Abstract
The nuclear export of the unspliced type D retrovirus mRNA depends on the cis-acting constitutive transport RNA element (CTE) that has been shown to interact with the human TAP (hTAP) protein promoting the export of the CTE-containing mRNAs. We report here that hTAP is a 619-amino-acid protein extending the previously identified protein by another 60 residues at the N terminus and that hTAP shares high homology with the predicted rat and mouse TAP proteins. We found that hTAP is a nuclear protein that accumulates in the nuclear rim and the nucleoplasm. We further demonstrated that hTAP is able to shuttle between the nucleus and the cytoplasm. Identification of the signals responsible for nuclear import (NLS) and export (NES) revealed that they are distinct but partially overlapping. NLS and NES of hTAP are active transferable signals that do not share similarities with known elements. The C-terminal portion contributes further to hTAP’s nuclear retention and contains a signal(s) for nuclear rim association. Taken together, our data show that hTAP is a dynamic protein capable of bidirectional trafficking across the nuclear envelope. These data further support hTAP’s role as an export factor of the CTE-containing mRNAs.
Posttranscriptional regulation is an essential regulatory step of many retroviruses and is necessary for virus production. This key regulatory step mediates the export of the unspliced, full-length viral RNA, which requires the interaction of viral and/or cellular factors. This controlled export of the viral RNA to the cytoplasm ensures the availability of the genomic RNA for packaging into the progeny virions and the production of the Gag/Pol polyproteins. Among the best-studied export systems are those used by the simian type D retroviruses (SRV/D) and the lentiviruses, such as human immunodeficiency virus type 1 (HIV-1) (for reviews see references 6, 14, and 28).
SRV/D expression is controlled by the essential cis-acting constitutive transport element (CTE) (5, 12, 54, 63). The SRV/D CTE (11, 55) and a related CTE-like element in a murine intracisternal A-particle retroelement (54) fold into an extended RNA stem-loop structure containing two conserved internal loops and an AAGA bulge. These loops and the bulge, the spacing of the loops within the RNA element, as well as the overall secondary structure of the element, have been shown to be essential features for CTE function (11, 55). Recently, we showed that the human TAP protein (hTAP) binds specifically to these internal loops and promotes nucleocytoplasmic transport of the CTE-containing intron lariat from the Xenopus oocyte nucleus (22). TAP had previously been identified as a factor binding to Tip, a herpesvirus saimiri protein responsible for cell transformation (60). The role of hTAP interaction with Tip is still unclear.
Whereas the SRV/D retroviruses have been proposed to utilize the cellular hTAP protein to export their unspliced mRNA (22), HIV-1 uses the viral Rev protein to promote the transport of the Rev responsive element (RRE)-containing mRNAs (7, 15, 23, 24, 30). Rev and several other shuttle proteins share a leucine-rich nuclear export signal (NES) (9, 18, 19, 21, 26, 31, 33, 37, 57, 61). The Rev export pathway was shown to be shared by some of these NES-containing proteins (9, 61), as well as by the putative factor(s) responsible for the export of spliceosomal U snRNAs and 5S rRNA (16). Essential for the export of proteins with the leucine-rich NES is the direct interaction with hCRM1, a member of the β-importin superfamily (2, 17, 20, 50). This interaction can be inhibited in the presence of the antibiotic leptomycin B (LMB) (29, 59), abolishing the export of the NES-containing protein. Recently, we and others also found that the nucleoporins Nup98 and Nup214 participate in Rev-hCRM1-mediated nuclear export in mammalian cells (3, 62).
The nucleocytoplasmic export pathway utilized by hTAP/CTE was shown to be distinct from that utilized by Rev/RRE (3, 39, 41, 43, 61). In agreement with this result, it was found that CTE function was not affected by LMB (reference 39 and unpublished observations), a finding which indicated that hTAP function does not involve binding to hCRM1. These findings suggested that hTAP function involves a yet-to-be-discovered mechanism. We therefore undertook a study to examine the molecular determinants of the hTAP protein in mammalian cells. In this report, we demonstrated that hTAP is a nuclear protein able to shuttle between the nucleus and the cytoplasm. Identification of the nuclear import (NLS) and export (NES) signals of TAP revealed that although these signals are partially overlapping, they are distinct. Importantly, the NES and NLS of hTAP do not share similarities with known signal sequences and appear to be novel. We also found that the C-terminal portion of hTAP contributes to its nuclear localization and contains signal(s) essential for the association with the nuclear rim. Taken together, these findings show that hTAP, like the HIV-1 Rev, is a shuttling protein. However, hTAP utilizes novel nuclear trafficking signals that are distinct from those of Rev and other shuttling proteins, suggesting that hTAP relies on novel routes of nuclear entry and exit. These data further support the previous finding that HIV and SRV/D utilize different RNA export pathways.
MATERIALS AND METHODS
Recombinant DNA.
The TAP61-619 cDNA clone was obtained from E. Izaurralde. Partial hTAP cDNAs were generated from the total RNA of human 293 cells by using reverse transcription and PCR amplification. The sense PCR primer was designed by using the human EST sequences predicted to encode the N terminus of the complete TAP cDNA and spanned nucleotides (nt) −64 to −41 (+1 corresponds to the first nucleotide of the coding sequence). The antisense primers were located at nt +382 to 402 and +459 to 480, respectively. The resulting cDNAs were cloned and sequenced. The coding sequence of the complete hTAP protein (amino acids [aa] 1 to 619) was assembled by using TAP61-619 cDNA and a partial cDNA providing the missing N-terminal peptide (aa 1 to 60). All cDNAs were subjected to double-stranded sequencing by the PCR-assisted fluorescent terminator method (ReadyReaction DyeDeoxy Terminator Cycle Sequencing Kit; ABI) according to the manufacturer’s instructions with the ABI Model 373A DNA Sequencing System. Sequencing data were analyzed by using Sequencher (Genes Code Corp.). hTAP contains the following changes compared to the sequence provided by Yoon et al. (60): N99D, W119C, and N256T (GenBank accession number AF126246). All eukaryotic expression vectors utilize the human cytomegalovirus (CMV) promoter. pCMV-GFPsg25 (53) was used to tag proteins with a strong mutant of green fluorescent protein (GFP). The GFP–β-galactosidase (β-Gal) expression vector was generated after insertion of the PCR-amplified GFP coding region from pFRED143 as a SacII-BamHI fragment into pFREDlacZ, generating a GFP–β-Gal open reading frame. pFRED143 (KH1035) and pFREDlacZ (KH1085) were kindly provided by K. Horie. pFRED143 contains a humanized version of a strong mutant of GFP (27a). hTAP and all fragments thereof were PCR amplified and cloned into the SacII and NheI sites located at the 5′ end of GFP. All TAP-GFP fusions contain the previously described Gly-Ala linker (51). The hemagglutinin (HA)-tagged hTAP was generated by PCR with a sense primer providing the HA sequence and was cloned into the BssHII and XbaI sites of pCMV37M1-10D replacing gag (45). The untagged expression vector contains the hTAP sequences from residues 1 to 619 and 61 to 619, respectively, cloned as BssHII-XbaI fragments into pCMV37M1-10D. To construct expression plasmids for the Tat-GFP export assay, Asp718-ClaI fragments spanning different TAP peptides were PCR amplified and inserted into pTat-GFP-NES (52), replacing the Rev NES. This generated a hybrid protein consisting of Tat–GFP–TAP peptides–GFP, which was verified by Western immunoblotting.
Cell culture, microscopy, and Western immunoblot analysis.
HLtat is a HeLa-derived cell line expressing HIV-1 Tat (46), human 293 is an embryonic kidney cell line, and A6 is a Xenopus laevis kidney cell line obtained through the American Type Culture Collection. Plasmid DNA was purified on Qiagen columns. Cells were transfected according to the calcium phosphate coprecipitation technique (293 cells) or SuperFect protocol (Qiagen) (HLtat and A6 cells). For microscopic analysis, HLtat cells were transfected in 35-mm glass-bottom plates. Cells were observed directly under an inverted microscope (Zeiss Axiovert 135TV), and the GFP fluorescence was detected by using a fluorescein isothiocyanate-fluorescent filter set. The fluorescent images were captured by using a digital charge-coupled device (CCD) camera (Photometrics) and processed by using IPLab Spectrum software. All pictures represent raw, unfiltered digital images. Confocal microscopy was performed as described previously (53). Treatment by actinomycin D and 5,6-dichlororibofuranosylbenzimidazole (DRB) was performed after pretreatment of the cells with cycloheximide as described earlier (1, 51). Western immunoblot analyses were performed with a rabbit anti-TAP antiserum (kindly provided by E. Izaurralde) or a 1:100 dilution of a mixture of anti-GFP antisera (Living Color Peptide antibody [Clontech] and anti-GFP antiserum [53]). Extracts from transfected 293 cells were separated on denaturing polyacrylamide gels (Novex) and blotted onto nitrocellulose and processed as described previously (23). For chloramphenicol acetyltransferase (CAT) expression, A6 cells were transfected with 2 μg of pDM138 (27) or pDM138CTE (39), together with 0.5 μg of the GFP-tagged TAP plasmids. CAT assays were performed 1 day posttransfection.
Fusion assay.
