Abstract
As a major metabolite of pyrethroids pesticide, 3-phenoxybenzoic acid (3-PBA) can be an indicator of health risk and human exposure assessment. Based on nanobodies (Nbs), we have developed a rapid flow-through dot enzyme linked immunosorbent assay (dot ELISA) and gold nanoparticles (GNPs) lateral-flow immunoassay for detecting 3-PBA. The limit of detection (LOD) values for detecting 3-PBA by flow-through dot ELISA and GNPs lateral-flow immunoassay were 0.01 ng mL−1 and 0.1 ng mL−1, respectively. The samples (urine and lake water) with and without 3-PBA were detected by both nanobody-based flow-through dot ELISA and GNPs lateral-flow immunoassay, as well as liquid chromatography-mass spectrometry (LC-MS) for validation. The results between immunoassays showed good consistency. It demonstrated that the two developed nanobody-based immunoassays are suitable for rapid detection of 3-PBA.
Keywords: 3-phenoxybenzoic acid, nanobody, gold nanoparticles, flow-through, lateral-flow, immunoassay
Graphical Abstract
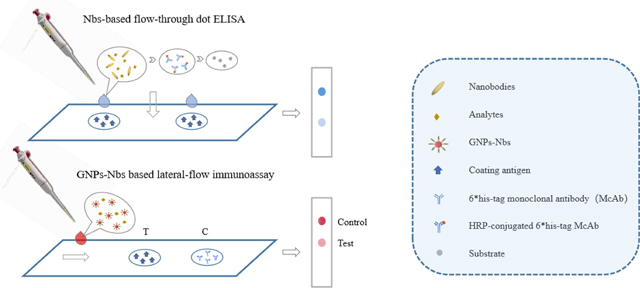
1. Introduction
3-PBA is a common metabolite of a class of pyrethroid pesticides, such as fenpropathrin, cyhalothrin, flucythrinate, permethrin, which can be used as a criterion of exposure assessment in the environment.1–3 Compared with the pyrethroid compounds, it is regarded relatively non-toxic. However, people may unconsciously ingest food contaminated by pyrethroids.4,5 Besides, people may also get exposure to 3-PBA through environmental media such as water. Research showed that 3-PBA mainly exists in human urine and blood, and the effect on estrogen of 3-PBA can cause the organism endocrine metabolism disorder.6 Therefore, people pay more attention to monitoring and assessing of 3-PBA.
Some methods have been established to detect 3-PBA, including instrumental analysis methods (e.g. high-performance liquid chromatography,7,8 supercritical fluid chromatography,9 gas chromatography-mass spectrometry10–13) and immunoassays (e,g. enzyme immunoassay,14–17 electrochemical immunoassay,18,19 fluorescence immunoassay20,21). Instrumental methods are of high sensitivity and easy automation. However, the cost is high and the sample’s clean-up is complex and time-consuming. Traditional ELISA, especially membrane-based immunoassay, is suitable for high throughput screening because of its sensitivity and visual evaluation.22 For the establishment of sensitive membrane-based immunoassay, the selection of antibodies plays a key role.
In the past, most of the antibodies (Abs) used for detecting 3-PBA were monoclonal antibodies (mAbs) and polyclonal antibodies (pAbs).23–25 Liu et al established membrane-based immunoassay which used colloidal gold labeled mAbs for detecting 3-PBA in river water and the LOD value was 1 μg mL-1.26 As the genetic engineering techniques developed, various small size Abs have been found. A new subclass of Abs in members of the camelid family was discovered and called as heavy-chain Abs.27 Recombinant expression of the heavy chain variable domains yield are known as nanobodies (Nbs).28 Compared with traditional Abs (pAbs and mAbs), Nbs have many advantages, including thermostability, accessibility and strong specificity.29,30 With the extensive development of Nbs, Nbs were gradually applied in the field of detection, such as pathogen diagnosis and pollutants detection.31,32 Kim14 et al first isolated VHH establishing VELISA for detecting 3-PBA, and half-maximal inhibitory concentration (IC50) could reach to 1.4 ng mL−1 after the fifth round of panning. To improve sensitivity of 3-PBA detection, Huo17 et al used nanobody-alkaline phosphatase fusion protein to develop the direct competitive fluorescence enzyme immunoassay (dc-FEIA), which achieved LOD as of 0.011 ng mL-1. Sun et al established a nanobody-based competitive dot ELISA for visual screening of ochratoxin A in cereals, and the cut-off level of this visualization assessment was 5 μg kg−1.33
