Abstract
Circulating tumor DNA (ctDNA) analysis is a promising non-invasive technique for active surveillance after chemoradiotherapy for locally advanced resectable esophageal carcinoma. In other malignancies false-positive results in ctDNA analysis have been reported due to clonal hematopoiesis. In this case, we present a 66-year-old male who had adenocarcinoma of the gastroesophageal junction for which he received neoadjuvant chemoradiotherapy and underwent a transhiatal esophagectomy. Postoperatively our patient received follow-up with ctDNA analysis using next generation sequencing (NGS) and droplet digital PCR (ddPCR). This case report illustrates a number of the current challenges in ctDNA diagnostics in esophageal carcinoma. Firstly, the TP53 c.524G>A; p.R175H mutation that was found in preoperative tumor biopsies became detectable in ctDNA only after distant metastases had already been confirmed by clinical symptoms and standard imaging- and biopsy techniques. Secondly our patient repeatedly had false-positive outcomes of ctDNA analysis. Genomic analysis of white blood cells revealed that the origin of these discordant mutations lies in clonal hematopoiesis. Failure to detect TP53 c.524G>A; p.R175H in cell-free DNA (cfDNA) is most likely due to the amount of ctDNA in the cfDNA fraction being below the limit of detection for NGS and ddPCR analyses. Clinicians should be aware of the possibility of finding mutations originating from clonal hematopoiesis when using ctDNA analysis during active surveillance for esophageal carcinoma. We recommend correlation of mutations in cfDNA with mutations in tumor biopsies.
Keywords: Esophageal cancer, active surveillance, clonal hematopoiesis, circulating tumor DNA (ctDNA), case report
Introduction
For patients with resectable locally advanced esophageal cancer, active surveillance after a clinically complete response to neoadjuvant chemoradiotherapy (nCRT) is currently being investigated as a novel treatment option (1-3). Limitations to organ-sparing approaches presently under investigation consist of a 10% risk of subclinical residual disease, and risk of patient non-compliance due to the high burden of invasive diagnostics (4,5). A promising new diagnostic approach in active surveillance is analysis of circulating tumor DNA (ctDNA). This involves detection of DNA fragments that enter the circulation by release from necrotic or apoptotic tumor tissue, or by active release from vital tumor cells (6). Tumor-specific genetic alterations can be identified in plasma-derived cell-free DNA (cfDNA) by next generation sequencing (NGS) (7-9). Ideally, residual or recurrent tumors after neoadjuvant treatment can be detected using these so-called “liquid biopsies”. For esophageal cancer, the application of liquid biopsies is still in its infancy. However, in some malignancies like non-small cell lung cancer and colorectal carcinoma it is already an accepted and valued modality for diagnosis of actionable mutations and detection of targeted treatment resistance mechanisms (10). Nevertheless, occasionally NGS of cfDNA yields mutations that cannot be matched with mutations found in tissue samples of the primary tumor. Hu et al. for example reported a series of thirty-three non-small cell lung cancer cases with a confirmed TP53 mutation in plasma genotyping of whom nineteen could not be identified in tumor biopsy (11). In that report, NGS of peripheral white blood cells reveals concordant somatic mutations within non-malignant hematopoietic cells associated with clonal hematopoiesis in five of these false-positive cases (12). Clonal hematopoiesis refers to clonal outgrowth of hematopoietic progenitor cells with somatic mutations. This phenomenon is not uncommon in the general population and affects 5–13% of people over 70 years old and 25% of cancer patients (12-15). White blood cells are the source of a significant proportion of circulating cfDNA (16). Therefore, clonal hematopoiesis potentially is a major factor in discordance between tumor genotyping and plasma cfDNA in not only non-small cell lung cancer, but other malignancies as well.
Herein we describe a patient with adenocarcinoma of the esophagogastric junction who underwent nCRT followed by surgical resection in whom genotyping of plasma cfDNA was discordant with matched tumor tissue due to clonal hematopoiesis. We present the following article in accordance with the CARE reporting checklist (available at https://dx.doi.org/10.21037/atm-21-525) (17).
