Abstract
Primary transcripts encoding the MADS box superfamily of proteins, such as MEF2 in animals and ZEMa in plants, are alternatively spliced, producing several isoformic species. We show here that murine serum response factor (SRF) primary RNA transcripts are alternatively spliced at the fifth exon, deleting approximately one-third of the C-terminal activation domain. Among the different muscle types examined, visceral smooth muscles have a very low ratio of SRFΔ5 to SRF. Increased levels of SRFΔ5 correlates well with reduced smooth muscle contractile gene activity within the elastic aortic arch, suggesting important biological roles for differential expression of SRFΔ5 variant relative to wild-type SRF. SRFΔ5 forms DNA binding-competent homodimers and heterodimers. SRFΔ5 acts as a naturally occurring dominant negative regulatory mutant that blocks SRF-dependent skeletal α-actin, cardiac α-actin, smooth α-actin, SM22α, and SRF promoter-luciferase reporter activities. Expression of SRFΔ5 interferes with differentiation of myogenic C2C12 cells and the appearance of skeletal α-actin and myogenin mRNAs. SRFΔ5 repressed the serum-induced activity of the c-fos serum response element. SRFΔ5 fused to the yeast Gal4 DNA binding domain displayed low transcriptional activity, which was complemented by overexpression of the coactivator ATF6. These results indicate that the absence of exon 5 might be bypassed through recruitment of transcription factors that interact with extra-exon 5 regions in the transcriptional activating domain. The novel alternatively spliced isoform of SRF, SRFΔ5, may play an important regulatory role in modulating SRF-dependent gene expression.
Alternative splicing is a commonly used molecular strategy for creating diverse gene products from a single genetic locus in most eucaryotic cells. The modular organization of transcription factor genes, through exon-encoded structural domains, may be conducive for forming a variety of alternatively spliced isoforms that affect DNA binding avidity and specificity, transactivation, subcellular localization, responsiveness to signaling pathways, and developmental regulation (reviewed in reference 31). For example, alternative splicing within the DNA binding domain of Pax-6 (9) and Wilm’s tumor-associated protein 1 (5, 21) alters their DNA binding specificities. Alternative splicing of exons encoding the transactivating domains in the paired family proteins Pax-3 (59), Pax-8 (24), and Pax-9 (39), the POU homeodomain family proteins Pit-1 (37), Oct-1 (8), Oct-2 (1), and Brn-3a (38), and the zinc finger transcription factor GATA-5 (32) results in isoforms possessing activation domains with different potencies. Splicing out of select exons from the activation domain as in AML1a (56), Oct-2 (29), and CREB family proteins CREB (reviewed in reference 11), Drosophila CREB/CREM (61), and CREM (12) produces isoforms with dominant negative activity. Similarly, several splice variants containing only the DNA binding domain act as dominant negative isoforms presumably by heterodimerizing with the wild-type isoforms and competing with the wild-type isoforms for DNA binding. Thus, additional genetic complexity might be further imposed by altering regulatory processes by the expression of alternatively spliced transcription factor isoforms.
Serum response factor (SRF) is a member of an ancient DNA binding protein superfamily whose evolutionarily divergent relatives shared a highly conserved DNA binding/dimerization domain of 90 amino acids, termed the MADS box (reviewed in reference 51). The complex and novel stratified organization of the MADS box domain recently resolved by X-ray crystallography (43) was probably assembled before the divergence of plants and animals, since identical MADS box structures were present in yeast transcription factors MCM1 and ARG80, a large number of homeotic-like plant proteins, and vertebrate SRFs (reviewed in references 46 and 52). All of these transcription factors, through their common MADS boxes, bind to virtually the same DNA sequences (reviewed in references 46 and 52) and interact with similar kinds of coaccessory regulatory factors (reviewed in references 52 and 58). SRF-related factors such as RSRF, also described as MEF2, contain the MADS box and the adjacent MEF2 box (44) but prefers binding to divergent MEF2 sites. Dissection of the human SRF revealed a modular organization of structural and functional domains, such as the central MADS box domain, the N-terminal domain, containing serine 103 phosphorylation sites that facilitate SRF DNA binding activity, and a putative transcriptional inhibitory domain (22, 47), while the C-terminal activation domain binds a diversity of protein factors such as the RAP74 subunit of TFIIF (23), ATF6 (64), and viral Tax1 (13) proteins.
MADS box proteins play a central role for a variety of regulatory activities. The mammalian MADS box proteins, SRF and MEF2, regulate the SRF-inducible genes and muscle-specific gene expression (19; reviewed in references 40, 45, 52, and 58). The regulation of muscle-specific gene expression appears to be mediated by the interaction of the SRF MADS box with striated muscle tissue-restricted factors such as the basic helix-loop-helix protein MyoD (18, 50), homeodomain protein Nkx-2.5 (6), and zinc finger protein GATA-4 (4). In addition, high levels of embryonic SRF expression coincide with the expression of cardiac, skeletal, and smooth muscle α-actins (3, 7), noted as early markers for terminal striated and smooth muscle cell (SMC) differentiation (49). We reported earlier that the skeletal muscle-enriched expression of SRF is under the control of a positive autoregulatory loop. Dominant negative mutants of SRF with point mutations in the DNA binding domain (SRFpm1) or truncation of the C-terminal activation domain (SRFΔC) repressed SRF promoter activity (3) and myogenesis. Vandromme et al. (60) showed that neutralizing SRF activity by microinjection of SRF antibodies, and blocking SRF gene expression through application of SRF antisense oligonucleotides (53) in C2C12 myoblasts, prevents terminal differentiation, thus underscoring the obligate role for SRF in myogenic differentiation and muscle-specific gene expression. These observations were confirmed by the recent analysis of SRF null mutants, which revealed an absolute dependence for SRF for the formation of embryonic myogenic mesoderm (2).
An array of MEF2 proteins are generated by alternative splicing (27, 33, 34, 35, 62). Furthermore, some of these splicing events are cell type restricted, and no specific function for these isoforms has been ascribed so far. We asked whether SRF, another closely related MADS box family member, is also alternatively spliced, and if it is, whether SRF splice variants have biological activity. We show here that murine SRF primary RNA transcripts are alternatively spliced. An SRF isoform with a deletion in the C-terminal activation domain acts as a naturally occurring dominant negative regulatory mutant that interferes with SRF-dependent promoter activity and muscle cell differentiation.
MATERIALS AND METHODS
Recombinant DNA clones.
Luciferase reporter constructs for Gal4-Luc (G5Luc), c-fos serum response element (SRE), cardiac α-actin, skeletal α-actin, SM22α, and −310 SRF promoters and the expression constructs pCGN, pCGNSRF, pCGNSRFΔC, pCGNATF6, GalDB (Gal4 DNA binding domain), and GalSRF 266-508 have been described elsewhere (3, 6, 22, 28). The smooth α-actin promoter-luciferase reporter was constructed by cloning a PCR-amplified −191 to +46 fragment between NheI and XhoI sites of pGL3 basic vector. To construct pCGNSRFΔ5, pCGNSRF was partially digested with ApalI followed by digestion to completion with BamHI. ApalI cuts SRF once within exon 4. The large ApalI-BamHI fragment containing the pCGN vector backbone including the β-globin intron and poly(A) signal sequences and SRF sequences from ATG to nucleotide (nt) 1446 was gel purified. Mouse skeletal muscle cDNA was amplified with an upstream primer (1317 to 1337) and a downstream primer overlapping the stop codon which contained an engineered BamHI site. The PCR-amplified product was digested with ApalI and BamHI and cloned into the large ApalI- and BamHI-digested vector fragment. GalSRF 266-508Δ5 was constructed by triple ligation of ApalI- and BamHI-digested PCR product containing the SRFΔ5 fragment (amino acids 350 to 508) described above, the XbaI-ApalI fragment containing SRF amino acids 266 to 350, and the XbaI and BamHI-cut Gal4 expression vector.
RNA isolation.
Total RNA was isolated from adult mouse tissues and cell lines by using Ultra Spec (Biotecx) according to the manufacturer’s recommendations.
RT-PCR.
