Abstract
The c-Myc oncoprotein induces cell proliferation and transformation through its activity as a transcription factor. Uncovering the genes regulated by c-Myc is an essential step for understanding these processes. We recently isolated the tumor-associated membrane protein gene, Tmp, from a c-myc-induced mouse brain tumor. Here we show that Tmp is specifically highly expressed in mammary tumors and T-cell lymphomas which develop in c-myc transgenic mice, suggesting that Tmp expression is a general characteristic of c-Myc-induced tumors. In addition, Tmp expression is induced upon serum stimulation of fibroblasts as shown in a time course closely correlated with c-myc expression. We have isolated the Tmp promoter region and identified a putative c-Myc binding element, CACGTG, located in the first intron of the gene. We show here that constructs containing the Tmp regulatory region fused to a reporter gene are activated by c-Myc through this CACGTG element and that the c-Myc–Max protein complex can bind to this element. Moreover, an inducible form of c-Myc, the MycER fusion protein, can activate the endogenous Tmp gene. We also show that Tmp-overexpressing fibroblasts induce rapidly growing tumors when injected into nude mice, suggesting that Tmp may possess a tumorigenic activity. Thus, TMP, a member of a novel family of membrane glycoproteins with a suggested role in cellular contact, is a c-Myc target and is possibly involved in c-Myc-induced transformation.
The c-Myc oncoprotein is a key regulator of cell growth and differentiation and is implicated in a variety of human tumors (30, 45, 48). c-Myc activity is associated with cell proliferation, and the expression of c-Myc can drive quiescent cells into S phase (15, 24). A role for c-Myc in the induction of apoptosis has also been established (27, 28).
c-myc encodes a basic helix-loop-helix leucine zipper (bHLH-LZ) protein, which heterodimerizes with the ubiquitous Max protein to form a nuclear transcription factor (54). A number of target genes for c-Myc transcriptional activation have been identified in recent years (reviewed in references 21, 30, and 34). The data regarding these genes do not, as yet, provide a clear understanding of the mechanisms of c-Myc activity, although initial attempts have been made to categorize the target genes into functional classes, either according to their cellular functions or to the c-Myc-related processes in which they appear to function (21, 30, 34). A role for c-Myc as a transcription repressor has also been suggested, and a number of negatively regulated genes have been described (30).
The c-Myc–Max complex activates transcription through a specific E-box hexamer, CACGTG, which acts as a binding site (11, 12), although binding to additional sequences, mainly to CATGTG, has also been observed (10, 35). In known c-Myc targets this binding site is typically located downstream to the transcription start site, most commonly in the first intron or in the 5′-untranslated region (7, 34).
We have recently isolated and characterized the Tmp (tumor-associated membrane protein) gene from mice and humans (5). Tmp was initially isolated from a cDNA library of a brain tumor which developed in a mouse transgenic for c-myc (7, 8). The gene was found to be highly expressed in the tumor but not in normal tissue (5). Tmp expression is associated with cell proliferation; it is highly expressed in proliferating fibroblasts but is downregulated when these cells are growth arrested (5). Tmp is also highly expressed in undifferentiated proliferating embryonic stem cells, but a much lower expression is observed after these cells are induced to differentiate (5). In both systems, Tmp expression closely correlates with that of c-myc. Tmp cDNAs from different mammals were isolated in parallel by several other groups (47, 57, 59, 69).
Tmp encodes a putative membrane glycoprotein of 160 amino acids, with four transmembrane domains. It is homologous to the Pmp22/Gas3 gene, which is involved in several human hereditary peripheral neuropathies (16, 51). Pmp22/Gas3 was initially characterized as a growth-arrest-specific gene, one upregulated in serum-starved fibroblasts (44). It thus displays an expression pattern inverse to that of Tmp in these cells (5). Overexpression of Pmp22/Gas3 in cultured cells can delay cell cycle progression and lead to apoptosis (29, 73, 74). The cellular function of PMP22 is, however, largely unknown, although a role in cell adhesion has been suggested (65). Recently, two additional novel genes, which are homologous to Tmp and PMP22, were identified both in mice and humans, and were named XMP/EMP2 and YMP/EMP3/HNMP-1 (5, 6, 13, 68). The four genes, Tmp, Pmp22, Xmp, and Ymp, thus define a novel gene family encoding membrane glycoproteins and are possibly involved in cellular contact.
The Tmp expression pattern, together with the fact that this gene was cloned from a c-myc-induced tumor, suggested that Tmp is a potential target for c-Myc activation. This is further supported here by the finding that high Tmp expression is a characteristic of Myc-induced tumors in the mouse. We present evidence that Tmp is a transcriptional target for c-Myc and that it may possess a tumorigenic activity.
MATERIALS AND METHODS
Cell lines and media.
NIH 3T3 and Rat1 cells were grown in Dulbecco modified Eagle medium supplemented with 50 U of penicillin per ml, 50 μg of streptomycin per ml, and 2 mM glutamine. Fetal calf serum (FCS; 10%) (Biological Industries) was added for cells in proliferation conditions, and 0.1% FCS was used for serum-starved cells. For growth induction assays, cells were grown to subconfluency, serum starved for 3 days, and then induced to proliferate with serum-rich medium. Rat1 cells stably transfected with the Tmp gene were grown in medium containing 200 μg of G418 per ml. Rat1 cells expressing the MycER and Δ106-143MycER constructs were kindly provided by Linda Z. Penn (46). The cells were grown in minimum essential medium (alpha modified) with the above supplements and 5 μg of puromycin per ml. For MycER induction experiments, cells were grown to confluency and serum starved for 3 days; then 100 nM 4-hydroxytamoxifen (4-OHT) was added to the medium. In cycloheximide experiments, 10 μg of cycloheximide per ml was added to the medium together with 4-OHT.
RNA extraction and Northern blot hybridization.
Total RNA from cells and tumors was extracted with a guanidium thiocyanate-phenol mixture as described previously (18). Northern blot hybridization was performed according to standard procedures with formaldehyde-containing gels and formamide-based hybridization solutions (58). Probes were radiolabeled by random priming (58).
Isolation of genomic clones and genomic structure analysis.
A mouse 129SVJ genomic library (Stratagene 946309) was screened with Tmp cDNA probes by standard procedures (58). The exon-intron structure was determined by sequencing and PCR with primers from within the cDNA. Sequencing of the Tmp regulatory region was performed by using an ABI automated sequencer according to standard procedures. Primer extension analysis of the Tmp transcript was performed with 1 μg of poly(A)+ RNA extracted from the BK-1 cell line, which is derived from a c-myc-induced brain tumor (7). A 1-ng portion of the primer described below (see Fig. 3B) was 32P end labeled and hybridized with the mRNA in the presence of KCl (133 mM). Avian myeloblastosis virus reverse transcriptase was used for the extension reaction under standard conditions (58).
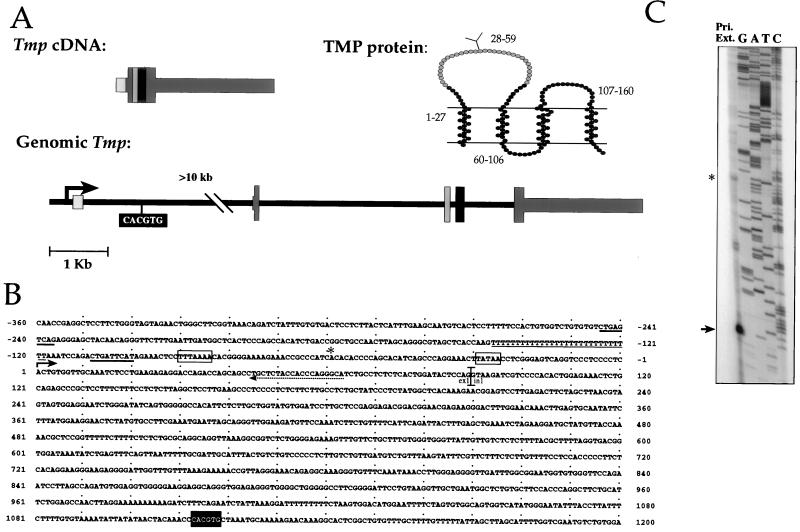
FIG. 3.