Case presentation
A 66-year-old male was referred to our tertiary referral hospital in February 2016 (Figure 1). All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee(s) and with the Helsinki Declaration (as revised in 2013). Written informed consent was obtained from the patient to perform comprehensive genomic testing and to publish his information. The patient had a medical history of gastroesophageal reflux disease and Barrett’s esophagus. He had no medical history of other malignancies. After standard diagnostic workup involving endoscopy with tumor biopsies plus endoscopic ultrasonography, diagnostic computed tomography (CT) and positron emission tomography with CT (PET-CT), the patient was diagnosed with cT3N0M0 (7th ed. UICC TNM staging manual) adenocarcinoma of the gastroesophageal junction (t=0). NGS of tumor biopsies revealed a TP53 missense mutation (c.524G>A; p.R175H) with a variant allele frequency (VAF) of 71% (Table 1). The patient was initially treated with nCRT according to the ChemoRadiotherapy for Oesophageal cancer followed by Surgery Study (CROSS)-regimen (18). The patient participated in the preSANO trial, which aimed to assess the accuracy of diagnostic response evaluations after nCRT (5). In context of this trial, 14 weeks (t=2) as well as 21 weeks after baseline (t=3), post-nCRT endoscopy was performed during which no macroscopic tumor could be found. At these points in time, biopsies were taken of the tumor site, showing no residual tumor cells.
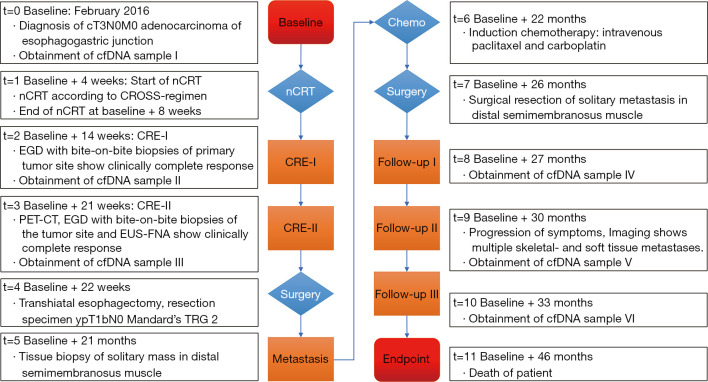
Figure 1.
Timeline of events from baseline to death of the patient. cfDNA, cell-free DNA; nCRT, neoadjuvant chemoradiotherapy; CROSS, ChemoRadiotherapy for Oesophageal cancer followed by Surgery Study; CRE, clinical response evaluation; EGD, esophago-gastro-duodenoscopy; PET-CT, positron emission tomography plus computed tomography; EUS-FNA, endoscopic ultrasonography with fine-needle aspiration; TRG, tumor regression grade.
Table 1. Characteristics of next generation sequencing outcomes of resection specimens, tumor biopsies and cfDNA per point in time.
| T | Moment of sample | Analysis | DNA concentration (ng/μL) | Total DNA (ng) | Allele | Allele Mol. Freq. | Mol. Cov. | Allele Mol. Cov. | Allele Read Freq. | Coverage | Allele Read Cov. | Remarks/allele call |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| t=0 (baseline) | Baseline, tissue | NGS with diagV5 panel | 1.57 | 4.7 | c.524G>A; p.R175H | – | – | – | 71% | n/a | n/a | – |
| c.646G>A; p.V216M | – | – | – | – | – | – | – | |||||
| c.659A>G; p.Y220C | – | – | – | – | – | – | – | |||||
| NGS with Oncomine | c.524G>A; p.R175H | 52.24% | 1,003 | 524 | 54.00% | 62,022 | 33,492 | Heterozygous | ||||
| cfDNA colon panel | c.646G>A; p.V216M* | 0.00%* | 876* | 0* | 0* | 74,997* | 0* | Absent | ||||
| c.659A>G; p.Y220C* | 0.00%* | 876* | 0* | 0* | 74,997* | 0* | Absent | |||||
| t=0 (baseline) | Baseline, cfDNA | NGS with Oncomine | 2.14 | 20 | c.524G>A; p.R175H* | 0.03%* | 2,968* | 1* | 0.05%* | 43,333* | 20* | No call |
| cfDNA Colon panel | c.646G>A; p.V216M | 0.37% | 3,006 | 11 | 0.30% | 54,264 | 162 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.00%* | 3,006* | 0* | 0.00%* | 54,264* | 0* | Absent | |||||
| t=2 (baseline + 14 w) | CRE-I | NGS with Oncomine | 2.07 | 21 | c.524G>A; p.R175H* | 0.04%* | 5,091* | 2* | 0.03%* | 71,538* | 22* | No call |
| cfDNA Colon panel | c.646G>A; p.V216M | 0.34% | 5,040 | 17 | 0.27% | 91,216 | 249 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.02%* | 5,040* | 1* | 0.01%* | 91,216* | 10* | Absent | |||||
| t=3 (baseline + 21 w) | CRE-II | NGS with Oncomine | 2.58 | 21 | c.524G>A; p.R175H* | 0.00%* | 4,433* | 0* | 0.00%* | 68,079* | 0* | Absent |
