Abstract
Ligand-dependent activation of the platelet-derived growth factor receptor (PDGFR) in fibroblasts in culture leads to the activation of the JAK family of protein-tyrosine kinases and of the transcription factors Stat1 and Stat3. To determine the biochemical mechanism of STAT activation by PDGFR, we devised a cell-free system composed of a membrane fraction from cells overexpressing PDGFR. When supplemented with crude cytosol, the membrane fraction supported PDGF- and ATP-dependent activation of both Stat1 and Stat3. However, the extent of Stat3 activation differed depending on the source of the cytosolic fraction. Using purified recombinant STAT proteins produced in Escherichia coli, we found that Stat1 could be activated by immunopurified PDGFR and showed no additional requirement for membrane- or cytosol-derived proteins. In contrast, activation of Stat3 exhibited a strong requirement for the cytosolic fraction. The activity present in the cytosolic fraction could be depleted with antibodies to JAK proteins. We conclude that the mechanisms of activation of Stat1 and Stat3 by PDGFR are distinct. Stat1 activation appears to result from a direct interaction with the receptor, whereas Stat3 activation additionally requires JAK proteins.
Proteins of the STAT family of transcription factors are activated by a wealth of cytokines and polypeptide growth factors. Activation by cytokine receptors requires members of the JAK family of protein tyrosine kinases (2–4, 8–10, 15, 17, 24). The general model for STAT activation was based on the paradigm of STAT activation by the interferon receptors (5, 18). Gamma interferon activates both Jak1 and Jak2, which are in turn required for the activation of Stat1 (12, 25). Alpha interferon activates Jak1 and Tyk2, leading to the activation of Stat1 and Stat2 (12, 21). For interferons and other cytokines that act through cell-surface receptors that lack intrinsic protein-tyrosine kinase activity, receptor-associated JAKs provide the catalytic activity responsible for receptor phosphorylation to create docking sites for signaling proteins such as STATs and presumably for phosphorylation of the proteins recruited to the phosphorylated receptor.
Polypeptide growth factors, such as epidermal growth factor and platelet-derived growth factor (PDGF), also activate JAKs and STATs (11, 22). Because the receptors for these factors harbor intrinsic protein-tyrosine kinase activity, the role of receptor-activated JAKs, if any, in STAT activation by these receptors remains unclear. We used a cell-free system and partially purified proteins to examine the biochemical steps in STAT activation by the PDGF receptor (PDGFR). We found that the mechanisms of activation of Stat1 and Stat3 are different and that JAKs may be required only for the activation of Stat3.
MATERIALS AND METHODS
Cell cultures and cellular extracts.
The cell line 2fTGH.PS1 (22), derived from the human fibrosarcoma HT1080 cells, was grown in Dulbecco modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS).
Cytosolic extracts were prepared from actively growing subconfluent cells starved for 15 to 18 h in DMEM supplemented with 0.5% FBS. Cells were scraped from the dishes into phosphate-buffered saline containing 2 mM Na3VO4 and resuspended in hypotonic buffer (20 mM HEPES [pH 7.9]; 20 mM NaF; 1 mM Na2P2O7; 1 mM EDTA; 1 mM EGTA; 0.25 mM ammonium molybdate; 2 mM Na3VO4; 3-μg/ml concentrations of leupeptin, aprotinin, and pepstatin; 0.5 mM phenylmethylsulfonyl fluoride [PMSF]; and 1 mM dithiothreitol [DTT]. Cells were lysed by repeated passage through a 25-gauge needle, and cytosolic extracts were prepared as described previously (22).
Membrane fractions were prepared from confluent 2fTGH.PS1 dishes starved for 24 h in DMEM supplemented with 0.5% FBS. Cells were scraped from the dishes into phosphate-buffered saline containing 2 mM Na3VO4, resuspended in hypotonic buffer, and lysed by repeated passage through a 25-gauge needle. The extract was spun twice for 1 min to eliminate crude nuclei and unbroken cells and supplemented with NaCl to 120 mM. Membranes were sedimented by centrifugation (20 min at 16,000 × g), and the membrane pellet was collected after two additional washes in hypotonic buffer. The 2fTGH.PS1 membrane pellets were resuspended in MRB buffer (20 mM Tris [pH 7.4]; 20 mM NaF; 1 mM EDTA; 150 mM NaCl; 10% glycerol; 2 mM Na3VO4; 1-μg/ml concentrations of leupeptin, aprotinin, and pepstatin; 0.5 mM PMSF; 1 mM DTT; and 0.25% Triton).
In vitro kinase assay.
The membrane fraction (20 μg/3 μl) was mixed with the STAT proteins and PDGF (PDGF-BB; Upstate Biotechnology Inc.) (6 ng). STAT proteins were either of cytosolic origin (12.5 to 25 μg of cytosolic extract) or purified from Escherichia coli (about 20 ng of the pure protein). The kinase reaction was initiated by the addition of a 5× kinase buffer (50 mM HEPES [pH 7.9], 50 mM MgCl2, 10 mM ATP). The reaction mixture (12.5 μl) was incubated at 30°C for 30 min and stopped by the addition of EDTA (15 mM final concentration). The reactions were spun for 5 min, and the supernatants (12 μl) were used for the electrophoretic mobility shift assay (EMSA).
EMSA.
Protein fractions activated in vitro were preincubated for 15 min at room temperature with 1 μg of poly(dI-dC) · poly(dI-dC) as nonspecific DNA competitor in the reaction buffer (13 mM HEPES [pH 7.9], 65 mM NaCl, 1 mM DTT, 0.15 mM EDTA, 2% Ficoll 400). The labeled probe (∼3 fmol) was then added to the reaction, which was incubated for another 20 min at room temperature. The protein-DNA complexes were separated by electrophoresis in 0.5× TBE (Tris-borate–EDTA) buffer through a 5% polyacrylamide gel (acrylamide-to-bis ratio, 39:1) containing 2.5% glycerol. Gels were dried and analyzed by autoradiography. The supershift experiment was performed using monoclonal antibody 12CA5 (7).
Immunoprecipitations and Western blots.
