Abstract
DNA polymerase α-primase is known to be phosphorylated in human and yeast cells in a cell cycle-dependent manner on the p180 and p68 subunits. Here we show that phosphorylation of purified human DNA polymerase α-primase by purified cyclin A/cdk2 in vitro reduced its ability to initiate simian virus 40 (SV40) DNA replication in vitro, while phosphorylation by cyclin E/cdk2 stimulated its initiation activity. Tryptic phosphopeptide mapping revealed a family of p68 peptides that was modified well by cyclin A/cdk2 and poorly by cyclin E/cdk2. The p180 phosphopeptides were identical with both kinases. By mass spectrometry, the p68 peptide family was identified as residues 141 to 160. Cyclin A/cdk2- and cyclin A/cdc2-modified p68 also displayed a phosphorylation-dependent shift to slower electrophoretic mobility. Mutation of the four putative phosphorylation sites within p68 peptide residues 141 to 160 prevented its phosphorylation by cyclin A/cdk2 and the inhibition of replication activity. Phosphopeptide maps of the p68 subunit of DNA polymerase α-primase from human cells, synchronized and labeled in G1/S and in G2, revealed a cyclin E/cdk2-like pattern in G1/S and a cyclin A/cdk2-like pattern in G2. The slower-electrophoretic-mobility form of p68 was absent in human cells in G1/S and appeared as the cells entered G2/M. Consistent with this, the ability of DNA polymerase α-primase isolated from synchronized human cells to initiate SV40 replication was maximal in G1/S, decreased as the cells completed S phase, and reached a minimum in G2/M. These results suggest that the replication activity of DNA polymerase α-primase in human cells is regulated by phosphorylation in a cell cycle-dependent manner.
DNA replication in eukaryotic cells takes place during a restricted period of the cell cycle, the S phase. The transition from G1 into S phase in vertebrate cells is regulated by at least two cyclin-dependent kinases, cyclin E/cdk2 and cyclin A/cdk2 (reviewed in references 37 and 38). Cyclin E/cdk2 activity peaks in late G1 (14, 26), while cyclin A/cdk2 activity appears later, with the onset of DNA synthesis (42, 44, 54). Microinjection of either an anti-cyclin A antibody, an antisense cyclin A expression plasmid (18, 40, 55, 63), or an anti-cyclin E antibody (39) prevented the entry of cells into S phase, documenting the importance of these cyclins for the G1-to-S transition. Interestingly, microinjection of anti-cyclin A antibodies after S-phase entry appeared to have no effect on DNA synthesis or S-phase progression (40), despite evidence that cyclin A/cdk2 resides in replication foci (5, 6, 47). However, cyclin A/cdk2 activity rises throughout S phase, and cyclin A is required again for the S/G2 transition (40). The requirement for cyclin E/cdk2 and cyclin A/cdk2 activities for entry into S phase implies that they are needed to modify protein substrates involved in initiation of DNA replication, but relatively little is known about how phosphorylation of physiological substrates triggers initiation of vertebrate DNA replication (reviewed in reference 60).
Initiation of DNA replication in eukaryotes is thought to involve stepwise assembly of a multiprotein complex at a replication origin that can then be activated in response to cyclin-dependent kinases at the G1/S transition (7, 23, 24; reviewed in reference 15). Events during activation probably include remodeling of the prereplication complex, recruitment of replication initiation proteins, such as DNA polymerase α-primase, and assembly of replication fork proteins (1, 53). Cyclin/cdk phosphorylation and/or proteolytic destruction of prereplication proteins after origin activation in yeast is proposed to prevent reassembly of prereplication complexes until the cyclins are destroyed during mitosis (reviewed in reference 25). However, recent evidence suggests that the destruction of prereplication proteins may not be entirely conserved in human cells (45, 62), implying the existence of other regulatory mechanisms.
Simian virus 40 (SV40) DNA replication, a model for eukaryotic DNA replication, also takes place in the nucleus during S phase and, except for SV40 T antigen, depends on cellular replication proteins (reviewed in references 4a, 19, and 21). SV40 T antigen serves many of the functions attributed to cellular prereplication complexes, but its replication activities are not inhibited by cyclin/cdks, allowing it to replicate multiple copies of the viral genome in a single S phase. It binds specifically to the viral replication origin, forming a multimeric complex, recruits replication initiation proteins to the complex through direct protein-protein interactions, and catalyzes bidirectional unwinding of the parental DNA strands. T antigen, replication protein A (RP-A), and DNA polymerase α-primase interact physically and functionally to direct primer synthesis and extension on the SV40 origin template (4, 9–13, 31, 34, 35, 46, 50). Subsequently, DNA polymerase δ and its auxiliary factors assemble at the DNA primer-template junctions to synthesize the leading strands (58, 59; reviewed in reference 19). SV40 T antigen continues to serve as a DNA helicase during elongation, and DNA polymerase α-primase, together with the DNA polymerase δ holoenzyme, replicates the lagging strands.
DNA polymerase α-primase is thus an essential factor in both initiation and elongation of SV40 DNA replication, as well as in yeast, and probably in all eukaryotes (reviewed in references 17 and 59). DNA polymerase α-primase also plays a key role in coordinating DNA replication, DNA repair, and cell cycle checkpoints (reviewed in reference 17). It is composed of four subunits: p180, the polymerase activity; p68 (also called the B subunit), whose function is presumed to be regulatory; and p58 and p48, which together constitute the primase (reviewed in references 17 and 59). DNA polymerase α-primase is a target for posttranslational modifications, including cell cycle-dependent phosphorylation of the p180 and p68 subunits in human cells at G2/M, the fission yeast p180 subunit in late S, and the p68 homolog in budding yeast at G1/S (16, 36, 41). Cell cycle-dependent phosphorylation of p58 and p48 subunits was not observed (16, 36). Phosphopeptide maps of human p180 and p68 suggested that cdc2 kinase could be responsible for the modification (36). Functional studies were not performed with the budding yeast enzyme to test for effects of phosphorylation, but the enzyme from fission yeast and human cells in G2/M was reported to have a slightly reduced affinity for single-stranded DNA compared to that from cells in G1/S (36, 41). More recently, purified recombinant human DNA polymerase α-primase was shown to be phosphorylated in vitro on p180 and p68, but not p58 or p48, by purified cyclin E/cdk2, cyclin A/cdk2, cyclin A/cdc2, and cyclin B/cdc2 (57) but not by cyclin D1/cdk4 (56). Phosphorylation by cyclin A/cdk2 and cyclin A/cdc2 in vitro strongly inhibited its ability to initiate SV40 DNA replication without affecting its primase and polymerase activities in simple enzyme assays (57).
In this study, we quantitated the inhibition of human DNA polymerase α-primase replication activity by cyclin A/cdk2, as well as its stimulation by cyclin E/cdk2. Phosphopeptide maps of the p180 and p68 subunits reveal a peptide in the p68 subunit (residues 141 to 160) that is modified well in vitro by cyclin A/cdk2 but poorly by cyclin E/cdk2. We demonstrate, by using a mutant p68 lacking four putative cyclin A/cdk2 sites in this peptide, that its modification is responsible for the inhibitory effects of cyclin A/cdk2 on the replication activity of DNA polymerase α-primase. We present evidence that p68 in DNA polymerase α-primase from human cells is modified in a cyclin E/cdk2-like pattern in G1/S and in a cyclin A-specific manner beginning in G2. Consistent with these results, we show that the capacity of DNA polymerase α-primase purified from human cells to initiate SV40 DNA replication is greatest in G1/S, decreases as the cells complete S phase, and reaches a minimum in G2/M. We propose that cyclin-dependent kinases regulate the activity of DNA polymerase α-primase in the cell cycle.
MATERIALS AND METHODS
Cell culture and synchronization.
Insect cells (High Five or SF9X) were cultured in monolayers in Grace’s medium (Gibco BRL, Gaithersburg, Md.) supplemented with 10% fetal calf serum (FCS; Hyclone, Logan, Utah) at 27°C. 293S cells were cultured as exponentially growing monolayers in Dulbecco modified Eagle (DME) medium (Gibco BRL) supplemented with 10% FCS (Gibco BRL) at 37°C.
To synchronize cells in G1/S, 5 or 10 mM thymidine (Sigma, St. Louis, Mo.) was added to the medium, which was then incubated for 24 h (43). Release from the G1/S block was accomplished by removal of the thymidine. The cells were washed twice in DME medium and then incubated for the indicated time periods in DME medium without thymidine; nocodazole (500 ng/ml; Sigma) was added 10 h after release to ensure that the cells did not pass through G2/M into G1. To obtain a G2/M block, cells were incubated with 500-ng/ml nocodazole for 16 h. Cells in G0/G1 were obtained by nutrient starvation. Cell cycle synchronization was verified by flow cytometry. Two million cells were directly stained with propidium iodide and analyzed by FACScan (Becton Dickinson, San Jose, Calif.) by J. Price at the Veteran’s Administration Hospital, Nashville, Tenn., or David McFarland at the HHMI Flow Cytometry Facility, Vanderbilt University Medical Center, Nashville, Tenn.
Metabolic phosphate labeling.
To label proteins in synchronized cells, the cells were either treated with 5 mM thymidine (for G1/S) or released for 9 h into culture medium without thymidine. Cells (107) were then labeled for 3 h with 1 mCi of [32P]orthophosphate (ca. 9,000 Ci/mmol; Dupont-NEN, Boston, Mass.) in 5 ml of phosphate-free medium (Gibco BRL) supplemented with 2% FCS. The cells in G1/S contained 5 mM thymidine during labeling. Nocodazole (500 ng/ml) was present during labeling of the released cells to prevent passage through mitosis.
Mutagenesis of p68 cDNA.