In this research, nanobodies-based flow-through dot enzyme linked immunosorbent assay (Nbs-based flow-through dot ELISA) and gold nanoparticles labeled nanobodies lateral-flow immunoassay (GNPs-Nbs lateral-flow immunoassay), which used nitrocellulose membrane as supporter, were established for sensitive and rapid detection of 3-PBA. To verify the reliability of the rapid assays, health volunteers’ urine and lake water were selected for analysis and the results were consistent with that of LC-MS method. Due to the advantages of high sensitivity, rapid detection and low-cost, these developed membrane-based immunoassays using Nbs can be applied as effective and convenient screening tools for monitoring 3-PBA residues in biological matrix or environment matrix.
2. Materials and Methods
2.1. Chemicals and Reagents
3-PBA standard was purchased from Aladdin (Shanghai, China). Bovine serum albumin (BSA) was purchased from Sigmal-Aldrich. Chloroauric acid (HAuCl4·4H2O), trisodium citrate, Tween-20 and methanol were obtained from Sinopharm (Shanghai Sinopharm Group Chemical reagent Co., Ltd.). 6*his-tag monoclonal antibody (McAb) (Cat No: 66005–1-1g) and HRP-conjugated 6*his-tag McAb (Cat No: HRP-66005) were purchased from Proteintech Group, Inc. 3,3’,5,5’-tetramethylbenzidine (TMB) was purchased from Huinuo Biotechnology (Shenzhen, China). The coating antigen (3-PBA-BSA) and Escherichia coli TOP10F’ strain used to express anti-3-PBA nanobodies containing plasmid were provided by Hammock Lab.14 After expression and purification, the concentration of obtained anti-3-PBA Nbs was 0.3 mg mL−1. Other chemicals were of analytical grade. Two types of nitrocellulose (NC) membrane were purchased from Merck Millipore Ltd (Cat No: HATF00010 and HF13502S25).
2.2. Nbs-based Flow-through Dot ELISA
The pre-treatment of NC membrane for flow-through dot ELISA was prepared with slight modification as the reference described.33 Briefly, slight marks were made at center areas to located the reaction zone on the membrane and the membrane was immersed into PBS buffer for activation. Five μL coating antigen of 3-PBA-BSA (75 μg mL−1) was dropped onto reaction zone. After the liquid flowed through, the membrane was blocked by immersing in PBS solution containing 3% non-powered milk (m/v) for 1 h. Finally, the membranes were washed, dried at room temperature, and stored at 4 °C until use.
Nbs-based flow-through dot ELISA for 3-PBA detection was performed as following procedures: firstly, different concentrations of 3-PBA solution and anti-3-PBA Nbs were pre-mixed, then dropped onto the reaction zone. After the liquid flowed through the membrane completely, the membrane was washed with PBST. Then 5 μL of 500- fold-diluted secondary antibody (HRP-conjugated 6*his-tag McAb) was added. Finally, the NC membrane was immersed in TMB substrate solution for coloration. After 10 min, color of each dot was visually judged by comparing with negative control (without 3-PBA).
2.3. Gold Nanoparticles Labeled Nbs (GNPs-Nbs)
Gold nanoparticles (GNPs) with diameter about 20 nm were prepared according to the procedure described by Frens.34 Briefly, l00 mL of 0.01% HAuCl4 solution (in Milli-Q purified water) was boiling thoroughly. Then 1% trisodium citrate solution was added under constant stirring. The generated GNPs colloidal was cooled to room temperature and stored at 4 °C until use.
For protein labeling of GNPs, protein has the best adsorption capacity when the pH value is closed to its isoelectric point (pH ≥ pI).35 The pH of GNPs was adjusted by adding different amounts of K2CO3. For 1 mL GNPs solution, 0.2 mol L−1 K2CO3 of different volume (5, 10, 15, 20, 25, 30, 50 and 100 μL) were separately added to adjust pH value. The excess Nbs were added into the GNPs solution containing different amounts of K2CO3 and incubated for 1 h. Then each tube was added 50 μL of 10% NaCl and kept for 30 min before scanning under 400–600 nm. The amount of K2CO3 corresponding to the value at λmax was chosen for regulation of GNPs solution before coupling with Nbs. The procedure performed three times repeatedly. Once the optimal pH condition was set, different amounts of 0.3 mg mL−1 Nbs (5, 10, 15, 20, 25 and 30 μL) were also optimized according to the same procedure. GNPs-Nbs were centrifuged for 30 min at 10000 rpm, the sediment was dissolved with storage buffer (containing 1% BSA and 0.25% Tween-20).