HLtat cells were seeded at 5 × 105 cells per 60-mm plate. The next day, the cells were transfected with 2 to 3 μg of the TAP expression vectors according to the SuperFect protocol (Qiagen). After 3 h the cells were washed, refed, and incubated for another 2 to 3 h. The cells were then trypsinized and placed into a total of 5 ml of complete Dulbecco Modified Eagle Medium. Aliquots of 0.5, 1, and 2 ml of transfected cells were mixed with 9 × 105 untransfected cells. After the volume was adjusted to 5 ml, 2.5 ml of the suspension was plated per 35-mm glass-bottom dish. After an overnight incubation the plates were inspected microscopically. For the fusion assay, only plates that had a low ratio of transfected to untransfected cells were chosen. The cells with or without pretreatment with cycloheximide were fused for 1.5 min according to the polyethylene glycol (PEG) protocol (1, 10).
RESULTS
Identification of complete human TAP protein.
By comparison to the database of human expressed sequence tags (ESTs), we found several human EST sequences (GenBank accession numbers T33563, R14280, AA307086, and AA173362) that overlapped the known hTAP cDNA (60) and extended beyond its 5′ end. These sequences also showed extensive homology to the recently sequenced murine and rat TAP cDNAs (GenBank accession numbers AF093140 and AF093139). We subjected cDNAs generated from RNA of human 293 cells to PCR with a sense primer derived from the alignment of these ESTs which were located upstream of the AUG initiation codon. Sequence analysis confirmed the presence of the extra 180 nt at the 5′ end of the cDNA, extending (GenBank accession number AF126246) the known hTAP sequence. The nucleotide sequence is highly homologous to the murine and the rat TAP cDNAs, as well as to the above-mentioned ESTs. This comparison revealed an open reading frame starting 60 aa upstream of the published hTAP initiator codon, similarly to the homologous murine and the rat open reading frames (Fig. 1A). Additionally, we found that the rat TAP cDNA contains two in-frame stop codons preceding the hTAP initiator codon, indicating that the published rat sequence encodes the complete TAP protein. Taken together, these data demonstrated that the extended open reading frame represented the complete human TAP protein.
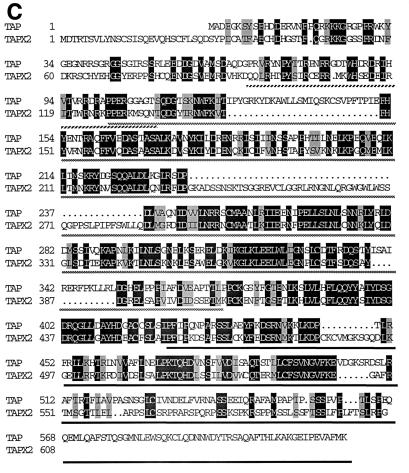
FIG. 1.
hTAP is conserved between species and contains homologies to other human proteins. (A) Comparison of amino acid sequences of human TAP and the predicted TAP proteins from mouse and rat. Lowercase letters indicate nonconserved residues. Boldface letters indicate the predicted classical NLS. The newly identified aa 1 to 60 of TAP are boxed. (B) Homology of the N-terminal TAP peptide to regions in the hnRNP K and X. laevis hnRNP C. Identical amino acids are shaded in black; similar amino acids are shaded in grey. The box indicates the position of the high-affinity bZip-like RBD identified in hnRNP C. (C) Comparison of hTAP to the predicted TAP-like human protein TAPX2. Identical and similar amino acids are indicated as in panel B. The NLS-NES region is indicated with a hatched bar, the RBD is shown with dotted bar, and the C-terminal portion is indicated with a black bar.
Interestingly, the sequence of aa 19 to 25 of the novel N-terminal sequence predicted a classical nuclear localization signal (for reviews, see references 8 and 38) (Fig. 1A). In addition, this N-terminal region shares sequence homology with some heterogeneous nuclear ribonucleoproteins (hnRNPs), such as hnRNP C and hnRNP K (Fig. 1B). The homologous region of hnRNP K (aa 230 to 287; GenBank accession number 585911) does not include the known functional determinants of this protein, such as nuclear localization signal (NLS), RNA binding motifs, or the nuclear shuttling domain (36). Interestingly, the hTAP homology in hnRNP C (aa 123 to 184; GenBank accession number 133263) includes its high-affinity bZIP-like RNA binding domain (aa 140 to 179) (32) implicated in binding to certain small nuclear RNAs (48). The relevance of these homologies to the function of the N-terminal portion of hTAP remains to be elucidated.
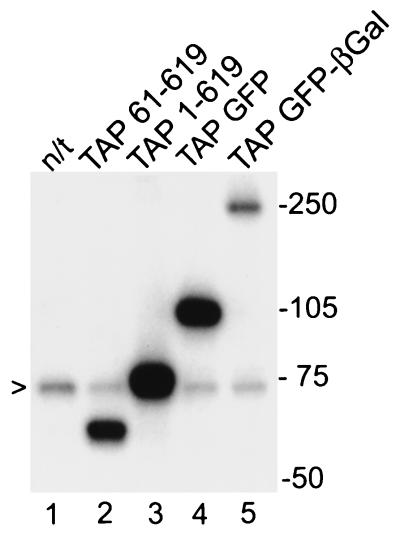
Western immunoblot with an antiserum raised against recombinant GST-TAP revealed that endogenous hTAP protein migrated with an apparent molecular mass of ∼70 kDa (Fig. 2, lane 1), which corresponds to the predicted size of the complete hTAP. No TAP-specific bands were found that matched the previously identified shorter TAP predicted to migrate at ∼63 kDa. To further support this conclusion, we analyzed the size of TAP1-619 and TAP61-619 expressed in human cells after transient transfections. We found that only TAP1-619 produced a protein of ∼70 kDa comigrating with the endogenous hTAP protein (lane 3), while TAP61-619 produced a smaller protein of ∼63 kDa (lane 2). Taken together, these data established that the genuine human TAP protein contains the newly identified 60 N-terminal aa residues in addition to the previously identified sequence (60). This finding also resolves the observed discrepancy in sizes between the recombinant hTAP protein (spanning aa 60 to 619) and the purified CTE binding activity (22).
FIG. 2.
The genuine hTAP is a 70-kDa protein. Extracts from human 293 cells transfected with the different TAP expression plasmids were separated on a 10% denaturing polyacrylamide gel, blotted onto nitrocellulose, and incubated with an anti-TAP antiserum and then 125I-labeled protein A. The relevant positions of the Rainbow molecular weight markers (Amersham) are indicated on the right. The arrow indicates the endogenous hTAP protein. Lane 1, not transfected (n/t); lanes 2 to 5, transfected with the indicated plasmids.
Subcellular localization of hTAP.
To study the localization of hTAP, we generated a fusion protein by tagging hTAP with a strong mutant of the GFP (53) or a GFP–β-Gal hybrid protein, which allowed us to visualize hTAP directly upon transfection. Both the GFP and GFP–β-Gal moieties do not contain signals for active nuclear import and export. The ∼24-kDa GFP can efficiently enter and exit the nucleus by diffusion due to its small size (53) and, as expected, it was found both in the nucleus and the cytoplasm (Fig. 3A). The size of GFP–β-Gal hybrid protein (∼143 kDa) is above the diffusion cutoff, and therefore it cannot enter the nucleus and localizes exclusively to the cytoplasm (Fig. 3B).
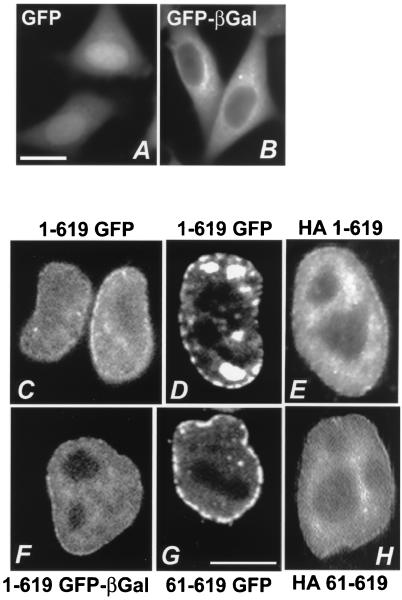
FIG. 3.
hTAP is a nuclear protein. HLtat cells were transfected with the indicated plasmids, and the tagged proteins were visualized the following day. While the GFP-tagging allows direct visualization in living cells, the HA-tagged proteins were visualized upon staining with anti-HA-antibody and rhodamine-labeled anti-mouse antiserum. The images were obtained by fluorescent microscopy and by use of a CCD camera (A and B; bar = 20 μm) and by confocal microscopy (C to H; bar = 10 μm). Panels: A, GFP; B, GFP–β-Gal; C and D, TAP1-619 GFP; E, HA-tagged TAP1-169; F, TAP1-619 GFP–β-Gal; G, TAP61-619 GFP; H, HA-tagged TAP61-619. Panels C and D show confocal images taken at different planes through the nucleus.
We took advantage of the properties of GFP and GFP–β-Gal to determine whether hTAP and TAP fragments contain NLSs. We found that TAP1-619 GFP (Fig. 3C and D) and TAP1-619 GFP–β-Gal (Fig. 3F; see also Fig. 4I) localized uniquely to the nucleus of HeLa cells. hTAP appeared to concentrate in the nuclear rim and the nucleoplasm but was excluded from the nucleoli. Similar localization of hTAP was found when the protein was tagged at the N terminus by the HA peptide (Fig. 3E). The rim association is slightly less pronounced when hTAP is tagged by GFP–β-Gal (compare Fig. 3F and C) which could probably be due to the bigger size of the β-Gal moiety. Similar subcellular localization of hTAP was found in 293 cells (data not shown). The tagging of hTAP was necessary since the available anti-TAP antiserum did not allow the detection of endogenous hTAP in the immunofluorescence assay.
