Abstract
Several species of unicellular eukaryotic algae exhibit relatively simple genomic and cellular architecture. Laboratory cultures of these algae grow faster than plants and often provide homogeneous cellular populations exposed to an almost equal environment. These characteristics are ideal for conducting experiments at the cellular and subcellular levels. Many microalgal lineages have recently become genetically tractable, which have started to evoke new streams of studies. Among such algae, the unicellular red alga Cyanidioschyzon merolae is the simplest organism; it possesses the minimum number of membranous organelles, only 4,775 protein-coding genes in the nucleus, and its cell cycle progression can be highly synchronized with the diel cycle. These properties facilitate diverse omics analyses of cellular proliferation and structural analyses of the intracellular relationship among organelles. C. merolae cells lack a rigid cell wall and are thus relatively easily disrupted, facilitating biochemical analyses. Multiple chromosomal loci can be edited by highly efficient homologous recombination. The procedures for the inducible/repressive expression of a transgene or an endogenous gene in the nucleus and for chloroplast genome modification have also been developed. Here, we summarize the features and experimental techniques of C. merolae and provide examples of studies using this alga. From these studies, it is clear that C. merolae—either alone or in comparative and combinatory studies with other photosynthetic organisms—can provide significant insights into the biology of photosynthetic eukaryotes.
Keywords: Cyanidiales, Cyanidioschyzon merolae, Photosynthetic eukaryotes, Red algae
Introduction
Algae capture light energy and produce organic compounds that are introduced into aquatic and terrestrial ecosystems. Algae include cyanobacteria and many different eukaryotic lineages (Fig. 1A) and exhibit diversity in metabolism and habitats (Graham and Wilcox 2000). Aside from their ecological and evolutionary importance, algae, especially unicellular ones, are a good alternative to plants for use as a model system for basic and applied studies for the following reasons. (i) In the laboratory, a unicellular algal culture often provides a homogeneous population of a single cell type; by contrast, many types of cells and tissues differentiate in plants. (ii) In an algal culture, each cell is exposed to a relatively homogeneous environment (light, inorganic nutrient concentrations, etc.); by contrast, each plant cell is exposed to different environments, even in the same individual. (iii) Some unicellular algal species in several different lineages possess relatively simple cellular and genome architecture. (iv) Many of them grow faster than plants. (v) Several of them can be cultured in a medium in which all of the chemical components are known. (vi) Some of them have been cultured on a large scale to produce feed, pigments and nutraceuticals, and some others are now being developed to obtain alternative energy (Khan et al. 2018). Furthermore, rapid progress has led to the very recent development of experimental tools, especially those for the genetic modification of eukaryotic algae, which until now had been limited.
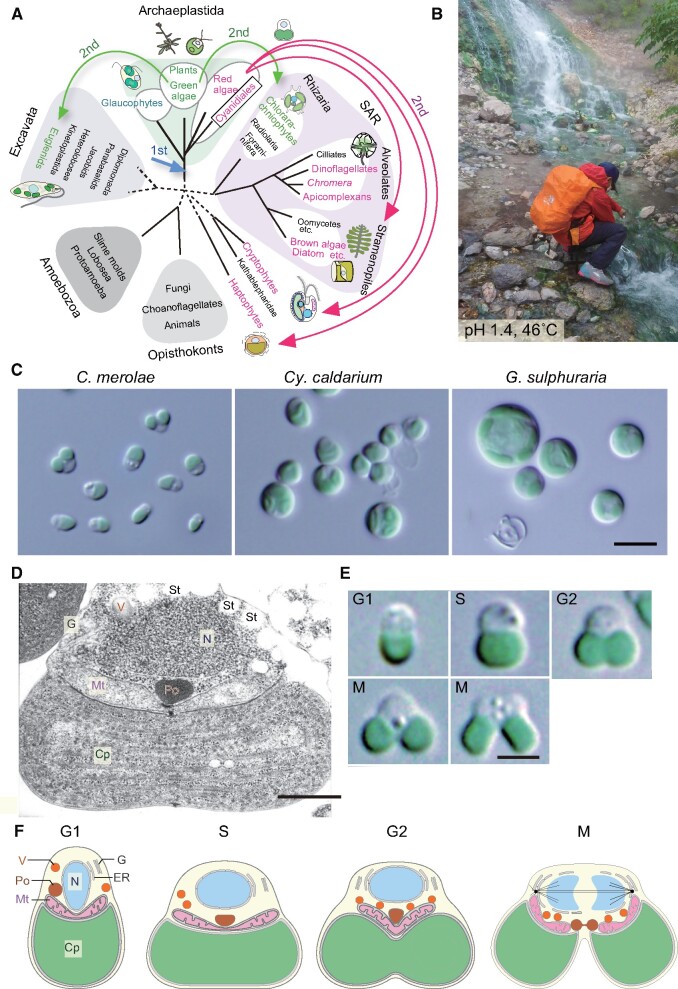
Fig. 1.
Phylogenetic position, habitat and classification of Cyanidiales and the cell cycle-dependent change in the cellular architecture of C. merolae. (A) Position of red algae and Cyanidiales in the eukaryotic tree. The broken lines denote the uncertainty of branch positions in the tree. Red algae and groups possessing chloroplasts (or nonphotosynthetic plastids) of red algal origin are shown in red. Viridiplantae (green algae and land plants) and groups possessing chloroplasts of green algal origin are shown in green. Arrows indicate the primary endosymbiotic event of a cyanobacterium (first) and secondary endosymbiotic events (second). The position and number of red algal secondary endosymbiotic events in the tree remain uncertain at present. This tree is according to Adl et al. (2012). (B) A sulfuric hot spring where cyanidialean red algae dominate (Kusatsu, Japan). (C) Interference-contrast microscopic images of C. merolae 10D, C. caldarium isolated in the hot spring shown in (B), and G. sulphuraria 074 G. Bar = 5 µm. (D) A transmission electron micrograph of C. merolae at the S phase. Bar = 500 nm. (E) Interference-contrast microscopic images of C. merolae showing the cell cycle progression. Bar = 2 µm. The images were reproduced from Miyagishima et al. (2012). (F) Schematic illustrations of C. merolae cells showing organelle inheritance. Cp, chloroplast; G, Golgi apparatus; Mt, mitochondrion; N, nucleus; Po, peroxisome; St, floridean starch granule; V, vacuole.
Evolution of eukaryotic algae and unicellular algal models
Oxygen-producing photosynthesis first occurred in cyanobacteria about 3 billion years ago and introduced into eukaryotes >1 billion years ago by establishing the chloroplast (plastid) in an ancient eukaryotic cell through cyanobacterial endosymbiosis (Fig. 1A;Elias and Archibald 2009). This ancestral eukaryotic alga evolved to glaucophytes, red algae, green algae and plants, all of which are grouped as Archaeplastida (Fig. 1A). After green and red algae had been established, chloroplasts were further spread into several other eukaryotic lineages independently through secondary and tertiary endosymbiotic events in which a eukaryotic alga was integrated into a previously heterotrophic eukaryote (Fig. 1A). The chloroplasts of red algae were distributed to stramenopiles, dinoflagellates, apicomplexans, haptophytes and cryptophytes, whereas those of green algae descended into euglenophytes and chlorarachniophytes (Fig. 1A;Elias and Archibald 2009). In addition, some species of dinoflagellates further replaced their original chloroplasts of red algal origin with those of green algae, cryptophytes, haptophytes or diatoms (Elias and Archibald 2009).
Among several eukaryotic algae, the unicellular green alga Chlamydomonas reinhardtii has been the most extensively studied because sexual reproduction in this alga has facilitated genetic studies—especially those on the cilium and photosynthesis, which are absent in yeast. As a result, genetic modification procedures were established in this alga >30 years ago (Harris et al. 2009, Tran and Kaldenhoff 2020). Recently, procedures for efficient genetic modification have also been developed in some other unicellular eukaryotic algae, such as the red alga C. merolae (described below), the green alga Ostreococcus tauri (Corellou et al. 2009, Sanchez et al. 2019), the diatom Phaeodactylum tricornutum (Butler et al. 2020) and the eustigmatophytes Nannochloropsis spp. (Poliner et al. 2018), which have promoted their uses as alternative model organisms, such as for evolutionary studies, metabolism and cell biology research (Table 1).
Table 1.
Comparison of nuclear genomes of C. merolae and other photosynthetic and nonphotosynthetic eukaryotes in which genetic modification has been widely applied for studies
| Species | C. merolae | C. reinhardtii | O. tauri | A. thaliana | P. tricornutum | Nannochloropsis oceanica | S. cerevisiae | Homo sapiens |
|---|---|---|---|---|---|---|---|---|
| Group | Rhodophyta | Chlorophyta | Chlorophyta | Streptophyta | Stramenopiles | Stramenopiles | Fungi | Animalia |
| Genome size (bp) | 16.5 M | ∼120 M | 12.6 M | ∼120 M | 27.4 Ma | 36.5 Mb | 12.5M | ∼2,900 M |
| G+C content (%) | 55 | 64 | 58 | 36 | 48.8a | 54b | 36 | 41 |
| Protein gene number | 4,775 | ∼15,000 | 8,166 | ∼25,500 | ∼10,400a | ∼11,000b | ∼6,100c | ∼21,000 |
| Genes with an intron (%) | 0.5 | 92 | 39 | 79 | 0.8a | 66b | 5.0 | 85 |
| Copy number of rRNA units (18S/5.8S/28S) | 3 | ND | 4 | 700–800 | ∼4d | ND | 200–400 | ∼400 |
| TEs in the genome (%) | 0.7 | ∼9 | ∼10 | ∼15 | 5.8a | >0.7b | 2.4 | >45% |
ND, not determined. TE, Transposable element.
Bowler et al. (2008).
Vieler et al. (2012); https://phycocosm.jgi.doe.gov/Nanoce1779.
Lin et al. (2013).
Gong and Marchetti (2019).
Other data are from Matsuzaki et al. (2004), Merchant et al. (2007) and Derelle et al. (2006).
Among them, C. merolae 10D (available from the Microbial Culture Collection at the National Institute of Environmental Studies in Japan; NIES-3377), which was isolated from a mixture of algae in sulfuric hot spring water in Italy, exhibits a very simple cellular architecture that comprises one mitochondrion, one chloroplast, and a minimal number of other membranous organelles (Fig. 1). These organelles divide in a cell cycle-dependent manner (Fig. 1F), and the cell cycle progression of cells in a culture can be highly synchronized to the diel cycle (Suzuki et al. 1994). These features have facilitated studies on the cell cycle and organelle inheritance. The complete organelle (Ohta et al. 1998, Ohta et al. 2003) and nuclear (Matsuzaki et al. 2004, Nozaki et al. 2007) genomes were sequenced for the first time in eukaryotic algae, and the nuclear genome content is very simple with very little genetic redundancy (16.5 Mbp; 4,775 protein-coding genes; Table 1). Hence, the essential features of a photosynthetic eukaryote are accommodated in the compact genome and simple cellular architecture of C. merolae. Because C. merolae cells lack a rigid cell wall (Merola et al. 1981), it is relatively easy to extract or isolate cellular contents. These characteristics are suitable for biochemical and several omics analyses. In addition, molecular genetic tools, such as gene targeting by highly efficient homologous recombination and inducible/repressive gene expression systems, have been recently developed (Fig. 2; described later). Based on these characteristics and techniques for genetic modification, C. merolae has started to be utilized as a model photosynthetic eukaryote for studies in several fields, such as cell biology, metabolism, structural and functional studies of the photosynthetic apparatus, epigenetics and comparative genomics with other related species.
Fig. 2.