To create mutations in the four potential cdk phosphorylation sites in residues 141 to 160 of human p68 cDNA (49), the corresponding serine or threonine codons were exchanged for alanine codons by overlap extension PCR (22). Primers (Table 1 contains all of the primer sequences) were designed to introduce the desired mutation together with a novel restriction site. The resulting mismatches were flanked on each side by properly base-paired sequences of at least nine nucleotides. Primers that were complementary to the mutation primer were designed and denoted by the suffix K. By using 2 ng of wild-type (wt) p68/pUC19 as a template, two PCRs were carried out. One reaction used an upstream primer (VER) that hybridizes to pUC19 sequences just upstream of the polylinker together with a primer complementary to the mutation to amplify one fragment, and the second PCR used a downstream primer (p68H/ME) at bp 860 of the p68 coding region together with the corresponding mutation primer to amplify a second, overlapping fragment. Each reaction mixture (100 μl) contained 20 pmol each of the two primers, 20 nmol of each deoxynucleoside triphosphate, and 2.5 U of Pwo polymerase (Boehringer Mannheim) in a buffer supplied by the company. After a hot start, the PCR was carried out (32 cycles of 45 s at 94°C to denature, 60 s at 50 to 60°C to anneal, and 90 s at 72°C to elongate). The two amplification products (ca. 400 bp, depending on the mutation) were isolated and used as templates (about 20 ng of each) for another amplification in which the base-paired overlap between the two fragments in the mutated region served to prime their extension to the regions complementary to the upstream or downstream primer. These initial products were then further amplified by the upstream and downstream primers present in the same reaction mixture. The PCR conditions were identical to those described above. The resulting amplified mutant fragment (856 bp) was isolated, digested with EcoRI (a unique site in the pUC19 polylinker just upstream of the p68 insert) and BstEII (a unique site downstream of the mutations in p68 cDNA), and inserted into wt p68/pUC19 recipient DNA from which the corresponding wt EcoRI/BstEII fragment (535 bp) had been excised. Clones containing the desired mutation were identified by their novel restriction site, and second-site mutations were excluded by DNA sequencing of the amplified EcoRI/BstEII fragment.
TABLE 1.
Oligonucleotides used for mutagenesisa
| Oligonucleotide | Sequence | Underlined site |
|---|---|---|
| S141A | dGTC AAC TCG TGC ACC CCA TCA G | ApaL-I |
| S141AK | dCTG ATG GGG TGC ACG AGT TGA C | |
| S147A | dCAG CTA CTC GCG CCG TCA AG | BstU-I |
| S147AK | dCTT GAC GGC GCG AGT AGC TG | |
| S152A | dGTC AAG TTT CGC ACC AAG TGC | DraIII |
| S152AK | dGCA CTT GGT GCG AAA CTT GAC | |
| T156A | dCCA AGT GCT GCG CCC TCC CAG | HhaI |
| T156AK | dCTG GGA GGG CGC AGC ACT TGG | |
| VER | dGTT GTA AAA CGA CGG CCA GT | |
| p68H/MEb | dTCG GTC GAC CTT GTT GTT CAG CTT CCC GTT GC | SalI |
| C |
Exchanged nucleotides are in boldface; novel restriction sites are underlined.
This primer is degenerate; the alternative nucleotide is shown below.
The quadruple mutant (4×A) was constructed by successive reiterations of this procedure to introduce the second, third, and fourth mutations into the cDNA.
Protein manipulations.
Protein concentrations were determined as described by Bradford (3) by using bovine immunoglobulin G as the standard (Bio-Rad, Richmond, Calif.). Sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) (27, 48) was carried out on 7.5% polyacrylamide gels (acrylamide-bisacrylamide ratio, 30:0.36) with prestained molecular weight marker proteins (Sigma). Western blot analysis was performed as described previously (49). Monoclonal antibody 9D5 against human p68 (28) and an alkaline phosphatase-conjugated anti-mouse antibody (Promega, Madison, Wis.) were used for detection.
Purification of DNA polymerase α-primase.
Untreated, mock-phosphorylated, and prephosphorylated forms of recombinant human DNA polymerase α-primase were purified from baculovirus-infected insect cells on antibody SJK 237-71–Sepharose beads, which bind to the p180 subunit (52), as described earlier (57). To ensure optimal phosphorylation, the antibody-bound substrate was subjected to two successive incubations with cyclin-dependent kinase (700 pmol/h per 2 μg of DNA polymerase α-primase) for 15 min each. This procedure yielded a preparation that was resistant to further phosphorylation by the same kinase, as assayed by using labeled ATP (data not shown), and was suitable for mass spectrometry (see below). After prephosphorylation reactions, cyclin/cdk2 was removed by washing the column with buffer containing 50 mM Tris-HCl (pH 7.8) and 150 mM KCl. The prephosphorylated DNA polymerase α-primase eluted from the column was free of histone H1 kinase activity (data not shown).
Human DNA polymerase α-primase from 293S cells was purified as described by Takada-Takayama et al. (51). An asynchronously growing or synchronized population of 293S cells was homogenized under hypotonic conditions with 1 volume of H2O–1% aprotinin–1 μM okadaic acid (both from Sigma)–5 mM NaF. The enzyme complex was purified by affinity chromatography with monoclonal antibody SJK 287-38–Sepharose (52) as described earlier (46).
DNA polymerase α assays were performed on gapped duplex (“activated”) DNA as described earlier (49). One unit of polymerase activity was defined as the incorporation of 1 nmol of dNMP in 1 h at 37°C. The specific polymerase activities were within the previously reported range of 4,900 to 6,300 U/mg (50).
The primase activities were determined by using single-stranded M13 DNA as the substrate and quantitating the radioactive oligoribonucleotide products with a PhosphorImager. The reaction mixtures (40 μl), containing either recombinant or natural human DNA polymerase α-primase, contained 200 ng of M13 DNA in 30 mM HEPES-KOH (pH 7.8)–7 mM Mg-acetate–4 mM EGTA (pH 7.8)–5 mM NaF–1 mM dithiothreitol (DTT)–0.2 mM UTP–0.2 mM GTP–0.01 mM CTP–4 mM ATP–40 mM creatine phosphate, 1 μg of creatine kinase–0.2-mg/ml BSA in the presence of 20 μCi [α32P]CTP (3,000 Ci/mmol; Dupont-NEN). After precipitation and separation by denaturing 20% PAGE, the amount of reaction product was quantitated by PhosphorImager analysis. These empirical values were termed primase units per microliter of enzyme preparation. To determine specific activities, the silver-stained p48 bands of primase after SDS-PAGE were scanned and their intensity was quantitated densitometrically. These values were defined as relative mass units per microliter of enzyme. The primase units and mass units per microliter were used to calculate the specific activities as primase units per mass unit. The variation in specific primase activity among these preparations ranged between 3.8 and 4.6 U/mass unit for prephosphorylated recombinant DNA polymerase α-primase and between 1.8 and 3.2 U/mass unit for DNA polymerase α-primase from 293S cells.
Purification of other proteins.
The recombinant cdc2- and cdk2-binding Schizosaccharomyces pombe protein suc1 was purified as described earlier (57), by using the expression vector pRK172-suc1 (33). Recombinant baculoviruses were kindly provided by D. O. Morgan, University of California San Francisco. Cyclin/cdc2 and cyclin/cdk2 complexes coexpressed in the baculovirus system were purified by suc1 affinity chromatography as described previously (57). The kinase activities of the different kinase preparations were tested with histone H1 as the substrate and are given as picomoles of phosphate incorporated per hour. A recombinant baculovirus for cdk4-glutathione S-transferase (GST) was a generous gift of E. Harlow, Harvard University, Boston, Mass. Cyclin D1/cdk4-GST coexpressed from baculovirus vectors was purified as a GST fusion protein by adsorption to glutathione-agarose and elution with 10 mM glutathione in 50 mM Tris, pH 8.
Human RP-A was purified as described previously (20). Recombinant SV40 T antigen was expressed in High Five insect cells (49) infected with recombinant baculovirus and purified by immunoaffinity chromatography as previously described (32). Topoisomerase I was a generous gift from I. Moarefi (32) and P. Taneja.
Protein kinase and phosphatase reactions.
Kinase reactions with histone H1 or DNA polymerase α-primase as the substrate were performed as described previously (57). Briefly, 2 μg of the substrate in histone kinase buffer (20 mM HEPES-KOH [pH 7.5], 1 mM DTT, 10 mM MgCl2, 4 mM EGTA, 5 mM NaF, 1 mM EDTA, 0.1-mg/ml bovine serum albumin, 0.1 mM ATP) was incubated with cyclin/cdk complex (200 pmol/h per 2 μg of substrate unless otherwise noted) at 37°C for 20 min. The proteins were separated by SDS-PAGE on 7.5% gels, and the p68 bands were detected by Western blotting with monoclonal antibody 9D5.
Phosphatase reactions were carried out by incubating DNA polymerase α-primase that was either in solution or immobilized on SJK 132-20–Sepharose beads with 500 U of λ-phosphatase (New England Biolabs, Beverly, Mass.) in phosphatase buffer (50 mM Tris-HCl [pH 7.5], 0.1 mM EDTA, 0.01% Nonidet P-40) for 1 h at 30°C. The reaction was carried out in either the absence or the presence of phosphatase inhibitors (5 mM Na3VO4, 50 mM NaF). Detection of the p68 bands was performed as described above.
Tryptic peptide mapping.
For analysis of in vitro-phosphorylated DNA polymerase α-primase, a kinase assay was performed as described above, except that the reaction mixture contained 10 μCi of [γ-32P]ATP (3,000 Ci/mmol) and the incubation time was extended to 30 min. After electrophoretic separation by denaturing SDS-PAGE, the protein bands were transferred to polyvinylidene difluoride (PVDF) membranes. After visualization of p180 and p68 of DNA polymerase α-primase by autoradiography, pieces of filter containing each subunit were excised.
For analysis of in vivo-phosphorylated DNA polymerase α-primase, the phosphate-labeled enzyme complex from extracts of 107 293S cells was isolated by immunoprecipitation with monoclonal antibody SJK 132-20–Sepharose. Separation of the p180 and p68 subunits and their transfer to membranes were carried out as with in vitro-phosphorylated DNA polymerase α-primase.
