Summary
Infectious diseases continually pose global medical challenges. The transcription factor GATA2 establishes gene networks and defines cellular identity in hematopoietic stem/progenitor cells and in progeny committed to specific lineages. GATA2-haploinsufficient patients exhibit a spectrum of immunodeficiencies associated with bacterial, viral, and fungal infections. Despite accumulating clinical knowledge of the consequences of GATA2 haploinsufficiency in humans, it is unclear how GATA2 haploinsufficiency compromises host anti-infectious defenses. To address this issue, we examined Gata2-heterozygous mutant (G2Het) mice as a model for human GATA2 haploinsufficiency. In vivo inflammation imaging and cytokine multiplex analysis demonstrated that G2Het mice had attenuated inflammatory responses with reduced levels of inflammatory cytokines, particularly IFN-γ, IL-12p40, and IL-17A, during lipopolysaccharide-induced acute inflammation. Consequently, bacterial clearance was significantly impaired in G2Het mice after cecal ligation and puncture-induced polymicrobial peritonitis. These results provide direct molecular insights into GATA2-directed host defenses and the pathogenic mechanisms underlying observed immunodeficiencies in GATA2-haploinsufficient patients.
Subject area: Molecular biology, Immunology
Graphical abstract

Highlights
-
•
Gata2Het mice exhibited reduced inflammation than WT mice in response to bacteria
-
•
After polymicrobial peritonitis, bacterial clearance was impaired in Gata2Het mice
-
•
During inflammation, IFN-γ, IL-12p40, and IL-17A were lower in Gata2Het mice
-
•
Inflammation imaging was performed during peritonitis and sepsis in hIL-6 Luc mice
Molecular biology; Immunology
Introduction
Although first-world countries have developed elaborate health care systems, we nonetheless continue to be confronted by long-standing or emerging infectious disease threats (Becker et al., 2006; Khabbaz et al., 2014). Many mechanistic aspects of the host immune defense system and its failures in the face of these recurrent challenges have long been topics of debate (Ibrahim et al., 2017). Clinical and genetic studies conducted over the last decade have revealed the inextricable link between the hematopoietic transcription factor GATA2 and the etiology of primary immunodeficiency disorder (Jung et al., 2020; Shimizu and Yamamoto, 2020).
GATA2 is a founding member of the GATA transcription factor family, which most avidly binds to a consensus (A/T) GATA (A/G) motif through two evolutionarily conserved C4 zinc fingers (Moriguchi and Yamamoto, 2014; Shimizu and Yamamoto, 2016; Tsai et al., 1989; Yamamoto et al., 1990). GATA2 has been established as a master regulator of hematopoiesis, since Gata2-mutant mice die around 9.5-day mouse embryos owing to complete hematopoietic failure (Tsai et al., 1994). Numerous subsequent studies have demonstrated that GATA2 is highly expressed in hematopoietic stem/progenitor cells (HSPCs) and is essential for normal hematopoietic development through its regulation of multiple target genes (Fujiwara et al., 2009; Lim et al., 2012; Suzuki et al., 2006, 2013; Tsai and Orkin, 1997).
Although abundant GATA2 expression has been detected in severe cases of acute myeloid leukemia, clinical studies of germline GATA2 mutations have established its causal role in an immunodeficiency syndrome, monocytopenia and mycobacterial infection (MonoMAC) syndrome, and dendritic cell, monocyte, and B and NK lymphocyte (DCML) deficiency, in which defects in monocytes, B cells, natural killer cells, and dendritic cells lead to mycobacterial, fungal, and viral infections (Bigley et al., 2011; Dickinson et al., 2011; Hsu et al., 2011; Luesink et al., 2012; Vinh et al., 2010). Patients with GATA2 mutations are also predisposed to myelodysplastic syndrome and acute myeloid leukemia (Hahn et al., 2011; Ostergaard et al., 2011). The germline GATA2 mutations detected in patients were monoallelic and were identified as missense, frameshift, or nonsense mutations mainly in the DNA-binding zinc-finger domains (Bresnick et al., 2020; Collin et al., 2015). Thus, haploinsufficiency of GATA2 underlies the pathogenesis of these conditions. Monoallelic complete gene deletion, mutations in the fifth intron cis-regulatory elements, and variations within the 5′ noncoding exons of GATA2 have also been found in diseased patients (Bresnick et al., 2020; Collin et al., 2015; Hsu et al., 2013; Johnson et al., 2012; Shimizu and Yamamoto, 2020). This spectrum of qualitative and quantitative mutations of GATA2 is believed to compromise the regulatory network of the hematopoietic system and the host defense system. Consequently, GATA2-haploinsufficient patients showed an increased prevalence of a variety of infections, i.e., severe viral infections in 70%, mycobacterial infections in 53%, fungal infections in 16%, and other severe bacterial infections in 49% of the 57 reported patients (Spinner et al., 2014). These data suggest that GATA2 plays a crucial role in pathogen clearance by reinforcing the host defense system. However, the molecular mechanisms underlying GATA2-directed host defense remain obscure.
Although GATA2 governs early hematopoietic development, mature hematopoietic and numerous other cell lineages in adult tissues sustain GATA2 expression as well (Harigae et al., 1998; Li et al., 2015; Ohmori et al., 2012; Tsai and Orkin, 1997; Watanabe-Asaka et al., 2020; Wilson et al., 1990; Yu et al., 2014; Zon et al., 1993). For example, our group and others have demonstrated that GATA2 plays a key role in the maintenance and function of mast cells and participates in allergic inflammatory responses (Li et al., 2017, 2021; Ohmori et al., 2015, 2019; Ohneda et al., 2019). We also showed that GATA2 exacerbated inflammatory kidney injury by promoting cytokine production from renal collecting duct cells (Yu et al., 2017). Host-pathogen interactions promote the production of inflammatory cytokines such as interleukins, chemokines, and interferons by a variety of cells, which subsequently facilitate leukocyte recruitment and clearance of pathogens (Kumar et al., 2011). Considering these features of inflammatory microenvironments, we hypothesized that GATA2 might promote inflammation and simultaneously accelerate pathogen clearance by regulating cytokine balance in adult animals.
To address the potential roles played by GATA2 in the inflammatory responses and pathogen clearance, we employed Gata2-heterozygous mutant mice (referred to as G2Het hereinafter) as a model for human GATA2 haploinsufficiency (Tsai et al., 1994). We found that G2Het mice had a less inflammatory status and lower cytokine production than wild-type (WT) mice upon lipopolysaccharide (LPS) administration. Of note, bacterial clearance was impaired in G2Het mice in comparison with WT mice upon polymicrobial peritonitis induced by cecal ligation and puncture (CLP). This study demonstrates that GATA2 is involved in LPS-induced inflammatory responses and bacterial clearance after CLP-induced polymicrobial peritonitis.
Results
Attenuation of LPS-induced inflammation in G2Het mice
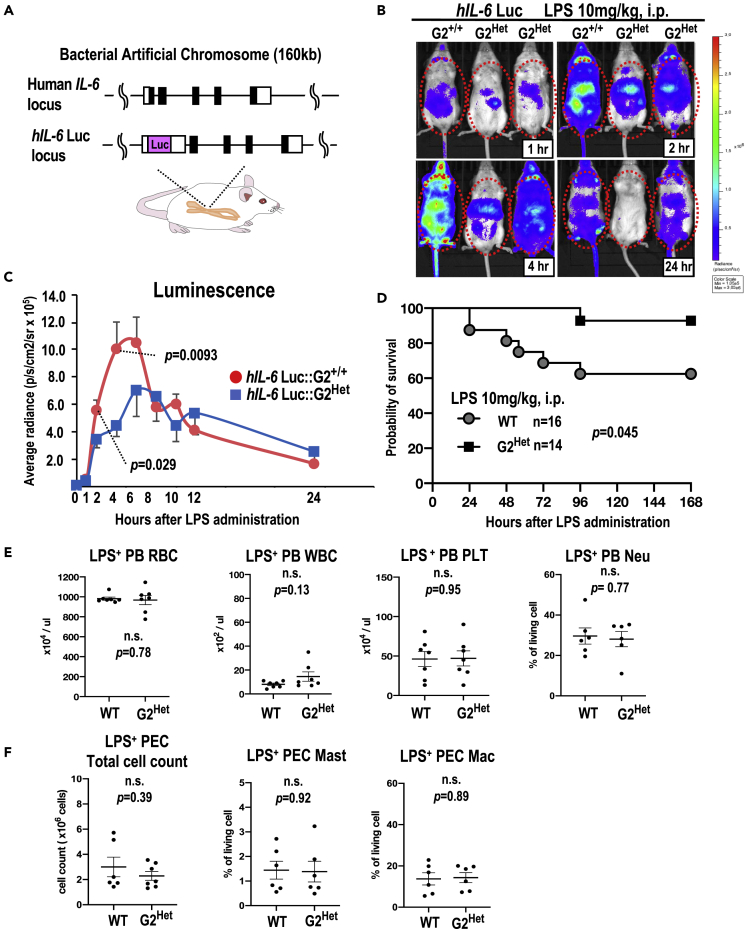
To test the hypothesis that GATA2 haploinsufficiency is responsible for impaired immune function, we examined whether Gata2 heterozygous mutant (G2Het) mice showed any alterations in inflammatory responses upon LPS administration. Previously, we developed an in vivo inflammation imaging strategy exploiting a transgenic mouse line carrying a human IL-6 bacterial artificial chromosome-driven luciferase reporter gene (hIL-6 Luc; Figure 1A) (Hayashi et al., 2015). By crossing these hIL-6 Luc transgenic mice with the G2Het mice, we were able to monitor the inflammatory status of G2Het mice up to 24 h after intraperitoneal (i.p.) LPS injection. We found that G2Het mice showed a diminished inflammatory response in comparison with Gata2 wild-type genetic background of mice harboring the hIL-6 Luc transgene (hIL-6 Luc::G2+/+ or G2+/+ background), particularly in the peritoneum during the early phases of the time course (see the 1-, 2-, and 4-h time points in Figure 1B). Quantification of these results clearly demonstrated that hIL-6 Luc luminescence was significantly diminished in the G2Het mice by 2–4 h after LPS administration: a 40% reduction after 2 h (p = 0.029) was followed by a 56% reduction at 4 h (p = 0.0093) relative to G2+/+ background (Figure 1C). Subcutaneous or low-dose i.p. administration of LPS exerted similar effects (Figures S1A and S1B). We also found that G2Het mice showed moderate and statistically significant increase of survival compared with wild-type (WT) mice after administration of LPS (p = 0.045), consistent with a lower inflammatory status in the G2Het mice (Figure 1D). A previous study reported that G2Het mice showed comparable populations of neutrophils, B lymphocytes, T lymphocytes, natural killer cells, macrophages, monocytes, and dendritic cells to WT mice in the spleen under steady-state conditions (Onodera et al., 2016). In this study, we further demonstrated that G2Het mice and WT mice harbored comparable populations of red blood cells, white blood cells, platelets, peripheral blood neutrophils, peritoneal mast cells, peritoneal macrophages, and other hematopoietic lineages, regardless of LPS administration (Figures 1E, 1F, and S2A–S2C). We also found that G2Het mice showed a reduced expression level of Gata2 mRNA in bone marrow progenitor fractions, as anticipated (Figures S2D–S2F). These results demonstrate that GATA2-haploinsufficiency mice have a lower inflammatory response than WT mice upon LPS-induced systemic inflammation and that this attenuation was not attributable to any gross hemocytopenia in the G2Het mice.
