Abstract
Bats are infected with several trypanosomatid species; however, assessing the diversity of this interaction remains challenging since there are species apparently unable to grow in conventional culture media. Accordingly, the ecology and biology of the Molecular Operational Taxonomic Units (MOTUs) Trypanosoma spp. Neobats are unknown. Therefore, we performed the molecular characterization targeting the 18S small subunit rDNA from the blood clot of 280 bats of three Brazilian regions (Paraíba, Rio de Janeiro and Acre states), bypassing the selective pressure of hemoculture. From 68 (24%) positive blood clot samples, we obtained 49 satisfactory sequences. Of these successfully sequenced results, T. spp. Neobats (1, 3 and 4) represented 67%, with the most abundant T. sp. Neobat 4 (53%). Our results show: (1) high abundance and wide geographic range of T. sp. Neobat 4, restricted to Carollia bats; (2) high infection rate of T. sp. Neobat 4 in Carollia perspicillata populations (mean 26%); (3) infection with the monoxenous Crithidia mellificae; and (4) a new MOTU (T. sp. Neobat 5) in Artibeus cinereus, positioning in the Trypanosoma wauwau clade. These data corroborate the importance of bats as hosts of many Trypanosoma species and C. mellificae. They also show that the diversity of the T. wauwau clade is underestimated and warn about the high magnitude of trypanosomes we overpass with the hemoculture. Our findings combined with previous data show that T. spp. Neobats include host-specific and host-generalist species, probably playing different ecological roles: T. sp. Neobat 1 shows broad host range; T. spp. Neobat 3 and 4 are restricted to Artibeus and Carollia, respectively. Finally, T. Neobat 4 seems to be a well-succeeded parasite, especially within C. perspicillata metapopulations across a wide geographical distribution. This work is a step forward to understand the biology and life history of T. spp. Neobats.
Keywords: Trypanosoma, Neobat, MOTU, Crithidia, Bats, Blood clot, Specificity
Graphical abstract
Highlights
-
•
Trypanossoma spp. Neobats 1–4 show distinct host ranges.
-
•
High abundance and wide geographic range of T. sp. Neobat 4, restricted to Carollia bats.
-
•
Discovery of a new MOTU within T. wauwau clade, detected in Artibeus cinereus.
-
•
Infection with Crithidia mellificae in bats from different localities.
-
•
Identification of mixed infections.
1. Introduction
Protozoan parasites of the trypanosomatid family are structured into monoxenous and dixenous genera. Although the former is considered restricted to invertebrate hosts (Kaufer et al., 2017), there are increasing reports of monogenetic trypanosomatids infecting mammals (Jansen et al., 1988; Kraeva et al., 2015; Hodo et al., 2016). This is the case of the honey bee-associated Crithidia mellificae, isolated from the blood of different bat species from the Rio de Janeiro Atlantic Forest, besides other mammal groups (Rangel et al., 2019; Dario et al., 2021). The Trypanosoma genus comprises parasites of vertebrates, mostly transmitted by a wide range of hematophagous arthropods (Kaufer et al., 2017). Among the trypanosomes infecting mammals, the species of the Trypanosoma cruzi clade (comprising species of the subgenus Schizotrypanum; e.g., T. cruzi cruzi, T. cruzi marinkellei, T. dionisii) are distributed worldwide associated with their putative ancestral bat hosts (Hamilton et al., 2012; Lima et al., 2015). Additionally, T. wauwau, T. madeirae, and the Molecular Operational Taxonomic Units (MOTUs) T. spp. Neobats were detected in neotropical bats (Cottontail et al., 2014; Lima et al., 2015; Barros et al., 2018; Rodrigues et al., 2019). These trypanosomes, together with T. janseni (Lopes et al., 2018) and the MOTU Trypanosoma sp. DID (Rodrigues et al., 2019), constitute the T. wauwau clade (Kostygov et al., 2021).
One of the main diagnostic methods of Trypanosoma infection includes the detection and isolation of the flagellates by hemoculture (Jansen et al., 2018). However, there are species apparently unable to grow in conventional culture media leading to nonrepresentative data and underestimation of the Trypanosoma diversity. In this context, the molecular detection of Trypanosoma DNA in mammalian blood clots showed to be a valuable and accessible approach to overcome the detection of non-cultivable species (Rodrigues et al., 2019).
Indeed, the advance of molecular tools and phylogenetic analysis improved the access of the remarkable genetic diversity within the Trypanosoma genus. High-sensitivity and effective techniques for species discrimination enabled the detection and characterization of scarce DNA fragments, including those from non-cultivable trypanosomes (Dario et al., 2017a). Regardless of the relevance and accuracy of molecular diagnosis, the description of a novel parasite species requires specimen isolation (Šlapeta, 2013). Thus, clusters of increasingly identified gene sequences of undescribed trypanosomes (named Molecular Operational Taxonomic Units – MOTUs) may be used as proxies for “species” (Blaxter et al., 2005).
The MOTUs designated as Trypanosoma spp. Neobat comprise a complex of unnamed trypanosomes divided in T. sp. Neobat 1–4 (Cottontail et al., 2014; Lima et al., 2015; Rodrigues et al., 2019). Each MOTU possibly represents a valid species requiring a formal taxonomic description. However, those unseen flagellates are apparently incapable of growing in conventional culture medium for Trypanosoma spp.
To date, T. spp. Neobat gene sequences were detected in phyllostomid bats from Panama, Colombia, and Brazil. T. sp. Neobat 1 was found in Artibeus jamaicensis and Trachops cirrhosis; T. spp. Neobat 2 and 3 in Artibeus jamaicensis and A. lituratus. T. sp. Neobat 2 was also detected in Artibeus planirostris (Cottontail et al., 2014; Lima et al., 2015; Rodrigues et al., 2019). T. sp. Neobat 4 was described in one Carollia perspicillata at the Rio de Janeiro Atlantic Forest (Rodrigues et al., 2019).