To analyze SRF RNA processing, total cellular RNA (5 μg) was denatured and reverse transcribed with 15 U of Superscript reverse transcriptase (RT; Bethesda Research Laboratories) in RT-PCR buffer (Stratagene) containing random primers and deoxynucleoside triphosphates. For negative controls, RT was not added. After 1 h of incubation at 42°C, 50 μl of water was added, and the RT was heat inactivated. Five microliters of the cDNA was used for PCR. The 50-μl PCR mixture contained the standard Taq polymerase buffer with 2.5 mM MgCl2, 0.2 mM deoxynucleoside triphosphates, and 100 pmol each of upstream (1317 to 1337) and downstream (1889 to 1909) primers. The reaction conditions were an initial denaturation at 95°C for 10 min, followed by 30 cycles of 1-min denaturation, 1-min annealing at 60°C, and 2-min extension at 72°C. A 5-μl aliquot of the reaction product was fractionated on a 1.5% agarose gel, stained with ethidium bromide, and photographed. A 0.5-μl aliquot of the RT-PCR product was fractionated on a 1.5% agarose gel and blotted simultaneously onto two pieces of a Zeta-Probe GT membrane (Bio-Rad). One blot was hybridized to a random-primed SRF cDNA probe, and the other was hybridized to an end-labeled exon 5-specific 25-mer oligonucleotide. Blots were washed under standard conditions and exposed to X-ray film for 10 to 30 min.
For semiquantitative RT-PCR, 32P-end-labeled primers were used, and the reaction was stopped in the linear range (20 to 25 cycles). The PCR products were resolved on a 5% polyacrylamide gel, dried, and autoradiographed.
Northern blotting.
Twenty micrograms of total RNA was denatured, resolved on a 0.8% formaldehyde-agarose gel, and transferred to a Zeta-Probe GT (Bio-Rad) membrane. The membrane was hybridized to random-primed mouse myogenin probe at 65°C in 0.25 M Na2HPO4 (pH 7.2)–7% sodium dodecyl sulfate (SDS) and washed at the same temperature in 20 mM Na2HPO4–1% SDS. Subsequently, the membrane was stripped and reprobed with a skeletal α-actin 3′ untranslated region probe.
RNase protection assays.
The antisense probe for RNase protection was synthesized from XhoI-linearized plasmid pBSSRF 1558-1842, by using T3 RNA polymerase. Radioactive probe (5 × 104 cpm) was hybridized to 40 μg of total RNA at 50°C overnight in a buffer containing 80% formamide, 10 mM sodium citrate (pH 6.4), 300 mM sodium acetate (pH 6.4), and 1 mM EDTA. Unhybridized probe and RNA were digested by adding a mixture of RNases A (1 μg) and T1 (20 U) for each sample and incubating the mixtures for 1 h at 37°C. RNase-resistant double-stranded RNA was resolved on a 5% polyacrylamide–8 M urea gel. The gel was dried and autoradiographed.
Cell culture and transfections.
NIH 3T3 and CV1 cells were maintained in Dulbecco modified Eagle medium (DMEM) containing 10% neonatal calf serum. 10T1/2 and C2C12 cells were maintained in DMEM containing a mixture of 10% neonatal calf serum and fetal bovine serum. For serum induction of transfected cells, 10T1/2 cells in 60-mm-diameter plates were cotransfected with 200 ng of SRELuc and 200 ng of pCGNSRF, pCGNSRFΔ5, pCGNSRFΔC, or the empty vector pCGN along with 200 ng of internal control plasmid pCMVβGal. Following 48 h of serum starvation, cells were induced by the addition of fetal bovine serum to 20% (final concentration), and cells were harvested 3 h later. CV1 cells in 60-mm-diameter plates were transfected with 1 μg of cardiac α-actin, skeletal α-actin, 3 kbp-Sm22α and −310 SRF promoter-luciferase reporter constructs along with 150 ng of pCGNSRF, pCGNSRFΔ5, and pCGNSRFΔC. To demonstrate the dose-dependent effect of SRFΔ5 on smooth α-actin promoter, 200 ng of smooth α-actin promoter-luciferase construct was cotransfected with 150 ng of SRF and various amounts of SRFΔ5. Cells were harvested 48 h posttransfection, and luciferase activity was measured according to standard methods in a luminometer. When cotransfected with internal control plasmid pCMVβGal, the luciferase values were normalized to β-galactosidase values.
For stable expression of SRFΔ5 and SRFpm1, C2C12 myoblasts in 60-mm-diameter plates were transfected with 250 ng of pSV2Neo and indicated plasmids. A day after transfection, cells were split into 10 100-mm-diameter plates and selected for neomycin resistance. DMEM containing 20% serum and 400 μg of G418 (GIBCO) per ml was changed every 5 days. Neomycin-resistant colonies were pooled after 2 weeks of selection, expanded, and analyzed.
Whole-cell extracts and electrophoretic gel mobility shift assays (EMSAs).
Whole-cell extracts were prepared as described earlier (26). Five micrograms of cell extract was preincubated with 1 μg of poly(dG-dC) at room temperature for 15 min in 20 μl of 1× buffer (50 mM NaCl, 20 mM HEPES-KOH [pH 7.5], 0.1 mM EDTA, 0.5 mM dithiothreitol, 10% glycerol). A 50-fold excess of specific and nonspecific competitors and 0.5 μl of antibodies were included in this reaction. Following the preincubation, 0.02 pmol of end-labeled proximal SRE1 probe from the SRF promoter (3) was added, and the mixtures were incubated at room temperature for a further 15 min. DNA-protein complexes were resolved on a 5% polyacrylamide gel cast and run in 0.5× Tris-borate-EDTA. For the demonstration of SRFΔ5-SRFΔC heterodimers, 2.5 μl of in vitro-translated SRFΔ5 was incubated with 2.5 μg of CV1 cell extract overexpressing SRFΔC in 20 μl of 1× binding buffer.
In vitro transcription and translation and GST pull-down assays.
For in vitro transcription, SRFΔ5 cloned in pBSSKII(+) was linearized with BamHI and transcribed with T3 RNA polymerase. Capped SRFΔ5 and luciferase RNA were translated in rabbit reticulocyte lysate (RRL; Promega) in the presence of [35S]methionine according to the manufacturer’s recommendations; 5-μl aliquots of the programmed lysates were incubated with either glutathione S-transferase (GST) or GST-SRF immobilized on beads. GST pull-down assays were performed as described earlier (6).
Western blot analysis.
NIH 3T3 cell extracts for Western blotting were prepared by extracting cells in phosphate-buffered saline containing 1% Nonidet P-40, 0.5% sodium deoxycholate, 0.1% SDS, and protease inhibitors. Soluble protein (20 μg) was resolved on an SDS–10% polyacrylamide gel, transferred to polyvinylidene difluoride membranes, and probed with SRF immune serum (a gift from R. Prywes). Whole-cell extracts prepared from 4-day embryoid bodies (EBs) were probed with a carboxy-terminus peptide epitope antibody (Santa Cruz Biotechnology) directed against amino acids 486 to 505 of human SRF. After visualization of the signal by enhanced chemiluminescence the blot was stripped and reprobed with an exon 5 epitope-specific antibody.
RESULTS
Alternatively splicing of SRF exon 5.
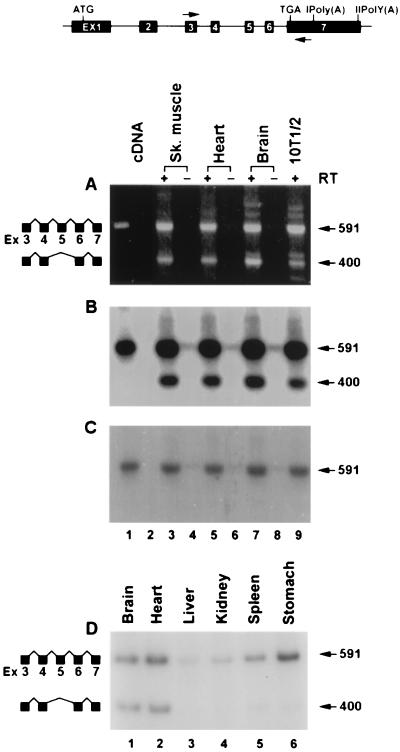
Members of the MADS box family of proteins such as MEF2 generate isoforms by alternative splicing of the primary transcripts. Some of these alternatively spliced isoforms are tissue type restricted. We asked if the primary transcript of SRF also undergoes alternative splicing. Analysis of adult mouse skeletal muscle RNA by RT-PCR using exon 3-specific (nt 1317 to 1337) and exon 7-specific (nt 1889 to 1909) primers detected two major amplified products 592 and 401 nt in length, (Fig. 1A). Based on the RNA chain lengths, the 592-bp band corresponds to constitutively spliced SRF RNA whereas the 401-bp band conforms to an alternatively spliced form of SRF RNA. Since the length of exon 5 is 192 bp, it is likely that the smaller amplified band of 401 bp arose from removal of exon 5. To confirm this observation, Southern blot analysis was performed on the RT-PCR-amplified products. These products, resolved on a 1.5% agarose gel, were transferred bidirectionally onto two membranes and probed with an SRF cDNA probe and an exon 5-specific oligonucleotide probe. As shown in Fig. 1B, both 592- and 401-bp DNA species hybridized to the SRF cDNA probe, indicating that both DNA species contain SRF sequences. However, the exon 5-specific oligonucleotide hybridized only to the 592-bp species, not the 401-bp species, indicating that the smaller amplified DNA species is devoid of exon 5 sequences (Fig. 1C). We confirmed that the 401-bp species lacked exon 5 by nucleic acid sequencing. The alternative splicing of exon 4 to exon 6 utilized the authentic splice donor and acceptor sites. The exon 5 skipping did not alter the reading frame but resulted in substitution of a single amino acid, glycine for valine at the splice junction. SRFΔ5 transcripts were expressed abundantly in brain, cardiac, and skeletal muscle and at very low levels in kidney, liver, spleen, and stomach (Fig. 1D).