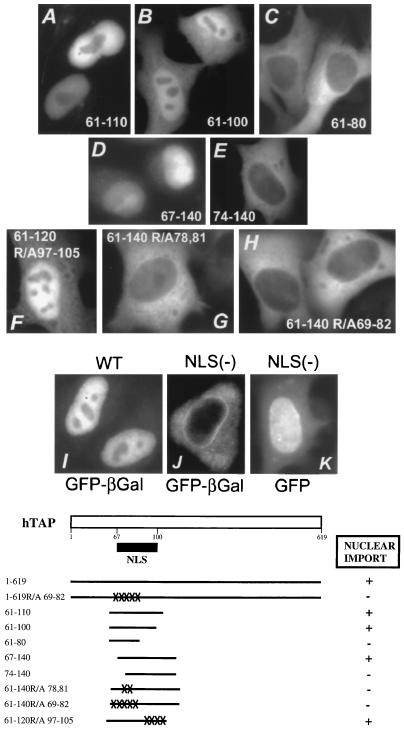
FIG. 4.
Identification of the nuclear localization signal of hTAP. (Top) HLtat cells were transfected with the indicated plasmids, and the TAP hybrid proteins were visualized as described in Fig. 3. All images were obtained by using fluorescent microscopy and a CCD camera. The TAP peptides or the intact hTAP protein were tagged with either GFP–β-Gal (panels A to C, E, and G to J) or GFP (panels D, F, and K). (Bottom) Schematic representation of hTAP indicating the location of the NLS. Different TAP mutants shown in Fig. 4A are represented.
Identification of the NLS.
We next generated a series of N- and C-terminal deletion mutants of hTAP to identify the NLS (Fig. 4, summarized in Table 1). TAP fragments were fused in frame to GFP, and the localization of the hybrid proteins was investigated upon transfection of HeLa cells. To distinguish active nuclear import from diffusion, the crucial TAP mutants were also tagged with the higher-molecular-weight moiety consisting of a GFP–β-Gal hybrid protein, which does not allow diffusion into the nucleus (Fig. 3B). Therefore, the use of the GFP–β-Gal tag warranted that only active transport into the nucleus could be observed. Western immunoblot analysis confirmed that TAP-GFP migrated at ∼100 kDa, whereas TAP–GFP–β-Gal migrated at ∼250 kDa according to their predicted molecular sizes (Fig. 2, lanes 4 and 5, respectively).
TABLE 1.
Identification of the nuclear import signal of hTAPa
| hTAP aa | GFP fusion
|
GFP–β-Gal fusion
|
||
|---|---|---|---|---|
| Nuclear accumulation | Nuclear rim association | Local-ization | Nuclear rim association | |
| 1–619 | Yes | Yes | N | Yes |
| 1–60 | No | No | C | No |
| 61–619 | Yes | Yes | N | Yes |
| 61–560 | Yes | No | ||
| 61–507 | Yes | No | ||
| 61–380 | Yes | No | N | No |
| 61–341 | Yes | No | ||
| 61–273 | Yes | No | ||
| 61–175 | Yes | No | ||
| 61–160 | Yes | No | ||
| 61–140 | Yes | No | ||
| 61–134 | Yes | No | ||
| 61–128 | Yes | No | N | No |
| 61–120 | Yes | No | N | No |
| 61–110 | N | No | ||
| 61–100 | Yes, impaired | No | N+C | No |
| 61–80 | No | No | C | No |
| 67–140 | Yes | No | ||
| 81–140 | No | No | C | No |
| 95–140 | No | No | ||
| 1–619 R/A 69–82 | Yes, impaired | Yes | C | Yes |
| 61–140 (Δ81–109) | Yes, impaired | No | C | No |
| 61–140 (Δ81–119) | Yes, impaired | No | C | No |
| 61–120 R/A 97–105 | N+C | No | ||
| 61–140 R/A 69–82 | No | No | C | No |
| 61–140 R/A 78,81 | C | No | ||
N, nucleus; C, cytoplasm.
The newly identified N-terminal fragment (TAP1-60) did not affect the localization of the GFP tags (Table 1), despite the presence of a predicted classic NLS (Fig. 1A). On the other hand, GFP- and HA-tagged TAP61-619 (Fig. 3G and H) localized to the nucleus such as the intact TAP1-619 (C to F). This observation further suggested that these newly identified N-terminal 60 residues were not essential for hTAP’s nuclear localization. Although this region is highly conserved among TAP proteins from different species (Fig. 1A), its role is still unclear.
We next analyzed a series of C-terminal deletions based on TAP61-619 GFP–β-Gal fusions (Table 1 and Fig. 4). All peptides with C termini ranging from aa 619 (Fig. 4I) to 110 (Fig. 4A) localized exclusively to the nucleus. TAP61-100 was found both in the nucleus and the cytoplasm (Fig. 4B), indicating that its nuclear accumulation is impaired, while TAP61-80 localized only to the cytoplasm (Fig. 4C), indicating that further deletion of 20 aa from TAP61-100 destroyed the NLS. This study identified residue 100 of hTAP as the C-terminal boundary of the core NLS. Since TAP61-110, but not TAP61-100, localized exclusively to the nucleus, residues 100 to 110 contributed to import and/or nuclear retention.
We next constructed a series of N-terminal deletions of GFP–β-Gal-tagged TAP61-140 (Table 1 and Fig. 4). TAP67-140 showed an intact NLS (panel D), while further deletions to aa 74 (panel E), 81, and 95 (see Table 1) abrogated NLS function. This analysis defined aa 67 as the N-terminal border of the NLS. In addition, two mutants of TAP61-140, which contain internal deletions of residues 81 to 109 and 81 to 119, respectively, are both unable to localize to the nucleus (Table 1). This finding further showed that residues 81 to 109 are essential for import. Taken together, the mutagenesis experiments demonstrated that the NLS of hTAP spans a relatively large region from aa 67 to 100.
Inspection of this region of hTAP did not reveal resemblance to any known NLS motifs. Notably, this region is very hydrophilic and has a high content of basic residues, especially arginine. To further characterize this NLS, we made alanine substitutions of several arginine residues. TAP61-120R/A97-105, a mutant containing four alanine substitutions at positions R97, R98, R100, and R105, was still able to localize to the nucleus when linked to GFP–β-Gal (Fig. 4F and Table 1). This result showed that these arginine residues were not essential for nuclear import. We noted, however, that the cells transfected with this mutant TAP fusion protein showed also significant cytoplasmic staining, revealing that substitution of these arginine residues led to impaired import and/or impaired nuclear retention. This finding further supported our results obtained with TAP61-100 (see above), which suggested that residues 100 to 110 contribute to nuclear accumulation mediated by the hTAP NLS.
We then made alanine substitutions of some arginine residues in the N-terminal portion of TAP61-140, which contains the intact NLS. TAP61-140R/A78,81 has two changes at R78 and R81, while TAP61-140R/A69-82 has five changes at R69, R71, R78, R81, and R82. Notably, mutations of either two or five arginine residues destroyed the NLS, resulting in exclusive cytoplasmic localization of GFP–β-Gal (Fig. 4G and H, respectively). These results demonstrated that residues R78 and R81 are essential for the NLS.
Role of the C-terminal portion in nuclear association of hTAP.
We explored whether there are additional NLS signals within hTAP (Table 2 and Fig. 5). TAP peptides 167 to 380 and 266 to 380 tagged with GFP did not affect GFP localization (data not shown). Interestingly, GFP-tagged TAP266-619, which lacks the above-described NLS, was also found to accumulate in the nucleus and the nuclear rim (Fig. 5A). To further delineate this signal, a series of N-terminal deletions of the GFP-tagged TAP266-619 were analyzed. TAP peptides 375 to 619 (Table 2) and 412 to 619 (panel B) also accumulated in the nucleus and the nuclear rim, whereas peptide 507 to 619 (panel C) and further N-terminal deletions thereof lost this property (Table 2). This finding pointed to an additional potential NLS located between residues 412 and 507. We further observed that removal of the C-terminal 59 aa, as in TAP375-560 and TAP412-560 (panel D), resulted in localization indistinguishable from that of GFP alone, suggesting an additional role of residues 560 to 619 for the nuclear accumulation. These findings, taken together, indicated that a combination of these two regions (residues 412 to 507 and 560 to 619) contributed to the nuclear accumulation of the GFP-tagged TAP fragments.
TABLE 2.
Identification of the nuclear retention and rim association determinants of hTAPa
| hTAP aa | GFP fusion
|
GFP–β-Gal fusion
|
||
|---|---|---|---|---|
| Nuclear accumulation | Nuclear rim association | Local-ization | Nuclear rim association | |
| 1–619 | Yes | Yes | N | Yes |
| 61–619 | Yes | Yes | N | Yes |
| 61–619 S585P | Yes | No | ||
| 61–619 S585A | Yes | Yes | ||
| 140–619 | C | Yes | ||
| 266–619 | Yes | Yes | C | Yes |
| 266–619 S585P | C | No | ||
| 266–619 S585A | C | Yes | ||
| 375–619 | Yes | Yes | ||
| 375–560 | No | |||
| 412–619 | Yes | Yes | C | Yes |
| 412–560 | No | |||
| 507–619 | No | |||
| 540–619 | No | |||
| 550–619 | No | |||
| 566–619 | No | |||
N, nucleus; C, cytoplasm.