While the study of the genetic diversity and phylogeny of the MOTUs T. spp. Neobats advances, their biological and ecological properties remain mysterious. Knowledge gaps such as their vectors, transmission cycles, as well as the interaction with their hosts and life strategies prevent us to understand their ecological role (i.e., the functions played by an organism on the ecosystem) and therefore the effects of these organisms on the ecosystem (Marcot and Vander Hayden, 2001). For instance, what are the impacts of the Neobat parasites on their host populations? What are the outcomes of mixed infection with the Neobat trypanosomes and other parasites?
Artibeus and Carollia include species that are among the most abundant and widespread in the Neotropics (Kalko and Handley, 2001; Gardner, 2008). The latter includes the sister species Carollia perspicillata and C. brevicauda, which occur in sympatry along most of their geographic distributions (Hoffmann and Baker, 2003). Both genera comprise predominantly frugivorous species (Giannini and Kalko, 2004) that may complement their diet with insects (Ayala and D'Alessandro, 1973; Willig et al., 1993; York and Billings, 2009; Guedes et al., 2020). Species in these genera use a wide variety of habitats and occupy different roost sites such as foliage of trees, human buildings, and caves (Fenton et al., 2001; Vilar et al., 2016; Carvalho et al., 2018).
Trypanosome species infecting bats remain an unsolved question. Herein, we expanded the knowledge on the diversity and life history of trypanosomatids of neotropical bats through molecular characterization directly from the blood clot, enabling the detection of non-cultivable species. We found high trypanosomatid diversity—including C. mellificae and a new MOTU, named T. sp. Neobat 5, positioning in the T. wauwau clade—, distinct host ranges of the T. spp. Neobats, and that T. sp. Neobat 4 is a well-established parasite apparently restricted to Carollia.
2. Materials and methods
2.1. Bat assemblage
We searched for trypanosomatid infection in 280 neotropical bats belonging to at least 29 species of 18 genera (Table 1), captured at three Brazilian Conservation Unities from 2014 to 2016: Guaribas Biological Reserve, Mamanguape municipality, Paraíba; Seringal Cachoeira (SC), Xapuri municipality, Acre; and Guapiaçu Ecological Reserve (REGUA), Cachoeiras de Macacu municipality, Rio de Janeiro (Fig. 1).
Table 1.
Number of sampled bat species per locality.
| Locality | Species (n) |
|---|---|
| Guaribas Biological Reserve, Mamanguape, Paraíba (N = 114; S = 10) | Artibeus cinereus (26), A. lituratus (18), A. planirostris (26), Carollia perspicillata (28), Desmodus rotundus (1), Glossophaga soricina (7), Myotis lavali (2), Phyllostomus discolor (1), Platyrrhinus lineatus (4), Sturnira lilium (1) |
| Seringal Cachoeira (SC), Xapuri, Acre (N = 50; S = 20) | Anoura caudifer (1), Artibeus cinereus (2), A. lituratus (2), A. obscurus (2), A. planirostris (3), Carollia brevicauda (2), C. benkeithi (1), C. perspicillata (17), Chiroderma villosum (1), Glossophaga soricina (1), Lasiurus blossevillii (1), Phyllostomus discolor (4), P. elongatus (1), P. hastatus (3), Rhinophylla fischerae (4), Sturnira tildae (1), Tonatia saurophila (1), Trachops cirrhosus (1), Uroderma bilobatum (1), Vampyressa thyone (1) |
| Guapiaçu Ecological Reserve (REGUA), Cachoeiras de Macacu, Rio de Janeiro (N = 116; S = 14) | Anoura caudifer (1), A. geoffroyi (2), Artibeus fimbriatus (2), A. lituratus (21), A. obscurus (2), Carollia perspicillata (50), Chrotopterus auritus (1), Desmodus rotundus (8), Lonchorhina aurita (1), Myotis sp. (1), Phyllostomus hastatus (4), Platyrrhinus recifinus (1), Sturnira lilium (20), S. tildae (2) |
N = total abundance; S = species richness; n = abundance.
Fig. 1.
Geographical location of the study areas. Bats were collected from 2014 to 2016 at three Brazilian Conservation Unities: Guaribas Biological Reserve, Mamanguape municipality, Paraíba; Seringal Cachoeira, Xapuri municipality, Acre; and Guapiaçu Ecological Reserve, Cachoeiras de Macacu municipality, Rio de Janeiro. (A) Political map of Brazil: areas filled in black indicate the study states; (B) Paraíba state; (C) Acre state; (D) Rio de Janeiro state. Red dots indicate the location of the Conservation Unities. Service Layer Credits Source: Esri, HERE, Garmin, © OpenStreetMap contributors, and the GIS user Community. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
For this, we took advantage of blood clot samples stored at the Laboratory of Trypanosomatid Biology previously collected by Pires (2017), dos Santos et al. (2018), and Rangel et al. (2019). These authors investigated the Trypanosoma diversity through the hemoculture technique and capture procedures are described by them (Pires, 2017; dos Santos et al., 2018; Rangel et al., 2019).
Bat samplings were conducted under permits of the Chico Mendes Institute for Biodiversity Conservation (ICMBio): licenses 41683-3 and 41683-5 (Guaribas); 47377-1 (SC); 19037–1 (REGUA). All procedures with the bats followed protocols approved by the Ethics Commission on Animal Use (CEUA): licenses P0007-99, P0179-03, P0292/06, L0015-07, e L81-12 (Guaribas); LW81-12 (SC); L-81/12 and LW-68/12 (REGUA).
2.2. Collection and storage of blood clots
The whole blood samples were obtained by cardiac puncture and centrifuged at 1180×g for 15 min. Blood clot samples were obtained by separation of the supernatant using sterile tips and stored at −20 °C with absolute ethanol (proportion 1:1).