Figure 1.
Attenuation of LPS-induced inflammation in Gata2 heterozygous mice
(A) Structure of the hIL-6 luciferase transgenic allele (hIL-6 Luc) used for in vivo inflammation monitoring.
(B and C) Time course monitoring of the inflammation induced by intraperitoneal (i.p.) LPS administration (n = 4–8 mice at each time point). Note that Gata2 heterozygous mutant (G2Het) mice showed a lower level of inflammation at 2–4 h after LPS administration than G2+/+.
(D) Kaplan-Meier survival curves of G2Het (n = 14) and WT (n = 16) mice after 10 mg/kg i.p. LPS administration. G2Het mice showed a higher survival rate.
(E) Hematological indices in the peripheral blood (PB): red blood cells (RBC), white blood cells (WBC), platelets (PLT) and Ly6G+CD11b+ neutrophils (Neu) 4 h after 10 mg/kg i.p. LPS administration (n = 6–7 for WT and n = 6–7 for G2Het).
(F) Total numbers of peritoneal cells, FcεR+c-kit+ peritoneal mast cells (PEC Mast), and CD11b+F4/80+ peritoneal macrophages (PEC Mac) in the peritoneal cavity 4 h after 10 mg/kg i.p. LPS administration (n = 6 for WT and n = 6–7 for G2Het).
The results were calculated from more than three experiments. Data in Figures 1C, 1E, and 1F are represented as the mean ± SEM. Dots in Figures 1E and 1F represent independent biological replicates. Dots in Figure 1D represent death events. p values were calculated with the unpaired t test for Figures 1C, 1E, and 1F and the log rank test for Figure 1D. n.s., not significant. See also Figures S1 and S2 for further information.
Reduced cytokine levels in G2Het mice
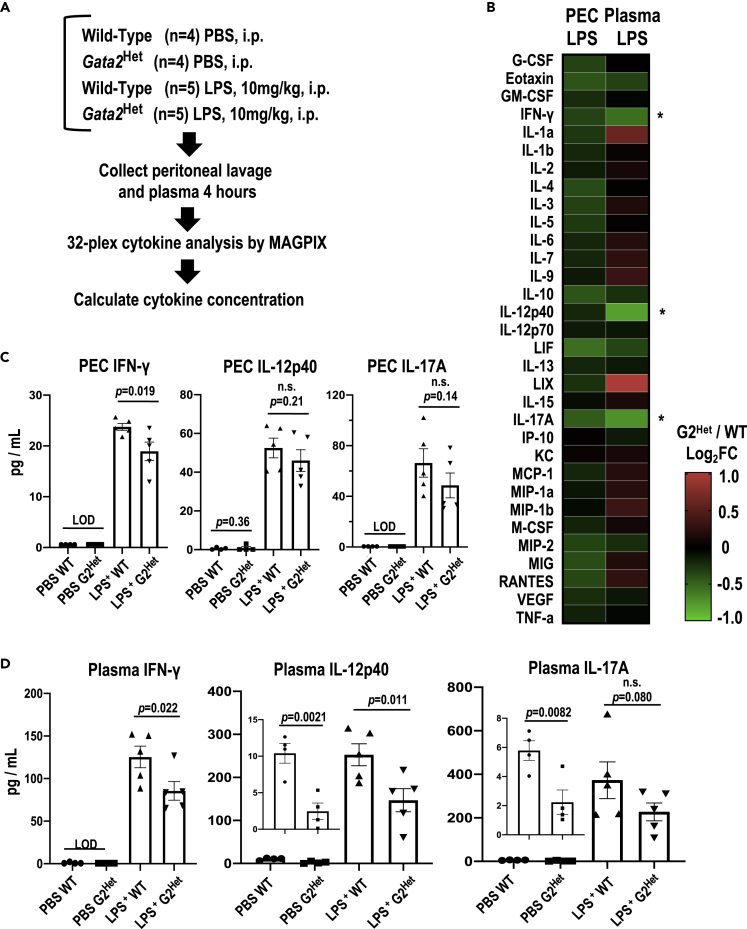
Given that G2Het mice exhibited a diminished inflammatory response while harboring normal hematopoietic populations, we hypothesized that cytokine production might be attenuated in the G2Het mice. To address this question, we subjected peritoneal lavage fluids and plasma recovered from LPS-treated G2Het and WT mice to multiple 32-plex cytokine immune analyses by MAGPIX (Figure 2A). We found that almost all of the cytokine levels examined (30/32) in G2Het peritoneal lavage fluid were lower than those in WT mice (left column in Figure 2B). We further noticed that IFN-γ, IL-12p40, and IL-17A were most reduced in the plasma of the G2Het mice, although almost one-third of the cytokines were higher in the G2Het than in the WT mice (∗ in right column of Figure 2B). The quantified results showed that IFN-γ, IL-12p40, and IL-17A levels in the plasma of LPS-treated G2Het mice were reduced to 58%–68% of the levels found in WT controls (Figure 2D). The same trends were also detected in the peritoneal lavage fluid of the G2Het mice (Figure 2C). Basal levels of IL-12p40 and IL-17A in the plasma of the vehicle (PBS)-treated G2Het mice were also decreased; IL-12p40 and IL-17A in G2Het mice were reduced by 76% and 61%, respectively, relative to the WT mice (insets in Figure 2D).
Figure 2.
Reduced cytokine levels in G2Het mice
(A) Experimental workflow of multiplex cytokine immune analysis by MAGPIX (n = 4 for the PBS-treated group and n = 5 for the LPS-treated group in WT and G2Het).
(B) Heatmaps visualizing the fold changes in mean cytokine concentration in the G2Het mice over the WT mice in peritoneal cavity lavage fluid (PEC) and plasma upon LPS stimulation. Note that almost all cytokines (30/32) in PEC were decreased in the G2Het mice compared with the WT mice (left column). IFN-γ, IL-12p40, and IL-17A were most diminished in the plasma of G2Het mice (∗ in right column).
(C and D) Quantification results show that IFN-γ, IL-12p40, and IL-17A levels upon LPS stimulation were diminished in peritoneal lavage (C) and plasma (D). Basal levels of IL-12p40 and IL-17A in the plasma of vehicle (PBS)-treated G2Het mice were also decreased (insets in D).
Results were calculated from two experiments. Data in Figures 2C and 2D are represented as the mean ± SEM. Dots in Figures 2C and 2D represent independent biological replicates. p Values were calculated with unpaired t tests for Figures 2C and 2D. LOD, limit of detection; n.s., not significant. See also Figure S3 for further information.
To evaluate the kinetics of cytokine production, we further examined representative cytokines, including IFN-γ, IL-12p40, IL-17A, IL-6, IL-1α, and MIP-1β, by MAGPIX over a period of time (0, 4, 8, and 24 h) after LPS administration. We found that plasma levels of IFN-γ, IL-12p40, and IL-17A were lower in the G2Het mice than in the WT mice during LPS-induced inflammation (Figure S3). In contrast, IL-1α and/or MIP-1β were transiently elevated in G2Het mice at the 8-h stage in peritoneal lavage fluid and at the 4- to 8-hr stage in plasma. Of note, most cytokines tended to be reduced in the G2Het mice at the 0- and 24-h stages (Figure S3).