| cfDNA colon panel | c.646G>A; p.V216M | 0.32% | 4,354 | 14 | 0.30% | 83,850 | 253 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.00%* | 4,354* | 0* | 0.00%* | 83,850* | 0* | Absent | |||||
| t=4 (baseline + 22 w) | Resection, specimen | n/a | – | – | c.524G>A; p.R175H | – | – | – | – | – | – | – |
| c.646G>A; p.V216M | – | – | – | – | – | – | – | |||||
| c.659A>G; p.Y220C | – | – | – | – | – | – | – | |||||
| t=5 (baseline + 21 m) | Biopsy, metastasis | NGS with diagV5 panel | 0.2 | 0.6 | c.524G>A; p.R175H | – | – | – | 26% | n/a | n/a | – |
| c.646G>A; p.V216M | – | – | – | – | – | – | – | |||||
| c.659A>G; p.Y220C | – | – | – | – | – | – | – | |||||
| t=7 (baseline + 26 m) | Resection, specimen, metastasis | NGS with diagV5 panel | 1.2 | 3.6 | c.524G>A; p.R175H | – | – | – | 59% | n/a | n/a | – |
| c.646G>A; p.V216M | – | – | – | – | – | – | – | |||||
| c.659A>G; p.Y220C | – | – | – | – | – | – | – | |||||
| NGS with Oncomine | c.524G>A; p.R175H | 39.52% | 2,715 | 1,073 | 39.38% | 59,754 | 23,533 | Heterozygous | ||||
| cfDNA Colon panel | c.646G>A; p.V216M* | 0.06%* | 3,260* | 2* | 0.08%* | 92,215* | 77* | Absent | ||||
| c.659A>G; p.Y220C* | 0.00%* | 3,260* | 0* | 0* | 92,215* | 0* | Absent | |||||
| t=8 (baseline + 27 m) | FU I | NGS with Oncomine | 1.76 | 35.2 | c.524G>A; p.R175H* | 0.04%* | 5,217* | 2* | 0.05%* | 64,998* | 29* | No call |
| cfDNA Colon panel | c.646G>A; p.V216M | 0.59% | 4,935 | 29 | 0.50% | 73,632 | 367 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.00%* | 4,935* | 0* | 0.00%* | 72,632* | 0* | Absent | |||||
| t=9 (baseline + 30 m) | FU II | NGS with Oncomine | 2.76 | 55.2 | c.524G>A; p.R175H | 0.23% | 4,325 | 10 | 0.22% | 48,465 | 105 | Heterozygous |
| cfDNA Colon panel | c.646G>A; p.V216M | 0.86% | 4,169 | 36 | 0.79% | 57,420 | 454 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.05%* | 4,169* | 2* | 0.07%* | 57,420* | 39* | No call | |||||
| WBCs-FU II# | NGS with Oncomine# | 3.82# | 359.08# | c.524G>A; p.R175H* | 0.00%* | 6,483* | 0* | 0.00%* | 48,676* | 0* | Absent | |
| cfDNA colon panel# | c.646G>A; p.V216M# | 0.77%# | 5,980# | 46# | 0.71%# | 51,804# | 367# | Heterozygous | ||||
| c.659A>G; p.Y220C# | 0.05%# | 5,980# | 3# | 0.04%# | 51,804# | 22# | Heterozygous | |||||
| t=10 (baseline + 33 m) | FU III | NGS with Oncomine | 1.16 | 15.66 | c.524G>A; p.R175H | 0.19% | 2,710 | 5 | 0.09% | 33,774 | 31 | Heterozygous |
| cfDNA Colon panel | c.646G>A; p.V216M | 0.83% | 2,166 | 18 | 1.02% | 32,718 | 333 | Heterozygous | ||||
| c.659A>G; p.Y220C* | 0.05%* | 2,166* | 1* | 0.05%* | 32,718* | 15* | No call | |||||
| WBCs-FU III# | NGS with Oncomine# | 105.6# | 1425.6# | c.524G>A; p.R175H* | 0.00%* | 3,513* | 0* | 0.00%* | 35,466* | 0* | Absent | |
| cfDNA colon panel# | c.646G>A; p.V216M# | 0.48%# | 3,108# | 15# | 0.49%# | 38,982# | 191# | Heterozygous | ||||
| c.659A>G; p.Y220C# | 0.10%# | 3,108# | 3# | 0.09%# | 38,982# | 36# | Heterozygous |
Reference sequence TP53 (NM_000546) was used for DNA variant annotation. *, measurements below quality control threshold (VAF <0.1%; Allele Mol. Cov. <3); #, analysis of white blood cells. VAF, variant allele frequency (percentage based on the number of variant alleles covered divided by the total amount of molecules covered); Mol. Cov., molecular coverage (total number of independent molecules sequenced); Allele Mol. Cov., allele molecular coverage: number of independent molecules with a variant; allele read freq., allele read frequency (percentage of variant allele reads in total coverage); Allele Read Cov., allele read coverage; Cov., coverage (number of unique reads); NGS, next generation sequencing; cfDNA, cell-free DNA; CRE, clinical response evaluation; WBC, white blood cells; FU, follow-up.