2.3. DNA extraction from blood clots
After discarding the absolute ethanol, a fragment of the blood clot (about 50 μl) was submitted to DNA extraction based on the ammonium acetate precipitation (Rodrigues et al., 2019). DNA was resuspended in 20 μl Tris-EDTA buffer (10 mM Tris-HCl pH 7.4; 1 mM EDTA pH 8.0) and the concentration and purity (OD260/OD280 ratio) were quantified using BioPhotometer® (Eppendorf).
2.4. Molecular diagnosis
Nested-PCR targeting the 18S small subunit of the ribosomal gene (SSU rDNA) was performed using two sets of primers specific for the trypanosomatid family (Smith et al., 2008). DNA extracts (150–200 ng) were used as template for a 25 μl reaction consisted of 8.5 μl GoTaq® Green Master Mix (Promega) and 16 pmol of the following external primers: TRY927F (5′ GAAACAAGAAACACGGGAG 3′) and TRY927R (5′ CTACTGGGCAGCTTGGA 3′). For the second round, the PCR products diluted at 1:10 in ultrapure sterile water were used as template following the same reaction protocol, with the internal primers: SSU561F (5′ TGGGATAACAAAGGAGCA 3′) and SSU561R (5′ CTGAGACTGTAACCTCAAAGC 3’). Ultrapure sterile water and T. cruzi cruzi DNA from positive hemoculture were used as negative and positive controls, respectively. The amplification was performed on Swift™ MiniPro Thermal Cycler (Esco) with the following cycling conditions: initial denaturation at 94 °C/3 min; followed by 30 cycles at 94 °C/30 s, 55 °C/60 s, and 72 °C/90 s; and a final elongation step at 72 °C/10 min. The PCR products were visualized on 1.5–2.0% agarose gels stained with GelRed (Biotium).
Both forward and reverse fragment strands were subjected to fluorescent dye-terminator cycle sequencing reactions with ABI 3730 BigDye Terminator v3.1 Cycle Sequencing Ready Reaction Kit (Applied Biosystems) using internal primers at 16 pmol and run on an ABI 3730 automated sequencer (Applied Biosystems) at the Oswaldo Cruz Foundation Sequencing Platform facility (PDTIS/FIOCRUZ). Chromatograms of both strands were visually inspected for possible double-peaks and correct nucleotide assignment, and assembled using SeqMan Lasergene v.7.0 (DNASTAR Inc.). Consensus sequences were compared against the NCBI database with the BLASTn algorithm (Altschul et al., 1990) for initial screening and then aligned and edited using MEGA-X v.10.1 (Kumar et al., 2018).
2.5. Phylogenetic analysis
The phylogenetic reconstructions were performed using both Bayesian and maximum-likelihood (ML) methods with the BEAST 2.6 platform (Bouckaert et al., 2019) and PhyML v.3.0 program (Guindon et al., 2010), respectively. The best-fitting models of base substitution were chosen with the Bayesian information criterion (BIC) in bModelTest 1.2 (Bouckaert and Drummond, 2017) and the same model was then applied to the dataset under ML analysis. Bayesian phylogenetic trees were inferred with three independent runs (2 × 107 generations) assuming the Birth-and-Death model of speciation, sampling parameters every 10,000 generations. We evaluated parameter convergence and proper mixing through visual inspection of chains and by checking that effective sample sizes (ESSs) were sufficiently large; in our case, all ESSs were ≥ 104. ML inferences were performed with random starting trees, subtree pruning and regrafting topological rearrangements for tree improvement. Statistical support for clades was inferred with posterior probability and 500 bootstraps replications for Bayesian and ML trees, respectively.
Apart from the samples sequenced in this study, we included into the phylogenetic analyses 14 sequences belonging to monogenetic species of Crithidia, Leptomonas and Lotmaria, and 50 sequences of the T. cruzi and T. wauwau clades (besides four sequences of T. livingstonei), all retrieved from the GenBank database. The sequences of T. madeirae were excluded since only 413bp overlapped our final 546bp alignment.
3. Results
We analyzed the blood clot of 280 bats, of which 68 (24%) amplified for the 18S target. Of these, we obtained 49 satisfactory sequencing results. Our results show: (1) high abundance and wide geographic range of T. sp. Neobat 4, restricted to Carollia spp.; (2) high infection rate of T. sp. Neobat 4 in C. perspicillata populations; (3) infection with the monoxenous C. mellificae; and (4) a new MOTU detected in Artibeus cinereus, named T. sp. Neobat 5, positioning in the T. wauwau clade.
The examined bats showed high richness of trypanosomatids with a predominance of T. sp. Neobat 4, restricted to the genus Carollia. In 10 bat species, we detected 8 taxa belonging to Trypanosoma and one to Crithidia (Fig. 2, Fig. 3). T. sp. Neobat 4 was the most abundant (53% of total trypanosomatids; n = 26) and geographically spread taxon, detected in all studied localities. Interestingly, among 18 studied host genera, T. sp. Neobat 4 was restricted to the genus Carollia. This MOTU was detected in 25 C. perspicillata individuals and in one C. brevicauda (Table 2 and S1 Table).
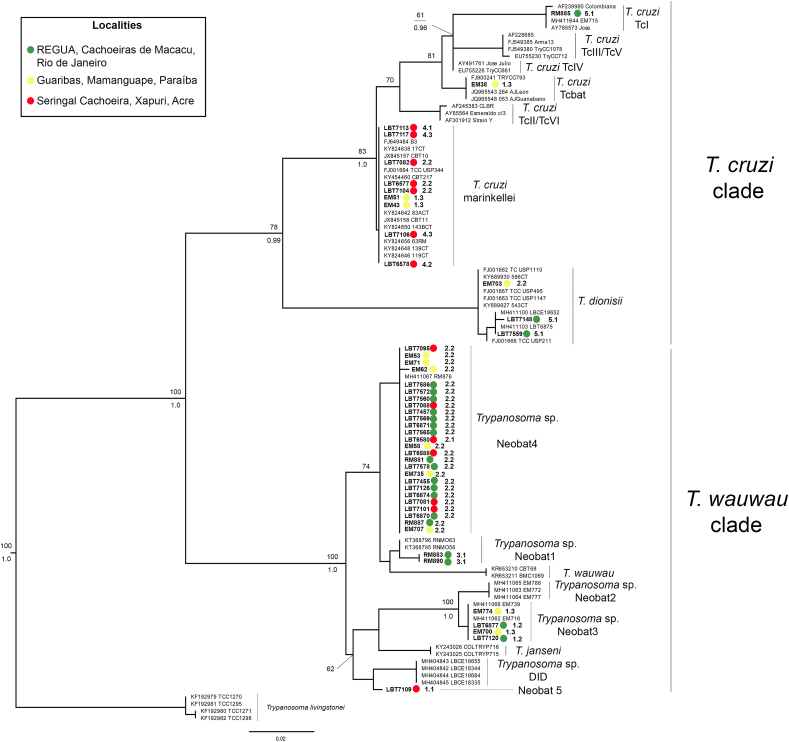
Fig. 2.