For immunoblot analysis of 2fTGH.PS1 or HeLa cytosolic extracts, proteins were denatured in Laemmli buffer, separated by sodium dodecyl sulfate–7.5% polyacrylamide gel electrophoresis and transferred to nitrocellulose. Western blots were performed with monoclonal antibodies against Stat1 or Stat3 N-terminal regions (Transduction Laboratories), phospho-Stat1 (Y701) (New England Biolabs, Inc.), or hemagglutinin (12CA5) (7). Immunoreactive bands were visualized with the epichemiluminescence Western blotting system (Amersham).
For the depletion of U3A (13) cytosol, protein extracts were incubated for 3 h at 4°C with polyclonal antibodies (Upstate Biotechnology Inc.) against JAK1, JAK2, Tyk2, or ERK2 prior to the addition of protein A Sepharose beads. For the immunoprecipitation of the PDGF-β receptor, membrane pellets from 2fTGH.PS1 cells were solubilized in MRB buffer containing 1% Triton and incubated with polyclonal antibodies against the human PDGF type B receptor (Upstate Biotechnology Inc.). Washes of the protein A Sepharose complexes were performed in the same buffer. Supernatant from the immunoprecipitation corresponded to the receptor-depleted membrane fraction whereas the bead pellet, further washed in the same buffer, provided the sample of immunoprecipitated receptor.
Cloning of histidine-tagged STATs for expression in E. coli: purification of the recombinant STATs.
Stat1 and Stat3 cDNAs were cloned downstream of a decahistidine tag in a pET-19b vector (Novagen). For Stat1, an additional hemagglutinin (Lerner) tag was inserted between the polyhistidine tag and the STAT ATG. Proteins were produced in E. coli and purified under native conditions on a nickel resin. For that purpose, BL21 strains expressing the recombinant STATs were grown in Luria-Bertani medium to an optical density at 600 nm of 0.8 and induced with 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) for 3 h. Cells were lysed by sonication in a buffer containing 50 mM Na-phosphate (pH 8.0); 300 mM NaCl; 0.25% Triton; 1-μg/ml concentrations of leupeptin, aprotinin, and pepstatin; 1 mM PMSF; and 3 mM β-mercaptoethanol. The sonication supernatant was adjusted to a final concentration of 40 mM imidazole and mixed with 200 μl of nickel packed beads for 30 min at 4°C. The beads were loaded onto a column and washed with 25 bed volumes of buffer containing 50 mM Na-phosphate (pH 8.0); 300 mM NaCl; 1-μg/ml concentrations of leupeptin, aprotinin, and pepstatin; 1 mM PMSF; 3 mM β-mercaptoethanol; 40 mM imidazole; and 10% glycerol. Elution of recombinant Stat1 and Stat3 was achieved with 120 mM and 200 mM imidazole, respectively. Stat3 fractions were immediately diluted twofold in washing buffer devoid of NaCl and imidazole. Protein fractions were kept at −70°C.
RESULTS
PDGF-dependent activation of Stat1 and Stat3 in vitro.
Stat1 and Stat3 are activated by the PDGF receptor in vivo (22, 23). When assayed by mobility shift assay on a DNA probe derived from the c-fos SIE (23), activated STATs form three specific complexes, termed SIF-A, SIF-B, and SIF-C, which correspond respectively to the binding of Stat3 homodimers, Stat1/Stat3 heterodimers, and Stat1 homodimers. We showed previously that in 2fTGH.PS1 cells (human fibrosarcoma cells engineered to express the PDGF-β receptor), binding of PDGF-BB to its receptor leads to the activation of Stat1 and Stat3 and the formation of all three SIF complexes (22).
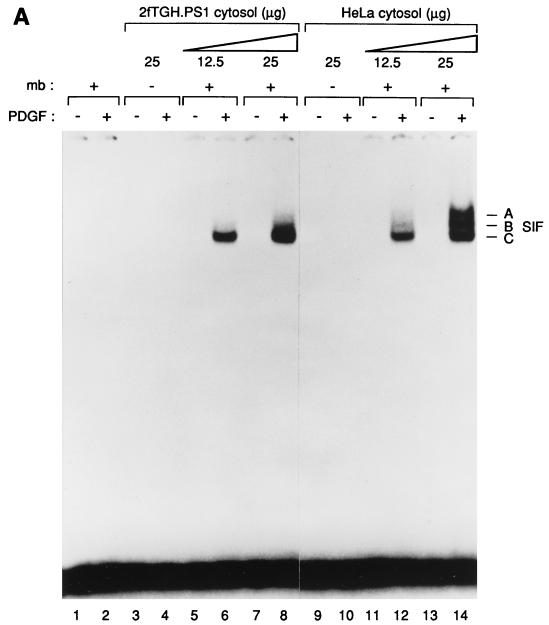
To examine the biochemical mechanism of STAT activation by the PDGF receptor in 2fTGH.PS1 cells, we extended a previous study (16) to develop a cell-free system composed of membrane and cytosol fractions derived from these cells. Addition of PDGF, magnesium, and ATP to reactions composed of 2fTGH.PS1 membranes and cytosol followed by incubation at 30°C for 30 min led to activation of STAT proteins and the appearance of SIF complexes on a mobility shift gel (Fig. 1A, lanes 1 to 8). Activation required both membrane and cytosol fractions and was dependent upon the addition of both ATP and PDGF (Fig. 1A and data not shown). These reaction conditions favored the activation of Stat1 over Stat3, as SIF-C, a Stat1 homodimer, was the primary product (lanes 6 and 8). This observation contrasts with the pattern observed following PDGF stimulation of the same cells in vivo, which leads primarily to activation of Stat3 and formation of the SIF-A complex (22).