2.4. GNPs-Nbs Lateral-flow Immunoassay
The NC membrane for GNPs-Nbs lateral-flow immunoassay was prepared according to the reference by Zhang.36 One microliter of 50 μg mL−1 3-PBA-BSA was coated as test zone and 1 μL of 6*his-tag McAb (20-fold diluted) was coated as control zone. The mixed solution containing GNPs-Nbs (10-fold diluted) and different concentrations of 3-PBA was dropped onto the beginning site of the membrane until the liquid migrated across test zone and control zone completely.
2.5. Cross-reactivity
To assess the specificity of the competitive immunoassay, cross-reactivity (CR) of anti-3-PBA Nbs with structural analogues (3-phenoxybenzaldehyde, 3-phenoxybenzyl alcohol and 4-(3-hydroxybenoxy) benzoic acid) were also determined by Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay.
2.6. Matrix Effect and Sample Analysis
To evaluate matrix effect, a series of concentrations of 3-PBA were prepared in 10% methanol/PBS (as standard) and negative sample matrix. They were measured using plate ELISA, flow-through dot ELISA and lateral-flow immunoassay. To facilitate the quantitative analysis of flow-through dot ELISA and lateral-flow immunoassay, we also used Adobe Photoshop CC software to analyze the images of standard solution, and calculated the inhibition ratio by comparing the grayscale difference of the dot’s color to get the standard curve.
We took urine samples from healthy volunteers and lake water from Jiangsu University with no exposure to pyrethroid insecticides for spiking analysis. Urine collection was performed following the guidelines and protocols of the Jiangsu University. They were proved to be non-3-PBA by LC-MS analysis. The urine samples were centrifuged at 10000 rpm for 10 min, and the supernatant was filtered by 0.22 μm filter membrane before immunoassays. The lake water was directly analyzed after filtration. Sample treatment for LC-MS analysis was the same as described by Huo.17
Negative samples confirmed to be free of 3-PBA by LC-MS were spiked with 3-PBA at concentrations of 0.1, 1, 10, 50 ng mL−1 for recovery analysis. The recovery analysis was done repeatedly four times. We also used software to quantitatively analyze the images of the visual results, and validated by LC-MS to get the correlation. Negative samples were also randomly spiked with 3-PBA and simultaneously analyzed using Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassays and LC-MS.
3. Results and Discussion
3.1. Nbs-based Flow-through Dot ELISA
To get better visual judgement, we first performed the optimization of experimental parameters. The concentration of Nbs and HRP-conjugated 6*his-tag McAb were optimized by checkerboard titration (Figure 1a). The color of dot faded down as the concentration of Nbs decreased from 25 μg mL−1 to 3.125 μg mL−1. With the dilution of HRP-conjugated 6*his-tag McAb increased, the dot color was getting weak. For flow-through dot ELISA, Nbs at the concentration of 25 μg mL−1 and 500-fold diluted HRP-conjugated 6*his-tag McAb were applied for the following analysis. A series of concentrations of 3-PBA were added for competitive immunoassay. In the method, the reaction is competitive combination of coating antigen and free 3-PBA with Nbs. When the concentration of free 3-PBA was too high, Nbs could combine with it completely in the mixture. There was no excess Nbs to combine with the coating antigen on the membrane, so the test zone was colorless. The intensity of dot’s color was inversely proportional to the increased concentration of 3-PBA. When the concentration of 3-PBA reached to 0.01 ng mL−1 (Figure 1b), the dot color could be still distinguished from negative control (without 3-PBA), which was defined as cutoff value for Nbs-based flow-through dot ELISA.
Figure 1.