Examples of genetic modification in C. merolae. (A) A schematic diagram of the targeted integration of a transgene and a URA selectable marker into a chromosomal locus and the subsequent removal of the URA marker (Takemura et al. 2018, Takemura et al. 2019). (B) Photos of transformed colonies 2 weeks after the transformation and liquid culture (10 ml) of a clone 2 weeks after the inoculation of a colony. For efficient colony formation, cells were spread on cornstarch beds (white circles) on a gel plate. (C) An inducible/repressive expression system using the NR promoter (Fujiwara et al. 2015). Although the expression of a transgene (GFP) is regulated by the NR promoter, this system is also applicable for an endogenous gene by integrating the NR promoter to the upstream of a gene of interest in the chromosome (Fujiwara et al. 2020). Red is the autofluorescence of the chloroplast, and green is the fluorescence of GFP in the cytosol. Bar = 5 µm.
Habitats, classification and taxonomic position of Cyanidiales, which comprise C. merolae, Cyanidium spp. and Galdieria spp.
Cyanidialean unicellular red algae form a monophyletic group estimated to have branched off from other red algae quite early in eukaryotic evolution (1.3–1.4 billion years ago; Yoon et al. 2004, Yoon et al. 2006). Similar to other red algae, Cyanidiales lack centrosomes and flagella. Furthermore, while their chloroplasts also contain phycocyanin and chlorophyll a as photosynthetic pigments (Graham and Wilcox 2000), they do, however, lack phycoerythrin (a red protein–pigment complex; Lin et al. 1990); thus, Cyanidiales exhibit a blue-green color (Fig. 1B, C). Also, while sexual reproduction has not been reported in Cyanidiales or in other unicellular red algae, by contrast multicellular red algae undergo sexual reproduction (Graham and Wilcox 2000).
Cyanidiales are the only photosynthetic organisms that dominate in sulfuric hot springs in geothermal areas globally (pH 0.05–5; 35–56°C), where sulfur reacts with oxygen and water producing sulfuric acid (Fig. 1B;Merola et al. 1981, Albertano et al. 2000, Pinto 2007). Despite such an extremely acidic environment, the intracellular pH of these algae is maintained in the pH range of 6.3–7.1 (Zenvirth et al. 1985, Enami et al. 2010). As observed in other acidophilic microbes, plasma membrane H+-ATPase is highly expressed in C. merolae, which likely pumps protons from the cytosol to the outside of the cell to avoid cytosolic acidification (Hirooka et al. 2017).
Cyanidiales have been classified into three genera, namely, Cyanidioschyzon, Cyanidium and Galdieria, based on the presence or absence of a cell wall, mode of cellular proliferation and molecular phylogenetic analyses (Fig. 1C;Merola et al. 1981, Albertano et al. 2000). Cyanidiococcus was newly described based on a morphological and molecular phylogenetic study (Liu et al. 2020). Cyanidioschyzon merolae does not possess a cell wall and proliferates by binary fission (Merola et al. 1981, Albertano et al. 2000). Cyanidium and Cyanidiococcus possess cell walls and proliferate by forming four daughter cells (autospores) in a mother cell by two successive cell divisions before hatching out of the mother cell (Fig. 1C;Merola et al. 1981, Albertano et al. 2000, Liu et al. 2020). Galdieria are also cell walled and the largest of the Cyanidiales and proliferate by forming 4–32 daughter cells in a mother cell by successive cell divisions (Fig. 1C;Merola et al. 1981, Albertano et al. 2000). Among Cyanidiales, only Galdieria can grow heterotrophically as well as photoautotrophically and can assimilate >50 different organic carbon sources by contrast to (ecologically) obligate photoautotrophic C. merolae and Cyanidium (and Cyanidiococcus) (Rigano et al. 1977, Gross 1999; Oesterhelt et al. 1999; also see below for the artificial heterotrophic cultivation of C. merolae in the laboratory).
Cellular Architecture and Proliferation Mode of C. merolae
The C. merolae cell is quite small (1.5–3 µm in diameter) compared to most other eukaryotic algae (Fig. 1C). As in other red algae, the photosynthetic products are stored as floridean starch granules in the cytosol of C. merolae (Fig. 1D) while, in green algae and plants, they are stored as starch granules in the chloroplasts (plastids; Hirabaru et al. 2010).
The C. merolae cell comprises a single nucleus, mitochondrion, chloroplast, Golgi apparatus, peroxisome, approximately four vacuoles and a single ER layer that extends from the outer nuclear membrane (Fig. 1C, D). The nuclear membrane and the single layer of endoplasmic reticulum (ER) dissolve partially in metaphase (semi-open mitosis; Fig. 1F;Fujiwara et al. 2013), while all other membranous organelles are also replicated in a cell cycle-dependent manner (Fig. 1E, F), which is the reason why cells possess a fixed number of mitochondrion, chloroplast and some other organelles. The chloroplast (Miyagishima et al. 2003), mitochondrion (Nishida et al. 2003) and peroxisome (Miyagishima et al. 1999; Imoto et al. 2013) divide in this order during S and M phases (Fig. 1E, F). The Golgi apparatus (Yagisawa et al. 2013) and small vacuoles (Fujiwara et al. 2010) are also somehow replicated before mitosis. At metaphase, a replicated Golgi apparatus and the lateral edge of a daughter mitochondrion are positioned near a spindle pole body (Fig. 1F;Yagisawa et al. 2013). Lysosomes attach to the mitochondria during the M phase and segregate to daughter cells with daughter mitochondria (Fig. 1F;Fujiwara et al. 2010). The chloroplast and peroxisome are attached to the mitochondrion (Fig. 1F;Miyagishima et al. 1999). Hence, these membranous organelles are apparently segregated with the daughter chromosomes directly or indirectly by spindle microtubules during mitosis. During cytokinesis, an actin-based contractile ring is not formed, unlike in its relative Cyanidium caldarium. A recent study has shown that, as in animal cells and archaea, ESCRT proteins are involved in C. merolae cytokinesis (Yagisawa et al. 2020).
The small cell size and minimal number of organelles make it easier to observe the entire structure of respective organelles and spatial relationships among organelles in a whole cell via conventional fluorescence microscopy. These characteristics have also facilitated the ultrastructural characterization of whole cells by a relatively small number of thin serial sections via transmission electron microscopy (Miyagishima et al. 1999) and recently via focused ion beam scanning microscopy (Ichinose and Iwane 2017).
C. merolae proliferates relatively faster than other unicellular algae; the doubling time is ∼9 h at optimal conditions, in an inorganic liquid medium (pH 1–2.5) under continuous light (Minoda et al. 2004). Some studies used time-lapse to observe the course of cell and organelle division within one round of the cell cycle (Sumiya et al. 2016), and a very recent study succeeded in the observation of single cells for 3 d in which cells divided three times on average (Ichinose and Iwane 2021). As in several other algae, cell cycle progression is synchronized with the diel cycle (Suzuki et al. 1994; Miyagishima et al. 2014). In the laboratory, C. merolae is often subjected to synchronous culture under the 12-h light/12-h dark (LD) cycle, in which cells grow and stay at the G1 phase during the light period and then, during the dark period, undergo S and M phases synchronously (Suzuki et al. 1994; Miyagishima et al. 2014). The G1–S transition is regulated by circadian rhythms and inhibited during (subjective) daytime (Miyagishima et al. 2014). Therefore, the timing of the G1–S transition (the period from the onset of the light period) changes little even when the length of the light period is changed (e.g. 8 h light/16 h dark and 16 h light/8 h dark; Miyagishima et al. 2014). However, the mitotic index varies depending on the cellular growth rate by photosynthesis during the light period (thus the length of the light period and light intensity) because only cells that have grown over a certain threshold enter the S phase early at (subjective) night and other cells stay at the G1 phase throughout the round of LD and wait for the G1–S transition until the next (subjective) night (Miyagishima et al. 2014). If the doubling time is <24 h under LD, some cells undergo two rounds of S/M phases during the dark period (Miyagishima et al. 2014). In appropriate conditions, almost 100% of cells undergo one round of cell division by binary fission. This is in contrast to some other algae, such as Cyanidium, Galdieria and the green alga C. reinhardtii, which undergo multiple fission by repeating two or more rounds of S/M phases in a relatively short time, thus making it difficult to characterize the progress of the cell cycle. In addition, cell cycle progression can be arrested at G1, S and M phases by the addition of certain chemical inhibitors to the culture (Nishida et al. 2007; Miyagishima et al. 2012) or by a conditional knockdown of cell cycle-regulating genes via genetic modification (Fujiwara et al. 2020). Hence, C. merolae is a powerful tool for investigating cell cycle, organelle division and the relationship between these and other cellular activities.
The C. merolae Genome
The C. merolae nuclear genome was the first fully sequenced eukaryotic algal genome (Matsuzaki et al. 2004) and the first fully complete eukaryotic genome assembled without gaps (Table 1; Nozaki et al. 2007). The genome is 16.5 Mbp in size and distributed amongst 20 chromosomes. The centromere (Kanesaki et al. 2015) and telomere (Nozaki et al. 2007) sequences of the respective chromosomes were also identified. The genome contents with relevant mRNA data have been summarized in the C. merolae genome project database (http://czon.jp/; last accessed on 14 April 2021.) (Matsuzaki et al. 2004).
In the nuclear genome, the number of protein-coding genes is 4,775, and the total number of genes, including RNA genes, is 5,331. This number is comparable with those of yeasts Saccharomyces cerevisiae (5,770) and Schizosaccharomyces pombe (4,929), although the C. merolae nuclear genome additionally encodes genes for photosynthesis, biogenesis and regulation of the chloroplast (Table 1). Recent comparative genomic studies have suggested that this compact genome of C. merolae resulted from substantial reductive evolution (Qiu et al. 2013). Similarly, recent studies have shown that several green algal lineages also underwent genome minimization independently, yielding nuclear genomes of 12–25 Mbp. However, the number of genes encoded in these minimized green algal genomes ranges from 6,340 to 11,000 (Foflonker et al. 2018, Lemieux et al. 2019). Thus, C. merolae possesses the smallest number of genes in photosynthetic eukaryotes thus far examined.
Genomes of free-living eukaryotes usually contain >100 copies of a tandemly repeated array of ribosomal RNA (rRNA) genes. By contrast, the C. merolae nuclear genome encodes only three copies of 18S, 5.8S and 28S rRNA units and 5S rRNA, respectively, in separate loci (Table 1; Matsuzaki et al. 2004). These small numbers of rRNA copies are likely consistent with the observation that the C. merolae nucleus contains only one small nucleolus (Matsuzaki et al. 2004). The percentage of transposable elements in the nuclear genome is also very low (0.7%) compared with other free-living eukaryotes (Table 1). The genome contains eight transposons and 26 retrotransposons, of which only three transcribe intact reverse transcriptases (Nozaki et al. 2007).
Of the total of 4,775 protein-coding nuclear genes, only 26 genes (0.5%) contain introns in which 25 genes possess a single intron and one gene possesses two (Table 1; Matsuzaki et al. 2004). Therefore, the construction of transgenes for genetic modification in most cases can be performed using the genomic DNA instead of cDNA. In addition, because the G + C content of the nuclear genome is 55% (Matsuzaki et al. 2004), PCR amplification of genomic and complementary DNA and reverse transcription of mRNA work efficiently, unlike in some green algae with nuclear genomes of higher G + C contents, such as C. reinhardtii (64%; Table 1; Merchant et al. 2007). In addition, the simple genome facilitates the resequencing of the genome of, e.g. mutant lines, chromatin immunoprecipitation sequencing, transcriptome or other related omics analyses based on high-throughput sequencing at a lower cost.
The C. merolae mitochondrial genome is 32.2 kb and encodes 35 proteins and 28 RNA genes (Ohta et al. 1998), which is the largest number in red algae thus far examined (Yang et al. 2015). Chloroplast genomes in red algae are the most gene rich in photosynthetic eukaryotes, possessing 223–250 genes (Janouskovec et al. 2013). The number of genes encoded in C. merolae is 234, of which 208 encode proteins (Ohta et al. 2003). Although the gene number is comparable with that in other red algae, the C. merolae chloroplast genome is highly compacted and the smallest in the red algae (150 kb). The compaction is mainly due to the short length of intergenic regions, which is only 10 bp on average (Ohta et al. 2003), and smaller than those in other red algae (60–85 bp; Janouskovec et al. 2013).
