-2 genomes by more than three times did not lead to an increase in the accuracy of the analysis (narrowing of the confidence intervals for dating of nodes).
Fig. 5.
Dated evolutionary tree of SARS-CoV-2, and closely related viruses of the genus Betacoronavirus, based on the complete coding regions of their genomes, was reconstructed applying exponential growth population size and relaxed clock with tip-dating (dating of the tree by the time of isolating strains). Numbers at nodes indicate statistical support values of tree topology and confidence intervals of molecular dating.
5. Discussion
Our results are consistent with the initial conclusions of some experts (Wu et al., 2020; Andersen et al., 2020; Velavan and Meyer, 2020) that the SARS-CoV-2 variant spreading during the first wave of the pandemic in the human population appeared in China in mid-2019. The initial stage of the pandemic has also been in China. However, the rapid spread of SARS-CoV-2 on a global scale (the Asian countries, the Middle East, the United States) had already begun in the autumn of 2019, probably before the first official reports from China about a new viral infection. In mid-February 2020, the SARS-CoV-2 spread around the world, and COVID-19 became a pandemic. This does not coincide with official media reports on the first cases of infection in a number of listed countries in March 2020. Most likely, these asymptomatic and mild forms of the disease allowed the virus to spread widely in these countries even before the introduction of mass diagnostics and preventive measures. Our conclusions on the mass spread of SARS-CoV-2 in the autumn of 2019 are confirmed by results of several retrospective studies on the detection of SARS-CoV-2 antibodies in patients’ blood samples: (1) in Italy in the autumn of 2019 (Apolone et al., 2020); (2) in donated blood in the USA in December 2019 (Basavaraju et al., 2020); (3) in France in November 2019 (Carrat et al., 2021). The WHO statements from 30 January 2020 on the emergency and threat of COVID-19, and reports on the beginning of the world pandemic on 11 March 2020, were made with one and a half to two months delay as shown by the results of the genomic data analysis. Therefore, the global community needs to review the standards of action to prevent new pandemics of infectious diseases.
Considering the hypothesis 'patient zero' (the first infected human), our results show that SARS-CoV-2 first was transmitted from animals to humans in the middle of 2019. Possibly, at the initial point, more than one person was infected. Therefore several genetically different strains of the virus appeared in the human population. In this case, our results indicate the time period of the existence of the closest common ancestor for this first-appeared group of strains. In this scenario, the time period when the pandemic began shifts to the end of 2019. So far, there is no definite evidence of the beginning of the 2019 COVID-19 pandemic due to the transmission of the virus from animals to humans. It is possible that SARS-CoV-2 circulated in the human population earlier. On that assumption, a mutation process resulted in the appearance of the more virulent and pathogenic strain of SARS-CoV-2 in China in mid-2019 that overcame the immune response developed in the human population against infection with previous strains of SARS-CoV-2. It was this new strain of SARS-CoV-2 that started the COVID-19 pandemic, displacing less virulent strains in the human population. This scenario could explain early detection of SARS-CoV-2 antibodies (September–November 2019) in human blood (Apolone et al., 2020; Carrat et al., 2021) in European countries. Detected SARS-CoV-2 antibodies were traces of the circulation of other, earlier variants of SARS-CoV-2, unnoticed by epidemiologists. In this case, our dating (summer-autumn 2019) indicates the point of the emergence of a new, more pathogenic and virulent strain of SARS-CoV-2, which caused the COVID-19 pandemic. Only a molecular clock dating of SARS-CoV-2 genomes from the wild animals’ populations, or sequencing of the SARS-CoV-2 genomes from human blood samples collected from September to November 2019, can confirm or deny these assumptions. All our results confirm the fact that the SARS-CoV-2 variant of the first wave of the pandemic has spread globally in a very short time – from late November 2019 to mid-February 2020 (during two and a half months).