Collectively, these results suggest that a broad spectrum of inflammatory cytokines, particularly IFN-γ, IL-12p40, and IL-17A, appear to be under intense direct or indirect regulatory influence by GATA2 upon LPS-induced inflammation, although there were differences in the induction kinetics of these cytokines.
Imaging of inflammation in vivo after induction of polymicrobial peritonitis by cecal ligation and puncture
Proinflammatory cytokines, including IFN-γ, IL-12, and IL-17A, play a role in bacterial clearance by recruiting and activating leukocytes (Cavalla et al., 2014; Muñoz-Carrillo et al., 2018). We thus hypothesized that the reduced cytokine production impaired bacterial clearance in the G2Het mice. To address this possibility, we conducted CLP as a model of polymicrobial peritonitis (Figure S5) (Rittirsch et al., 2009). When we monitored the inflammatory status of polymicrobial peritonitis by exploiting hIL-6 Luc transgenic mice on a G2+/+ background, luciferase luminescence was first detected around the peritoneum as early as 4 h after the CLP procedure (Figure 3A). Thereafter, the luciferase luminescence expanded throughout the body by 12 h (Figure 3A). Quantitative analysis demonstrated that the luciferase luminescence reached a peak at 24 h and thereafter gradually decreased; by 72 h, the inflammatory status was approximately as it had been in the early stage (Figure 3B). It has been reported that bacterial proliferation peaked 24 h after CLP surgery, which is in agreement with our current luminescence time course data demonstrated in hIL-6 Luc transgenic mice (Meng et al., 2012). We next asked whether the hIL-6-controlled luminescence level correlated with the number of bacterial colony-forming units (CFUs), which were counted by inoculating peritoneal lavage fluid onto agar plates. We found a positive correlation (R2 = 0.705) between the luminescence intensity and CFU counts in the peritoneal lavage fluid 24 h after the CLP procedure, indicating that the luminescence level positively correlated with the robustness of bacterial colonization in the G2+/+ mice (Figures 3C and 3D). Collectively, the hIL-6 Luc transgenic mouse system enabled us to precisely monitor the inflammation status along with bacterial infiltration in mice in response to CLP-induced peritonitis.
Figure 3.
Inflammation imaging after cecal ligation and puncture (CLP)
(A and B) Time course inflammation imaging of the polymicrobial peritonitis induced by CLP, exploiting hIL-6 Luc transgenic mice on a G2+/+ background (n = 6–10 mice at each time point). Representative images are shown in Figure 3A.
(C) Inflammation imaging of CLP-induced polymicrobial peritonitis 24 h after CLP. The number of bacterial colony-forming units (CFU) in the peritoneum of each mouse is indicated. Note that the mouse with robust luminescence shows a high number of CFUs.
(D) A correlation between luminescence intensity and bacterial CFU counts upon CLP-induced polymicrobial peritonitis in hIL-6 Luc transgenic mice on a G2+/+ background (n = 18). The coefficient of determination, at 0.705, shows a positive correlation between luminescence and CFU counts.
The results in Figures 3B and 3D were calculated from more than three experiments. Data in Figure 3B are represented as the mean ± SEM. Dots in Figure 3D represent independent biological replicates. See also Figure S5A for details of the CLP procedure.
Impaired bacterial clearance in G2Het mice upon CLP-induced polymicrobial peritonitis
In order to further examine the role of GATA2 in bacterial clearance, G2Het and G2+/+ mice crossed with hIL-6 Luc transgenic mice were subjected to CLP. We found that the bacterial CFU count in the peritoneal lavage fluid 24 h after CLP was significantly higher in the G2Het group than in the G2+/+ group (G2Het: 1.2 × 106 CFU; G2+/+: 1.7 × 104 CFU, p = 0.0014: Figures 4A and 4C). CFU counts in the peripheral blood were also moderate and statistically significant higher in the G2Het group than in the G2+/+ group, suggesting the prevalence of bacteremia in the G2Het mice (G2Het: 9.4 × 102 CFU versus G2+/+: 4.1 × 102 CFU, p = 0.047: Figures 4A and 4C). We next monitored the inflammatory status in the G2Het and G2+/+ mice by employing the hIL-6 Luc imaging system described above. We found that hIL-6-directed luciferase luminescence showed a moderate and statistically significant decrease (p = 0.040) at 4 h after CLP and a slight increase at 24 h after CLP in the G2Het group (Figures 4D and 4E). With the hIL-6 Luc imaging system, the luminescence level positively correlated with bacterial counts in mice with the G2+/+ genetic background (Figure 3D). Therefore, we normalized the luminescence level to the number of bacterial CFUs to estimate the inflammatory response per unit of bacterial count. Of note, we found that G2Het showed a significantly lower level of luminescence per CFU than G2+/+, indicating a diminished inflammatory response to bacteria in G2Het mice (G2Het: 0.27 versus G2+/+: 12.5, p = 0.0024; Figure 4F). These results indicate that the inflammatory immune responses to bacterial infection failed to fully activate in the G2Het mice, which subsequently impaired bacterial clearance in the CLP model.
Figure 4.
Impaired bacterial clearance in G2Het mice upon CLP
(A) Bacterial CFU in the peritoneal cavity at 24 h after CLP (n = 7 for sham G2+/+, n = 6 for sham G2Het, n = 10 for CLP G2+/+, and n = 11 for CLP G2Het).
(B) Bacterial colony-forming units (CFU) in the peripheral blood at 24 h after CLP (n = 7 for sham G2+/+, n = 6 for sham G2Het, n = 10 for CLP G2+/+, and n = 11 for CLP G2Het).
(C) Representative agar plates with cultures from peritoneal cavity lavage fluid (PEC) and peripheral blood (PB) from G2+/+ and G2Het at 24 h after CLP.
(D) Whole-body inflammation monitoring using hIL-6 Luc at 4 or 24 h after CLP or the sham controls in the G2+/+ and G2Het groups.
(E) Quantification of the luminescence at 4 or 24 h after the CLP in the G2+/+ and G2Het groups (n = 10–12 mice at each time point).
(F) Luminescence per CFU was calculated to evaluate the response to inflammation per bacterial count (n = 10 for G2+/+ and n = 11 for G2Het). Note that G2Het showed significantly diminished luminescence per CFU.
The results in Figures 4A, 4B, 4E, and 4F were calculated from more than three experiments. The results in Figures 4C and 4D are representative images from Figures 4A, 4B, and 4E, respectively. Data in Figures 4A, 4B, 4E, and 4F are represented as the median and interquartile range. Dots in Figures 4A, 4B, 4E, and 4F represent independent biological replicates. p Values were calculated with the Wilcoxon-Mann-Whitney test for Figures 4A, 4B, 4E, and 4F. n.s., not significant.
Roles of macrophages and mast cells in GATA2-directed bacterial clearance
GATA2 was reported to be expressed in differentiated mast cells and macrophages, both of which are involved in the host immune system (Cassado et al., 2015; Johnzon et al., 2016; Ohmori et al., 2015; Wu et al., 2013). We found that bright GFP signal was detected in the c-kit+FcεR+ peritoneal mast cells (MCs), whereas there was virtually no GFP fluorescence in F4/80+CD11b+ peritoneal macrophages (Macs) in the Gata2 GFP knock-in mice, in which a GFP expression cassette was inserted into the translational start site (Figures 5A and 5B) (Suzuki et al., 2006). To further assess the endogenous Gata2 mRNA profile in the isolated c-kit+ bone marrow progenitors, peritoneal macrophages, and peritoneal mast cells of WT mice, we performed reverse transcription quantitative PCR (RT-qPCR) using a Gata2-specific primer pair. Consistent with the GFP expression profile, the Gata2 mRNA level in peritoneal macrophages was more than 100-fold lower than that in c-kit+ bone marrow progenitors or peritoneal mast cells (Figure 5C).
Figure 5.
Roles of macrophages and mast cells in GATA2-directed bacterial clearance
(A) Schematic diagrams of Gata2 GFP knock-in (Gata2GFP), Gata2 floxed (G2F) alleles, and ROSA26-tdTomato allele (R26-tdT).
(B) Histograms depicting GFP fluorescence in peritoneal macrophages (Mac) and peritoneal mast cells (MC) of Gata2GFP/+ mice and WT mice.
(C) The relative Gata2 mRNA levels were quantified by RT-qPCR using a Gata2 exon 3–4 primer set and normalized to glyceraldehyde-3-phosphate dehydrogenase (Gapdh) in the WT mice. Gata2 mRNA levels in Mac and MC were calculated by defining the average Gata2 mRNA level in BM c-kit+ progenitors as 1.0.
(D) Td-tomato-gating histogram of Mac and MC in R26-tdT crossed with lysozyme-Cre (LysM-Cre) mice.
(E) Td-tomato-gating histogram of Mac and MC in R26-tdT crossed with histidine decarboxylase-Cre (Hdc-Cre) mice.
(F) Cell frequencies of Mac and MC in the Gata2-sufficient control (control; n = 5), G2F/+::LysM-Cre (n = 6), and G2F/F::LysM-Cre (n = 5) groups.
(G) Bacterial colony-forming units (CFU) in the peritoneal cavity at 24 h after CLP in the control (n = 6), G2F+::LysM-Cre (n = 8), and G2F/F::LysM-Cre groups (n = 5).
