Abstract
Mutations in SSY1 and PTR3 were identified in a genetic selection for components required for the proper uptake and compartmentalization of histidine in Saccharomyces cerevisiae. Ssy1p is a unique member of the amino acid permease gene family, and Ptr3p is predicted to be a hydrophilic protein that lacks known functional homologs. Both Ssy1p and Ptr3p have previously been implicated in relaying signals regarding the presence of extracellular amino acids. We have found that ssy1 and ptr3 mutants belong to the same epistasis group; single and ssy1 ptr3 double-mutant strains exhibit indistinguishable phenotypes. Mutations in these genes cause the nitrogen-regulated general amino acid permease gene (GAP1) to be abnormally expressed and block the nonspecific induction of arginase (CAR1) and the peptide transporter (PTR2). ssy1 and ptr3 mutations manifest identical differential effects on the functional expression of multiple specific amino acid transporters. ssy1 and ptr3 mutants have increased vacuolar pools of histidine and arginine and exhibit altered cell growth morphologies accompanied by exaggerated invasive growth. Subcellular fractionation experiments reveal that both Ssy1p and Ptr3p are localized to the plasma membrane (PM). Ssy1p requires the endoplasmic reticulum protein Shr3p, the amino acid permease-specific packaging chaperonin, to reach the PM, whereas Ptr3p does not. These findings suggest that Ssy1p and Ptr3p function in the PM as components of a sensor of extracellular amino acids.
Amino acids are transported into the yeast Saccharomyces cerevisiae by both general and specific transport systems. These transport proteins are members of the conserved amino acid permease (AAP) gene family that includes 18 members (3). The AAP genes are differentially expressed. The nitrogen-regulated permeases, the general AAP (GAP1 [34]), and the proline-specific permease (PUT4 [63]) are high-capacity systems that are induced by growth on low-quality nitrogen sources, their expression enables cells to use amino acids as sole nitrogen sources (14, 27, 45). The majority of other AAP family members are low-capacity, high-affinity amino acid permeases, each exhibiting characteristic narrow substrate specificities (31). The transcriptional regulatory mechanisms governing the expression of the low-capacity permeases are not well understood, but in general their expression requires induction and appears less dependent on nitrogen regulation (48). Despite different modes of transcriptional regulation, the functional expression of AAPs depends on the endoplasmic reticulum (ER) packaging chaperonin Shr3p. In cells lacking SHR3, AAPs specifically accumulate in the ER and are not transported to the plasma membrane (PM) (38, 43).
At least five proteins (Ure2p, Dal80p, Gln3p, Nil1p, and Nil2p) function coordinately to control the transcription of many nitrogen-regulated genes, including GAP1 (9, 16, 59, 60). Dal80p, Gln3p, and Nil1p are DNA binding proteins that bind to upstream regulatory sequences that contain a motif surrounding a core GATA sequence. Dal80p is a transcriptional repressor that competes with the transcriptional activators Gln3p and Nil1p for binding to GATA regulatory sequences (13, 15). In cells grown in the presence of ammonium, Ure2p physically interacts with Gln3p and prevents it from acting as an activator (6). Similarly, when glutamine is present in the growth environment, Nil2p represses NIL1 expression (55, 57). Thus, it is only when both of these high-quality nitrogen sources are absent that Gln3p and Nil1p are able to compete for binding to GATA regulatory sequences and activate transcription. The mechanisms that independently control or modulate Ure2p in response to ammonium and Nil2p activity in response to amino acids have not been elucidated.
In addition to regulation by GATA factors, the expression of many nitrogen-regulated genes is controlled by both specific and nonspecific induction mechanisms. For example, in cells grown in the presence of arginine, arginase gene (CAR1) expression is greatly induced. This specific induction is dependent on cis-acting sequences recognized by the Arg80p-Mcm1p activation complex (18, 37). The ability of Arg80p-Mcm1p to induce CAR1 expression is known to be controlled by intracellular levels of arginine. CAR1 expression is nonspecifically induced in cells grown in the presence of micromolar concentrations of a variety of amino acids (19). The mechanisms controlling the nonspecific induction of CAR1 have not been defined.
There is evidence that some members of the sugar and amino acid transport gene families, though structurally similar to the transporters, act as nutritional sensors. For example, the hexose transporter (HXT) gene family is composed of more than 20 proteins related in sequence (3). Two unique members of this family, Snf3p and Rgt2p, control glucose uptake (41, 51a). These proteins differ from other family members; they possess unusually long C-terminal domains, are poorly expressed compared to the other known functional hexose transporters, and pleiotropically affect the function of multiple HXTs. The C-terminal domain alone of Snf3p has been shown to complement a number of snf3 mutant phenotypes, suggesting that Snf3p does not function as an HXT but rather transduces nutritional signals regarding extracellular glucose availability (11).
The AAP gene family also contains a unique member, Ssy1p (32, 35), that differs from the other members of the family in that it contains an unusually long N-terminal domain. The induced expression of a broad-specificity AAP gene (AGP1), several specific branched-chain AAP genes (BAP2, TAT1, and BAP3), and the peptide transporter gene (PTR2) have been shown to require Ssy1p (17, 32). The Ssy1p-dependent induction occurs in response to extracellular amino acids and in the absence of detectable amino acid uptake (17, 32). In gap1 null mutant strains, ssy1 mutations pleiotropically affect the uptake of other, nonbranched amino acids (17, 32). Based on its amino acid sequence and similarity to known amino acid transporters, Ssy1p has been suggested to act at the PM as a sensor of extracellular amino acids exhibiting high sensitivity to hydrophobic amino acids, e.g., leucine (17, 32). In addition to Ssy1p, the induced expression of PTR2 and BAP2 has been shown to be dependent on Ptr3p (5). Ptr3p is predicted to be a hydrophilic protein that lacks identifiable homologs of known function; these features have made its localization and function less clear.
In this report, we describe the isolation and characterization of mutations in SSY1 and PTR3 and demonstrate that ptr3 mutant phenotypes are similar to those of ssy1 despite differences in the structure, membrane association, and intracellular trafficking of the two proteins. ssy1Δ and ptr3Δ mutants exhibit altered patterns of expression of two nitrogen-regulated genes, GAP1 and CAR1. In response to alternative nitrogen sources, ssy1 and ptr3 mutations exert both negative and positive effects on the transcription of a diverse spectrum of specific amino acid and peptide transport proteins. Additionally, ssy1Δ and ptr3Δ mutants have increased vacuolar pools of histidine and arginine and exhibit enhanced haploid-specific invasive growth. Subcellular fractionation experiments demonstrate that Ssy1p localizes to the PM in an Shr3p-dependent manner and that Ptr3p is a peripheral membrane protein that localizes to the cytosolic face of the PM. Our results suggest that Ssy1p and Ptr3p function together at the PM, or within the same pathway, to transmit signals regarding the availability of extracellular amino acids.
MATERIALS AND METHODS
Strains and media.
Yeast strains used are listed in Table 1. PLY2 was constructed from PLY1; the mating type was switched by transformation with plasmid pGAL-HO (29). PLY1 was transformed with a linear SalI/SpeI fragment of pHK030 (Table 2; plasmids are described below) containing ssy1Δ12::hisG-URA3-hisG, and PLY2 was transformed with a linear EcoRI fragment of pPL341 containing ptr3Δ14::hisG-URA3-hisG; Ura+ transformants were propagated on medium containing 5-fluoroorotic acid (5-FOA) (28), resulting in strains HKY37 and HKY38, respectively. A spontaneous Ade+ revertant of AA280 was isolated and crossed to AA288 (4), the resulting diploid was sporulated, and tetrads were dissected. PLY122, PLY124, and PLY125 are segregants from this cross. The diploid strain HKDY1, obtained by mating PLY122 and PLY125, was transformed with the ssy1Δ12::hisG-URA3-hisG cassette (pHK030). A Ura+ transformant was sporulated, and tetrads were dissected. ssy1Δ12 segregants were propagated on medium containing 5-FOA to attain strains with the unmarked ssy1Δ13 deletion (HKY20 and HKY21). A strain with the unmarked ptr3Δ15 deletion (HKY31) was similarly constructed by using the ptr3Δ14::hisG-URA3-hisG cassette (pPL341). The diploid strain HKDY5, obtained by mating HKY20 and HKY21, was used to construct ssy1Δ13 ptr3Δ15 and ssy1Δ13 shr3Δ6 double-mutant strains. One allele of SHR3 in HKDY5 was replaced with a linear SalI/EcoRI fragment of pPL288 containing shr3Δ5::hisG-URA3-hisG. After tetrad analysis, an shr3Δ5 segregant was propagated on medium containing 5-FOA to attain HKY29. Similarly, one allele of PTR3 in HKDY5 was replaced with ptr3Δ14 to create strain HKY33. SHR3 in strain HKY31 was replaced, and the resulting ptr3Δ15 shr3Δ5 double-mutant strain was propagated on medium containing 5-FOA to create strain HKY51. Strain FGY58 is a meiotic segregant from a diploid strain derived from the mating of PLY129 with PLY214. ssy1Δ12 and ptr3Δ14 alleles were introduced directly into strain FGY58 by transformation, resulting in strains HKY63 and HKY65, respectively. The ptr3Δ14, ssy1Δ12, and shr3Δ5 alleles were introduced into the Σ1278b background strain 10480-5C to create strains HKY39, HKY41, and HKY55. HKY40, HKY42, and HKY56 were similarly derived from strain 10480-5D. HKY39 and HKY41 were propagated on 5-FOA, resulting in strains HKY43 and HKY45, respectively. All gene replacements were confirmed by Southern analysis. Diploid strains were constructed by crossing strains as indicated in Table 1.
TABLE 1.