Nbs-based flow-through dot ELISA. (a) Optimization of Nbs and HRP-conjugated 6*his-tag McAb; (b) a series of concentrations of 3-PBA detected by Nbs-based flow-through dot ELISA.
3.2. Optimization of Gold Nanoparticles Labeled Nbs
In the process of labeling, the pH value played a key role. The pH value of GNPs solution was adjusted by adding different amounts of K2CO3. Figure 2a shows the pH effect on labeling, along with the volume of K2CO3 increased from 0 to 100 μL, the absorbance of λmax reaches maximum value with the amount of 20 μL K2CO3, and the conjugation between GNPs and Nbs reached the best stable state. With gradually increased amount of Nbs (Figure 2b) at the optimum pH, the maximum absorption peak and its corresponding absorbance almost kept constant and reached the optimal conjugation state when the amount of Nbs was 15 μL (at the concentration of 0.3 mg mL−1). Based on these parameters, for 1 mL GNPs solution, the optimum labeling conditions were 20 μL K2CO3 (0.2 mol L−1) for pH adjustment and 15 μL Nbs (0.3 mg mL−1) for conjugation. The GNPs and GNPs labeled Nbs were characterized by wavelength scanning from 400 to 600 nm. After conjugation, λmax of GNPs had a redshift from 520 nm to 525 nm, which could preliminarily indicate successful conjugation (Figure 3a). The characterizations of GNPs and GNPs-Nbs analyzed by the transmission electron microscope (TEM) and particle size analyzer are shown in Figure S1. From the images of TEM, both GNPs and GNPs-Nbs are well-dispersed without aggregation. The particle size of GNPs-Nbs reached to 30 nm, while the size of bare GNPs was about 20 nm (Figure 3b). To further verify the successful conjugation, both GNPs and GNPs-Nbs were selected for lateral-flow immunoassay (Figure 3c). GNPs did not form red spots on the NC membrane (Figure 3c–A), and GNPs-Nbs showed red spots which indicated specific binding between coating antigen and GNPs-Nbs (Figure 3c–B) for successful conjugation.
Figure 2.
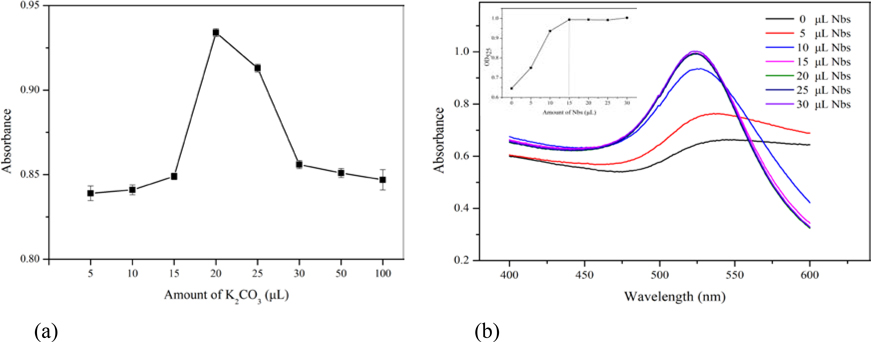
Optimization of GNPs labeling Nbs. (a) amount of K2CO3 (n=3); (b) amount of Nbs.
Figure 3.
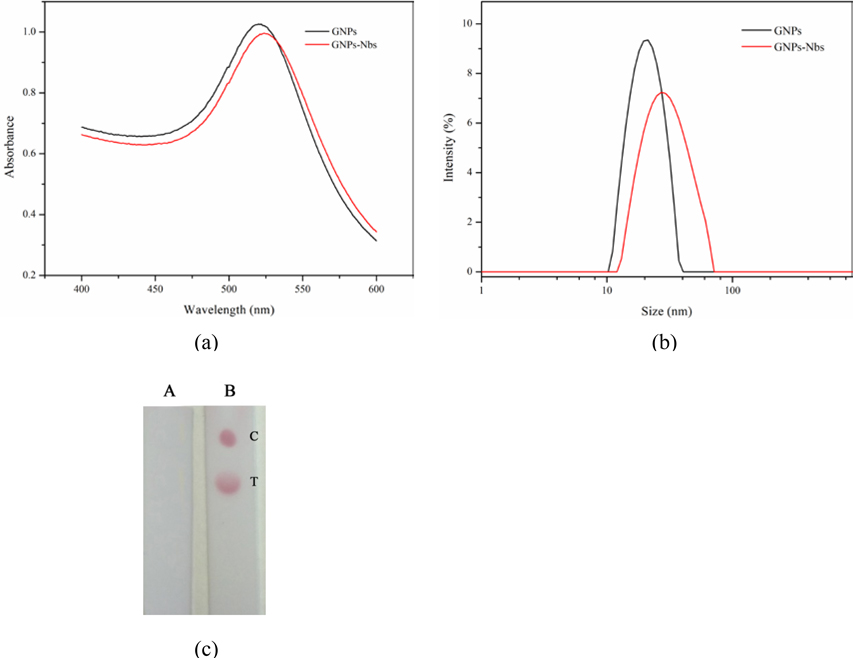
Verification of GNPs-Nbs. (a) scanning spectra of GNPs and GNPs-Nbs; (b) particle size analysis of GNPs and GNPs-Nbs; (c) verification of GNPs-Nbs by lateral-flow immunoassay: GNPs (A); GNPs-Nbs (B).
3.3. GNPs-Nbs Based Lateral-flow Immunoassay
Figure 4 shows the competitive GNPs-Nbs lateral-flow immunoassay for 3-PBA detection. The Nbs firstly mixed with 3-PBA, and along with the increasing concentration of 3-PBA, the color of test zone gradually faded down. When the concentration of 3-PBA was more than 0.1 ng mL−1, the color of test zone could be easily distinguished from the color of test zone without 3-PBA. Thus, the LOD of GNPs-Nbs lateral-flow immunoassay was determined to be 0.1 ng mL−1.
Figure 4.