(H) Cell frequencies of Mac and MC in the Gata2-sufficient control (control; n = 8), G2F/+::Hdc-Cre (n = 6), and G2F/F::Hdc-Cre. Note the reduced frequencies of MC in the G2F/F:: Hdc-Cre mice.
(I) Bacterial CFU in the peritoneal cavity at 24 h after CLP in the control (n = 9), G2F/+::Hdc-Cre (n = 7), and G2F/F::Hdc-Cre (n = 5) groups.
One representative data point from each of two experiments is presented in Figures 5B, 5D, and 5E. Results in Figures 5C and 5F–5I were calculated from more than three experiments. Dots in Figures 5C and 5F–5I represent independent biological replicates. Data in Figures 5C, 5F, and 5H are represented as the means ± SEM. Data in Figures 5G and 5I are represented as the median and interquartile range. p Values were calculated with the unpaired t test for Figures 5F and 5H and with the Wilcoxon-Mann-Whitney test for Figures 5G and 5I. Gata2F/F mice without the cre transgene or Gata2+/+ mice with the cre transgene were used as control mice in Figures 5F–5I. See also Figure S6 for further information.
We next examined whether GATA2 expression in peritoneal macrophages and mast cells might contribute to bacterial clearance. To this end, we employed Gata2 floxed mice (G2F) crossed with lysozyme (LysM)-Cre mice or histidine decarboxylase (Hdc)-Cre transgenic mice that express cre recombinase specifically in macrophages or mast cells, respectively (Figure 5A) (Charles et al., 2006; Clausen et al., 1999). We tested and confirmed that LysM-Cre and Hdc-Cre activity was specifically detected in Mac and MC, respectively, utilizing the ROSA26 tdTomato (R26-tdT) reporter system (Figures 5D and 5E). We further validated the residual unrecombined Gata2 floxed allele in Mac and ΜC from LysM- and Hdc-Cre mice, respectively, by quantitative genomic PCR using a Gata2 floxed allele-specific primer. In support of results using the R26-tdT reporter system, we found that 87% of copies of the Gata2 floxed allele were excised specifically in Mac of G2F/F::LysM-Cre, but there was little or no excision in Mac of G2F/F::Hdc-Cre (Figure S6). Meanwhile, we failed to determine the residual unrecombined Gata2 floxed allele in MC due to an insufficient MC fraction (approximately 104 cells per G2F/F mouse) in the peritoneum (Figure S6). Notably, G2F/F::Hdc-Cre mice had a significant 76% reduction in their mast cell counts compared with Gata2-sufficient control mice, whereas LysM-Cre-mediated Gata2 excision rarely affected the peritoneal macrophage population (Figures 5F and 5H). Considering the MC-specific activity in Hdc-cre and the reduced frequency of MC in G2F/F::Hdc-Cre mice, we inferred that the Gata2 floxed allele was excised effectively in G2F/F::Hdc-Cre mice (Figures 5E and 5H). Subsequent CLP experiments demonstrated that bacterial clearance was not significantly changed in macrophage-specific Gata2-deficient mice (Figure 5G). In contrast, the bacterial count tended to be increased, albeit by a statistically nonsignificant margin (p = 0.12), in mast-cell-specific Gata2-deficient mice compared with Gata2-sufficient control mice (Figure 5I). Collectively, these results indicate that GATA2 expression in peritoneal macrophages and mast cells affected bacterial clearance only marginally, if at all.
Discussion
One of the major clinical manifestations of syndromic GATA2 haploinsufficiency is primary immunodeficiency with marked susceptibility to weakly virulent bacterial, viral, and fungal infections, suggesting that GATA2 haploinsufficiency compromises the innate immune system of the host (Jung et al., 2020; Shimizu and Yamamoto, 2020). However, immunological insights and molecular explanations for GATA2-haploinsufficient patients are largely lacking. We demonstrate here that Gata2 heterozygous mutant mice show attenuated inflammatory responses to LPS and suffer from susceptibility to bacterial peritonitis in a CLP model.
The reported human GATA2 mutations thus far have been categorized into two groups, i.e., quantitative and qualitative GATA2 haploinsufficiency, both of which are associated with a highly penetrant phenotype of immunodeficiency. There is a cohort of patients showing reduced or abnormal GATA2 transcripts from one allele, which are caused by mutations in the coding sequences, complete gene deletion, regulatory element mutations, or nonsense-mediated decay of mRNA (Hsu et al., 2013). Indeed, we found that G2Het mice showed an attenuated inflammatory response upon LPS stimulation and that this attenuation was accompanied by reduced cytokine levels (Figures 1 and 2). The CLP-induced polymicrobial peritonitis study employed here revealed that GATA2 is required for sufficient bacterial clearance and that even haploid mutation of Gata2 significantly weakened the elaborate immune system (Figure 4). Therefore, G2Het mice recapitulate the bacterial infection susceptibility observed in human GATA2 haploinsufficiency.
The multiple 32-plex cytokine immune analyses revealed that IFN-γ, IL-12p40, and IL-17A were most significantly diminished in the plasma of LPS-treated G2Het mice (Figures 2B and 2D). IL-12 (heterodimeric complex of IL-12p35 and IL-12p40) secreted from macrophages and dendritic cells stimulates T cells and NK cells through a specific receptor complex and promotes the production of IFN-γ (Teng et al., 2015). IFN-γ confers activity on macrophages, enabling them to kill intracellular bacteria (Ivashkiv, 2018). Mutation of IFN-γ and IL-12 signaling molecules in humans leads to impaired IFN-γ signaling and results in susceptibility to intracellular bacterial infections (Kong et al., 2018; Martínez-Barricarte et al., 2018; Tangye et al., 2020). In addition, the IL-17 protein family contains the highly homologous members IL-17A and IL-17F, which are preferentially produced by Th17 cells (Iwakura et al., 2011). The IL-17A/IL-17F cytokines induce proinflammatory cytokine output, neutrophil infiltration, and antimicrobial peptide production, thereby participating in host defense against bacterial infection (Iwakura et al., 2011). Indeed, mice with compound IL-17A and IL-17F deficiency exhibit increased susceptibility to opportunistic infection by S. aureus (Ishigame et al., 2009). As the IL-12/IFN-γ regulatory axis and IL-17 cytokines play important functions in innate immunity against low-virulence bacterial infections, one would predict that the simultaneous suppression of both cytokine systems would severely disrupt host immune defense in the event of GATA2 haploinsufficiency (Figure 6). Importantly, IFN-γ, IL-12p40 and IL-17A have not been linked previously to GATA2 function. Since these cytokines are produced mainly by T cells, NK cells, macrophages, and dendritic cells in which GATA2 is rarely expressed, as previously reported and shown in this study (Figures 1E, 1F, and 5A), non-cell-autonomous mechanisms of GATA2 haploinsufficiency must be operative in these processes (Onodera et al., 2016).
Figure 6.
A proposed model for the GATA2-directed host defense system and the pathogenesis of immunodeficiency resulting from human GATA2 haploinsufficiency
A sufficient amount of intact GATA2 promotes an adequate inflammatory response; thus, GATA2 confers host defense against pathogens (left panel). Insufficient intact GATA2 in the whole body leads to a reduced inflammatory response against pathogens, thereby impairing pathogen clearance (right panel).
Plasma IFN-γ, IL-12p40, and IL-17A were usually diminished during LPS stimulation in G2Het mice compared with WT mice, whereas some other cytokines, including IL-1α and MIP-1β, were transiently elevated in G2Het mice during the middle stages (i.e., 8 h in peritoneal lavage fluid and 4–8 h in plasma) of LPS stimulation (Figures 2 and S3). Given these results, we surmise that cytokine levels dynamically change in the middle stages depending on the regulatory kinetics of each cytokine, which can be reciprocally regulated by other cytokines. Therefore, in the middle stage, the levels of some cytokines are increased, whereas others are decreased. However, at 0 h (i.e., basal level) and 24 h, most of the cytokines were diminished in the G2Het mice relative to the WT mice. These results suggest that GATA2 may exert its most dominant effects at the basal to early stage and the terminal stage of LPS stimulation.
The phenotypic complexity of GATA2 haploinsufficiency might reflect the consequences of a dual defect between hematopoietic cells and non-hematopoietic cells expressing GATA2. To discriminate between these possibilities, we employed macrophage- or mast-cell-specific conditional deletion of Gata2 in this study. Although macrophage-specific Gata2 deletion hardly altered the efficiency of bacterial clearance, bacterial clearance was diminished slightly in the mice with mast-cell-specific GATA2 defects (Figure 5). Given these results, we speculate that GATA2 in mast cells might be partially responsible for host defense against bacteria (Figure 5). Notably, however, the bacterial clearance of mice with mast-cell-specific Gata2 deletion was less affected than that of the G2Het mice. These data raise the possibility that GATA2 expression in other cell lineages also participates in establishing the host immune defenses. GATA2 is expressed in a number of other somatic tissues, including vascular endothelium, urogenital tissues, and the kidney, in which GATA2 is known to play crucial developmental roles (Kanki et al., 2011; Khandekar et al., 2004; Linnemann et al., 2011; Yu et al., 2014). In particular, vascular endothelial cells are well known to produce cytokines in response to inflammatory stimuli (Mai et al., 2013). In fact, public data from ChIP-Atlas (https://chip-atlas.org) indicate that the chromosomal human IL-6 locus contains a number of GATA2 binding peaks in human umbilical vein endothelial cells. Therefore, it would be quite interesting to address whether GATA2 directly regulates cytokine gene expression in endothelial cells and thereby plays a role in bacterial clearance through the vascular system.