S. cerevisiae strains used
| Strain | Genotype | Reference |
|---|---|---|
| Isogenic derivatives of PLY1 | ||
| PLY1 | MATa ura3-52 his4Δ29 | 43 |
| PLY2 | MATα ura3-52 his4Δ29 | This work |
| PLY4 | MATα ura3-52 his4Δ29 ade2Δ1::URA3 | 43 |
| PLAS7-4C | MATa ura3-52 his4Δ29 shr10-7 (ssy1-107) | This work |
| PLAS6-4D | MATa ura3-52 his4Δ29 shr6-6 (ptr3-66) | This work |
| PLAS14-1A | MATa ura3-52 his4Δ29 shr6-14 (ptr3-614) | This work |
| HKY37 | MATa ura3-52 his4Δ29 ssy1Δ13 | This work |
| HKY38 | MATα ura3-52 his4Δ29 ptr3Δ15 | This work |
| Isogenic derivatives of AA280 | ||
| AA280 | MATα ura3-52 lys2Δ201 his3Δ200 ade2 | 4 |
| AA288 | MATa ura3-52 lys2Δ201 leu2-3,112 ade2 | 4 |
| PLY122 | MATa ura3-52 lys2Δ201 leu2-3,112 | This work |
| PLY125 | MATα ura3-52 lys2Δ201 his3Δ200 | This work |
| PLY126 | MATa ura3-52 lys2Δ201 | This work |
| PLY129 | MATa ura3-52 lys2Δ201 leu2-3,112 ade2 gap1Δ::LEU2 | 43 |
| PLY214 | MATα ura3-52 lys2Δ201 ade2Δ1 | 38 |
| FGY58 | MATα ura3-52 lys2Δ201 leu2-3,112 ade2 gap1Δ::LEU2 | This work |
| HKY20 | MATa ura3-52 lys2Δ201 ssy1Δ13 | This work |
| HKY21 | MATα ura3-52 lys2Δ201 his3Δ200 ssy1Δ13 | This work |
| HKY29 | MATa ura3-52 lys2Δ201 ssy1Δ13 shr3Δ6 | This work |
| HKY31 | MATa ura3-52 lys2Δ201 ptr3Δ15 | This work |
| HKY33 | MATa ura3-52 lys2Δ201 ssy1Δ13 ptr3Δ15 | This work |
| HKY51 | MATa ura3-52 lys2Δ201 ptr3Δ15 shr3Δ6 | This work |
| HKY63 | MATα ura3-52 lys2Δ201 leu2-3,112 ade2 gap1Δ::LEU2 ssy1Δ12::hisG-URA3-kanr-hisG | This work |
| HKY65 | MATα ura3-52 lys2Δ201 leu2-3,112 ade2 gap1Δ::LEU2 ptr3Δ14::hisG-URA3-kanr-hisG | This work |
| Isogenic Σ background strains | ||
| 10480-5C | MATa ura3-52 | 42 |
| 10480-5D | MATα ura3-52 | 42 |
| HKY39 | MATa ura3-52 ptr3Δ14::hisG-URA3-kanr-hisG | This work |
| HKY40 | MATα ura3-52 ptr3Δ14::hisG-URA3-kanr-hisG | This work |
| HKY41 | MATa ura3-52 ssy1Δ12::hisG-URA3-kanr-hisG | This work |
| HKY42 | MATα ura3-52 ssy1Δ12::hisG-URA3-kanr-hisG | This work |
| HKY43 | MATa ura3-52 ptr3Δ15 | This work |
| HKY45 | MATa ura3-52 ssy1Δ13 | This work |
| HKY55 | MATa ura3-52 shr3Δ5::hisG-URA3-kanr-hisG | This work |
| HKY56 | MATα ura3-52 shr3Δ5::hisG-URA3-kanr-hisG | This work |
| HKDY11 | HKY39 × HKY40 | This work |
| HKDY12 | HKY41 × HKY42 | This work |
| HKDY14 | 10480-5C × 10480-5D | This work |
| HKDY15 | HKY55 × HKY56 | This work |
TABLE 2.
Plasmids and oligonucleotides used
| Name | Description or sequence | Reference |
|---|---|---|
| Plasmids | ||
| pPL156 | 14-kb fragment containing SSY1 in YCp50 | This work |
| pPL160 | 12.5-kb fragment containing PTR3 in YCp50 | This work |
| pPL193 | 5 kb-EcoRI fragment containing PTR3 in pRS316 | This work |
| pPL288 | shr3Δ5::hisG-URA3-kanr-hisG in pBluescript II SK(+) | 38 |
| pPL341 | ptr3Δ14::hisG-URA3-kanr-hisG in pUC118 (ΔBamHI-ΔHindIII) | This work |
| pPL356 | 3.9-kb SpeI/ClaI fragment containing SSY1 in pRS316 | This work |
| pHK002 | pPL356 ΔXbaI in polylinker | This work |
| pHK003 | pHK002 without native XbaI in SSY1 | This work |
| pHK004 | pHK003 with XbaI site introduced into SSY1 at nucleotides 100–105 | This work |
| pHK005 | pHK003 with XbaI site introduced into SSY1 at nucleotides 202–207 | This work |
| pHK006 | pHK003 with XbaI site introduced into SSY1 at nucleotides 616–621 | This work |
| pHK010 | SSY1-HA1 in pRS316 (functional) | This work |
| pHK013 | SSY1-HA1 in pRS202 (functional) | This work |
| pHK017 | pPL193 ΔXbaI in polylinker | This work |
| pHK018 | PTR3-HA1 in pRS316 (functional) | This work |
| pHK024 | pHK017 with XbaI site introduced into PTR3 | This work |
| pHK030 | ssy1Δ12::hisG-URA3-kanr-hisG in pRS316 | This work |
| pHK033 | ssy1-HA2 in pRS316 (nonfunctional) | This work |
| pHK034 | ssy1-HA3 in pRS316 (nonfunctional) | This work |
| Oligonucleotides | ||
| POL93-025 | 5′-CGAAATACACAACTGATAGGCGGATCCAAGCTTGCATCTAAATATATACGTATGTTTAAGG-3′ | |
| POL95-036 | 5′-GGACTGGCTTCCAGATTTGTAGAGGATGC-3′ | |
| POL95-037 | 5′-CTTCGTCCAGCCTTTCTAGAAATGATACAGACGG-3′ | |
| POL95-038 | 5′-GACGCTGAGTTATCTAGAAGTTCAATAC-3′ | |
| POL95-039 | 5′-GATTGAGTTGAATTCTTCTACCACTAC-3′ | |
| POL96-045 | 5′-GACACTATCAGACTCTAGAGTTAAGGAGAAATC-3′ | |
| POL97-038 | 5′-AGAAGATGTTTACCAGCACG-3′ | |
| POL97-039 | 5′-TGTTTTCGTCTTTACCATCT GG-3′ | |
| POL97-042 | 5′-ACGCTTGGTAATAGACGCC-3′ | |
| POL97-043 | 5′-TTCTGTTCTTGTTCTTGTGGTG-3′ | |
| POL97-048 | 5′-TCAGATGATGCTCAGGACG-3′ | |
| POL97-049 | 5′-CTGTCTCTGTTACCAACGG-3′ | |
| POL98-012 | 5′-ATGCTTAAGCATGGTCTGC-3′ | |
| POL98-013 | 5′-TTGTCAACGTGGTACATGG-3′ |
Standard yeast media were prepared as described in reference 28. Nonstandard synthetic media with alternative nitrogen sources proline (SPD), leucine (SLD), glutamine (SQD), glutamate (SED), and urea (SUD) were prepared as follows. The nitrogen source (4 g/liter) and Difco Yeast Nitrogen Base (26.8 g/liter) were combined to make 4× stock solutions that were filter sterilized. Other components were autoclaved as separate stock solutions (40% glucose, 4% Difco Bacto Agar). Stock solutions and sterile water were mixed to make a 2× solution, and an equal volume of molten 4% agar was added. Where required, SPD was supplemented with 30 mM l-histidine or 30 mM l-lysine. The concentration of Difco Yeast Nitrogen Base in these synthetic media is fourfold higher than the amount used in other standard synthetic media. Yeast transformations were performed as described by Ito et al. (33), using 50 μg of heat-denatured calf thymus DNA. Transformants were selected on solid synthetic complete (SC) dextrose medium lacking uracil.
Plasmids.
Plasmids and oligonucleotides used are listed in Table 2. A 5-kb EcoRI fragment from pPL160 containing PTR3 was inserted into EcoRI-digested pRS316 (56) to create pPL193. The 3.9-kb SpeI/ClaI fragment from pPL156 containing SSY1 was inserted into SpeI/ClaI-digested pRS316 to create pPL356. The ptr3Δ14 deletion cassette in pPL341 was constructed as follows. The 5-kb EcoRI fragment in pPL193 was inserted into the EcoRI site of a modified pUC118 (64) lacking BamHI and HindIII endonuclease sites, resulting in plasmid pPL334. Using single-stranded pPL334 as a template, site-directed mutagenesis (oligonucleotide POL93-025) was used to delete the entire PTR3 coding sequence and to simultaneously create a unique BamHI site (pPL340) (39). A 5-kb BglII/BamHI hisG-URA3-kanr-hisG cassette isolated from pSE1076 (1) was inserted into the BamHI site of pPL340 to create pPL341. pHK030 was constructed by inserting a blunt-end 5-kb BglII/BamHI hisG-URA3-kanr-hisG cassette into BlpI/HindIII-digested pPL356 made blunt by treatment with Klenow fragment. This ssy1Δ construct removes coding sequences for amino acids (aa) 95 to 783. A hemagglutinin (HA) epitope-tagged version of SSY1 was created in multiple steps. Plasmid pPL356 was digested with EagI and SpeI, and the ends were made blunt by treatment with Klenow enzyme and religated to form pHK002; this removed the XbaI site in the polylinker. Site-directed single-stranded mutagenesis was used to remove an XbaI site in SSY1 (oligonucleotide POL95-036) (pHK003) and to introduce new XbaI-sites (oligonucleotides POL95-037, POL95-038, and POL95-039) at the desired positions, creating pHK004, pHK005, and pHK006. An XbaI-flanked cloning cassette, encoding the HA epitope (66) reiterated three times (HA3), was inserted into these unique XbaI sites to form pHK010, pHK033, and pHK034. pHK013 was constructed by ligating a KpnI/SacII fragment containing SSY1-HA1 from pHK010 into KpnI/SacII-digested pRS202 (10). Similarly, the XbaI-flanked HA cloning cassette was inserted into a unique XbaI site (pHK024) previously introduced into PTR3 (oligonucleotide POL96-045), creating plasmid pHK018. The HA3 epitope is placed between amino acids 157 and 158 of Ptr3p.
Genetic analysis.
Strain PLY1 was used to isolate spontaneous mutants resistant to 30 mM histidine (43). The super-high-histidine-resistant (shr) mutants were backcrossed to PLY4 (MATα his4Δ29 ura3-52 ade2Δ1::URA3), an isogenic derivative of PLY1. Tetrad analysis indicated that the mutant phenotypes segregated 2:2. Strains PLAS7-4C (shr10-7), PLAS6-4D (shr6-6), and PLAS14-1A (shr6-14) were obtained as meiotic segregants from these crosses. SHR6 and SHR10 were cloned by complementation of the 30 mM histidine-resistant phenotype exhibited by strains PLAS14-1A (shr6-14) and PLAS7-4C (shr10-7), respectively. These strains were transformed with a plasmid library (54), and Ura+ transformants unable to grow on selective media containing 30 mM histidine were identified. The complementing plasmids were isolated and further analyzed. Both strands of a 5-kb EcoRI fragment (pPL193) that complemented all available shr6 alleles were sequenced and shown to contain a single complete open reading frame (ORF) (YFR029w). One strand of a 3.9-kb SpeI/ClaI fragment (pPL356) that complemented all available shr10 alleles was sequenced and shown to contain one ORF (YDR160w).
Subcellular fractionation.