Liquid biopsies were obtained at baseline (t=0) as well as 14 weeks after baseline (t=2; i.e., 8 weeks after completion of nCRT) and 21 weeks after baseline (t=3; i.e., 13 weeks after completion of nCRT). Analysis of cfDNA with Oncomine NGS at these points in time did not identify TP53 c.524G>A; p.R175H at baseline (t=0) and 14 weeks after baseline (t=2), in 3 or more independent molecules and a VAF of >0.1% (for 20 ng input). NGS did, however, consistently result in identification of a TP53 c.646G>A; p.V216M (VAF 0.37%, 0.34% and 0.32%, respectively) mutation. Twenty-two weeks after baseline (t=4; i.e., 14 weeks after completion of nCRT) the patient underwent a standard transhiatal esophagectomy. Resection specimen showed ypT1bN0 (all 19 removed lymph nodes negative) with Mandard’s Tumor Regression Grade 2 (1–10% residual vital tumor cells). The resection specimen did not contain sufficient residual tumor tissue for NGS.
Twenty-one months after initial diagnosis (t=5), biopsy from a solitary mass in the distal semimembranosus muscle revealed adenocarcinoma proving the patient had metastatic disease. Tissue mutation analysis revealed a TP53 c.524G>A; p.R175H mutation, confirming it being a metastasis from the forementioned esophageal carcinoma. The patient received palliative chemotherapy (t=6) with 6 cycles of intravenous paclitaxel and carboplatin. PET-CT imaging post-chemotherapy continued to show only this solitary soft tissue metastasis, therefore, the patient underwent surgical resection of this mass (t=7). Pathological analysis of metastatic tissue did not reveal remarkable mitotic activity. TP53 c.524G>A; p.R175H mutation again was identified after NGS of the resected metastasis. Twenty-seven months after baseline (t=8), during follow-up at the surgical outpatient clinic, liquid biopsies were obtained revealing TP53 c.646G>A; p.V216M (VAF 0.59%) mutation. The patient experienced a significant progression of symptoms 30 months after baseline (t=9) reporting pain, weight loss and malaise. Another set of liquid biopsies was obtained and at this point in time NGS identified TP53 c.524G>A; p.R175H (VAF 0.23%) as well as TP53 c.646G>A; p.V216M (VAF 0.86%) in cfDNA. Imaging studies showed a total of 12 skeletal- and soft-tissue metastases located in the vertebral column, the dorsal musculature, the pelvis and the bones and muscles of the right lower extremity (Figure 2). The patient received palliative treatment with analgesic medication and localized external beam radiation therapy on symptomatic bone metastases.
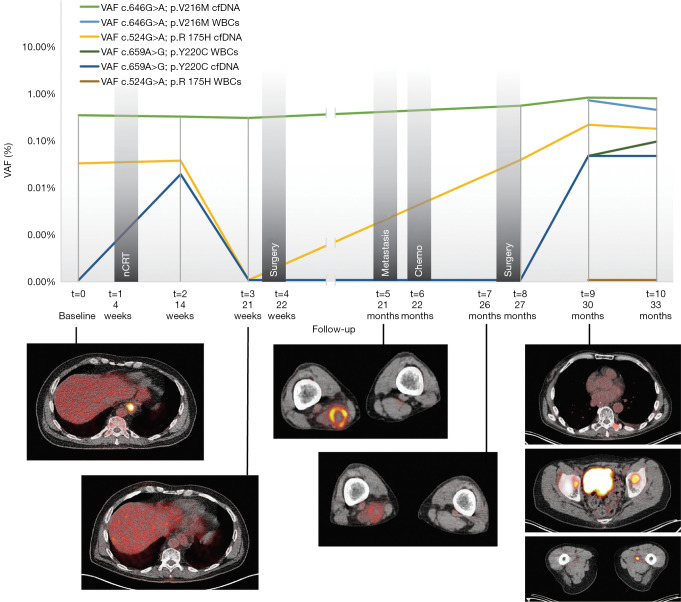
Figure 2.