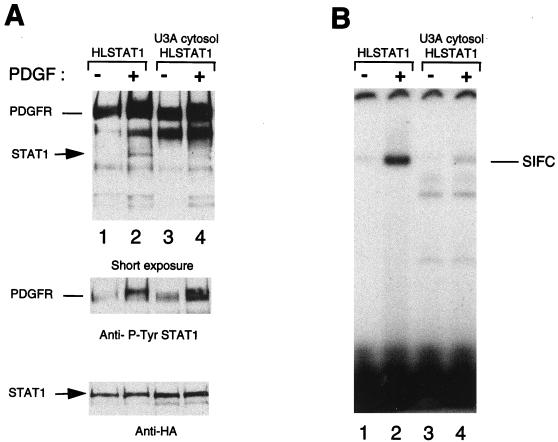
FIG. 1.
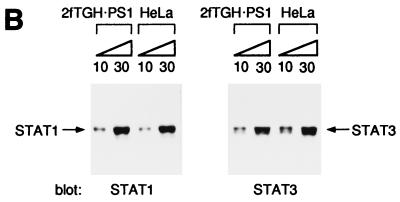
(A) PDGF-dependent activation of Stat1 and Stat3 in vitro. Detergent-solubilized membrane proteins from 2fTGH.PS1 cells, which overexpress the PDGF-β receptor, were mixed with cytosolic extracts from 2fTGH.PS1 or HeLa cells. The proteins were subjected to an in vitro kinase reaction in the presence of ATP and magnesium, and the activated STATs were analyzed by mobility shift assay using the high-affinity SIEm67 32P-labeled probe. Lanes: 1 and 2, no cytosolic extract; 3 to 8, 2fTGH.PS1 cytosol; 9 to 14, HeLa cytosol. The kinase reactions were done with (+) or without (−) membranes (mb) as indicated and in the presence (+) or absence (−) of PDGF as indicated. The three specific complexes SIF-A, SIF-B, and SIF-C are indicated at right. (B) 2fTGH.PS1 and HeLa cells express similar concentrations of Stat1 as well as of Stat3 proteins. Equal amounts (10 and 30 μg, respectively) of cytosolic extracts from 2fTGH.PS1 or HeLa cells were analyzed by Western blotting using antibodies directed against either Stat1 or Stat3. The STAT protein recognized by the antibodies is indicated.
When cytosol from HeLa cells was substituted for 2fTGH.PS1 cytosol, the pattern of STAT activation changed. With HeLa cytosol, significant amounts of the SIF-A and -B complexes were formed, indicating efficient activation of Stat3 as well as Stat1 (Fig. 1A, lane 14). The ability of HeLa cytosol to support the activation of Stat3 to a significantly greater extent than 2fTGH.PS1 cytosol could be due to a correspondingly higher concentration of Stat3 protein in the HeLa cytosol fraction. To test this idea, we compared the concentrations of Stat1 and Stat3 in 2fTGH.PS1 and HeLa cytosol by Western blotting (Fig. 1B). No significant differences were observed for either protein, suggesting that another difference in the constitution of the HeLa cytosol fraction accounts for its ability to support Stat3 activation in vitro.
Thus, this cell-free system is able to support the activation of both Stat1 and Stat3 by PDGF. Stat1 was activated with similar efficiency by using cytosolic fractions from either 2fTGH.PS1 or HeLa cells. In contrast, only HeLa cell extracts supported the activation of Stat3. These observations suggest that the biochemical mechanisms of activation of Stat1 and Stat3 may differ.
In vitro activation of a purified recombinant Stat1 protein.
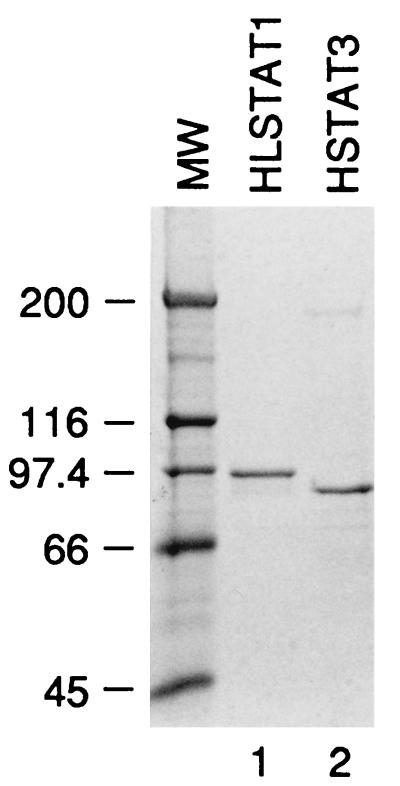
To begin to dissect the mechanisms of activation of Stat1 and Stat3 in vitro, we produced recombinant Stat1 and Stat3 in E. coli. Both proteins carried N-terminal histidine tags, allowing them to be purified on nickel resin, leading to the protein preparations shown in Fig. 2. The recombinant Stat1 protein also carried an epitope tag immediately C-terminal to the histidine tag. A Stat3 protein containing the epitope tag could not be produced.
FIG. 2.
Purification of Stat1 and Stat3. Histidine-tagged Stat1 and Stat3 proteins were expressed in E. coli and purified on nickel. The purified proteins were analyzed by polyacrylamide gel electrophoresis and revealed by Coomassie staining. Stat3 harbors the histidine tag at its N terminus (HSTAT3) while Stat1 contains an additional hemagglutinin tag between the polyhistidine sequence and the N terminus of the protein (HLSTAT1). The molecular weight (MW) markers are indicated (in thousands) on the left of the gel.