Transcriptome analyses in C. merolae performed in the studies introduced below have generated datasets showing transcriptomic changes under several environmental conditions. In addition, the C. merolae genome dataset has been effectively used as representative information of a basal red alga in evolutionary studies. For example, a study using C. merolae contributed to the elucidation of the evolutionary history of laforin, a carbohydrate-binding phosphatase, which is mutated in patients with Lafora disease (Gentry et al. 2007).
After the genome project of C. merolae 10D, draft genome sequences of Galdieria sulphuraria 074 W (Schönknecht et al. 2013) and Galdieria phlegrea (Qiu et al. 2013) were published. Recently, with the development of high-throughput sequencing technologies, genome sequences of several other strains of Cyanidiales started to be published (Rossoni et al. 2019). These genomic data also have facilitated comparative genomic studies, especially those about speciation and adaptation to different environments in which respective strains reside.
Techniques for Genetic Manipulation in C. merolae
Since the generation and isolation of a uracil auxotrophic mutant of C. merolae (Minoda et al. 2004), many techniques for genetic modifications, such as gene knockout and knock-in, and inducible/repressive expression of a transgene, have been developed in C. merolae (Fig. 2). By introducing an exogenous linear DNA using polyethylene glycol, transient transformants can be prepared and analyzed (Ohnuma et al. 2008). From that population, in several cases, stable transformants have been selected and cultured for further analyses (Fig. 2; Imamura et al. 2009, Kobayashi et al. 2011). In addition to the original system for the selection of stable transformants using the URA gene as a selectable marker and the uracil auxotrophic mutant as a host (Fig. 2A), another system using a chloramphenicol-resistant gene as a marker and the wild-type strain as a host was later developed (Fujiwara et al. 2017, Zienkiewicz et al. 2017a). More recently, a selectable marker recycling system was developed to edit multiple loci (Fig. 2A;Takemura et al. 2018, Takemura et al. 2019). After several improvements to date, colonies of a transformant on a plate medium can usually be obtained within 2 weeks (Fig. 2). After checking the insertion of a transgene into a targeted locus, a transformant liquid culture is usually obtained after an additional 2 weeks (Fig. 2B). As necessary, transformants can be stored as frozen stock (Ohnuma et al. 2011). The notable features of genetic manipulation in C. merolae are (i) a transgene(s) with a selectable marker gene (for the expression of transgenes) or only a selection marker (for gene disruption) is integrated into a desired chromosomal locus by highly efficient homologous recombination (Fig. 2); (ii) transgenes are stably expressed without any silencing activity so far tested; and (iii) besides the above-mentioned genetic modification of the nuclear genome, a procedure for the manipulation of the chloroplast genome was also reported recently (Zienkiewicz et al. 2017b).
Transformant selection systems using uracil auxotrophic mutants
The C. merolae URA (URA5.3) gene (named according to its relevant yeast genes URA5 and URA3) encodes a fusion protein of N-terminal orotidine-5′-phosphoribosyl transferase (corresponding to yeast Ura5p) and C-terminal orotidine 5′-monophosphate (OMP) decarboxylase (corresponding to yeast Ura3p). As a host strain for genetic manipulation, C. merolae M4, which is a derivative of C. merolae 10D and contains a point mutation (one nucleotide insertion) in the midportion of the URA gene, was originally generated (Minoda et al. 2004). The spontaneous mutant line M4 was isolated as a uracil auxotrophic and 5-fluoroorotic acid (5-FOA)-resistant clone. 5-FOA itself is not toxic to the cells while it is converted to cytotoxic 5-fluorouracil by OMP decarboxylase. Therefore, URA mutants, but not the wild type, are resistant to 5-FOA. Recently, another uracil auxotrophic mutant (T1), in which the URA locus was completely deleted via genetic manipulation, was established and is being used as a host for genetic manipulation (Taki et al. 2015).
Regarding the URA marker for selecting transformants, an artificial URACm-Gs marker was used initially so that URACm-Gs, which was integrated into the genome, could be easily distinguished from the endogenous URA gene with the point mutation (Imamura et al. 2009, Kobayashi et al. 2011). In the URACm-Gs, the C-terminal OMP decarboxylase domain of C. merolae was replaced with that of G. sulphuraria. However, it later turned out that the selection of transformants by the URACm-Gs marker resulted in the multicopy insertion of the marker and transgene at high frequencies accompanied by recombination events at the targeted locus. The number of copies varies among transformed clones (Fujiwara et al. 2013, Fujiwara et al. 2019). By contrast, the selection of transformants by the URA marker (C. merolae wild-type gene) resulted in the efficient single-copy insertion of the marker and transgene (Fujiwara et al. 2013, Fujiwara et al. 2019). Although the reason for the multiple insertions of URACm-Gs into the genome is unclear at this point, the activity of URACm-Gs is likely lower than that of normal URA; hence, only clones in which URACm-Gs and a transgene are multiplicated can grow and form colonies on a uracil-depleted medium. The multiple integrations of the transgene by URACm-Gs result in a higher expression of transgene than single-copy insertion by URA and thus useful for the overexpression of transgenes (Fujiwara et al. 2013, Fujiwara et al. 2019).
Transformant selection systems using chloramphenicol
Aside from transformant selection using uracil auxotrophic mutants, an antibiotic-based system for transformant selection has also been developed. In this case, the host can be the wild type or its derivatives, and the bacterial chloramphenicol acetyltransferase (CAT) gene is used as a selectable marker (Fujiwara et al. 2017, Zienkiewicz et al. 2017a).
Selectable marker recycling system
Although additional selectable markers other than CAT and URA will undoubtedly be developed in the future, the same markers can at present be used in a stepwise manner on their own or in combination to enable the manipulation of multiple chromosomal loci. A procedure that was very recently developed involves recycling of the URA marker (Fig. 2A; Takemura et al. 2018, Takemura et al. 2019). In this system, the promoter and orf of the URA marker (with or without transgene for knock-in or knockout, respectively) are sandwiched between a directly repeated URA terminator (Fig. 2A). After selecting the transformant with the URA marker, the marker can be eliminated through homologous recombination between the two copies of the URA terminator sequence and 5-FOA selection (Fig. 2A). After removing the URA marker, another construct with the URA marker can subsequently be integrated into any chromosomal locus. Therefore, using this system, any number of chromosomal loci can be edited in a stepwise manner.
Inducible and repressive expression systems
The simple nature of the cellular architecture and genome content of C. merolae should facilitate the characterization of intracellular phenomena, which is essential and common to photosynthetic eukaryotes. However, an inactivation or modification of such essential mechanisms by gene manipulation often leads to cell death. Thus, conditionally inducible/repressive expression systems of a transgene or endogenous gene are desired in such cases. Until now, two kinds of techniques have been developed to conditionally induce transgene expression or repress endogenous gene expression (Sumiya et al. 2014, Fujiwara et al. 2015).
The inducible system uses the promoter of the C. merolae gene encoding a small heat shock protein (Hsp20). The promoter is inactive at a lower temperature (≤40°C) but highly activated at a higher temperature (≥46°C; Sumiya et al. 2014). This system has been used to overexpress normal or dominant-negative forms of proteins upon heat shock treatment (Sumiya et al. 2016, Yagisawa et al. 2020). The inducible/repressive system uses a promoter of nitrate assimilation genes, such as those encoding nitrate reductase, nitrite reductase and nitrate/nitrite transporter (Fujiwara et al. 2015). These promoters are active in ammonium ( )-depleted medium (for cellular growth, the medium is supplemented with nitrate as an inorganic nitrogen source) but inactivated by the addition of
)-depleted medium (for cellular growth, the medium is supplemented with nitrate as an inorganic nitrogen source) but inactivated by the addition of  into the medium (Fig. 2C). For example, one of these promoters is integrated into the upstream of a gene of interest in the nitrate medium. When
into the medium (Fig. 2C). For example, one of these promoters is integrated into the upstream of a gene of interest in the nitrate medium. When  is added to the transformant culture, the expression of the gene is depressed. When cells are transferred to the fresh nitrate medium, the gene becomes expressed again. This system is used to conditionally inactivate cyclin-dependent kinase to arrest cells at the G1 phase (Fujiwara et al. 2020).
is added to the transformant culture, the expression of the gene is depressed. When cells are transferred to the fresh nitrate medium, the gene becomes expressed again. This system is used to conditionally inactivate cyclin-dependent kinase to arrest cells at the G1 phase (Fujiwara et al. 2020).
Genetic manipulation of the chloroplast genome
All of the above are procedures for the genetic manipulation of the nuclear genome. However, a recent study has shown that the chloroplast genome can also be edited by the chloramphenicol and CAT system (Zienkiewicz et al. 2017b). In this case, the selectable marker comprises a chloroplast gene promoter and CAT gene, which was codon-optimized to the chloroplast genome. The marker is integrated into the chloroplast genome by homologous recombination, and the CAT protein is expressed in the chloroplast conferring cellular chloramphenicol resistance. Although the application of this procedure has not yet been extended to mutate/modify chloroplast genes or express transgenes, it should nonetheless further facilitate studies in C. merolae.
Notable Biochemical Features of C. merolae
A notable feature of C. merolae is that its proteins are relatively thermostable compared with those in other eukaryotes given its acidothermophile nature. In fact, many proteins of C. merolae were chosen for the three-dimensional ultrastructural determination of eukaryotic proteins (Markley et al. 2009). Until now, the ultrastructures of several proteins, such as a dynamin-related protein (DRP3), involved in mitochondrial and peroxisomal division and described below (Bohuszewicz and Low 2018), and protein complexes of the photosynthetic apparatus (also described below) have been characterized in C. merolae.
In addition, evidence to date suggests that metabolic pathways are also likely very simple in C. merolae. Although the contents of C. merolae metabolites have not yet been extensively examined, recent studies have shown that, relative to other eukaryotes, the lipid profile in C. merolae is quite simple and appears to lack phosphatidylserine and cardiolipin (Mori et al. 2016, Sato et al. 2017, Mori et al. 2019). Collectively, the simple characteristics of C. merolae render it suitable for many diverse studies, as exemplified in the section below.
Examples of Studies Using C. merolae
C. merolae has long been used in studies on the division and inheritance of mitochondria and chloroplasts and the relationship between the states of these endosymbiotic organelles and the host cell cycle progression (Fig. 3). Many of these studies have focused on how the activities of the eukaryotic host cell and those of endosymbiotic organelles are coordinated. Recent studies have also studied responses to environmental changes and the photosynthetic apparatus in this alga. In addition, researchers in several other fields have started to use this alga. Here, we introduce some examples of studies using C. merolae.
Fig. 3.

Mechanism of cell cycle progression and its relationship with the chloroplast DNA replication and chloroplast division in C. merolae. (A) A diagram showing the structure of chloroplast division machinery in C. merolae. For reference, the mitochondrial division machinery in C. merolae is also shown. DRP5B, the outer and inner PD rings and two types of FtsZ proteins also participate in chloroplast (plastid) division in plants. DRP3 and MDA1 (called Mdv1p in yeasts) also participate in mitochondrial division in many other eukaryotic lineages. In some lineages of eukaryotes, including C. merolae, the outer and inner MD rings and two types of FtsZ proteins also participate in mitochondrial division. (B) A schematic diagram of the interactive synchronization of the cell cycle progression and the chloroplast division in C. merolae. The host cell restricts the onset of chloroplast division to the S phase by the S phase-specific expression of nucleus-encoded chloroplast division proteins. The formation and constriction of the chloroplast division ring lift the prophase arrest so that the host cell enters into the metaphase only when chloroplast division progresses. (C) A schematic diagram of the G1–S transition, which is regulated by the retrograde signaling from the chloroplast to the host cell cycle and circadian rhythms. The chloroplast DNA replication somehow leads to the accumulation of Mg-Proto in the cell, which binds Fbox3. This binding inhibits the ubiquitination and degradation of G1 cyclin (CYC1) by Fbox3, leading to the accumulation of G1 cyclin-bound CDKA. During the G1 phase, RB-related protein (RBR) inhibits the transcriptional activity of E2F-DP. E2F is phosphorylated in a circadian rhythm-dependent manner, and E2F phosphorylation peaks during the evening. E2F phosphorylation permits CDKA-CYC1 to phosphorylates RBR. RBR phosphorylation inactivates RBR, thereby enabling E2F-DP to transcribe S-phase genes.