Phylogenetic analysis of sequences of the 18S SSU rDNA gene of Trypanosoma detected in the blood clot of bats. ML phylogenetic tree of Trypanosoma sequences using 18S SSU rDNA from blood clot of bats captured at Guaribas Biological Reserve, Seringal Cachoeira, and Guapiaçu Ecological Reserve. Upper and lower numbers on tree nodes are posterior probabilities (>0.95) and bootstrap (>60), respectively. Accession numbers of sequences retrieved from Genbank are shown on tip labels. Localities are represented by colors - yellow circles: Guaribas Biological Reserve, Mamanguape, Paraíba; red circles: Seringal Cachoeira, Xapuri, Acre; and green circles: Guapiaçu Ecological Reserve, Cachoeiras de Macacu, Rio de Janeiro. Sample number codes on the circles right represent parasite hosts. The decimal point divides the host genus (left number) and the host species (right number) codes. Host genera: 1. Artibeus; 2. Carollia; 3. Desmodus; 4. Phyllostomus; 5. Sturnira. Host species: 1.1. Artibeus cinereus; 1.2. Artibeus lituratus; 1.3. Artibeus planirostris; 2.1. Carollia brevicauda; 2.2. Carollia perspicillata; 3.1. Desmodus rotundus; 4.1. Phyllostomus discolor; 4.2. Phyllostomus elongatus; 4.3. Phyllostomus hastatus; 5.1. Sturnira lilium. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
Fig. 3.
Phylogenetic analysis of sequences of the 18S SSU rDNA gene of Crithidia detected in the blood clot of bats. Bayesian phylogenetic tree of Crithidia sequences using 18S SSU rDNA from blood clot of Carollia perspicillata captured at Guaribas Biological Reserve (EM 69) and Sturnira lilium at Guapiaçu Ecological Reserve (RM 872). Upper and lower numbers on tree nodes are posterior probabilities (>0.95) and bootstrap (>60), respectively. Accession numbers of sequences retrieved from Genbank are shown on tip labels. Yellow circle: Guaribas Biological Reserve, Mamanguape, Paraíba; green circle: Guapiaçu Ecological Reserve, Cachoeiras de Macacu, Rio de Janeiro. Sample number codes on the circles right represent parasite hosts. The decimal point divides the host genus (left number) and the host species (right number) codes. Host genera: 2. Carollia; 5. Sturnira. Host species: 2.2. Carollia perspicillata; 5.1. Sturnira lilium. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
Table 2.
Trypanosomatid species detected from 49 blood clots of bats from Guaribas Biological Reserve/Paraíba, Seringal Cachoeira/Acre, Brazil, and Guapiaçu Ecological Reserve/Rio de Janeiro.
| Taxon (Proportion %) | Locality | Species; Infection rate % (n) |
|---|---|---|
| T. sp. Neobat 4 (53%) | Guaribas | Carollia perspicillata 21% (28) |
| SC | C. perspicillata 29% (17), C. brevicauda 50% (2) | |
| REGUA | C. perspicillata 28% (50) | |
| T. c. marinkellei (19%) | Guaribas | Artibeus planirostris 8% (26) |
| SC | C. perspicillata 18% (17), Phyllostomus elongatus 100% (1), P. hastatus 67% (3), P. discolor 25% (4) | |
| T. sp. Neobat 3 (8%) | Guaribas | A. planirostris 8% (26) |
| REGUA | A. lituratus 10% (21) | |
| T. dionisii (6%) | Guaribas | C. perspicillata 4% (28) |
| REGUA | Sturnira lilium 10% (20) | |
| T. sp. Neobat 1 (4%) | REGUA | Desmodus rotundus 22% (9) |
| C. mellificae (4%) | Guaribas | C. perspicillata 4% (28) |
| REGUA | S. lilium 5% (20) | |
| T. c. cruzi DTU TcI (2%) | REGUA | S. lilium 5% (21) |
| T. c. cruzi DTU TcBat (2%) | Guaribas | A. planirostris 3% (29) |
| T. sp. Neobat 5 (2%) | SC | A. cinereus 50% (2) |
DTU = Discrete Typing Unit; SC = Seringal Cachoeira; REGUA = Guapiaçu Ecological Reserve; n = number of evaluated bats.
Trypanosoma cruzi marinkellei was the second most abundant taxon (19% of total trypanosomatids; n = 9) and the most dispersed among bat species (A. planirostris, C. perpicillata, Phyllostomus elongatus, P. hastatus, and P. discolor; Table 2). The monoxenous C. mellificae was detected in C. perspicillata at Guaribas and Sturnira lilium at REGUA (Fig. 3). Finally, our phylogenetic results revealed a new MOTU detected in A. cinereus, named T. sp. Neobat 5, positioning in the T. wauwau clade (GenBank accession number MW618923; Fig. 2).