Genomic structure of Tmp and sequence of the Tmp regulatory region. (A) Tmp is composed of five exons, which are marked by differently shaded gray boxes. Exon 1 contains most of the 5′ untranslated sequence. Exons 2 to 5 contain the 480-bp coding sequence, which is marked by wide boxes. The codons of the TMP protein encoded by each exon are indicated by numbers in the protein diagram. The lines in the protein diagram represent the membrane, and a putative glycosylation site is marked on the first outer loop. (B) Sequence of the Tmp regulatory region. The start site of transcription as determined by primer extension is marked by an arrow. An asterisk marks a possible alternative start site corresponding to the asterisk-marked product in panel C. Putative TATA boxes are marked by a rectangles. The primer used for primer extension is marked by a dashed arrow. The CACGTG element is marked by a black background. AP-1 binding elements are marked by a thick line, and the poly(T) element is marked by a thin line. ex1, exon1; in1, intron1. GenBank accession number, AF055577. (C) Determination of the transcription start site. A primer extension reaction was performed with the primer indicated in panel B on RNA from a c-myc-induced brain tumor cell line used as a template. Products were run on a 4% polyacrylamide gel in parallel with a sequencing reaction of the genomic region, which was initiated with the same primer. The prominent product, 63 bases long, is marked by an arrow. An asterisk marks a product corresponding in size to a previously published cDNA sequence (43).
Construction of chimeric genes and expression vectors.
To create chimeric Tmp-CAT constructs (see Fig. 4A, construct a), an ∼5-kb AccI fragment encompassing the Tmp promoter region, exon 1, and ∼3 kb of intron 1, including the CACGTG element, was cloned into the AccI site of the pCAT-Basic Vector (Promega E1041). A 1.5-kb BbrpI-SmaI fragment was excised from construct a, beginning at the CACGTG element and extending to the 3′ end of the original fragment, to create construct b. Construct c was created by inserting a point mutation in the CACGTG element in construct a. Vectors were restricted with BbrpI, which cuts in the unique CACGTG element. A terminal transferase reaction was performed on the linearized plasmid by using Taq polymerase and dATP+dTTP nucleotides, and the vectors were then religated and restricted with BbrpI. Bacteria were transformed with the total reaction mix, and the colonies were analyzed. For the construction of the Tmp expression vector, a BamHI-XhoI 850-bp cDNA fragment encompassing the full coding sequence of the mouse Tmp gene was cloned into the corresponding sites of the pCA1038 expression vector (71), which contains the phosphoglycerate-kinase-1 (PGK1) promoter and poly(A) signal. Plasmid PGK-neo (71) was used as a selection marker for stable transfections.
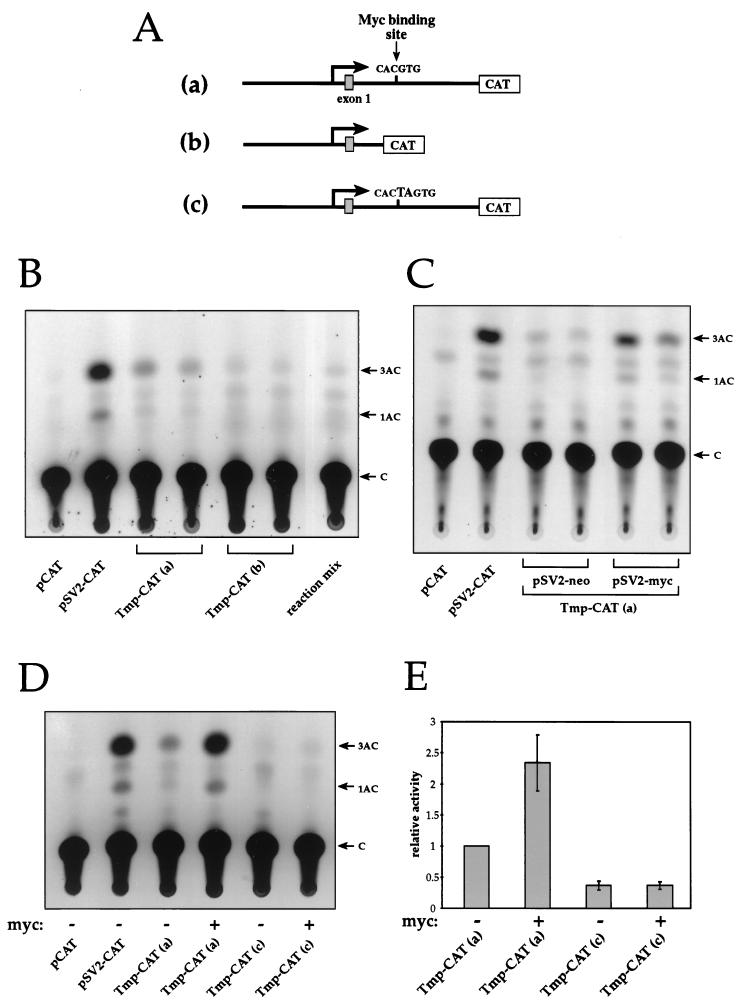
FIG. 4.
Activation of the Tmp promoter by c-Myc. (A) Schematic representation of Tmp-CAT constructs. Construct a contains a 5-kb fragment from the Tmp regulatory region cloned upstream of the CAT reporter gene. In construct b a 1.5-kb fragment containing the CACGTG element and the downstream region of intron 1 was excised from construct a. In construct c, a point mutation was introduced into the CACGTG element. (B) Activity of Tmp promoter. NIH 3T3 cells were transfected with Tmp-CAT construct a or b, and the CAT activity was studied. Activity was observed only upon transfection with construct a. C, free chloramphenicol; 1AC and 3AC, 1- and 3-acetylated forms of chloramphenicol, respectively. (C) c-Myc activates the Tmp promoter. Cells were cotransfected with Tmp-CAT construct a and with either the pSV2-myc expression vector or the pSV2-neo vector as a control. (D) The integrity of the CACGTG element is necessary for Tmp promoter activity and c-Myc activation. Cells were transfected with Tmp-CAT construct a or c, with or without the pSV2-myc vector. (E) Histogram bars represent the mean CAT activities from four to seven experiments ± the standard error. The results are presented relative to the activity level of Tmp-CAT construct a.
Transfection conditions and CAT assays.
For chloramphenicol acetyltransferase (CAT) assays, NIH 3T3 cells were transiently transfected with the described Tmp-CAT constructs. c-Myc activation was studied by cotransfection with the pSV2-myc expression vector containing the wild-type human c-myc. Transient transfections were performed by electroporating cells at 960 μF and 260 V, as described previously (53), with 20 μg of plasmid DNA. The pCAT-Basic vector was used as a negative control, and the pSV2-CAT vector was used as a positive control. The pSV2-neo vector was used to control for promoter competition. Cells were harvested 48 h after transfection, and a CAT activity assay was performed according to standard procedures (33). To control for transfection efficiency, plasmid DNA was extracted from transfected cells by lysis in Tris-EDTA buffer containing 0.6% sodium dodecyl sulfate (SDS). Genomic DNA was precipitated in the presence of 1 M NaCl. Supernatants were treated with proteinase K, followed by phenol extraction and ethanol precipitation. Samples were dot blotted and hybridized with a CAT probe. Stable transfections were performed by calcium phosphate-DNA coprecipitation in HEPES buffer (pH 6.95) (17). Rat1 cells were cotransfected with 40 μg of the PGK-Tmp plasmid and 1 μg of the PGK-neo plasmid as a selection marker. Resistant colonies were selected in medium containing 400 μg of G418 per ml.
Electrophoretic mobility shift assays.
A double-stranded oligonucleotide of the sequence 5′-ACAAACCCACGTGCTAAATGC-3′, representing the Tmp CACGTG sequence and flanking nucleotides, was radiolabeled by a fill-in reaction with [32P]dCTP. A mutant form of the oligonucleotide containing a CTCGAG element with identical flanking nucleotides served as a control. The labeled oligonucleotide was incubated with purified recombinant maltose-binding protein (MBP)-Myc and MBP-Max proteins, which were kindly provided by Thanos D. Halazonetis (22). These contain either the bHLH-LZ region of human c-Myc (amino acids 346 to 439) or the bHLH-LZ region of Max (amino acids 7 to 114) fused to MBP. Concentrations of recombinant proteins were determined by size separation by SDS-polyacrylamide gel electrophoresis (PAGE) and comparison to known bovine serum albumin (BSA) reference concentrations. Protein (10 to 50 ng) was used in the reactions as indicated. Incubation was performed in 100 mM HEPES buffer (pH 7.9), 10 mM MgCl2, 5 mM dithiothreitol, 2.5 mM EGTA, 40 mM KCl, and 1.5 mg of BSA per ml, with 2 μg of poly(dI-dC), in a final volume of 25 μl for 30 min at room temperature. For the competition assays, unlabeled oligonucleotide, either wild type or mutant, was added in a 25- or 100-fold molar excess to the mixture. The reaction mixture was run on a 6% polyacrylamide gel in 0.5× Tris-borate-EDTA buffer, dried, and subjected to autoradiography for 16 h.