FIG. 1.
Alternative splicing of SRF RNA removes exon 5. The exon-intron organization of the SRF genome and positions of the primers used for PCR are shown at the top. (A) Ethidium bromide-stained agarose gel showing the RT-PCR products from skeletal muscle (lanes 3 and 4), heart (lanes 5 and 6), brain (lane 7 and 8), and 10T1/2 cells (lane 9). Cloned mSRF cDNA was used as the template in lane 1. Lane 2 contained no input cDNA. RT was omitted during cDNA synthesis for lanes 4, 6, and 8. Constitutively spliced and alternatively spliced forms are diagrammatically represented on the left, and sizes of the bands are shown in base pairs on the right. Southern blots of the gel probed with the mSRF cDNA and exon 5 oligonucleotide are shown in panels B and C, respectively. End-labeled primers were used for PCR shown in panel D.
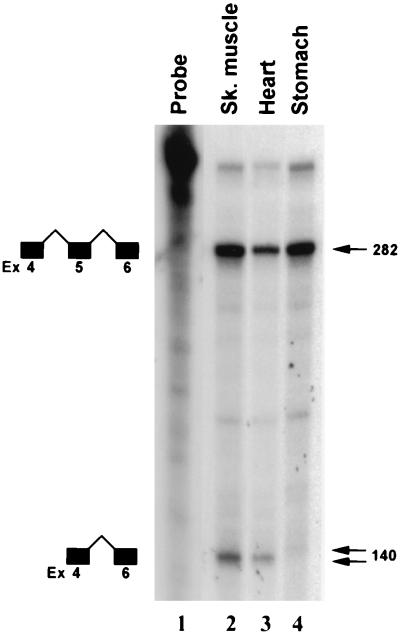
Recently, we reported that SRF is expressed predominantly in mouse brain, skeletal, cardiac, and smooth muscle tissues (3). The relative proportions of the constitutively and alternatively spliced SRF RNA species in three different types of muscle tissues were assessed by RNase protection assays. The antisense SRF probe consisted of a cDNA fragment spanning exons 5 through 7. Total RNA prepared from skeletal, cardiac, and visceral smooth muscle tissues protected 284- and 144-nt fragments, representing the constitutively and alternatively spliced RNA species, respectively (Fig. 2). The relative proportion of the constitutively spliced RNA to alternatively spliced RNA varied for the three muscle types examined, as shown in Fig. 2. Stomach, representing visceral smooth muscle, expressed the highest proportion of constitutively spliced RNA.
FIG. 2.
Tissue-restricted expression of alternatively spliced SRFΔ5 RNA, determined by RNase protection analysis of total cellular RNA isolated from skeletal muscle (lane 2), heart (lane 3), and stomach (lane 4) with the antisense SRF probe. The undigested probe was run in lane 1. Protected unspliced and spliced products are diagrammatically represented to the left; sizes of the protected fragments are indicated in base pairs on the right.
SRFΔ5 is expressed in a rostrocaudal gradient in the neonatal rat pup aorta.
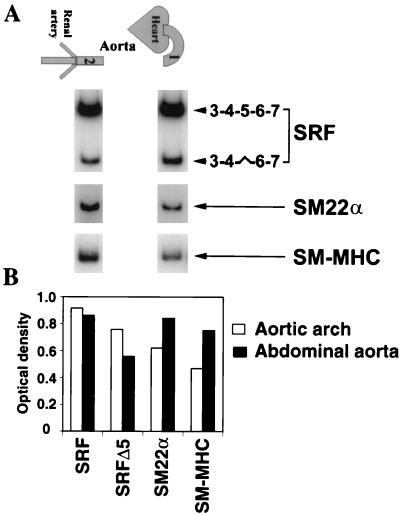
We asked whether SRFΔ5 was expressed in aortic SMCs and whether the cardiac neural crest derived aortic arch SMCs differed from mesoderm-derived abdominal aortic SMCs in the expression of SRFΔ5. As shown in Fig. 3, SRFΔ5 was expressed in a rostrocaudal gradient, with the highest level of SRFΔ5 in aortic arch and the lowest level in the abdominal aorta. We further analyzed the expression and distribution of SMC markers such as SM-MHC, SM22α, and SRFΔ5 in the same aortic segments. Both SM-MHC and SM22α were expressed in a gradient along the aorta, with the lowest level of expression in the elastic aortic arch and the highest in the contractile abdominal aorta (Fig. 3). Conversely, the highest levels of SRFΔ5 was in the segments of the aortic arch and was reduced in the segments of abdominal aorta. Thus, there appears to be an inverse relationship between the expression of these SRF-regulated genes and levels of SRFΔ5 (Fig. 3).
FIG. 3.
Inverse gradient of expression of SRFΔ5, SM22α, and SM-MHC along the aorta. (A) Total RNA isolated from the aortic arch (lane 1) and abdominal aorta (lane 2) analyzed by semiquantitative RT-PCR with the indicated end-labeled primers. SRF was amplified for 25 cycles, and SM22α and SM-MHC were amplified for 20 cycles. The linearity of amplification was confirmed by harvesting the amplified products at 20, 23, 25, 28, and 30 cycles. Constitutively and alternatively spliced isoforms of SRF are diagrammatically represented to the right. (B) Relative optical densities of bands.
Identification of cellular SRFΔ5-expressed protein.
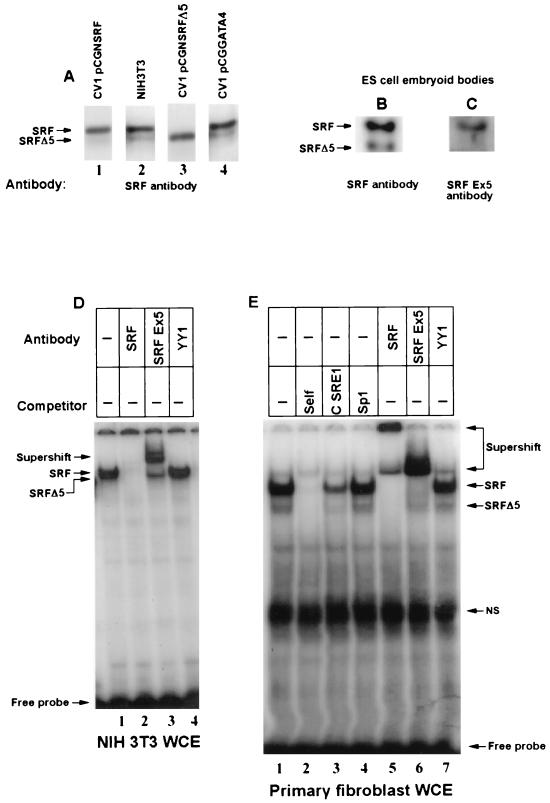
We asked whether the alternatively spliced SRF transcript was translated into cellular protein. Protein extracts from SRF, SRFΔ5, and GATA-4 expression vector-transfected CV1 fibroblasts, which served as ectopically expressed internal standards, were mixed with NIH 3T3 cell extracts, electrophoresed, and then assayed by Western blotting with anti-SRF antibodies. As shown in Fig. 4A (lane 2), SRF antiserum reacted with 67- and 57-kDa proteins. The 67-kDa band was more intense in the extracts spiked with SRF-enriched cell extract (Fig. 4A, lane 1). Similarly, the 57-kDa band was more intense when the extract was mixed with the SRFΔ5-enriched cell extract (Fig. 4A, lane 3). The GATA-4 containing extract did not markedly alter the anti-SRF reactivity (Fig. 4A, lane 4). We also evaluated soluble extracts prepared from day 4 EBs with a C-terminal epitope (common to both SRF and SRFΔ5) and the exon 5 epitope-specific peptide antibodies. The C-terminal SRF antibody reacted with two proteins of 67 and 57 kDa. However, stripping and reprobing of this blot with the exon 5-specific antibody resulted in immunoreactivity of only the 67-kDa protein (Fig. 4B and C). Thus, based on several criteria including predicted size, comigration with exogenously expressed SRF and SRFΔ5 proteins, and immune specificity, the 67- and 57-kDa proteins were identified as the full-length and alternatively spliced SRFΔ5 protein isoforms.