For two-dimensional mapping of phosphate-labeled tryptic peptides (2), the membrane pieces bearing in vitro- or in vivo-labeled DNA polymerase α-primase subunits were blocked with polyvinylpyrrolidone 360 (0.5% in 100 mM acetic acid) and then subjected to trypsin digestion (2 × 10 μg of trypsin for 2 h). Phosphorylated p180 and p68 tryptic peptides were separated on thin-layer chromatography plates by electrophoresis in pH 1.9 buffer in the first dimension (anode on the left, cathode on the right) by using an HTLE-7000 apparatus (CBS Scientific, Del Mar, Calif.) and ascending chromatography in phosphochromatography buffer (37.5% n-butanol, 25% pyridine, 7.5% acetic acid) in the second dimension. The phosphorylated peptides were visualized by PhosphorImager analysis.
Mass spectrometry of p68 phosphopeptides.
Recombinant DNA polymerase α-primase was prephosphorylated in a reaction with two successive additions of cyclin/cdk for 15 min each and eluted from the antibody matrix, and the p68 subunit was isolated and digested with trypsin as described above. Tryptic peptides of cyclin A/cdk2- or cyclin E/cdk2-phosphorylated p68 were separated by reverse-phase high-performance liquid chromatography (HPLC) (data not shown), and fractions that were preferentially modified by cyclin A/cdk2, as indicated by 32P labeling, were analyzed by matrix-assisted laser desorption ionization (MALDI) time-of-flight (TOF) mass spectrometry. A Voyager Elite TOF instrument (PerSeptive Biosystems, Framingham, Mass.) equipped with a nitrogen laser (337 nm, 3-ns pulse) was used in linear mode. The accelerating voltage in the ion source was 23 kV. Data were acquired with a transient recorder with 2-ns resolution. The matrix was a saturated solution of α-cyano-4-hydroxycinnamic acid in 0.1% aqueous trifluoroacetic acid-acetonitrile (1:1, vol/vol). To prepare the samples for mass analysis, 1 μl of the peptide fraction (1 to 10 pmol/μl in 0.1% trifluoroacetic acid) was mixed with 1 μl of the matrix solution on a stainless steel plate and air dried before introduction into the instrument. Each spectrum was produced by accumulating data from 64 to 256 laser pulses. Time-to mass conversion was achieved by internal or external calibration with bradykinin (MH+ at m/z 1,061.2) and insulin (MH+ at m/z 5,734.6). The accuracy of the mass assignments was approximately ±0.1% (±1 Da/1,000 Da). MSU MassMap software (29) was used to calculate the average masses of possible peptide and phosphopeptide fragments of the protein and the m/z value of the mass spectral peak for the corresponding MH+ ion.
SV40 initiation reactions.
Initiation reactions on SV40 DNA (57 and references therein) contained 250 ng of pUC-HS DNA, 300 ng of SV40 T antigen, 600 ng of RP-A, and 300 ng of topoisomerase I in initiation buffer (30 mM HEPES-KOH [pH 7.8], 7 mM Mg-acetate, 4 mM EGTA [pH 7.8], 5 mM NaF, 1 mM DTT, 0.2 mM UTP, 0.2 mM GTP, 0.01 mM CTP, 4 mM ATP, 40 mM creatine phosphate, 1 μg of creatine kinase, 0.2-mg/ml bovine serum albumin) in the presence of 20 μCi of [α32P]CTP (3,000 Ci/mmol; Dupont NEN). The reaction was carried out with either recombinant or natural human DNA polymerase α-primase, as indicated in the figure legends. The reaction products were precipitated with 0.8 M LiCl–10 μg of sonicated salmon sperm DNA (Sigma)–120 μl of ethanol for 30 min at −70°C. The washed and dried products were redissolved in loading buffer (45% formamide, 5 mM EDTA, 0.08% xylene cyanol FF, 0.08% bromphenol blue) at 65°C for 30 min and separated by denaturing 20% PAGE for 3 to 4 h at 500 V. The reaction products were visualized by PhosphorImager analysis and quantitated as empirical SV40 initiation units.
SV40 monopolymerase replication reaction.
Monopolymerase reactions to assay primer synthesis and elongation on SV40 DNA by prephosphorylated and mock-phosphorylated DNA polymerase α-primase (35) contained 100 ng of pUC-HS DNA, 3 μg of SV40 T antigen, 400 ng of RP-A, 300 ng of topoisomerase I, and increasing amounts of DNA polymerase α-primase (5 to 15 primase units) in 30 mM Tris-acetate (pH 7.5)–7 mM Mg-acetate–4 mM EGTA (pH 7.8)–5 mM NaF–1 mM DTT–0.2 mM each UTP, CTP, and GTP–0.1 mM each dTTP, dGTP, and dATP–25 μM dCTP–4 mM ATP–40 mM creatine phosphate–1 μg of creatine kinase–0.2-mg/ml BSA in the presence of 3 μCi of [α-32P]dCTP (3,000 Ci/mmol; Dupont NEN). The reaction was carried out with DNA polymerase α-primase that had been prephosphorylated with cyclin E-cdk2 or cyclin A-cdk2 or mock phosphorylated. The reaction was stopped by addition of proteinase K (0.1-mg/ml proteinase K, 1% SDS, 1 mM EDTA). After gel filtration through G-50 Sephadex columns (Boehringer Mannheim), the reaction products were precipitated with 0.8 M LiCl–3 μg of sonicated salmon sperm DNA–160 μl of ethanol for 30 min at −70°C. The washed and dried products were redissolved in 10 μl of H2O and separated by alkaline PAGE. The reaction products were quantitated by PhosphorImager analysis.
RESULTS
Differential effects of cyclin-dependent kinases on the replication activity of recombinant human DNA polymerase α-primase.
Preliminary studies suggested that phosphorylation of recombinant human DNA polymerase α-primase by cyclin A-dependent kinases suppressed its ability to initiate SV40 DNA replication in vitro, while cyclin E/cdk2-treated polymerase α-primase remained fully active (57). To quantitate the effects of phosphorylation on the replication activity of recombinant DNA polymerase α-primase, the enzyme bound to antibody beads during immunoaffinity purification was prephosphorylated with purified cyclin A/cdk2 or cyclin E/cdk2 or, as a control, mock phosphorylated without kinase, as described in Materials and Methods. The amounts of the two kinases used had equal activity with histone H1 as the substrate (data not shown). The baculovirus-expressed recombinant human DNA polymerase α-primase used as the substrate for these experiments appeared to be essentially unphosphorylated (data not shown). After the kinase reaction, the antibody beads were thoroughly washed, each prephosphorylated DNA polymerase α-primase was eluted free of kinase activity (data not shown), and its activity was compared with that of the mock-phosphorylated enzyme.
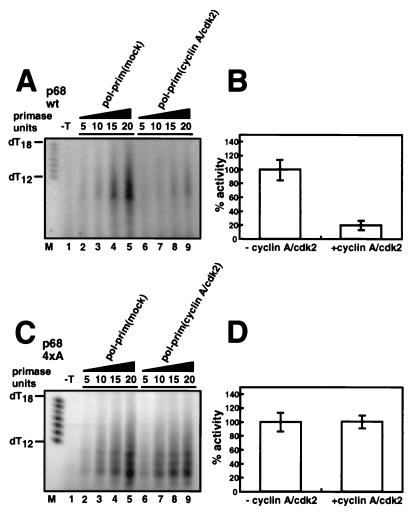
All three preparations were able to synthesize RNA primers on a single-stranded DNA template (57; data not shown), and their specific primase activities were quite similar (see Materials and Methods). Increasing volumes of each preparation of DNA polymerase α-primase with known primase activity were tested for the ability to synthesize primers on double-stranded supercoiled DNA bearing the SV40 origin of replication in the presence of purified RP-A, T antigen, and topoisomerase I. The initiation activity of DNA polymerase α-primase prephosphorylated with cyclin A/cdk2 was inhibited relative to that of the mock-phosphorylated control preparation (Fig. 1A, compare lanes 4 to 6 with lanes 1 to 3). Prephosphorylation with cyclin E/cdk2 stimulated initiation activity (Fig. 1A, compare lanes 7 to 9 with lanes 1 to 3). The signals in Fig. 1A were quantitated and divided by the primase enzyme activity of the corresponding preparation of DNA polymerase α-primase, and the results are expressed in Fig. 1B as the average SV40 initiation activity per primase unit for three or more reactions with each preparation. We estimate that initiation activity was inhibited 5 to 10-fold by phosphorylation with cyclin A/cdk2 and stimulated at least twofold by phosphorylation with cyclin E/cdk2.
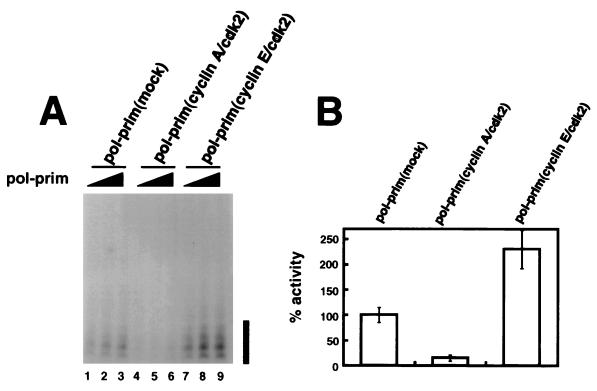
FIG. 1.
Initiation of SV40 DNA replication in vitro by DNA polymerase α-primase (pol-prim) prephosphorylated by cyclin A/cdk2 or cyclin E/cdk2. (A) Increasing amounts of the indicated preparations of DNA polymerase α-primase were tested for initiation of SV40 DNA replication. The initiation products of each reaction were separated by denaturing PAGE and quantitated by PhosphorImager analysis, and the background (data not shown) was subtracted to give the number of SV40 initiation units per microliter of DNA polymerase α-primase. The bar to the right indicates primers of 8 to 10 nucleotides. (B) The SV40 initiation activity in each reaction in A was divided by the number of primase units per microliter (data not shown) determined for the corresponding preparation of phosphorylated DNA polymerase α-primase. The bar graph shows the average relative initiation activity of each preparation; that of the mock-phosphorylated preparation was set to 100%. The error bars represent the variation obtained from three different reactions with each preparation of DNA polymerase α-primase.
Mapping of the sites of phosphorylation by cyclin-dependent kinases.