Cells expressing functional epitope-tagged SSY1-HA1 and PTR3-HA1 were grown in SC lacking uracil to an optical density at 600 nm (OD600) of 0.8. Cells were harvested by centrifugation; protein extracts were prepared and fractionated on 12 to 60% sucrose gradients as described by Egner et al. (20). Fractions were collected from the bottom of the gradients by using a fraction recovery system (Beckman). Proteins from equal aliquots of the collected fractions were concentrated by trichloroacetic acid precipitation, separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and blotted onto nitrocellulose membranes. Blots were incubated for 2 h with primary antibody in blocking buffer diluted as follows: 12CA5 ascites fluid (anti-HA monoclonal), 1:1,500; rabbit anti-Dap2p, 1:2,000; rabbit anti-Pma1p, 1:3,000; rabbit anti-Wbp1p, 1:1,000; rabbit anti-Kex2p, 1:1,000. Blots were washed three times for 15 min each with washing buffer (phosphate-buffered saline, 0.1% Tween-20), and incubated with horseradish peroxidase-coupled secondary antibody, either anti-mouse (Amersham) or anti-rabbit (Jackson), diluted 1:5,000 or 1:10,000 in washing buffer. Blots were washed three times for 15 min each with washing buffer, and immunoreactive proteins were visualized by chemiluminescence detection reagents (ECL or ECL-PLUS Western blotting detection systems; Amersham).
Protein manipulations.
Protein was determined by the method of Markwell et al. (47); whole-cell protein was determined in lysates of cells boiled in 0.1 M NaOH. Ptr3p membrane association was examined in whole-cell lysates prepared from strain HKY31 transformed with pHK018 grown in SC lacking uracil as described by Chang and Slayman (7). Aliquots of lysate (100 μg of protein) in low-salt BB buffer (0.3 M sorbitol, 5 mM MgCl2, 5 mM Tris [pH 7.5]) were diluted 1:1 with either H2O, 1.6 M urea, 0.6 M NaCl, 0.2 M Na2CO3 (pH 11.3), 2 mM EDTA, or low-salt BB buffer, mixed, and incubated on ice for 30 min. Samples were centrifuged at 100,000 × g for 1 h at 4°C, and protein pellets were resuspended in 2× sample buffer. After sonication, denaturation, and incubation for either 10 min at 50°C or 3 min at 95°C, aliquots (10 μg of protein) were resolved by SDS-PAGE and analyzed by immunoblotting. To determine whether Ptr3p is an intracellular protein, cells were grown in SC lacking uracil to an OD600 of 0.8, harvested, washed once, resuspended in spheroplasting buffer containing 10 mM NaN3 and 0.3 mg of Zymolyase-100T per ml, and incubated at 30°C. Spheroplasts (corresponding to 100 μg of protein) were placed on ice and treated for 10 min with 0 to 100 μg of proteinase K per ml in the presence or absence of 1% Triton X-100. Proteolysis was stopped by the addition of 2 mM phenylmethylsulfonyl fluoride (PMSF), and proteins resolved by SDS-PAGE were analyzed by immunoblotting.
Amino acid uptake and pool size determination.
Cells were grown in SUD containing uracil and histidine to an OD600 of 0.8, and amino acid uptake rates were assayed as described by Ljungdahl et al. (43). The initial uptake rates were determined at substrate concentrations of 10 and 0.004 mM; two 14C-labeled stock solutions (0.25 and 125 mCi mmol−1) were used to obtain the desired final concentrations. Citrulline uptake was measured at a substrate concentration of 0.017 mM (57.8 mCi mmol−1) in cells grown to an OD600 as described in the figure legends in either YPD, SC, SQD, or SED. Subsamples were removed at 30, 90, and 180 s; the uptake rate for each amino acid and citrulline was linear throughout the subsampling period. Uniformly 14C-labeled l-amino acids and l-ureido-14C]citrulline were obtained from Amersham and NEN-Dupont, respectively. Whole-cell and vacuolar amino acid pool concentrations were determined in cells grown in YPD to an OD600 of ≈1 essentially as described by Ohsumi et al. (51). Appropriate quantities of cultures (3 × 108 cells) were harvested by centrifugation; cell pellets were washed twice with 1.5 ml of water and resuspended in 1.5 ml of AA buffer (0.6 M sorbitol, 2.5 mM potassium phosphate buffer [pH 6]) containing 10 mM glucose. For the determination of vacuolar amino acid pools, the cells were resuspended in the same buffer containing 0.8 mM CuCl2 and incubated 10 min at 30°C. One-milliliter aliquots of cell suspensions were filtered (Whatman GF/F filters), and filters were washed four times with AA buffer. The washed filters were boiled in 3 ml of water for 15 min; 1-ml aliquots were centrifuged to remove particles of filter. The concentrations of amino acids in 30-μl aliquots were determined.
Northern analysis.
Strains PLY1, HKY37, and HKY38 were grown to OD600 of 0.8 in SC, washed once with water, resuspended in a 10× volume of YPD, SC, SD, SPD, SLD, SQD, SED, or SUD, and grown to an OD600 of 0.8. Total RNA was prepared as described by Elder et al. (21). Ten-microgram aliquots of denatured RNA were separated by agarose electrophoresis using a formaldehyde buffer system essentially as described in reference 46 and transferred to a nylon membrane (Hybond-N+; Amersham) in 10× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate). Blots were rinsed with 4× SSC and prehybridized for 5 h at 55°C in Church buffer (7% SDS, 1% bovine serum albumin, 1 mM EDTA, 250 mM NaPi [pH 7.2]) (8). Radioactive probes were prepared as follows. An 800-bp EcoRI fragment of GAP1, excised from pPL247 (43), and a 1.65-kb BamHI/HindIII fragment containing ACT1 (50) were purified from agarose gels (GELase; Amersham). The following DNA fragments were obtained by PCR (the oligonucleotides used to prime the reactions are indicated in parentheses; 25 ng of genomic DNA isolated from yeast strain S288C served as the template): a 222-bp fragment from the N terminus of DIP5 (POL97-038 and POL97-039), a 419-bp fragment from the N-terminal region of GNP1 (POL97-042 and POL97-043), and a 569-bp fragment of CAR1 (POL98-012 and POL98-013). Twenty nanograms of a plasmid genomic yeast DNA library was used as the template for amplifying a 530-bp fragment from PTR2 (POL97-048 and POL97-049). PCR products were analyzed by restriction analysis to ensure their identity and gel purified. DNA fragments were labeled with [α-32P]dCTP (3,000 Ci/mmol; Amersham), using a random-primed DNA labeling kit (MBI Fermentas Molecular Biology). Hybridizations were carried out in Church buffer at 55°C overnight (106 cpm/ml). Blots were rinsed once with 5× SSC, washed twice for 20 min each time with 5× SSC, and washed twice for 20 min each time with 0.5× SSC at 55°C. The amount of radioactivity was quantitated with a Fujix BAS1500 bioimage analyzer (Fuji Photo Film Co., Ltd., Tokyo, Japan). After background correction, signal strengths were normalized to the levels of actin mRNA present in RNA preparations.
Invasive growth assay.
Strains were patched on 1-day-old YPD plates (2% agar) and incubated at 30°C. After 2 days of incubation, plates were washed under a stream of running water as the surface was gently rubbed with a finger to remove cells not in the agar, and invasive growth was scored (24). Morphologies of invasive cells were microscopically examined after growth on YPD for 5 days.
RESULTS
ssy1 and ptr3 mutations result in histidine resistance.
Yeast strains carrying mutations in SHR6 and SHR10 were isolated in a genetic selection for shr mutants resistant to 30 mM histidine (43) and exhibit an identical level of resistance. SHR6 and SHR10 were cloned by complementation of the 30 mM histidine-resistant phenotype exhibited by strains PLAS14-1A (shr6-14) and PLAS7-4C (shr10-7), respectively (see Materials and Methods). Subsequent sequence analysis indicated that SHR6 is identical to ORF YFR029w, also known as PTR3 (5) and SSY3 (35), and SHR10 is identical to ORF YDR160w, previously identified as SSY1 (35).
ssy1 and ptr3 null mutations are synthetic lethal in combination with leu2.
Deletion alleles of SSY1 and PTR3 were created by removal of protein coding sequences and replacement of deleted segments with the selectable marker URA3. These constructs, ssy1Δ12::hisG-URA3-kanr-hisG and ptr3Δ14::hisG-URA3-kanr-hisG, were individually introduced into the diploid strain HKDY1 (HIS3/his3Δ200 LEU2/leu2-3,112 ura3-52/ura3-52 lys2Δ201/lys2Δ201) by transformation. Stable Ura+ transformants were selected and sporulated. Tetrads were dissected on both YPD and synthetic minimal dextrose medium (SD; minimal medium supplemented only with auxotrophic requirements). Spore viability was excellent on SD medium. The URA3 marker segregated 2:2 and in Leu+ spores was 100% linked to resistance to 30 mM histidine, showing that the deletion of either SSY1 or PTR3 leads to a resistance phenotype. Spore-derived colonies containing either ssy1Δ12 or ptr3Δ14 and the leu2 auxotrophic marker were not resistant to 30 mM histidine and did not grow on YPD. The synthetic lethality of ssy1Δ12 and ptr3Δ14 null mutations in combination with the leu2 auxotrophic allele was reflected in the pattern of spore inviability observed when tetrads were dissected onto medium with high concentrations of all amino acids, either YPD or SC.
A hallmark of mutations exhibiting pleiotropic effects on amino acid transport is that they manifest synthetic lethality on complex media when combined with mutations in genes encoding enzymes of amino acid biosynthetic pathways (23, 35, 43, 49). We surmise that the synthetic lethality on both YPD and SC is due to their high amino acid content. On these media, the overabundance of competing amino acids must interfere with the residual uptake mechanisms, effectively inhibiting uptake of the required amino acid. Additionally, Gap1p in cells grown on complex media is inactivated by posttranslational mechanisms (58). Thus, when grown on either YPD or SC, leu2 auxotrophic ssy1 or ptr3 mutants cannot synthesize leucine, nor can they import leucine from the external environment at rates sufficient to support growth.
SSY1 encodes a unique member of the AAP gene family that localizes to the PM in an SHR3-dependent manner.
As previously shown (32, 35), SSY1 encodes a unique member of the AAP gene family. Ssy1p is comprised of 852 aa, whereas the other 17 members of AAP family average 604 (±24) aa in length (range, 558 to 663). The sequence of Ssy1p is 22 to 28% identical to those of the other AAP gene family members. The homology between Ssy1p and the other AAPs begins with aa 278 of Ssy1p and stretches throughout the remaining 574 aa. The function of this unique N-terminal domain is not known.
We constructed HA3 epitope-tagged alleles of SSY1 inserted in frame between aa 34 and 35 (ssy1-HA2), 68 and 69 (ssy1-HA3), and 206 and 207 (SSY1-HA1). Cell lysates prepared from strain HKY20 individually transformed with CEN plasmid carrying these epitope-tagged constructs (pHK033, pHK034, and pHK010) expressed similar levels of HA-tagged proteins (data not shown). Although similar levels of HA-tagged proteins were detected, the ssy1-HA2 and ssy1-HA3 tagged alleles did not complement ssy1 mutations. The findings that in-frame insertions within the extended N-terminal region abolish Ssy1p activity without decreasing expressed protein levels suggest that the N-terminal domain of Ssy1p is important for function. In contrast, the SSY1-HA1 allele was judged functional based on its ability, whether inserted into a low-copy (pHK010) or a multicopy 2μm (pHK013) plasmid, to complement ssy1 mutant phenotypes. Specifically, SSY1-HA1 complements the 30 mM histidine-resistant phenotype, sensitivity to 30 mM lysine, and the synthetic lethality in combination with leu2 exhibited by ssy1 mutants.