Variant allele frequency dynamics over time in cell-free DNA and white blood cells with corresponding PET-CT images. Images from left to right: baseline PET-CT image showing FDG-avid esophageal tumor; CRE-II (t=3) PET-CT image showing radiologically complete response to nCRT; PET-CT image at 21 months after baseline (t=5) showing FDG-avid lesion suspect for metastasis within the semimembranosus muscle; PET-CT image 26 months after baseline (t=7) of semimembranosus lesion showing decreased intensity after induction chemotherapy; PET-CT image 30 months after baseline (t=9) showing FDG-avid lesions suspect for metastasis within the adductor muscles of the right lower extremity, bilaterally in the pelvic acetabulum and in the transverse process of a thoracic vertebra. VAF, variant allele frequency; cfDNA, cell-free deoxyribonucleic acid; WBCs, white blood cells; PET-CT, positron emission tomography-computed tomography; nCRT, neoadjuvant chemoradiotherapy; FDG, 18-fluoride-fluorodeoxyglucose; CRE-II clinical response evaluation-II.
NGS of white blood cell DNA obtained at 30 months (t=9) and 33 months after baseline (t=10) revealed the presence of the TP53 c.646G>A; p.V216M mutation (VAF 0.77%), which was identical to the mutation consistently identified in previous plasma genotyping. Furthermore, a TP53 c.659A>G; p.Y220C mutation was identified in two independent white blood cell samples (Table 1). Analysis of cfDNA samples by droplet digital PCR (ddPCR) confirmed earlier results from NGS with Oncomine colon panel (Table 2). At the time of writing this report the patient has passed away, 46 months after initial diagnosis of esophageal cancer.
Table 2. Characteristics of ddPCR outcomes of cfDNA per point in time.
| T | Moment of sample | Analysis | DNA concentration (ng/μL) | Total DNA (ng) | Allele name | Wildtype concentration (copy/μL) | Variant concentration (copy/μL) | Variant percentage | Remark |
|---|---|---|---|---|---|---|---|---|---|
| t=0 (baseline) | Baseline cfDNA | ddPCR | 2.14 | 25.70 | c.524G>A; p.R175H | 198.5 | 0 | 0 | Negative |
| t=2 (baseline + 14 w) | CRE-I | ddPCR | 2.07 | 39.12 | c.524G>A; p.R175H | 499.3 | 0 | 0 | Negative |
| t=3 (baseline + 21 w) | CRE-II | ddPCR | 2.58 | 38.70 | c.524G>A; p.R175H | 425.8 | 0 | 0 | Negative |
| t=8 (baseline + 27 m) | FU I | ddPCR | 1.76 | 3.52 | c.524G>A; p.R175H | 51.0 | 0 | 0 | Negative |
| t=9 (baseline + 30 m) | FU II | ddPCR | 2.76 | 16.56 | c.524G>A; p.R175H | 217.0 | 0 | 0 | Negative |
| WBCs-FU II | ddPCR | 3.82 | 38.20 | c.524G>A; p.R175H | 508.0 | 0 | 0 | Negative | |
| t=10 (baseline + 33 m) | FU III | ddPCR | 1.02 | 19.28 | c.524G>A; p.R175H | 234.4 | 0.74 | 0.31 | Positive |
| WBCs-FU III | ddPCR | 120 | 30.00 | c.524G>A; p.R175H | 282.0 | 0 | 0 | Negative |
TP53 p.R175H (chr17:007578406; c.524G>A). cfDNA, cell free deoxynucleic acid; CRE, clinical response evaluation; WBCs, white blood cells; FU, follow-up.