Molecular mechanism of mitochondrial and chloroplast division and their coordination with the host cell cycle
Mitochondria and chloroplasts, double-membraned energy-converting organelles in eukaryotic cells, arose in this order by serial endosymbiotic events (Elias and Archibald 2009). To maintain permanently sustainable endosymbiotic associations, several regulatory systems evolved for constraining both the endosymbionts and the host cells (Rodriguez-Ezpeleta and Philippe 2006). One major and obvious requirement is the coordinated proliferation of the host and endosymbiotic organelles. It is believed that this coordinated proliferation was established originally by the synchronization of endosymbiotic cell division with host cell division, as seen in the relationship between algal cells and their chloroplasts (Rodriguez-Ezpeleta and Philippe 2006). The majority of algae have just one or, at most, only a few chloroplasts per cell, and it is obvious that a direct and precise relationship exists between the host cell and chloroplast division (Miyagishima et al. 2012). That is, chloroplasts must divide once per cell cycle before the host cell completes cytokinesis. Studies using C. merolae and other model organisms have explained how the chloroplast and mitochondrion divide coordinately with the host cell.
As do their bacterial ancestors, mitochondria and chloroplasts proliferate by growth and division of preexisting ones (Kuroiwa et al. 1998). In chloroplasts of plants and algae, early electron microscopic studies identified a ring-like structure called the plastid-dividing (PD) ring surrounding the division site of chloroplasts. The PD ring comprises the inner (at the stromal side of the inner envelope) and the outer (at the cytosolic side of the outer envelope) structures (Kuroiwa et al. 1998). The structure and behavior of these PD rings were examined most extensively in C. merolae (Miyagishima et al. 2011). At the time, a homolog of the bacterial division gene, ftsZ, was also identified in the nuclear genome of plants and algae (Osteryoung and Vierling 1995). FtsZ is a self-assembling GTPase protein. In bacteria, it forms a ring structure beneath the cell membrane at the cell division site, By contrast, the algal and plant FtsZ homologs of cyanobacterial endosymbiotic origin, which are nuclear-encoded, form a ring structure at the stromal side of the chloroplast division site, where it is involved in its division (Chen et al. 2018). Furthermore, in C. merolae the FtsZ ring forms at the stromal side of the inner PD ring, but the composition of the inner ring itself still remains unknown (Miyagishima et al. 2011).
Further studies using C. merolae and Arabidopsis thaliana plants have shown the following (Miyagishima et al. 2011, Chen et al. 2018; Fig. 3A): (i) Unlike cyanobacteria, which possess a single type of FtsZ, in plants and algae, the gene that descended from cyanobacteria was duplicated and evolved into two types of FtsZ (cpFtsZA and cpFtsZB in C. merolae) that play different roles in chloroplast division. (ii) Dynamin-related protein 5B (DRP5B; also known as ARC5) of eukaryotic host origin, a member of the dynamin family of self-assembling GTPase, localizes at the cytosolic side of the chloroplast division site where it is involved in the division process. Eukaryotes possess several members of DRPs, which are involved in membrane fission and fusion. Each member is involved in endocytosis, mitochondrial/peroxisomal division, mitochondrial fusion, chloroplast division and cytokinesis. For example, both A. thaliana and C. reinhardtii genomes encode 16 DRPs. By contrast, the C. merolae genome encodes only two DRPs (DRP5B and DRP3); DRP3 is involved in mitochondrial/peroxisomal division (Nishida et al. 2003, Imoto et al. 2013), and DRP5B is involved in chloroplast division (Miyagishima et al. 2003). (iii) By isolating and characterizing the outer PD ring of C. merolae, it was shown that the ring is a bundle of glucan filaments synthesized by the PDR1 protein of eukaryotic host origin (Yoshida et al. 2010).
In addition to the chloroplast-targeted FtsZ, some eukaryotic lineages, including C. merolae, possess a nucleus-encoded mitochondrial-targeted FtsZ that descended from an alphaproteobacterial endosymbiont, the progenitor of mitochondria. However, this mitochondrial FtsZ was secondarily lost independently in several of the eukaryotic lineages, such as green algae, plants, fungi and animals (Kiefel et al. 2006). In C. merolae mitochondrial division, FtsZ, mitochondrial-dividing (MD) and DRP3 rings form in this order (Nishida et al. 2003; Fig. 3A); this is similar to chloroplast division in which FtsZ, PD and DRP5 rings form in a subsequential manner (Miyagishima et al. 2003). Like chloroplast FtsZ, the C. merolae mitochondrial FtsZ was duplicated, resulting in mtFtsZA and mtFtsZB (Miyagishima et al. 2011). The MD ring in C. merolae is also a glucan filament synthesized by MDR1, a mitochondrial paralog of PDR1 (Yoshida et al. 2017; Fig. 3A). Therefore, it is suggested that the division machinery of alphaproteobacterial and cyanobacterial endosymbionts was modified similarly during the establishment of mitochondrion and chloroplast, although these modifications occurred independently several billion years apart (Miyagishima et al. 2011).
The above understanding of the molecular mechanism of chloroplast division further led to the elucidation of the mechanism in algae that synchronizes chloroplast division with the cell cycle. Namely, chloroplast division is restricted to the S phase in algae (possessing both primary and secondary endosymbiotic origins) by the cell cycle-regulated expression of nucleus-encoded chloroplast division genes (Fig. 3B;Miyagishima et al. 2012). In C. merolae, when the chloroplast division was hampered before the division site constriction by conditional overexpression of FtsZ or a dominant-negative form of DRP5B, cell cycle progression was arrested at prophase (Sumiya et al. 2016). Hence, chloroplast division restricts host cell cycle progression so that the cell cycle progresses to the metaphase only when chloroplast division has commenced (Fig. 3B). In addition, the cell cycle arrest upon the blockage of chloroplast division was also observed in the glaucophyte alga Cyanophora paradoxa, raising the possibility that the chloroplast division checkpoint was established early in algal evolution (Sumiya et al. 2016). These results collectively suggested that chloroplast division and host cell cycle progression are synchronized by an interactive restriction between the nucleus and the chloroplast (Fig. 3B).
Coordination of chloroplast and mitochondrial DNA replication with the host cell cycle
As with mitochondrial and chloroplast division, genome replication in these organelles should also be coordinated with host cell cycle progression (Kuroiwa et al. 1998). In C. merolae, DNA replication in both organelles [organelle DNA replication (ODR)] occurs before nuclear DNA replication (NDR), and the inhibition of ODR led to the blockage of NDR (Fig. 3C;Kobayashi et al. 2009). Further analyses have shown that chloroplast DNA replication results in the accumulation of Mg-protoporphyrin IX (Mg-Proto; an intermediate metabolite in the chlorophyll biosynthetic pathway) in the cell, which binds an F-box protein (Fbox3). This binding inhibits the ubiquitination and degradation of G1 cyclin by Fbox3, which leads to the accumulation of G1 cyclin and subsequent G1–S transition and NDR by circadian gating (Fig. 3C;Kobayashi et al. 2009, Kobayashi et al. 2011). The circadian gating of the G1–S transition was shown to be regulated by the retinoblastoma (RB)–E2F–DP complex (Fig. 3C;Miyagishima et al. 2014). According to the studies in other eukaryotes, such as animal and plant cells and unicellular algae, the G1 cyclin–CDK complex most likely phosphorylates/inactivates RB, which, in turn, leads to the transcription of S-phase genes by the E2F-DP heterodimeric transcription factor (Fig. 3C;Miyagishima et al. 2014).
Retrograde signaling from the chloroplast to the nucleus
In eukaryotes, it has been suggested that signals derived from organelles are transmitted to the nucleus to modify the expression of nuclear genes depending on the states of the organelles. Such signals from organelles to the nucleus are called retrograde signals (or plastid signals in the case of plastids) in contrast to anterograde signals from the nucleus to organelles (Chan et al. 2016). Mg-Proto produced in the chloroplast has been suggested to act as a retrograde signal to modulate the expression of certain nuclear genes in Arabidopsis. Also, in C. merolae, Mg-Proto acts as a retrograde signal to coordinate the DNA replication of organelles and nucleus as introduced above, although this example does not constitute transcriptional regulation (Fig. 3C;Kobayashi et al. 2009, Kobayashi et al. 2011).
Abscisic acid (ABA) is a well-known plant hormone involved in abiotic stress responses and dormancy (Vishwakarma et al. 2017). C. merolae was also found to produce ABA upon high salt stress (Kobayashi et al. 2016). In C. merolae, ABA, produced in the chloroplast, induces nuclear gene expression for heme biosynthesis and tryptophan-rich sensory protein-related protein O (TSPO). In addition, heme was shown to be required for the G1–S transition. These results suggested that cell cycle arrest at the G1 phase upon heme quenching by TSPO prevents stress-labile cell cycle progression (Kobayashi et al. 2016). Because heme is produced only in the mitochondrion in C. merolae, the chloroplast-derived ABA and the mitochondrion-derived heme likely constitute a signaling network to control the G1–S transition, which is another example of the retrograde signaling system in C. merolae (Kobayashi et al. 2016). In vascular plants, ABA plays a role in intercellular signaling. By contrast, ABA is unstable in a sulfur acidic medium (Vishwakarma et al. 2017) and thus unlikely to act as an intercellular hormone but rather an intracellular signaling molecule in C. merolae.
Relationship between diel changes in metabolic activities and cell cycle progression
The growth of photosynthetic eukaryotes depends on photosynthesis. Thus, their metabolism and several other cellular activities are greatly affected by the diel cycle. In photosynthetic organisms, most metabolic genes exhibited changes with a periodicity of 24 h at the mRNA level according to the diel cycle. In addition, cell division in eukaryotic algae is restricted to a certain time range within a day; in many cases, S and M phases progress at evening and night (Johnson 2010). Thus, correlations have been observed between diel metabolic changes and cell cycle progression. However, it is unclear whether transcriptomic changes in metabolic pathways are regulated by the cell cycle or whether metabolic pathways and cell cycle progression are independently synchronized to the diel cycle. It is also poorly understood what kind of advantage is conferred by restricting cell division at night.
To address these issues using C. merolae, diel transcriptomic changes were compared between wild type and two transgenic lines in which the cell cycle was uncoupled from the diel cycle by the conditional depletion of cyclin-dependent kinase A (G1 arrest) or knockout of the RB-related gene (G1–S transition occurs anytime; Fujiwara et al. 2020). The results showed that the periodic expression patterns of most metabolic genes were not affected in the transgenic lines, indicating that cell cycle and transcriptomic changes in most metabolic pathways are independently synchronized to the diel cycle (Fujiwara et al. 2020).
This independence of diel rhythms in most metabolic pathways from cell cycle progression is a likely scenario under natural habitats. As described above, the timing of the G1–S transition is restricted at night, but only cells that have grown beyond a certain threshold size enter the S phase, and other cells stay at the G1 phase during the night (Miyagishima et al. 2014). In contrast to laboratory culture conditions with nutrient-rich medium and a regular illumination intensity, under natural conditions, low levels of inorganic nutrients are present and the light strength changes depending on the weather. Therefore, cellular growth by photosynthesis during the daytime is relatively slow, and only some cells undergo cell division during the night, whereas all cells are subjected to diel rhythms of photosynthesis and growth regardless of the G1–S transition.