Tumor growth assays.
Four clones of Rat1 cells expressing the PGK-Tmp expression vector and four clones expressing only the PGK-neo selection marker were used for injection. Cells (∼107), suspended in 200 to 400 μl of phosphate-buffered saline, were injected subcutaneously into athymic nude (nu/nu) mice as described previously (20). Animals were monitored at least twice a week for the appearance and measurement of tumors.
RESULTS
Tmp is highly expressed in tumors induced by c-myc in transgenic mice.
Tmp was initially isolated from a brain tumor that developed in a mouse that was transgenic for c-myc (5, 7). To examine whether high Tmp expression is a general phenomenon in c-myc-induced tumors, we tested Tmp expression in several cell lines derived from adenocarcinomas of the mammary gland, which developed in mice transgenic for c-myc (63), c-neu (50), or v-Ha-ras (14). These lines present an excellent experimental system because the development of the tumor can be ascribed primarily to the expression of a specific oncogene. Impressively, the highest expression levels of Tmp were observed in c-myc-induced mammary tumors; these levels were approximately 10- to 20-fold higher than in normal mammary gland and epithelium (Fig. 1A). In c-neu-induced mammary tumors, Tmp was also expressed at higher levels than in the normal tissues but to a much lower extent (Fig. 1A). The level of Tmp expression in v-Ha-ras-induced mammary tumors was equal to or lower than that in the normal tissues (Fig. 1A).
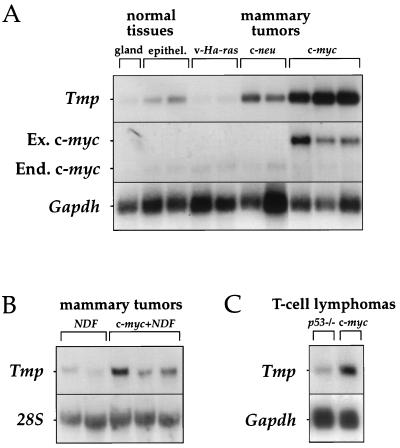
FIG. 1.
Tmp is specifically expressed in c-myc-induced tumors. (A) Northern blot hybridization was performed on poly(A)+ RNA from normal mammary gland and epithelium and from cell lines derived from adenocarcinomas of the mammary gland, which developed in mice transgenic for v-Ha-ras, c-neu, or c-myc (26). Each lane contains RNA from a different cell line. Ex. c-myc, exogenous transcript from transgene; End. c-myc, endogenous transcript. (B) Northern blot hybridization was performed on total RNA from cell lines of mammary tumors which developed in mice transgenic for NDF or for both NDF and c-myc (39). (C) Northern blot hybridization was performed on poly(A)+ RNA from cell lines derived from T-cell lymphomas, which developed in p53−/− mice and in c-myc transgenic mice (25). Hybridizations were performed with Tmp, Gapdh, and 28S rRNA probes, as indicated.
We also examined the expression of Tmp in cell lines from mammary tumors that developed in mice transgenic for the Neu differentiation factor gene (NDF) or for both NDF and c-myc (39). Tmp expression was detected in all tumors but was significantly higher in tumors from mice transgenic for both genes (Fig. 1B).
The correlation between c-myc and Tmp expression was also observed in T-cell lymphoma cells. We compared Tmp expression in cell lines derived from lymphomas which developed in c-myc transgenic mice and in mice homozygous for a null mutation in the p53 gene (25). Once again, a higher level of Tmp expression was observed in c-myc-induced lymphomas than in p53−/− lymphomas (Fig. 1C). Thus, Tmp is highly expressed in at least three different types of tumors induced by c-Myc in the mouse, suggesting that it is activated in the c-Myc-induced tumorigenic pathway.
Tmp expression is induced in serum-stimulated cells.
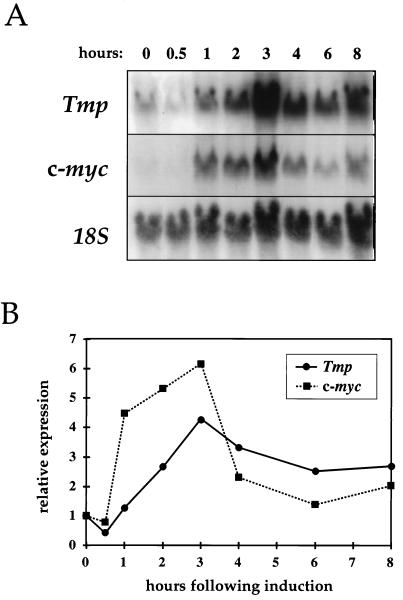
We have previously reported that Tmp expression is high in dividing cells but much lower in quiescent cells, a finding correlating c-myc expression (5). We studied the time course of Tmp activation in Rat1 cells after growth stimulation by serum. Activation of Tmp transcription was seen as early as 2 h after serum stimulation, and expression peaked after 3 h (Fig. 2). This time course closely matches that of c-myc in these cells, showing a rise in c-myc expression that is observed as early as 1 h after stimulation and lasting 2 to 3 h (Fig. 2). Tmp expression thus follows the characteristic activation pattern of c-Myc during the G0-to-G1 transition. It is noteworthy that Tavtigian et al. (67) isolated a transcript designated I-8-9, later identified by us as Tmp, in a screen for mid-G1 serum response genes (67).
FIG. 2.
Tmp activation in serum-stimulated cells. (A) Rat1 fibroblasts were grown in low-serum conditions for 3 days and then transferred to a high-serum medium. Total RNA was extracted at indicated time points. Northern blot hybridizations were performed with Tmp, c-myc, and 18S rRNA probes. (B) Quantitation of Tmp and c-myc activation levels, calculated by comparison to 18S rRNA levels and normalized relative to the expression level at the time of serum stimulation.
Isolation of the genomic Tmp gene and analysis of the promoter region.
To isolate Tmp genomic clones and analyze the Tmp promoter region, we screened a 129SVJ mouse genomic DNA library with cDNA probes encompassing the entire Tmp transcript. Four genomic clones were isolated and analyzed. We determined the exon-intron boundaries of the Tmp gene by using sequencing and PCR performed with primers from within the cDNA sequence. Our analysis indicated that Tmp is composed of five exons (Fig. 3A). Exons 2 to 5 contain the coding region and a 3′-untranslated region of 2.1 kb, as has been previously reported by Lobsiger et al. (43). One of the isolated genomic clones contained the Tmp promoter region and exon 1. Exon 1 contains most of the 5′-untranslated region and is separated from exon 2 by an intron of at least 10 kb. The genomic structure of Tmp is similar to that of its homolog Pmp22. Pmp22 is also composed of 5 exons, of similar sizes, which encode corresponding regions of the homologous protein (43, 66).
We determined the sequence of approximately 1.6 kb of the Tmp genomic region, including the promoter region, exon 1, and part of intron 1 (Fig. 3B). To determine the start site of transcription, we performed a primer extension assay, with a primer from the 3′ region of exon 1, as determined from the cDNA sequence and from genomic analysis. A prominent band, 63 bases long, was detected (Fig. 3C, arrow), representing a putative start site at an adenine residue marked 1 (Fig. 3B). A putative TATA box (TATAA) is located at position −30 relative to this site. However, as shown, there are additional minor bands in this region (Fig. 3C). Interestingly, a cDNA sequence extending to position −60 has been previously published (43). We detected a minor band corresponding to exactly this point (Fig. 3C, asterisk), suggesting that this is an alternative start site. This would be supported by the finding of an additional putative TATA box (TTTAAAA) 31 bp upstream of this point. The sequences of both start sites conform to the start site consensus: an adenine residue preceded by a cytosine and surrounded by pyrimidine residues (19).