Analyses
Blood samples drawn at t=0, t=2 and t=3 were collected using EDTA tubes. Plasma was separated, isolated and frozen within one hour after collection in order to provide stable cfDNA levels. All other samples were collected using CellSave tubes in which cfDNA levels are stable up to 96 hours after collection and were isolated and frozen within 48 hours (19). Isolation of cfDNA from plasma samples was performed with QIAamp Circulating Nucleic Acid kits (Qiagen, Hilden, Germany). DNA from white blood cells was extracted by a nucleospin kit. Concentrations of cfDNA were quantified using Quant-iT™ dsDNA High-Sensitivity Assay Kit (Thermo Fisher Scientific, Waltham, Massachusetts, USA) and Qubit® 2.0 fluorometer (Thermo Fisher Scientific). Sequencing was performed on the Ion Torrent S5 GeneStudio Prime system (Thermo Fisher Scientific). Sequencing of tumor tissue DNA was completed using a dedicated custom-made cancer NGS panel for detection of TP53 aberrations. The entire coding region and the exon-intron boundaries of TP53 are covered by this panel. TP53 mutations in cfDNA of plasma samples were analyzed with cfDNA NGS kits with unique molecular identifiers (UMI: single molecule barcoding, Oncomine cfDNA assays, Thermo Fisher Scientific). Mean read coverage depths were >1,000 reads and >20,000 for diagnostic tissue and cfDNA NGS assays, respectively. In the cfDNA analysis identified variants were considered reliable when detected with at least 3 independent molecules (UMIs). To confirm our findings, tumor tissue DNA was also analyzed with Oncomine cfDNA assays and cfDNA in plasma samples were also analyzed using digital polymerase chain reaction (Crystal digital PCR, Stilla Technology, Villejuif, France).
Discussion
In this case report we describe the long-term follow-up of an esophageal carcinoma patient with false-positive plasma genotyping due to clonal hematopoiesis. In our patient a TP53 c.524G>A; p.R175H mutation was identified in the primary tumor via endoscopic biopsy. This is a well-described pathogenic variant associated with a variety of malignancies (20). Mutant p53 has been demonstrated to result in enhanced integrin and epidermal growth factor receptor (EGFR) trafficking causing increased cell scattering and invasion (21). Frequent plasma-derived cfDNA genotyping was performed aiming to find an association between disease activity and traceability of TP53 c.524G>A; p.R175H in plasma cfDNA. This mutation became detectable only after distant metastases had already been confirmed by clinical symptoms and standard imaging- and biopsy techniques. Surprisingly, plasma genotyping in our case consistently identified a TP53 c.646G>A; p.V216M mutation that was not found in the tumor tissues. TP53 c.646G>A; p.V216M variants have been associated with various malignancies. Although pathogenicity has as yet not been clearly established, it has been mentioned as a recurrent clonal hematopoietic variant (22,23). The VAF of the p.V216M variant was constant between baseline and 21 weeks while the patient developed a clinically complete response, which would suggest a clonal hematopoietic origin. Of interest, the VAF of TP53 c.646G>A; p.V216M doubled at 27 months and increased further at 30 months. This would have made it hard to ascribe it to clonal hematopoiesis if leukocyte analysis had not been performed. Eventual NGS of white blood cells identified this latter mutation in normal appearing white blood cells of the patient. Analysis of circulating tumor cells was performed which resulted in the finding of no epithelial cell adhesion molecule (EpCAM)-positive cells, rendering contamination of white blood cells by circulating tumor cells improbable and confirming the TP53 c.646G>A; p.V216M mutation was of clonal hematopoietic origin. In the present case-report we focused on TP53 mutation analysis.
Our case illustrates a number of challenges currently associated with cfDNA diagnostics in esophageal carcinoma. The tumor suppressor gene 53 (TP53) is one of the most frequently mutated genes in human cancers and codes for the p53 protein which has been referred to as “the guardian of the genome” (24,25). Esophageal adenocarcinoma displays an exceptionally high rate of TP53 mutations that reaches up to 70% (26). TP53, apart from frequently being altered in malignancy, is 1 out of 6 frequently mutated genes in clonal hematopoiesis as well (14). Over the past few years there has been an increased attention for clonal hematopoiesis. Acquisition of somatic mutations in hematopoietic cells results in abnormal clonal expansion that is associated with an increased risk of myeloid and lymphoid malignancies. Presence of clonal hematopoiesis in other types of malignancy is associated with worse outcomes and an increased risk of all-cause mortality (12). Clonal hematopoiesis is nevertheless most frequently a phenomenon of indeterminate malignant potential and can be found in 1 out of 10 people of 70 years and older and is present in a quarter of patients with any type of malignancy (13,14). This means that if cfDNA analysis in the future possibly becomes a diagnostic option in esophageal cancer, clinicians will have to consider the possibility of finding mutations that originate from clonal hematopoiesis rather than from a malignant lesion. This risk of false-positive test results can be readily averted by screening for clonal hematopoiesis in patients who have cfDNA mutations that cannot be matched with mutations in tumor biopsies.