Clinical studies of GATA2-haploinsufficient patients have shown that hematopoietic stem cell transplantation is a potentially curative therapy to circumvent severe infections and hematological malignancies (Hofmann et al., 2020; Parta et al., 2018). However, a clinical survey of 79 French and Belgian GATA2-haploinsufficient patients reported that severe infections still emerged in some transplant recipients within five years after transplantation (Donadieu et al., 2018). Thus, the clinical evidence implies that defects in nonhematopoietic tissue may participate in the etiology of the immunodeficient phenotypes observed in some GATA2-haploinsufficient patients.
IL-6 is a prototypical proinflammatory cytokine that is promptly and transiently induced in response to infection. The induction of IL-6 evokes acute-phase inflammation and subsequently reinforces host immune defense (Tanaka et al., 2014). In addition to the previously reported LPS-induced inflammation model, we performed successful inflammation imaging of CLP-induced polymicrobial peritonitis and subsequent systemic inflammation by employing the hIL-6 Luc reporter system (Hayashi et al., 2015). Therefore, we reasoned that the hIL-6 Luc reporter would reliably monitor the inflammatory response and reasonably represent the host immune defense status in G2Het mice. G2Het mice crossed with hIL-6 Luc reporter mice appear to have considerable merit for further unveiling disease mechanisms and may potentially serve as a robust screening system for therapeutic development.
Given the diverse functions of GATA2 in multiple tissue contexts, it should be carefully considered whether GATA2 germline mutations uniformly disrupt GATA2 expression in all tissues or whether GATA2 haploinsufficiency affects only a subset of GATA2-regulated processes. More detailed studies on the innate immune system and pathogen clearance in G2Het mice will offer a deeper understanding of immune deficiency and open new therapeutic avenues for GATA2 haploinsufficiency diseases.
Limitations of the study
This study demonstrated that G2Het mice exhibited a weakened inflammatory response and impaired bacterial clearance. Although these phenotypes were consistent with the clinical symptoms observed in GATA2-haploinsufficient patients, further prospective studies are needed to confirm whether GATA2-haploinsufficient patients manifest a weakened inflammatory response. In addition, the molecular mechanisms underlying the regulation of inflammatory cytokines by GATA2 have not been identified. Statistically significant differences were observed in Figures 1D, 4B, and 4E; however, the biological importance of these results needs to be further validated.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Purified Anti-Mouse CD16 / CD32 (Fc Shield) (2.4G2) | TONBO | 70-0161 |
| Fixable Viability Dye eFluor™ 450 | Thermo Fisher Scientific | 65-0863- |
| CD117 (c-Kit) Monoclonal Antibody (2B8), APC | Thermo Fisher Scientific | Catalog # 17-1171; AB_469430 |
| FceR1 alpha Monoclonal Antibody (MAR-1), PE | Thermo Fisher Scientific | Catalog # 12-5898; AB_466028 |
| CD11b Monoclonal Antibody (M1/70), APC-eFluor 780 | Thermo Fisher Scientific | Catalog # 47-0112; AB_1603193 |
| Ly-6G Monoclonal Antibody (1A8-Ly6g), APC | Thermo Fisher Scientific | Catalog # 17-9668; AB_2573307 |
| Ly-6G Monoclonal Antibody (1A8-Ly6g), Super Bright 600 | Thermo Fisher Scientific | Catalog # 63-9668-82; AB_2762785 |
| F4/80 Monoclonal Antibody (BM8), PE-Cyanine7 | Thermo Fisher Scientific | Catalog # 25-4801-82; AB_469653 |
| Chemicals, peptides, and recombinant proteins | ||
| Lipopolysaccharide | Sigma-Aldrich | L2880 |
| VivoGlo™ Luciferin, In Vivo Grade | Promega | P1043 |
| Critical commercial assays | ||
| MILLIPLEX MAP Mouse Cytokine/Chemokine Magnetic Bead Panel - Premixed 32 Plex | Merk | MCYTMAG-70K-PX32 |
| MILLIPLEX MAP Mouse Cytokine/Chemokine Magnetic Bead Panel – 6 Plex | Merk | MCYTOMAG-70K-06 |
| MagextractorTM -Genome- | TOYOBO | NPK-101 |
| Agencourt® RNAdvanceTM Tissue Kit | BECKMAN COULTER | A32645 |
| ReverTra Ace® qPCR RT Master Mix with gDNA Remove | TOYOBO | FSQ-301 |
| THUNDERBIRD® Next SYBR® qPCR Mix | TOYOBO | QPX-201 |
| Experimental models: Organisms/strains | ||
| Gata2 heterozygous mutant mice/ C57B6N-Tyrc-Brd/BrdCrCrl | Tsai et al., 1994 | N.A. |
| human IL-6 luciferase transgenic mice/ C57B6N-Tyrc-Brd/BrdCrCrl | Hayashi et al., 2015 | N.A. |
| Gata2 Floxed mice/C57BL/6J | Charles et al., 2006 | N.A. |
| Gata2 GFP knockin mice/C57BL/6J | Suzuki et al., 2006 | N.A. |
| lysozyme-Cre mice/C57BL/6J | Clausen et al., 1999 | N.A. |
| Histidine decarboxylase–Cre mice/C57BL/6J | Mutant Mouse Resource & Research Centers | RRID: MMRRC_037409-UCD |
| ROSA26-tdTomato mice/C57BL/6J | Madisen et al., 2010 | N.A. |
| Software and algorithms | ||
| Living Image software | PerkinElmer | N.A. |
| FlowJo | BD Biosciences | N.A. |
| xPONENT | Merk | N.A. |
| Excel | Microsoft | N.A. |
| Prism 8 | GraphPad Software | N.A. |
Resource availability
Lead contact
Further information and requests should be directed by the Lead Contact, Takashi Moriguchi (moriguchi@tohoku-mpu.ac.jp).
Material availability
No unique materials were generated in this study.
Data code and availability
The data generated or analyzed in this study are available upon request from the Lead Contact, Takashi Moriguchi (moriguchi@tohoku-mpu.ac.jp).
Experimental models and subject details
Mice
The Gata2 heterozygous mutant mice, human IL-6 luciferase (hIL-6 Luc) transgenic mice, Gata2Flox mice, Gata2 GFP knock-in mice, lysozyme-Cre mice and ROSA26-tdTomato mice used for this study have been described previously (Charles et al., 2006; Clausen et al., 1999; Madisen et al., 2010; Suzuki et al., 2006; Tsai et al., 1994). Histidine decarboxylase–Cre mice (RRID: MMRRC_037409-UCD) were purchased from the Mutant Mouse Resource & Research Centers (https://www.mmrrc.org). Littermate mice of genotypes, namely, Gata2+/+, Gata2Flox/Flox without Cre recombinase transgene and Gata2+/+ with the Cre recombinase transgene, were used as controls in each analysis. All mutant mice were crossed with a C57B6N-Tyrc-Brd/BrdCrCrl (B6 albino mouse) or C57BL/6J genetic background in this study. Both male and female mice at 10–16 weeks were mainly used for this study. All mice were handled according to the regulations of the Standards for Humane Care and Use of Laboratory Animals of Tohoku Medical and Pharmaceutical University and the Guidelines for the Proper Conduct of Animal Experiments from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan. All animal experiments were approved by the Tohoku Medical Pharmaceutical University Animal Experiment Committee (registration number: 20002-cn).
Methods details
LPS-induced inflammation and analysis of the mortality rate
LPS-induced inflammation was executed by intraperitoneal (i.p.) or subcutaneous (s.c.) injection of lipopolysaccharide (LPS; Sigma-Aldrich, St. Louis, Missouri) at a dose of 1 mg/kg or 10 mg/kg body weight. To analyze the mortality rate after LPS administration, we used a sublethal dose of LPS at 10 mg/kg as previously reported (Genster et al., 2017) and monitored the viability of the mice twice a day (morning and evening) for 168 hr (7 days) after LPS administration.
Imaging of luminescence in vivo
In vivo imaging was conducted utilizing an in vivo imaging system (IVIS; PerkinElmer, Waltham, MA) as previously described (Hayashi et al., 2015). Briefly, hIL-6 Luc transgenic mice were intraperitoneally injected with 75 mg/kg D-luciferin for analysis of LPS-induced inflammation or subcutaneously injected with 150 mg/kg D-luciferin for analysis of cecal ligation and puncture (CLP)-induced polymicrobial peritonitis. Subsequently, the anesthetized mice were placed in a light-sealed chamber, and the luminescence was imaged for 60 s to monitor LPS-induced inflammation or for 180 s to monitor CLP-induced polymicrobial peritonitis. Luminescence emitted from various regions of the mouse was quantified with Living Image software (PerkinElmer, Waltham, MA).