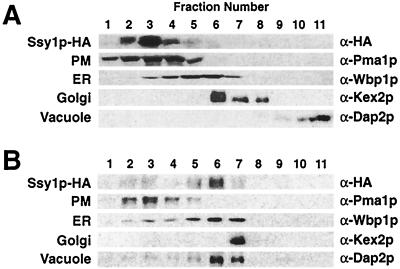
The functional HA epitope-tagged allele of SSY1 was used to monitor the subcellular fractionation of Ssy1p. Cell lysates were prepared from strain HKY20 carrying the CEN vector pHK010. The lysates were fractionated on 12 to 60% step sucrose gradients (20). Ssy1p-HA1 cofractionated with the PM marker protein, the plasma membrane ATPase (Pma1p [Fig. 1A]), indicating that Ssy1p is a component of the PM. Similar results were obtained for SSY1-HA1 inserted into the 2μm plasmid pHK013.
FIG. 1.
Ssy1p is a component of the PM that requires the AAP-specific packaging chaperone Shr3p to exit the ER. Cell lysates from strains HKY20 (A) and HKY29 (shr3Δ6) (B) expressing a functional HA epitope-tagged allele of SSY1 (pHK010) were fractionated on 12 to 60% step sucrose gradients. Proteins within fractions 1 to 11 were separated by SDS-PAGE and analyzed by immunoblotting. The antibodies recognizing marker proteins Pma1p, Wbp1p, Kex2p, and Dap2p were used to identify fractions containing PM, ER, Golgi, and vacuolar proteins, respectively. Although most membrane marker proteins reproducibly localized to specific fractions in fractionation experiments, the location of the vacuolar marker protein Dap2p occasionally varied in gradients; Dap2p localized to lighter fractions (as in panel A) or more dense fractions (as in panel B). The unpredictable behavior of Dap2p in gradients was not affected by Shr3p function, and Dap2p localization did not correlate with Ssy1p.
We examined whether the PM localization of Ssy1p was dependent on the AAP-specific packaging chaperonin Shr3p (38, 43). A cell lysate prepared from the shr3Δ6 ssy1Δ13 double-mutant strain HKY29 transformed with pHK010 was fractionated on a 12 to 60% step sucrose gradient. In these lysates, the bulk of the immunodetectable Ssy1p-HA1 was found in fractions 5 and 6, fractions that contained the highest levels of the ER marker protein Wbp1p (Fig. 1B). Thus, Ssy1p behaves similarly to the other members of the AAP gene family in that it requires Shr3p function for its localization to the PM.
Ptr3p is a peripheral membrane component of the PM.
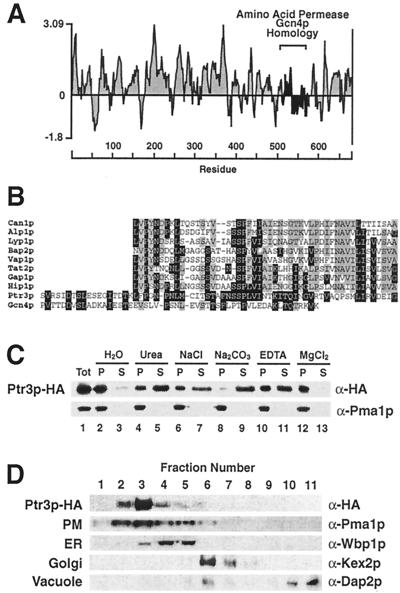
The PTR3 ORF encodes a protein of 678 aa with a calculated molecular mass of 76.4 kDa (5). Ptr3p is predicted to be a hydrophilic protein (Fig. 2A) that contains a small region (aa 511 to 575) exhibiting protein sequence homology to several AAPs and Gcn4p (Fig. 2B). Sequence similarities were found by comparing the last 200 aa of Ptr3p with the proteins in the yeast database, using the BLAST program (2) and the PAM120 similarity matrix at the Saccharomyces genome database, Stanford University. In pairwise comparisons using the Ptr3p sequence as the reference sequence, protein similarities were found to be statistical significant (≥5.5 standard deviations above the mean of 20 randomizations, using the comparison algorithm BESTFIT in the Genetics Computer Group Wisconsin Sequence Analysis Package). The homologous region of Gcn4p (aa 129 to 180) lies between the two well-characterized transcriptional activation (aa 107 to 125) and bZIP DNA binding (aa 226 to 281) domains (22, 30). Each of the homologous regions within AAPs (see Fig. 2 legend for amino acid coordinates) lies between putative membrane-spanning domains 7 and 8 and is predicted to be on the extracellular side of the PM.
FIG. 2.
PTR3 encodes a peripheral component of the PM. (A) Hydrophilicity plot of the predicted Ptr3p protein calculated using a window size of 11 amino acid residues (40); (B) Ptr3p shares protein sequence homology with Gcn4 and several members of the AAP family. The sequence alignments between Ptr3p (aa 511 to 575), the arginine permease (Can1p; aa 342 to 390), putative basic AAP (Alp1p; aa 326 to 374), lysine permease (Lyp1p, aa 364 to 412), branched-chain AAP (Bap2p; aa 346 to 395), valine/tyrosine permease (Vap1p; aa 339 to 389), tryptophan permease (Tat2p; aa 325 to 372), general AAP (Gap1p; aa 340 to 389), histidine permease (Hip1p; aa 340 to 390), and the general control regulatory protein Gcn4p (aa 129 to 180). The amino acid coordinates refer to amino acid residues in the proteins from which they originate. Amino acid residues identical to those in the Ptr3p sequence are indicated by white lettering on a black background; identical amino acid residues found in at least four sequences but not present in Ptr3p are highlighted with gray shading. The membrane association of Ptr3p was examined in whole-cell lysates prepared from strain HKY31 expressing PTR3-HA1 (C). Aliquots of lysate were diluted 1:1 with H2O, 1.6 M urea, 0.6 M NaCl, 0.2 M Na2CO3 (pH 11.3), 2 mM EDTA, or buffer containing 5 mM MgCl2, mixed, and incubated on ice for 30 min. Membrane pellet (P) and soluble (S) fractions were resolved by SDS-PAGE and analyzed by immunoblotting. As a control the membrane association of the PM ATPase (Pma1p) was monitored. The cellular localization of Ptr3p was determined by subcellular fractionation (D). A cell lysate from strain HKY31 expressing PTR3-HA1 was fractionated on a 12 to 60% step sucrose gradient and analyzed as described in Fig. 1.
A functional epitope-tagged allele of PTR3 (PTR3-HA1) that complemented the 30 mM histidine-resistant and 30 mM lysine-sensitive ptr3 mutant phenotypes was constructed. Cell lysates prepared from strain HKY31 transformed with pHK018 (PTR3-HA1) were fractionated. Although the predicted amino acid sequence of Ptr3p is characteristic of a soluble hydrophilic protein, Ptr3p-HA1 clearly fractionated as a membrane protein (Fig. 2C, lanes 1 to 3). We examined the membrane association of Ptr3p by treating lysates with reagents known to extract peripherally associated membrane proteins. The results (Fig. 2C, lanes 4 to 9) show that Ptr3p is extracted from membranes in the presence of 0.1 M Na2CO3 (pH 11.3); however, significant amounts of Ptr3p remain membrane associated in the presence of 0.8 M urea or 0.3 M NaCl, indicating that Ptr3p is a tightly associated peripheral membrane protein. During the subcellular fractionation experiments, we noticed that the membrane association of Ptr3p was sensitive to EDTA. Incubation of membrane preparations in the presence of 1 mM EDTA facilitated the dissociation of Ptr3p (Fig. 2C, lanes 10 and 11); conversely, the membrane association was stabilized in the presence of Mg2+ (Fig. 2C, lanes 12 and 13).
The intracellular location of Ptr3p was determined. A cell extract was prepared from strain HKY31 expressing PTR3-HA1 (pHK018). The prompt processing of the extract together with maintenance of high protein concentrations in the extract stabilized the membrane interaction. The lysate was fractionated on a 12 to 60% step sucrose gradient. Ptr3p-HA1 cofractionated with Pma1p, the PM marker protein (Fig. 2D), indicating that Ptr3p is a component of the PM. In additional experiments, we found that Ptr3p localized to the PM in both ssy1Δ and shr3Δ mutant strains (data not shown), indicating that the primary targeting of Ptr3p to the PM is independent of either Ssy1p or Shr3p function.
To determine whether Ptr3p was localized to the extra- or intracellular side of the PM, spheroplasts prepared from strain HKY31 expressing PTR3-HA1 were treated with proteinase K (data not shown). In the absence of detergent, Ptr3p was refractory to protease treatment. In the presence of 1% Triton X-100, the lowest concentration of proteinase K (1 μg ml−1) degraded all of the immunodetectable Ptr3p. These results suggest that Ptr3p is localized to the cytosolic face of the PM. Consistent with these observations Ptr3p lacks a recognizable signal sequence.
Mutations in SSY1 and PTR3 exhibit pleiotropic effects on amino acid uptake.
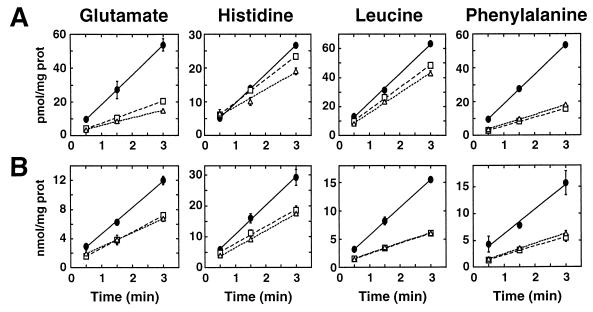
Amino acid uptake into isogenic wild-type and ptr3Δ15 and ssy1Δ13 mutant cells was determined in strains pregrown in SUD. Uptake rates were determined at low (4 μM) and high (10 mM) substrate concentrations. The initial rates of uptake of four representative amino acids are shown in Fig. 3, and a quantitative summary of the data is presented in Table 3. At 4 μM substrate (Fig. 3A), ptr3 and ssy1 mutations exhibited a pleiotropic effect on several specific uptake systems. Most notably, glutamate and phenylalanine uptake was reduced by 70% in ptr3 and ssy1 mutants. Glutamate-specific transport is mediated by the dicarboxylic acid permease (Dip5p [53]). The phenylalanine-specific permease has not been defined. Reductions in the rates of histidine, leucine, and lysine uptake were also observed (Table 3), indicating that the histidine-specific permease (Hip1p [62]), the branched-chain AAP (Bap2p [26]), and the lysine-specific permease (Lyp1p [61]), respectively, are affected by mutations in SSY1 or PTR3. At 10 mM substrate (Fig. 3B; Table 3), the uptake of amino acids occurs predominantly through Gap1p. ssy1 and ptr3 mutations reduce Gap1p activity by 50%. The rates of arginine uptake were relatively unaffected by ssy1 and ptr3 mutations (Table 3).