FIG. 4.
SRFΔ5 protein is expressed in NIH 3T3 cells and several other mouse cell lines. (A to C) Western blot analysis of SRF and SRFΔ5 proteins. Protein extracts (20 μg) prepared from NIH 3T3 cells and day 4 EBs were used for Western analysis. NIH 3T3 cell extract (lane 2) was mixed with CV1 whole-cell extract overexpressing SRF (lane 1), SRFΔ5 (lane 3), and GATA-4 (lane 4) and probed with the polyclonal immune serum raised against bacterially expressed SRF protein. Lanes 2 and 4 were exposed three to four times longer than lanes 1 and 3. In panel B, the C-terminal epitope-specific antibody (Santa Cruz) was used. This blot was stripped and subsequently reacted with an exon 5 epitope-specific immune serum (C). Expression of SRFΔ5 proteins in NIH 3T3 cells (D) and mouse embryonic fibroblasts (E) was demonstrated by EMSA. Whole-cell extracts (WCE; 5 μg) were preincubated with a 50-fold excess of the indicated specific (self and cardiac α-actin SRE1) and nonspecific (Sp1) competitors and 0.5 μl of polyclonal immune serum. Annealed SRE1 oligonucleotide from the SRF promoter was the probe. Positions of SRF, SRFΔ5, and supershifted and nonspecific (NS) complexes and of the free probe are indicated. The gel in panel E was run for a longer time to resolve the SRF and SRFΔ5 complexes.
EMSAs were used to confirm the presence of two SRF protein isoforms and to evaluate their DNA binding activities. Incubation of whole-cell extracts prepared from NIH 3T3 cells, CV1 cells, and primary mouse embryo fibroblasts with the SRE probe resulted in two EMSA complexes (Fig. 4D and E), both of which were competed by a 50-fold molar excess of unlabeled SREs but not by Sp1 sites. Both SRF protein-DNA complexes were either supershifted and or abolished by SRF antiserum raised against the full-length SRF (Fig. 4D and E), indicating the presence of SRF in these complexes. However, the exon 5 epitope-specific antibody supershifted the slower-migrating but not the faster-migrating SRF-DNA complex, suggesting that only the former contains exon 5-encoded SRF sequences.
SRFΔ5 protein binds DNA independently and heterodimerizes with other SRF species.
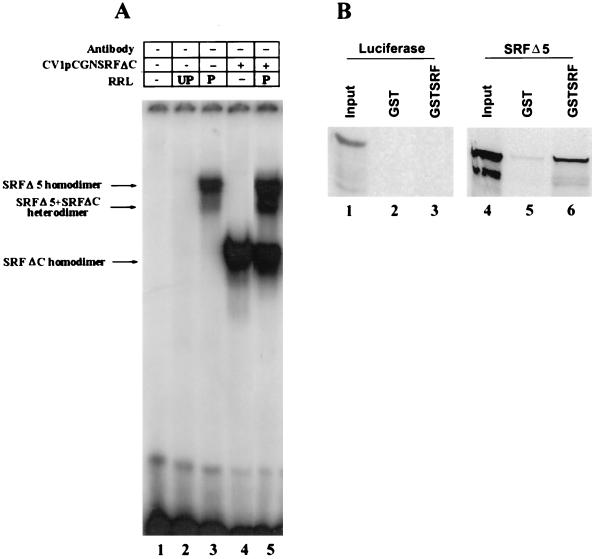
Although the SRF DNA binding and dimerization domains overlap the MADS box, sequences outside the MADS box have been shown to influence the protein’s DNA binding activities. We wanted to determine whether removal of exon 5 (amino acids 384 to 448) perturbs SRFΔ5 DNA binding and dimerization activities. An EMSA with in vitro-translated SRFΔ5 bound the SRE probe with high affinity, as shown in Fig. 5A, indicating that SRFΔ5 can bind DNA as a homodimer. Because we could not resolve heterodimers formed between wild-type SRF and SRFΔ5, we mixed SRFΔ5 with truncated SRFΔC. SRFΔC was recruited into a novel complex displaying an intermediate mobility, demonstrating the capability of SRFΔ5 to heterodimerize with other SRF species (Fig. 5A). To directly assess heterodimerization of SRFΔ5 with SRF, we used GST pull-down assays. SRFΔ5 and luciferase translated in vitro in the presence of 35S-labeled methionine were incubated with purified GST or GST-SRF immobilized on glutathione-linked beads. After extensive washing, SRFΔ5 but not luciferase was retained by the immobilized GST-SRF, demonstrating that SRFΔ5 can heterodimerize with SRF (Fig. 5B); in control assays, we detected only background levels of SRFΔ5 binding to glutathione-linked beads.
FIG. 5.
SRFΔ5 forms DNA binding-competent homodimers and heterodimers. (A) The EMSA conditions were as described for Fig. 4D and E. The binding reaction mixture contained unprogrammed (UP) RRL (lane 2), 2.5 μl of in vitro-translated SRFΔ5 in programmed (P) RRL (lanes 3 and 5), and 2.5 μg of CV1 whole-cell extract overexpressing SRFΔC (lanes 4 and 5). In vitro-translated SRFΔ5 was preincubated with CV1 whole-cell extract overexpressing SRFΔC in lane 5. SRFΔ5 and SRFΔC homodimers and SRFΔ5-SRFΔC heterodimers are indicated by arrows to the left. (B) SRFΔ5 heterodimerizes with SRF independent of DNA binding. [35S]methionine-labeled in vitro-translated luciferase and SRFΔ5 were incubated with GST (lanes 2 and 5) or GST-SRF (lanes 3 and 6) immobilized on glutathione beads, washed extensively, and analyzed on an SDS–10% polyacrylamide gel. Lane 1 and 4 contained 10% of the input luciferase and SRFΔ5, respectively.
SRFΔ5 is a potent repressor of SRE-dependent myogenic promoters.
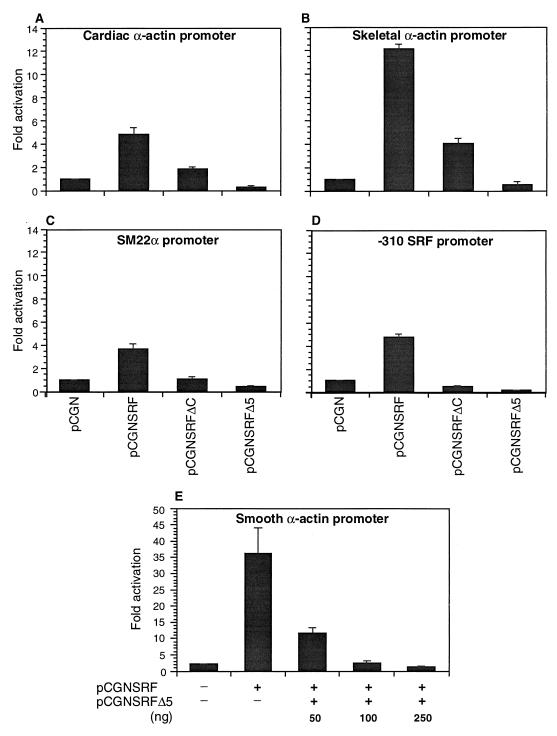
The activation domain of SRF was mapped previously to the C terminus (22, 26, 30). Earlier studies have shown that the mutation of the DNA binding domain or deletion of the entire transactivating domain of SRF confers a dominant negative phenotype on the mutant SRF (3, 7, 15, 16). Because approximately one-third of the transactivating region of SRF is encoded by exon 5, we asked if whether SRFΔ5 also acted as a dominant negative mutant. We evaluated the role of SRFΔ5 on the cardiac, skeletal, smooth α-actin, Sm22α, and SRF gene promoters, because these promoters contain several SREs and promoter activity is highly dependent on SRF binding. Plasmids encoding the wild-type SRF (pCGNSRF), SRFΔ5 (pCGNSRFΔ5), a C-terminal truncated SRF mutant in which amino acids 266 to 508 were deleted (pCGNSRFΔC), and or the empty vector (pCGN) were cotransfected with cardiac and skeletal α-actin, Sm22α, and SRF promoter-reporter constructs into CV1 cells, and reporter activity was assayed 48 h posttransfection. As shown in Fig. 6, all of the promoters were activated severalfold by forced expression of SRF; in contrast to wild-type SRF, SRFΔ5 repressed promoter activity by nearly 75 to 90%. As reported previously, SRFΔC, in which all of the previously mapped transactivation domain was deleted, repressed SRF promoter activity by 50% (Fig. 6D). Surprisingly, this plasmid activated both the skeletal actin and cardiac actin promoters to a lower extent (4.5-fold for the skeletal actin promoter and 2-fold for the cardiac actin promoter). SRFΔ5 repressed the SRF-stimulated activity of smooth α-actin promoter activity in a dose-dependent manner (Fig. 6E). These results indicate that SRFΔ5 acts as a potent repressor of SRF-dependent promoters.