Hematological analysis, cell counting and flow cytometry analyses
Single-cell suspensions were prepared from bone marrow, peripheral blood and peritoneal lavage fluid, and then these suspensions were used for subsequent experiments. Hematological indices were automatically measured using a Celltac α-MEK-6550 (Nihonkoden, Tokyo, Japan). Cell counts were examined using a TC20™ Automated Cell Counter (Bio-Rad Laboratories, Hercules, CA). Red blood cells were lysed using RBC lysis buffer (Thermo Fisher Scientific, Waltham, MA). Fc receptor blocking was performed with anti-CD16/32 (2.4G2 hybridoma culture supernatant, dilution 1:400). Immunostaining was performed with each anti-body for 30 min at 4°C and subsequently cell suspensions were washed with phosphate-buffered saline (PBS) containing 2% fetal bovine serum (FACS buffer) as previously described (Takai et al., 2019). Fixable Viability Dye eFluor™ 450 (Thermo Fisher Scientific, Waltham, MA)–negative live cells were subjected to all of the flow cytometry analyses. Flow cytometry analysis was performed using a BD LSRFortessa™ X-20 (BD Biosciences, San Jose, CA), a FACSAria Fusion (BD Biosciences, San Jose, CA) and an Attune NxT Flow Cytometer (Thermo Fisher Scientific, Waltham, MA). Cell sorting was conducted with a FACSAria Fusion (BD Biosciences, San Jose, CA). The data were analyzed with FlowJo software (BD Biosciences, San Jose, CA). The following flow cytometry antibodies were diluted and used with FACS buffer: CD11b (M1/70, dilution 1:400), FcεR (MAR-1, dilution 1:200), c-kit (2B8, dilution 1:200), F4/80 (BM8, dilution 1:200) and Ly6G (1A8, dilution 1:200). The gating strategies are shown in Figure S4.
Cytokine multiplex analysis
After the intraperitoneal injection of PBS or LPS, peritoneal lavage fluid was collected with 1 mL PBS. Plasma was also collected from submandibular blood. Multiplex cytokine analysis was performed with diluted samples and a 6-plex/32-plex Mouse Cytokine/Chemokine Magnetic Bead Panel (Merck Millipore, Barrington, MA) on a MAGPIX (Merck Millipore, Barrington, MA) according to the manufacturer’s instructions. The concentration of each cytokine was automatically calculated by the xPONENT system on a MAGPIX.
CLP-induced polymicrobial peritonitis and counting of colony-forming units (CFU)
Cecal ligation and puncture (CLP) was performed as a model of polymicrobial peritonitis as previously described, with minor modifications (Rittirsch et al., 2009). Each step of the CLP procedure is shown in Figure S5. Peritoneal bacteria were collected by lavage of the peritoneal cavity with 10 mL PBS. Peritoneal lavage fluid or whole blood was plated on brain-heart infusion agar plates (BD Biosciences, San Jose, CA) and cultured at 37°C overnight. Based on serial dilution, bacterial colony counts were calculated as CFU per 10 mL for peritoneal lavage fluid or per mL for peripheral blood.
Quantitative real-time PCR
Genomic DNA was extracted from the mouse tails using Magextractor™ -Genome- (TOYOBO, Osaka, Japan), and the hematopoietic cells were sorted by FACS according to the manufacturer’s instructions. Total RNA from the sorted hematopoietic cells was extracted using the Agencourt® RNAdvance™ Tissue Kit (BECKMAN COULTER, Brea, CA) according to the manufacturer’s instructions. cDNA was synthesized using ReverTra Ace® qPCR RT Master Mix with gDNA Remover (TOYOBO, Osaka, Japan). Quantitative real-time PCR was performed using THUNDERBIRD® Next SYBR® qPCR Mix (TOYOBO, Osaka, Japan) on a CFX96 Touch™ Detection System (Bio-Rad Laboratories, Hercules, CA). The genomic DNA was normalized to the value of the Gata2 -2.8 kb locus. The mRNA expression level was normalized to the Gapdh expression level. The primers used in the quantitative real-time PCR are listed in Table S1.
Quantification and statistical analysis
Comparisons between the 2 groups were made using the unpaired t-test and the Mann-Whitney U test as described in the figure legends. The data are presented as the mean ± SEM or median ± interquartile range as described in the figure legends. Since a reduced inflammatory response in G2Het mice was demonstrated in Figure 1, one-tailed tests were adopted in Figures 2, Fig 4 and Fig 5. For all analyses, the criterion for statistical significance was defined as p < 0.05. For correlation analysis, a correlation coefficient ranging from 0.50 to 0.70 was defined as moderate correlation, and a value of 0.70 to 0.90 was defined as high correlation, as previously described (Mukaka, 2012). Data management and statistical analysis were performed using Excel (Microsoft, Redmond, WA) and GraphPad Prism 8 software (GraphPad Software, San Diego, CA).
Acknowledgments
We thank Naoki Yokose, Yuka Sagae, Yukihiro Kaneyasu, and the Technical Service Division of Tohoku Medical and Pharmaceutical University for providing technical support. We also thank Satoshi Uemura, Shinya Ohmori, Kinuko Ohneda, and the Department of Medical Biochemistry, Tohoku University Graduate School of Medicine for providing technical support and suggestions. This study was supported in part by JSPS KAKENHI Grants JP 18K15068 (to J.T.), 19K07388 (to T.M. and J.T.), Innovative Areas (grant 18H05041 to T.M.), and the Takeda Science Foundation (to J.T.).
Author contributions
J.T. performed all the experiments and contributed to the project design. T.S. and T.N. performed animal experiments. T.M. and J.D.E. contributed to the project design and supervised the study. J.T., J.E., and T.M. wrote the main manuscript text. All authors contributed to the integration and discussion of the results.
Declaration of interests
The authors declare no competing interests.
Published: July 29, 2021
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2021.102836.
Supplemental information
References
- Becker K., Hu Y., Biller-Andorno N. Infectious diseases – a global challenge. Int. J. Med. Microbiol. 2006;296:179–185. doi: 10.1016/j.ijmm.2005.12.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bigley V., Haniffa M., Doulatov S., Wang X.-N., Dickinson R., McGovern N., Jardine L., Pagan S., Dimmick I., Chua I. The human syndrome of dendritic cell, monocyte, B and NK lymphoid deficiency. J. Exp. Med. 2011;208:227–234. doi: 10.1084/jem.20101459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bresnick E.H., Jung M.M., Katsumura K.R. Human GATA2 mutations and hematologic disease: how many paths to pathogenesis? Blood Adv. 2020;4:4584–4592. doi: 10.1182/bloodadvances.2020002953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cassado A.d.A., D’Império Lima M.R., Bortoluci K.R. Revisiting mouse peritoneal macrophages: heterogeneity, development, and function. Front. Immunol. 2015;6 doi: 10.3389/fimmu.2015.00225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavalla F., Araujo-Pires A.C., Biguetti C.C., Garlet G.P. Cytokine networks regulating inflammation and immune defense in the oral cavity. Curr. Oral Health Rep. 2014;1:104–113. doi: 10.1007/s40496-014-0016-9. [DOI] [Google Scholar]
- Charles M.A., Saunders T.L., Wood W.M., Owens K., Parlow A.F., Camper S.A., Ridgway E.C., Gordon D.F. Pituitary-specific Gata2 knockout: effects on gonadotrope and thyrotrope function. Mol. Endocrinol. (Baltimore, Md.) 2006;20:1366–1377. doi: 10.1210/me.2005-0378. [DOI] [PubMed] [Google Scholar]
- Clausen B.E., Burkhardt C., Reith W., Renkawitz R., Förster I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 1999;8:265–277. doi: 10.1023/a:1008942828960. [DOI] [PubMed] [Google Scholar]
- Collin M., Dickinson R., Bigley V. Haematopoietic and immune defects associated with GATA2 mutation. Br. J. Haematol. 2015;169:173–187. doi: 10.1111/bjh.13317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickinson R.E., Griffin H., Bigley V., Reynard L.N., Hussain R., Haniffa M., Lakey J.H., Rahman T., Wang X.-N., McGovern N. Exome sequencing identifies GATA-2 mutation as the cause of dendritic cell, monocyte, B and NK lymphoid deficiency. Blood. 2011;118:2656–2658. doi: 10.1182/blood-2011-06-360313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donadieu J., Lamant M., Fieschi C., de Fontbrune F.S., Caye A., Ouachee M., Beaupain B., Bustamante J., Poirel H.A., Isidor B. Natural history of GATA2 deficiency in a survey of 79 French and Belgian patients. Haematologica. 2018;103:1278–1287. doi: 10.3324/haematol.2017.181909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., O'Geen H., Keles S., Blahnik K., Linnemann A.K., Kang Y.-A., Choi K., Farnham P.J., Bresnick E.H. Discovering hematopoietic mechanisms through genome-wide analysis of GATA factor chromatin occupancy. Mol. Cell. 2009;36:667–681. doi: 10.1016/j.molcel.2009.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genster N., Ostrup O., Schjalm C., Eirik Mollnes T., Cowland J.B., Garred P. Ficolins do not alter host immune responses to lipopolysaccharide-induced inflammation in vivo. Sci. Rep. 2017;7:3852. doi: 10.1038/s41598-017-04121-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hahn C.N., Chong C.-E., Carmichael C.L., Wilkins E.J., Brautigan P.J., Li X.-C., Babic M., Lin M., Carmagnac A., Lee Y.K. Heritable GATA2 mutations associated with familial myelodysplastic syndrome and acute myeloid leukemia. Nat. Genet. 2011;43:1012–1017. doi: 10.1038/ng.913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harigae H., Takahashi S., Suwabe N., Ohtsu H., Gu L., Yang Z., Tsai F.Y., Kitamura Y., Engel J.D., Yamamoto M. Differential roles of GATA-1 and GATA-2 in growth and differentiation of mast cells. Genes Cells. 1998;3:39–50. doi: 10.1046/j.1365-2443.1998.00166.x. [DOI] [PubMed] [Google Scholar]