Two putative AP-1 binding elements (41) are located at positions −245 to −237 and −108 to −100. In addition, a poly(dT) tract, 29 residues long, is located at position −147 to −119. This element is common in yeast promoters (36) and has recently also been detected in mammalian promoters and shown to be a functional transcription promoting element (72).
A putative c-Myc binding element, namely, a canonical CACGTG E-box, was found in intron 1, approximately 1.1 kb downstream of the start site of transcription (Fig. 3B). This position is analogous to the position of functional c-Myc binding sites identified to date (9, 34). In known c-Myc targets, the binding site is typically located downstream of the start site of transcription, most commonly either in the 5′-untranslated region or in intron 1. The location of the CACGTG element in Tmp thus suggested that it may be a functional c-Myc binding element and that Tmp transcription is directly regulated by c-Myc.
c-Myc activates the Tmp promoter.
To test the activity of the Tmp promoter, we prepared several chimeric constructs in which the Tmp regulatory region was placed upstream of the CAT reporter gene (Fig. 4A). A Tmp fragment containing the promoter, exon 1, and a part of intron 1 including the c-Myc binding element, was cloned upstream to the CAT gene (Fig. 4A, construct a). This construct induced a low level of CAT activity when transfected into NIH 3T3 cells (Fig. 4B). Construct b was created by excising a fragment containing the CACGTG element from construct a. No CAT activity was observed upon transfection of this construct (Fig. 4B), indicating that an element, or elements, contained in the excised fragment, among them possibly the CACGTG element, are necessary for Tmp promoter activity.
We wished to test whether c-Myc can transcriptionally activate the Tmp promoter and whether the CACGTG element plays a role in this activation. Cotransfection of the pSV2-myc expression vector with Tmp-CAT construct a into NIH 3T3 cells resulted in an increase in CAT activity (Fig. 4C and D). An average 2.3-fold increase in activity was observed in seven experiments (Fig. 4E). We introduced a point mutation into the element in the original, active, construct a, changing its sequence from CACGTG to CACTAGTG, thus creating construct c. This mutation resulted in a significant reduction of CAT activity induced by the Tmp promoter (Fig. 4D and E), indicating that the CACGTG element is a functional transcription-promoting element that is necessary for Tmp promoter activity. When the pSV2-myc vector was cotransfected with construct c, no increase in CAT activity was detected (Fig. 4D and E), suggesting that c-Myc activation is mediated through this element.
These results suggest that the CACGTG sequence in intron 1 of Tmp is a functional transcription-promoting element and that c-Myc can activate the Tmp promoter through this element.
The c-Myc–Max complex can bind the CACGTG element found in Tmp.
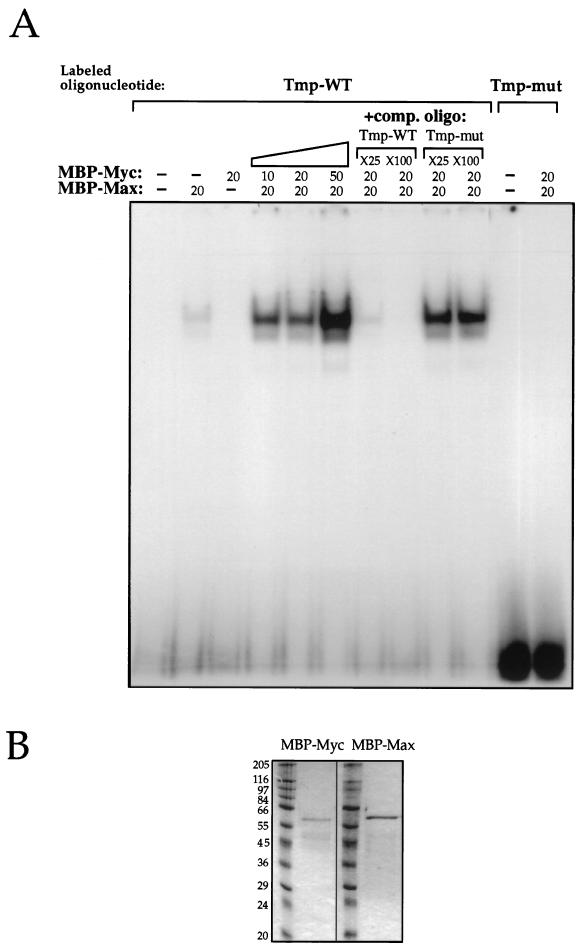
To test whether the c-Myc–Max complex can bind the CACGTG element found in intron 1 of Tmp, we performed an electrophoretic mobility shift assay. A labeled double-stranded oligonucleotide containing the CACGTG element and flanking nucleotides from the Tmp intron 1 sequence was incubated with purified recombinant truncated c-Myc and Max proteins. These proteins contain the bHLH-LZ region of either c-Myc or Max, fused to MBP. Incubation with both proteins resulted in a strong shifted band of the labeled oligonucleotide (Fig. 5A). The intensity of the shifted band was increased with increasing c-Myc/Max ratios (0.5:1, 1:1, and 2.5:1) (Fig. 5A). Incubation with c-Myc alone did not produce a shifted band, while incubation with Max alone produced a weak shifted band (Fig. 5A). Since the MBP-Myc and MBP-Max proteins used in this assay are of similar molecular sizes (Fig. 5B), Max-Max and c-Myc–Max complexes cannot be distinguished by size. However, a 1:1 ratio of c-Myc–Max proteins produced a much stronger shifted band than that observed with Max alone (7.6 [±0.7]-fold increase, average of three experiments), a finding consistent with the known increased DNA binding activity of c-Myc–Max complexes (3). This indicates that both proteins are probably present in the complexes. Incubation of c-Myc and Max proteins with the labeled oligonucleotide in the presence of an unlabeled competitor oligonucleotide resulted in a significant reduction of the labeled shift, while no reduction was observed upon incubation with an unlabeled competitor oligonucleotide carrying a mutated Myc binding element, CTCGAG (Fig. 5A). Incubation of c-Myc and Max proteins with a labeled mutated oligonucleotide did not result in a shifted band (Fig. 5A), indicating the specificity of the binding.
FIG. 5.
The c-Myc–Max complex binds the CACGTG element in Tmp. (A) A labeled, 21-bp oligonucleotide containing the CACGTG element and flanking nucleotides from intron 1 (Tmp-WT) or an identical oligonucleotide carrying a mutated element, CTCGAG (Tmp-mut), was incubated with purified recombinant MBP-Myc and MBP-Max proteins containing the bHLH-LZ region of either c-Myc or Max fused to MBP. Binding was tested by electrophoretic mobility shift assay of the labeled oligonucleotide. Where indicated, 20 ng of each protein was used, except under the diagonal bar, where increasing concentrations of MBP-Myc protein were used as follows: 10, 20, and 50 ng. Unlabeled competitor oligonucleotide, either Tmp-WT or Tmp-mut, was used in 25× and 100× molar excess, as indicated, of labeled Tmp-WT. (B) MBP-Myc and MBP-Max proteins as they appear on SDS-PAGE gel. The numbers indicate the molecular masses of ladder proteins in kilodaltons.
Tmp is activated by the MycER fusion protein.