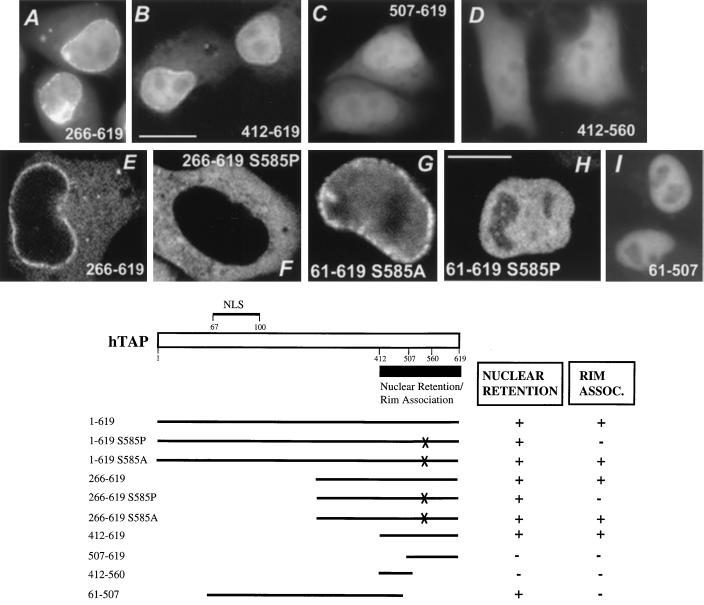
FIG. 5.
Analysis of the C-terminal portion of TAP. (Top) HLtat cells were transfected with the indicated plasmids, and the TAP hybrid proteins were visualized as described in Fig. 3. The TAP peptides were tagged with either GFP (panels A to D and panel I) or GFP–β-Gal (panels E to H). All images were obtained by using fluorescent microscopy and a CCD camera (bar = 20 μm) except panels E to H (bar = 10 μm), for which confocal microscopy was used. Tagging of C-terminal peptides of TAP with GFP (panels A to D) revealed the presence of nuclear retention signal(s), while tagging of the same peptides with GFP–β-Gal (panels E and F) demonstrates that this region lacks an active NLS. (B) Schematic representation of hTAP indicating the locations of the NLS (Fig. 4) and the signal for nuclear retention and rim association. Different TAP mutants shown in Fig. 5A are represented.
To distinguish between active transport and nuclear retention resulting from diffusion of these relatively small GFP-tagged hybrid proteins into the nucleus, we also analyzed GFP–β-Gal-tagged peptides. Notably, all these peptides failed to affect the cytoplasmic localization of GFP–β-Gal (Table 2; see also Fig. 5E, TAP266-619), demonstrating that the C-terminal portion of hTAP did not contain an active import signal. However, our analysis showed that this region contains signal(s) that can contribute to the nuclear retention of hTAP. The role of such retention signal(s) for the localization of the complete protein was unclear, since the C-terminal portion is not essential for the nuclear accumulation of hTAP mediated by its NLS (see above, Fig. 4 and Table 1).
However, through an independent line of experiments, we found that the C-terminal portion is essential for the nuclear rim association of hTAP in human cells. We made the observation that TAP61-619 with a fortuitous serine-to-proline change of aa 585 lost its association to the nuclear rim, although it still localized exclusively to the nucleus (Fig. 5H). However, an alanine substitution at S585 did not affect the nuclear rim association (panel G). We concluded that the S585P mutation might affect the folding of the protein, thereby abolishing hTAP’s rim association, whereas the S585 residue itself is not required for the rim association. In further support of this finding, we noted that all C-terminal deletion mutants, including TAP61-560 and TAP61-507 (Fig. 5I) failed to show the nuclear rim association (see also Table 2). This result independently pointed to a role of the region containing residues 560 to 619 in the nuclear rim association of hTAP. We also observed that the GFP–β-Gal-tagged mutants (which lack the NLS) such as TAP140-619 (Table 1), TAP266-619 (Fig. 5E), and TAP412-619 (Table 2) localized to the cytoplasm, as well as to the nuclear envelope, resembling the nuclear rim association observed for the intact hTAP. As observed with TAP61-619, mutation of S585P (panel F) but not S585A abolished the rim association (see also Table 2).
Taken together, these findings showed that the C-terminal portion of hTAP contains signals for nuclear retention and, importantly, for nuclear rim association. Our experiments also showed that these signals function independently of hTAP’s trafficking signals (see also below).
We further employed a functional test to support a role for the identified signals in hTAP. hTAP is present in all mammalian cells and CTE functions in all mammalian cells tested. Cotransfection of exogenous TAP did not further increase CTE function in these cells, indicating that TAP is present at saturating levels. However, we identified a X. laevis kidney cell line (A6) where the CTE is inactive. These studies were performed by using the previously published CAT indicator system DM138 in which CAT is only expressed from the unspliced mRNA and thus CAT production reflects posttranscriptional regulation (27). In a permissive cell line such as HeLa, transfection of a plasmid containing a functional CTE (pDM138CTE) led to an at least 20-fold-increased CAT production compared to pDM138. Upon cotransfection of A6 cells with DM138CTE plus tagged or untagged hTAP, we observed a three- to fourfold activation of CAT production. In contrast, the presence of a TAP mutant (TAP61-380) lacking the C terminus or of a TAP mutant (TAP266-619) lacking the NLS and NES and the CTE RNA binding domain (RBD) did not have any effect. These findings demonstrate that the above-identified signals of hTAP are important for function.
Role of the signals for nuclear localization and for rim association in import of the complete hTAP protein.
We have identified two signals, the NLS and the nuclear retention signal, which mediate the nuclear accumulation of hTAP. We next investigated the contribution of these signals when present within the complete hTAP protein. To do this, we introduced the five alanine substitutions shown to destroy the NLS in the TAP61-140 (Fig. 4) into the otherwise intact hTAP, generating the NLS(−) TAP1-619R/A69-82.
Upon tagging with GFP–β-Gal, we found that this TAP mutant was found exclusively in the cytoplasm and accumulated strongly in the nuclear rim (Fig. 4J). Thus, the localization of the mutant TAP1-619R/A69-82 was indistinguishable from that of a N-terminal deletion mutant lacking the NLS (such as TAP266-619 [Fig. 5E]), demonstrating that the identified NLS is necessary and sufficient for the active import of the complete hTAP. These data further showed that there is no other active NLS present within hTAP. As expected from the data shown above, the signal(s) essential for nuclear rim association can act independently of the NLS, resulting in prominent accumulation at the nuclear envelope.
To understand the effect of the NLS elimination within the complete hTAP protein more closely resembling the natural protein, we also studied the same NLS(−) TAP mutant tagged with GFP only. We found that the GFP-tagged mutant TAP no longer localized exclusively to the nucleus but could be detected in the nucleus, the nuclear rim, and the cytoplasm (Fig. 4K). Again, as observed for the GFP–β-Gal-tagged mutant, its localization was indistinguishable from those of N-terminal deletion mutants that lack the NLS such as TAP266-619 (Fig. 5A).
Taken together, these data showed that upon elimination of the NLS by introducing amino acid changes or by deletion of the N-terminal portion, the GFP-tagged (but not the GFP–β-Gal-tagged) mutant TAP was still able to accumulate in the nucleus, although its import was clearly impaired. This finding suggested that although the size of the hybrid protein is above the expected diffusion cutoff, some diffusion into the nucleus and the nuclear retention contributed significantly to the observed nuclear accumulation of the GFP-tagged NLS(−) TAP1-619. Since hTAP accumulated very strongly in the nuclear rim independent of the presence of an active NLS, it is possible that hTAP’s diffusion into the nucleus is enhanced due to its concentration in the vicinity of the diffusion channels. Taken together, these findings led us to conclude that the unique active NLS, as well as the nuclear retention signal(s), contributes to the nuclear accumulation of hTAP.
hTAP is a nuclear shuttle protein.
Since hTAP had been shown to be essential for the export of the CTE RNA from the nucleus to the cytoplasm of Xenopus oocytes (22), we examined whether hTAP can shuttle between the nucleus and the cytoplasm. To address this question, we studied the trafficking of hTAP and its mutants by using a PEG-mediated fusion assay. Briefly, HeLa cells were transfected with expression vectors for GFP-tagged TAP and then mixed with an excess of untransfected cells. The following day, the cells were subjected to fusion, and hTAP’s ability to translocate to neighboring nuclei within the syncytia was monitored. As shown in Fig. 6A (top panels), TAP1-619 could be exported from the transfected nucleus (as indicated by an arrow) and was efficiently imported into the acceptor nuclei. Similar data were obtained independent of pretreatment of the cells with cycloheximide and independent of the tagging system (GFP or GFP–β-Gal). To demonstrate the specificity of hTAP shuttling, we cotransfected cells with GFP-tagged TAP and the blue variant of GFP (BFP)-tagged NES(−) Rev (RevM10BL-BFP) (51, 53). Upon fusion, we found only hTAP to accumulate in the acceptor nuclei, whereas the NES(−) Rev remained in the donor nucleus (Fig. 6A, bottom panels). Taken together, these findings demonstrated that hTAP is a bona fide shuttle protein.