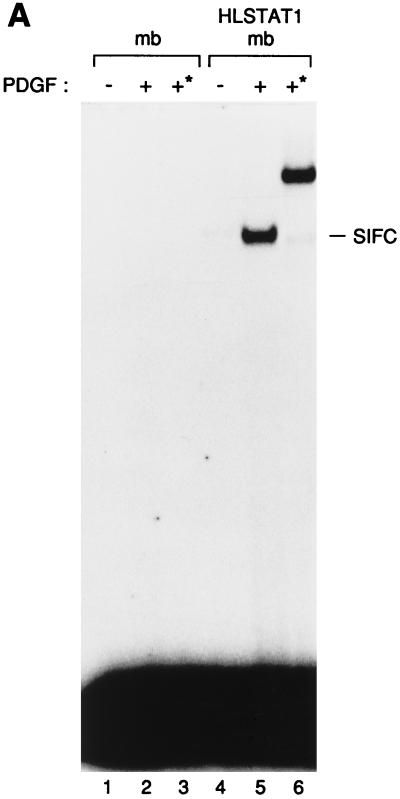
To determine whether purified recombinant Stat1 (HLSTAT1) could substitute for the cytosolic fraction in our PDGF-dependent system, we added HLSTAT1 to a membrane fraction from 2fTGH.PS1 cells. The addition of PDGF to reactions composed of the membrane fraction and purified HLSTAT1 led to the appearance of a SIF-C complex (Fig. 3A, lane 5). This complex was not observed when HLSTAT1 was omitted from the reaction (lane 2). Furthermore, the addition of a monoclonal antibody that recognizes an N-terminal epitope tag incorporated into HLSTAT1 shifted the mobility of the induced SIF-C complex (lane 6). Thus, this complex contains the recombinant Stat1 protein, and these data suggest that the 2fTGH.PS1 membrane fraction is sufficient to support the activation of Stat1 in vitro.
FIG. 3.
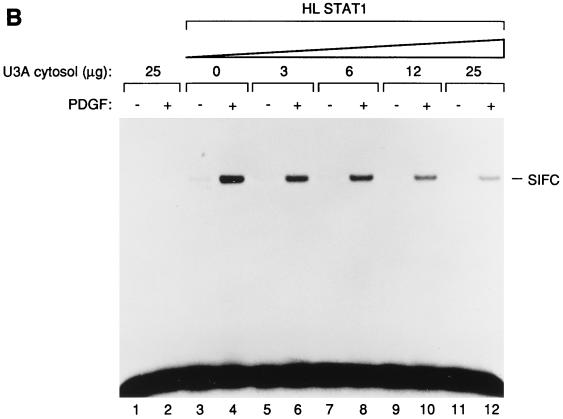
(A) In vitro activation of the purified Stat protein. The purified Stat1 protein was mixed with 2fTGH.PS1 membranes (mb), activated in vitro, and tested by EMSA on an SIE probe (lanes 4 to 6). Lanes 1 to 3, mock reactions with membranes only. The kinase reactions were done in the presence (+) or absence (−) of PDGF. The addition of the 12CA5 monoclonal antibody during the gel shift reaction is indicated by an asterisk. The SIF-C complex corresponding to the binding of Stat1 to the SIE site is indicated at the right. (B) The PDGF-dependent activation of Stat1 is inhibited by cytosolic proteins. The purified HLSTAT1 protein was mixed with increasing concentrations of U3A cytosolic extracts and subjected to the in vitro kinase reaction using 2fTGH.PS1 membrane fractions as the source of PDGF-β receptor. As a control, U3A cytosolic proteins (25 μg) were tested in the kinase reaction in the absence of recombinant Stat1 protein (lanes 1 and 2). The kinase reactions were done in the absence (−) or presence (+) of PDGF. The SIF-C complex is indicated.
To determine whether additional activities in the cytosol might also be required for Stat1 activation by PDGF, we added cytosolic extracts from U3A cells, a derivative of 2fTGH that lacks endogenous Stat1 (13). This fraction failed to stimulate PDGF induction of HLSTAT1 DNA-binding activity (Fig. 3B) or phosphorylation of HLSTAT1 on tyrosine-701 (Fig. 4). Rather, it exhibited dose-dependent inhibition of HLSTAT1 activation. Taken together, these observations suggest that membrane-associated proteins from 2fTGH.PS1 cells are sufficient for activation of Stat1 by PDGFR in vitro and that no additional activities from the cytosol are required.
FIG. 4.
Stat1 tyrosine phosphorylation and DNA binding activity are both inhibited by cytosolic proteins. The purified HLSTAT1 protein was subjected to the in vitro kinase reaction using 2fTGH.PS1 membranes (40 μg), in the absence (−) or presence (+) of U3A cytosolic extract (100 μg). Following the kinase reaction, half of the reaction mixture was analyzed by Western blotting using Stat1 phosphotyrosine (Y701) antibodies (A), the second half of the reaction mixture was analyzed by EMSA on an SIE probe (B). The phosphorylated Stat1 protein is indicated. Reprobing of the blot with an antihemagglutinin antibody (12CA5) shows equal Stat1 concentrations in the various lanes.
To determine whether membrane proteins other than PDGFR itself are required for activation of HLSTAT1 by PDGF, we purified PDGFR from detergent-solubilized 2fTGH.PS1 membrane fractions by specific immunoprecipitation. Incubation of HLSTAT1 with immunopurified PDGFR led to ATP-dependent and partially ligand-dependent Stat1 activation (Fig. 5, lanes 4 to 6). The background activation of Stat1 in the absence of PDGF (lane 5) could be due to cross-linking of the PDGFR by antibody. In contrast, incubation of HLSTAT1 with immune complexes obtained with normal rabbit serum did not lead to Stat1 activation (lanes 13 to 15). HLSTAT1 was not activated by a membrane fraction that had been depleted of PDGFR with the receptor antibody (lanes 7 to 9), indicating that PDGFR was required for activation of Stat1 by PDGF, whereas activation by the membrane fraction was not affected by depletion with control serum (lanes 16 to 18). Furthermore, activation of HLSTAT1 by immunopurified PDGFR was not further augmented by the addition of the receptor-depleted membrane fraction (lanes 10 to 12). These observations suggest that purified PDGFR is sufficient for the activation of HLSTAT1 in vitro. We cannot formally rule out the possibility that another factor tightly associated with the receptor might act as an intermediate in Stat1 activation by PDGFR. However, any such protein would have to be present in excess in the receptor complex, since supplementation with the receptor-depleted membrane fraction did not further increase HLSTAT1 activation. We provisionally conclude that PDGFR is both necessary and sufficient for Stat1 activation in vitro.