FIG. 3.
Amino acid uptake into wild-type and ssy1Δ13 and ptr3Δ15 null mutant strains. Wild-type (PLY1; ●), ssy1Δ13 (HKY37; □), and ptr3Δ15 (HKY38; ▵) strains were grown in SUD containing histidine and uracil. The uptakes of the indicated amino acids were assayed as described in Materials and Methods. (A) High-affinity amino acid uptake determined at amino acid concentrations of 4 μM; (B) low-affinity amino acid uptake determined at amino acid concentrations of 10 mM. Rate measurements were determined in duplicate samples; error bars represent 1 standard deviation.
TABLE 3.
Amino acid uptake into wild-type, ssy1Δ, and ptr3Δ strains
| Amino acid | Uptake
|
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| With 4 μM substrate (pmol mg−1 min−1)
|
Folda
|
With 10 mM substrate (nmol mg−1 min−1)
|
Fold
|
|||||||
| Wild type | ssy1Δ13 | ptr3Δ15 | ssy1Δ | ptr3Δ | Wild type | ssy1Δ13 | ptr3Δ15 | ssy1Δ | ptr3Δ | |
| Leucine | 20.2 ± 0.3 | 15.5 ± 1.0 | 13.9 ± 0.6 | 0.8 | 0.7 | 4.9 ± 0.1 | 1.8 ± 0.0 | 1.8 ± 0.1 | 0.4 | 0.4 |
| Phenylalanine | 17.5 ± 0.4 | 5.1 ± 0.4 | 5.7 ± 0.0 | 0.3 | 0.3 | 4.6 ± 1.4 | 1.7 ± 0.5 | 1.9 ± 0.2 | 0.4 | 0.4 |
| Glutamate | 17.6 ± 1.1 | 6.5 ± 0.2 | 4.5 ± 0.3 | 0.4 | 0.3 | 3.6 ± 0.3 | 2.2 ± 0.3 | 1.9 ± 0.1 | 0.6 | 0.5 |
| Histidine | 8.6 ± 0.4 | 6.9 ± 0.3 | 5.1 ± 0.9 | 0.8 | 0.6 | 9.3 ± 1.1 | 5.5 ± 0.0 | 5.4 ± 0.2 | 0.6 | 0.6 |
| Lysine | 22.8 ± 0.7 | 18.9 ± 3.0 | 13.2 ± 0.2 | 0.8 | 0.6 | 2.3 ± 0.2 | 2.3 ± 0.2 | 2.3 ± 0.3 | 1.0 | 1.0 |
| Arginine | 245 ± 7.5 | 281 ± 5.5 | 281 ± 7.0 | 1.1 | 1.1 | 1.2 ± 0.2 | 1.3 ± 0.2 | 1.3 ± 0.2 | 1.1 | 1.1 |
Ratio with respect to wild type.
ssy1 and ptr3 mutations affect mRNA levels of multiple genetically distinct AAPs and the peptide transporter (PTR2).
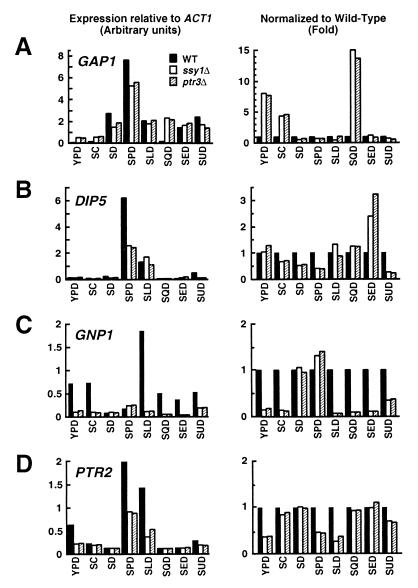
We examined the steady-state mRNA levels of GAP1, GNP1 (encoding glutamine permease [67]), DIP5, and PTR2 (peptide transporter gene [52]) in isogenic wild-type and ssy1Δ13 and ptr3Δ15 mutant cells. Strains grown in ammonium-containing SC were used to inoculate media containing various nitrogen sources. The starting OD600 in each medium was adjusted to 0.08, and RNA was isolated from cells when cultures reached an OD600 of 0.8. Expression levels were analyzed by Northern analysis. Under the growth conditions used, the mutations in SSY1 and PTR3 did not adversely affect growth. In all media except SED (see Fig. 6), wild-type and mutants strains adjusted to the shift in nitrogen source similarly and grew at similar rates. The levels of expression were quantitated by phosphorimaging, and ACT1 transcript levels were used to standardize quantitations. Regardless of the nitrogen source used in the growth media, the levels of ACT1 transcripts per OD260 of RNA were similar in RNA isolated from wild-type and mutant cells. Results are presented in two formats: relative to the levels of ACT1 mRNA (Fig. 4, panels on left) and normalized to wild-type levels of expression (Fig. 4, panels on right).
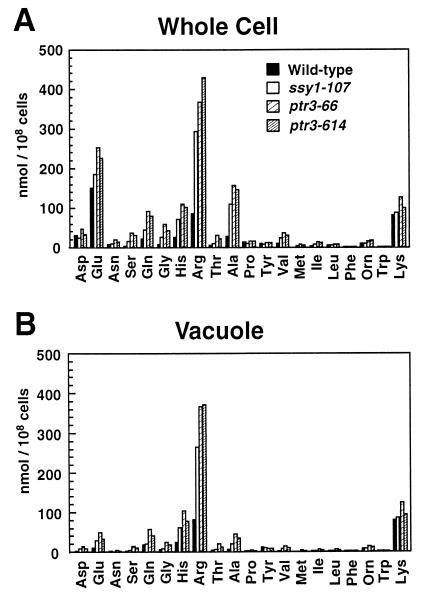
FIG. 6.
Amino acid concentrations in whole cells and in vacuolar pools of wild-type (PLAS1-7D), ssy1-107 (PLAS7-4C), ptr3-66 (PLAS6-4D), and ptr3-614 (PLAS14-1A) strains grown to a density of 2 × 107 cells/ml in YPD. Whole-cell (A) and vacuolar (B) amino acid concentrations were determined as described in Materials and Methods.
FIG. 4.
Mutations in SSY1 and PTR3 affect the steady-state mRNA levels of multiple permeases. Total RNA isolated from PLY1 (wild type [WT]), HKY37 (ssy1Δ), and HKY38 (ptr3Δ) was analyzed by Northern analysis. Expression levels of GAP1 (A), DIP5 (B), GNP1 (C), and PTR2 (D) were determined in the strains grown to an OD600 to 0.8 in media containing alternate nitrogen sources as indicated. Signal strengths (arbitrary units) relative to the levels of actin mRNA (ACT1) after background correction are plotted in the panels to the left; mRNA levels normalized to the expression observed in wild-type cells are replotted in the panels on the right.
When grown on YPD, SC, and SQD, ptr3 and ssy1 mutant strains expressed 4- to 15-fold higher levels of GAP1 mRNA than the isogenic wild-type strain (Fig. 4A). Conversely, when grown on SD, SPD, and to a lesser extent SUD, mutant strains have lower levels of GAP1 mRNA (Fig. 4A). Similarly, DIP5 expression in the mutants increased or decreased depending on the nitrogen source (Fig. 4B). On SED, ssy1 and ptr3 mutants have two- to threefold more DIP5 mRNA than the wild-type strain; on SPD, the mutants contain only 50% of the wild-type DIP5 transcripts (Fig. 4B). In most instances, the expression of GNP1 was found to depend on Ssy1p and Ptr3p function. The wild-type strain had 2- to 18-fold higher levels of GNP1 mRNA compared to either ssy1 or ptr3 null mutant strains (Fig. 4C). GNP1 was expressed in wild-type strains only when grown on media containing amino acids; no significant GNP1 expression was observed on SD lacking amino acids. The expression of PTR2 was also found to be substantially reduced in mutant strains grown in YPD, SPD, and SLD (Fig. 4D). Didion et al. (17) and Iraqui et al. (32) have observed similar Ssy1p-dependent expression of AGP1, BAP2, BAP3, TAT1, and TAT2.
In summary, our data indicate that mutations in either SSY1 and PTR3 exert identical, direct or indirect effects on the steady-state transcript levels of multiple permeases. Based on our results and studies with gap1Δ ssy1Δ double-deletion mutants (17, 32), the expression of at least a dozen different, genetically distinct AAPs and the peptide transporter (PTR2) appear to be affected by the combined function of Ssy1p and Ptr3p.
Amino acid-dependent nonspecific induction of arginase (CAR1) expression requires Ssy1p and Ptr3p.
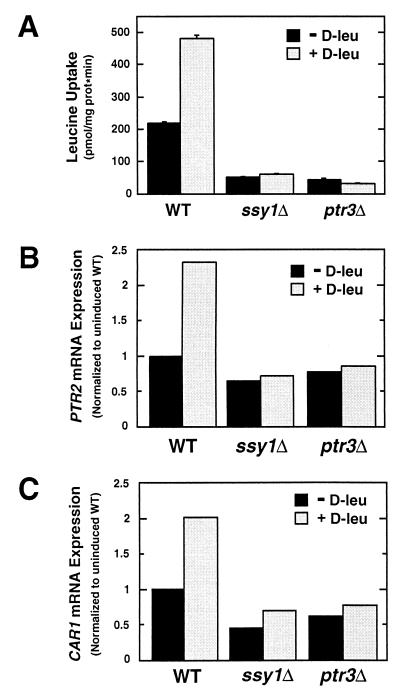
We examined d-leucine-stimulated l-leucine uptake, peptide transporter (PTR2) expression, and arginase (CAR1) expression in isogenic wild-type and ssy1 and ptr3 null mutant strains carrying a gap1 null mutation (Fig. 5). FGY58 (wild type), HKY63 (ssy1Δ12), and HKY65 (ptr3Δ14) were grown in ammonia-based SD containing uracil, lysine, and adenine to an OD600 of 1. Cells were harvested, resuspended in fresh medium lacking or containing 0.15 mM d-leucine, and incubated for 30 min at 30°C. As previously described (5, 17), preincubation of wild-type cells in the presence of d-leucine resulted in a twofold stimulation of l-leucine uptake (Fig. 5A) and accumulation of PTR2 transcripts (Fig. 5B). Similarly, d-leucine induced a twofold increase in the level of CAR1 transcripts in wild-type cells (Fig. 5C). In contrast to wild-type cells, preincubation with d-leucine did not stimulate l-leucine uptake or increase PTR2 or CAR1 transcript levels in either ssy1 or ptr3 mutant cells. These results provide the first indication that cells may use Ssy1p- and Ptr3p-derived signals to regulate the expression of non-transport-related genes. The gap1 null mutant strains used in these experiments, although unable to grow on citrulline as the sole nitrogen source, exhibited substantial d-leucine uptake. Thus, we were unable to determine whether the observed regulatory affects occurred in the absence of d-leucine transport.