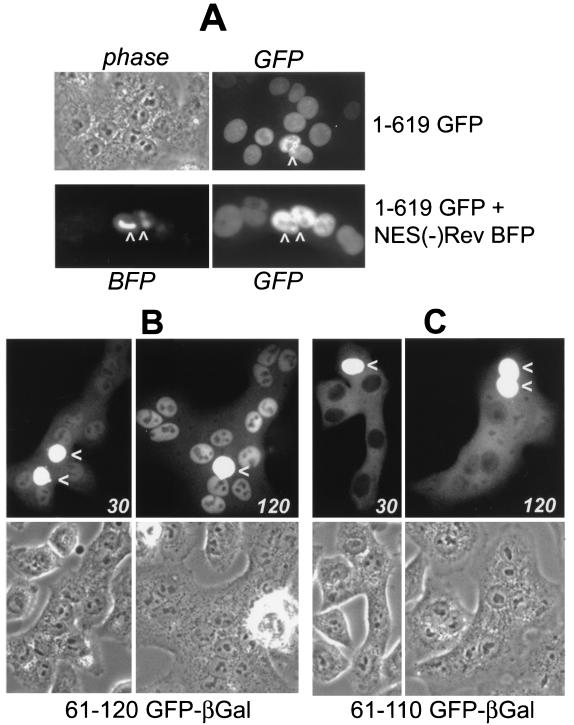
FIG. 6.
hTAP is a shuttle protein. Transfected HLtat cells were mixed with an excess of untransfected HLtat cells. The next day, the cells were fused by using PEG, and the GFP fluorescence was detected by using fluorescent microscopy and a CCD camera. (A) Fusion of cells transfected with GFP-tagged TAP1-619 in the absence (top panels) and presence (bottom panels) of the NES(−) Rev M10BL-BFP. (B and C) Fusion of cells transfected with GFP–β-Gal-tagged TAP61-120 and TAP61-110 fixed with 3.7% formaldehyde after 30 and 120 min, respectively. The arrows indicate the donor nuclei.
Since some NLS determinants, such as those found in hnRNP A1, hnRNP K, HuR, and HIV-1 Rev (13, 34–36), were shown to exhibit transcription-dependent import activity, we subjected cells transfected with GFP-tagged TAP1-619 to treatment with either actinomycin D or DRB. We did not observe any change in hTAP’s nuclear localization as a result of these treatments (data not shown). In contrast, in control plates cells transfected with a Rev expression plasmid, but not those that contain a NES(−) Rev mutant (RevM10BL), Rev translocated to the cytoplasm as expected (34, 51). We concluded that the NLS-mediated import of hTAP was not dependent on active RNA polymerase II transcription.
Identification of NES, a signal distinct from the NLS.
To characterize the NES, we tested the ability of different hTAP peptides for their ability to shuttle between the nucleus and the cytoplasm (Fig. 6B and C). To exclude any effect due to diffusion of the protein, we examined only those mutants that were tagged by GFP–β-Gal. Fusion proteins containing TAP61-128 and TAP61-120 (Fig. 6B) were able to translocate to the acceptor nuclei as efficiently as the intact hTAP (see also Table 3 and Fig. 7B). Remarkably, although TAP61-110 (Fig. 6C) was exported efficiently, it accumulated in the cytoplasm and appeared more slowly in the acceptor nuclei. While the import of TAP61-120 was clearly detectable within 30 min of fusion, it took more than 2 h for TAP61-110 to accumulate in the acceptor nuclei (Fig. 6). We concluded that although TAP61-110 contains an intact NLS (Fig. 4B) leading to exclusive nuclear accumulation at steady state, it may have a slightly reduced import rate or impaired nuclear retention detectable only in a real-time experiment. Taken together, these data demonstrated that residues 61 to 110 are sufficient to promote nuclear export and indicate that NES and NLS are partially overlapping.
TABLE 3.
Identification of the nuclear export signal of hTAP
| hTAP aa | NLS | Export
|
|
|---|---|---|---|
| Shuttling in fusion assay | Tat-GFP-hybrid | ||
| 1–619 | Yes | Yes | NAa |
| 61–128 | Yes | Yes | |
| 61–140 | Yes | No | |
| 61–120 | Yes | Yes | |
| 61–110 | Yes | Yes | |
| 61–100 | Yes, impaired | No | |
| 81–140 | No | Yes | |
| 90–128 | Yes, impaired | ||
| 61–140 (Δ81–119) | No | No | |
| 61–140 R/A 78,81 | No | Yes, impaired | |
| 61–140 R/A 69–82 | No | Yes | |
| 412–619 | No | No | |
| 540–619 | No | Yes | |
NA, not applicable.
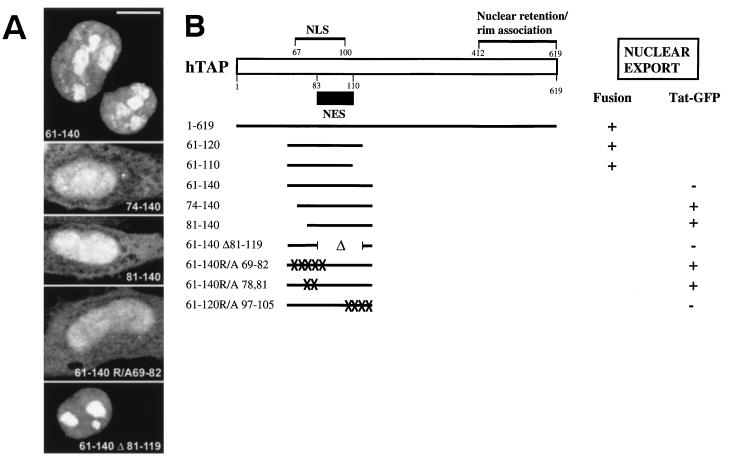
FIG. 7.
Identification of the NES of hTAP. (A) Different peptides of hTAP, as indicated, were inserted between two moieties of GFP in pTat-GFP (52). HLtat cells were transfected and analyzed as described above. All images were obtained by confocal microscopy. Bar = 10 μm. (B) Schematic representation of hTAP indicating the locations of the NLS (Fig. 4), the signal(s) for nuclear retention and rim association (Fig. 5), and the NES. Different hTAP mutants shown in the fusion assay (Fig. 6) and the export assay (Fig. 7A) are represented.
Since the NLS and NES of hTAP appeared to overlap, we sought an export assay, which would allow us to dissect these two determinants. Recently, Stauber and Pavlakis (52) published an elegant assay for the identification of NES elements. This assay takes advantage of HIV-1 Tat providing an NLS linked to two moieties of GFP, generating a protein of ∼64 kDa that localizes to the nucleus. Upon insertion of the Rev NES, this Tat-GFP-NES hybrid was found exclusively in the cytoplasm (52), since it is exported more efficiently than it is imported. Therefore, this test system allows the detection of a strong NES able to counteract the nuclear import mediated by Tat NLS. We tested whether different peptides of hTAP were able to affect the localization of Tat-GFP (Table 3 and Fig. 7). Insertion of TAP residues 61 to 140 (Fig. 7A) and 61 to 100 did not have any effect on the subcellular localization of the Tat-GFP fusion proteins. This result was expected, since this peptide also contained hTAP’s intact NLS, which could additionally contribute to the nuclear accumulation of the Tat-GFP hybrid protein. Interestingly, the presence of peptides 74 to 140 and 81 to 140, both of which lack the functional hTAP NLS, resulted in considerable export of the Tat-GFP hybrid protein (Fig. 7A). These results identified a potent NES located between residues 81 and 140 of hTAP, which was able to counteract Tat’s NLS and to relocalize a heterologous nuclear protein to the cytoplasm. Since the TAP peptide 81-140 lacks a functional NLS (Table 1), these data demonstrated further that NLS and NES are distinct and can be structurally dissected. This finding was supported by the analysis of Tat-GFP proteins containing the NLS(−) TAP peptides 61-140R/A69-82 (Fig. 7A) or 61-140R/A78,81 (Table 3) having either five or two alanine substitutions, respectively. Both of these peptides promoted the export of Tat-GFP. On the other hand, TAP61-140Δ81-119 (Fig. 7A), which lacks both residues 81 to 119 and a functional NLS (Table 1), did not export. Taken together, the data from the fusion assay and the Tat-GFP export assay demonstrated that residues ∼83 to 110 of hTAP are sufficient for NES function (Fig. 7B). To support this conclusion, peptide 81-110 was fused to the Tat-GFP and was shown to promote export (Table 3). Therefore, these studies have revealed that the NLS (ca. residues 67 to 100) and the NES (ca. residues 83 to 110) are distinct but partially overlapping elements within the N-terminal portion of hTAP.
We also observed that the TAP peptide 540-619, but not a bigger peptide spanning residues 412 to 619, was able to translocate Tat-GFP to the cytoplasm (Table 3). This finding led us to hypothesize that peptide 540-619 would be also unable to execute export in the context of the complete protein. Therefore, these results do not support the presence of a bona fide NES between residues 540 and 619. Taken together with the finding that export occurred independently of the C-terminal portion, these data indicated that this region was not essential for hTAP’s nuclear export. These results further demonstrated that hTAP contains a potent signal able to mediate active nuclear export that is located between ca. residues 83 to 110.