A series of concentrations of 3-PBA detected by GNPs-Nbs based lateral-flow immunoassay.
3.4. Cross-reactivity Analyzed by Nbs-based Flow-through Dot ELISA and GNPs-Nbs Lateral-flow Immunoassay
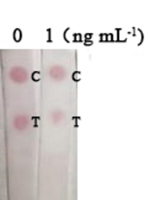
Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay were also applied for the detection of other structural analogues (3-phenoxybenzaldehyde, 3-phenoxybenzyl alcohol and 4-(3-hydroxybenoxy) benzoic acid). The specificity and validation of the developed assays were studied. As shown in Table 1, the concentrations of 0.1 ng mL−1 and 1 ng mL−1 were selected for cross-reactivity analysis. The analogues of 3-phenoxybenzaldehyde and 4-(3-hydroxybenoxy) benzoic acid had the cross-reactivity of 75.1% and 13.5% by plate ELISA, and 3-phenoxybeneyl alcohol showed no cross-reactivity with 3-PBA (<0.1%). The results also indicated that the higher cross-reactivity leaded to the more similar detection images by Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay.
Table 1.
Cross-reactivity detected by Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassay and plate ELISA.
| Analytes | Chemical Structures | Cross-reactivity |
||
|---|---|---|---|---|
| Flow-throughdot ELISA | Lateral-flow Immunoassay | Plate ELISA | ||
| 3-phenoxybenzoic acid |