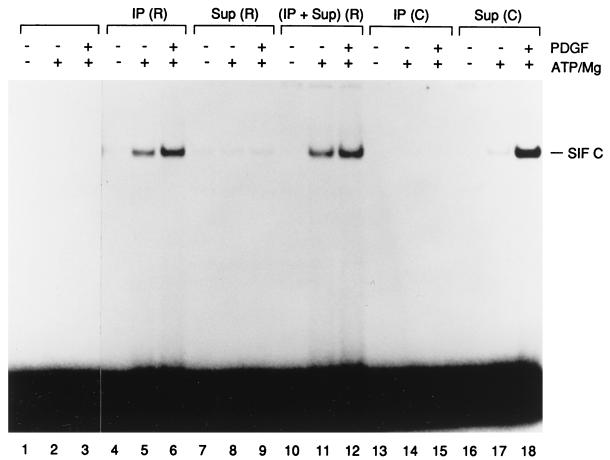
FIG. 5.
Activation of Stat1 protein by the immunoprecipitated PDGF-β receptor. The PDGF-β receptor was immunoprecipitated from detergent-solubilized membranes of 2fTGH.PS1 cells. The purified Stat1 was incubated in the in vitro kinase reaction with the immunoprecipitated PDGFR [IP (R); lanes 4 to 6], the membrane fraction depleted of PDGFR [Sup (R); lanes 7 to 9] or a mixture of the two [(IP + Sup) (R); lanes 10 to 12] and tested by EMSA on an SIE probe. In the mock immunoprecipitations [IP (C); lanes 13 to 15] or depletions [Sup (C); lanes 16 to 18], normal rabbit serum was used instead of specific anti-PDGFR antibodies. Lanes 1 to 3, control reactions where Stat1 was incubated without membrane proteins. The kinase reactions were done in the presence (+) or absence (−) of PDGF as well as of ATP/magnesium. The SIF-C complex corresponding to Stat1 binding to the SIE site is indicated.
In vitro activation of Stat3 requires additional cytoplasmic factors.
We performed a similar analysis using purified recombinant Stat3 (HSTAT3). In contrast to our observations with Stat1, the 2fTGH.PS1 membrane fraction alone was insufficient to support PDGF-dependent activation of Stat3 (Fig. 6A, lane 4). However, the addition of U3A cytosol, which was unable on its own to support SIF-A activation (lane 2) and inhibited activation of Stat1, led to a dose-dependent induction of the SIF-A complex, indicating activation of HSTAT3 (lanes 6 and 8). Thus, the combination of 2fTGH.PS1 membranes and U3A cytosol was required to reconstitute PDGF-dependent activation of purified recombinant Stat3 in vitro.
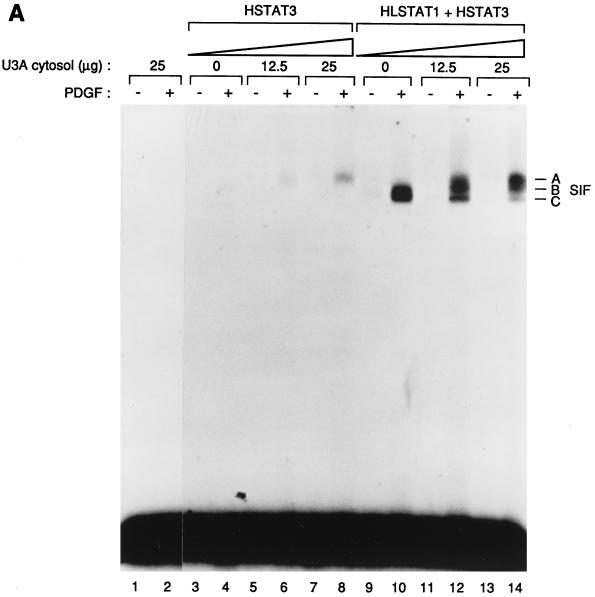
FIG. 6.
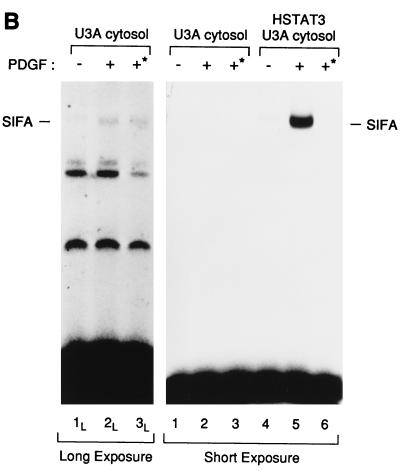
(A) In vitro activation of Stat3 by the PDGF receptor requires cytosolic proteins. The purified Stat3 protein (HSTAT3; lanes 3 to 8) or a mixture of Stat1 and Stat3 (HLSTAT1 + HSTAT3; lanes 9 to 14) were incubated with increasing concentrations of U3A cytosol, activated in vitro, and tested by EMSA on an SIE probe. As a control, the kinase reaction was done with U3A cytosol (25 μg), in the absence of recombinant STATs (lanes 1 and 2). Kinase reactions were done in the presence (+) or absence (−) of PDGF. The three specific complexes SIF-A, SIF-B, and SIF-C are indicated. (B) The SIF-A complex generated in vitro corresponds to the activation of the recombinant Stat3 protein. HSTAT3 was incubated with U3A cytosol for the kinase assay and tested by EMSA (lanes 4 to 6). Control reactions were performed in the absence of recombinant Stat3 (lanes 1 to 3). In lanes 3 and 6, protein extracts obtained by the kinase reactions were depleted on nickel resin of histidine-containing proteins prior to the EMSA. Depletion on nickel resin is indicated by an asterisk. The SIF-A complex obtained in the presence (+) of PDGF is indicated. The left panel shows a longer exposure of the gel (lanes 1L to 3L).
Because the addition of U3A cytosol appeared to have opposite effects on the activation of recombinant Stat1 and Stat3, respectively, we added U3A cytosol to a reaction containing a mixture of HLSTAT1 and HSTAT3. In the absence of cytosol, we mainly observed the induction of SIF-C (lane 10), whereas the addition of U3A cytosol led to the disappearance of SIF-C with the concomitant appearance of SIF-A (lanes 12 and 14). This observation reinforces the conclusion that activation of Stat3 but not Stat1 requires activities present in U3A cytosol and, therefore, that the mechanisms of activation of Stat1 and Stat3 must differ.