Carollia perspicillata displayed the highest parasite richness and abundance throughout sampled regions. In 16 infected animals we detected trypanosomatids belonging to three taxa of Trypanosoma and one Crithidia. T. sp. Neobat 4 was the most prevalent taxon in C. perspicillata (84% of trypanosomatids found in this bat species), showing high infection rates in C. perspicillata populations (21% in Guaribas, 29% in SC, 28% in REGUA; mean 26%). Trypanosoma cruzi marinkellei was the second most prevalent taxon (10%), followed by T. dionisii (3%) and C. mellificae (3%) (Table 2 and S1 Table).
The evaluation of trypanosomatid infection by direct molecular detection of the blood clot revealed mixed infections. Nine animals previously considered by the hemoculture single-infected with T. c. cruzi TcI or T. dionisii (Pires, 2017; dos Santos et al., 2018; Rangel et al., 2019) showed mixed infections when tested through direct examination of the blood clot (Table 3).
Table 3.
Mixed Trypanosoma infection detected by direct molecular analysis of the blood clot and hemoculture.
| Region | Voucher | Species | Hemoculture | Blood clot |
|---|---|---|---|---|
| Guaribas (Pires, 2017) | EM 38 | Artibeus planirostris | T. c. cruzi TcI | T. c. cruzi TcBat |
| SC (dos Santos et al., 2018) | LBT 7081 | Carollia perspicillata | T. c. cruzi TcI | T. sp. Neobat 4 |
| LBT 7082 | C. perspicillata | T. c. cruzi TcI | T. c. marinkellei | |
| LBT 7088 | C. perspicillata | T. c. cruzi TcI | T. sp. Neobat 4 | |
| LBT 7095 | C. perspicillata | T. c. cruzi TcI | T. sp. Neobat 4 | |
| LBT 7101 | C. perspicillata | T. c. cruzi TcI | T. sp. Neobat 4 | |
| LBT 7104 | C. perspicillata | T. c. cruzi TcI | T. c. marinkellei | |
| REGUA (Rangel et al., 2019) | LBT 7455 | C. perspicillata | T. dionisii | T. sp. Neobat 4 |
SC = Seringal Cachoeira; REGUA = Guapiaçu Ecological Reserve.
4. Discussion
Bats are infected with several species of trypanosomatids; however, assessing the diversity remains challenging due to limitations of the hemoculture technique. Non-cultivable species are rather detected through the evaluation of the blood clot (Rodrigues et al., 2019). Therefore, we investigated the diversity of trypanosomatids in bats from three Brazilian regions through molecular characterization targeting the 18S SSU rDNA directly from the blood clot, bypassing the selective pressure of hemoculture.
We came across noticeable findings: (1) T. sp. Neobat 4 showed the highest abundance and wide geographical distribution infecting a significant portion of C. perspicillata populations from Guaribas, REGUA and SC; (2) a new MOTU (T. sp. Neobat 5) detected in the A. cinereus, positioning in the T. wauwau clade, and (3) bats infected with the monoxenous C. mellificae.
Parasites impact on individual, population, and ecosystem levels (Frainer et al., 2018). They modulate allergic, autoimmune (Allaerts and Chang, 2017) and evolution of their hosts (Morand, 2015); they promote biodiversity, modify community structure, trophic networks, and energy flow (Thompson et al., 2005; Lafferty, 2012; Hatcher et al., 2012; Dougherty et al., 2016). Trypanosomes must be regarded as pivotal components of ecosystem processes as they constitute a widely dispersed group, infecting all vertebrate groups. This is especially the case of trypanosomes associated with bats—a mammalian group worldwide distributed that includes important components of ecological networks. However, the ecological role of those trypanosomes, especially the Neobat parasites, remains totally unknown.
Functional traits are the characteristics of organisms that impact their fitness thus allowing to unveil their effects on ecosystem processes (Violle et al., 2007). A key functional trait of parasites is the level of host specificity (Llopis-Belenguer et al., 2019). Parasite specificity is the result of ecological and biological attributes of parasite and host, together with emergent properties of their ecological and evolutionary interactions.
In this sense, our results show that the so-called T. spp. Neobats comprise a heterogeneous and complex assemblage of parasites that display distinct life strategies— including host-specific and host-generalist species—, probably playing different ecological roles. Herein, we expanded the knowledge of the host ranges of T. spp. Neobat 1, 3, and 4. The former shows the broadest range, capable of infecting five species of distinct genera (Artibeus jamaicensis, Trachops cirrhosis, Sturnira tildae, Desmodus rotundus, and Carollia perspicillata). T. sp. Neobat 3 is observed solely in Artibeus (A. jamaicensis, A. lituratus, and A. cinereus), likewise T. sp. Neobat 2 (A. jamaicensis, A. lituratus, and A. planirostris) (Cottontail et al., 2014; Lima et al., 2015; Rodrigues et al., 2019). Finally, T. sp. Neobat 4 is restricted to Carollia, detected in the closely related species C. brevicauda and C. perspicillata, with high infection rates in C. perspicillata populations.
The highest abundance and richness of trypanosomatids in C. perspicillata can be partially explained by the sample size since this species represents 34% of the total evaluated bats. However, we cannot rule out that their generalist behavior regarding their diet and habitats makes them highly exposed to various potential vector species (Dario et al., 2017b).
The infection with the monogenetic C. mellificae in bats showed a broader mammalian host and geographical range than previously described (Rangel et al., 2019; Dario et al., 2021). We detected this species in S. lilium (in REGUA) and in a new locality (Guaribas Biological Reserve), indicating that the presence of Crithidia mellificae in a mammal host is not incidental or at a local scale, but a host-parasite interaction that deserves to be better explored. Further, the direct diagnosis of these parasites through the blood clot rejects the hypothesis of a putative contamination regarding Crithidia growth in culture medium.
The molecular diagnosis based on the blood clot showed to be an informative tool to investigate non-cultivable trypanosome species and trypanosomes at low parasitemia levels in bats. Nonetheless, it was ineffective to detect mixed infections, which were only discovered when our analyzes were combined with previous results from the hemoculture technique (Table 3). The possible reasons are (1) PCR bias, (2) non-representativeness of the fragment of the blood clot collected to DNA extraction, and (3) higher amount of representatives of a species in the blood, leading to the amplification of the most abundant DNA template.