FIG. 6.
SRFΔ5 inhibits SRE-dependent promoter activity. Subconfluent CV1 cells were cotransfected with 1 μg of cardiac α-actin (A), skeletal α-actin (B), SM22α (C) and −310 SRF (D) promoter-luciferase reporter plasmids and 150 ng of expression vectors for SRF (pCGNSRF), SRFΔ5 (pCGNSRFΔ5), C-terminally truncated mutant of SRF (pCGNSRFΔC), or the empty vector pCGN. For panel E, 200 ng of smooth α-actin reporter was cotransfected with 150 ng of pCGNSRF or a combination of pCGNSRF and the indicated amounts of pCGNSRFΔ5. Cells were harvested 48 h posttranscription, and the luciferase activity was measured. Results shown are mean ± standard error of the mean for three duplicate experiments.
SRFΔ5 interferes with myogenic differentiation of C2C12 cells.
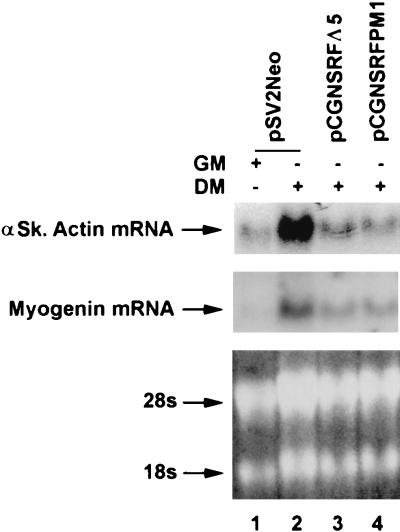
Stably transfected C2C12 cell lines were generated with expression vectors for SRFΔ5 and SRFpm1 along with pSV2neo. Three days following the addition of low-serum-containing differentiation medium wild-type C2C12 cells showed the morphological characteristics of fused, multinucleated, differentiated myotubes, while SRFΔ5 and SRFpm1 cells remained as replicating mononucleated, stellate myoblasts (data not shown). As shown in Fig. 7, total RNA Northern blot analysis detected the appearance of the muscle-specific skeletal α-actin and myogenin RNA transcripts, using random-primed cDNA probes in wild-type C2C12 fused myotubes. Expression of both the endogenous skeletal α-actin and myogenin genes was significantly inhibited in both mutant cell lines. Since myogenin is not thought to be regulated by SRF, these results indicate both direct and indirect inhibitory activities of SRFΔ5, consistent with its proposed role as a myogenic dominant negative inhibitor.
FIG. 7.
SRFΔ5 inhibits differentiation of myogenic C2C12 cells. C2C12 cells were cotransfected with SRFΔ5 and SRFpm1 expression plasmids and pSV2neo vector. G418-resistant cell populations were grown to 50% confluence in growth medium (GM) and induced to differentiate for 3 days by adding differentiation medium (DM). Total RNA (20 μg) was subjected to Northern analysis to detect expression of the skeletal muscle-specific factors skeletal α-actin (αSk actin) and myogenin, using random-primed probes. 28S and 18S rRNA bands were visualized by ethidium bromide staining to show RNA loading. Expression of skeletal α-actin and myogenin was significantly inhibited in both mutant cell lines.
SRFΔ5 repressed the serum-induced activity of the c-fos SRE.
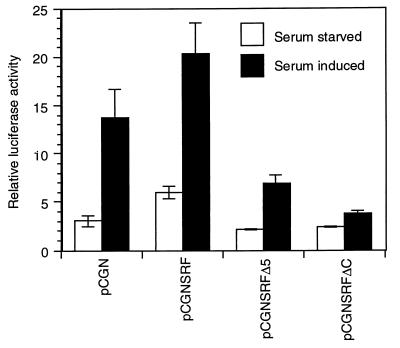
Since SREs have contextural differences and are not equivalent, we wanted to evaluate the role of SRFΔ5 on a minimal c-fos SRE promoter. Rapid induction and subsequent downregulation of c-fos response following serum growth factor stimulation have been shown to be mediated by the dyad symmetry element, the core of which binds SRF (10, 17). We assessed the effect of overexpression of SRFΔ5 on the serum responsiveness of c-fos SRE. 10T1/2 mouse fibroblasts were cotransfected with the c-fos SRE luciferase reporter, the expression vectors for SRF and SRFΔ5, and the empty vector pCGN. After 48 h of serum starvation, cells were induced with 20% serum for 3 h. As shown in Fig. 8, overexpression of SRF increased both basal and serum-induced responses of the c-fos SRE. Overexpression of SRFΔC and SRFΔ5 did not affect the basal activity of the c-fos SRE. However, serum responsiveness was significantly reduced but not eliminated by SRFΔ5. In contrast to SRFΔ5, SRFΔC attenuated the serum inducibility of the c-fos SRE. Essentially similar results were obtained for NIH 3T3 cells (data not shown). These results indicate that the absence of exon 5 may be bypassed to some extent through recruitment of transcription factors that interact with extra-SRFΔ5 regions of the transcriptional activating domain.
FIG. 8.
SRFΔ5 represses serum-induced activity of c-fos SRE. 10T1/2 murine fibroblasts were transfected with 200 ng each of the internal control plasmid pCMVβGal, luciferase reporter plasmid c-fos SRELuc, and expression plasmids pCGNSRF, pCGNSRFΔ5, pCGNSRFΔC or the empty vector pCGN. After transfection, cells were maintained in DMEM containing 10% serum for 16 h. Cells were serum starved for 48 h in DMEM containing 0.5% serum and then induced with 20% serum for 3 h. Luciferase activity was normalized to β-galactosidase activity. Similar results were obtained for NIH 3T3 cells.
ATF6 complements Gal4-SRFΔ5 transcriptional activity.
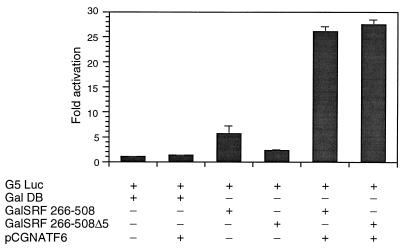
To further determine the role of exon 5 and extra-exon 5 regions of the transactivation domain, we then evaluated the role of two Gal4-SRF fusion constructions; plasmid Gal4SRF contains the C-terminal transcriptional activation domain of SRF (amino acids 266 to 508) fused to the Gal4 DNA binding domain, while Gal4SRFΔ5 contains the precise deletion of exon 5-encoded sequences within the C-terminal domain. Expression plasmids GalDB, Gal4SRF, and Gal4SRFΔ5 were cotransfected with a luciferase reporter containing five copies of the upstream activation sequence cloned upstream of the minimal adenovirus E1B promoter into CV1 cells. Cotransfection of the reporter with the Gal4DB alone did not increase reporter activity (Fig. 9). Plasmid Gal4SRF, encoding the SRF transactivation domain, stimulated reporter activity modestly (sixfold). In comparison, Gal4SRFΔ5 transactivated reporter activity only two- to threefold, indicating that the transcriptional activating region of SRF appears to be encoded primarily within exon 5. The coactivator ATF6 (64) was then used in cotransfection experiments with plasmids Gal4SRF and GAL4SRFΔ5 to determine whether the absence of exon 5 might be bypassed through recruitment of ATF6. As shown in Fig. 9, we observed a 5-fold activation of GAL4-SRF activity and a robust 14-fold stimulation of GAL4-SRFΔ5 activity. These results indicate that alternatively spliced SRFΔ5 can be rescued by ATF6 coactivation through sequences which lie outside exon 5.
FIG. 9.
ATF6 complements Gal4-SRF C-terminal Δ5 transcriptional activity. CV1 cells were transfected with 1 μg of the Gal4 luciferase reporter (G5luc), Gal4DB, Gal4SRF 266-508, SRFΔ5, and ATF6 expression vectors in a variety of combinations as indicated. Cells were harvested 48 h posttransfection, and luciferase activity was measured. The results shown are from three experiments.
DISCUSSION
Alternative splicing is a commonly used strategy for creating a functionally diverse pool of gene products derived from a single gene. The primary transcripts encoding the MADS box superfamily of proteins from animals, such as MEF2 (reviewed in reference 40), and plants, such as ZEMa (reviewed in reference 46), were alternatively spliced to produce several isoforms. Further, some of these splicing events were cell type restricted. The splicing in of the alternative exon A for MEF2C was restricted to brain (35), and skipping of the SEEEELEL miniexon was restricted to vascular smooth muscles and liver (55). However, no specific function for any of the MEF2 spliced isoforms has been described so far. Here we have reported the alternative splicing of an exon from the C-terminal transactivating domain of SRF. The C-terminally truncated MEF2 proteins were deficient for transactivation and functioned as dominant negative mutants (41). Similarly, transactivation domain-deleted SRF was also a potent repressor of SRE-dependent promoters. The transrepressing activity of SRF could be attributed to the presence of two potent inhibitory domains in SRF (10, 22).