Although U3A cytosol without supplementation with recombinant Stat3 protein was unable to support SIF-A activation (Fig. 6A, lane 2; see also Fig. 6B, lane 2), we performed an additional experiment to verify that the SIF-A activity observed in the presence of U3A cytosol and HSTAT3 was indeed derived from the recombinant protein. An in vitro reaction containing both cytosol and HSTAT3 led to the appearance of the expected SIF-A complex (Fig. 6B, lane 5). If this activity contains the histidine-tagged recombinant protein, we expected to be able to deplete this activity with nickel resin. Therefore, following completion of the activation reaction, but prior to mobility shift analysis, the reaction was incubated with nickel resin, and the unbound material was analyzed by mobility shift assay. Incubation with nickel resin led to complete disappearance of SIF-A (lane 6), consistent with the idea that the SIF-A activity generated in this reaction was composed of recombinant protein.
To verify that the effect of the nickel resin was specific, we performed a parallel depletion on U3A cytosol alone. Although SIF-A activation by cytosol alone was not detectable at the same exposure level used to visualize activation of the recombinant protein, exposure of the same gel for several days allowed visualization of a weak SIF-A band derived from endogenous U3A Stat3 protein (lane 2L). This band was not affected by incubation with nickel resin (lane 3L). Thus, we conclude that membranes and cytosol are necessary and sufficient to activate purified recombinant Stat3.
JAK proteins are required for the activation of Stat3 by the PDGF receptor.
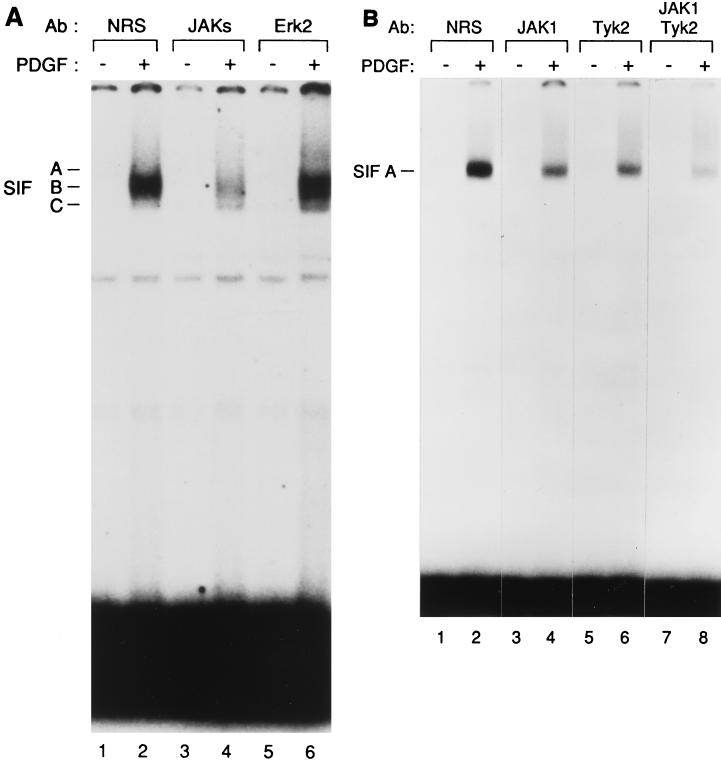
Our results suggest that activation of Stat3 by PDGFR in vitro requires proteins in the cytosolic fraction. One obvious candidate is the JAK family. To determine whether JAK proteins were required for the activity present in U3A cytosol, we treated the cytosol with a cocktail of antibodies specific for JAK1, JAK2, and Tyk2, respectively, with the goal of depleting the cytosol of all JAK activity. We conducted in vitro activation assays composed of 2fTGH.PS1 membranes, a mixture of recombinant Stat1 and Stat3, and U3A cytosol depleted with various antibodies. Treatment of U3A cytosol with normal rabbit serum led to a pattern of SIF activation similar to that seen with untreated cytosol (compare Fig. 7A, lane 2, to Fig. 6A, lane 14)—the predominant activities observed were SIF-A and -B. Treatment of the cytosol with antibody to Erk2 also did not affect induction of SIF-A and -B (Fig. 7A, lane 6). In contrast, when U3A cytosol was depleted with the cocktail of anti-JAK antibodies, activation of SIF-A and -B was significantly reduced (lane 4). This observation suggests that JAK proteins are required for U3A cytosol to support Stat3 activation.
FIG. 7.
(A) JAKs are required for Stat3 activation. A mixture of purified Stat1 and Stat3 recombinant proteins was incubated and activated in vitro with U3A cytosolic extracts that had been previously incubated with a cocktail of antibodies (Ab) to JAKs (lanes 3 and 4), with antibodies to ERK2 (lanes 5 and 6), or, as a control, with normal rabbit serum (NRS) (lanes 1 and 2). The activated STATs were then tested by EMSA on an SIE probe. The kinase reactions were done in the presence (+) or absence (−) of PDGF. The three specific complexes SIF-A, SIF-B, and SIF-C are indicated. (B) Individual JAKs contribute to Stat3 activation by the PDGF-β receptor. The purified Stat3 protein was incubated with U3A cytosol that had been previously depleted using antibodies to JAK1 (lanes 3 and 4), Tyk2 (lanes 5 and 6), a mixture of these two antibodies (lanes 7 and 8), or, as a control, with normal rabbit serum (lanes 1 and 2). The kinase reactions were done in the presence (+) or absence (−) of PDGF. The SIF-A complex obtained by EMSA on an SIE probe is indicated.
To confirm that JAKs were required for Stat3 activation in vitro, we repeated the depletion assay with antibodies against individual JAKs. Specific depletion of the U3A cytosol with antibodies to JAK1 (Fig. 7B, lane 4), Tyk2 (lane 6), and JAK2 (data not shown) each led to a partial inhibition of SIF-A induction. The effect of each antibody appeared to be additive, as seen by the results shown in lane 8. Thus, we believe that the observed requirement for JAKs can be satisfied by any of the JAK proteins.