In C. merolae cultured under LD, the levels of photosynthetic and respiratory activities peaked in the morning and then decreased toward the evening and at night, whereas the pathways for the anaerobic consumption of pyruvate, produced by glycolysis, were upregulated during the evening and at night (Miyagishima et al. 2019), as also reported for the green alga C. reinhardtii (Strenkert et al. 2019). Also, the day/night rhythm of respiration depends on the photosynthetic activity. In addition, when cells were forced to enter the S phase while performing photosynthesis, the frequency of nuclear DNA double-strand breaks (DSB) was elevated (Miyagishima et al. 2019). These results suggested that the temporal separation of oxygenic energy metabolism (photosynthesis and respiration), which causes oxidative stress, from NDR reduces the risk of DSB during cell proliferation. In other words, this sequestration enables the safe proliferation of cells and allows the coexistence of the chloroplast/mitochondrion and the eukaryotic host.
Responses to environmental changes
Responses to various environmental cues are important for survival in natural habitats, and some of these responses have been analyzed at the gene expression level in C. merolae. Representative examples are responses to nutritional starvations.  is a favorable inorganic nitrogen source in C. merolae, and the depletion of
is a favorable inorganic nitrogen source in C. merolae, and the depletion of  or the substitution of
or the substitution of  with
with  induces the transcription of a set of nitrogen-responsive genes, including nitrate reductase and nitrite reductase (Imamura et al. 2009, Imamura et al. 2010). Based on transcriptome analyses of cells in the nitrogen-depleted medium, it was shown that one of the R2R3-type MYB family of transcription factors, MYB1, is responsible for inducing nitrogen-responsive genes (Imamura et al. 2009). The target of rapamycin (TOR) kinase, which is highly conserved in eukaryotes and known to act as an amino acid and nutrient sensor, is also involved in response to nitrogen depletion in C. merolae (Imamura et al. 2015, Pancha et al. 2020). However, the mechanism that senses the cellular nitrate status remains mostly elusive.
induces the transcription of a set of nitrogen-responsive genes, including nitrate reductase and nitrite reductase (Imamura et al. 2009, Imamura et al. 2010). Based on transcriptome analyses of cells in the nitrogen-depleted medium, it was shown that one of the R2R3-type MYB family of transcription factors, MYB1, is responsible for inducing nitrogen-responsive genes (Imamura et al. 2009). The target of rapamycin (TOR) kinase, which is highly conserved in eukaryotes and known to act as an amino acid and nutrient sensor, is also involved in response to nitrogen depletion in C. merolae (Imamura et al. 2015, Pancha et al. 2020). However, the mechanism that senses the cellular nitrate status remains mostly elusive.
In response to decreases in CO2 concentration, sets of nuclear genes, including those involved in CO2 assimilation and photorespiration, are known to be induced (Rademacher et al. 2017); however, no information is presently available for the underlying mechanism.
Because C. merolae is a photoautotrophic organism, light is one of the most important environmental cues for survival. In plants, photoreceptors, such as phytochrome, cryptochrome and phototropin, which respond to specific light wavelengths, have been identified, and their downstream responses have been well elucidated (Jing and Lin 2020). Among known plant photoreceptors, C. merolae possesses only cryptochrome, and the blue light-specific responses presumably initiated by cryptochrome were identified (Tardu et al. 2016). Other than such photoreceptor-mediated responses, photosynthetic activity in plant chloroplasts modulates gene expression through various signal mediators, such as redox state and reactive oxygen species. Such mechanisms also likely work in C. merolae because red light also greatly affects the transcription of many nuclear genes despite the apparent lack of red light-specific receptors (Tardu et al. 2016). In addition, another study in C. merolae showed that the transcription of the chloroplast-encoded Rubisco genes (rbcL-rbcS-cbbX operon) is activated by the chloroplast-encoded transcription factor Ycf30 upon illumination, in a nucleus-independent manner (Minoda et al. 2010).
Ycf30 is one of the four transcription factors (others are Ycf27,28,29) coded by the chloroplast genomes of red algae, which descended from a cyanobacterial ancestor (Tanaka 2012). Ycf28 and Ycf30 correspond to cyanobacterial NtcA and RbcR, which are involved in responses to nitrogen and CO2 depletion in cyanobacteria, respectively (Tanaka 2012). Ycf27 and Ycf29 are the counterparts of cyanobacterial response regulators, which constitute two-component systems with histidine kinases that sense specific environmental stimuli (Puthiyaveetil and Allen 2009). Although the function of these chloroplast-encoded transcription factors—other than Ycf30—have not been reported, they are likely involved in autonomous responses of the chloroplast independent of nuclear control.
The chloroplast genome encodes most of the subunits of RNA polymerase, whereas the sigma subunits of the complex, required for the recognition of the promoters, are encoded by the nuclear genome, translated in the cytosol and targeted to the chloroplast (Tanaka et al. 1996). The array of sigma subunits and their differential specificities to subsets of chloroplast genes are suggestive of nuclear control of chloroplast gene expression in plants (Chi et al. 2015). C. merolae possess four types of chloroplast sigma subunits, namely, SIG1, SIG2, SIG3 and SIG4. Among them, SIG2 and SIG4 were shown to prefer promoters of phycobilisomal and high-light-responsive genes, respectively (Fujii et al. 2013, Fujii et al. 2015). The nucleus-regulated sigma subunits and the nucleus-independent chloroplast-encoded transcription factors likely coordinately regulate the chloroplast gene expression according to environmental changes. Chloroplast-encoded transcription factors are conserved among glaucophyte algae, red algae and organisms possessing the chloroplast of red algal secondary endosymbiotic origin. Further studies on these transcription factors in C. merolae will reveal how chloroplast gene expression is regulated by the chloroplast and host cell coordinately.
Other studies using C. merolae
Photosynthetic apparatus
Although the photosynthetic apparati in eukaryotic chloroplasts descended from a cyanobacterial ancestor, they are remodeled differently. Red algae and cyanobacteria possess phycobilisomes as light-harvesting antennas associated with photosystem II (PS II) unlike in green algae and plants, which lost them during evolution. However, the luminal side of PS II in red algae is structurally different from that in cyanobacteria (Ohta et al. 2003, Krupnik et al. 2013). Unlike cyanobacteria, red algae possess the light-harvesting complex associated with photosystem I, similar to green algae and plants (together grouped as Viridiplantae; Busch et al. 2010, Pi et al. 2018, Antoshvili et al. 2019). Thus, the red algal photosynthetic apparatus is believed to be an intermediate between that of cyanobacteria and Viridiplantae. In addition, the C. merolae PS II retains significant photosynthetic activity in an unusually broad range of pH, temperatures, and light intensities (Krupnik et al. 2013), probably reflecting the fact that C. merolae can survive in a broad range of environments. Recent structural and biochemical analyses in C. merolae have started to elucidate similarities and differences in the photosynthetic apparatus of red algae, cyanobacteria and Viridiplantae and how it is adapted to thermoacidophilic environments and copes with environmental changes (Krupnik et al. 2013, Nilsson et al. 2014, Nikolova et al. 2017, Pham et al. 2019).
Because C. merolae and Cyanidium spp. (and Cyanidiococcus) are (ecologically) obligate photoautotrophs, unlike Galdieria spp., which can grow both photoautotrophically and heterotrophically (Albertano et al. 2000), any mutation that abolishes photosynthesis leads to cell death in C. merolae. However, a recent study has shown that C. merolae could grow heterotrophically in the dark when the medium is supplemented with a very high concentration (≥200 mM) of glycerol (Moriyama et al. 2015). Another study has shown that a C. merolae strain, in which a Galdieria plasma membrane sugar transporter was expressed, can grow heterotrophically with glucose even when photosynthesis was inhibited in the dark or by the addition of 3-(3,4-dichlorophenyl)-1,1-dimethylurea in the light (Fujiwara et al. 2019). Hence, these procedures likely enable the generation and analyses of photosynthesis-deficient mutants.
Spliceosome
The spliceosome is a large ribonucleoprotein complex that catalyzes pre-mRNA splicing. In humans, it comprises five small nuclear RNAs and >200 proteins. Because of this complexity, much study has focused on the budding yeast spliceosome, which comprises fewer than half as many splicing factors as humans. However, it has still been difficult to elucidate the functions of respective factors or even identify which are critical for catalyzing the splicing reaction (Hudson et al. 2015). A recent study has shown that C. merolae has a highly reduced set of identifiable core splicing proteins (43 proteins) compared with about 90 proteins in budding yeast and about 140 proteins in humans, suggesting that C. merolae may be a more tractable system for studying the central processes of pre-mRNA splicing (Stark et al. 2015, Reimer et al. 2017).
Epigenetic regulation
In eukaryotic cells, the transcription state is dependent on the underlying chromatin state, which is altered by posttranslational modifications in histones (Batista and Kohler 2020). Among several histone modifications, the trimethylation of lysine 27 on histone H3 (H3K27me3) is associated with transcriptional silencing. The deposition of H3K27me3 is mediated by polycomb group protein complexes. The function of H3K27me3 and polycomb group proteins has been extensively studied in animals and plants, and they have been shown to control developmentally regulated processes in multicellular organisms (Batista and Kohler 2020). In contrast to these multicellular organisms, the function of H3K27me3 in unicellular eukaryotes has been poorly understood, although there have been some studies in diatoms (Veluchamy et al. 2015). This is largely because yeasts lack polycomb group proteins. In addition, monomethylation and dimethylation of lysine 27 on histone H3 were detected in the green alga C. reinhardtii, but trimethylation was not convincingly detected (Shaver et al. 2010). Thus, the function of H3K27me cannot be investigated in these unicellular eukaryotic models. A recent study has shown that C. merolae possesses a polycomb complex, PRC2, and that H3K27me3 is enriched in telomeric and subtelomeric regions and has unique preferential binding toward intein-containing genes involved in protein splicing (Mikulski et al. 2017). Further studies in C. merolae should provide evolutionary insights into chromatin repression.
Industrial application
C. merolae contains higher concentrations of proteins and vitamins K, C and E than other microalgae that have been used commercially (Supplementary Table S1). In addition, the phycocyanin of Cyanidiales is stable at lower pH and higher temperature compared with that of Spirulina (Arthrospira platensis), which has been used as a natural blue colorant in certain food products (Rahman et al. 2017). Thus, C. merolae phycocyanin is likely tolerant to pasteurization. The low pH at which C. merolae grows minimizes the microbial contamination risk, especially those feeding on microalgae. A recent study has shown that C. merolae could grow in a seawater-based acidic medium, which reduces the risk of contamination in nonsterile open cultures more efficiently than acidic conditions alone in freshwater-based medium (Hirooka et al. 2020). Due to C. merolae lacking a cell wall, cellular contents can be easily extracted hence raising the possibility that this organism can be used as a source of phycocyanin, proteins, feeds and functional food production.
Perspectives
Based on its simple cellular and genomic architecture, genome information availability and genetic modification techniques, C. merolae is becoming an attractive model for studies in several different disciplines, including synthetic biology, systems biology and experimental evolution.
Although several genetic modification techniques have been developed in C. merolae, additional tools are desired. The C. merolae genome does not encode RNA interference machinery components, such as Dicer (Matsuzaki et al. 2004). Because of this limitation, experiments based on gene knockdown have been performed by expressing antisense RNA from transiently introduced DNA (Ohnuma et al. 2009). However, the efficiency of DNA introduction into cells is relatively low. To overcome this issue, techniques, such as those combining CRISPR interference (Qi et al. 2013) and the above-introduced inducible expression systems in stable transformants, can be applied for conditional gene knockdown in C. merolae. Alternatively, a protein knockdown system (Yesbolatova et al. 2020) could be applied. Another desired experimental system to be developed is that for random mutagenesis and forward genetics. In this regard, the small genome size of C. merolae should facilitate the resequencing of mutant genomes.