When fused to the heterologous GAL4 DNA binding domain, the C terminus of SRF comprising exons 3 through 7 activated transcription in CV1 cells. A portion of the transactivation function was localized to exon 5 because the precise deletion of its encoded sequences severely reduced transcriptional activity. Our results differ from those of Johansen and Prywes (22), who mapped the activation domain to SRF amino acids 339 to 508 in HeLa cells and 414 to 508 in NIH 3T3 cells. Further, in their assays a C-terminal fragment of SRF from amino acids 204 to 465 comprising the complete exon 5 sequences in addition to part of exon 2, exons 3 and 4, and part of exon 6 was incapable of transcriptional activation. In contrast, in HuT-12 cells, amino acids 406 to 476 were sufficient for transactivation (30). The disparity between these results could be explained by our hypothesis that the SRF activation domain probably arose from the assembly of several functional protein interactive domains (exons) during evolution. The correlation between the evolutionarily conserved organization of exon 5 and the surrounding introns between mouse and Xenopus (and possibly human) SRF (3, 36) and the loss of transactivation function upon deletion of this exon (Fig. 6 and 9) suggested that the activation function of SRF was modular and encoded in part by exon 5. Further, the requirement for the SRF activation domain could differ between different cell types and the context of SRF binding site in the promoter. This possibility was supported by the observation that SRF activation domains have been mapped to different regions in various cell types (amino acids 339 to 508 for HeLa cells, 414 to 508 for NIH 3T3 cells, 406 to 476 for HuT-12 cells, and 388 to 452 for CV1 cells) and a C-terminally deleted SRF (SRFΔC) was a weak activator for skeletal and cardiac α-actin promoters and a repressor for the SRF promoter (Fig. 6A, B, and D). In addition, SRFΔ5 reduced but did not abolish serum-induced activation of c-fos SRE. Thus, c-fos transactivation required both exon 5 and extra-exon 5 regions of the transactivating domain. In addition, the cell type dependence of SRF activation domain could be explained by its differential interaction with components of basal transcription machinery such as TFIID (63), RAP74 subunit of TFIIF (23), accessory proteins such as human T-cell leukemia virus type 1 Tax protein (13), ATF6 (64), or cell-type-specific SRF coactivators. As shown in Fig. 9, coactivators such ATF6 (64) complemented Gal4-SRFΔ5 transcriptional activity, indicating that the absence of exon 5 might be circumvented through procuring cofactors that interact with extra-exon 5 regions.
SRFΔ5 repressed the activity of SRE-dependent promoters in transient transfection assays (Fig. 6) and interfered with the fusion and myogenic differentiation of C2C12 cells upon stable expression (Fig. 7). In earlier reports, interference with the activity or expression of SRF was shown to block the myogenic differentiation of C2C12 cells and the expression of myogenic regulatory factors such as MyoD and myogenin and muscle-specific markers such as skeletal α-actin and troponin T (7, 16, 60). The repression of SRE-dependent promoters by SRFΔ5 appears to be due to the formation of SRF-SRFΔ5 heterodimers and SRFΔ5 homodimers at the SRE (Fig. 5). In addition to repression caused by exclusion of SRF binding, SRFΔ5 homodimers might actively repress transcription through their repressor domains (10, 22). Two SREs present in the SRF promoter mediated both the positive autoregulation of the promoter in skeletal myotubes (3) and serum induction in NIH 3T3 cells (54). Repression of the SRF promoter activity in both cell types is likely to be mediated by binding of SRFΔ5 to the same SREs. Therefore, the SRF promoter is both positively and negatively autoregulated by alternatively spliced products of SRF.
The relative proportion of constitutively spliced RNA to the alternatively spliced RNA varied among the tissues examined. SRFΔ5 RNA was expressed at a lower level in visceral and vascular SMCs (Fig. 2 and 3). Further, SRFΔ5 was expressed in a rostro-caudal gradient along the aorta, with the highest level of SRFΔ5 in the aortic arch and the lowest in the abdominal aorta. The SMC subpopulations within these segments are derived from distinct lineages and differ from each other in their growth potential and response to transforming growth factor β (57) and expression of differentiation markers such as tropoelastin (14). Interestingly, SM22α and SM-MHC, which are regulated by SRF, showed an inversely graded expression to that of SRFΔ5. We suspect this inverse gradient of SM22α and SM-MHC expression is a consequence of SRFΔ5-mediated repression because overexpression of SRFΔ5 inhibited both endogenous (25) and transfected SM22α promoter activities. SMCs differ from striated muscle cells in that they readily dedifferentiate and reenter cell cycle in response to various signals (reviewed in reference 42). Thus, SMCs may require higher levels of SRF to be maintained in the nonproliferating contractile state. Alternatively, upregulation of SRFΔ5 could repress the SRE-dependent genes and induce the preproliferative synthetic state in these cells. Induction of the synthetic state is an important step in the pathogenesis of atherosclerosis (reviewed in reference 48). Studies are under way to investigate if the signals that induce dedifferentiation of SMCs increase the level of SRFΔ5 RNA and protein.
In addition to SRFΔ5, we have detected several other alternatively spliced isoforms of SRF during differentiation of embryonic stem cells into EBs, a process which precisely recapitulates the events that occur during early mouse embryogenesis. Based on the SRF cDNA sequence, splice variants lacking individual exons 4 and 6 and double exons 4-5 and 5-6 are predicted to express isoforms of SRF with different C termini. Since EBs contain diverse population of cells from different lineages, different SRF isoforms could serve different functions among these cell lineages. Work is under way to characterize these novel isoforms of SRF.
Several antagonistic protein isoforms derived from the same primary transcript by alternative splicing or alternative translation initiation have been reported (reviewed in reference 31). Generation of functionally antagonistic proteins by regulated alternative splicing of a primary transcript would facilitate rapid termination of the cellular response or switching to the alternate modes of response. An important role for SRFΔ5 can be envisioned in silencing the activity of immediate-early genes in differentiated cells and the rapid termination of their responses to mitogens and growth factors. Previous studies have suggested that both the induction and termination of c-fos response are mediated by the SRE (17). In quiescent cells, c-fos expression could be kept repressed by the occupation of SRE by SRFΔ5. The growth factor stimuli could result in the rapid displacement of SRFΔ5 with SRF, allowing the expression of c-fos. This could also explain why SRF appeared to be constitutively bound to SRE irrespective of serum stimulation (20). Termination of the response could again be mediated by displacement of SRF by SRFΔ5. Possibly, the induction of SRFΔ5 could also play an important role in the dedifferentiation and proliferation of SMCs in wound repair, neovascularization, and pathological processes such as atherosclerosis and heart failure. We believe that the novel alternatively spliced isoform of SRF, SRFΔ5, that we have reported here could serve as a valuable tool for dissection of the role of SRF in the activation of immediate-early genes and muscle-specific genes and ultimately play a regulatory role in modulating contractile protein gene expression that is dependent on SRF.
ACKNOWLEDGMENT
This study was supported by National Institutes of Health grant PO1HL49953.
REFERENCES
- 1.Annweiler A, Muller-Immergluck M, Wirth T. Oct2 transactivation from a remote enhancer position requires a B-cell-restricted activity. Mol Cell Biol. 1992;12:3107–3116. doi: 10.1128/mcb.12.7.3107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Arsenian S, Weinhold B, Oelgeschlager M, Ruther U, Nordheim A. Serum response factor is essential for mesoderm formation during mouse embryogenesis. EMBO J. 1998;17:6289–6299. doi: 10.1093/emboj/17.21.6289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Belaguli N S, Schildmeyer L A, Schwartz R J. Organization and myogenic restricted expression of the murine serum response factor gene. J Biol Chem. 1997;272:18222–18231. doi: 10.1074/jbc.272.29.18222. [DOI] [PubMed] [Google Scholar]
- 4.Belaguli, N. S., J. L. Sepulveda, V. Nigam, and R. J. Schwartz. Unpublished data.