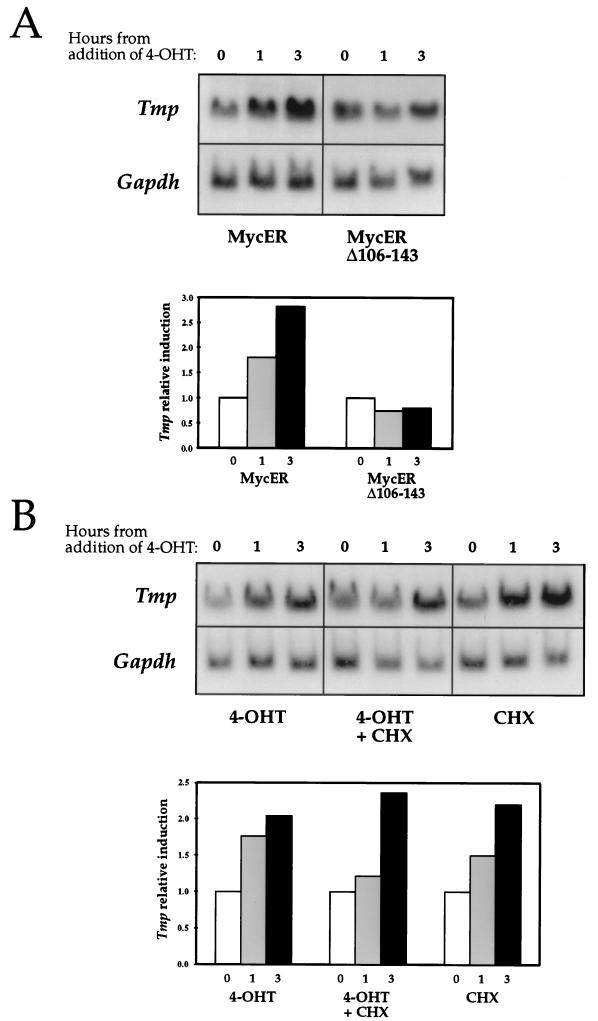
We utilized the MycER-inducible system to test the ability of c-Myc to upregulate Tmp expression. The MycER construct produces a fusion protein of c-Myc and the estrogen receptor ligand-binding domain (24). We used the MycER version of the construct, which carries a G252R mutation in the TAF-2 transcription-activating domain of the estrogen receptor (42). The transcriptional activity of the MycER product is specifically induced by 4-OHT. Rat1 cells expressing either the wild-type MycER or the transcriptionally inactive deletion mutant Δ106-143MycER (46) were cultured in low-serum conditions for 3 days, after which 4-OHT was added to the medium. A two- to threefold increase in Tmp expression was observed within 3 h of the induction of the MycER cells (Fig. 6A). Tmp expression in the Δ106-143MycER cells was unaffected by the addition of 4-OHT (Fig. 6A), indicating that a transcriptionally active c-Myc protein is required for Tmp activation. This mutant form of the c-Myc protein has also been characterized as deficient in transformation activity (64). The transcription of the endogenous Tmp gene is thus upregulated when c-Myc activity is induced, supporting the hypothesis that it is a direct c-Myc target. The time course and level of upregulation are similar to those found for other c-Myc targets which were tested in this system (31, 35, 70).
FIG. 6.
Tmp expression is induced by the MycER fusion protein. (A) Rat1 cells expressing either the wild-type MycER or deletion mutant Δ106-143MycER (42, 46) were serum starved for 3 days. 4-OHT was added to the medium, and total RNA was extracted at different time points. Northern blot hybridizations were performed with a Tmp probe and with a probe for the glyceraldehyde-3-phosphate dehydrogenase (Gapdh) gene. The histogram represents Tmp expression relative to Gapdh expression, as determined by densitometry. Results were normalized relative to the expression level at the time of the addition of 4-OHT. (B) The experiment in panel A was repeated in the presence of cycloheximide (CHX) and with cycloheximide alone. The histogram represents Tmp expression relative to Gapdh expression.
Tmp activation was also observed when MycER cells were treated with 4-OHT in the presence of the protein synthesis inhibitor cycloheximide (Fig. 6B). This finding is consistent with the suggestion that Tmp is activated directly by c-Myc in a way that is independent of protein synthesis. Cycloheximide alone also caused an elevation of Tmp mRNA levels (Fig. 6B), as was observed for several other c-Myc targets in which this issue was addressed (35, 70). Yet, the fact that Tmp induction by MycER is observed in the presence of cycloheximide and that it is higher than the induction observed with cycloheximide or 4-OHT alone (Fig. 6B) supports the notion that Tmp is a direct target for c-Myc activity.
Cells overexpressing the Tmp gene are tumorigenic in nude mice.
To examine the effects of Tmp overexpression, a vector containing the mouse Tmp gene under the control of the PGK1 promoter was transfected into Rat1 fibroblasts. These cells express the endogenous Tmp gene when they are proliferating, and expression is downregulated during growth arrest (Fig. 2A and Fig. 7A). Four clones stably expressing the exogenous transfected Tmp gene were isolated (Fig. 7A). The level of expression in these cells is ca. two- to threefold higher than in control cells (Fig. 7A).
FIG. 7.
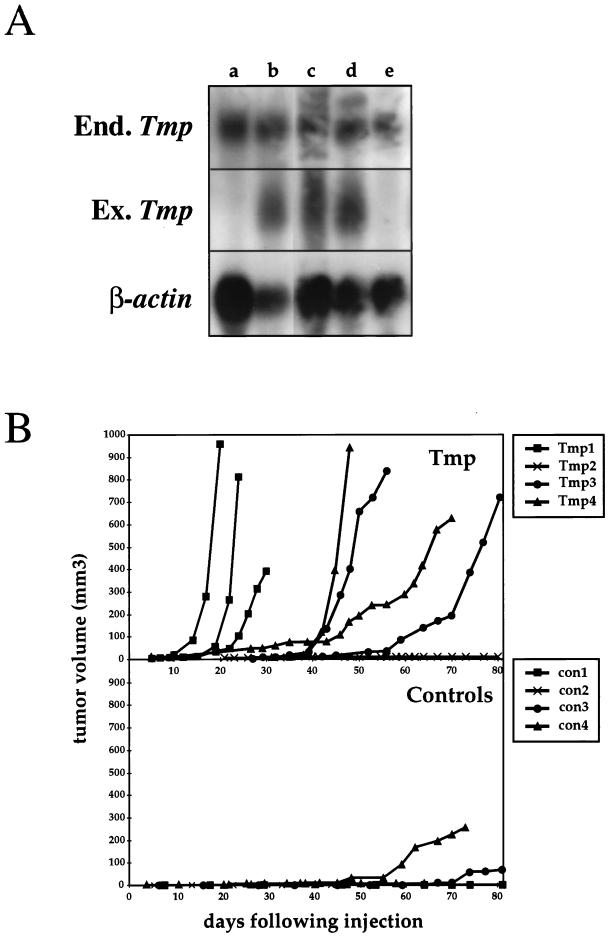
Tmp-overexpressing cells are tumorigenic in nude mice. (A) Rat1 fibroblasts were cotransfected with the PGK-Tmp and the PGK-neo constructs. Total RNA was extracted from G418 resistant colonies, and Northern blot hybridization was performed with a Tmp probe, thereby detecting both endogenous (End.) and exogenous (Ex.) transcripts (2.7 and 1.2 kb, respectively), and with a β-actin probe. Examples of three positive clones (lanes b, c, and d) and two negative clones (lanes a and e) are shown. (B) Cells from four PGK-Tmp-expressing clones (Tmp1 to Tmp4) and four clones expressing only the PGK-neo selection marker (con1 to con4) were injected subcutaneously into athymic nude mice. The graphs display the tumor volumes at various times after injection for PGK-Tmp-expressing cells (top panel) and for control cells (bottom panel).
To test whether Tmp overexpression increases the degree of tumorigenicity of these cells, cells from four different PGK-Tmp clones and from four different control clones expressing only the neo resistance selection marker were injected subcutaneously into nude mice. Cells from three of the four PGK-Tmp clones resulted in rapidly developing subcutaneous tumors in all injections (Fig. 7B). In total, tumors developed as a result of 7 of 9 injections of these cells. Of the control cells, tumors developed in only two of seven injections, and these appeared relatively late after injection (Fig. 7B). These results thus suggest that overexpression of Tmp can enhance the tumorigenicity of Rat1 cells and that Tmp may possess a function which can contribute to the transformation process.
DISCUSSION
To understand the mechanisms by which c-Myc induces cells to proliferate and by which it can lead to malignant transformation, it is essential to identify the genes regulated by c-Myc. In this study, we establish, based on the following evidence, that the Tmp gene is directly activated by c-Myc. (i) Tmp carries a putative c-Myc binding element located in the first intron, a typical location for this element in c-Myc targets. (ii) The Tmp promoter can activate a reporter gene, depending on the integrity of the CACGTG element, indicating that this is a functional element. (iii) c-Myc can activate the Tmp promoter when cotransfected into cells, and this activation is mediated by the c-Myc binding element. (iv) The c-Myc–Max protein complex can bind the CACGTG element. (v) The endogenous Tmp gene is activated when MycER transcriptional activity is induced, and this occurs also in the absence of protein synthesis.