The differential effects of cyclin A/cdk2 and cyclin E/cdk2 on the initiation activity of DNA polymerase α-primase suggested that the sites modified by the two kinases differ. To test whether one or both of the phosphorylated subunits (p180 and p68) might be phosphorylated differently by the two kinases, tryptic phosphopeptide maps of both subunits were prepared. Purified recombinant DNA polymerase α-primase was phosphorylated with labeled ATP and either cyclin A/cdk2 or cyclin E/cdk2, and the p180 and p68 subunits were separated by denaturing PAGE, transferred to membranes, and digested with trypsin. The resulting peptides were separated in two dimensions and visualized by PhosphorImager analysis (Fig. 2).
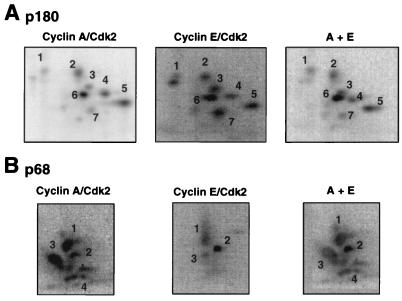
FIG. 2.
Tryptic phosphopeptide maps of in vitro-phosphorylated DNA polymerase α-primase. Two micrograms of DNA polymerase α-primase bound to SJK 132-20–Sepharose was phosphorylated by either cyclin A/cdk2 or cyclin E/cdk2 at 200 pmol/h for 30 min. After removal of the cyclin/cdk complexes, the subunits were separated by SDS-PAGE and transferred to a PVDF membrane. The labeled subunits were detected by autoradiography and excised from the membrane. The p180 (A) and p68 (B) subunits were digested twice with 10 μg of tolylsulfonyl phenylalanyl chloromethyl ketone-treated trypsin. The digested peptides were loaded onto thin-layer chromatography plates and separated by electrophoresis at pH 1.9 in the first dimension, followed by ascending chromatography in the second dimension. The labeled peptides were detected by PhosphorImager analysis. To verify identical peptides, the digests of cyclin E/cdk2- and cyclin A/cdk2-treated p180 were loaded onto the same spot and subjected to two-dimensional separation (A, far right), or the maps from the differently phosphorylated p68 subunits were superimposed (B, far right).
The phosphopeptide patterns observed for the p180 subunit were very similar with both kinases (Fig. 2A). Seven prominent phosphopeptides were generated by both kinases, with some differences in relative intensity, but all seven peptides comigrated when both digests were loaded on the same thin-layer plate (Fig. 2A, A+E). In contrast, the patterns obtained for p68 with cyclin A/cdk2 and cyclin E/cdk2 differed significantly (Fig. 2B). Cyclin A/cdk2 generated three prominent phosphopeptides (peptides 1 to 3) and several peptides migrating on a diagonal, labeled 4. Cyclin E/cdk2 also generated phosphopeptides 1 to 3, although peptide 3 was much weaker than with cyclin A/cdk2, but little or no peptide 4 was detected. Superimposition of the two maps indicated that peptides 1, 2, and 3 comigrated, suggesting that they were identical (Fig. 2B, A+E). Tryptic peptides of p68 that were phosphorylated well by cyclin A/cdk2 but poorly by cyclin E/cdk2 were also fractionated by HPLC (data not shown) and analyzed by MALDI-TOF mass spectrometry. The results demonstrated that these fractions contained the same tryptic peptide (residues 141 to 160), which occurred with either one, two, or three phosphates (Table 2). This peptide corresponded to phosphopeptide family 4, as confirmed by mutagenesis as described below (see Fig. 4A). Taken together, these data indicate that one or more sites in p68 peptide 4 were phosphorylated by well cyclin A/cdk2 and poorly by cyclin E/cdk2.
TABLE 2.
Mass spectrometry of a cyclin A/cdk2-modified tryptic peptidea
| Fraction | Observed mass (MH+) (m/z) | Residuesb | No. of sites phosphorylatedb | Expected mass (MH+) (m/z) |
|---|---|---|---|---|
| 17.5 | 2,244.74 | 141–160 or 124–140 | 2 or 5 | 2,244.25 or 2,245.01 |
| 17.5 | 2,324.10 | 141–160 or 124–140 | 3 or 6 | 2,324.23 or 2,324.99 |
| 18.0 | 2,163.96 | 141–160 or 124–140 | 1 or 4 | 2,164.27 or 2,165.03 |
| 18.0 | 2,244.13 | 141–160 or 124–140 | 2 or 5 | 2,244.25 or 2,245.01 |
| 17.5 + PPase | 2,084.83 | 141–160 | 0 | 2,084.29 |
| 18.0 + PPase | 2,084.51 | 141–160 | 0 | 2,084.29 |
HPLC fractions 17.5 and 18.0 containing the tryptic p68 peptides phosphorylated well by cyclin A/cdk2 but not by cyclin E/cdk2 were analyzed by MALDI-TOF mass spectrometry, and the observed masses were compared with those calculated for predicted tryptic peptides and phosphopeptides of p68 with MSU MassMap software (29). Mass spectrometric analyses were repeated with alkaline phosphatase-treated fractions, as indicated (+PPase).
A peptide containing residues 124 to 140 would represent an incomplete tryptic digestion product that failed to undergo hydrolysis at Lys134 and Arg135. It contains two Thr-Pro motifs. The peptide containing residues 141 to 160 would be a complete digestion product and contains one Thr-Pro and three Ser-Pro motifs. Although the observed masses would be consistent with either of the two possibilities listed, the dephosphorylation results favored the peptide containing residues 141 to 160, which was therefore tested further by mutagenesis of the predicted sites (Fig. 4A).
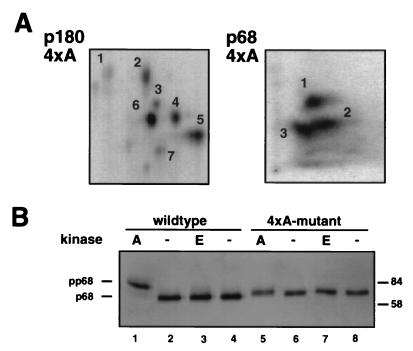
FIG. 4.
The 4× alanine mutant form of p68 lacks peptide 4 in the cyclin A/cdk2-generated pattern and shows no mobility shift after phosphorylation. (A) Purified recombinant 4× alanine (4×A) DNA polymerase α-primase was phosphorylated with cyclin A/cdk2 at 200 pmol/h, and tryptic peptide maps of p180 and p68-4×A were prepared as described in the legend to Fig. 2. (B) Wild-type or mutant polymerase α-primase (4 μg) was incubated with cyclin A/cdk2 (A) or cyclin E/cdk2 (E) (700 pmol/h) or without kinase (−). The subunits were separated by SDS–7.5% PAGE, and the p68 bands were detected by immunoblotting using monoclonal antibody 9D5. The values on the right are molecular sizes in kilodaltons.
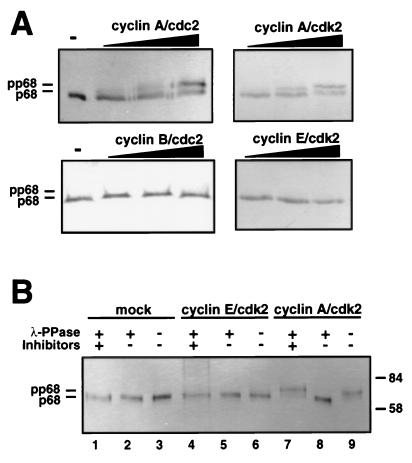
As a second, more rapid assay to detect the differential phosphorylation of p68 by cyclin-dependent kinases, we employed an electrophoretic mobility shift assay. Purified recombinant DNA polymerase α-primase was phosphorylated with increasing concentrations of cyclin-dependent kinases, electrophoresed on gels with reduced cross-linking (48), and immunoblotted to detect the p68 subunit (Fig. 3A). A slower-migrating form of p68 (termed pp68) was generated by both cyclin A/cdk2 and cyclin A/cdc2 and became more prominent as the amount of kinase complex was increased. Little difference was noted between the two kinases, despite their different kinase subunits. In contrast, pp68 was not detected after incubation with cyclin B/cdc2 or cyclin E/cdk2, when equivalent amounts of histone H1 kinase activity were used. To verify that the mobility shift was indeed caused by phosphorylation of p68 by the kinases, prephosphorylated DNA polymerase α-primase was treated with λ-phosphatase (Fig. 3B). The phosphatase had no effect on the mobility of p68 from mock phosphorylated or cyclin E/cdk2-phosphorylated DNA polymerase α-primase (lanes 1 to 6), but the slow-migrating form pp68 (lane 7) was restored to the same mobility as mock-phosphorylated p68 by phosphatase treatment (lane 8) and remained unaffected by phosphatase treatment in the presence of phosphatase inhibitors (lane 9). These results demonstrate that the shift of p68 to slower mobility was caused by phosphorylation specific to cyclin A-dependent kinases.
FIG. 3.
Cyclin A-dependent kinases induce a p68 mobility shift which is reversible with λ-phosphatase. (A) Purified recombinant DNA polymerase α-primase (2 μg) was incubated without (−) or with increasing amounts (200, 400, and 600 pmol/h) of cyclin/cdk kinases for 20 min. The subunits of DNA polymerase α-primase were separated by SDS–7.5% PAGE, and the p68 subunit was detected by Western blotting using monoclonal antibody 9D5. (B) Two micrograms of DNA polymerase α-primase, immobilized on SJK 132-20–Sepharose, was prephosphorylated with the indicated cyclin/cdk complexes or mock phosphorylated (mock), washed with phosphate-buffered saline, and incubated with (+) λ-phosphatase (λ-PPase) in the presence (+) or absence (−) of phosphatase inhibitors, as indicated, or without phosphatase (−). The p68 bands were separated by SDS-PAGE and detected by immunoblotting with monoclonal antibody 9D5. The values on the right are molecular sizes in kilodaltons.
Phosphorylation of p68 tryptic phosphopeptide 4 by cyclin A/cdk2 inhibits replication activity of DNA polymerase α-primase.