The mutation rate of SARS-CoV-2 is less than or comparable to the rate of evolution of such RNA-containing viruses as measles morbillivirus (Furuse et al., 2010), rubella virus (Zhu et al., 2012), and poliovirus (Enterovirus C) (Smura et al., 2014). In our study, the calculations of the rate of evolution were only applied to the whole virus genome. Recent studies have shown (Garvin et al., 2020; Wang et al., 2021) that some parts of the gene encoding the surface protein S of the virus undergo natural selection, with the effect of increasing the mutation rate. Due to this, in just one year of the SARS-CoV-2 spread, more contagious genotypes of the virus, termed as “variants of concern” (such as B.1.1.7, B.1351, P.1), have appeared in the human population (Rambaut et al., 2020). There is evidence that some of these variants are able to avoid the immune response developed in patients infected with earlier variants of the virus (Wibmer et al., 2021). The same acceleration in the accumulation of nucleotide and amino acid substitutions in envelope proteins, compared to other genomic regions, is observed in other human viruses (Hepacivirus C, Human immunodeficiency virus 1, Human immunodeficiency virus 2) (Berry et al., 2007; Skar et al., 2010; Yuan et al., 2013).
The obtained time period of the divergence of SARS-CoV-2 strains collected in the first wave of COVID-19 with closely related viruses of the genus Betacoronavirus ranged from 31 to 72 years ago. This dating corresponds to the divergence time of different strains within the same virus species, such as already mentioned: measles morbillivirus, rubella virus, and poliovirus (Furuse et al., 2010; Zhu et al., 2012; Smura et al., 2014). The genetic distances between SARS-CoV-2, bat coronavirus RaTG13, bat coronavirus RmYN0, bat coronavirus RShSTT182, and bat coronavirus RShSTT200 are similar to the level of intraspecific genetic polymorphism for other species of viruses. The bat coronavirus RaTG13 strain was isolated from bats in 2013 in an abandoned mine in Mojiang Hani Autonomous County, Yunnan Province, China after three mineworkers developed a fatal viral pneumonia of unknown etiology in 2012 (Wu et al., 2014; Andersen et al., 2020). At that time, it was not possible to identify the pathogen from the diseased people. The latest study (Zech et al., 2021) shows that the bat coronavirus RaTG13 strain lyses both bats’ and humans’ cell cultures. In this case, the one amino acid substitution in the surface protein S of bat coronavirus RaTG13 participating in the interaction with the cell receptor increases the affinity of the virus to the human cell culture. It is likely, that SARS-CoV-2 diverged from its closest relatives of the genus Betacoronavirus recently, no later than 43 years ago. Single amino acid substitutions in the genome of an ancestral form of SARS-CoV-2 could increase the pathogenicity and infectivity for humans.
Our estimates for the first wave of the COVID-19 pandemic showed that there was a low level of genetic polymorphism in complete SARS-CoV-2 genomes. If the number of parsimony-informative sites is less than the number of sequences applied in a phylogenetic analysis, then we will always face low topology support and ambiguous clustering results. In our dataset, we face the problem of unreliable estimates of the topology of the reconstructed phylogenetic tree. This could not be improved by applying more data. When we increased the number of genomes in the analysis, this did not lead to an increase in the ratio of the number of parsimony-informative sites to the total number of analyzed genomes. The low genetic polymorphism of SARS-CoV-2 is associated with three factors: (1) the mutation rate in the genome is relatively low for viruses; (2) the time period of virus persistence in the human population is short on an evolutionary scale; (3) the effective population size of the virus growths exponentially. All these factors do not allow us to distinguish any individual genotypes of SARS-CoV-2, which circulated specifically in period of the first wave of the pandemic. Although such attempts have been made by researchers (Tang et al., 2020; Liu et al., 2020), stable virus lineages likely began to form later during the pandemic from March 2020, when widespread isolation between countries was launched. At this point, isolation barriers appeared, forming separate lineages of SARS-CoV-2 in each country, which is confirmed by the data of the online resource https://cov-lineages.org (Rambaut et al., 2020). Also, the consequence of low genetic polymorphism of the virus is the incapability of genome data to provide information on the virus pathways during the first wave of the pandemic which probably occurred due to the movement of patients with asymptomatic or mild forms of COVID-19 some-time before the introduction of mass accurate PCR-diagnostics for the disease. Although attempts to conduct genomic analysis to identify the first COVID-19 cases with the account epidemiological data in any country of the world are being made by researchers (Munnink et al., 2020), they may be inaccurate due to the unreliability of the primary information. In addition, the results of these studies may be questioned due to new data (Apolone et al., 2020) on the earlier spread of SARS-CoV-2 in Europe, long before the first genomic data were obtained.