|

|

|
100% |
| 3-phenoxybenzaldehyde |

|

|

|
75.1% |
| 4-(3-hydroxybenoxy) benzoic acid |

|

|

|
13.5% |
| 3-phenoxybenzyl alcohol |

|

|

|
<0.1% |
3.5. Sample Analysis
We studied the influence of negative urine and lake water as matrix on Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassay and plate ELISA. A series of concentrations of 3-PBA were separately dissolved in 10% methanol/PBS and negative samples matrix. The detection results of the three immunoassays were compared and showed good consistency (Figure S2). For Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay, the influence of matrix almost can be ignored. The images were captured by smartphone and analyzed by software, the gray scale and standard curve of two methods were shown in Figure S3. The cutoff levels which defined as the concentration corresponding to 10% inhibition ratio, were 0.01 ng mL−1 and 0.1 ng mL−1, respectively. The results were also consistent with that of visual judgement results.
To evaluate the validation of Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay, we spiked 3-PBA in negative urine samples at the concentrations of 0.1, 1, 10, 50 ng mL−1. Figure 5 shows that the dot color faded down when the spiked concentration increased. The recoveries by LC-MS ranged from 97% to 103% (Table 2). The quantitatively analysis of the visual results showed in Table S1. And correlation curves of flow-through dot ELISA (R2=0.982) and GNPs-Nbs lateral-flow immunoassay (R2=0.973) between LC-MS showed good consistent (Figure S4). We also used the data to obtain ROC curves in Supplementary Materials (Figure S5). The results showed that the cut off value of flow-through dot ELISA and lateral-flow immunoassay were 0.011 ng mL−1 and 0.107 ng mL−1, which was consistent with visual results. We also randomly spiked 3-PBA on negative samples and analyzed by Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay (Table 3). The visual results by flow-through dot ELISA (3-PBA>0.1 ng mL−1) showed good consistency with LC-MS. As well, the results by lateral-flow immunoassay (3-PBA>0.1 ng mL−1) showed good consistency with LC-MS. Therefore, the developed Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay demonstrated to be the practicable biological monitoring methods for rapid screening of 3-PBA.
Figure 5.
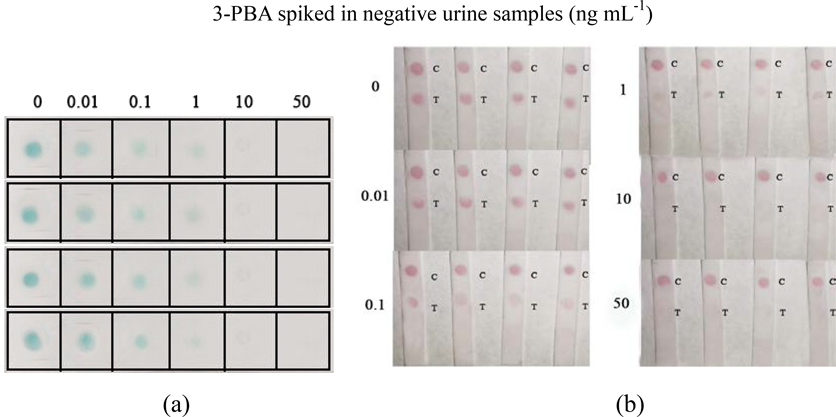
Spiked samples analysis (n=4). (a) Nbs-based flow-through dot ELISA; (b) GNPs-Nbs lateral-flow immunoassay.
Table 2.
Spiked negative samples detected by Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassay and LC-MS (n = 4).
| 3-PBA Spiked (ng mL−1) | Visual results a |
LC-MSb (ng mL−1) | LC-MS Recovery (%) | |
|---|---|---|---|---|
| Flow-through Dot ELISA | Lateral-flow Immunoassay | |||
| 0.1 | +, +, +, + | −/+, +, −/+, + | 0.097 ± 0.070 | 97 |
| 1 | +, +, +, + | +, +, +, + | 0.981 ± 0.045 | 98 |
| 10 | +, +, +, + | +, +, +, + | 10.3 ± 0.8 | 103 |
| 50 | +, +, +, + | +, +, +, + | 50.7 ± 2.6 | 101 |
+: positive, the dot color is weak than control; −: negative, the dot color is bright than control; −/+: negative/positive, the dot color is around control
qualitative detection by Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassay
quantitative analysis by LC-MS
Table 3.
Randomly spiked samples analyzed by Nbs-based flow-through dot ELISA, GNPs-Nbs lateral-flow immunoassay and LC-MS.
| Sample | Visual results (n = 3) |
LC-MS (ng mL−1) | RSD (%) | |
|---|---|---|---|---|
| Flow-through dot ELISA | Lateral-flow Immunoassay | |||
| U1 |