To investigate whether phosphorylation of p68 tryptic phosphopeptide 4 by cyclin A/cdk2 inhibits the SV40 replication activity of DNA polymerase α-primase, the four potential cyclin/cdk modification motifs Ser/Thr-Pro (38) in residues 141 to 160 of p68 were changed to Ala-Pro by mutagenesis of the corresponding cDNA. The mutations in the cDNA were verified by DNA sequencing and transferred into recombinant baculovirus vectors for coexpression with the other three subunits of DNA polymerase α-primase. Recombinant enzyme containing the mutant p68 (termed 4×A) assembled into a stable heterotetramer with normal yields and had polymerase and primase specific activities comparable to those of the wild-type recombinant enzyme (data not shown).
To confirm the proposed assignment of peptide 4 to residues 141 to 160 of p68 (Table 2), tryptic phosphopeptide maps of the mutant p68 subunit of cyclin A/cdk2-phosphorylated 4×A DNA polymerase α-primase were prepared (Fig. 4A). If cyclin A/cdk2 modifies one or more of these sites within residues 141 to 160 of p68, peptide 4 should be absent in the mutant p68 map. Since it is also conceivable that mutations in p68 could influence the modification pattern of the p180 subunit, tryptic phosphopeptide mapping of the p180 subunit from the 4×A DNA polymerase α-primase was carried out in parallel. The phosphopeptide pattern observed with p180 from the 4×A mutant DNA polymerase α-primase was identical to that observed with the wild-type enzyme (compare Fig. 4A with Fig. 2A), demonstrating that the mutations in p68 had no major effect on the phosphorylation of the p180 subunit. As predicted, the phosphopeptide pattern of the mutant p68 subunit differed from that of the wild-type subunit (compare Fig. 4A with Fig. 2B). Cyclin A/cdk2 still modified peptides 1 to 3 in the mutant p68 subunit, but the signal in the position of phosphopeptide 4 was clearly diminished. This result confirmed that residues 141 to 160 of p68 contain cyclin A/cdk2 sites. In agreement with this conclusion, phosphorylation of mutant DNA polymerase α-primase with cyclin A/cdk2 had little effect on the electrophoretic mobility of the mutant subunit, while that of wild-type p68 was reduced after phosphorylation, as expected (Fig. 4B).
The replication activity of the mutant enzyme complex was then compared with that of the wild-type enzyme (Fig. 5). SV40 initiation activity was assayed with increasing amounts of the purified wild-type enzyme as a positive control, either mock phosphorylated or prephosphorylated with cyclin A/cdk2. Phosphorylation inhibited its initiation activity about fivefold compared to the mock-phosphorylated preparation (Fig. 5A and B). However, the initiation activity of the 4×A mutant enzyme complex tested in parallel was largely resistant to inhibition by phosphorylation (Fig. 5C and D). These results strongly suggest that phosphorylation of one or more sites in p68 peptide 4 by cyclin A-dependent kinase inhibited the replication activity of DNA polymerase α-primase.
FIG. 5.
The SV40 initiation activity of DNA polymerase α-primase (pol-prim) containing the 4× alanine mutant form of p68 is resistant to inhibition by cyclin A/cdk2. Prephosphorylated and mock-phosphorylated wt (A) and mutant (C) forms of DNA polymerase α-primase were assayed for the ability to initiate SV40 DNA replication. The reaction products were separated on 20% urea gels as described in Materials and Methods and analyzed by PhosphorImager. A control reaction was carried out in the absence of SV40 T antigen (−T). The SV40 initiation activities of the mock- and cyclin A/cdk2-phosphorylated forms of DNA polymerase α-primase (units per microliter) were divided by the number of primase units per microliter of the corresponding preparation. The bar graphs show the SV40 initiation activities of the cyclin A/cdk2-phosphorylated wt (B) and mutant (D) forms of DNA polymerase α-primase relative to the activity of the corresponding mock-phosphorylated preparation, which was set to 100%. Error bars indicate the variation observed in three different reactions with each preparation of DNA polymerase α-primase.
Cell cycle-dependent phosphorylation of DNA polymerase α-primase in human cells regulates its replication activity.
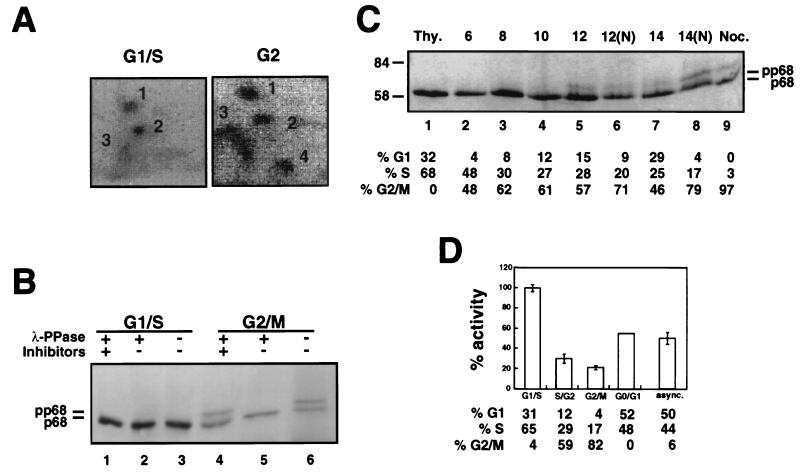
The effect of in vitro phosphorylation of DNA polymerase α-primase on its replication activity raised the question of whether cyclin-dependent kinases could also regulate its activity in vivo. To approach this question, we first investigated whether the sites modified in p68 by cyclin-dependent kinases in vitro were also modified in human cells in a cell cycle-related manner. Thus, phosphopeptide mapping was performed on the p68 subunit from DNA polymerase α-primase metabolically labeled in vivo with phosphate in G1/S and in G2. Human cells were synchronized in G1/S by thymidine addition to the medium and in G2 by release from a thymidine block and continued incubation for 9 h. Flow cytometry confirmed that the thymidine-blocked cell population was primarily in early S phase, and the blocked-and-released population was primarily in G2 (data not shown). The synchronized cells were labeled with [32P]orthophosphate for 3 h prior to preparation of cell extracts and immunoprecipitation of DNA polymerase α-primase. The p68 subunit was isolated by denaturing PAGE, blotted onto a membrane, and digested with trypsin, and the digest was separated in two dimensions to examine the phosphopeptide pattern (Fig. 6A). Two prominent phosphopeptides (1 and 2), a small amount of phosphopeptide 3, and little peptide 4 were observed in p68 from cells labeled in G1/S (Fig. 6A). This pattern strongly resembled that of p68 phosphorylated in vitro with cyclin E/cdk2 (compare Fig. 6A and 2B). Four prominent phosphopeptides were detected in p68 from cells labeled in late S/G2 (Fig. 6A). Their migration and relative intensities were nearly identical with those observed for p68 phosphorylated in vitro with cyclin A/cdk2 (compare Fig. 6A and 2B). These results confirm and extend an early report that DNA polymerase α-primase is modified on p68 in a cell cycle-dependent manner (36) and show that the sites of modification in G1/S and G2 closely resemble those modified in vitro by purified cyclin E/cdk2 and cyclin A/cdk2, respectively.
FIG. 6.
Phosphorylation pattern and replication activity of DNA polymerase α-primase purified from human cells at different stages of the cell cycle. (A) 293S cells were blocked in G1/S or blocked and released into fresh medium for 9 h prior to labeling. Cells in G1/S and late G2 were then labeled with [32P]orthophosphate for 3 h prior to preparation of cell extracts. DNA polymerase α-primase was precipitated from the extracts on SJK 132-20–Sepharose beads, separated by denaturing PAGE, and transferred to a PVDF membrane. Membrane slices containing phosphorylated p68 were digested with trypsin. Peptides were resolved by electrophoresis at pH 1.9 (left to right), followed by ascending chromatography, and detected by PhosphorImager (4 days). (B) 293S cells synchronized in G1/S with thymidine and in G2/M with nocodazole were lysed under hypotonic conditions, and DNA polymerase α-primase was immunoprecipitated with antibody SJK 132-20–Sepharose. After washing, the beads were incubated with λ-phosphatase, as indicated (+), in either the absence (−) or the presence (+) of phosphatase inhibitors. The p68 bands were separated by SDS-PAGE and detected by immunoblotting with monoclonal antibody 9D5. (C) 293S cells were blocked in G1/S and released into S phase for the times indicated. At 10 h after release, 500-ng/ml nocodazole was added to the indicated cultures (N) to prevent passage through mitosis into G1. DNA polymerase α-primase was isolated on SJK 132-20–Sepharose beads, and the p68 bands were detected after separation on denaturing 7.5% gels and immunoblotting with monoclonal antibody 9D5. Flow cytometry was performed on parallel cultures at each time point, and the cell cycle distributions are given under each lane. Thy., thymidine. The values to the left are molecular sizes in kilodaltons. (D) DNA polymerase α-primase purified from cells at different stages of the cell cycle was tested for SV40 initiation activity. Primase assays and SV40 initiation assays were performed and quantitated as described in the legends to Fig. 1 and 5 (data not shown). The bar graph shows the average SV40 initiation activity per primase unit determined for each preparation, relative to the activity of DNA polymerase α-primase in G1/S, which was set to 100%. The error bars represent the variation observed in three separate SV40 initiation assays with each preparation, except the G1/G0 preparation, which was tested only once. The cell cycle distribution of the cells from which DNA polymerase α-primase was purified was determined by flow cytometry and is indicated under each bar. async., asynchronous.
To further confirm these results, we analyzed the electrophoretic mobility of p68 from DNA polymerase α-primase isolated from unlabeled human cells synchronized in G1/S by addition of thymidine and in G2/M by addition of nocodazole. DNA polymerase α-primase was immunoprecipitated, the subunits were separated by denaturing PAGE, and the p68 subunit was visualized by immunoblotting (Fig. 6B). The p68 subunit from cells in G1/S migrated as a single band at the position expected for the unmodified or cyclin E/cdk2-modified polypeptide (lane 3), and its mobility remained unchanged after incubation with λ-phosphatase (compare lanes 1 and 2 with lane 3). In contrast, p68 from cells in G2/M migrated in two clearly distinguishable bands approximately equal in intensity (Fig. 6B, lane 6). The band with slower mobility was converted to the more rapidly migrating species by incubation with λ-phosphatase (lane 5), and the conversion was inhibited by phosphatase inhibitors (lane 6). Since the appearance of the slower-migrating form pp68 was characteristic for cyclin A/cdk2 and cyclin A/cdc2 modification in vitro (Fig. 3A), these results suggest that about half of the DNA polymerase α-primase in human cells in G2/M was specifically phosphorylated by cyclin A-dependent kinases.