The hemoculture technique was also incapable of detecting mixed infections of cultivable trypanosomes: in three bat individuals, T. c. cruzi TcI was detected through the hemoculture while T. c. marinkellei and T. c. cruzi TcBat were detected through the blood clot (Table 3). In these cases, we hypothesize positive selection of T. c. cruzi TcI, outcompeting T. c. marinkellei and T. c. cruzi TcBat during amplification in culture medium. Mixed infections are probably more common than are noticed as the diagnosis methods exert selective pressure already from the blood puncture on, to hemoculture follow-up schedule (Deane et al., 1984; Yurchenko et al., 2009; Grybchuk-Ieremenko et al., 2014).
Besides the expected findings concerning the detection of T. c. cruzi, T. c. marinkellei and T. dionisii—as these trypanosome species are recognized as widely distributed among several bat species of different families (Jansen et al., 2018; Clément et al., 2019)—, the detection of a new MOTU (T. sp. Neobat 5) endorses our prediction about the underestimation of the diversity of the trypanosomes. Further, the detection of high rates of T. sp. Neobat 4 illustrates the magnitude of trypanosomes we overpass using the hemoculture technique.
This work corroborates the importance of bats as hosts of many Trypanosoma species of the T. cruzi clade—including the Neobat parasites—, and possibly the monoxenous Crithidia mellificae. Through molecular detection from the blood clot, we demonstrate that the unseen species within the T. wauwau clade is underestimated. Accordingly, T. sp. Neobat 4 seems to be a well-succeeded parasite, especially within C. perspicillata metapopulations across a wide geographical distribution. This work is an additional step forward to understand the biological aspects and life history of Trypanosoma spp. Neobats.
Declaration of competing interest
The authors declare that they have no conflict of interest.
Acknowledgments
We are grateful to Bruno Alves, Ewerth Janssen, Francisco Charles dos Santos, Natan Feitas, Rair Verde, Renan de França Souza, Renata de Cássia Pires, and Roberto Leonan Morim Novaes for the invaluable assistance with field work; to the ICMbio team “Juliao” Jorge Nascimento and Getulio Freitas for infrastructure support at the Guaribas Biological Reserve; to Rui Cerqueira, Ricardo Finotti, and Pedro Cordeiro-Estrela for PPBIO project coordination; to Caio Alves for the map construction. We thank Research Capacity Network (REDe-Brazil), Sustainable Science Institute, and ZikaPlan for the tutoring sessions provided to FMA during the Scientific Manuscript Writing Workshop 2021. This work was financially supported through grants from Fiocruz; Rio de Janeiro Carlos Chagas Filho Research Foundation (Faperj): JCNE (E-26/202.794/2019) to ALRR; and National Council for Scientific and Technological Development (CNPq): Productivity Grant to ALRR and AMJ, and CNPq/Universal (425293/2018-1) to ALRR. Bat and parasite survey in Guaribas Biological Reserve was funded by CNPq PPBIO Rede BioM.A. Inventários: Padrões de diversidade, biogeografia e endemismo de espécies de mamíferos, aves, anfíbios, drosófilas e parasitos na Mata Atlântica (Proc.: 457524/2012-0).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.ijppaw.2021.09.003.
Appendix A. Supplementary data
The following is the Supplementary data to this article:
References
- Allaerts W., Chang T.W. Skewed exposure to environmental antigens complements hygiene hypothesis in explaining the rise of allergy. Acta Biotheor. 2017;65:117–134. doi: 10.1007/s10441-017-9306-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Ayala S.C., D'Alessandro A. Insect feeding behavior of some Colombian fruit-eating bats. J. Mammal. 1973;54:266–267. [PubMed] [Google Scholar]
- Barros J.H.S., Lima L., Schubach A.O., Teixeira M.M.G. Trypanosoma madeira sp. n.: a species of the clade T. cruzi associated with the neotropical common vampire bat Desmodus rotundus. Int J Parasitol Parasites Wildl. 2018;8:71–81. doi: 10.1016/j.ijppaw.2018.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blaxter M., Mann J., Chapman T., Thomas F., Whitton C., Floyd R., Abebe E. Defining operational taxonomic units using DNA barcode data. Philos Trans R Soc London [Biol] 2005;360:1935–1943. doi: 10.1098/rstb.2005.1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouckaert R., Drummond A.J. bModelTest: Bayesian phylogenetic site model averaging and model comparison. BMC Evol. Biol. 2017;17:42. doi: 10.1186/s12862-017-0890-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouckaert R., Vaughan T.G., Barido-Sottani J., Duchene S., Fourment M., Gavryushkina A. BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 2019;15 doi: 10.1371/journal.pcbi.1006650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carvalho W.D., Silvestres S.M., Mustin K., Hilário R.R., Toledo J.J. Predation of an American fruit-eating bat (Artibeus sp.) by an Amazon tree boa (Corallus hortulanus) in the northern Brazilian Amazon. Acta Amazonica. 2018;49:24–27. [Google Scholar]
- Clément L., Dietrich M., Markotter W., Fasel N.J., Monadjem A., López-Baucells A., Scaravelli D., Théou P., Pigeault R., Ruedi M., Christe P. Out of Africa: the origins of the protozoan blood parasites of the Trypanosoma cruzi clade found in bats from Africa. Mol. Phylogenet. Evol. 2020;145 doi: 10.1016/j.ympev.2019.106705. [DOI] [PubMed] [Google Scholar]