- 5.Bickmore W A, Oghene K, Little M H, Seawright A, VanHeyningen V, Hastie N D. Modulation of DNA binding specificity by alternative splicing of the Wilms tumor wt1 gene transcript. Science. 1992;257:235–238. doi: 10.1126/science.1321494. [DOI] [PubMed] [Google Scholar]
- 6.Chen C-Y, Schwartz R J. Recruitment of the tinman homolog Nkx-2.5 by serum response factor activates cardiac α-actin gene transcription. Mol Cell Biol. 1996;16:6372–6384. doi: 10.1128/mcb.16.11.6372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Croissant J D, Kim J H, Eichele G, Goering L, Lough J, Prywes R, Schwartz R J. Avian serum response factor expression restricted primarily to muscle cell lineages is required for alpha-actin gene transcription. Dev Biol. 1996;177:250–264. doi: 10.1006/dbio.1996.0160. [DOI] [PubMed] [Google Scholar]
- 8.Das G, Herr W. Enhanced activation of the human histone H2B promoter by an Oct-1 variant generated by alternative splicing. J Biol Chem. 1993;268:25026–25032. [PubMed] [Google Scholar]
- 9.Epstein J A, Glaser T, Cai J, Jepeal L, Walton D S, Mass R L. Two independent and interactive DNA-binding subdomains of the Pax6 paired domain are regulated by alternative splicing. Genes Dev. 1994;8:2022–2034. doi: 10.1101/gad.8.17.2022. [DOI] [PubMed] [Google Scholar]
- 10.Ernst W H, Janknecht R, Cahill M A, Nordheim A. Transcriptional repression mediated by the serum response factor. FEBS Lett. 1995;357:45–49. doi: 10.1016/0014-5793(94)01321-q. [DOI] [PubMed] [Google Scholar]
- 11.Foulkes N S, Sassone-Corsi P. More is better: activators and repressors from the same gene. Cell. 1992;68:411–414. doi: 10.1016/0092-8674(92)90178-f. [DOI] [PubMed] [Google Scholar]
- 12.Foulkes N S, Mellstrom B, Benusiglio E, Sassone-Corsi P. Developmental switch of CREM function during spermatogenesis: from antagonist to activator. Nature. 1992;355:80–84. doi: 10.1038/355080a0. [DOI] [PubMed] [Google Scholar]
- 13.Fujii M, Tsuchiya H, Chuhjo T, Akizawa T, Seiki M. Interaction of HTLV-1 Tax1 with p67SRF causes the aberrant induction of cellular immediate early genes through CArG boxes. Genes Dev. 1992;6:2066–2076. doi: 10.1101/gad.6.11.2066. [DOI] [PubMed] [Google Scholar]
- 14.Gadson P F, Jr, Rossignol C, McCoy J, Rosenquist T H. Expression of elastin, smooth muscle alpha-actin, and c-jun as a function of the embryonic lineage of vascular smooth muscle cells. In Vitro Cell Dev Biol Anim. 1993;29A:773–781. doi: 10.1007/BF02634344. [DOI] [PubMed] [Google Scholar]
- 15.Gauthier-Rouviere C, Cai Q Q, Lautredou N, Fernandez A, Blanchard J M, Lamb N J. Expression and purification of the DNA-binding domain of SRF: SRF-DM, a part of a DNA-binding protein which can act as a dominant negative mutant in vivo. Exp Cell Res. 1993;209:208–215. doi: 10.1006/excr.1993.1303. [DOI] [PubMed] [Google Scholar]
- 16.Gauthier-Rouviere C, Vandromme M, Tuil D, Lautredou N, Morris M, Soulez M, Kahn A, Fernandez A, Lamb N J. Expression and activity of serum response factor is required for expression of the muscle-determining factor MyoD in both dividing and differentiating mouse C2C12 myoblasts. Mol Biol Cell. 1996;7:719–729. doi: 10.1091/mbc.7.5.719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gius D, Cao X M, Rauscher F J, Cohen D R, Curran T, Sukhatme V P. Transcriptional activation and repression by Fos are independent functions: the C terminus represses immediate-early gene expression via CArG elements. Mol Cell Biol. 1990;10:4243–4255. doi: 10.1128/mcb.10.8.4243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Groisman R, Masutani H, Leibovitch M-P, Robin P, Soudant I, Trouche D, Harel-Bellan A. Physical interaction between the mitogen-responsive serum response factor and myogenic basic-helix-loop-helix proteins. J Biol Chem. 1996;271:5258–5264. doi: 10.1074/jbc.271.9.5258. [DOI] [PubMed] [Google Scholar]
- 19.Han T-H, Prywes R. Regulatory role of MEF2D in serum induction of the c-jun promoter. Mol Cell Biol. 1995;15:2907–2915. doi: 10.1128/mcb.15.6.2907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Herrera R E, Shaw P E, Nordheim A. Occupation of the c-fos serum response element in vivo by a multi-protein complex is unaltered by growth factor induction. Nature. 1989;340:68–70. doi: 10.1038/340068a0. [DOI] [PubMed] [Google Scholar]
- 21.Hewitt S M, Fraizer G C, Wu Y-J, Rauscher III F J, Saunders G F. Differential function of Wilms’ tumor gene WT1 splice isoforms in transcriptional regulation. J Biol Chem. 1996;271:8588–8592. doi: 10.1074/jbc.271.15.8588. [DOI] [PubMed] [Google Scholar]
- 22.Johansen F-E, Prywes R. Identification of transcriptional activation and inhibitory domains in serum response factor (SRF) by using GAL4-SRF constructs. Mol Cell Biol. 1993;18:4640–4647. doi: 10.1128/mcb.13.8.4640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jollot V, Demma M, Prywes R. Interaction with RAP74 subunit of TFIIF is required for transcriptional activation by serum response factor. Nature. 1995;373:632–635. doi: 10.1038/373632a0. [DOI] [PubMed] [Google Scholar]
- 24.Kozmik Z, Kurzbauer R, Dorfler P, Busslinger M. Alternative splicing of Pax-8 gene transcripts is developmentally regulated and generates isoforms with different transactivation properties. Mol Cell Biol. 1993;13:6024–6035. doi: 10.1128/mcb.13.10.6024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Landerholm, T. E., X. R. Dong, J. Lu, N. S. Belaguli, R. J. Schwartz, and M. Majesky. A role for serum response factor in coronary smooth muscle differentiation from proepicardial cell. Development 126:2053–2062. [DOI] [PubMed]
- 26.Lee T-C, Shi Y, Schwartz R J. Displacement of BrdUrd-induced YY1 by serum response factor activates skeletal alpha-actin transcription in embryonic myoblasts. Proc Natl Acad Sci USA. 1992;89:9814–9818. doi: 10.1073/pnas.89.20.9814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Leifer D, Krainc D, Yu Y-T, McDermott J, Breitbart R E, Heng J, Neve R L, Kosofsky B, Nadal-Ginard B, Lipton B. MEF2C, a MADS/MEF2-family transcription factor expressed in a laminar distribution in cerebral cortex. Proc Natl Acad Sci USA. 1992;90:1546–1550. doi: 10.1073/pnas.90.4.1546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li L, Liu Z-C, Mercer B, Overbeek P, Olson E N. Evidence for serum response factor-mediated regulatory networks governing SM22α transcription in smooth, skeletal, and cardiac muscle cells. Dev Biol. 1997;187:311–321. doi: 10.1006/dbio.1997.8621. [DOI] [PubMed] [Google Scholar]
- 29.Lillycrop K A, Latchman D S. Alternative splicing of the Oct-2 transcription factor is differentially regulated in B cells and neuronal cells and results in protein isoforms with opposite effects on the activity of octamer/TAATGARAT-containing promoters. J Biol Chem. 1992;267:24960–24966. [PubMed] [Google Scholar]
- 30.Liu S H, Ma J T, Yueh A Y, Lees-Miller S P, Anderson C W, Ng S Y. The carboxyl-terminal transactivation domain of human serum response factor contains DNA-activated protein kinase phosphorylation sites. J Biol Chem. 1993;268:21147–21154. [PubMed] [Google Scholar]
- 31.Lopez J A. Developmental role of transcription factor isoforms generated by alternative splicing. Dev Biol. 1995;172:396–411. doi: 10.1006/dbio.1995.8050. [DOI] [PubMed] [Google Scholar]
- 32.Macneil C, Ayres B, Laverriere A C, Burch J E. Transcripts for functionally distinct isoforms of chicken GATA-5 are differentially expressed from alternative first exons. J Biol Chem. 1997;272:8396–8401. doi: 10.1074/jbc.272.13.8396. [DOI] [PubMed] [Google Scholar]