Conservation of hTAP signals in TAP-related open reading frames.
Database comparison of hTAP suggested two additional genes encoding TAP homologs in the human genome which are located on chromosome X as noted previously (44). The model mRNAs were constructed from these genomic sequences by using exon search and assembly programs at the Sanger Centre (43a). These two mRNAs contained predicted coding sequences which we termed TAPX1 (363 aa) and TAPX2 (607 aa). Further analyses indicated that TAPX2 coding sequences, but not those of TAPX1, are found in human EST database (GenBank accession numbers AI028725, AA918605, AI208506, and AI283407), indicating that the TAPX2 gene is transcribed. Sequence comparison of the predicted TAPX2 protein and hTAP revealed a high degree of homology, especially in the recently identified RNA-binding domain (RBD) (4) and the C-terminal region (Fig. 1C). Interestingly, the region spanning hTAP’s NLS-NES shows also significant homology and shares the arrangement of some of the arginine residues. Importantly, the R78 and R81 residues of hTAP, which we identified to be necessary for the NLS function are conserved in TAPX2. These data led us to speculate that TAPX2 may also contain an NLS that is similar to that of hTAP, supporting our data from the mutagenesis analysis. These data further suggest that hTAP may be a member of a protein family with a related function.
DISCUSSION
The CTE utilizes a nuclear export pathway that is distinct from the hCRM1-mediated export (3, 39, 41, 43, 61) and was shown to be shared by cellular mRNAs (4, 22). The recent finding that hTAP directly bound to the CTE RNA element and mediated the CTE-directed nucleocytoplasmic export, identified hTAP as the first component of this pathway (4, 22). As a first step to elucidate the mechanism of hTAP-mediated export, we studied the subcellular localization of hTAP. Here we found that hTAP tagged by either an HA-peptide or by GFP is found on the nuclear rim and in the nucleoplasm of human cells. hTAP is able to shuttle between the nucleus and the cytoplasm, a finding which is in agreement with its proposed nuclear export function. In this report, we present the identification of signal sequences (the NLS, the NES, and the signals for nuclear retention and rim association) that mediate subcellular localization and nucleocytoplasmic trafficking of hTAP (summarized in Fig. 8).
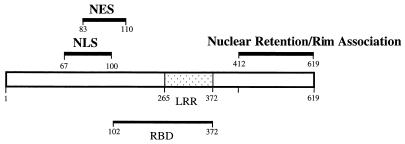
FIG. 8.
Signals identified on hTAP. Schematic representation of hTAP shows the NLS and NES are distinct elements that are partially overlapping. In addition, residues 100 to 110 were shown to contribute to nuclear import or nuclear retention. The NLS and NES are located next to, but are distinct from, the recently characterized RBD (4). The RBD encompasses the LRR (47). The nuclear retention and rim association signal(s) are located in the C-terminal portion of hTAP.
The signals mediating active nuclear import (ca. residues 67 to 100) and export (ca. residues 83 to 110) are partially overlapping in the N-terminal part of hTAP, a region rich in basic residues, especially arginine. We demonstrated that some of these arginine residues are essential for NLS function. The NLS and NES could be dissected by mutagenesis, supporting the model that this region contains distinct binding sites for the import and for the export factors. However, their partial overlap points to the possibility that the binding of import and export factors to hTAP could be mutually exclusive. We also noted that residues 100 to 110 contribute to the nuclear accumulation, affecting either nuclear entry or nuclear retention. Inspection of the primary structure of the NLS-NES region did not reveal similarities to known signals of nucleocytoplasmic trafficking, including the bidirectional trafficking signals found in hnRNP A1 (35, 49), hnRNP K (36), and HuR (13, 42). We also found that the import of hTAP is insensitive to inhibitors of RNA polymerase II transcription, a feature distinguishing hTAP from other shuttling proteins, such as hnRNP A1 (35, 49, 58), HuR (42), or Rev (34, 51), which suggested that its import likely involves different factors. This region did also not reveal matches to the previously described importin-α or importin-β recognition signals (25, 40, 56). It remains to be determined whether the novel shuttling signals of hTAP represent noncanonical binding sites of the currently known receptors or, alternatively, whether they interact with novel components of the trafficking machinery. Clearly, the NES of hTAP does not share any similarity with the signature motif of the leucine-rich NES found in HIV-1 Rev and other shuttling proteins (9, 18, 19, 21, 26, 31, 33, 37, 57, 61) and therefore is not thought to bind to hCRM-1. This finding was anticipated, since we and others had previously demonstrated that hTAP does not utilize the hCRM-1 export pathway (3, 39, 41, 43, 61). By analogy to known mechanisms of nuclear export, we speculate that hTAP may serve as an adapter between CTE and an unknown export receptor that interacts with its NES in the nucleus and tethers the resultant complex to the nuclear pore complex (NPC) for export. Our identification of the NES of hTAP provides a first lead to identify the components of this pathway.
Importantly, the function of NLS and NES is not dependent on the presence of the recently identified intact core RBD (residues 102 to 372) of hTAP (4) (Fig. 8). We concluded that the ability of hTAP to shuttle is not dependent on, or regulated by, the RNA binding status of the protein. Since coexpression of a CTE-containing mRNA did not affect hTAP’s localization in mammalian cells (data not shown), its role in trafficking was not further pursued. Braun et al. (4) demonstrated that, although TAP102-372 was able to bind and stimulate the export of CTE RNA, the presence of residues 61 to 101 further contributed to the binding of the core RBD to CTE RNA. Therefore, it is still possible that RNA binding could modulate the activity of NES and/or NLS in the context of the whole protein. The RBD includes hTAP’s leucine-rich repeat domain (LRR; residues 265 to 372 [47]). Such LRR domains in other proteins have been implicated in protein-protein interactions. Here, we showed that the presence of the LRR domain did not contribute to hTAP’s nucleocytoplasmic trafficking or nuclear rim association and thus appeared dispensable for these processes.
A signal(s) that anchors hTAP to the nuclear rim is found in a location that is distinct from the NES, NLS, and RBD and was shown to act independently of these signals. It is important to note that pMex67p, the homolog of hTAP in yeast cells, was also found to localize to the nuclear pores (47). hTAP’s anchoring region did not show similarities to known domains implicated in NPC targeting and may contain a novel signal. The anchoring of hTAP to the nuclear rim may reflect its interactions with the components of the NPC, as suggested for Mex67p (44, 47). Since these interactions occur independently of the NLS or NES, we excluded that the putative receptors of the NLS and NES of hTAP play a significant role in nuclear rim association. We found also that the C-terminal portion was necessary for hTAP function, as determined by activating CAT production in A6 cell line. Notably, a recombinant hTAP protein lacking this region was able to bind to the CTE RNA element and to promote the export of the RNA upon microinjection into Xenopus oocyte nuclei (4). This finding demonstrated that at least in that system the C-terminal portion of hTAP region was not essential for function. Taken together, in addition to an overall similarity, including the LRR regions and the ability to bind to RNA (4, 22, 47), hTAP and Mex67p share a similar subnuclear localization (reference 47 and the present study). Since we were unable to detect homologies in the region encompassing the NLS and the NES, the trafficking of these two proteins is likely controlled through distinct mechanisms. This observation is further supported by the identification of a Rev-like NES in Mex67p, which would also predict that hTAP and Mex67p are subjected to different export mechanisms.
In summary, our studies revealed that hTAP has distinct molecular determinants promoting its subcellular localization (Fig. 8). These signals (the NLS, the NES, and the nuclear retention/rim anchoring signals) contribute independently to hTAP’s nucleocytoplasmic trafficking. These determinants do not share homology with binding sites of known transport factors. The identification of the cellular factors mediating hTAP’s trafficking will shed light onto these novel mechanisms.
ACKNOWLEDGMENTS
We thank E. Izaurralde for sharing unpublished data, reagents, and discussions. We are grateful to E. Afonina, K. Horie, A. Gragerov, G. N. Pavlakis, and T. Hope for reagents and discussions, to G. Gragerova for technical assistance, and to C. Rhoderick for secretarial assistance.
This research sponsored by the National Cancer Institute, Department of Health and Human Services, under a contract with ABL.
J.B., W.T., and A.S.Z. contributed equally to this work.
ADDENDUM IN PROOF
In agreement with our data, a recent paper by Katahira et al. (EMBO J. 18:2593–2609, 1999) also describes that hTAP associates with the nuclear rim, where they find TAP to interact with nucleoporins.