NGS of cfDNA isolated from peripheral blood is a relatively new diagnostic modality that is already employed in a variety of cancers and has fueled great expectations of broad applicability. Application of this technique in active surveillance of esophageal carcinoma potentially replaces modalities such as endoscopy and PET-CT. This could have important benefits with regard to cost-effectiveness, invasiveness, and patient compliance. cfDNA analysis does however not yet capitalize on its potential value in the case we describe. Failure to detect TP53 c.524G>A; p.R175H in cfDNA is most likely due to the amount of ctDNA in the cfDNA fraction being below the limit of detection for NGS and ddPCR analyses. Recent data from esophageal carcinoma indicate that analysis using Cancer Personalized Profiling by deep sequencing (CAPP-seq, analysis of multiple tumor-specific DNA variants) can detect ctDNA down to 0.07%, well before tumor progression can be radiologically detected (27). Our case does however demonstrate the possibility of finding mutations originating from clonal hematopoiesis. We recommend correlation of mutations in cfDNA with mutations in tumor biopsies. If mutations in cfDNA cannot be matched with tumor mutations, performing NGS of white blood cells can confirm clonal hematopoiesis and preclude false-positive outcomes.
Supplementary
The article’s supplementary files as
Acknowledgments
The authors thank Bo Noordman, Sjoerd Lagarde, Bianca Mostert for their contributions in obtainment of tissue- and blood samples and Ronald van Marion for his help with the NGS analyses. The authors would also like to acknowledge the contribution of Katharina Biermann in pathological analysis of the resection specimens.
Funding: None.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee(s) and with the Helsinki Declaration (as revised in 2013). Written informed consent was obtained from the patient to perform comprehensive genomic testing and to publish his information.
Footnotes
Reporting Checklist: The authors have completed the CARE reporting checklist. Available at https://dx.doi.org/10.21037/atm-21-525
Peer Review File: Available at https://dx.doi.org/10.21037/atm-21-525
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://dx.doi.org/10.21037/atm-21-525). The authors have no conflicts of interest to declare.
References
- 1.Noordman BJ, Wijnhoven BPL, Lagarde SM, et al. Neoadjuvant chemoradiotherapy plus surgery versus active surveillance for oesophageal cancer: a stepped-wedge cluster randomised trial. BMC Cancer 2018;18:142. 10.1186/s12885-018-4034-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.van der Wilk BJ, Noordman BJ, Neijenhuis LKA, et al. Active Surveillance Versus Immediate Surgery in Clinically Complete Responders After Neoadjuvant Chemoradiotherapy for Esophageal Cancer: A Multicenter Propensity Matched Study. Ann Surg 2019. [Epub ahead of print]. doi: . 10.1097/SLA.0000000000003636 [DOI] [PubMed] [Google Scholar]
- 3.Castoro C, Scarpa M, Cagol M, et al. Complete clinical response after neoadjuvant chemoradiotherapy for squamous cell cancer of the thoracic oesophagus: is surgery always necessary? J Gastrointest Surg 2013;17:1375-81. 10.1007/s11605-013-2269-3 [DOI] [PubMed] [Google Scholar]
- 4.Chirieac LR, Swisher SG, Ajani JA, et al. Posttherapy pathologic stage predicts survival in patients with esophageal carcinoma receiving preoperative chemoradiation. Cancer 2005;103:1347-55. 10.1002/cncr.20916 [DOI] [PubMed] [Google Scholar]
- 5.Noordman BJ, Spaander MCW, Valkema R, et al. Detection of residual disease after neoadjuvant chemoradiotherapy for oesophageal cancer (preSANO): a prospective multicentre, diagnostic cohort study. Lancet Oncol 2018;19:965-74. 10.1016/S1470-2045(18)30201-8 [DOI] [PubMed] [Google Scholar]
- 6.Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 2011;11:426-37. 10.1038/nrc3066 [DOI] [PubMed] [Google Scholar]
- 7.Creemers A, Krausz S, Strijker M, et al. Clinical value of ctDNA in upper-GI cancers: A systematic review and meta-analysis. Biochim Biophys Acta Rev Cancer 2017;1868:394-403. 10.1016/j.bbcan.2017.08.002 [DOI] [PubMed] [Google Scholar]