Furthermore, the Tmp expression pattern in several cellular systems is closely correlated with that of c-myc. Tmp is expressed in proliferating fibroblasts but is downregulated when these cells are quiescent. When cells are serum stimulated to progress from G0 to G1, Tmp is strongly activated, as shown by a time course that closely matches that of c-myc. Tmp is expressed in nondifferentiated, highly proliferative, ES cells, but when these cells differentiate into embryoid bodies Tmp expression is greatly reduced, again mimicking the behavior of c-myc (5). Most impressively, we show that Tmp is highly expressed in c-myc-induced tumors. This is demonstrated for mammary tumors and T-cell lymphomas that develop in c-myc transgenic mice (Fig. 1) and also for c-myc-induced brain tumors, from which Tmp was initially isolated (5). The levels of Tmp expression in these tumors greatly exceed its expression levels in the corresponding normal tissues. Such a dramatic degree of Tmp activation does not appear in mammary tumors induced by other oncogenes.
Overexpression of Tmp in Rat1 cells results in increased tumorigenicity of these cells when injected into nude mice, suggesting that Tmp function is related to cell transformation. Combined with the high level of expression of Tmp in c-myc-induced tumors, these findings suggest that Tmp may play a role in the c-Myc-induced transformation pathway. We were unable to detect significant phenotypes of the Tmp-overexpressing cells regarding their proliferation rate and cell-cycle-stage distribution under high-serum and low-serum conditions. We also did not detect an increase in the anchorage-independent growth ability of the PGK-Tmp cells. It is possible that the activity of Tmp does not affect these characteristics in this experimental system. An additional explanation, however, could be that Rat1 cells express the endogenous Tmp and that this expression was elevated only ca. two- to threefold in the transfected clones. While possibly reflecting a more physiologically relevant system for studying Tmp function, this expression level might not have been sufficient to induce a strong effect regarding these characteristics. It is interesting to note that Keath et al. (37) reported similar results for c-myc-overexpressing rodent fibroblasts. The most prominent phenotype of these cells was their ability to form tumors in nude mice, while no dramatic changes were observed in the proliferation characteristics in high- or low-serum conditions and in their anchorage-independent growth ability (37). Various phenotypes were reported, however, by other groups, probably reflecting differences in the levels of c-myc overexpression and in transfected cell lines (38, 55, 61).
Tmp belongs to a novel family of genes that encode membrane glycoproteins with four transmembrane domains (5, 68, 69). All four family members display widespread expression in adult and embryo tissues, in rodents and in humans, indicating that these proteins probably perform a basic function in cell life (5, 68, 69). Several possible roles have been suggested for PMP22, the most studied member of this family (65). The PMP22 protein carries the L2/HNK1 epitope, suggesting a role in cell adhesion (62). A role for PMP22 in cell cycle modulation has also been suggested, based on its induction in growth-arrested cells (44) and on the phenotypes of PMP22 overexpression, i.e., a delay in cell cycle progression and an induction of apoptosis (29, 73, 74). These results, combined with ours, suggest that Tmp and Pmp22 not only are inversely expressed during proliferation and growth arrest but also modulate cell proliferation in opposite directions.
Uncovering the c-Myc target genes and their cellular functions should reveal which specific cellular processes are controlled by c-Myc and how their combined action leads to the end results of cell proliferation, transformation, or apoptosis. In recent years a number of potential c-Myc targets have been presented, and attempts to link their functions with the c-Myc-induced pathways have been made. Some of these genes, such as p53 and cdc25A, encode nuclear proteins that are directly involved in cell cycle control (31, 56). Others, such as Eca39/Bcat1, ODC, cad, and LDH-A, encode metabolic enzymes whose function, in some cases, can be linked with cell growth and division (4, 7, 23, 49, 52, 60). Tmp is the first isolated c-Myc target that encodes a membrane protein, thus extending the cellular range of c-Myc-controlled effects and adding a possible link to cell contact processes.
The c-Myc target genes can also be tested in their ability to induce cellular effects parallel to those induced by c-Myc itself. It has been demonstrated, for example, that the Cdc25A, ODC, and eIF-4E genes, when overexpressed, can induce cellular transformation (1, 2, 32, 40). The tumorigenic effect of Tmp overexpression places this gene among the c-Myc targets which can mimic some of the transformation abilities of c-Myc.
ACKNOWLEDGMENTS
This research was supported by grant 3811a from The Council for Tobacco Research, by grant 93-00017 from The U.-S.-Israel Binational Science Foundation, Jerusalem, Israel, and by a grant from The Israel Science Foundation, funded by The Israel Academy of Sciences and Humanities.
We thank the following investigators: Linda Z. Penn for the Rat1-MycER cells, Ari Elson and Ian Krane for the RNA blots of tumor cell lines from transgenic mice, Thanos Halazonetis for the MBP-Myc and MBP-Max proteins, and Chaim Kahana for advice with the experiments. We thank Ayellet Falcovitz and Irit Marbach for assistance with the experiments. We thank Shoshana Klein, Amir Eden, and Yuval Dor for critical reading of the manuscript.
REFERENCES
- 1.Auvinen M, Laine A, Paasinen-Sohns A, Kangas A, Kangas L, Saksela O, Andersson L C, Holtta E. Human ornithine decarboxylase-overproducing NIH3T3 cells induce rapidly growing, highly vascularized tumors in nude mice. Cancer Res. 1997;57:3016–3025. [PubMed] [Google Scholar]
- 2.Auvinen M, Paasinen A, Andersson L C, Holtta E. Ornithine decarboxylase activity is critical for cell transformation. Nature. 1992;360:355–358. doi: 10.1038/360355a0. [DOI] [PubMed] [Google Scholar]
- 3.Ayer D E, Kretzner L, Eisenman R N. Mad: a heterodimeric partner for Max that antagonizes Myc transcriptional activity. Cell. 1993;72:211–222. doi: 10.1016/0092-8674(93)90661-9. [DOI] [PubMed] [Google Scholar]
- 4.Bello-Fernandez C, Packham G, Cleveland J L. The ornithine decarboxylase gene is a transcriptional target of c-Myc. Proc Natl Acad Sci USA. 1993;90:7804–7808. doi: 10.1073/pnas.90.16.7804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ben-Porath I, Benvenisty N. Characterization of a tumor-associated gene, a member of a novel family of genes encoding membrane glycoproteins. Gene. 1996;183:69–75. doi: 10.1016/s0378-1119(96)00475-1. [DOI] [PubMed] [Google Scholar]
- 6.Ben-Porath I, Kozak C A, Benvenisty N. Chromosomal mapping of Tmp (Emp1), Xmp (Emp2), and Ymp (Emp3), genes encoding membrane proteins related to Pmp22. Genomics. 1998;49:443–447. doi: 10.1006/geno.1998.5238. [DOI] [PubMed] [Google Scholar]
- 7.Benvenisty N, Leder A, Kuo A, Leder P. An embryonically expressed gene is a target for c-Myc regulation via the c-Myc-binding sequence. Genes Dev. 1992;6:2513–2523. doi: 10.1101/gad.6.12b.2513. [DOI] [PubMed] [Google Scholar]