REFERENCES
- 1.Afonina E, Stauber R, Pavlakis G N. The human poly(A) binding protein 1 shuttles between the nucleus and the cytoplasm. J Biol Chem. 1998;273:13015–13021. doi: 10.1074/jbc.273.21.13015. [DOI] [PubMed] [Google Scholar]
- 2.Askjaer P, Jensen T H, Nilsson J, Englmeier L, Kjems J. The specificity of the CRM1-Rev nuclear export signal interaction is mediated by RanGTP. J Biol Chem. 1998;273:33414–33422. doi: 10.1074/jbc.273.50.33414. [DOI] [PubMed] [Google Scholar]
- 3.Bogerd H P, Echarri A, Ross T M, Cullen B R. Inhibition of human immunodeficiency virus Rev and human T-cell leukemia virus Rex function, but not Mason-Pfizer monkey virus constitutive transport element activity, by a mutant human nucleoporin targeted to CRM1. J Virol. 1998;72:8627–8635. doi: 10.1128/jvi.72.11.8627-8635.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Braun I, Rohrbach E, Schmitt C, Izaurralde E. TAP binds to the constitutive transport element through a novel RNA-binding motif that is sufficient to promote CTE-dependent RNA export from the nucleus. EMBO J. 1999;18:1953–1965. doi: 10.1093/emboj/18.7.1953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bray M, Prasad S, Dubay J W, Hunter E, Jeang K-T, Rekosh D, Hammarskjold M-L. A small element from the Mason-Pfizer monkey virus genome makes human immunodeficiency virus type 1 expression and replication Rev-independent. Proc Natl Acad Sci USA. 1994;91:1256–1260. doi: 10.1073/pnas.91.4.1256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cullen B. Retroviruses as model systems for the study of nuclear RNA export pathways. Virology. 1998;248:203–210. doi: 10.1006/viro.1998.9331. [DOI] [PubMed] [Google Scholar]
- 7.Dayton A I, Terwilliger E F, Potz J, Kowalski M, Sodroski J G, Haseltine W A. cis-Acting sequences responsive to the rev gene product of the human immunodeficiency virus. J Acquired Immune Defic Syndr. 1988;1:441–452. [PubMed] [Google Scholar]
- 8.Dingwall C, Laskey R. Nuclear targeting sequences—a consensus? Trends Biochem Sci. 1991;16:478–481. doi: 10.1016/0968-0004(91)90184-w. [DOI] [PubMed] [Google Scholar]
- 9.Dobbelstein M, Roth J, Kimberly W T, Levine A J, Shenk T. Nuclear export of the E1B 55-kDa and E4 34-kDa adenoviral oncoproteins mediated by a rev-like signal sequence. EMBO J. 1997;16:4276–4284. doi: 10.1093/emboj/16.14.4276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Drysdale C M, Pavlakis G N. Rapid activation and subsequent downregulation of the human immunodeficiency virus type 1 promoter in the presence of Tat: possible mechanisms contributing to latency. J Virol. 1991;65:3044–3051. doi: 10.1128/jvi.65.6.3044-3051.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ernst R K, Bray M, Rekosh D, Hammarskjold M-L. Secondary structure and mutational analysis of the Mason-Pfizer monkey virus RNA constitutive transport element. RNA. 1997;3:210–222. [PMC free article] [PubMed] [Google Scholar]
- 12.Ernst R K, Bray M, Rekosh D, Hammarskjold M-L. A structured retroviral RNA element that mediates nucleocytoplasmic export of intron-containing RNA. Mol Cell Biol. 1997;17:135–144. doi: 10.1128/mcb.17.1.135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Fan X C, Steitz J A. HNS, a nuclear-cytoplasmic shuttling sequence in HuR. Proc Natl Acad Sci USA. 1998;95:15293–15298. doi: 10.1073/pnas.95.26.15293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Felber B K. Viral regulatory structures and their degeneracies. XXVIII. Reading, Mass: Addison-Wesley Publishing Co., Inc.; 1998. Posttranscriptional control: a general and important regulatory feature of HIV-1 and other retroviruses; pp. 101–122. [Google Scholar]
- 15.Felber B K, Hadzopoulou-Cladaras M, Cladaras C, Copeland T, Pavlakis G N. rev protein of human immunodeficiency virus type 1 affects the stability and transport of the viral mRNA. Proc Natl Acad Sci USA. 1989;86:1495–1499. doi: 10.1073/pnas.86.5.1495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fischer U, Huber J, Boelens W C, Mattaj I W, Luhrmann R. The HIV-1 Rev activation domain is a nuclear export signal that accesses an export pathway used by specific cellular RNAs. Cell. 1995;82:475–483. doi: 10.1016/0092-8674(95)90436-0. [DOI] [PubMed] [Google Scholar]
- 17.Fornerod M, Ohno M, Yoshida M, Mattaj I W. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell. 1997;90:1051–1060. doi: 10.1016/s0092-8674(00)80371-2. [DOI] [PubMed] [Google Scholar]
- 18.Fridell R A, Fischer U, Luhrmann R, Meyer B E, Meinkoth J L, Malim M H, Cullen B R. Amphibian transcription factor IIIA proteins contain a sequence element functionally equivalent to the nuclear export signal of human immunodeficiency virus type 1 rev. Proc Natl Acad Sci USA. 1996;93:2936–2940. doi: 10.1073/pnas.93.7.2936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fridell R A, Partin K M, Carpenter S, Cullen B R. Identification of the activation domain of equine infectious anemia virus rev. J Virol. 1993;67:7317–7323. doi: 10.1128/jvi.67.12.7317-7323.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fukuda M, Asano S, Nakamura T, Adachi M, Yoshida M, Yanagida M, Nishida E. CRM1 is responsible for intracellular transport mediated by the nuclear export signal. Nature. 1997;390:308–311. doi: 10.1038/36894. [DOI] [PubMed] [Google Scholar]
- 21.Fukuda M, Gotoh I, Gotoh Y, Nishida E. Cytoplasmic localization of mitogen-activated protein kinase kinase directed by its NH2-terminal, leucine-rich short amino acid sequence, which acts as a nuclear export signal. J Biol Chem. 1996;271:20024–20028. doi: 10.1074/jbc.271.33.20024. [DOI] [PubMed] [Google Scholar]
- 22.Grüter P, Tabernero C, von Kobbe C, Schmitt C, Saavedra C, Bachi A, Wilm M, Felber B K, Izaurralde E. TAP, the human homolog of Mex67p, mediates CTE-dependent RNA export from the nucleus. Mol Cell. 1998;1:649–659. doi: 10.1016/s1097-2765(00)80065-9. [DOI] [PubMed] [Google Scholar]
- 23.Hadzopoulou-Cladaras M, Felber B K, Cladaras C, Athanassopoulos A, Tse A, Pavlakis G N. The rev (trs/art) protein of human immunodeficiency virus type 1 affects viral mRNA and protein expression via a cis-acting sequence in the env region. J Virol. 1989;63:1265–1274. doi: 10.1128/jvi.63.3.1265-1274.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hammarskjöld M L, Heimer J, Hammarskjöld B, Sangwan I, Albert L, Rekosh D. Regulation of human immunodeficiency virus env expression by the rev gene product. J Virol. 1989;63:1959–1966. doi: 10.1128/jvi.63.5.1959-1966.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Henderson B R, Percipalle P. Interactions between HIV Rev and nuclear import and export factors: the Rev nuclear localisation signal mediates specific binding to human importin-beta. J Mol Biol. 1997;274:693–707. doi: 10.1006/jmbi.1997.1420. [DOI] [PubMed] [Google Scholar]
- 26.Hope T J, Bond B L, McDonald D, Klein N P, Parslow T G. Effector domains of human immunodeficiency virus type 1 Rev and human T-cell leukemia virus type 1 Rex are functionally interchangeable and share an essential peptide motif. J Virol. 1991;65:6001–6007. doi: 10.1128/jvi.65.11.6001-6007.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hope T J, Huang X, McDonald D, Parslow T G. Steroid-receptor fusion of the human immunodeficiency virus type 1 Rev transactivator: mapping cryptic functions of the arginine-rich motif. Proc Natl Acad Sci USA. 1990;87:7787–7791. doi: 10.1073/pnas.87.19.7787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27a.Horie, K., and G. N. Pavlakis. Unpublished data.