- Hayashi M., Takai J., Yu L., Motohashi H., Moriguchi T., Yamamoto M. Whole-body in vivo monitoring of inflammatory diseases exploiting human interleukin 6-luciferase transgenic mice. Mol. Cell Biol. 2015;35:3590–3601. doi: 10.1128/MCB.00506-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofmann I., Avagyan S., Stetson A., Guo D., Al-Sayegh H., London W.B., Lehmann L. Comparison of outcomes of myeloablative allogeneic stem cell transplantation for pediatric patients with bone marrow failure, myelodysplastic syndrome and acute myeloid leukemia with and without germline GATA2 mutations. Biol. Blood Marrow Transplant. 2020;26:1124–1130. doi: 10.1016/j.bbmt.2020.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu A.P., Johnson K.D., Falcone E.L., Sanalkumar R., Sanchez L., Hickstein D.D., Cuellar-Rodriguez J., Lemieux J.E., Zerbe C.S., Bresnick E.H., Holland S.M. GATA2 haploinsufficiency caused by mutations in a conserved intronic element leads to MonoMAC syndrome. Blood. 2013;121:3830–3837. doi: 10.1182/blood-2012-08-452763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu A.P., Sampaio E.P., Khan J., Calvo K.R., Lemieux J.E., Patel S.Y., Frucht D.M., Vinh D.C., Auth R.D., Freeman A.F. Mutations in GATA2 are associated with the autosomal dominant and sporadic monocytopenia and mycobacterial infection (MonoMAC) syndrome. Blood. 2011;118:2653–2655. doi: 10.1182/blood-2011-05-356352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ibrahim M.K., Zambruni M., Melby C.L., Melby P.C. Impact of childhood malnutrition on host defense and infection. Clin. Microbiol. Rev. 2017;30:919. doi: 10.1128/CMR.00119-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishigame H., Kakuta S., Nagai T., Kadoki M., Nambu A., Komiyama Y., Fujikado N., Tanahashi Y., Akitsu A., Kotaki H. Differential roles of interleukin-17a and -17F in host defense against mucoepithelial bacterial infection and allergic responses. Immunity. 2009;30:108–119. doi: 10.1016/j.immuni.2008.11.009. [DOI] [PubMed] [Google Scholar]
- Ivashkiv L.B. IFNγ: signalling, epigenetics and roles in immunity, metabolism, disease and cancer immunotherapy. Nat. Rev. Immunol. 2018;18:545–558. doi: 10.1038/s41577-018-0029-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwakura Y., Ishigame H., Saijo S., Nakae S. Functional specialization of interleukin-17 family members. Immunity. 2011;34:149–162. doi: 10.1016/j.immuni.2011.02.012. [DOI] [PubMed] [Google Scholar]
- Johnson K.D., Hsu A.P., Ryu M.-J., Wang J., Gao X., Boyer M.E., Liu Y., Lee Y., Calvo K.R., Keles S. Cis-element mutated in GATA2-dependent immunodeficiency governs hematopoiesis and vascular integrity. J. Clin. Invest. 2012;122:3692–3704. doi: 10.1172/JCI61623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnzon C.-F., Rönnberg E., Pejler G. The role of mast cells in bacterial infection. Am. J. Pathol. 2016;186:4–14. doi: 10.1016/j.ajpath.2015.06.024. [DOI] [PubMed] [Google Scholar]
- Jung S., Gies V., Korganow A.-S., Guffroy A. Primary immunodeficiencies with defects in innate immunity: focus on orofacial manifestations. Front. Immunol. 2020;11:1065. doi: 10.3389/fimmu.2020.01065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanki Y., Kohro T., Jiang S., Tsutsumi S., Mimura I., Suehiro J., Wada Y., Ohta Y., Ihara S., Iwanari H. Epigenetically coordinated GATA2 binding is necessary for endothelium-specific endomucin expression. EMBO J. 2011;30:2582–2595. doi: 10.1038/emboj.2011.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khabbaz R.F., Moseley R.R., Steiner R.J., Levitt A.M., Bell B.P. Challenges of infectious diseases in the USA. The Lancet. 2014;384:53–63. doi: 10.1016/S0140-6736(14)60890-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khandekar M., Suzuki N., Lewton J., Yamamoto M., Engel J.D. Multiple, distant Gata2 enhancers specify temporally and tissue-specific patterning in the developing urogenital system. Mol. Cell Biol. 2004;24:10263. doi: 10.1128/MCB.24.23.10263-10276.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong X.-F., Martinez-Barricarte R., Kennedy J., Mele F., Lazarov T., Deenick E.K., Ma C.S., Breton G., Lucero K.B., Langlais D. Disruption of an antimycobacterial circuit between dendritic and helper T cells in human SPPL2a deficiency. Nat. Immunol. 2018;19:973–985. doi: 10.1038/s41590-018-0178-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar H., Kawai T., Akira S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 2011;30:16–34. doi: 10.3109/08830185.2010.529976. [DOI] [PubMed] [Google Scholar]
- Li Y., Gao J., Kamran M., Harmacek L., Danhorn T., Leach S.M., O’Connor B.P., Hagman J.R., Huang H. GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super-enhancers. Nat. Commun. 2021;12:494. doi: 10.1038/s41467-020-20766-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Liu B., Harmacek L., Long Z., Liang J., Lukin K., Leach S.M., O'Connor B., Gerber A.N., Hagman J. The transcription factors GATA2 and microphthalmia-associated transcription factor regulate Hdc gene expression in mast cells and are required for IgE/mast cell-mediated anaphylaxis. J. Allergy Clin. Immunol. 2017 doi: 10.1016/j.jaci.2017.10.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Qi X., Liu B., Huang H. The STAT5-GATA2 pathway is critical in basophil and mast cell differentiation and maintenance. J. Immunol. 2015;194:4328–4338. doi: 10.4049/jimmunol.1500018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim K.-C., Hosoya T., Brandt W., Ku C.-J., Hosoya-Ohmura S., Camper S.A., Yamamoto M., Engel J.D. Conditional Gata2 inactivation results in HSC loss and lymphatic mispatterning. J. Clin. Invest. 2012;122:3705–3717. doi: 10.1172/JCI61619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linnemann A.K., O’Geen H., Keles S., Farnham P.J., Bresnick E.H. Genetic framework for GATA factor function in vascular biology. Proc. Natl. Acad. Sci. U S A. 2011;108:13641. doi: 10.1073/pnas.1108440108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luesink M., Hollink I.H.I.M., van der Velden V.H.J., Knops R.H.J.N., Boezeman J.B.M., de Haas V., Trka J., Baruchel A., Reinhardt D., van der Reijden B.A. High GATA2 expression is a poor prognostic marker in pediatric acute myeloid leukemia. Blood. 2012;120:2064–2075. doi: 10.1182/blood-2011-12-397083. [DOI] [PubMed] [Google Scholar]
- Madisen L., Zwingman T.A., Sunkin S.M., Oh S.W., Zariwala H.A., Gu H., Ng L.L., Palmiter R.D., Hawrylycz M.J., Jones A.R. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 2010;13:133–140. doi: 10.1038/nn.2467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mai J., Virtue A., Shen J., Wang H., Yang X.-F. An evolving new paradigm: endothelial cells--conditional innate immune cells. J. Hematol. Oncol. 2013;6:61. doi: 10.1186/1756-8722-6-61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martínez-Barricarte R., Markle J.G., Ma C.S., Deenick E.K., Ramírez-Alejo N., Mele F., Latorre D., Mahdaviani S.A., Aytekin C., Mansouri D. Human IFN-γ immunity to mycobacteria is governed by both IL-12 and IL-23. Sci. Immunol. 2018;3:eaau6759. doi: 10.1126/sciimmunol.aau6759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng W., Paunel-Görgülü A., Flohé S., Hoffmann A., Witte I., MacKenzie C., Baldus S.E., Windolf J., Lögters T.T. Depletion of neutrophil extracellular traps in vivo results in hypersusceptibility to polymicrobial sepsis in mice. Crit. Care. 2012;16:R137. doi: 10.1186/cc11442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moriguchi T., Yamamoto M. A regulatory network governing Gata1 and Gata2 gene transcription orchestrates erythroid lineage differentiation. Int. J. Hematol. 2014;100:417–424. doi: 10.1007/s12185-014-1568-0. [DOI] [PubMed] [Google Scholar]