- Cottontail V.M., Kalko E.K., Cottontail I., Wellinghausen N., Tschapka M., Perkins S.L., Miguel Pinto C. High local diversity of Trypanosoma in a common bat species, and implications for the biogeography and taxonomy of the T. cruzi clade. PloS One. 2014;9 doi: 10.1371/journal.pone.0108603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dario M.A., Moratelli R., Schwabl P., Jansen A.M., Llewellyn M.S. Small subunit ribosomal metabarcoding reveals extraordinary trypanosomatid diversity in Brazilian bats. PLoS Neglected Trop. Dis. 2017;11 doi: 10.1371/journal.pntd.0005790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dario M.A., Lisboa C.V., Costa L.M., Moratelli R., Nascimento M.P., Costa L.P., Leite Y.L.R., Llewellyn M.S., Xavier S.C.C., Roque A.L.R., Jansen A.M. High Trypanosoma spp. diversity is maintained by bats and triatomines in Espírito Santo state, Brazil. PloS One. 2017;12 doi: 10.1371/journal.pone.0188412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dario M.A., Lisboa C.V., Silva M.V., Herrera H.M., Rocha F.L., Furtado M.C., Moratelli R., Roque A.L.R., Jansen A.M. Crithidia mellificae infection in different mammalian species in Brazil. Int J Parasitol Parasites Wildl. 2021;15:58–69. doi: 10.1016/j.ijppaw.2021.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deane M.P., Jansen A.M., Mangia R.H.R., Gonçalves A.M., Morel C.M. Are our laboratory “strains” representative samples of Trypanosoma cruzi populations that circulate in nature? Mem. Inst. Oswaldo Cruz. 1984;79:19–24. [Google Scholar]
- dos Santos F.C.B., Lisboa C.V., Xavier S.C.C., Dario M.A., Verde R.S., Calouro A.M., Roque A.L.R., Jansen A.M. Trypanosoma sp. diversity in amazonian bats (chiroptera; mammalia) from Acre state, Brazil. Parasitology. 2018;145:828–837. doi: 10.1017/S0031182017001834. [DOI] [PubMed] [Google Scholar]
- Dougherty E.R., Carlson C.J., Bueno V.M., Burgio K.R., Cizauskas C.A., Clements C.F., Seidel D.P., Harris N.C. Paradigms for parasite conservation. Conserv. Biol. 2016;30(4):724–733. doi: 10.1111/cobi.12634. [DOI] [PubMed] [Google Scholar]
- Fenton M.B., Bernard E., Bouchard S. The bat fauna of Lamanai, Belize: roosts and trophic roles. J. Trop. Ecol. 2001;17:511–524. [Google Scholar]
- Frainer A., McKie B.G., Amundsen P., Knudsen R., Lafferty K.D. Parasitism and the biodiversity-functioning relationship. Trends Ecol. Evol. 2018;33:260–268. doi: 10.1016/j.tree.2018.01.011. [DOI] [PubMed] [Google Scholar]
- Gardner A.L. University of Chicago Press; Chicago: 2008. Mammals of South America, Volume 1: Marsupials, Xenarthrans, Shrews, and Bats. [Google Scholar]
- Giannini N.P., Kalko E.K.V. Trophic structure in a large assemblage of phyllostomid bats in Panama. Oikos. 2004;105:109–120. [Google Scholar]
- Grybchuk-Ieremenko A., Losev A., Kostygov A.Y., Lukeš J., Yurchenko V. High prevalence of trypanosome co-infections in freshwater fishes. Folia Parasitol. 2014;61:495–504. [PubMed] [Google Scholar]
- Guedes P.G., da Silva S.S.P., Santos K.S., Carneiro A.M.S., Lopes L.S., de Carvalho J.F. Padrão reprodutivo, dieta e parasitologia de Artibeus lituratus (Olfers, 1818) (Mammalia, Chiroptera) em parques urbanos do município do Rio de Janeiro (Rio de Janeiro, Brasil) Biotemas. 2020;33:1–16. [Google Scholar]
- Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 2010;59:307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- Hamilton P.B., Teixeira M.M., Stevens J.R. The evolution of Trypanosoma cruzi: the 'bat seeding' hypothesis. Trends Parasitol. 2012;28:136–141. doi: 10.1016/j.pt.2012.01.006. [DOI] [PubMed] [Google Scholar]
- Hatcher M.J., Dick J.T.A., Dunn A.M. Diverse effects of parasites in ecosystems: linking interdependent processes. Front. Ecol. Environ. 2012;10:186–194. [Google Scholar]
- Hodo C.L., Goodwin C.C., Mayes B.C., Mariscal J.A., Waldrup K.A., Hamer S.A. Trypanosome species, including Trypanosoma cruzi, in sylvatic and peridomestic bats of Texas, USA. Acta Trop. 2016;164:259–266. doi: 10.1016/j.actatropica.2016.09.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann F.G., Baker R.J. Comparative phylogeography of short‐tailed bats (Carollia: phyllostomidae) Mol. Ecol. 2003;12:3403–3414. doi: 10.1046/j.1365-294x.2003.02009.x. [DOI] [PubMed] [Google Scholar]
- Jansen A.M., Carreira J.C., Deane M.P. Infection of a mammal by monogenetic insect trypanosomatids (Kinetoplastida, trypanosomatidae) Mem. Inst. Oswaldo Cruz. 1988;83:271–272. doi: 10.1590/s0074-02761988000300001. [DOI] [PubMed] [Google Scholar]
- Jansen A.M., Xavier S.C.C., Roque A.L.R. Trypanosoma cruzi transmission in the wild and its most important reservoir hosts in Brazil. Parasites Vectors. 2018;11:502. doi: 10.1186/s13071-018-3067-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalko E.K.V., Handley C.O., Jr. Neotropical bats in the canopy: diversity, community structure, and implications for conservation. Plant Ecol. 2001;153:319–333. [Google Scholar]