The results of Fig. 6B indicate that cyclin A-dependent kinases begin to modify DNA polymerase α-primase sometime between G1/S and G2/M in human cells. To define more precisely when modification takes place, cells were blocked in G1/S with thymidine or blocked, released, and cultured for different time periods. Nocodazole was added to half of the cultures, released for 12 or 14 h, to prevent cells from progressing through mitosis into G1. The cell cycle distribution of each culture was determined by flow cytometry. DNA polymerase α-primase was isolated from each set of cultures by immunoprecipitation, and cyclin A-specific modification of p68 was detected by its altered electrophoretic mobility in denaturing gels and immunoblotting (Fig. 6C). The p68 subunit migrated as a single band in DNA polymerase α-primase from thymidine-blocked cells, as expected (lane 1), and remained unaltered until 12 h after release from the block (lanes 2 to 5), when a slower band of pp68 became prominent. Most of the cell population at this time point had progressed through the S phase into G2/M. Traces of pp68 were barely detectable in samples from cells harvested 8 to 10 h after release from the block (lanes 3 and 4), although most of the cells had also completed S phase at these time points. The amount of pp68 relative to the p68 band was unaffected by the presence of nocodazole at 12 h after release from the block (compare lanes 5 and 6) but increased at 14 h after release (compare lanes 7 and 8). Taken together, the results indicate that the cyclin A-specific modification of p68 occurred predominantly during G2 and M phase.
The differential effects of p68 phosphorylation in vitro by cyclin E/cdk2 and cyclin A/cdk2 on its replication activity and the cell cycle-dependent modification patterns of p68 in DNA polymerase α-primase from human cells suggested that differential phosphorylation of p68 may regulate the replication activity of the enzyme in the cell cycle. To test this prediction, 2 × 109 human cells were synchronized at different times in the cell cycle, and DNA polymerase α-primase was purified from them. The cell cycle distribution of each culture was determined by flow cytometry. The purified complexes from each culture contained all four subunits in approximately equimolar amounts, as expected (data not shown), and their specific activities in primase assays differed by less than twofold (see Materials and Methods). Each preparation of DNA polymerase α-primase was tested for the ability to initiate SV40 DNA replication as shown in Fig. 1. To calculate the specific initiation activity of each different preparation, the amount of reaction product obtained in several SV40 initiation reactions with each preparation (data not shown) was quantitated. To facilitate comparison, the results were expressed as a ratio of the average number of SV40 initiation units per primase unit for each preparation (Fig. 6D). The activity of DNA polymerase α-primase from cells in G1/S was maximal and was therefore set to 100%. Initiation activity of DNA polymerase α-primase dropped as the cells progressed through S phase into G2, reached a minimum in nocodazole-blocked cells (G2/M), and increased again as the cells exited from mitosis and moved into G1 (asynchronous and G0/G1). These findings demonstrate that the replication activity of DNA polymerase α-primase is regulated during the cell cycle.
DISCUSSION
Phosphorylation of human DNA polymerase α-primase p68 by cyclin-dependent kinases in vitro and in vivo.
We have identified a family of tryptic peptides from p68 that are modified well by cyclin A/cdk2 but poorly by cyclin E/cdk2 (Fig. 2B). We have shown by mass spectrometry that this family contains residues 141 to 160 in different states of phosphorylation (Table 2). The presence of this phosphopeptide family correlated with a slow-electrophoretic-mobility form of p68 generated by phosphorylation with sufficient amounts of cyclin A-dependent kinases in vitro but not by cyclin E/cdk2 (Fig. 3). Interestingly, p68 did become phosphorylated on peptide 4 by small amounts of cyclin A/cdk2 (Fig. 2B) (57), even though a pp68 band was generated only with greater amounts of kinase (Fig. 3A). This observation suggests that the slower-mobility form of p68 may depend on phosphorylation of a specific site that is modified only when kinase activity is high, or multiple sites may need to be modified. Neither phosphopeptide family 4 nor the slow-mobility form was generated by cyclin A/cdk2 phosphorylation of a mutant p68 lacking the four putative sites of phosphorylation in the peptide containing residues 141 to 160 (Fig. 4). Taken together, these data demonstrate that one or more sites within residues 141 to 160 of p68 were preferentially modified by cyclin A-dependent kinases in vitro. In contrast, phosphopeptide mapping of p180 revealed no qualitative differences between the patterns generated by cyclin A/cdk2 and cyclin E/cdk2 in vitro.
Human DNA polymerase α-primase labeled in vivo with phosphate during G1/S contained p68 whose tryptic phosphopeptides (1 to 3) closely resembled those of p68 modified in vitro by cyclin E/cdk2 (Fig. 6A). Since cyclin E/cdk2 activity was shown to be maximal in G1/S (14, 26), the simplest interpretation of the mapping data is that this kinase modified p68 in vivo. However, since these same peptides were also modified later in the cell cycle, when cyclin E/cdk2 activity had declined, it is conceivable that low levels of cyclin A/cdk2 present in G1/S generated the G1/S phosphopeptide pattern. On the other hand, since only traces of phosphopeptide 4 and the pp68 band, both characteristic for cyclin A-specific modification of p68 in vitro, were detected in G1/S (Fig. 6A and B), this interpretation would require some additional mechanism to prevent significant modification of peptide 4 during G1/S.
Both the pp68 band (Fig. 6B and C) and phosphopeptide cluster 4 (Fig. 6A) were prominent in vivo during G2, when cyclin A/cdc2 is most active (40, 42, 44, 54). Taken together with the in vitro phosphorylation data, these results provide indirect evidence that one or more sites in the p68 tryptic peptide containing residues 141 to 160 was specifically phosphorylated in vivo by a cyclin A-dependent kinase, probably cyclin A/cdc2. Although cyclin B/cdc2 activity is also maximal in G2/M (38, 42), it failed to generate the slower-mobility form of p68 in vitro when tested at the same activity as cyclin A kinases (Fig. 3A) and inhibited SV40 DNA replication less significantly in vitro (57), making cyclin A/cdc2 the more likely candidate. This interpretation is consistent with an earlier suggestion that cdc2 participates in phosphorylation of DNA polymerase α-primase in G2/M (36).
Why did cyclin A/cdk2 apparently fail to phosphorylate phosphopeptide 4 of p68 during S phase in vivo, when it was capable of modifying it quite well in vitro? One possible explanation is that since the modification of peptide 4 is dependent on the kinase concentration (Fig. 3A), the activity of cyclin A/cdk2 during S phase may be too low to catalyze the reaction efficiently or at multiple sites. Alternatively, it may be that modification occurs but is rapidly removed by a phosphatase active during S phase, such as PP2A (30). Another intriguing possibility is that the cyclin A-specific sites in p68 are masked during S phase by other replication proteins that associate with DNA polymerase α-primase (61). SV40 T antigen, for example, binds to sequences within p68 residues 1 to 240, and this interaction is proposed to play a role in SV40 DNA replication (9). On the other hand, physical interactions of DNA polymerase α-primase with T antigen in enzyme-linked immunosorbent assays and coimmunoprecipitation assays were not significantly diminished by prephosphorylation with cyclin A/cdk2 (data not shown), arguing that the phosphorylation of p68 peptide 4 probably disrupts initiation functions rather than physical association with T antigen.
Phosphorylation of human DNA polymerase α-primase regulates its initiation activity in vitro and in vivo.
RNA primer synthesis was taken as a measure of initiation activity at the SV40 origin, since primer synthesis in the absence of deoxyribonucleotides remains origin proximal regardless of the reaction time (4a), whereas in the presence of both ribo- and deoxyribonucleotides, elongation of RNA primers by DNA polymerase α-primase in the monopolymerase replication reaction is no longer restricted to the origin region and also uses the initial primer elongation products as substrates for multiple elongation events (4a, 34, 35). However, even in the monopolymerase reaction, DNA polymerase α-primase prephosphorylated by cyclin A/cdk2 synthesized two- to threefold less reaction product than the mock-phosphorylated control enzyme (data not shown). Similarly, the stimulation observed with cyclin E/cdk2-treated DNA polymerase α-primase was more modest in the monopolymerase assay than in the initiation assay (data not shown). This result suggests that, at least in vitro, phosphorylation of DNA polymerase α-primase by cyclin-dependent kinases probably regulates RNA primer synthesis rather than elongation of these primers.
The differential phosphorylation patterns observed with p68 in vitro correlate well with differences in the ability of DNA polymerase α-primase to initiate DNA replication at the SV40 origin (57; Fig. 1 and 5). In vitro, the initiation activity of recombinant DNA polymerase α-primase prephosphorylated with cyclin E/cdk2 increased at least twofold (Fig. 1). Our data do not distinguish whether stimulation of initiation activity by cyclin E/cdk2 was caused by modification of p180 or p68 or both. Specific modification of phosphopeptide 4 of the p68 subunit of DNA polymerase α-primase by cyclin A-dependent kinase correlated with a 5- to 10-fold decrease in replication initiation activity (Fig. 1 to 3). Mutagenesis of the four cdk phosphorylation motifs in phosphopeptide 4 to alanine rendered DNA polymerase α-primase resistant to inhibition of replication by phosphorylation (Fig. 5). However, attempts to mimic the negative charge of phosphate at these four sites by substituting aspartate for serine and threonine were not successful, since the replication activity of 4× aspartate mutant DNA polymerase α-primase resembled that of the unphosphorylated wild-type enzyme (data not shown). The results indicate that phosphorylation of one or more sites in phosphopeptide 4 of p68 in vitro was necessary to inhibit replication initiation.