Although cyanidialean red algae have been cultured often in suspension in the laboratory, in natural habitats, they adhere to the bottom of shallow hot pools and streams (Fig. 1B) and on rocks exposed to fumes above the water surface (Pinto 2007). In addition, some strains inhabit an endolithic environment (Pinto 2007). A previous study has shown that the hot spring water itself and light were not sufficient, and the addition of an inorganic nitrogen source was required for the growth of Cyanidiales (Hirooka and Miyagishima 2016). Because a metagenomic analysis has shown that Cyanidiales form a mat with other archaea and bacterial species (Walker et al. 2005), their growth likely requires nitrogen sources supplied by other microbes in their natural habitats. In addition, it was recently shown in laboratory cultures that C. merolae cells move over surfaces toward the light (Ohnuma et al. 2011, Maschmann et al. 2020). To apply C. merolae to ecological and environmental studies, further information would be required.
Although there is still scope for gaining more information of C. merolae and expanding the repertoire of experimental techniques and tools described above, studies focusing on this organism, as well as comparative and combinatory studies using C. merolae and other photosynthetic organisms, have thus far provided significant insights into the lives of photosynthetic eukaryotes.
Supplementary Data
Supplementary data are available at PCP online.
Supplementary Material
Acknowledgments
Our apologies to those whose work could not be cited because of space constraints. We would like to thank Drs. Takayuki Fujiwara and Shunsuke Hirooka for their critical reading of the manuscript.
Funding
The Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (20H00477 to S.-Y.M.); the Japan Science and Technology Agency (JST)-Mirai Program (to S.-Y.M.) and JST-ALCA Program (to K.T.); and the Nagase Science Technology Foundation (to K.T.).
Disclosures
The JST has filed patent applications related to Supplementary Table S1 and a reference (Hirooka et al. 2020) on behalf of S.-Y.M. The other author declared no conflict of interest.
Contributor Information
Shin-Ya Miyagishima, Department of Gene Function and Phenomics, National Institute of Genetics, Mishima, Shizuoka, 411-8540 Japan.
Kan Tanaka, Institute of Innovative Research, Tokyo Institute of Technology, Yokohama, Kanagawa, 226-8503 Japan.
References
- Adl S.M., Simpson A.G., Lane C.E., Lukes J., Bass D., Bowser S.S., et al. (2012) The revised classification of eukaryotes. J. Eukaryot. Microbiol. 59: 429–493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albertano P., Ciniglia C., Pinto G., Pollio A. (2000) The taxonomic position of Cyanidium, Cyanidioschyzon and Galdieria: an update. Hydrobiologia 433: 137–143. [Google Scholar]
- Antoshvili M., Caspy I., Hippler M., Nelson N. (2019) Structure and function of photosystem I in Cyanidioschyzon merolae. Photosynth. Res. 139: 499–508. [DOI] [PubMed] [Google Scholar]
- Batista R.A., Kohler C. (2020) Genomic imprinting in plants-revisiting existing models. Genes Dev. 34: 24–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohuszewicz O., Low H.H. (2018) Structure of a mitochondrial fission dynamin in the closed conformation. Nat. Struct. Mol. Biol. 25: 722–731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowler C., Allen A.E., Badger J.H., Grimwood J., Jabbari K., Kuo A., et al. (2008) The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456: 239–244. [DOI] [PubMed] [Google Scholar]
- Busch A., Nield J., Hippler M. (2010) The composition and structure of photosystem I-associated antenna from Cyanidioschyzon merolae. Plant J. 62: 886–897. [DOI] [PubMed] [Google Scholar]
- Butler T., Kapoore R.V., Vaidyanathan S. (2020) Phaeodactylum tricornutum: a diatom cell factory. Trends Biotechnol. 38: 606–622. [DOI] [PubMed] [Google Scholar]
- Chan K.X., Phua S.Y., Crisp P., McQuinn R., Pogson B.J. (2016) Learning the languages of the chloroplast: retrograde signaling and beyond. Annu. Rev. Plant Biol. 67: 25–53. [DOI] [PubMed] [Google Scholar]
- Chen C., MacCready J.S., Ducat D.C., Osteryoung K.W. (2018) The molecular machinery of chloroplast division. Plant Physiol. 176: 138–151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi W., He B., Mao J., Jiang J., Zhang L. (2015) Plastid sigma factors: their individual functions and regulation in transcription. Biochim. Biophys. Acta 1847: 770–778. [DOI] [PubMed] [Google Scholar]
- Corellou F., Schwartz C., Motta J.P., Djouani-Tahri el B., Sanchez F., Bouget F.Y. (2009) Clocks in the green lineage: comparative functional analysis of the circadian architecture of the picoeukaryote Ostreococcus. Plant Cell 21: 3436–3449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Derelle E., Ferraz C., Rombauts S., Rouze P., Worden A.Z., Robbens S., et al. (2006) Genome analysis of the smallest free-living eukaryote Ostreococcus tauri unveils many unique features. Proc. Natl. Acad. Sci. USA 103: 11647–11652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elias M., Archibald J.M. (2009) Sizing up the genomic footprint of endosymbiosis. Bioessays 31: 1273–1279. [DOI] [PubMed] [Google Scholar]
- Enami I., Adachi J., Shen J.-R. (2010) Mechanisms of acido-tolerance and characteristics of photosystems in an acidophilic and thermophilic red alga, Cyanidium caldarium. In Red Algae in the Genomic Age. Edited by Seckbach J., Chapman D.J. pp. 373–389. Springer, Dordrecht; New York. [Google Scholar]
- Foflonker F., Mollegard D., Ong M., Yoon H.S., Bhattacharya D. (2018) Genomic analysis of Picochlorum species reveals how microalgae may adapt to variable environments. Mol. Biol. Evol. 35: 2702–2711. [DOI] [PubMed] [Google Scholar]
- Fujii G., Imamura S., Era A., Miyagishima S.Y., Hanaoka M., Tanaka K. (2015) The nuclear-encoded sigma factor SIG4 directly activates transcription of chloroplast psbA and ycf17 genes in the unicellular red alga Cyanidioschyzon merolae. FEMS Microbiol. Lett. 362: fnv063. [DOI] [PubMed] [Google Scholar]
- Fujii G., Imamura S., Hanaoka M., Tanaka K. (2013) Nuclear-encoded chloroplast RNA polymerase sigma factor SIG2 activates chloroplast-encoded phycobilisome genes in a red alga, Cyanidioschyzon merolae. FEBS Lett. 587: 3354–3359. [DOI] [PubMed] [Google Scholar]
- Fujiwara T., Hirooka S., Mukai M., Ohbayashi R., Kanesaki Y., Watanabe S., et al. (2019) Integration of a Galdieria plasma membrane sugar transporter enables heterotrophic growth of the obligate photoautotrophic red alga Cyanidioschyzon merolae. Plant Direct 3: e00134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Hirooka S., Ohbayashi R., Onuma R., Miyagishima S.Y. (2020) Relationship between cell cycle and diel transcriptomic changes in metabolism in a unicellular red alga. Plant Physiol. 183: 1484–1501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Kanesaki Y., Hirooka S., Era A., Sumiya N., Yoshikawa H., et al. (2015) A nitrogen source-dependent inducible and repressible gene expression system in the red alga Cyanidioschyzon merolae. Front Plant Sci. 6: 657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Kuroiwa H., Yagisawa F., Ohnuma M., Yoshida Y., Yoshida M., et al. (2010) The coiled-coil protein VIG1 is essential for tethering vacuoles to mitochondria during vacuole inheritance of Cyanidioschyzon merolae. Plant Cell 22: 772–781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Ohnuma M., Kuroiwa T., Ohbayashi R., Hirooka S., Miyagishima S.Y. (2017) Development of a double nuclear gene-targeting method by two-step transformation based on a newly established chloramphenicol-selection system in the red alga Cyanidioschyzon merolae. Front Plant Sci. 8: 343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Ohnuma M., Yoshida M., Kuroiwa T., Hirano T. (2013) Gene targeting in the red alga Cyanidioschyzon merolae: single- and multi-copy insertion using authentic and chimeric selection markers. PLoS One 8: e73608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Tanaka K., Kuroiwa T., Hirano T. (2013) Spatiotemporal dynamics of condensins I and II: evolutionary insights from the primitive red alga Cyanidioschyzon merolae. Mol. Biol. Cell 24: 2515–2527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gentry M.S., Dowen R.H. 3rd, Worby C.A., Mattoo S., Ecker J.R., Dixon J.E. (2007) The phosphatase laforin crosses evolutionary boundaries and links carbohydrate metabolism to neuronal disease. J. Cell Biol. 178: 477–488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gong W., Marchetti A. (2019) Estimation of 18S gene copy number in marine eukaryotic plankton using a next-generation sequencing approach. Front. Mar. Sci. 6: 219. [Google Scholar]
- Graham L.D., Wilcox L.W. (2000) Algae. Prentice Hall, Upper Saddle River, NJ. [Google Scholar]
- Gross W. (1999) Revision of comparative traits for the acido- and thermophilic red algae Cyanidium and Galdieria. InEvolutionary Pathways and Enigmatic Algae: Cyanidium caldarium (Rhodophyta) and Related Cells. Edited by Seckbach J.. pp. 437–446. Springer, Netherlands. [Google Scholar]
- Harris E.H., Stern D.B., Witman G.B. (2009) The Chlamydomonas Sourcebook. Elsevier/Academic Press, Amsterdam. [Google Scholar]
- Hirabaru C., Izumo A., Fujiwara S., Tadokoro Y., Shimonaga T., Konishi M., et al. (2010) The primitive rhodophyte Cyanidioschyzon merolae contains a semiamylopectin-type, but not an amylose-type, alpha-glucan. Plant Cell Physiol. 51: 682–693. [DOI] [PubMed] [Google Scholar]
- Hirooka S., Hirose Y., Kanesaki Y., Higuchi S., Fujiwara T., Onuma R., et al. (2017) Acidophilic green algal genome provides insights into adaptation to an acidic environment. Proc. Natl. Acad. Sci. USA 114: E8304–E8313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirooka S., Miyagishima S.Y. (2016) Cultivation of acidophilic algae Galdieria sulphuraria and Pseudochlorella sp. YKT1 in media derived from acidic hot springs. Front. Microbiol. 7: 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirooka S., Tomita R., Fujiwara T., Ohnuma M., Kuroiwa H., Kuroiwa T., et al. (2020) Efficient open cultivation of cyanidialean red algae in acidified seawater. Sci. Rep. 10: 13794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hudson A.J., Stark M.R., Fast N.M., Russell A.G., Rader S.D. (2015) Splicing diversity revealed by reduced spliceosomes in C. merolae and other organisms. RNA Biol. 12: 1–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ichinose T.M., Iwane A.H. (2017) Cytological analyses by advanced electron microscopy. InCyanidioschyzon merolae: A New Model Eukaryote for Cell and Organelle Biology. Edited by Kuroiwa T., Miyagishima S.Y., Matsunaga S., Sato N., Nozaki H., Tanaka K.. et al. Springer Nature, Singapore. [Google Scholar]
- Ichinose T.M., Iwane A.H. (2021) Long-term live cell cycle imaging of single Cyanidioschyzon merolae cells. Protoplasma., in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imamura S., Kanesaki Y., Ohnuma M., Inouye T., Sekine Y., Fujiwara T., et al. (2009) R2R3-type MYB transcription factor, CmMYB1, is a central nitrogen assimilation regulator in Cyanidioschyzon merolae. Proc. Natl. Acad. Sci. USA 106: 12548–12553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imamura S., Kawase Y., Kobayashi I., Sone T., Era A., Miyagishima S.Y., et al. (2015) Target of rapamycin (TOR) plays a critical role in triacylglycerol accumulation in microalgae. Plant Mol. Biol. 89: 309–318. [DOI] [PubMed] [Google Scholar]