|

|
45.1 ± 1.1 | 2.4 |
| U2 |

|
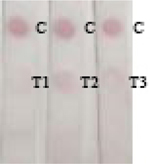
|
1.11 ± 0.07 | 6.3 |
| U3 |

|

|
22.6 ± 0.5 | 2.2 |
| U4 |

|

|
0.67 ± 0.04 | 5.9 |
| L1 |

|
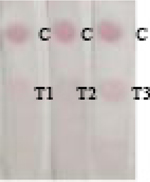
|
4.63 ± 0.26 | 5.6 |
| L2 |

|

|
11.7 ± 0.8 | 6.8 |
| L3 |

|

|
2.47 ± 0.12 | 4.8 |
| L4 |

|
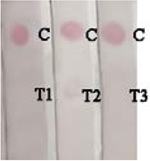
|
50.7 ± 1.6 | 3.2 |
U and L: randomly spiked in urine (U) and lake water (L) samples, respectively
C: control; T1-T3: repeated sample detection
4. Conclusion
We developed two formats (flow-through and lateral-flow) of rapid and convenient Nbs-based immunoassays for 3-PBA detection. The results can be evaluated by the color change of reaction zone which could be directly judged by naked eyes. The LOD value of GNPs-Nbs based lateral-flow immunoassay for 3-PBA detection was 0.1ng mL−1, which was 100-fold sensitive than mAbs-based lateral-flow immunoassay reported by Liu.26 The flow-through dot ELISA was more sensitive, but the process of GNPs-Nbs lateral-flow immunoassay was more convenient because there was no step for substrate participation and the detection time is within 10 min. The spiked samples were tested by Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay, which showed consistence with the results of LC-MS. As Nbs have the advantage for anti-matrix interference, samples can be analyzed without complicated treatments. Moreover, the membrane-based immunoassays (flow-through and lateral-flow) perform the operation more quickly than instrumental methods for high throughput sample analysis. Therefore, Nbs-based flow-through dot ELISA and GNPs-Nbs lateral-flow immunoassay are suitable for visual evaluation and qualitatively on-site sensitive detection of 3-PBA in biological matrix and environmental matrix.
Supplementary Material
HIGHLIGHTS.
Flow-through dot ELISA and gold nanoparticles lateral-flow immunoassay based on nanobodies were developed for detecting 3-PBA.
The sensitivity of nanobody-based flow-through dot ELISA is 10-fold higher than that of gold nanoparticles lateral-flow immunoassay.
The nanobody-based gold nanoparticles lateral-flow immunoassay is 100-fold sensitive than monoclonal antibody-based gold nanoparticles lateral-flow immunoassay.
Acknowledgments
We are very grateful to volunteers for providing samples. We are very grateful to Dr Wan for LC-MS analysis. This work was financially supported by the National Institute of Environmental Health Science Superfund Research Program (P42ES004699) and the National Academy of Sciences and USAID (NAS, Sub-Award No. 2000009144). Any opinions, findings, conclusions, or recommendations expressed in such article are those of the authors alone, and do not necessarily reflect the views of USAID or NAS.
Footnotes
Conflicts of interest
There are no conflicts to declare.
Ethical Approval
All applicable procedures for the use of urines were approved by the Ethical Committee of Jiangsu University and performed following the guidelines and protocols of Jiangsu University.
References
- 1.Shan GM, Huang HZ, Stoutamire DW, Gee SJ, Leng G and Hammock BD, Chem. Res. Toxicol, 2004, 17(2), 218–225. [DOI] [PubMed] [Google Scholar]
- 2.Sudakin DL, Clin. Toxicol, 2006, 44(1), 31–37. [DOI] [PubMed] [Google Scholar]
- 3.Chen L, Zhao TF, Pan CP, Ross JH and Krieger RI, J. Agric. Food Chem, 2012, 60(36), 9342–9351. [DOI] [PubMed] [Google Scholar]
- 4.Richardson JR, Taylor MM, Shalat SL, Guillot TS, Caudle WM, Hossain MM, Mathews TA, Jones SR, Cory-Slechta DA and Miller GW, FASEB J, 2015, 29(5), 1960–1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Erstfeld KM, Chemosphere, 1999, 39(10), 1737–1769. [DOI] [PubMed] [Google Scholar]
- 6.Morgan M, Jones P, Sobus J and Boyd DB, Int. J. Environ. Res. Public Health, 2016, 13(11), 1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ding Y, White CA, Muralidhara S, Bruckner JV and Bartlett MG, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci, 2004, 810(2), 221–227. [DOI] [PubMed] [Google Scholar]
- 8.Abu-Qare AW and Abou-Donia MB, J. Anal. Toxicol, 2001, 25(4), 275–279. [DOI] [PubMed] [Google Scholar]