The differential in vivo phosphorylation pattern of DNA polymerase α-primase isolated from human cells also correlated well with its replication initiation activity on the SV40 origin. In G1/S, when cyclin E/cdk2 activity peaks, DNA polymerase α-primase displayed a cyclin E/cdk2-like phosphorylation pattern on p68 and the maximal initiation activity (Fig. 6D). In G2 and G2/M, the cyclin A-specific phosphorylation of p68 on peptide 4 (Fig. 6A, B, and C) and the decline in replication initiation activity of DNA polymerase α-primase (Fig. 6D) coincided temporally with cyclin A-dependent kinase activity in the cell cycle. The correlation between the cyclin A-like phosphorylation pattern of p68 and the decline in replication activity suggests that phosphorylation of DNA polymerase α-primase on p68 peptide 4 by a cyclin A-dependent kinase may suppress its replication activity in vivo.
The relative difference between the maximal and minimal initiation activities observed with DNA polymerase α-primase prephosphorylated in vitro with cyclin E/cdk2 and cyclin A/cdk2 was 10- to 20-fold (Fig. 1B), whereas that observed with in vivo-phosphorylated DNA polymerase α-primase was only about 5-fold (Fig. 6D). Why was the effect in vivo less dramatic? One explanation is that under the conditions used for prephosphorylation with cyclin A/cdk2 in vitro, most of the p68 was maximally modified on peptide 4, as evidenced by the appearance of pp68 (Fig. 3B and 4B), whereas only one-third to one-half of the p68 was shifted to slower mobility in vivo in G2/M (Fig. 6B, lanes 4 and 6; Fig. 6C). This comparison suggests that despite the presence of phosphopeptide 4 in vivo, less than half of the enzyme was maximally modified on this peptide. If we assume that the fully modified half of the enzyme is inhibited 10-fold in G2/M and the other half retains full activity, then a rough calculation suggests that the overall initiation activity would be expected to fall to about 50%. Our data do not distinguish what fraction of DNA polymerase α-primase was modified in vitro by cyclin E/cdk2, or in vivo in G1/S, but if we assume that it was nearly complete in vitro and about half in vivo and that full modification corresponds to 2-fold stimulation in activity, then we might expect an overall stimulation in G1/S of about 1.5-fold. This would correspond to an expected difference between the maximal and minimal activities in vivo of approximately threefold, which is remarkably close to the observed difference of fivefold, given the uncertainty of the assumptions.
Based on the results presented here, we suggest that phosphorylation of DNA polymerase α-primase by cyclin-dependent kinases may regulate its activity in the cell cycle. It is easy to imagine that phosphorylation of DNA polymerase α-primase activity in G1/S by cyclin E/cdk2 could facilitate its assembly in prereplication complexes or their remodeling into elongation complexes, but it remains to be determined whether initiation at cellular origins is affected by the phosphorylation state of DNA polymerase α-primase. The importance of suppressing replication activity in G2/M is less obvious. Since DNA polymerase α-primase is implicated in coordination of DNA replication with cell cycle checkpoints (reviewed in reference 17), one might speculate that phosphorylation and down-regulation of DNA polymerase α-primase in G2/M is one of the events signaling completion of S phase. Inhibition of DNA polymerase α-primase activity by phosphorylation in G2/M could be one of several mechanisms that cooperate to prevent rereplication or, like the inhibition of transcription and RNA processing enzymes by phosphorylation in G2/M (8 and references therein), might also contribute to the major structural and functional reorganization of chromatin that occurs in preparation for mitosis (reviewed in reference 38).
ACKNOWLEDGMENTS
We thank K. Gould, C. Prives, D. Reese, M. Becker, M. Westfall, D. Lane, D. Morgan, T. Wang, E. Harlow, and the members of the Fanning lab for sharing reagents and advice with us. We thank V. Podust and A. Altman for helpful criticism of the manuscript.
We gratefully acknowledge the financial support of the NIH (RO1 GM 52948 to E.F. and RR00480 to the MSU-NIH Mass Spectrometry Facility), Vanderbilt University, a Boehringer Ingelheim Fonds predoctoral scholarship for C. Rehfuess, and the NSF (Shared Instrumentation Grant BIR-941667). Mutagenesis of the p68 cDNA was supported in part by the DFG (Fa138/6-1 and Na190/6-3) and the European Community (CHRX-CT93-0248 DG 12).
REFERENCES
- 1.Aparicio O M, Weinstein D M, Bell S P. Components and dynamics of DNA replication complexes in S. cerevisiae: redistribution of MCM proteins and Cdc45p during S phase. Cell. 1997;91:59–69. doi: 10.1016/s0092-8674(01)80009-x. [DOI] [PubMed] [Google Scholar]
- 2.Boyle W J, van der Geer P, Hunter T. Phosphopeptide mapping and phosphoamino acid analysis by two-dimensional separation on thin-layer cellulose plates. Methods Enzymol. 1991;201:110–149. doi: 10.1016/0076-6879(91)01013-r. [DOI] [PubMed] [Google Scholar]
- 3.Bradford M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- 4.Braun K A, Lao Y, He Z, Ingles C J, Wold M S. Role of protein-protein interactions in the function of replication protein A (RP-A): RP-A modulates the activity of DNA polymerase α by multiple mechanisms. Biochemistry. 1997;36:8443–8454. doi: 10.1021/bi970473r. [DOI] [PubMed] [Google Scholar]
- 4a.Bullock P A. The initiation of simian virus 40 DNA replication in vitro. Crit Rev Biochem Mol Biol. 1997;32:503–568. doi: 10.3109/10409239709082001. [DOI] [PubMed] [Google Scholar]
- 5.Canella D, Roberts J M, Fotedar R. Association of cyclin A and cdk2 with SV40 DNA in replication initiation complexes is cell cycle dependent. Chromosoma. 1997;105:349–359. doi: 10.1007/BF02529750. [DOI] [PubMed] [Google Scholar]
- 6.Cardoso M C, Leonhardt H, Nadal-Ginard B. Reversal of terminal differentiation and control of DNA replication: cyclin A and cdk2 specifically localize at subnuclear sites of DNA replication. Cell. 1993;74:979–992. doi: 10.1016/0092-8674(93)90721-2. [DOI] [PubMed] [Google Scholar]
- 7.Coleman T R, Carpenter P B, Dunphy W G. The Xenopus Cdc6 protein is essential for the initiation of a single round of DNA replication in cell-free extracts. Cell. 1996;87:53–63. doi: 10.1016/s0092-8674(00)81322-7. [DOI] [PubMed] [Google Scholar]
- 8.Colgan D F, Murthy K G K, Zhao W, Prives C, Manley J L. Inhibition of poly(A) polymerase requires p34cdc2/cyclin B phosphorylation of multiple consensus and non-consensus sites. EMBO J. 1998;17:1053–1062. doi: 10.1093/emboj/17.4.1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Collins K L, Russo A A R, Tseng B Y, Kelly T J. The role of the 70 kDa subunit of human polymerase α in DNA replication. EMBO J. 1993;12:4555–4566. doi: 10.1002/j.1460-2075.1993.tb06144.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Collins K L, Kelly T J. Effects of T antigen and replication protein A on the initiation of DNA synthesis by DNA polymerase α-primase. Mol Cell Biol. 1991;11:2108–2115. doi: 10.1128/mcb.11.4.2108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dornreiter I, Höß A, Arthur A K, Fanning E. SV40 T antigen binds directly to the catalytic subunit of DNA polymerase α. EMBO J. 1990;9:3329–3336. doi: 10.1002/j.1460-2075.1990.tb07533.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dornreiter I, Erdile F L, Gilbert I, von Winkler D, Kelly T J, Fanning E. Interaction of DNA polymerase α-primase with replication protein A and SV40 T antigen. EMBO J. 1992;11:769–776. doi: 10.1002/j.1460-2075.1992.tb05110.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dornreiter I, Copeland W C, Wang T S-F. Initiation of simian virus 40 DNA replication requires the interaction of a specific domain of human DNA polymerase α with large T antigen. Mol Cell Biol. 1993;13:809–820. doi: 10.1128/mcb.13.2.809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dulic V, Lees E, Reed S I. Association of human cyclin E with a periodic G1-S phase protein kinase. Science. 1992;257:1958–1961. doi: 10.1126/science.1329201. [DOI] [PubMed] [Google Scholar]
- 15.Dutta A, Bell S. Initiation of DNA replication in eukaryotic cells. Annu Rev Cell Dev Biol. 1997;13:293–352. doi: 10.1146/annurev.cellbio.13.1.293. [DOI] [PubMed] [Google Scholar]
- 16.Foiani M, Liberi G, Lucchini G, Plevani P. Cell cycle-dependent phosphorylation and dephosphorylation of the yeast DNA polymerase α-primase B subunit. Mol Cell Biol. 1995;15:883–891. doi: 10.1128/mcb.15.2.883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Foiani M, Lucchini G, Plevani P. The DNA polymerase α-primase complex couples DNA replication, cell-cycle progression and DNA-damage response. Trends Biochem Sci. 1997;11:411–452. doi: 10.1016/s0968-0004(97)01109-2. [DOI] [PubMed] [Google Scholar]
- 18.Girard F, Strausfeld U, Fernandez A, Lamb N. Cyclin A is required for the onset of DNA replication in mammalian fibroblasts. Cell. 1991;67:1169–1179. doi: 10.1016/0092-8674(91)90293-8. [DOI] [PubMed] [Google Scholar]
- 19.Hassell J A, Brinton B T. SV40 and polyoma DNA replication. In: DePamphilis M L, editor. DNA replication in eukaryotic cells. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1996. pp. 461–493. [Google Scholar]
- 20.Henricksen L A, Wold M S. Recombinant replication protein A: expression, complex formation, and functional characterization. J Biol Chem. 1994;269:11121–11131. [PubMed] [Google Scholar]
- 21.Herendeen D, Kelly T J. SV40 DNA replication. In: Blow J J, editor. Eukaryotic DNA replication. New York, N.Y: Oxford University Press; 1997. pp. 29–65. [Google Scholar]
- 22.Horton R M, Pease L R. Recombination and mutagenesis of DNA sequences using PCR. In: McPherson M J, editor. Directed mutagenesis—a practical approach. New York, N.Y: Oxford University Press; 1991. pp. 217–249. [Google Scholar]
- 23.Hua H X, Newport J. Identification of a preinitiation step in DNA replication that is independent of origin recognition complex and cdc6, but dependent on cdk2. J Cell Biol. 1998;140:271–281. doi: 10.1083/jcb.140.2.271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hua H X, Yan H, Newport J. A role for Cdk2 kinase in negatively regulating DNA replication during S phase of the cell cycle. J Cell Biol. 1997;137:183–192. doi: 10.1083/jcb.137.1.183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jallepalli P, Kelly T J. Cyclin-dependent kinase and initiation at eukaryotic origins: a replication switch? Curr Opin Cell Biol. 1997;9:358–363. doi: 10.1016/s0955-0674(97)80008-7. [DOI] [PubMed] [Google Scholar]
- 26.Koff A, Giordano A, Desai D, Katsumi Y, Harper J W, Elledge S, Nishimoto T, Morgan D O, Franza B R, Roberts J M. Formation and activation of a cyclin E-cdk2 complex during the G1 phase of the human cell cycle. Science. 1992;257:1689–1694. doi: 10.1126/science.1388288. [DOI] [PubMed] [Google Scholar]
- 27.Laemmli U K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 28.Lane, D. Unpublished data.