Our results showed that an increase in the number of genomes in a dataset does not lead to an increase in the accuracy of the analysis when calculating the rates of evolution and dating the lifetime of a common ancestor for all SARS-CoV-2 variants of the pandemic first wave. Datasets of 252, 513 and 777 SARS-CoV-2 genomes give approximately the same estimates of these parameters (overlapping confidence intervals of the estimates). Confidence intervals do not narrow with increasing sample size. Three assumptions can explain this result: (1) insufficient polymorphism of the genomic data of SARS-CoV-2 of the pandemic first wave, leading to the fact that the number of parsimony-informative sites is always less than the sample size of genomes; (2) the possible presence of incorrectly identified nucleotides in the sequences, increasing the scatter of estimated values; (3) a possible incorrect indication of dates of the virus isolation in the genome submissions to the databases, which complicates accurate estimation of the evolutionary rates. Analysis of various datasets leads to different estimates of the time of the appearance of the first virus phylogenetic lineages in each of the studied countries. This is due to the low level of polymorphism in the genomes of SARS-CoV-2 of the pandemic first wave, and to the fact that the samples and datasets of sequenced genomes of complete databases used in the analysis make up an insignificant part of the total number of COVID-19 cases during this period of time. GISAID database accessed on 12 August 2020 accumulated 80189 genome records - 0.38% from official statistics in 20692140 confirmed cases (https://covid19.who.int) around the world on this date. At the same time, the official statistics are significantly less than the real number of cases due to asymptomatic and mild forms of the disease.
Calculations in the program "BEAST v. 2.6.2", when testing phylogenetic hypotheses and reconstructing the trees required extreme computational resources, in our case provided with access to the supercomputer" Akademik Matrosov " at the Irkutsk Supercomputer Center SB RAS. The analysis required about three months of testing and basic calculations on three computing nodes of the cluster, each of which was equipped with two 18-core 36-thread Intel Xeon E5-2695 v4 "Broadwell" processors. Parallel calculations using the BEAGLE v library.3.1.0 (Ayres et al., 2012) showed that the calculation speed could be increased by choosing the options with four threads and the division each of partitions (1 + 2 and 3rd codon position) by two. For the dataset of 252 genomes, 1,000,000 Markov chain generations required 5-7 minutes of estimation time, depending on the selected reconstruction model. A further increase in the number of threads and instances did not improve the speed, and after applying eight threads, the calculation speed, on the contrary, decreased. Calculations using the BEAGLE library on GPU (NVIDIA GTX 1080 ti) led to a slowdown in the calculation speed, to 25-30 minutes per 1,000,000 of the Markov chain generations. A run for 513 the SARS-CoV-2 genomes applying 1,000,000 Markov chain generations required at least 24 minutes of estimation time (optimal parameters were 4 threads and 2 instances). The converging value of the ESS statistic > 200 was achieved with 1.2 × 109 generations of Markov chains. Analysis of the dataset consisting of the 777 SARS-CoV-2 genomes for 1,000,000 Markov chain generations required at least 56 minutes of estimation time (with the optimal parameters of 4 threads and 2 instances). The converging value of the ESS statistic > 200 was achieved with 1.9 × 109 generations of Markov chains. Thus, studies using the Bayesian phylogenetic approach for several thousand genomes may require many years of calculations on high-performance computing systems. Developers are trying to improve computational methods (Miura et al., 2020) to help researchers involved in the full-genome phylogeny of SARS-CoV-2 and other viruses.
Our estimates of SARS-CoV-2 evolution rates and the time of occurrence of the first cases in the human population are consistent with the results of other studies analyzing the initial stages of the spread of the virus in different regions (Giovanetti et al., 2020; Farah et al., 2020) or in the world (Benvenuto et al., 2020; Koyama et al., 2020). The studies (Giovanetti et al., 2020; Farah et al., 2020; Benvenuto et al., 2020) were based on limited data sets, including several tens to hundreds of SARS-CoV-2 genomes available at that time. The analysis in the BEAST program described by Koyama et al. (2020) included 2,000 complete genomes, but a reconstruction model with strict clocks and without differentiation of codon positions was not optimal for data analysis as shown in our study. In addition, it is unlikely that with the large dataset of 2,000 genomes authors managed to achieve convergent results of ESS statistics of MCMC modelling in such a short time of calculations (article published online on May 13, 2020).