In summary, we find that the biochemical mechanisms of activation of Stat1 and Stat3 by PDGFR are distinct. Stat1 activation does not require any activities present in the cytosol. Indeed, the purified PDGFR appears to be sufficient for Stat1 activation. In contrast, Stat3 activation requires activities present in the cytosol. One of those required activities is supplied redundantly by the JAK proteins.
DISCUSSION
We have developed a cell-free system that reproduces PDGF-dependent activation of Stat1 and Stat3. This system is composed of a membrane fraction derived from a cell line that overexpresses PDGFR (22) and was supplemented with either recombinant purified STAT proteins or with complex cytosolic fractions from tissue culture cells. Analysis of this system revealed unexpected differences in the mechanisms of activation of the two STAT proteins. Activation of Stat1 could be reconstituted with immunopurified PDGFR and purified recombinant Stat1. The addition of neither membrane nor cytosol fractions augmented Stat1 activation, indicating that PDGFR alone is probably sufficient for Stat1 activation in vitro. Although membrane fractions were prepared from extensively starved 2fTGH.PS1 cells to minimize any preexisting association of PDGFR with signaling intermediates, we cannot unambiguously rule out a requirement for an additional receptor-associated factor. Nevertheless, we favor the hypothesis that Stat1 is a direct substrate for PDGFR kinase.
We found that cytosolic fractions inhibited the activation of Stat1 by purified PDGFR. Several activities in the cytosol could be responsible for the observed inhibition of Stat1 tyrosine-701 phosphorylation. The cytosol could contain high levels of a phosphatase that reverses Stat1 phosphorylation. A second possibility is that the cytosol provides SH2-containing proteins that compete with Stat1 for its putative binding sites on the PDGFR, as has been shown for other STATs (1, 6, 14, 20).
In contrast to the apparently direct interaction between PDGFR and Stat1, PDGF-dependent activation of Stat3 displays a strong requirement for the cytosolic fraction. Depletion of JAK proteins from the cytosol abolishes this activity, suggesting that JAKs are required for activation of Stat3 by PDGFR. We have previously shown that all three ubiquitously expressed JAKs are associated with the activated PDGFR (22), although their role in PDGF signaling was not established. JAKs could be performing several functions in linking PDGFR to Stat3 activation in our cell-free system. First, receptor-associated JAKs could phosphorylate the receptor to generate Stat3-specific recruitment sites that are not phosphorylated by the receptor itself. Second, the JAKs could directly phosphorylate Stat3 itself, an activity presumably not enabled in PDGFR. Third, JAKs could provide Stat3 recruitment sites in the receptor complex, as proposed for Jak2 in response to growth hormone receptor activation (19).
Although PDGF treatment of intact 2fTGH.PS1 cells leads to activation of both Stat1 and Stat3 in roughly equal measure, a cell-free system composed of membrane and cytosol fractions from the same cells activates Stat1 almost exclusively. What accounts for the selective loss of Stat3 activation in the cell-free system? One simple explanation is that Stat1 activation is presumably a two-component reaction, comprising just PDGFR and Stat1, whereas Stat3 activation requires at least a third component present in the cytosol. Thus, Stat3 activation, which requires the association of at least three factors, should be much more sensitive to changes in the concentration of the reactants. Since the cell-free reaction is probably substantially more dilute with respect to the reactants than intact cells, Stat3 activation would be preferentially lost in vitro. That Stat3 activation is not substantially lost when HeLa cytosol is used in place of 2fTGH.PS1 cytosol suggests that one component of the Stat3 activation reaction, presumably the JAKs, is present at either a higher concentration or higher specific activity in the HeLa fraction.
Finally, we showed previously that PDGF induced the phosphorylation of all three JAKs in 2fTGH.PS1 cells but that no single JAK was uniquely required for STAT activation by PDGF (22). We reached this conclusion based on the use of cell lines that lacked individual JAKs. Because we had no cell lines lacking all of the JAKs, we could not rule out the possibility that the three JAKs present in these cells were functionally redundant for STAT activation but that at least one was required. The biochemical experiments we describe here allowed us to address this possibility, since we could deplete the JAKs both individually and in combination. Indeed, we found that depletion of all three JAK proteins from the cytosolic fraction substantially eliminated Stat3 activation in vitro, whereas depletion of individual JAKs yielded intermediate effects. Thus, we believe that each JAK is independently capable of mediating Stat3 activation by PDGFR.
ACKNOWLEDGMENTS
We thank Theresa Nahreini for help with protein expression and Mandy Cunningham for manuscript preparation.
This work was supported by Public Health Service grant CA45642 from the National Cancer Institute.