FIG. 5.
d-Leucine-stimulated transport and nonspecific induction of arginase expression require Ssy1p and Ptr3p. gap1 null mutant strains FGY58 (wild type [WT]), HKY63 (ssy1Δ), and HKY65 (ptr3Δ) were grown in SD containing uracil, lysine, and adenine to an OD600 of 1. Cells were harvested and resuspended in fresh medium lacking or containing 0.15 mM d-leucine (D-leu). After incubation for 30 min at 30°C, leucine uptake (A) and PTR2 (B) and CAR1 (C) mRNA transcript levels were measured.
Physiological consequences of ptr3 and ssy1 mutations. (i)ssy1 and ptr3 mutants have increased vacuolar pools of histidine and arginine.
The pool sizes of amino acids were measured in whole cells and vacuoles in wild-type (PLY1), ssy1 (PLAS7-4C), and ptr3 (PLAS6-4D and PLAS14-1) strains (Fig. 6). In wild-type cells, glutamate was the predominant amino acid, whereas in the mutant strains, arginine was found to accumulate at the highest concentrations. Arginine levels in both ssy1 and ptr3 mutants were three- to fivefold higher than those in the wild type (Table 4). Histidine levels were also higher (two- to fourfold), whereas lysine levels were unaffected (Table 4). In addition to the increased levels of histidine and arginine, the mutant strains contained substantially higher concentrations of serine, glutamine, glycine, and alanine. The levels of other amino acids remained relatively unchanged.
TABLE 4.
Concentrations of basic amino acids in vacuoles of wild-type, ssy1, and ptr3 strains
| Amino acid | PLY1 (wild type) (nmol/108 cells) | PLAS7-4C (ssy1-107)
|
PLAS6-4D (ptr3-66)
|
PLAS14-1A (ptr3-614)
|
|||
|---|---|---|---|---|---|---|---|
| nmol/ 108 cells | Folda | nmol/ 108 cells | Fold | nmol/ 108 cells | Fold | ||
| Histidine | 24.0 | 61.9 | 2.6 | 104.9 | 4.4 | 77.7 | 3.2 |
| Arginine | 81.1 | 265.9 | 3.3 | 366.8 | 4.5 | 370.8 | 4.6 |
| Lysine | 79.9 | 85.4 | 1.1 | 124.3 | 1.6 | 94.7 | 1.2 |
Ratio with respect to wild-type concentrations.
As expected, greater than 90% of the basic amino acids (arginine, histidine, and lysine) were recovered in the vacuolar fraction (Fig. 6B). The acidic amino acids (aspartate and glutamate) were excluded from the vacuole and maintained in cytosolic pools. The remaining amino acids were found relatively evenly distributed between the cytosol and vacuole. The intracellular distribution of amino acids that we observed is consistent with previous studies (36). Our data indicate that ssy1 and ptr3 mutations result in an increased capacity to compartmentalize basic amino acids, but do not affect the intracellular distribution of amino acids between the vacuole and cytosol.
(ii) Growth on glutamate as sole nitrogen source.
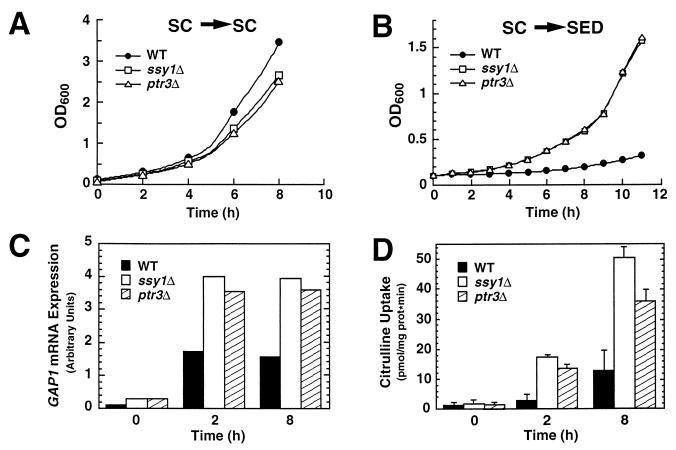
We examined the consequences of shifting wild-type (PLY1), ptr3Δ15 (HKY38), and ssy1Δ13 (HKY37) cells from media containing ammonia to media containing various amino acids as sole nitrogen sources. A particularly striking effect was observed when cells were shifted to SED (Fig. 7). Strains PLY1 (wild type), HKY38 (ptr3Δ) and HKY37 (ssy1Δ) were pregrown in SC (OD600 of 0.8). Cells washed once with water were used to inoculate fresh SC and SED to an OD600 of ≈0.15. Wild-type and mutant strains grew at approximately the same rate in SC (Fig. 7A). However, when shifted to glutamate (SED), the mutant strains grew without substantial delay, whereas the wild-type strain required a period of greater than 10 h before noticeable growth was observed (Fig. 7B).
FIG. 7.
Physiological consequences of ptr3Δ15 and ssy1Δ13 null mutations. Strains PLY1 (wild type [WT]), HKY37 (ssy1Δ), and HKY38 (ptr3Δ) were pregrown in SC. Cells, washed once with water, were used to inoculate either fresh SC (A) or SED (B); growth was monitored spectrophotometrically (OD600). GAP1 mRNA levels (C) and Gap1p activity (D) in cells shifted to SED were determined at the times indicated.
Cells require a functional Gap1p to sustain high growth rates on media containing amino acids as the predominant nitrogen sources (12, 36, 45, 65). It is known that although wild-type cells express high levels of GAP1 mRNA in SED (compare GAP1 mRNA levels in Fig. 4A, left panel), Gap1p activity is maintained low, presumably posttranslational regulatory circuits maintain inactivate Gap1p (58). In our experiments, mutant strains growing on SED initially had higher levels of GAP1 mRNA (mutant cells have twofold more GAP1 mRNA [Fig. 7C, time zero) and perhaps other nitrogen-regulated genes required for glutamate utilization. Additionally, mutant strains more rapidly expressed functionally active Gap1p (at 2 h, citrulline transport occurred at rates sixfold greater than in wild-type cells [Fig. 7D]). Thus, the absence of growth inhibition of mutant cells when shifted from ammonium-based SC to SED correlates with increased levels of GAP1 transcription and a concomitant increase in Gap1p activity.
(iii) ptr3 and ssy1 mutants exhibit enhanced haploid invasive but not diploid pseudohyphal growth.
In response to nutrient availability, growing yeast cells engage distinct developmental pathways leading to vegetative growth or filamentous-like growth. To examine whether Ssy1p- or Ptr3p-derived signals affect developmental outcomes, we constructed Σ1278b-derived ptr3 (HKY43) and ssy1 (HKY45) null mutant strains. The Σ1278b-derived mutants are resistant to 30 mM histidine. When propagated on YPD, haploid Σ1278b-derived ptr3 (HKY43) and ssy1 (HKY45) mutant strains exhibited enhanced invasive growth compared to an isogenic wild-type strain (10480-5C) (Fig. 8A, upper sectors). The invasive growth was dependent on the Σ genetic background; ssy1 and ptr3 strains in an S288C background did not invade the agar (Fig. 8A, lower sectors). The invasive growth phenotype exhibited by strains HKY43 and HKY45 was accompanied by changes in cell morphology. Invasively growing cells were elongated and have an increased axial (length/width) ratio. These elongated cells remained attached, leading to the formation of filaments of cells (Fig. 8B). We quantitated this phenotype by taking photographs of cells remaining in the agar after washing away surface growth. Cells present in areas of similar cell densities were counted, and the number of cells with an axial ratio greater than 2 was noted (Table 5). In mutant strains, there was a >20-fold increase in the numbers of elongated cells.
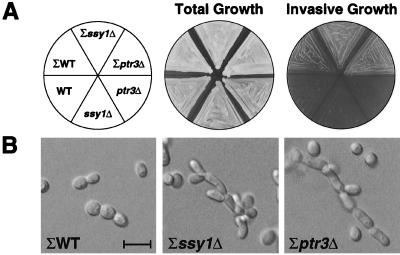
FIG. 8.
ssy1Δ and ptr3Δ mutants exhibit enhanced haploid-specific invasive growth. (A) Σ background strains 10480-5C (wild type [ΣWT]), HKY45 (Σssy1Δ), and HKY43 (Σptr3Δ) and S288C background strains PLY1 (WT), HKY37 (ssy1Δ), and HKY38 (ptr3Δ) were patched on solid YPD and incubated for 2 days at 30°C. The plate was photographed before (total growth) and after (invasive growth) cells were washed off the agar surface. (B) Σ background strains (10480-5C, HKY45, and HKY43) were grown on solid YPD incubated at 30°C for 5 days and invasively growing cells were photographed. The 10-μm scale bar applies to all three photographs.
TABLE 5.
Percentage of elongated (axial [length/width] ratio > 2) cells during invasive growth
| Genotype (strain) | No. of cells counted | No. of elongated cells | % Elongated cells |
|---|---|---|---|
| Wild type (10480-5C) | 537 | 3 | 0.6 |
| ssy1Δ12 (HKY45) | 518 | 97 | 19 |
| ptr3Δ14 (HKY43) | 451 | 83 | 18 |
Homozygous shr3 diploid mutants exhibit extensive pseudohyphal growth (25). We examined whether the enhanced pseudohyphal growth exhibited by shr3 mutant strains is due to decreased levels of Ssy1p (Fig. 1B) or Ptr3p in the PM. Homozygous ssy1/ssy1 and ptr3/ptr3 diploid mutant strains were constructed in the Σ background, these mutant strains did not exhibit enhanced filamentous growth (data not shown). Under similar conditions, extensive filamentous growth surrounding colonies of a Shr3− diploid strain HKDY15 was observed. These results indicate that the enhanced pseudohyphal growth exhibited by shr3/shr3 strains is not a consequence of reduced PM levels of Ssy1p or Ptr3p.
DISCUSSION
We have found that mutations in two genes, SSY1 (SHR10) and PTR3 (SHR6), cause the nitrogen-regulated general AAP gene (GAP1) to be aberrantly expressed and block the nonspecific induction of arginase (CAR1). ssy1 and ptr3 mutations also pleiotropically affect the steady-state levels of multiple specific amino acid transporter mRNA transcripts and diminish the expression of the peptide transporter (PTR2). Additionally, ssy1Δ and ptr3Δ mutants have increased vacuolar pools of histidine and arginine. The resistance of these mutants to high concentrations of histidine is likely a consequence of the altered uptake and increased capacity to compartmentalize histidine. The observations that mutations in SSY1 and PTR3 manifest identical phenotypes and ssy1Δ ptr3Δ double mutants do not exhibit additive effects suggest that Ssy1p and Ptr3p function in the same pathway.