- 29.Liao P C, Leykam J, Andrews P C, Gage D A, Allison J. An approach to locate phosphorylation sites in a phosphoprotein: mass mapping by combining specific enzymatic degradation with matrix-assisted laser desorption/ionization mass spectrometry. Anal Biochem. 1994;219:9–20. doi: 10.1006/abio.1994.1224. [DOI] [PubMed] [Google Scholar]
- 30.Ludlow J W. Selective ability of S phase cell extracts to dephosphorylate large T antigen in vitro. Oncogene. 1992;7:1011–1014. [PubMed] [Google Scholar]
- 31.Melendy T, Stillman B. An interaction between replication protein A and SV40 T antigen appears essential for primosome assembly during SV40 DNA replication. J Biol Chem. 1993;268:3389–3395. [PubMed] [Google Scholar]
- 32.Moarefi I F, Small D, Gilbert I, Höpfner M, Randall S K, Schneider C, Russo A A R, Ramsperger U, Arthur A K, Stahl H, Kelly T J, Fanning E. Mutation of the cyclin-dependent kinase phosphorylation site in simian virus 40 (SV40) large T antigen specifically blocks SV40 origin DNA unwinding. J Virol. 1993;67:4992–5002. doi: 10.1128/jvi.67.8.4992-5002.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Moreno S, Hayles J, Nurse P. Regulation of p34cdc2 kinase during mitosis. Cell. 1989;58:361–372. doi: 10.1016/0092-8674(89)90850-7. [DOI] [PubMed] [Google Scholar]
- 34.Murakami Y, Hurwitz J. Functional interactions between SV40 T antigen and other replication proteins at the replication fork. J Biol Chem. 1993;268:11008–11017. [PubMed] [Google Scholar]
- 35.Murakami Y, Hurwitz J. DNA polymerase α stimulates the ATP-dependent binding of simian virus 40 T antigen to the SV40 origin of replication in vitro. J Biol Chem. 1993;268:11018–11027. [PubMed] [Google Scholar]
- 36.Nasheuer H P, Moore A, Wahl A F, Wang T S F. Cell cycle dependent phosphorylation of human DNA polymerase α. J Biol Chem. 1991;266:7893–7903. [PubMed] [Google Scholar]
- 37.Nasmyth K. Control of S phase. In: DePamphilis M L, editor. DNA replication in eukaryotic cells. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1996. pp. 331–386. [Google Scholar]
- 38.Nigg E. Cyclin-dependent protein kinases: key regulators of the eukaryotic cell cycle. BioEssays. 1995;17:471–480. doi: 10.1002/bies.950170603. [DOI] [PubMed] [Google Scholar]
- 39.Ohtsubo M, Theodoras A M, Schumacher J, Roberts J M, Pagano M. Human cyclin E, a nuclear protein essential for the G1-to-S phase transition. Mol Cell Biol. 1995;15:2612–2624. doi: 10.1128/mcb.15.5.2612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pagano M, Pepperkok R, Verde F, Ansorge W, Draetta G. Cyclin A is required at two points in the human cell cycle. EMBO J. 1992;11:961–971. doi: 10.1002/j.1460-2075.1992.tb05135.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Park H, Davis R, Wang T S F. Studies of Schizosaccharomyces pombe DNA polymerase α at different stages of the cell cycle. Nucleic Acids Res. 1995;23:4337–4344. doi: 10.1093/nar/23.21.4337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pines J, Hunter T. p34cdc2, the S and M kinase? New Biol. 1990;2:389–401. [PubMed] [Google Scholar]
- 43.Rao P T, Johnson R T. Mammalian cell fusion: studies on the regulation of DNA synthesis and mitosis. Nature. 1970;225:159–164. doi: 10.1038/225159a0. [DOI] [PubMed] [Google Scholar]
- 44.Rosenblatt J, Gu Y, Morgan D O. Human cyclin-dependent kinase 2 is activated during the S and G2 phases of the cell cycle and associates with cyclin A. Proc Natl Acad Sci USA. 1992;89:2824–2828. doi: 10.1073/pnas.89.7.2824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Saha P, Chen J, Thome K C, Lawlis S J, Hou Z-H, Hendricks M, Parvin J D, Dutta A. Human CDC6/Cdc18 associates with Orc1 and cyclin-cdk and is selectively eliminated from the nucleus at the onset of S phase. Mol Cell Biol. 1998;18:2758–2767. doi: 10.1128/mcb.18.5.2758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Schneider C, Weißhart K, Guarino L A, Dornreiter I, Fanning E. Species-specific functional interactions of DNA polymerase α-primase with simian virus 40 (SV40) T antigen require SV40 origin DNA. Mol Cell Biol. 1994;14:3176–3185. doi: 10.1128/mcb.14.5.3176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sobczak-Thepot J, Harper F, Florentin Y, Zindy F, Brechot C, Puvion E. Localization of cyclin A at the sites of cellular DNA replication. Exp Cell Res. 1993;206:43–48. doi: 10.1006/excr.1993.1118. [DOI] [PubMed] [Google Scholar]
- 48.Sorger P K, Pelham H R. Yeast heat shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell. 1988;54:855–864. doi: 10.1016/s0092-8674(88)91219-6. [DOI] [PubMed] [Google Scholar]
- 49.Stadlbauer F, Brückner A, Rehfuess C, Eckerskorn C, Lottspeich F, Förster V, Tseng B Y, Nasheuer H-P. DNA replication in vitro by recombinant DNA-polymerase-α-primase. Eur J Biochem. 1994;222:781–793. doi: 10.1111/j.1432-1033.1994.tb18925.x. [DOI] [PubMed] [Google Scholar]
- 50.Stadlbauer F, Voitenleitner C, Brückner A, Fanning E, Nasheuer H-P. Species-specific replication of simian virus 40 DNA in vitro requires the p180 subunit of human DNA polymerase α-primase. Mol Cell Biol. 1996;16:94–104. doi: 10.1128/mcb.16.1.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Takada-Takayama R, Tada S, Hanaoka F, Ui M. Peptide mapping of the four subunits of the mouse DNA polymerase-α primase complex. Biochem Biophys Res Commun. 1990;170:589–595. doi: 10.1016/0006-291x(90)92132-j. [DOI] [PubMed] [Google Scholar]
- 52.Tanaka S, Hu S-Z, Wang T S F, Korn D. Preparation and preliminary characterization of monoclonal antibodies against human DNA polymerase alpha. J Biol Chem. 1982;257:8386–8390. [PubMed] [Google Scholar]
- 53.Tanaka T, Knapp D, Nasmyth K. Loading of an Mcm protein onto DNA replication origins is regulated by Cdc6p and CDKs. Cell. 1997;90:649–660. doi: 10.1016/s0092-8674(00)80526-7. [DOI] [PubMed] [Google Scholar]
- 54.Tsai L H, Harlow E, Meyerson M. Isolation of the human cdk2 gene that encodes the cyclin A- and adenovirus E1A-associated p33 kinase. Nature. 1991;353:174–177. doi: 10.1038/353174a0. [DOI] [PubMed] [Google Scholar]
- 55.Tsai L H, Lees E, Faha B, Harlow E, Riabowol K. The cdk2 kinase is required for the G1-to-S transition in mammalian cells. Oncogene. 1993;8:1593–1602. [PubMed] [Google Scholar]
- 56.Voitenleitner, C., D. Reese, and E. Fanning. Unpublished data.
- 57.Voitenleitner C, Fanning E, Nasheuer H-P. Phosphorylation of DNA polymerase α-primase by cyclin A-dependent kinases regulates initiation of DNA replication in vitro. Oncogene. 1997;14:1611–1615. doi: 10.1038/sj.onc.1200975. [DOI] [PubMed] [Google Scholar]
- 58.Waga S, Stillman B. Anatomy of a DNA replication fork revealed by reconstitution of SV40 DNA replication in vitro. Nature. 1994;369:207–212. doi: 10.1038/369207a0. [DOI] [PubMed] [Google Scholar]
- 59.Wang T S F. Cellular DNA polymerases. In: DePamphilis M L, editor. DNA replication in eukaryotic cells. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1996. pp. 461–493. [Google Scholar]
- 60.Weisshart K, Fanning E. Phosphorylation of replication proteins. In: DePamphilis M L, editor. DNA replication in eukaryotic cells. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1996. pp. 295–330. [Google Scholar]
- 61.Wittmeyer J, Formosa T. The Saccharomyces cerevisiae DNA polymerase α catalytic subunit interacts with Cdc68/Spt16 and with Pob3, a protein similar to an HMG1-like protein. Mol Cell Biol. 1997;17:4178–4190. doi: 10.1128/mcb.17.7.4178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Yan Z, DeGregori J, Shohet R, Leone G, Stillman B, Nevins J R, Williams R S. Cdc6 is regulated by E2F and is essential for DNA replication in mammalian cells. Proc Natl Acad Sci USA. 1998;95:3603–3608. doi: 10.1073/pnas.95.7.3603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zindy F, Lamas F, Chenivesse X, Sobczak J, Wang J, Fesquet D, Henglein B, Brechot C. Cyclin A is required in S phase in normal epithelial cells. Biochem Biophys Res Commun. 1992;182:1144–1154. doi: 10.1016/0006-291x(92)91851-g. [DOI] [PubMed] [Google Scholar]