- 9.El-Saeid MH and Khan HA, Int. J. Food Prop, 2015, 18(5), 1119–1127. [Google Scholar]
- 10.Mudiam MK, Chauhan A, Jain R, Dhuriya YK, Saxena PN and Khanna VK, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci, 2014, 945–946, 23–30. [DOI] [PubMed] [Google Scholar]
- 11.Saito S, Ueyama J, Kondo T, Saito I, Shibata E, Gotoh M, Nomura H, Wakusawa S, Nakai K and Kamijima M, J. Exposure Sci. Environ. Epidemiol, 2014, 24(2), 200–207. [DOI] [PubMed] [Google Scholar]
- 12.Bragança I, Lemos PC, Delerue-Matos C and Domingues VF, Environ. Sci. Pollut. Res, 2019, 26(3), 2987–2997. [DOI] [PubMed] [Google Scholar]
- 13.Kavvalakis MP, Tzatzarakis MN, Alegakis AK, Vynias D, Tsakalof AK and Tsatsakis AM, Drug Test. Anal, 2014, 6 (Suppl 1), 9–16. [DOI] [PubMed] [Google Scholar]
- 14.Kim HJ, McCoy MR, Majkova Z, Dechant JE, Gee SJ, Tabares-da Rosa S, Gonzalez-Sapienza GG and Hammock BD, Anal. Chem, 2012, 84(2), 1165–1171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Thiphom S, Prapamontol T, Chantara S, Mangklabruks A, Suphavilai C, Ahn KC, Gee SJ and Hammock BD, J. Environ. Sci. Health, Part B, 2014, 49(1), 15–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ahn KC, Gee SJ, Kim HJ, Aronov PA, Vega H, Krieger RI and Hammock BD, Anal. Bioanal. Chem, 2011, 401(4), 1285–1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huo JQ, Li ZF, Wan DB, Li DY, Qi M, Barnych B, Vasylieva N, Zhang JL and Hammock BD, J. Agric. Food Chem, 2018, 66(43), 11284–11290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.El-Moghazy AY, Huo JQ, Amaly N, Vasylieva N, Hammock BD and Sun G, ACS Appl. Mater. Interfaces, 2020, 12(5), 6159–6168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pali M, Bever CRS, Vasylieva N, Hammock BD and Suni II, Electroanalysis, 2018, 30(11), 2653–2659. [Google Scholar]
- 20.Wang YL, Li ZF, Barnych B, Huo JQ, Wan D, Vasylieva N, Xu JL, Li P, Liu BB, Zhang CZ and Hammock BD, J. Agric. Food Chem, 2019, 67(41), 11536–11541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nichkova M, Dosev D, Davies AE, Gee SJ, Kennedy IM and Hammock BD, Anal. Lett, 2007, 40(7), 1423–1433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhao CY, Si Y, Pan BF, Taha AY, Pan TR and Sun G, Talanta, 2020, 217, 121054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kim HJ, Ahn KC, González-Techera A, González-Sapienza GG, Gee SJ and Hammock BD, Anal. Biochem, 2009, 386(1), 45–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kim HJ, Gee SJ, Li QX and Hammock BD, ACS Symp. Ser, 2007, 966, 171–185. [Google Scholar]
- 25.Lu Y, Xu N, Zhang Y, Liu B, Song Y and Wang S, Food Agric. Immunol, 2010, 21(1), 27–45. [Google Scholar]
- 26.Liu Y, Wu AH, Hu J, Lin MM, Wen MT, Zhang X, Xu CX, Hu XD, Zhong JF, Jiao LX, Xie YJ, Zhang CZ, Yu XY, Liang Y and Liu XJ, Anal. Biochem, 2015, 483, 7–11. [DOI] [PubMed] [Google Scholar]
- 27.Hamers-Casterman C, Atarhouch T, Muyldermans S, Robinson G, Hamers C, Songa EB, Bendahman N and Hamers R, Nature, 1993, 363, 446–448. [DOI] [PubMed] [Google Scholar]
- 28.Maass DR, Sepulveda J, Pernthaner A and Shoemaker CB, J. Immunol. Methods, 2008, 324(1–2), 13–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liang L, Hu ZX, Huang YY, Duan SL, He J, Huang Y, Zhao YX and Lu XL, J. Nanosci. Nanotechnol, 2016, 16(12), 12099–12111. [Google Scholar]
- 30.Meyer TD, Muyldermans S and Depicker A, Trends in Biotechnology, 2014, 32(5), 263–270. [DOI] [PubMed] [Google Scholar]
- 31.Sroga P, Safronetz D and Stein DR, Future Virol, 2020, 15(3), 195–205. [Google Scholar]
- 32.Chen Q, Wang YY, Mao FJ, Su BC, Bao KL, Zhang ZL, Xie GF and Liu X, RSC Adv, 2020, 10(56), 33700–33705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sun ZC, Duan ZH, Liu X, Deng X and Tang ZW, Food Anal. Methods, 2017, 10, 3558–3564. [Google Scholar]
- 34.Frens G, Nature (London), Phys. Sci, 1973, 241(105), 20–22. [Google Scholar]
- 35.Thobhani S, Attree S, Boyd R, Kumarswami N, Noble J, Szymanski M and Porter RA, J. Immunol. Methods, 2010, 356(1–2), 60–69. [DOI] [PubMed] [Google Scholar]
- 36.Zhang C, Zhang Y and Wang S, J. Agric. Food Chem, 2006, 54(7), 2502–2507. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