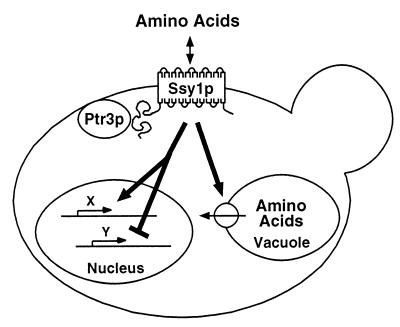
Our data extend previous work (5, 17, 32), suggesting that Ssy1p and Ptr3p are components of an amino acid sensing system. The assignment of Ssy1p and Ptr3p as components of an extracellular amino acid sensor rests on a number of observations. First, mutations in these genes, unlike those in the AAP genes, manifest identical pleiotropic alterations in amino acid uptake (Fig. 3; Table 3) and vacuolar amino acid pools (Fig. 6; Table 4). Second, although Ssy1p is clearly a member of the AAP family and requires Shr3p to localize to the PM (Fig. 1), it contains a functionally important N-terminal extension absent from other family members. Third, the proper induction of many AAPs requires Ssy1p and Ptr3p (Fig. 4 and 5) and occurs without detectable amino acid uptake (17, 32). Our finding that Ptr3p, which is not predicted to be a hydrophobic protein, fractionates as a component of the PM (Fig. 2C and D) directly implicates this protein as a constituent of this sensing system. As both Ssy1p and Ptr3p are localized to the plasma membrane, they could interact, although there is as yet no evidence for a physical association between them. According to this model (Fig. 9), yeast cells use the PM Ssy1p/Ptr3p sensing system to regulate diverse metabolic processes important for proper amino acid uptake and compartmentalization, two processes that enable cells to maintain cytosolic amino acid pools.
FIG. 9.
Model for Ssy1p and Ptr3p function. See text for details.
The localization of Ptr3p to the cytosolic face of the PM and the observed sequence homology between Ptr3p and several AAPs and Gcn4p (Fig. 2B) raises several possibilities. The homologous region within Ptr3p may function as part of an amino acid binding site that regulates Ptr3p activity. It is possible that Ssy1p transports regulatory amounts of amino acids into the cell, and that this transport is coupled so that amino acids are directed to this putative regulatory domain. Alternatively, Ptr3p may function in sensing cytoplasmic levels of amino acids, providing a regulatory loop that modulates the signals generated by Ssy1p. Finally, we have found that under certain conditions Ptr3p dissociates from the PM (Fig. 2C), thus the possibility exists that the regulatory events controlled by the Ssy1p/Ptr3p sensor require that Ptr3p disengage from the membrane. After activation, presumably the consequence of a Ssy1p-dependent event, regulatory amounts Ptr3p could localize to other regions of the cell to directly exert a controlling function.
ssy1 and ptr3 mutants express elevated levels of GAP1 mRNA (Fig. 4A) when grown on amino acid-rich medium (either YPD or SC) and in medium containing glutamine as the sole nitrogen source (SQD). These results indicate that Ssy1p/Ptr3p are part of a pathway that can negatively regulate GAP1 expression. Two pathways converge to control nitrogen-regulated genes, one sensitive to the presence of ammonium (Ure2p and Gln3p) and the other sensitive to the presence of amino acids (Nil2p and Nil1p) (6, 55, 57). Regulation by Ssy1p/Ptr3p-derived signals is independent of ammonia repression: ssy1/ptr3 mutants grown in SD without amino acids but the same level of ammonia as SC express similar levels of GAP1 as wild-type cells (Fig. 4A). As the expression pattern of Gap1p in ssy1 and ptr3 mutants mimics that found in a strain lacking the Nil2p repressor (55), the Ssy1p/Ptr3p-derived signals may be mediated through the Nil2p/Nil1p pathway.
The observation that haploid ssy1 and ptr3 strains exhibit enhanced invasive growth is consistent with these proteins being components of an amino acid sensor. Presumably, the enhanced invasiveness is a consequence of the erroneous sensing in mutant cells of the availability of amino acids in the extracellular environment. Our findings suggest that wild-type cells use Ssy1p- and Ptr3p-derived signals to moderate invasive growth. Similarly, the high-affinity ammonium permease (MEP2) is thought to act as an ammonium sensor that influences the frequency at which diploid cells enter the pseudohyphal growth pathway (44). The observation that nutrient sensors acting at the PM are required to initiate proper growth responses raises the possibility that yeast cells do not rely solely on intracellularly derived nutritional signals for making decisions affecting developmental outcomes.
ACKNOWLEDGMENTS
We thank Tom Stevens, Robert Fuller, Carolyn Slayman, and Stephan te Heesen for their generous gifts of antibodies to Dap2p, Kex2p, Pma1p, and Wbp1p, respectively. We thank C. Fredrik Gilstring for S. cerevisiae FGY58 and members of Ljungdahl laboratory for constructive comments, especially Marten Hammar for critical review of the manuscript.
This work was supported by NIH grants GM40266 and GM35010 (G. R. Fink) and the Ludwig Institute for Cancer Research (P. O. Ljungdahl).
REFERENCES
- 1.Allen J B, Elledge S J. A family of vectors that facilitate transposon and insertional mutagenesis of cloned genes in yeast. Yeast. 1994;10:1267–1272. doi: 10.1002/yea.320101003. [DOI] [PubMed] [Google Scholar]
- 2.Altschul S F, Gish W, Miller W, Myers E W, Lipman D J. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 3.André B. An overview of membrane transport proteins in Saccharomyces cerevisiae. Yeast. 1995;11:1575–1611. doi: 10.1002/yea.320111605. [DOI] [PubMed] [Google Scholar]
- 4.Antebi A, Fink G R. The yeast Ca2+-ATPase homologue, PMR1, is required for normal Golgi function and localizes in a novel Golgi-like distribution. Mol Biol Cell. 1992;3:633–654. doi: 10.1091/mbc.3.6.633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Barnes D, Lai W, Breslav M, Naider F, Becker J M. PTR3, a novel gene mediating amino acid-inducible regulation of peptide transport in Saccharomyces cerevisiae. Mol Microbiol. 1998;29:297–310. doi: 10.1046/j.1365-2958.1998.00931.x. [DOI] [PubMed] [Google Scholar]
- 6.Blinder D, Coschigano P W, Magasanik B. Interaction of the GATA factor Gln3p with the nitrogen regulator Ure2p in Saccharomyces cerevisiae. J Bacteriol. 1996;178:4734–4736. doi: 10.1128/jb.178.15.4734-4736.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chang A, Slayman C W. Maturation of the yeast plasma membrane [H+]ATPase involves phosphorylation during intracellular transport. J Cell Biol. 1991;115:289–295. doi: 10.1083/jcb.115.2.289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Church G M, Gilbert W. Genomic sequencing. Proc Natl Acad Sci USA. 1984;81:1991–1995. doi: 10.1073/pnas.81.7.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Coffman J A, Rai R, Loprete D M, Cunningham T, Svetlov V, Cooper T G. Cross regulation of four GATA factors that control nitrogen catabolic gene expression in Saccharomyces cerevisiae. J Bacteriol. 1997;179:3416–3429. doi: 10.1128/jb.179.11.3416-3429.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Connelly C, Hieter P. Budding yeast SKP1 encodes an evolutionarily conserved kinetichore protein required for cell cycle progression. Cell. 1996;86:275–285. doi: 10.1016/S0092-8674(00)80099-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Coons D M, Vagnoli P, Bisson L F. The C-terminal domain of Snf3p is sufficient to complement the growth defect of snf3 null mutations in Saccharomyces cerevisiae: SNF3 functions in glucose recognition. Yeast. 1997;13:9–20. doi: 10.1002/(SICI)1097-0061(199701)13:1<9::AID-YEA51>3.0.CO;2-U. [DOI] [PubMed] [Google Scholar]
- 12.Cooper T. Transport in Saccharomyces cerevisiae. In: Strathern J N, Jones E W, Broach J R, editors. The molecular biology of the yeast Saccharomyces: metabolism and gene expression. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1982. pp. 399–461. [Google Scholar]
- 13.Coornaert D, Vissers S, André B, Grenson M. The UGA43 negative regulatory gene of Saccharomyces cerevisiae contains both a GATA-1 type zinc finger and a putative leucine zipper. Curr Genet. 1992;21:301–307. doi: 10.1007/BF00351687. [DOI] [PubMed] [Google Scholar]
- 14.Courchesne W E, Magasanik B. Ammonia regulation of amino acid permeases in Saccharomyces cerevisiae. Mol Cell Biol. 1983;3:672–683. doi: 10.1128/mcb.3.4.672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cunningham T S, Cooper T G. The Saccharomyces cerevisiae DAL80 repressor protein binds to multiple copies of GATAA-containing sequences (URSGATA) J Bacteriol. 1993;175:5851–5861. doi: 10.1128/jb.175.18.5851-5861.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Daugherty J R, Rai R, Berry H M, Cooper T G. Regulatory circuit for response of nitrogen catabolic gene expression to the GLN3 and DAL80 proteins and nitrogen catabolic repression in Saccharomyces cerevisiae. J Bacteriol. 1993;175:64–73. doi: 10.1128/jb.175.1.64-73.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Didion T, Regenberg B, Jørgensen M U, Kielland-Brandt M C, Andersen H A. The permease homologue Ssy1p controls the expression of amino acid and peptide transporter genes in Saccharomyces cerevisiae. Mol Microbiol. 1998;27:643–650. doi: 10.1046/j.1365-2958.1998.00714.x. [DOI] [PubMed] [Google Scholar]
- 18.Dubois E, Messenguy F. Integration of the multiple controls regulating the expression of the arginase gene CAR1 of Saccharomyces cerevisiae in response to different nitrogen signals: role of Gln3p, ArgRp-Mcm1p, and Ume6p. Mol Gen Genet. 1997;253:568–580. doi: 10.1007/s004380050359. [DOI] [PubMed] [Google Scholar]
- 19.Dubois E L, Wiame J-M. Non specific induction of arginase in Saccharomyces cerevisiae. Biochimie. 1976;58:207–211. doi: 10.1016/s0300-9084(76)80371-9. [DOI] [PubMed] [Google Scholar]
- 20.Egner R, Mahé Y, Pandjaitan R, Kuchler K. Endocytosis and vacuolar degradation of the plasma membrane-localized Pdr5 ATP-binding cassette multidrug transporter in Saccharomyces cerevisiae. Mol Cell Biol. 1995;15:5879–5887. doi: 10.1128/mcb.15.11.5879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Elder R T, Loh E Y, Davis R W. RNA from the yeast transposable element TY1 has both ends in the direct repeats, a structure similar to retrovirus RNA. Proc Natl Acad Sci USA. 1983;80:2432–2436. doi: 10.1073/pnas.80.9.2432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ellenberger T E, Brandl C J, Struhl K, Harrison S C. The GCN4 basic region leucine zipper binds DNA as a dimer of uninterrupted α helices: a crystal structure of the protein-DNA complex. Cell. 1992;71:1223–1237. doi: 10.1016/s0092-8674(05)80070-4. [DOI] [PubMed] [Google Scholar]