- Imamura S., Terashita M., Ohnuma M., Maruyama S., Minoda A., Weber A.P., et al. (2010) Nitrate assimilatory genes and their transcriptional regulation in a unicellular red alga Cyanidioschyzon merolae: genetic evidence for nitrite reduction by a sulfite reductase-like enzyme. Plant Cell Physiol. 51: 707–717. [DOI] [PubMed] [Google Scholar]
- Imoto Y., Kuroiwa H., Yoshida Y., Ohnuma M., Fujiwara T., Yoshida M., et al. (2013) Single-membrane-bounded peroxisome division revealed by isolation of dynamin-based machinery. Proc. Natl. Acad. Sci. USA 110: 9583–9588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janouskovec J., Liu S.L., Martone P.T., Carre W., Leblanc C., Collen J., et al. (2013) Evolution of red algal plastid genomes: ancient architectures, introns, horizontal gene transfer, and taxonomic utility of plastid markers. PLoS One 8: e59001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jing Y., Lin R. (2020) Transcriptional regulatory network of the light signaling pathways. New Phytol. 227: 683–697. [DOI] [PubMed] [Google Scholar]
- Johnson C.H. (2010) Circadian clocks and cell division: what's the pacemaker? Cell Cycle 9: 3864–3873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanesaki Y., Imamura S., Matsuzaki M., Tanaka K. (2015) Identification of centromere regions in chromosomes of a unicellular red alga, Cyanidioschyzon merolae. FEBS Lett. 589: 1219–1224. [DOI] [PubMed] [Google Scholar]
- Khan M.I., Shin J.H., Kim J.D. (2018) The promising future of microalgae: current status, challenges, and optimization of a sustainable and renewable industry for biofuels, feed, and other products. Microb. Cell Fact. 17: 36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiefel B.R., Gilson P.R., Beech P.L. (2006) Cell biology of mitochondrial dynamics. Int. Rev. Cytol. 254: 151–213. [DOI] [PubMed] [Google Scholar]
- Kobayashi Y., Ando H., Hanaoka M., Tanaka K. (2016) Abscisic acid participates in the control of cell cycle initiation through heme homeostasis in the unicellular red alga Cyanidioschyzon merolae. Plant Cell Physiol. 57: 953–960. [DOI] [PubMed] [Google Scholar]
- Kobayashi Y., Imamura S., Hanaoka M., Tanaka K. (2011) A tetrapyrrole-regulated ubiquitin ligase controls algal nuclear DNA replication. Nat. Cell Biol. 13: 483–487. [DOI] [PubMed] [Google Scholar]
- Kobayashi Y., Kanesaki Y., Tanaka A., Kuroiwa H., Kuroiwa T., Tanaka K. (2009) Tetrapyrrole signal as a cell-cycle coordinator from organelle to nuclear DNA replication in plant cells. Proc. Natl. Acad. Sci. USA 106: 803–807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krupnik T., Kotabova E., van Bezouwen L.S., Mazur R., Garstka M., Nixon P.J., et al. (2013) A reaction center-dependent photoprotection mechanism in a highly robust photosystem II from an extremophilic red alga, Cyanidioschyzon merolae. J. Biol. Chem. 288: 23529–23542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroiwa T., Kuroiwa H., Sakai A., Takahashi H., Toda K., Itoh R. (1998) The division apparatus of plastids and mitochondria. Int. Rev. Cytol. 181: 1–41. [DOI] [PubMed] [Google Scholar]
- Lemieux C., Turmel M., Otis C., Pombert J.F. (2019) A streamlined and predominantly diploid genome in the tiny marine green alga Chloropicon primus. Nat. Commun. 10: 4061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin D., Yin X., Wang X., Zhou P., Guo F.B. (2013) Re-annotation of protein-coding genes in the genome of Saccharomyces cerevisiae based on support vector machines. PLoS One 8: e64477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin S., Offner G.D., Troxler R.F. (1990) Studies on Cyanidium caldarium phycobiliprotein pigment mutants. Plant Physiol. 93: 772–777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu S.-L., Chiang Y.-R., Yoon H.S., Fu H.-Y. (2020) Comparative genome analysis reveals Cyanidiococcus gen. nov., a new extremophilic red algal genus sister to Cyanidioschyzon (Cyanidioschyzonaceae, Rhodophyta). J. Phycol. 56: 1428–1442. [DOI] [PubMed] [Google Scholar]
- Markley J.L., Aceti D.J., Bingman C.A., Fox B.G., Frederick R.O., Makino S., et al. (2009) The center for eukaryotic structural genomics. J. Struct. Funct. Genomics 10: 165–179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maschmann S., Ruban K., Wientapper J., Walter W.J. (2020) Phototaxis of the unicellular red alga Cyanidioschyzon merolae is mediated by novel actin-driven tentacles. Int. J. Mol. Sci. 21: 6209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuzaki M., Misumi O., Shin I.T., Maruyama S., Takahara M., Miyagishima S.Y., et al. (2004) Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 428: 653–657. [DOI] [PubMed] [Google Scholar]
- Merchant S.S., Prochnik S.E., Vallon O., Harris E.H., Karpowicz S.J., Witman G.B., et al. (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245–250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merola A., Castaldo R., De Luca P., Gambardella R., Musacchio A., Taddei R. (1981) Revision of Cyanidium caldarium: three species of acidophilic algae. Gio. Bot. Ital. 115: 189–195. [Google Scholar]
- Mikulski P., Komarynets O., Fachinelli F., Weber A.P.M., Schubert D. (2017) Characterization of the polycomb-group mark H3K27me3 in unicellular algae. Front Plant Sci. 8: 607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minoda A., Sakagami R., Yagisawa F., Kuroiwa T., Tanaka K. (2004) Improvement of culture conditions and evidence for nuclear transformation by homologous recombination in a red alga, Cyanidioschyzon merolae 10D. Plant Cell Physiol. 45: 667–671. [DOI] [PubMed] [Google Scholar]
- Minoda A., Weber A.P., Tanaka K., Miyagishima S.Y. (2010) Nucleus-independent control of the rubisco operon by the plastid-encoded transcription factor Ycf30 in the red alga Cyanidioschyzon merolae. Plant Physiol. 154: 1532–1540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagishima S.Y., Era A., Hasunuma T., Matsuda M., Hirooka S., Sumiya N., et al. (2019) Day/night separation of oxygenic energy metabolism and nuclear DNA replication in the unicellular red alga Cyanidioschyzon merolae. mBio 10: e00833-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagishima S.Y., Fujiwara T., Sumiya N., Hirooka S., Nakano A., Kabeya Y., et al. (2014) Translation-independent circadian control of the cell cycle in a unicellular photosynthetic eukaryote. Nat. Commun. 5: 3807. [DOI] [PubMed] [Google Scholar]
- Miyagishima S.Y., Itoh R., Toda K., Kuroiwa H., Nishimura M., Kuroiwa T. (1999) Microbody proliferation and segregation cycle in the single-microbody alga Cyanidioschyzon merolae. Planta 208: 326–336. [Google Scholar]
- Miyagishima S.Y., Nakanishi H., Kabeya Y. (2011) Structure, regulation, and evolution of the plastid division machinery. Int. Rev. Cell Mol. Biol. 291: 115–153. [DOI] [PubMed] [Google Scholar]
- Miyagishima S.Y., Nishida K., Mori T., Matsuzaki M., Higashiyama T., Kuroiwa H., et al. (2003) A plant-specific dynamin-related protein forms a ring at the chloroplast division site. Plant Cell 15: 655–665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagishima S.Y., Suzuki K., Okazaki K., Kabeya Y. (2012) Expression of the nucleus-encoded chloroplast division genes and proteins regulated by the algal cell cycle. Mol. Biol. Evol. 29: 2957–2970. [DOI] [PubMed] [Google Scholar]
- Mori N., Moriyama T., Sato N. (2019) Uncommon properties of lipid biosynthesis of isolated plastids in the unicellular red alga Cyanidioschyzon merolae. FEBS Open Bio 9: 114–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mori N., Moriyama T., Toyoshima M., Sato N. (2016) Construction of global acyl lipid metabolic map by comparative genomics and subcellular localization analysis in the red alga Cyanidioschyzon merolae. Front Plant Sci. 7: 958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moriyama T., Mori N., Sato N. (2015) Activation of oxidative carbon metabolism by nutritional enrichment by photosynthesis and exogenous organic compounds in the red alga Cyanidioschyzon merolae: evidence for heterotrophic growth. SpringerPlus 4: 559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolova D., Weber D., Scholz M., Bald T., Scharsack J.P., Hippler M. (2017) Temperature-induced remodeling of the photosynthetic machinery tunes photosynthesis in the thermophilic alga Cyanidioschyzon merolae. Plant Physiol. 174: 35–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson H., Krupnik T., Kargul J., Messinger J. (2014) Substrate water exchange in photosystem II core complexes of the extremophilic red alga Cyanidioschyzon merolae. Biochim. Biophys. Acta 1837: 1257–1262. [DOI] [PubMed] [Google Scholar]
- Nishida K., Takahara M., Miyagishima S.Y., Kuroiwa H., Matsuzaki M., Kuroiwa T. (2003) Dynamic recruitment of dynamin for final mitochondrial severance in a primitive red alga. Proc. Natl. Acad. Sci. USA 100: 2146–2151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishida K., Yagisawa F., Kuroiwa H., Yoshida Y., Kuroiwa T. (2007) WD40 protein Mda1 is purified with Dnm1 and forms a dividing ring for mitochondria before Dnm1 in Cyanidioschyzon merolae. Proc. Natl. Acad. Sci. USA 104: 4736–4741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nozaki H., Takano H., Misumi O., Terasawa K., Matsuzaki M., Maruyama S., et al. (2007) A 100%-complete sequence reveals unusually simple genomic features in the hot-spring red alga Cyanidioschyzon merolae. BMC Biol. 5: 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oesterhelt C., Schnarrenberger C., Gross W. (1999) Characterization of a sugar/polyol uptake system in the red alga Galdieria sulphuraria. Eur. J. Phycol. 34: 271–277. [Google Scholar]
- Ohnuma M., Kuroiwa T., Tanaka K. (2011) Optimization of cryopreservation conditions for the unicellular red alga Cyanidioschyzon merolae. J. Gen. Appl. Microbiol. 57: 137–143. [DOI] [PubMed] [Google Scholar]
- Ohnuma M., Misumi O., Fujiwara T., Watanabe S., Tanaka K., Kuroiwa T. (2009) Transient gene suppression in a red alga, Cyanidioschyzon merolae 10D. Protoplasma 236: 107–112. [DOI] [PubMed] [Google Scholar]
- Ohnuma M., Misumi O., Kuroiwa T. (2011) Phototaxis in the unicellular red algae Cyanidioschyzon merolae and Cyanidium caldarium. Cytologia 76: 295–300. [Google Scholar]
- Ohnuma M., Yokoyama T., Inouye T., Sekine Y., Tanaka K. (2008) Polyethylene glycol (PEG)-mediated transient gene expression in a red alga, Cyanidioschyzon merolae 10D. Plant Cell Physiol. 49: 117–120. [DOI] [PubMed] [Google Scholar]
- Ohta H., Suzuki T., Ueno M., Okumura A., Yoshihara S., Shen J.R., et al. (2003) Extrinsic proteins of photosystem II: an intermediate member of PsbQ protein family in red algal PS II. Eur. J. Biochem. 270: 4156–4163. [DOI] [PubMed] [Google Scholar]
- Ohta N., Matsuzaki M., Misumi O., Miyagishima S.Y., Nozaki H., Tanaka K., et al. (2003) Complete sequence and analysis of the plastid genome of the unicellular red alga Cyanidioschyzon merolae. DNA Res. 10: 67–77. [DOI] [PubMed] [Google Scholar]