6. Conclusion
Our analysis of SARS-CoV-2 genomic data allows us to draw the following conclusions: (1) as an independent evolutionary line, SARS-CoV-2 appeared in nature at the end of the 20th Century; (2) that human infection with the SARS-CoV-2 variant of the COVID-19 pandemic first wave occurred in China in mid-2019; (3) that the spread of the virus from China to almost all countries across the world occurred in the period from the autumn of 2019, before the actual discovery of the SARS-CoV-2 virus, to mid-February 2020, and there was an exponential increase in the effective population size of the virus; 4) that the rate of evolution of the coding part of the SARS-CoV-2 genome is comparable to other human RNA-containing viruses (Measles morbillivirus, Rubella virus, Enterovirus C); 5) that in the first wave of the SARS-CoV-2 pandemic, genomic data had a low level of polymorphism, which does not allow us to track the exact pathways and spread of the virus in different regions or in the world as a whole without using additional accurate data from epidemiological observations.
CRediT authorship contribution statement
Yu.S. Bukin: Conceptualization, Visualization, Formal analysis, Writing – review & editing. A.N. Bondaryuk: Conceptualization, Visualization, Funding acquisition, Formal analysis, Writing – review & editing. N.V. Kulakova: Conceptualization, Visualization, Formal analysis, Writing – review & editing. S.V. Balakhonov: Conceptualization, Visualization. Y.P. Dzhioev: Conceptualization, Visualization, Writing – review & editing. V.I. Zlobin: Conceptualization, Visualization, Writing – review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
The work was supported by the Russian State Project No. 121032300196-8. The authors are grateful to the Irkutsk Supercomputer Center SB RAS and its administrator Ivan Sidorov for providing access to the high-performance cluster “Akademik Matrosov” for computing calculation facilities. We are very grateful to academic of the Russian Academy of Sciences - Mikhail Aleksandrovich Grachev for valuable recommendations when writing the manuscript. We thank Wai Ling Kok and Rachel Wilkins for proofreading.
Reference
- Andersen K.G., Rambaut A., Lipkin W.I., Holmes E.C., Garry R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020;26(4):450–452. doi: 10.1038/s41591-020-0820-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen K.G., Rambaut A., Lipkin W.I., Holmes E.C., Garry R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020;26(4):450–452. doi: 10.1038/s41591-020-0820-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apolone G., Montomoli E., Manenti A., Boeri M., Sabia F., Hyseni I., Pastorino U. Unexpected detection of SARS-CoV-2 antibodies in the prepandemic period in Italy. Tumori J. 2020;11 doi: 10.1177/0300891620974755. 0300891620974755. doi:10.1177%2F0300891620974755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ayres D.L., Darling A., Zwickl D.J., Beerli P., Holder M.T., Lewis P.O., Suchard M.A. BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Syst. Biol. 2012;61(1):170–173. doi: 10.1093/sysbio/syr100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baele G., Lemey P., Bedford T., Rambaut A., Suchard M.A., Alekseyenko A.V. Improving the accuracy of demographic and molecular clock model comparison while accommodating phylogenetic uncertainty. Mol. Biol. Evol. 2012;29(9):2157–2167. doi: 10.1093/molbev/mss084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basavaraju S.V., Patton M.E., Grimm K., Rasheed M.A.U., Lester S., Mills L., Stramer S.L. Serologic testing of US blood donations to identify severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)–reactive antibodies: December 2019–January 2020. Clin. Infect. Dis. 2020;72(12):e1004–e1009. doi: 10.1093/cid/ciaa1785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benson D.A., Cavanaugh M., Clark K., Karsch-Mizrachi I., Lipman D.J., Ostell J., Sayers E.W. GenBank. Nucleic Acids Res. 2012;41(D1):D36–D42. doi: 10.1093/nar/gks1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D. Benvenuto, M. Giovanetti, M. Salemi, M. Prosperi, C. De Flora, L.C. Junior Alcantara, & M. Ciccozzi, 2020. The global spread of 2019-nCoV: a molecular evolutionary analysis. Pathog. Glob. Health 114(2), 64-67. 10.1080%2F20477724.2020.1725339. [DOI] [PMC free article] [PubMed]