- Mukaka M.M. Statistics corner: a guide to appropriate use of correlation coefficient in medical research. Malawi Med. J. 2012;24:69–71. [PMC free article] [PubMed] [Google Scholar]
- Muñoz-Carrillo J.L., Contreras-Cordero J.F., GutiÈrrez-Coronado O., Villalobos-GutiÈrrez P.T., Ramos-Gracia L.G., Hern·ndez-Reyes V.E. In: Immune Response Activation and Immunomodulation. Tyagi R.K., Bisen P.S., editors. IntechOpen); 2018. Cytokine profiling plays a crucial role in activating immune system to clear infectious pathogens; pp. 9–38. [DOI] [Google Scholar]
- Ohmori S., Moriguchi T., Noguchi Y., Ikeda M., Kobayashi K., Tomaru N., Ishijima Y., Ohneda O., Yamamoto M., Ohneda K. GATA2 is critical for the maintenance of cellular identity in differentiated mast cells derived from mouse bone marrow. Blood. 2015;125:3306–3315. doi: 10.1182/blood-2014-11-612465. [DOI] [PubMed] [Google Scholar]
- Ohmori S., Takai J., Ishijima Y., Suzuki M., Moriguchi T., Philipsen S., Yamamoto M., Ohneda K. Regulation of GATA factor expression is distinct between erythroid and mast cell lineages. Mol. Cell Biol. 2012;32:4742–4755. doi: 10.1128/MCB.00718-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohmori S.Y., Ishijima Y., Numata S., Takahashi M., Sekita M., Sato T., Chugun K., Yamamoto M., Ohneda K. GATA2 and PU.1 collaborate to activate the expression of the mouse <em>Ms4a2</em> gene, encoding FcεRIβ, through distinct mechanisms. Mol. Cell Biol. 2019;39 doi: 10.1128/MCB.00314-19. e00314-00319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohneda K., Ohmori S., Yamamoto M. Mouse tryptase gene expression is coordinately regulated by GATA1 and GATA2 in bone marrow-derived mast cells. Int. J. Mol. Sci. 2019;20 doi: 10.3390/ijms20184603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onodera K., Fujiwara T., Onishi Y., Itoh-Nakadai A., Okitsu Y., Fukuhara N., Ishizawa K., Shimizu R., Yamamoto M., Harigae H. GATA2 regulates dendritic cell differentiation. Blood. 2016;128:508–518. doi: 10.1182/blood-2016-02-698118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ostergaard P., Simpson M.A., Connell F.C., Steward C.G., Brice G., Woollard W.J., Dafou D., Kilo T., Smithson S., Lunt P. Mutations in GATA2 cause primary lymphedema associated with a predisposition to acute myeloid leukemia (Emberger syndrome) Nat. Genet. 2011;43:929–931. doi: 10.1038/ng.923. [DOI] [PubMed] [Google Scholar]
- Parta M., Shah N.N., Baird K., Rafei H., Calvo K.R., Hughes T., Cole K., Kenyon M., Schuver B.B., Cuellar-Rodriguez J. Allogeneic hematopoietic stem cell transplantation for GATA2 deficiency using a busulfan-based regimen. Biol. Blood Marrow Transplant. 2018;24:1250–1259. doi: 10.1016/j.bbmt.2018.01.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rittirsch D., Huber-Lang M.S., Flierl M.A., Ward P.A. Immunodesign of experimental sepsis by cecal ligation and puncture. Nat. Protoc. 2009;4:31–36. doi: 10.1038/nprot.2008.214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimizu R., Yamamoto M. GATA-related hematologic disorders. Exp. Hematol. 2016;44:696–705. doi: 10.1016/j.exphem.2016.05.010. [DOI] [PubMed] [Google Scholar]
- Shimizu R., Yamamoto M. Quantitative and qualitative impairments in GATA2 and myeloid neoplasms. IUBMB Life. 2020;72:142–150. doi: 10.1002/iub.2188. [DOI] [PubMed] [Google Scholar]
- Spinner M.A., Sanchez L.A., Hsu A.P., Shaw P.A., Zerbe C.S., Calvo K.R., Arthur D.C., Gu W., Gould C.M., Brewer C.C. GATA2 deficiency: a protean disorder of hematopoiesis, lymphatics, and immunity. Blood. 2014;123:809–821. doi: 10.1182/blood-2013-07-515528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki M., Kobayashi-Osaki M., Tsutsumi S., Pan X., Ohmori S.y., Takai J., Moriguchi T., Ohneda O., Ohneda K., Shimizu R. GATA factor switching from GATA2 to GATA1 contributes to erythroid differentiation. Genes Cells. 2013;18:921–933. doi: 10.1111/gtc.12086. [DOI] [PubMed] [Google Scholar]
- Suzuki N., Ohneda O., Minegishi N., Nishikawa M., Ohta T., Takahashi S., Engel J.D., Yamamoto M. Combinatorial Gata2 and Sca1 expression defines hematopoietic stem cells in the bone marrow niche. Proc. Natl. Acad. Sci. United States America. 2006;103:2202–2207. doi: 10.1073/pnas.0508928103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takai J., Ohtsu H., Sato A., Uemura S., Fujimura T., Yamamoto M., Moriguchi T. Lipopolysaccharide-induced expansion of histidine decarboxylase-expressing Ly6G+ myeloid cells identified by exploiting histidine decarboxylase BAC-GFP transgenic mice. Scientific Rep. 2019;9:15603. doi: 10.1038/s41598-019-51716-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka T., Narazaki M., Kishimoto T. IL-6 in inflammation, immunity, and disease. Cold Spring Harb Perspect. Biol. 2014;6:a016295. doi: 10.1101/cshperspect.a016295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tangye S.G., Al-Herz W., Bousfiha A., Chatila T., Cunningham-Rundles C., Etzioni A., Franco J.L., Holland S.M., Klein C., Morio T. Human inborn errors of immunity: 2019 update on the classification from the international union of immunological societies expert committee. J. Clin. Immunol. 2020;40:24–64. doi: 10.1007/s10875-019-00737-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teng M.W.L., Bowman E.P., McElwee J.J., Smyth M.J., Casanova J.-L., Cooper A.M., Cua D.J. IL-12 and IL-23 cytokines: from discovery to targeted therapies for immune-mediated inflammatory diseases. Nat. Med. 2015;21:719–729. doi: 10.1038/nm.3895. [DOI] [PubMed] [Google Scholar]
- Tsai F.-Y., Keller G., Kuo F.C., Weiss M., Chen J., Rosenblatt M., Alt F.W., Orkin S.H. An early haematopoietic defect in mice lacking the transcription factor GATA-2. Nature. 1994;371:221–226. doi: 10.1038/371221a0. [DOI] [PubMed] [Google Scholar]
- Tsai F.-Y., Orkin S.H. Transcription factor GATA-2 is required for proliferation/survival of early hematopoietic cells and mast cell formation, but not for erythroid and myeloid terminal differentiation. Blood. 1997;89:3636–3643. doi: 10.1182/blood.V89.10.3636. [DOI] [PubMed] [Google Scholar]
- Tsai S.-F., Martin D.I.K., Zon L.I., D'Andrea A.D., Wong G.G., Orkin S.H. Cloning of cDNA for the major DNA-binding protein of the erythroid lineage through expression in mammalian cells. Nature. 1989;339:446–451. doi: 10.1038/339446a0. [DOI] [PubMed] [Google Scholar]
- Vinh D.C., Patel S.Y., Uzel G., Anderson V.L., Freeman A.F., Olivier K.N., Spalding C., Hughes S., Pittaluga S., Raffeld M. Autosomal dominant and sporadic monocytopenia with susceptibility to mycobacteria, fungi, papillomaviruses, and myelodysplasia. Blood. 2010;115:1519–1529. doi: 10.1182/blood-2009-03-208629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe-Asaka T., Hayashi M., Engel J.D., Kawai Y., Moriguchi T. GATA2 functions in adrenal chromaffin cells. Genes Cells. 2020;25:607–614. doi: 10.1111/gtc.12795. [DOI] [PubMed] [Google Scholar]
- Wilson D.B., Dorfman D.M., Orkin S.H. A nonerythroid GATA-binding protein is required for function of the human preproendothelin-1 promoter in endothelial cells. Mol. Cell Biol. 1990;10:4854. doi: 10.1128/MCB.10.9.4854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu T.-T., Tai Y.-T., Cherng Y.-G., Chen T.-G., Lin C.-J., Chen T.-L., Chang H.-C., Chen R.-M. GATA-2 transduces LPS-induced il-1β gene expression in macrophages via a toll-like receptor 4/MD88/MAPK-dependent mechanism. PLoS One. 2013;8:e72404. doi: 10.1371/journal.pone.0072404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto M., Ko L.J., Leonard M.W., Beug H., Orkin S.H., Engel J.D. Activity and tissue-specific expression of the transcription factor NF-E1 multigene family. Genes Dev. 1990;4:1650–1662. doi: 10.1101/gad.4.10.1650. [DOI] [PubMed] [Google Scholar]
- Yu L., Moriguchi T., Kaneko H., Hayashi M., Hasegawa A., Nezu M., Saya H., Yamamoto M., Shimizu R. Reducing inflammatory cytokine production from renal collecting duct cells by inhibiting GATA2 ameliorates acute kidney injury. Mol. Cell Biol. 2017;37:e00211–e00217. doi: 10.1128/MCB.00211-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu L., Moriguchi T., Souma T., Takai J., Satoh H., Morito N., Engel J.D., Yamamoto M. GATA2 regulates body water homeostasis through maintaining aquaporin 2 expression in renal collecting ducts. Mol. Cell Biol. 2014;34:1929–1941. doi: 10.1128/MCB.01659-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zon L.I., Yamaguchi Y., Yee K., Albee E.A., Kimura A., Bennett J.C., Orkin S.H., Ackerman S.J. Expression of mRNA for the GATA-binding proteins in human eosinophils and basophils: potential role in gene transcription. Blood. 1993;81:3234–3241. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.