- 8.Benvenisty N, Ornitz D M, Bennett G L, Sahagan B G, Kuo A, Cardiff R D, Leder P. Brain tumours and lymphomas in transgenic mice that carry HTLV-I LTR/c-myc and Ig/tax genes. Oncogene. 1992;7:2399–2405. [PubMed] [Google Scholar]
- 9.Ben-Yosef T, Yanuka O, Benvenisty N. ECA39 is regulated by c-Myc in human and by a Jun/Fos homolog, Gcn4, in yeast. Oncogene. 1996;13:1859–1866. [PubMed] [Google Scholar]
- 10.Blackwell T, Huang J, Ma A, Kretzner L, Alt F, Eisenman R, Weintraub H. Binding of Myc proteins to canonical and noncanonical DNA sequences. Mol Cell Biol. 1993;13:5216–5224. doi: 10.1128/mcb.13.9.5216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blackwell T K, Kretzner L, Blackwood E M, Eisenman R N, Weintraub H. Sequence-specific DNA binding by the c-Myc protein. Science. 1990;250:1149–1151. doi: 10.1126/science.2251503. [DOI] [PubMed] [Google Scholar]
- 12.Blackwood E M, Eisenman R N. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science. 1991;251:1211–1217. doi: 10.1126/science.2006410. [DOI] [PubMed] [Google Scholar]
- 13.Bolin L, McNeil T, Lucian L, DeVaux B, Franz-Bacon K, Gorman D, Zurawski S, Murray R, McClanahan T. HNMP-1: a novel hematopoietic and neural membrane protein differentially regulated in neural development and injury. J Neurosci. 1997;17:5493–5502. doi: 10.1523/JNEUROSCI.17-14-05493.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cardiff R, Leder A, Kuo A, Pattengale P, Leder P. Multiple tumor types appear in a transgenic mouse with the ras oncogene. Am J Pathol. 1993;142:1199–1207. [PMC free article] [PubMed] [Google Scholar]
- 15.Cavalieri F, Goldfarb M. Growth factor-deprived BALB/c 3T3 murine fibroblasts can enter the S phase after induction of c-myc gene expression. Mol Cell Biol. 1987;7:3554–3560. doi: 10.1128/mcb.7.10.3554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chance P F, Fischbeck K H. Molecular genetics of Charcot-Marie-Tooth disease and related neuropathies. Hum Mol Genet. 1994;3:1503–1507. doi: 10.1093/hmg/3.suppl_1.1503. [DOI] [PubMed] [Google Scholar]
- 17.Chen C, Okayama H. High-efficiency transformation of mammalian cells by plasmid DNA. Mol Cell Biol. 1987;7:2745–2752. doi: 10.1128/mcb.7.8.2745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162:156–159. doi: 10.1006/abio.1987.9999. [DOI] [PubMed] [Google Scholar]
- 19.Corden J, Wasylyk B, Buchwalder A, Sassone C P, Kedinger C, Chambon P. Promoter sequences of eukaryotic protein-coding genes. Science. 1980;209:1406–1414. doi: 10.1126/science.6251548. [DOI] [PubMed] [Google Scholar]
- 20.Cox A, Der C. Biological assays for cellular transformation. Methods Enzymol. 1994;238:277–294. doi: 10.1016/0076-6879(94)38026-0. [DOI] [PubMed] [Google Scholar]
- 21.Dang C. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol. 1999;19:1–11. doi: 10.1128/mcb.19.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Davis L, Halazonetis T. Both the helix-loop-helix and the leucine zipper motifs of c-Myc contribute to its dimerization specificity with Max. Oncogene. 1993;8:125–132. [PubMed] [Google Scholar]
- 23.Eden A, Simchen G, Benvenisty N. Two yeast homologs of ECA39, a target for c-Myc regulation, code for cytosolic and mitochondrial branched-chain amino acid aminotransferases. J Biol Chem. 1996;271:20242–20245. doi: 10.1074/jbc.271.34.20242. [DOI] [PubMed] [Google Scholar]
- 24.Eilers M, Picard D, Yamamoto K R, Bishop J M. Chimaeras of myc oncoprotein and steroid receptors cause hormone-dependent transformation of cells. Nature. 1989;340:66–68. doi: 10.1038/340066a0. [DOI] [PubMed] [Google Scholar]
- 25.Elson A, Deng C, Campos-Torres J, Donehower L, Leder P. The MMTV/c-myc transgene and p53 null alleles collaborate to induce T-cell lymphomas, but not mammary carcinomas in transgenic mice. Oncogene. 1995;11:181–190. [PubMed] [Google Scholar]
- 26.Elson A, Leder P. Protein-tyrosine phosphatase epsilon. An isoform specifically expressed in mouse mammary tumors initiated by v-Ha-ras or neu. J Biol Chem. 1995;270:26116–26122. doi: 10.1074/jbc.270.44.26116. [DOI] [PubMed] [Google Scholar]
- 27.Evan G, Harrington E, Fanidi A, Land H, Amati B, Bennett M. Integrated control of cell proliferation and cell death by the c-myc oncogene. Philos Trans R Soc Lond B Biol Sci. 1994;345:269–275. doi: 10.1098/rstb.1994.0105. [DOI] [PubMed] [Google Scholar]
- 28.Evan G I, Wyllie A H, Gilbert C S, Littlewood T D, Land H, Brooks M, Waters C M, Penn L Z, Hancock D C. Induction of apoptosis in fibroblasts by c-myc protein. Cell. 1992;69:119–128. doi: 10.1016/0092-8674(92)90123-t. [DOI] [PubMed] [Google Scholar]
- 29.Fabbretti E, Edomi P, Brancolini C, Schneider C. Apoptotic phenotype induced by overexpression of wild-type gas3/PMP22: its relation to the demyelinating peripheral neuropathy CMT1A. Genes Dev. 1995;9:1846–1856. doi: 10.1101/gad.9.15.1846. [DOI] [PubMed] [Google Scholar]
- 30.Facchini L M, Penn L Z. The molecular role of Myc in growth and transformation: recent discoveries lead to new insights. FASEB J. 1998;12:633–651. [PubMed] [Google Scholar]
- 31.Galaktionov K, Chen X, Beach D. Cdc25 cell-cycle phosphatase as a target of c-myc. Nature. 1996;382:511–517. doi: 10.1038/382511a0. [DOI] [PubMed] [Google Scholar]
- 32.Galaktionov K, Lee A K, Eckstein J, Draetta G, Meckler J, Loda M, Beach D. CDC25 phosphatases as potential human oncogenes. Science. 1995;269:1575–1577. doi: 10.1126/science.7667636. [DOI] [PubMed] [Google Scholar]
- 33.Gorman C M, Moffat L F, Howard B H. Recombinant genomes which express chloramphenicol acetyltransferase in mammalian cells. Mol Cell Biol. 1982;2:1044–1051. doi: 10.1128/mcb.2.9.1044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Grandori C, Eisenman R. Myc target genes. Trends Biochem Sci. 1997;22:177–181. doi: 10.1016/s0968-0004(97)01025-6. [DOI] [PubMed] [Google Scholar]
- 35.Grandori C, Mac J, Siebelt F, Ayer D, Eisenman R. Myc-Max heterodimers activate a DEAD box gene and interact with multiple E box-related sites in vivo. EMBO J. 1996;15:4344–4357. [PMC free article] [PubMed] [Google Scholar]
- 36.Iyer V, Struhl K. Poly(dA:dT), a ubiquitous promoter element that stimulates transcription via its intrinsic DNA structure. EMBO J. 1995;14:2570–2579. doi: 10.1002/j.1460-2075.1995.tb07255.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Keath E J, Caimi P G, Cole M D. Fibroblast lines expressing activated c-myc oncogenes are tumorigenic in nude mice and syngeneic animals. Cell. 1984;39:339–348. doi: 10.1016/0092-8674(84)90012-6. [DOI] [PubMed] [Google Scholar]
- 38.Kelekar A, Cole M. Tumorigenicity of fibroblast lines expressing the adenovirus E1a, cellular p53, or normal c-myc genes. Mol Cell Biol. 1986;6:7–14. doi: 10.1128/mcb.6.1.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Krane I, Leder P. NDF/heregulin induces persistence of terminal end buds and adenocarcinomas in the mammary glands of transgenic mice. Oncogene. 1996;12:1781–1788. [PubMed] [Google Scholar]
- 40.Lazaris-Karatzas A, Montine K, Sonenberg N. Malignant transformation by a eukaryotic initiation factor subunit that binds to mRNA 5′ cap. Nature. 1990;345:544–547. doi: 10.1038/345544a0. [DOI] [PubMed] [Google Scholar]
- 41.Lee W, Haslinger A, Karin M, Tjian R. Activation of transcription by two factors that bind promoter and enhancer sequences of the human metallothionein gene and SV40. Nature. 1987;325:368–372. doi: 10.1038/325368a0. [DOI] [PubMed] [Google Scholar]