REFERENCES
- 1.Aman M J, Leonard W J. Cytokine signaling: cytokine-inducible signaling inhibitors. Curr Biol. 1997;7:784–788. doi: 10.1016/s0960-9822(06)00405-2. [DOI] [PubMed] [Google Scholar]
- 2.Bach E A, Auguet M, Schreiber R D. The IFNγ receptor: a paradigm for cytokine receptor signaling. Annu Rev Immunol. 1997;15:563–591. doi: 10.1146/annurev.immunol.15.1.563. [DOI] [PubMed] [Google Scholar]
- 3.Briscoe J, Kohlhuber F, Müller M. JAKs and STATs branch out. Trends Cell Biol. 1996;6:336–340. doi: 10.1016/0962-8924(96)10028-3. [DOI] [PubMed] [Google Scholar]
- 4.Darnell J E., Jr STATs and gene regulation. Science. 1997;277:1630–1635. doi: 10.1126/science.277.5332.1630. [DOI] [PubMed] [Google Scholar]
- 5.Darnell J J, Kerr I M, Stark G R. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science. 1994;264:1415–1421. doi: 10.1126/science.8197455. [DOI] [PubMed] [Google Scholar]
- 6.Endo T A, Masuhara M, Yokouchi M, Suzuki R, Sakamoto H, Mitsui K, Matsumoto A, Tanimura S, Ohtsubo M, Misawa H, Miyazaki T, Leonor N, Taniguchi T, Fujita T, Kanakura Y, Komiya S, Yoshimura A. A new protein containing an SH2 domain that inhibits JAK kinases. Nature. 1997;387:921–924. doi: 10.1038/43213. [DOI] [PubMed] [Google Scholar]
- 7.Field J, Nikawa J, Broek D, MacDonald B, Rodgers L, Wilson I A, Lerner R A, Wigler M. Purification of a RAS-responsive adenylyl cyclase complex from Saccharomyces cerevisiae by use of an epitope addition method. Mol Cell Biol. 1988;8:2159–2165. doi: 10.1128/mcb.8.5.2159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ihle J N. STATs: signal transducers and activators of transcription. Cell. 1996;84:331–334. doi: 10.1016/s0092-8674(00)81277-5. [DOI] [PubMed] [Google Scholar]
- 9.Ivashkiv L B. Cytokines and STATs: how can signals achieve specificity? Immunity. 1995;3:1–4. doi: 10.1016/1074-7613(95)90152-3. [DOI] [PubMed] [Google Scholar]
- 10.Johnston J A, Bacon C M, Riedy M C, O’Shea J J. Signaling by IL-2 and related cytokines: JAKs, STATs, and relationship to immunodeficiency. J Leukoc Biol. 1996;60:441–452. doi: 10.1002/jlb.60.4.441. [DOI] [PubMed] [Google Scholar]
- 11.Leaman D W, Leung S, Li X, Stark G R. Regulation of STAT-dependent pathways by growth factors and cytokines. FASEB J. 1996;10:1578–1588. [PubMed] [Google Scholar]
- 12.Müller M, Briscoe J, Laxton C, Guschin D, Ziemiecki A, Silvennoinen O, Harpur A G, Barbieri G, Witthuhn B A, Schindler C, Pellegrini S, Wilks A F, Ihle J N, Stark G R, Kerr I M. The protein tyrosine kinase JAK1 complements defects in interferon-alpha/beta and -gamma signal transduction. Nature. 1993;366:129–135. doi: 10.1038/366129a0. [DOI] [PubMed] [Google Scholar]
- 13.Müller M, Laxton C, Briscoe J, Schindler C, Improta T, Darnell J E, Jr, Stark G R, Kerr I M. Complementation of a mutant cell line: central role of the 91 kDa polypeptide of ISGF3 in the interferon-α and -γ signal transduction pathways. EMBO J. 1993;12:4221–4228. doi: 10.1002/j.1460-2075.1993.tb06106.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Naka T, Narazaki M, Hirata M, Matsumoto T, Minamoto S, Aono A, Nishimoto N, Kajita T, Taga T, Yoshizaki K, Akira S, Kishimoto T. Structure and function of a new STAT-induced STAT inhibitor. Nature. 1997;387:924–928. doi: 10.1038/43219. [DOI] [PubMed] [Google Scholar]
- 15.O’Shea J J. Jaks, STATs, cytokine signal transduction and immunoregulation: are we there yet? Immunity. 1997;7:1–11. doi: 10.1016/s1074-7613(00)80505-1. [DOI] [PubMed] [Google Scholar]
- 16.Sadowski H B, Gilman M Z. Cell-free activation of a DNA-binding protein by epidermal growth factor. Nature. 1993;362:79–83. doi: 10.1038/362079a0. [DOI] [PubMed] [Google Scholar]
- 17.Schindler C, Darnell J E., Jr Transcriptional responses to polypeptide ligands: the JAK-STAT pathway. Annu Rev Biochem. 1995;64:621–651. doi: 10.1146/annurev.bi.64.070195.003201. [DOI] [PubMed] [Google Scholar]
- 18.Shuai K. Interferon-activated signal transduction to the nucleus. Curr Opin Cell Biol. 1994;6:253–259. doi: 10.1016/0955-0674(94)90144-9. [DOI] [PubMed] [Google Scholar]
- 19.Sotiropoulos A, Moutoussamy S, Binart N, Kelly P A, Finidori J. The membrane proximal region of the cytoplasmic domain of the growth hormone receptor is involved in the activation of Stat 3. FEBS Lett. 1995;369:169–172. doi: 10.1016/0014-5793(95)00734-q. [DOI] [PubMed] [Google Scholar]
- 20.Starr R, Willson T A, Viney E M, Murray L J L, Rayner J R, Jenkins B J, Gonda T J, Alexander W S, Metcalf D, Nicola N A, Hilton D J. A family of cytokine-inducible inhibitors of signaling. Nature. 1997;387:917–921. doi: 10.1038/43206. [DOI] [PubMed] [Google Scholar]
- 21.Velazquez L, Fellous M, Stark G R, Pellegrini S. A protein tyrosine kinase in the interferon alpha/beta signaling pathway. Cell. 1992;70:313–322. doi: 10.1016/0092-8674(92)90105-l. [DOI] [PubMed] [Google Scholar]
- 22.Vignais M-L, Sadowski H B, Watling D, Rogers N C, Gilman M Z. Platelet-derived growth factor induces phosphorylation of multiple JAK family kinases and STAT proteins. Mol Cell Biol. 1996;16:1759–1769. doi: 10.1128/mcb.16.4.1759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wagner B J, Hayes T E, Hoban C J, Cochran B H. The SIF binding element confers sis/PDGF inducibility onto the c-fos promoter. EMBO J. 1990;9:4477–4484. doi: 10.1002/j.1460-2075.1990.tb07898.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Watanabe S, Arai K-I. Roles of the JAK-STAT system in signal transduction via cytokine receptors. Curr Opin Genet Dev. 1996;6:587–596. doi: 10.1016/s0959-437x(96)80088-8. [DOI] [PubMed] [Google Scholar]
- 25.Watling D, Guschin D, Müller M, Silvennoinen O, Witthuhn B A, Quelle F W, Rogers N C, Schindler C, Stark G R, Ihle J N. Complementation by the protein tyrosine kinase JAK2 of a mutant cell line defective in the interferon-gamma signal transduction pathway. Nature. 1993;366:166–170. doi: 10.1038/366166a0. [DOI] [PubMed] [Google Scholar]