- 33.Martin J F, Schwarz J J, Olson E N. Myocyte enhancer factor (MEF) 2C: a tissue-restricted member of the MEF-2 family of transcription factors. Proc Natl Acad Sci USA. 1993;90:5282–5286. doi: 10.1073/pnas.90.11.5282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Martin J F, Miano J M, Hustad C M, Copeland N G, Jenkins N A, Olson E N. A Mef2 gene that generates a muscle-specific isoform via alternative mRNA splicing. Mol Cell Biol. 1994;14:1647–1656. doi: 10.1128/mcb.14.3.1647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.McDermott J C, Cordoso M C, Yu Y-T, Andres V, Leifer D, Krainc D, Lipton S A, Nadal-Ginard B. HMEF2C gene encodes skeletal muscle- and brain-specific transcription factors. Mol Cell Biol. 1993;13:2564–2577. doi: 10.1128/mcb.13.4.2564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Mohun T J, Chambers A E, Towers N, Taylor M V. Expression of genes encoding the transcription factor SRF during early development of Xenopus laevis: identification of a CArG box-binding activity as SRF. EMBO J. 1991;10:933–940. doi: 10.1002/j.1460-2075.1991.tb08027.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Morris A E, Kloss B, McChesney R E, Bancroft C, Chasin L A. An alternatively spliced Pit-1 isoform altered in its ability to transactivate. Nucleic Acids Res. 1992;20:1355–1361. doi: 10.1093/nar/20.6.1355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Morris P J, Theil T, Ring C J A, Lillycrop K A, Moroy T, Latchman D S. The opposite and antagonistic effects of the closely related POU family transcription factors on the activity of a target promoter are dependent upon differences in the POU domain. Mol Cell Biol. 1994;14:6907–6914. doi: 10.1128/mcb.14.10.6907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Nornes S, Mikkola I, Krauss S, Delghandi M, Perander M, Johansen T. Zebrafish Pax9 encodes two proteins with distinct C-terminal transactivating domains of different potency negatively regulated by adjacent N-terminal sequences. J Biol Chem. 1996;271:26914–26923. doi: 10.1074/jbc.271.43.26914. [DOI] [PubMed] [Google Scholar]
- 40.Olson E N, Perry M, Schulz R A. Regulation of muscle differentiation by the MEF2 family of MADS box transcription factors. Dev Biol. 1995;172:2–14. doi: 10.1006/dbio.1995.0002. [DOI] [PubMed] [Google Scholar]
- 41.Ornatsky O I, Andreucci J J, McDermott J C. A dominant-negative form of transcription factor MEF2 inhibits myogenesis. J Biol Chem. 1997;272:33271–33278. doi: 10.1074/jbc.272.52.33271. [DOI] [PubMed] [Google Scholar]
- 42.Owens G K. Regulation of differentiation of vascular smooth muscle cells. Physiol Rev. 1995;75:487–517. doi: 10.1152/physrev.1995.75.3.487. [DOI] [PubMed] [Google Scholar]
- 43.Pellegrini L, Tans S, Richmond T J. Structure of serum response factor core bound to DNA. Nature. 1995;376:490–498. doi: 10.1038/376490a0. [DOI] [PubMed] [Google Scholar]
- 44.Pollock R, Treisman R. Human SRF-related proteins: DNA-binding properties and potential regulatory targets. Genes Dev. 1991;5:2327–2341. doi: 10.1101/gad.5.12a.2327. [DOI] [PubMed] [Google Scholar]
- 45.Reecy J, Belaguli N S, Schwartz R J. SRF/homeobox protein interactions. In: Harvey R, Rosenthal N, editors. Heart development. San Diego, Calif: Academic Press; 1998. pp. 273–290. [Google Scholar]
- 46.Reichmann J L, Meyerowitz E M. MADS domain proteins in plant development J. Biol Chem. 1997;378:1079–1101. [PubMed] [Google Scholar]
- 47.Rivera V M, Miranti C K, Misra R P, Ginty D D, Chen R-H, Blenis J, Greenberg M E. A growth factor-induced kinase phosphorylates the serum response factor at a site that regulates its DNA-binding activity. Mol Cell Biol. 1993;13:6260–6273. doi: 10.1128/mcb.13.10.6260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ross R. Atherosclerosis: a defence mechanism gone awry. Am J Pathol. 1993;143:987–1002. [PMC free article] [PubMed] [Google Scholar]
- 49.Ruzicka D L, Schwartz R J. Sequential activation of alpha-actin genes during avian cardiogenesis: vascular smooth muscle alpha-actin gene transcripts mark the onset of cardiomyocyte differentiation. J Cell Biol. 1988;107:2575–2586. doi: 10.1083/jcb.107.6.2575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Sartorelli V, Webster K, Kedes L. Muscle-specific expression of the cardiac α-actin gene requires MyoD1, CArG-box binding factor, and SP1. Genes Dev. 1990;4:1811–1822. doi: 10.1101/gad.4.10.1811. [DOI] [PubMed] [Google Scholar]
- 51.Schwarz-Sommer Z, Huijser P, Nacken W, Seadler H, Sommer H. Genetic control of flower development by homeotic genes in Antirrhinum majus. Science. 1990;250:931–936. doi: 10.1126/science.250.4983.931. [DOI] [PubMed] [Google Scholar]
- 52.Shore P, Sharrocks A D. The MADS-box family of transcription factors. Eur J Biochem. 1995;229:1–13. doi: 10.1111/j.1432-1033.1995.tb20430.x. [DOI] [PubMed] [Google Scholar]
- 53.Soulez M, Gauthier-Rouviere C, Chafey P, Hentzen D, Vandromme M, Lautredou N, Lamb N, Kahn N, Tuil D. Growth and differentiation of C2 myogenic cells are dependent on serum response factor. Mol Cell Biol. 1996;16:6065–6074. doi: 10.1128/mcb.16.11.6065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Spencer J A, Misra R P. Expression of the serum response factor gene is regulated by serum response factor binding sites. J Biol Chem. 1996;271:16535–16543. doi: 10.1074/jbc.271.28.16535. [DOI] [PubMed] [Google Scholar]
- 55.Suzuki E, Guo K M, Kolman M, Yu Y-T, Walsh K. Serum induction of MEF2/RSRF in vascular myocytes is mediated at the level of translation. Mol Cell Biol. 1995;15:3415–3423. doi: 10.1128/mcb.15.6.3415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tanaka T, Tanaka K, Ogawa S, Kurokawa M, Mitani K, Nishida J, Shibata Y, Yazaki Y, Hirai H. An acute myeloid leukemia gene, AML1, regulates hemopoietic myeloid cell differentiation and transcriptional activation antagonistically by two alternatively spliced forms. EMBO J. 1995;14:341–350. doi: 10.1002/j.1460-2075.1995.tb07008.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Topouzis S, Majesky M. Smooth muscle lineage diversity in the chick embryo. Two types of aortic smooth muscle cell differ in growth and receptor-mediated transcriptional responses to transforming growth factor-beta. Dev Biol. 1996;178:430–445. doi: 10.1006/dbio.1996.0229. [DOI] [PubMed] [Google Scholar]
- 58.Treisman R. Journey to the surface of the cell: Fos regulation and the SRE. EMBO J. 1995;14:4905–4913. doi: 10.1002/j.1460-2075.1995.tb00173.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tsukamoto K, Nakamura Y, Niikawa N. Isolation of two isoforms of the PAX3 gene transcripts and their tissue-specific alternative expression in human adult tissues. Hum Genet. 1994;93:270–274. doi: 10.1007/BF00212021. [DOI] [PubMed] [Google Scholar]
- 60.Vandromme M, Gauthier-Rouviere C, Carnac G, Lamb N, Fernandez A. Serum response factor p67SRF is expressed and required during myogenic differentiation of both mouse C2 and rat L6 muscle cell lines. J Cell Biol. 1992;118:1489–1500. doi: 10.1083/jcb.118.6.1489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yin J C P, Wallach J S, Wilder E L, Klingensmith J, Dang D, Perrimon N, Zhou H, Tully T, Quinn W G. A Drosophila CREB/CREM homolog encodes multiple isoforms, including a cyclic AMP-dependent protein kinase-responsive transcriptional activator and antagonist. Mol Cell Biol. 1995;15:5123–5130. doi: 10.1128/mcb.15.9.5123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Yu Y-T, Breitbart R E, Smoot L B, Lee Y, Mahdavi V, Nadal-Ginard B. Human myocyte-specific enhancer factor 2 comprises a group of tissue-restricted MADS box transcription factors. Genes Dev. 1992;6:1783–1798. doi: 10.1101/gad.6.9.1783. [DOI] [PubMed] [Google Scholar]
- 63.Zhu H, Roy A L, Roeder R G, Prywes R. Serum response factor affects preinitiation complex formation by TFIID in vitro. New Biol. 1991;3:455–464. [PubMed] [Google Scholar]
- 64.Zhu C, Johansen F-E, Prywes R. Interaction of ATF6 and serum response factor. Mol Cell Biol. 1997;17:4957–4966. doi: 10.1128/mcb.17.9.4957. [DOI] [PMC free article] [PubMed] [Google Scholar]