- Kaufer A., Ellis J., Stark D., Barratt J. The evolution of trypanosomatid taxonomy. Parasites Vectors. 2017;10:287. doi: 10.1186/s13071-017-2204-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kostygov A.Y., Karnkowska A., Votýpka J., Tashyreva D., Maciszewski K., Yurchenko V., Lukeš J. Euglenozoa: taxonomy, diversity and ecology, symbioses and viruses. Open Biol. 2021;11 doi: 10.1098/rsob.200407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraeva N., Butenko A., Hlaváčová J., Kostygov A., Myškova J., Grybchuk D., Leštinová T., Votýpka J., Volf P., Opperdoes F., Flegontov P., Lukeš J., Yurchenko V. Leptomonas seymouri: adaptations to the dixenous life cycle analyzed by genome sequencing, transcriptome profiling and Co-infection with Leishmania donovani. PLoS Pathog. 2015;11 doi: 10.1371/journal.ppat.1005127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar S., Stecher G., Li M., Knyaz C., Tamura K. Mega X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018;35(6):1547–1549. doi: 10.1093/molbev/msy096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lafferty K.D. Biodiversity loss decreases parasite diversity: theory and patterns. Phil. Trans. Roy. Soc. Lond. 2012;367:2814–2827. doi: 10.1098/rstb.2012.0110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lima L., Espinosa-Álvarez O., Pinto C.M., Cavazzana M., Jr., Pavan A.C., Carranza J.C., Kim B.K., Campaner M., Takata C.S.A., Camargo E.P., Hamilton P.B., Teixeira M.M.G. New insights into the evolution of the Trypanosoma cruzi clade provided by a new trypanosome species tightly linked to Neotropical Pteronotus bats and related to an Australian lineage of trypanosomes. Parasites Vectors. 2015;8:657. doi: 10.1186/s13071-015-1255-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Llopis-Belenguer C., Balbuena J.A., Lange K., de Bello F., Blasco-Costa I. Towards a unified functional trait framework for parasites. Trends Parasitol. 2019;35:972–982. doi: 10.1016/j.pt.2019.09.003. [DOI] [PubMed] [Google Scholar]
- Lopes C.M.T., Menna-Barreto R.F.S., Pavan M.G., Pereira M.C.S., Roque A.L.R. Trypanosoma janseni n. sp. (trypanosomatida: trypanosomatidae) isolated from Didelphis aurita (mammalia: didelphidae) in the atlantic rainforest of Rio de Janeiro, Brazil: integrative taxonomy and phylogeography within the Trypanosoma cruzi clade. Mem. Inst. Oswaldo Cruz. 2018;113:45–55. doi: 10.1590/0074-02760170297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcot B.G., Vander Hayden M. In: Wildlife-Habitat Relationships in Oregon and Washington. Johnson D.H., O'Neil T.A., editors. Oregon State University Press; Corvallis: 2001. Key ecological functions of wildlife species; pp. 168–186. [Google Scholar]
- Morand S. (macro-) Evolutionary ecology of parasite diversity: from determinants of parasite species richness to host diversification. Int J Parasitol Parasites Wildl. 2015;4:80–87. doi: 10.1016/j.ijppaw.2015.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pires R.C. Oswaldo Cruz Institute; Rio de Janeiro, Rio de Janeiro: 2017. Natural Infection of Trypanosomatid Species in Bats (Mammalian: Chiroptera) Captured in Different Regions of Brazil and Development of Multiplex PCR as a Toll Diagnosis for Leishmania Spp. infection in wild mammals (Doctoral dissertation) [Google Scholar]
- Rangel D.A., Lisboa C.V., Novaes R.L.M., Silva B.A., Souza R.F., Jansen A.M., Moratelli R., Roque A.L.R. Isolation and characterization of trypanosomatids, including Crithidia mellificae, in bats from the Atlantic Forest of Rio de Janeiro, Brazil. PLoS Neglected Trop. Dis. 2019;13 doi: 10.1371/journal.pntd.0007527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigues M.S., Lima L., Xavier S.C.C., Herrera H.M., Rocha F.L., Roque A.L.R., Teixeira M.M.G., Jansen A.M. Uncovering Trypanosoma spp. diversity of wild mammals by the use of DNA from blood clots. Int J Parasitol Parasites Wildl. 2019;8:171–181. doi: 10.1016/j.ijppaw.2019.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Šlapeta J. Ten simple rules for describing a new (parasite) species. Int J Parasitol Parasites Wildl. 2013;2:152–154. doi: 10.1016/j.ijppaw.2013.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith A., Clark P., Averis S., Lymbery A.J., Wayne A.F., Morris K.D., Thompson R.C.A. Trypanosomes in a declining species of threatened Australian marsupial, the brush-tailed bettong Bettongia penicillate (Marsupialia: potoroidae) Parasitology. 2008;35:1329–1335. doi: 10.1017/S0031182008004824. [DOI] [PubMed] [Google Scholar]
- Thompson R.M., Mouritsen K.N., Poulin R. Importance of parasites and their life cycle characteristics in determining the structure of a large marine food web. J. Anim. Ecol. 2005;74:77–85. [Google Scholar]
- Vilar E.M., Silva-Filho T.P., da Silva R.R., Gomes E.S., da Silva L.A.M. Anthropic shelters used by bats in the semi-arid region of Pernambuco. Bol Soc Bras Mastozool. 2016;77:79–86. [Google Scholar]
- Violle C., Navas M.L., Vile D., Kazakou E., Fortunel C., Hummel I., Garnier E. Let the concept of trait be functional! Oikos. 2007;116:882–892. [Google Scholar]
- Willig M.R., Camilo G.R., Noble S.J. Dietary overlap in frugivorous and insectivorous bats from edaphic cerrado habitats of Brazil. J. Mammal. 1993;74:117–128. [Google Scholar]
- York H.A., Billings S.A. Stable-isotope analysis of diets of short-tailed fruit bats (chiroptera: phyllostomidae: Carollia) J. Mammal. 2009;90:1469–1477. [Google Scholar]
- Yurchenko V.Y., Lukes J., Jirku M., Maslov D.A. Selective recovery of the cultivation-prone components from mixed trypanosomatid infections: a case of several novel species isolated from Neotropical Heteroptera. Int. J. Syst. Evol. Microbiol. 2009;59:893–909. doi: 10.1099/ijs.0.001149-0. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.