- Ohta N., Sato N., Kuroiwa T. (1998) Structure and organization of the mitochondrial genome of the unicellular red alga Cyanidioschyzon merolae deduced from the complete nucleotide sequence. Nucleic Acids Res. 26: 5190–5198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osteryoung K.W., Vierling E. (1995) Conserved cell and organelle division. Nature 376: 473–474. [DOI] [PubMed] [Google Scholar]
- Pancha I., Chokshi K., Tanaka K., Imamura S. (2020) Microalgal target of rapamycin (TOR): a central regulatory hub for growth, stress response and biomass production. Plant Cell Physiol. 61: 675–684. [DOI] [PubMed] [Google Scholar]
- Pham L.V., Janna Olmos J.D., Chernev P., Kargul J., Messinger J. (2019) Unequal misses during the flash-induced advancement of photosystem II: effects of the S state and acceptor side cycles. Photosynth. Res. 139: 93–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pi X., Tian L., Dai H.E., Qin X., Cheng L., Kuang T., et al. (2018) Unique organization of photosystem I-light-harvesting supercomplex revealed by cryo-EM from a red alga. Proc. Natl. Acad. Sci. USA 115: 4423–4428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinto G. (2007) Cyanidiophyceae: looking back–looking forward. InAlgae and Cyanobacteria in Extreme Environments. Edited by Seckbach J. pp. 387–397. Springer, Netherlands. [Google Scholar]
- Poliner E., Farre E.M., Benning C. (2018) Advanced genetic tools enable synthetic biology in the oleaginous microalgae Nannochloropsis sp. Plant Cell Rep. 37: 1383–1399. [DOI] [PubMed] [Google Scholar]
- Puthiyaveetil S., Allen J.F. (2009) Chloroplast two-component systems: evolution of the link between photosynthesis and gene expression. Proc. Biol. Sci. 276: 2133–2145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi L.S., Larson M.H., Gilbert L.A., Doudna J.A., Weissman J.S., Arkin A.P., et al. (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152: 1173–1183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu H., Price D.C., Weber A.P., Reeb V., Yang E.C., Lee J.M., et al. (2013) Adaptation through horizontal gene transfer in the cryptoendolithic red alga Galdieria phlegrea. Curr. Biol. 23: R865–R866. [DOI] [PubMed] [Google Scholar]
- Rademacher N., Wrobel T.J., Rossoni A.W., Kurz S., Brautigam A., Weber A.P.M., et al. (2017) Transcriptional response of the extremophile red alga Cyanidioschyzon merolae to changes in CO2 concentrations. J. Plant Physiol. 217: 49–56. [DOI] [PubMed] [Google Scholar]
- Rahman D.Y., Sarian F.D., van Wijk A., Martinez-Garcia M., van der Maarel M. (2017) Thermostable phycocyanin from the red microalga Cyanidioschyzon merolae, a new natural blue food colorant. J Appl. Phycol. 29: 1233–1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reimer K.A., Stark M.R., Aguilar L.C., Stark S.R., Burke R.D., Moore J., et al. (2017) The sole LSm complex in Cyanidioschyzon merolae associates with pre-mRNA splicing and mRNA degradation factors. RNA 23: 952–967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigano C., Aliotta G., Rigano V.D., Fuggi A., Vona V. (1977) Heterotrophic growth patterns in the unicellular alga Cyanidium caldarium. A possible role for threonine dehydrase. Arch. Microbiol. 113: 191–196. [DOI] [PubMed] [Google Scholar]
- Rodriguez-Ezpeleta N., Philippe H. (2006) Plastid origin: replaying the tape. Curr. Biol. 16: R53–R56. [DOI] [PubMed] [Google Scholar]
- Rossoni A.W., Price D.C., Seger M., Lyska D., Lammers P., Bhattacharya D., et al. (2019) The genomes of polyextremophilic cyanidiales contain 1% horizontally transferred genes with diverse adaptive functions. eLife 8: e45017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanchez F., Geffroy S., Norest M., Yau S., Moreau H., Grimsley N. (2019) Simplified transformation of Ostreococcus tauri using polyethylene glycol. Genes 10: 399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato N., Moriyama T., Mori N., Toyoshima M. (2017) Lipid metabolism and potentials of biofuel and high added-value oil production in red algae. World J. Microbiol. Biotechnol. 33: 74. [DOI] [PubMed] [Google Scholar]
- Schönknecht G., Chen W.H., Ternes C.M., Barbier G.G., Shrestha R.P., Stanke M., et al. (2013) Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote. Science 339: 1207–1210. [DOI] [PubMed] [Google Scholar]
- Shaver S., Casas-Mollano J.A., Cerny R.L., Cerutti H. (2010) Origin of the polycomb repressive complex 2 and gene silencing by an E(z) homolog in the unicellular alga Chlamydomonas. Epigenetics 5: 301–312. [DOI] [PubMed] [Google Scholar]
- Stark M.R., Dunn E.A., Dunn W.S., Grisdale C.J., Daniele A.R., Halstead M.R., et al. (2015) Dramatically reduced spliceosome in Cyanidioschyzon merolae. Proc. Natl. Acad. Sci. USA 112: E1191–E1200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strenkert D., Schmollinger S., Gallaher S.D., Salome P.A., Purvine S.O., Nicora C.D., et al. (2019) Multiomics resolution of molecular events during a day in the life of Chlamydomonas. Proc. Natl. Acad. Sci. USA 116: 2374–2383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumiya N., Fujiwara T., Era A., Miyagishima S.Y. (2016) Chloroplast division checkpoint in eukaryotic algae. Proc. Natl. Acad. Sci. USA 113: E7629–E7638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumiya N., Fujiwara T., Kobayashi Y., Misumi O., Miyagishima S.Y. (2014) Development of a heat-shock inducible gene expression system in the red alga Cyanidioschyzon merolae. PLoS One 9: e111261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki K., Ehara T., Osafune T., Kuroiwa H., Kawano S., Kuroiwa T. (1994) Behavior of mitochondria, chloroplasts and their nuclei during the mitotic cycle in the ultramicroalga Cyanidioschyzon merolae. Eur. J. Cell Biol. 63: 280–288. [PubMed] [Google Scholar]
- Takemura T., Imamura S., Kobayashi Y., Tanaka K. (2018) Construction of a selectable marker recycling system and the use in epitope tagging of multiple nuclear genes in the unicellular red alga Cyanidioschyzon merolae. Plant Cell Physiol. 59: 2308–2316. [DOI] [PubMed] [Google Scholar]
- Takemura T., Imamura S., Kobayashi Y., Tanaka K. (2019) Multiple modification of chromosomal loci using URA5.3 selection marker in the unicellular red alga Cyanidioschyzon merolae. Bio Protoc. 9: 3204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taki K., Sone T., Kobayashi Y., Watanabe S., Imamura S., Tanaka K. (2015) Construction of a URA5.3 deletion strain of the unicellular red alga Cyanidioschyzon merolae: a backgroundless host strain for transformation experiments. J. Gen. Appl. Microbiol. 61: 211–214. [DOI] [PubMed] [Google Scholar]
- Tanaka K. (2012) Chloroplast transcription machinery of red algae: conservation of four types of transcriptional regulators in non-green chloroplasts. In Porphyra Yezoensis, Frontiers in Physiological and Molecular Biological Research. Edited by Mikami K. pp. 39–59. Nova Science Publishers, New York, NY. [Google Scholar]
- Tanaka K., Oikawa K., Ohta N., Kuroiwa H., Kuroiwa T., Takahashi H. (1996) Nuclear encoding of a chloroplast RNA polymerase sigma subunit in a red alga. Science 272: 1932–1935. [DOI] [PubMed] [Google Scholar]
- Tardu M., Dikbas U.M., Baris I., Kavakli I.H. (2016) RNA-seq analysis of the transcriptional response to blue and red light in the extremophilic red alga, Cyanidioschyzon merolae. Funct. Integr. Genomics 16: 657–669. [DOI] [PubMed] [Google Scholar]
- Tran N.T., Kaldenhoff R. (2020) Achievements and challenges of genetic engineering of the model green alga Chlamydomonas reinhardtii. Algal. Res. 50: 101986. [Google Scholar]
- Veluchamy A., Rastogi A., Lin X., Lombard B., Murik O., Thomas Y., et al. (2015) An integrative analysis of post-translational histone modifications in the marine diatom Phaeodactylum tricornutum. Genome Biol. 16: 102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieler A., Wu G., Tsai C.H., Bullard B., Cornish A.J., Harvey C., et al. (2012) Genome, functional gene annotation, and nuclear transformation of the heterokont oleaginous alga Nannochloropsis oceanica CCMP1779. PLoS Genet. 8: e1003064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vishwakarma K., Upadhyay N., Kumar N., Yadav G., Singh J., Mishra R.K., et al. (2017) Abscisic acid signaling and abiotic stress tolerance in plants: a review on current knowledge and future prospects. Front Plant Sci. 8: 161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker J.J., Spear J.R., Pace N.R. (2005) Geobiology of a microbial endolithic community in the Yellowstone geothermal environment. Nature 434: 1011–1014. [DOI] [PubMed] [Google Scholar]
- Yagisawa F., Fujiwara T., Ohnuma M., Kuroiwa H., Nishida K., Imoto Y., et al. (2013) Golgi inheritance in the primitive red alga, Cyanidioschyzon merolae. Protoplasma 250: 943–948. [DOI] [PubMed] [Google Scholar]
- Yagisawa F., Fujiwara T., Takemura T., Kobayashi Y., Sumiya N., Miyagishima S.Y., et al. (2020) ESCRT machinery mediates cytokinetic abscission in the unicellular red alga Cyanidioschyzon merolae. Front Cell Dev. Biol. 8: 169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang E.C., Kim K.M., Kim S.Y., Lee J., Boo G.H., Lee J.H., et al. (2015) Highly conserved mitochondrial genomes among multicellular red algae of the florideophyceae. Genome Biol. Evol. 7: 2394–2406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yesbolatova A., Saito Y., Kitamoto N., Makino-Itou H., Ajima R., Nakano R., et al. (2020) The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice. Nat. Commun. 11: 5701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoon H.S., Hackett J.D., Ciniglia C., Pinto G., Bhattacharya D. (2004) A molecular timeline for the origin of photosynthetic eukaryotes. Mol. Biol. Evol. 21: 809–818. [DOI] [PubMed] [Google Scholar]
- Yoon H.S., Müller K.M., Sheath R.G., Ott F.D., Bhattacharya D. (2006) Defining the major lineages of red algae (Rhodophyta). J. Phycol. 42: 482–492. [Google Scholar]
- Yoshida Y., Kuroiwa H., Misumi O., Yoshida M., Ohnuma M., Fujiwara T., et al. (2010) Chloroplasts divide by contraction of a bundle of nanofilaments consisting of polyglucan. Science 329: 949–953. [DOI] [PubMed] [Google Scholar]
- Yoshida Y., Kuroiwa H., Shimada T., Yoshida M., Ohnuma M., Fujiwara T., et al. (2017) Glycosyltransferase MDR1 assembles a dividing ring for mitochondrial proliferation comprising polyglucan nanofilaments. Proc. Natl. Acad. Sci. USA 114: 13284–13289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zenvirth D., Volokita M., Kaplan A. (1985) Photosynthesis and inorganic carbon accumulation in the acidophilic alga Cyanidioschyzon merolae. Plant Physiol. 77: 237–239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zienkiewicz M., Krupnik T., Drozak A., Golke A., Romanowska E. (2017a) Chloramphenicol acetyltransferase-a new selectable marker in stable nuclear transformation of the red alga Cyanidioschyzon merolae. Protoplasma 254: 587–596. [DOI] [PubMed] [Google Scholar]
- Zienkiewicz M., Krupnik T., Drozak A., Golke A., Romanowska E. (2017b) Transformation of the Cyanidioschyzon merolae chloroplast genome: prospects for understanding chloroplast function in extreme environments. Plant Mol. Biol. 93: 171–183. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.