- 8.van der Wilk BJ, Eyck BM, Spaander MCW, et al. Towards an Organ-Sparing Approach for Locally Advanced Esophageal Cancer. Dig Surg 2019;36:462-9. 10.1159/000493435 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chin RI, Chen K, Usmani A, et al. Detection of Solid Tumor Molecular Residual Disease (MRD) Using Circulating Tumor DNA (ctDNA). Mol Diagn Ther 2019;23:311-31. 10.1007/s40291-019-00390-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ignatiadis M, Lee M, Jeffrey SS. Circulating Tumor Cells and Circulating Tumor DNA: Challenges and Opportunities on the Path to Clinical Utility. Clin Cancer Res 2015;21:4786-800. 10.1158/1078-0432.CCR-14-1190 [DOI] [PubMed] [Google Scholar]
- 11.Hu Y, Ulrich BC, Supplee J, et al. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin Cancer Res 2018;24:4437-43. 10.1158/1078-0432.CCR-18-0143 [DOI] [PubMed] [Google Scholar]
- 12.Steensma DP, Bejar R, Jaiswal S, et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 2015;126:9-16. 10.1182/blood-2015-03-631747 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Jaiswal S, Fontanillas P, Flannick J, et al. Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med 2014;371:2488-98. 10.1056/NEJMoa1408617 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Coombs CC, Zehir A, Devlin SM, et al. Therapy-Related Clonal Hematopoiesis in Patients with Non-hematologic Cancers Is Common and Associated with Adverse Clinical Outcomes. Cell Stem Cell 2017;21:374-382.e4. 10.1016/j.stem.2017.07.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Razavi P, Li BT, Brown DN, et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med 2019;25:1928-37. 10.1038/s41591-019-0652-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Xia L, Li Z, Zhou B, et al. Statistical analysis of mutant allele frequency level of circulating cell-free DNA and blood cells in healthy individuals. Sci Rep 2017;7:7526. 10.1038/s41598-017-06106-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gagnier JJ, Kienle G, Altman DG, et al. The CARE guidelines: consensus-based clinical case reporting guideline development. J Med Case Rep 2013;7:223. 10.1186/1752-1947-7-223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.van Hagen P, Hulshof MC, van Lanschot JJ, et al. Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med 2012;366:2074-84. 10.1056/NEJMoa1112088 [DOI] [PubMed] [Google Scholar]
- 19.van Dessel LF, Beije N, Helmijr JC, et al. Application of circulating tumor DNA in prospective clinical oncology trials - standardization of preanalytical conditions. Mol Oncol 2017;11:295-304. 10.1002/1878-0261.12037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.National Center for Biotechnology Information. ClinVar; [VCV000012374.13]. Available online: https://www.ncbi.nlm.nih.gov/clinvar/variation/VCV000012374.13 (accessed March 30, 2021).
- 21.Grugan KD, Vega ME, Wong GS, et al. A common p53 mutation (R175H) activates c-Met receptor tyrosine kinase to enhance tumor cell invasion. Cancer Biol Ther 2013;14:853-9. 10.4161/cbt.25406 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.National Center for Biotechnology Information. ClinVar; [VCV000182965.10]. Available online: https://www.ncbi.nlm.nih.gov/clinvar/variation/VCV000182965.10 (accessed March 30, 2021).
- 23.Yaung SJ, Fuhlbrück F, Peterson M, et al. Clonal Hematopoiesis in Late-Stage Non–Small-Cell Lung Cancer and Its Impact on Targeted Panel Next-Generation Sequencing. JCO Precis Oncol 2020:1271-9. 10.1200/PO.20.00046 [DOI] [PubMed] [Google Scholar]
- 24.Petitjean A, Mathe E, Kato S, et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat 2007;28:622-9. 10.1002/humu.20495 [DOI] [PubMed] [Google Scholar]
- 25.Lane DP. Cancer. p53, guardian of the genome. Nature 1992;358:15-6. 10.1038/358015a0 [DOI] [PubMed] [Google Scholar]
- 26.Dulak AM, Stojanov P, Peng S, et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat Genet 2013;45:478-86. 10.1038/ng.2591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Azad TD, Chaudhuri AA, Fang P, et al. Circulating Tumor DNA Analysis for Detection of Minimal Residual Disease After Chemoradiotherapy for Localized Esophageal Cancer. Gastroenterology 2020;158:494-505.e6. 10.1053/j.gastro.2019.10.039 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The article’s supplementary files as