- Berry I.M., Ribeiro R., Kothari M., Athreya G., Daniels M., Lee H.Y., Leitner T. Unequal evolutionary rates in the human immunodeficiency virus type 1 (HIV-1) pandemic: the evolutionary rate of HIV-1 slows down when the epidemic rate increases. J. Virol. 2007;81(19):10625–10635. doi: 10.1128/JVI.00985-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouckaert R., Heled J., Kühnert D., Vaughan T., Wu C.H., Xie D., Drummond A.J. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 2014;10(4) doi: 10.1371/journal.pcbi.1003537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradshaw D., Vasylyeva T.I., Davis C., Pybus O.G., Thézé J., Thomson E.C., Nelson M. Transmission of hepatitis C virus in HIV-positive and PrEP-using MSM in England. J. Viral Hepat. 2020;27(7):721–730. doi: 10.1111/jvh.13286. [DOI] [PubMed] [Google Scholar]
- Cacciapaglia G., Cot C., Sannino F. Second wave COVID-19 pandemics in Europe: a temporal playbook. Sci. Rep. 2020;10(1):1–8. doi: 10.1038/s41598-020-72611-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrat F., Figoni J., Henny J., Desenclos J.C., Kab S., de Lamballerie X., Zins M. Evidence of early circulation of SARS-CoV-2 in France: findings from the population-based “CONSTANCES” cohort. Eur. J. Epidemiol. 2021;36(2):219–222. doi: 10.1007/s10654-020-00716-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chatterjee P., Nagi N., Agarwal A., Das B., Banerjee S., Sarkar S., Gangakhedkar R.R. The 2019 novel coronavirus disease (COVID-19) pandemic: A review of the current evidence. Indian J. Med. Res. 2020;151(2-3):147. doi: 10.4103/ijmr.IJMR_519_20. 10.4103%2Fijmr.IJMR_519_20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duchene S., Lemey P., Stadler T., Ho S.Y.W., Duchene D.A., Dhanasekaran V., Baele G. Bayesian evaluation of temporal signal in measurably evolving populations. Mol. Biol. Evol. 2020;37:3363–3379. doi: 10.1093/molbev/msaa163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farah S., Atkulwar A., Praharaj M.R., Khan R., Gandham R., Baig M. Phylogenomics and phylodynamics of SARS-CoV-2 genomes retrieved from India. Future Virol. 2020;15(11):747–753. doi: 10.2217/fvl-2020-0243. [DOI] [Google Scholar]
- Furuse Y., Suzuki A., Oshitani H. Origin of measles virus: divergence from rinderpest virus between the 11 th and 12 th centuries. Virol. J. 2010;7(1):1–4. doi: 10.1186/1743-422X-7-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao Z., Xu Y., Sun C., Wang X., Guo Y., Qiu S., Ma K. A systematic review of asymptomatic infections with COVID-19. J. Microbiol. Immunol. Infect. 2020;54:12–16. doi: 10.1016/j.jmii.2020.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garvin M.R., Prates E.T., Pavicic M., Jones P., Amos B.K., Geiger A., Jacobson D. Potentially adaptive SARS-CoV-2 mutations discovered with novel spatiotemporal and explainable AI models. Genome Biol. 2020;21(1):1–26. doi: 10.1186/s13059-020-02191-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giovanetti M., Benvenuto D., Angeletti S., Ciccozzi M. The first two cases of 2019-nCoV in Italy: where they come from? J. Med. Virol. 2020;92(5):518–521. doi: 10.1002/jmv.25699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 2010;59(3):307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- R. He, F. Dobie, M. Ballantine, A. Leeson, Y. Li, N. Bastien, X. Li, 2004. Analysis of multimerization of the SARS coronavirus nucleocapsid protein. Biochem. Biophys. Res. Commun. 316(2), 476-483. 10.1016/j.bbrc.2004.02.074. [DOI] [PMC free article] [PubMed]