- 42.Littlewood T, Hancock D, Danielian P, Parker M, Evan G. A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 1995;23:1686–1690. doi: 10.1093/nar/23.10.1686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lobsiger C, Magyar J, Taylor V, Wulf P, Welcher A, Suter U Amgen EST Program. Identification and characterization of a cDNA and the structural gene encoding the mouse epithelial membrane protein-1. Genomics. 1996;36:379–387. doi: 10.1006/geno.1996.0482. [DOI] [PubMed] [Google Scholar]
- 44.Manfioletti G, Ruaro M E, Del Sal G, Philipson L, Schneider C. A growth arrest-specific (gas) gene codes for a membrane protein. Mol Cell Biol. 1990;10:2924–2930. doi: 10.1128/mcb.10.6.2924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Marcu K B, Bossone S A, Patel A J. myc function and regulation. Annu Rev Biochem. 1992;61:809–860. doi: 10.1146/annurev.bi.61.070192.004113. [DOI] [PubMed] [Google Scholar]
- 46.Marhin W, Chen S, Facchini L, Fornace J, Penn L. Myc represses the growth arrest gene gadd45. Oncogene. 1997;14:2825–2834. doi: 10.1038/sj.onc.1201138. [DOI] [PubMed] [Google Scholar]
- 47.Marvin K W, Fujimoto W, Jetten A M. Identification and characterization of a novel squamous cell-associated gene related to PMP22. J Biol Chem. 1995;270:28910–28916. doi: 10.1074/jbc.270.48.28910. [DOI] [PubMed] [Google Scholar]
- 48.Meichle A, Philipp A, Eilers M. The functions of Myc proteins. Biochim Biophys Acta. 1992;1114:129–146. doi: 10.1016/0304-419x(92)90011-m. [DOI] [PubMed] [Google Scholar]
- 49.Miltenberger R, Sukow K, Farnham P. An E-box-mediated increase in cad transcription at the G1/S-phase boundary is suppressed by inhibitory c-Myc mutants. Mol Cell Biol. 1995;15:2527–2535. doi: 10.1128/mcb.15.5.2527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Muller W J, Sinn E, Pattengale P K, Wallace R, Leder P. Single-step induction of mammary adenocarcinoma in transgenic mice bearing the activated c-neu oncogene. Cell. 1988;54:105–115. doi: 10.1016/0092-8674(88)90184-5. [DOI] [PubMed] [Google Scholar]
- 51.Patel P I, Lupski J R. Charcot-Marie-Tooth disease: a new paradigm for the mechanism of inherited disease. Trends Genet. 1994;10:128–133. doi: 10.1016/0168-9525(94)90214-3. [DOI] [PubMed] [Google Scholar]
- 52.Pena A, Reddy C D, Wu S, Hickok N J, Reddy E P, Yumet G, Soprano D R, Soprano K J. Regulation of human ornithine decarboxylase expression by the c-Myc.Max protein complex. J Biol Chem. 1993;268:27277–27285. [PubMed] [Google Scholar]
- 53.Potter H, Weir L, Leder P. Enhancer-dependent expression of human kappa immunoglobulin genes introduced into pre-B lymphocytes by electroporation. Proc Natl Acad Sci USA. 1984;81:7161–7165. doi: 10.1073/pnas.81.22.7161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Prendergast G C, Ziff E B. A new bind for Myc. Trends Genet. 1992;8:91–96. doi: 10.1016/0168-9525(92)90196-b. [DOI] [PubMed] [Google Scholar]
- 55.Ray R, Thomas S, Miller D. Mouse fibroblasts transformed with the human c-myc gene express a high level of mRNA but a low level of c-myc protein and are non-tumorigenic in nude mice. Oncogene. 1989;4:593–600. [PubMed] [Google Scholar]
- 56.Reisman D, Elkind N B, Roy B, Beamon J, Rotter V. c-Myc transactivates the p53 promoter through a required downstream CACGTG motif. Cell Growth Differ. 1993;4:57–65. [PubMed] [Google Scholar]
- 57.Ruegg C L, Wu H Y, Fagnoni F F, Engleman E G, Laus R. B4B, a novel growth-arrest gene, is expressed by a subset of progenitor/pre-B lymphocytes negative for cytoplasmic mu-chain. J Immunol. 1996;157:72–80. [PubMed] [Google Scholar]
- 58.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 59.Schiemann S, Ruckels M, Engelholm L H, Schwirzke M, Brunner N, Weidle U H. Differential gene expression in human mammary carcinoma cells: identification of a new member of a receptor family. Anticancer Res. 1997;17:13–20. [PubMed] [Google Scholar]
- 60.Shim H, Dolde C, Lewis B C, Wu C S, Dang G, Jungmann R A, Dalla-Favera R, Dang C V. c-Myc transactivation of LDH-A: implications for tumor metabolism and growth. Proc Natl Acad Sci USA. 1997;94:6658–6663. doi: 10.1073/pnas.94.13.6658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Small M B, Hay N, Schwab M, Bishop J M. Neoplastic transformation by the human gene N-myc. Mol Cell Biol. 1987;7:1638–1645. doi: 10.1128/mcb.7.5.1638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Snipes G J, Suter U, Shooter E M. Human peripheral myelin protein-22 carries the L2/HNK-1 carbohydrate adhesion epitope. J Neurochem. 1993;61:1961–1964. doi: 10.1111/j.1471-4159.1993.tb09840.x. [DOI] [PubMed] [Google Scholar]
- 63.Stewart T A, Pattengale P K, Leder P. Spontaneous mammary adenocarcinomas in transgenic mice that carry and express MTV/myc fusion genes. Cell. 1984;38:627–637. doi: 10.1016/0092-8674(84)90257-5. [DOI] [PubMed] [Google Scholar]
- 64.Stone J, de Lange T, Ramsay G, Jakobovits E, Bishop J M, Varmus H, Lee W. Definition of regions in human c-myc that are involved in transformation and nuclear localization. Mol Cell Biol. 1987;7:1697–1709. doi: 10.1128/mcb.7.5.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Suter U, Snipes G J. Peripheral myelin protein 22: facts and hypotheses. J Neurosci Res. 1995;40:145–151. doi: 10.1002/jnr.490400202. [DOI] [PubMed] [Google Scholar]
- 66.Suter U, Snipes G J, Schoener S R, Welcher A A, Pareek S, Lupski J R, Murphy R A, Shooter E M, Patel P I. Regulation of tissue-specific expression of alternative peripheral myelin protein-22 (PMP22) gene transcripts by two promoters. J Biol Chem. 1994;269:25795–25808. [PubMed] [Google Scholar]
- 67.Tavtigian S V, Zabludoff S D, Wold B J. Cloning of mid-G1 serum response genes and identification of a subset regulated by conditional myc expression. Mol Biol Cell. 1994;5:375–388. doi: 10.1091/mbc.5.3.375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Taylor V, Suter U. Epithelial membrane protein-2 and epithelial membrane protein-3: two novel members of the peripheral myelin protein 22 gene family. Gene. 1996;175:115–120. doi: 10.1016/0378-1119(96)00134-5. [DOI] [PubMed] [Google Scholar]
- 69.Taylor V, Welcher A A, Suter U Amgen EST Program. Epithelial membrane protein-1, peripheral myelin protein 22, and lens membrane protein 20 define a novel gene family. J Biol Chem. 1995;270:28824–28833. doi: 10.1074/jbc.270.48.28824. [DOI] [PubMed] [Google Scholar]
- 70.Wagner A J, Meyers C, Laimins L A, Hay N. c-Myc induces the expression and activity of ornithine decarboxylase. Cell Growth Differ. 1993;4:879–883. [PubMed] [Google Scholar]
- 71.Wulf G, Adra C, Lim B. Inhibition of hematopoietic development from embryonic stem cells by antisense vav RNA. EMBO J. 1993;12:5065–5074. doi: 10.1002/j.1460-2075.1993.tb06200.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Yang W, Wang H, Fliegel L. Regulation of Na+/H+ exchanger gene expression. Role of a novel poly(dA · dT) element in regulation of the NHE1 promoter. J Biol Chem. 1996;271:20444–20449. doi: 10.1074/jbc.271.34.20444. [DOI] [PubMed] [Google Scholar]
- 73.Zoidl G, Blass K S, D’Urso D, Schmalenbach C, Muller H W. Retroviral-mediated gene transfer of the peripheral myelin protein PMP22 in Schwann cells: modulation of cell growth. EMBO J. 1995;14:1122–1128. doi: 10.1002/j.1460-2075.1995.tb07095.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Zoidl G, D’Urso D, Blass K S, Schmalenbach C, Kuhn R, Muller H W. Influence of elevated expression of rat wild-type PMP22 and its mutant PMP22Trembler on cell growth of NIH3T3 fibroblasts. Cell Tissue Res. 1997;287:459–470. doi: 10.1007/s004410050770. [DOI] [PubMed] [Google Scholar]