- 28.Izaurralde E, Adam S. Transport of macromolecules between the nucleus and the cytoplasm. RNA. 1998;4:351–364. [PMC free article] [PubMed] [Google Scholar]
- 29.Kudo N, Wolff B, Sekimoto T, Schreiner E P, Yoneda Y, Yanagida M, Horinouchi S, Yoshida M. Leptomycin B inhibition of signal-mediated nuclear export by direct binding to CRM1. Exp Cell Res. 1998;242:540–547. doi: 10.1006/excr.1998.4136. [DOI] [PubMed] [Google Scholar]
- 30.Malim M H, Hauber J, Le S-Y, Maizel J V, Cullen B R. The HIV-1 rev trans-activator acts through a structured target sequence to activate nuclear export of unspliced viral mRNA. Nature. 1989;338:254–257. doi: 10.1038/338254a0. [DOI] [PubMed] [Google Scholar]
- 31.Malim M H, McCarn D F, Tiley L S, Cullen B R. Mutational definition of the human immunodeficiency virus type 1 Rev activation domain. J Virol. 1991;65:4248–4254. doi: 10.1128/jvi.65.8.4248-4254.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McAfee J G, Shahied-Milam L, Soltaninassab S R, LeStourgeon W M. A major determinant of hnRNP C protein binding to RNA is a novel bZIP-like RNA binding domain. RNA. 1996;2:1139–1152. [PMC free article] [PubMed] [Google Scholar]
- 33.Mermer B, Felber B K, Campbell M, Pavlakis G N. Identification of trans-dominant HIV-1 Rev protein mutants by direct transfer of bacterially produced proteins into human cells. Nucleic Acids Res. 1990;18:2037–2044. doi: 10.1093/nar/18.8.2037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Meyer B E, Malim M H. The HIV-1 Rev trans-activator shuttles between the nucleus and the cytoplasm. Genes Dev. 1994;8:1538–1547. doi: 10.1101/gad.8.13.1538. [DOI] [PubMed] [Google Scholar]
- 35.Michael W M, Choi M, Dreyfuss G. A nuclear export signal in hnRNP A1: a signal-mediated, temperature-dependent nuclear protein export pathway. Cell. 1995;83:415–422. doi: 10.1016/0092-8674(95)90119-1. [DOI] [PubMed] [Google Scholar]
- 36.Michael W M, Eder P S, Dreyfuss G. The K nuclear shuttling domain: a novel signal for nuclear import and nuclear export in the hnRNP K protein. EMBO J. 1997;16:3587–3598. doi: 10.1093/emboj/16.12.3587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Murphy R, Wente S R. An RNA-export mediator with an essential nuclear export signal. Nature. 1996;383:357–360. doi: 10.1038/383357a0. [DOI] [PubMed] [Google Scholar]
- 38.Nakielny S, Dreyfuss G. Nuclear export of proteins and RNAs. Curr Opin Cell Biol. 1997;9:420–429. doi: 10.1016/s0955-0674(97)80016-6. [DOI] [PubMed] [Google Scholar]
- 39.Otero G C, Harris M E, Donello J E, Hope T J. Leptomycin B inhibits equine infectious anemia virus Rev and feline immunodeficiency virus rev function but not the function of the hepatitis B virus posttranscriptional regulatory element. J Virol. 1998;72:7593–7597. doi: 10.1128/jvi.72.9.7593-7597.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Palmeri D, Malim M H. Importin beta can mediate the nuclear import of an arginine-rich nuclear localization signal in the absence of importin alpha. Mol Cell Biol. 1999;19:1218–1225. doi: 10.1128/mcb.19.2.1218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pasquinelli A E, Ernst R K, Lund E, Grimm C, Zapp M L, Rekosh D, Hammarskjöld M-L, Dahlberg J E. The constitutive transport element (CTE) of Mason-Pfizer monkey virus (MPMV) accesses an RNA export pathway utilized by cellular messenger RNAs. EMBO J. 1997;16:7500–7510. doi: 10.1093/emboj/16.24.7500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Peng S S, Chen C Y, Xu N, Shyu A B. RNA stabilization by the AU-rich element binding protein, HuR, an ELAV protein. EMBO J. 1998;17:3461–3470. doi: 10.1093/emboj/17.12.3461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Saavedra C, Felber B, Izaurralde E. The simian retrovirus-1 constitutive transport element, unlike the HIV-1 RRE, uses factors required for cellular mRNA export. Curr Biol. 1997;7:619–628. doi: 10.1016/s0960-9822(06)00288-0. [DOI] [PubMed] [Google Scholar]
- 43a.Sanger Centre. 1997–1999, copyright date. [Online.] http://genomic.sanger.ac.uk/gf/gf.shtml. [12 July 1999, last date accessed.]
- 44.Santos-Rosa H, Moreno H, Simos G, Segref A, Fahrenkrog B, Pante N, Hurt E. Nuclear mRNA export requires complex formation between Mex67p and Mtr2p at the nuclear pores. Mol Cell Biol. 1998;18:6826–6838. doi: 10.1128/mcb.18.11.6826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Schneider R, Campbell M, Nasioulas G, Felber B K, Pavlakis G N. Inactivation of the human immunodeficiency virus type 1 inhibitory elements allows Rev-independent expression of Gag and Gag/Protease and particle formation. J Virol. 1997;71:4892–4903. doi: 10.1128/jvi.71.7.4892-4903.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Schwartz S, Felber B K, Benko D M, Fenyö E M, Pavlakis G N. Cloning and functional analysis of multiply spliced mRNA species of human immunodeficiency virus type 1. J Virol. 1990;64:2519–2529. doi: 10.1128/jvi.64.6.2519-2529.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Segref A, Sharma K, Doye V, Hellwig A, Huber J, Luhrmann R, Hurt E. Mex67p, a novel factor for nuclear mRNA export, binds to both poly(A)+ RNA and nuclear pores. EMBO J. 1997;16:3256–3271. doi: 10.1093/emboj/16.11.3256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shahied-Milam L, Soltaninassab S R, Iyer G V, LeStourgeon W M. The heterogeneous nuclear ribonucleoprotein C protein tetramer binds U1, U2, and U6 snRNAs through its high affinity RNA binding domain (the bZIP-like motif) J Biol Chem. 1998;273:21359–21367. doi: 10.1074/jbc.273.33.21359. [DOI] [PubMed] [Google Scholar]
- 49.Siomi H, Dreyfuss G. A nuclear localization domain in the hnRNP A1 protein. J Cell Biol. 1995;129:551–560. doi: 10.1083/jcb.129.3.551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Stade K, Ford C S, Guthrie C, Weis K. Exportin 1 (Crm1p) is an essential nuclear export factor. Cell. 1997;90:1041–1050. doi: 10.1016/s0092-8674(00)80370-0. [DOI] [PubMed] [Google Scholar]
- 51.Stauber R, Gaitanaris A S, Pavlakis G N. Analysis of trafficking of Rev and transdominant Rev proteins in living cells using green fluorescent protein fusions: transdominant Rev blocks the export of Rev from the nucleus to the cytoplasm. Virology. 1995;213:439–454. doi: 10.1006/viro.1995.0016. [DOI] [PubMed] [Google Scholar]
- 52.Stauber R, Pavlakis G. Intracellular trafficking and interactions of the HIV-1 Tat protein. Virology. 1998;252:126–136. doi: 10.1006/viro.1998.9400. [DOI] [PubMed] [Google Scholar]
- 53.Stauber R H, Horie K, Carney P, Hudson E A, Tarasova N I, Gaitanaris G A, Pavlakis G N. Development and applications of enhanced green fluorescent protein mutants. BioTechniques. 1998;24:462–471. doi: 10.2144/98243rr01. [DOI] [PubMed] [Google Scholar]
- 54.Tabernero C, Zolotukhin A S, Bear J, Schneider R, Karsenty G, Felber B K. Identification of an RNA Sequence Within an intracisternal-A particle element able to replace Rev-mediated posttranscriptional regulation of human immunodeficiency virus type 1. J Virol. 1997;71:95–101. doi: 10.1128/jvi.71.1.95-101.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tabernero C, Zolotukhin A S, Valentin A, Pavlakis G N, Felber B K. The posttranscriptional control element of the simian retrovirus type 1 forms an extensive RNA secondary structure necessary for its function. J Virol. 1996;70:5998–6011. doi: 10.1128/jvi.70.9.5998-6011.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Truant R, Cullen B R. The arginine-rich domains present in human immunodeficiency virus type 1 tat and rev function as direct importin beta-dependent nuclear localization signals. Mol Cell Biol. 1999;19:1210–1217. doi: 10.1128/mcb.19.2.1210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Weichselbraun I, Farrington G, Rusche J, Bohnlein E, Hauber J. Definition of the human immunodeficiency virus type 1 Rev and human T-cell leukemia virus type I Rex protein activation domain by functional exchange. J Virol. 1992;66:2583–2587. doi: 10.1128/jvi.66.4.2583-2587.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Weighardt F, Biamonti G, Riva S. Nucleo-cytoplasmic distribution of human hnRNP proteins: a search for the targeting domains in hnRNP A1. J Cell Sci. 1995;108:545–555. doi: 10.1242/jcs.108.2.545. [DOI] [PubMed] [Google Scholar]
- 59.Wolff B, Sanglier J-J, Wang Y. Leptomycin B is an inhibitor of nuclear export: inhibition of nucleo-cytoplasmic translocation of the human immunodeficiency virus type 1 (HIV-1) Rev protein and Rev-dependent mRNA. Chem Biol. 1997;4:139–147. doi: 10.1016/s1074-5521(97)90257-x. [DOI] [PubMed] [Google Scholar]
- 60.Yoon D W, Lee H, Seol W, DeMaria M, Rosenzweig M, Jung J U. Tap: a novel cellular protein that interacts with tip of herpesvirus saimiri and induces lymphocyte aggregation. Immunity. 1997;6:571–582. doi: 10.1016/s1074-7613(00)80345-3. [DOI] [PubMed] [Google Scholar]
- 61.Zolotukhin A S, Felber B K. Mutations in the nuclear export signal of human Ran-binding protein RanBP1 block the Rev-mediated posttranscriptional regulation of human immunodeficiency virus type 1. J Biol Chem. 1997;272:11356–11360. doi: 10.1074/jbc.272.17.11356. [DOI] [PubMed] [Google Scholar]
- 62.Zolotukhin A S, Felber B K. Nucleoporins Nup98 and Nup214 participate in nuclear export of human immunodeficiency virus type 1 Rev. J Virol. 1999;73:120–127. doi: 10.1128/jvi.73.1.120-127.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zolotukhin A S, Valentin A, Pavlakis G N, Felber B K. Continuous propagation of RRE(−) and Rev(−)RRE(−) human immunodeficiency virus type 1 molecular clones containing a cis-acting element of simian retrovirus type 1 in human peripheral blood lymphocytes. J Virol. 1994;68:7944–7952. doi: 10.1128/jvi.68.12.7944-7952.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]