- Hul V., Delaune D., Karlsson E.A., Hassanin A., Tey P.O., Baidaliuk A., Duong V. A novel SARS-CoV-2 related coronavirus in bats from Cambodia. BioRxiv. 2021 doi: 10.1101/2021.01.26.428212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalyaanamoorthy S., Minh B.Q., Wong T.K.F., von Haeseler A., Jermiin L.S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods. 2017;14:587–589. doi: 10.1038/nmeth.4285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koyama T., Platt D., Parida L. Variant analysis of SARS-CoV-2 genomes. Bull. World Health Organ. 2020;98(7):495–504. doi: 10.2471/BLT.20.253591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larget B., Simon D. L. Markov chain Monte Carlo algorithms for the Bayesian analysis of phylogenetic trees. Mol. Biol. Evol. 1999;16(6):750–759. doi: 10.1093/oxfordjournals.molbev.a026160. [DOI] [Google Scholar]
- Li J., Wang H., Mao L., Yu H., Yu X., Sun Z., Wang X. Rapid genomic characterization of SARS-CoV-2 viruses from clinical specimens using nanopore sequencing. Sci. Rep. 2020;10(1):1–10. doi: 10.1038/s41598-020-74656-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q., Zhao S., Shi C.M., Song S., Zhu S., Su Y., Chen H. Genom. Proteom. Bioinform. 2020. Population genetics of SARS-CoV-2: disentangling effects of sampling bias and infection clusters. In Press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minh B.Q., Nguyen M.A.T., von Haeseler A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 2013;30(5):1188–1195. doi: 10.1093/molbev/mst024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miura S., Tamura K., Tao Q., Huuki L.A., Pond S.L.K., Priest J., Kumar S. A new method for inferring timetrees from temporally sampled molecular sequences. PLoS Comput. Biol. 2020;16(1) doi: 10.1371/journal.pcbi.1007046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munnink B.B.O., Nieuwenhuijse D.F., Stein M., O'Toole Á., Haverkate M., Mollers M., Koopmans M. Rapid SARS-CoV-2 whole-genome sequencing and analysis for informed public health decision-making in the Netherlands. Nat. Med. 2020;26(9):1141–1405. doi: 10.1038/s41591-020-0997-y. [DOI] [PubMed] [Google Scholar]
- Nguyen L.T., Schmidt H.A., Von Haeseler A., Minh B.Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015;32(1):268–274. doi: 10.1093/molbev/msu300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- J.H.O. Pettersson, J. Bohlin, M. Dupont-Rouzeyrol, O.B. Brynildsrud, K. Alfsnes, V.M. Cao-Lormeau, & E.A. Gould, 2018. Re-visiting the evolution, dispersal and epidemiology of Zika virus in Asia. Emerg. Microbes Infect. 7(1), 1-8. 10.1038/s41426-018-0082-5. [DOI] [PMC free article] [PubMed]
- Paradis E., Claude J., Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20(2):289–290. doi: 10.1093/bioinformatics/btg412. [DOI] [PubMed] [Google Scholar]
- Pybus O.G., Rambaut A. Evolutionary analysis of the dynamics of viral infectious disease. Nat. Rev. Genet. 2009;10:540–550. doi: 10.1038/nrg2583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rambaut A., Holmes E.C., O'Toole Á., Hill V., McCrone J.T., Ruis C., Pybus O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020;5(11):1403–1407. doi: 10.1038/s41564-020-0770-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rambaut A., Lam T.T., Max Carvalho L., Pybus O.G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen) Virus Evol. 2016;2(1) doi: 10.1093/ve/vew007. vew007. 10.1093/ve/vew007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapiro B., Rambaut A., Drummond A.J. Choosing appropriate substitution models for the phylogenetic analysis of protein-coding sequences. Mol. Biol. Evol. 2006;23(1):7–9. doi: 10.1093/molbev/msj021. [DOI] [PubMed] [Google Scholar]