- 23.Garrett J M. Characterization of AAT1: a gene involved in the regulation of amino acid transport in Saccharomyces cerevisiae. J Gen Microbiol. 1989;135:2429–2437. doi: 10.1099/00221287-135-9-2429. [DOI] [PubMed] [Google Scholar]
- 24.Gimeno C J, Fink G R. Induction of pseudohyphal growth by overexpression of PHD1, a Saccharomyces cerevisiae gene related to transcriptional regulators of fungal development. Mol Cell Biol. 1994;14:2100–2112. doi: 10.1128/mcb.14.3.2100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gimeno C J, Ljungdahl P O, Styles C A, Fink G R. Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell. 1992;68:1077–1090. doi: 10.1016/0092-8674(92)90079-r. [DOI] [PubMed] [Google Scholar]
- 26.Grauslund M, Didion T, Kielland-Brandt M C, Andersen H A. BAP2, a gene encoding a permease for branched-chain amino acids in Saccharomyces cerevisiae. Biochim Biophys Acta. 1995;1269:275–280. doi: 10.1016/0167-4889(95)00138-8. [DOI] [PubMed] [Google Scholar]
- 27.Grenson M. Amino acid transporters in yeast: structure, function and regulation. In: Pont J J L L M D, editor. Molecular aspects of transport proteins. Amsterdam, The Netherlands: Elsevier Science; 1992. pp. 219–245. [Google Scholar]
- 28.Guthrie C, Fink G R, editors. Methods in enzymology. 194. Guide to yeast genetics and molecular biology. San Diego, Calif: Academic Press, Inc.; 1991. [PubMed] [Google Scholar]
- 29.Herskowitz I, Jensen R E. Putting the HO gene to work: practical uses for mating-type switching. Methods Enzymol. 1991;194:132–146. doi: 10.1016/0076-6879(91)94011-z. [DOI] [PubMed] [Google Scholar]
- 30.Hope I A, Mahadevan S, Struhl K. Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein. Nature. 1988;333:635–640. doi: 10.1038/333635a0. [DOI] [PubMed] [Google Scholar]
- 31.Horák J. Yeast nutrient transporters. Biochim Biophys Acta. 1997;1331:41–79. doi: 10.1016/s0304-4157(96)00015-9. [DOI] [PubMed] [Google Scholar]
- 32.Iraqui I, Vissers S, Bernard F, De Craene J-O, Boles E, Urrestarazu A, André B. Amino acid signaling in Saccharomyces cerevisiae: a permease-like sensor of external amino acids and F-box protein Grr1p are required for transcriptional induction of the AGP1 gene, which encodes a broad-specificity amino acid permease. Mol Cell Biol. 1999;19:989–1001. doi: 10.1128/mcb.19.2.989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ito H, Fukuda Y, Murata K, Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983;153:163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jauniaux J C, Grenson M. GAP1, the general amino acid permease gene of Saccharomyces cerevisiae. Nucleotide sequence, protein similarity with the other bakers yeast amino acid permeases, and nitrogen catabolite repression. Eur J Biochem. 1990;190:39–44. doi: 10.1111/j.1432-1033.1990.tb15542.x. [DOI] [PubMed] [Google Scholar]
- 35.Jørgensen M U, Bruun M B, Didion T, Kielland-Brandt M C. Mutations in five loci affecting GAP1-independent uptake of neutral amino acids in yeast. Yeast. 1998;14:103–114. doi: 10.1002/(SICI)1097-0061(19980130)14:2<103::AID-YEA203>3.0.CO;2-C. [DOI] [PubMed] [Google Scholar]
- 36.Kitamoto K, Yoshizawa K, Ohsumi Y, Anraku Y. Dynamic aspects of vacuolar and cytosolic amino acid pools of Saccharomyces cerevisiae. J Bacteriol. 1988;170:2683–2686. doi: 10.1128/jb.170.6.2683-2686.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kovari L, Sumrada R, Kovari I, Cooper T G. Multiple positive and negative cis-acting elements mediate induced arginase (CAR1) gene expression in Saccharomyces cerevisiae. Mol Cell Biol. 1990;10:5087–5097. doi: 10.1128/mcb.10.10.5087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kuehn M J, Schekman R, Ljungdahl P O. Amino acid permeases require COPII components and the ER resident membrane protein Shr3p for packaging into transport vesicles in vitro. J Cell Biol. 1996;135:585–595. doi: 10.1083/jcb.135.3.585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kunkel T A, Roberts J D, Zakour R A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- 40.Kyte J, Doolittle R F. A simple method of displaying the hydropathic character of a protein. J Mol Biol. 1982;157:105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- 41.Liang H, Gaber R F. A novel signal transduction pathway in Saccharomyces cerevisiae defined by Snf3-regulated expression of HXT6. Mol Biol Cell. 1996;7:1953–1966. doi: 10.1091/mbc.7.12.1953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Liu H, Styles C A, Fink G R. Saccharomyces cerevisiae S288C has a mutation in FLO8, a gene required for filamentous growth. Genetics. 1996;144:967–978. doi: 10.1093/genetics/144.3.967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ljungdahl P O, Gimeno C J, Styles C A, Fink G R. SHR3: a novel component of the secretory pathway specifically required for the localization of amino acid permeases in yeast. Cell. 1992;71:463–478. doi: 10.1016/0092-8674(92)90515-e. [DOI] [PubMed] [Google Scholar]
- 44.Lorenz M C, Heitman J. The MEP2 ammonium permease regulates pseudohyphal differentiation in Saccharomyces cerevisiae. EMBO J. 1998;17:1236–1247. doi: 10.1093/emboj/17.5.1236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Magasanik B. Regulation of nitrogen utilization. In: Broach J R, Jones E W, Pringle J R, editors. The molecular and cellular biology of the yeast Saccharomyces: gene expression. Vol. 2. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1992. pp. 283–317. [Google Scholar]
- 46.Maniatis T, Fritsch E F, Sambrook J. Molecular cloning: a laboratory manual. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1982. [Google Scholar]
- 47.Markwell M K, Haas S M, Bieber L L, Tolbert N E. A modification of the Lowry procedure to simplify protein determination in membrane and lipoprotein samples. Anal Biochem. 1978;87:206–210. doi: 10.1016/0003-2697(78)90586-9. [DOI] [PubMed] [Google Scholar]
- 48.Marzluf G A. Genetic regulation of nitrogen metabolism in the fungi. Microbiol Mol Biol Rev. 1997;61:17–32. doi: 10.1128/mmbr.61.1.17-32.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.McCusker J H, Haber J E. Mutations in Saccharomyces cerevisiae which confer resistance to several amino acid analogs. Mol Cell Biol. 1990;10:2941–2949. doi: 10.1128/mcb.10.6.2941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ng R, Abelson J. Isolation and sequence of the gene for actin in Saccharomyces cerevisiae. Proc Natl Acad Sci USA. 1980;77:3912–3916. doi: 10.1073/pnas.77.7.3912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Ohsumi Y, Kitamoto K, Anraku Y. Changes induced in the permeability barrier of the yeast plasma membrane by cupric ion. J Bacteriol. 1988;170:2676–2682. doi: 10.1128/jb.170.6.2676-2682.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51a.Özcan S, Dover J, Rosenwald A G, Woelfl S, Johnston M. Two glucose transporters in Saccharomyces cerevisiae are glucose sensors that generate a signal for induction of gene expression. Proc Natl Acad Sci USA. 1996;93:12428–12432. doi: 10.1073/pnas.93.22.12428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Perry J R, Basral M A, Steiner H-Y, Naider F, Becker J M. Isolation and characterization of a Saccharomyces cerevisiae peptide transport gene. Mol Cell Biol. 1994;14:104–115. doi: 10.1128/mcb.14.1.104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Regenberg B, Holmberg S, Olsen L D, Kielland-Brandt M C. Dip5p mediates high-affinity and high-capacity transport of l-glutamate and l-aspartate in Saccharomyces cerevisiae. Curr Genet. 1998;33:171–177. doi: 10.1007/s002940050324. [DOI] [PubMed] [Google Scholar]
- 54.Rose M D, Novick P, Thomas J H, Botstein D, Fink G R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60:237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- 55.Rowen D W, Esiobu N, Magasanik B. Role of GATA factor Nil2p in nitrogen regulation of gene expression in Saccharomyces cerevisiae. J Bacteriol. 1997;179:3761–3766. doi: 10.1128/jb.179.11.3761-3766.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Sikorski R S, Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989;122:19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Soussi-Boudekou S, Vissers S, Urrestarazu A, Jauniaux J C, André B. Gzf3p, a fourth GATA factor involved in nitrogen-regulated transcription in Saccharomyces cerevisiae. Mol Microbiol. 1997;23:1157–1168. doi: 10.1046/j.1365-2958.1997.3021665.x. [DOI] [PubMed] [Google Scholar]
- 58.Stanbrough M, Magasanik B. Transcriptional and posttranscriptional regulation of the general amino acid permease of Saccharomyces cerevisiae. J Bacteriol. 1995;177:94–102. doi: 10.1128/jb.177.1.94-102.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Stanbrough M, Magasanik B. Two transcription factors, Gln3p and Nil1p, use the same GATAAG sites to activate the expression of GAP1 of Saccharomyces cerevisiae. J Bacteriol. 1996;178:2465–2468. doi: 10.1128/jb.178.8.2465-2468.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Stanbrough M, Rowen D W, Magasanik B. Role of the GATA factors Gln3p and Nil1p of Saccharomyces cerevisiae in the expression of nitrogen-regulated genes. Proc Natl Acad Sci USA. 1995;92:9450–9454. doi: 10.1073/pnas.92.21.9450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sychrova H, Chevallier M R. Cloning and sequencing of the Saccharomyces cerevisiae gene LYP1 coding for a lysine-specific permease. Yeast. 1993;9:771–782. doi: 10.1002/yea.320090711. [DOI] [PubMed] [Google Scholar]
- 62.Tanaka J, Fink G R. The histidine permease gene (HIP1) of Saccharomyces cerevisiae. Gene. 1985;38:205–214. doi: 10.1016/0378-1119(85)90219-7. [DOI] [PubMed] [Google Scholar]
- 63.Vandenbol M, Jauniaux J C, Grenson M. Nucleotide sequence of the Saccharomyces cerevisiae PUT4 proline-permease-encoding gene: similarities between CAN1, HIP1 and PUT4 permeases. Gene. 1989;83:153–159. doi: 10.1016/0378-1119(89)90413-7. [DOI] [PubMed] [Google Scholar]
- 64.Vieira J, Messing J. Production of single-stranded plasmid DNA. Methods Enzymol. 1987;153:3–11. doi: 10.1016/0076-6879(87)53044-0. [DOI] [PubMed] [Google Scholar]
- 65.Wiame J M, Grenson M, Arst H N., Jr Nitrogen catabolite repression in yeasts and filamentous fungi. Adv Microb Physiol. 1985;26:1–88. doi: 10.1016/s0065-2911(08)60394-x. [DOI] [PubMed] [Google Scholar]
- 66.Wilson I A, Niman H L, Houghton R A, Cherenson A R, Connolly M L, Lerner R A. The structure of an antigenic determinant in a protein. Cell. 1984;37:767–778. doi: 10.1016/0092-8674(84)90412-4. [DOI] [PubMed] [Google Scholar]
- 67.Zhu X, Garrett J, Schreve J, Michaeli T. GNP1, the high-affinity glutamine permease of S. cerevisiae. Curr Genet. 1996;30:107–114. doi: 10.1007/s002940050108. [DOI] [PubMed] [Google Scholar]