- Shu Y., McCauley J. GISAID: Global initiative on sharing all influenza data–from vision to reality. Euro Surveill. 2017;22(13):30494. doi: 10.1038/s41598-020-74656-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skar H., Borrego P., Wallstrom T.C., Mild M., Marcelino J.M., Barroso H., Albert J. HIV-2 genetic evolution in patients with advanced disease is faster than that in matched HIV-1 patients. J. Virol. 2010;84(14):7412–7415. doi: 10.1128/JVI.02548-09. 0.1128/JVI.02548-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smura T., Blomqvist S., Vuorinen T., Ivanova O., Samoilovich E., Al-Hello H., Roivainen M. Recombination in the evolution of enterovirus C species sub-group that contains types CVA-21, CVA-24, EV-C95, EV-C96 and EV-C99. PLoS One. 2014;9(4):e94579. doi: 10.1371/journal.pone.0094579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabari P., Amini M., Moghadami M., Moosavi M. International public health responses to COVID-19 outbreak: a rapid review. Iran. J. Med. Sci. 2020;45(3):157. doi: 10.30476/ijms.2020.85810.1537. 10.30476%2Fijms.2020.85810.1537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang X., Wu C., Li X., Song Y., Yao X., Wu X., Lu J. On the origin and continuing evolution of SARS-CoV-2. Natl. Sci. Rev. 2020;7(6):1012–1023. doi: 10.1093/nsr/nwaa036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Boheemen S., de Graaf M., Lauber C., Bestebroer T.M., Raj V.S., Zaki A.M., Fouchier R.A. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. MBio. 2012;3(6):1–9. doi: 10.1128/mBio.00473-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- T.P. Velavan, C.G. Meyer, 2020. The COVID-19 epidemic. Trop. Med. Int. Health 25(3), 278. 10.1111/tmi.13383. [DOI] [PMC free article] [PubMed]
- Wang R., Chen J., Gao K., Hozumi Y., Yin C., Wei G.W. Analysis of SARS-CoV-2 mutations in the United States suggests presence of four substrains and novel variants. Commun. Biol. 2021;4(1):1–14. doi: 10.1038/s42003-021-01754-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wibmer C.K., Ayres F., Hermanus T., Madzivhandila M., Kgagudi P., Oosthuysen B., Moore P.L. SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. Nat. Med. 2021:1–4. doi: 10.1038/s41591-021-01285-x. [DOI] [PubMed] [Google Scholar]
- Worobey M., Watts T.D., McKay R.A., Suchard M.A., Granade T., Teuwen D.E., Jaffe H.W. 1970s and ‘Patient 0’HIV-1 genomes illuminate early HIV/AIDS history in North America. Nature. 2016;539(7627):98–101. doi: 10.1038/nature19827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu F., Zhao S., Yu B., Chen Y.M., Wang W., Song Z.G., Zhang Y.Z. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579(7798):265–269. doi: 10.1038/s41591-020-0820-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Z., Yang L., Yang F., Ren X., Jiang J., Dong J., Jin Q. Novel henipa-like virus, mojiang paramyxovirus, in rats, China. Emerg. Infect. Dis. 2014;20(6):10642012. doi: 10.3201/eid2006.131022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan M., Lu T., Li C., Lu L. The evolutionary rates of HCV estimated with subtype 1a and 1b sequences over the ORF length and in different genomic regions. PLoS One. 2013;8(6):e64698. doi: 10.1371/journal.pone.0064698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zech F., Schniertshauer D., Jung C., Herrmann A., Xie Q., Nchioua R., Kirchhoff F. Spike mutation T403R allows bat coronavirus RaTG13 to use human ACE2. bioRxiv. 2021 doi: 10.1101/2021.05.31.446386. [DOI] [Google Scholar]
- Zhao H., Lu X., Deng Y., Tang Y., Lu J. COVID-19: asymptomatic carrier transmission is an underestimated problem. Epidemiol. Infect. 2020;148:e116. doi: 10.1017/S0950268820001235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu Z., Cui A., Wang H., Zhang Y., Liu C., Wang C., Xu W. Emergence and continuous evolution of genotype 1E rubella viruses in China. J. Clin. Microbiol. 2012;50(2):353–363. doi: 10.1128/JCM.01264-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou H., Chen X., Hu T., Li J., Song H., Liu Y., Shi W. A novel bat coronavirus closely related to SARS-CoV-2 contains natural insertions at the S1/S2 cleavage site of the spike protein. Curr. Biol. 2020;30:2196–2203. doi: 10.1016/j.cub.2020.05.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All supplementary materials such as the genomes with GISAID and GenBank numbers used in the study, the xml files for the BEAST v. 2.6.2 program, and original reconstructed phylogenetic trees are available at https://doi.org/10.6084/m9.figshare.14830899.