Abstract
Growth and differentiation of Candida albicans over a broad pH range underlie its ability to infect an array of tissues in susceptible hosts. We identified C. albicans RIM101, RIM20, and RIM8 based on their homology to components of the one known fungal pH response pathway. PCR product-disruption mutations in each gene cause defects in three responses to alkaline pH: filamentation, induction of PRA1 and PHR1, and repression of PHR2. We find that RIM101 itself is an alkaline-induced gene that also depends on Rim20p and Rim8p for induction. Two observations indicate that a novel pH response pathway also exists. First, PHR2 becomes an alkaline-induced gene in the absence of Rim101p, Rim20p, or Rim8p. Second, we created strains in which Rim101p activity is independent of Rim20p and Rim8p; in these strains, filamentation remains pH dependent. Thus, pH governs gene expression and cellular differentiation in C. albicans through both RIM101-dependent and RIM101-independent pathways.
Candida albicans is the most common human fungal pathogen. In most individuals, it is a commensal fungus that can colonize environmentally diverse niches, such as the oral and vaginal cavities (19). In susceptible hosts, C. albicans can infect virtually any tissue (9). Thus, both colonization and infection by C. albicans require its adaptation to diverse environments.
One environmental variable, extracellular pH, governs C. albicans cellular morphology. At acidic pH, C. albicans grows in the yeast form; at alkaline pH, it grows primarily in the filament form (19). Alkaline-induced filamentation correlates with a number of physiological changes, such as alterations in cell wall architecture and adhesion properties (4). These responses to alkaline pH are thought to be dependent on changes in gene expression.
Differential expression screens have led to the identification of two alkaline-induced genes, PRA1 and PHR1. PRA1 specifies a cell wall protein that has a minor role in alkaline-induced filamentation (22). PHR1 also specifies a cell wall protein; however, it is required for both growth and filamentation at alkaline pH (21). Ectopic expression of PHR2, a PHR1 homolog that is not normally expressed at alkaline pH, permits both filamentation and growth of the phr1−/phr1− mutant (18). Since PHR2 does not promote filamentation when it is normally expressed, the phr1−/phr1− alkaline-induced filamentation defect appears to be a result of the growth defect. Thus, the functions of known alkaline-induced genes do not explain how alkaline pH induces filamentation.
A number of regulators that control the yeast-to-filament transition have been identified, including Efg1p, Cph1p, and Tup1p (3, 12, 14), but these factors are not thought to be specific for alkaline responses. A conserved alkaline response pathway has been identified in the fungi Aspergillus nidulans, Saccharomyces cerevisiae, and Yarrowia lipolytica (6, 10, 11, 15, 20, 25, 26). In this pathway, the zinc finger transcription factor Rim101p/PacC, from S. cerevisiae and A. nidulans, respectively, stimulates expression of alkaline response genes and represses acidic response genes (26). Rim101p/PacC activity is controlled by proteolytic processing. In acidic conditions, Rim101p exists primarily in a full-length “long” form which has no known function (11, 20). In alkaline conditions, a C-terminal portion is cleaved to yield the active “short” form. Proteolysis is controlled by pH through the action of a number of gene products, including Rim20p/PalA, Rim8p/PalF, Rim13p/PalB, and Rim9p/PalI (5, 6, 11, 15). Here we have used the partial C. albicans genomic database to identify RIM101 pathway members in C. albicans and to determine their role in alkaline responses. Our studies demonstrate that the RIM101 pathway in C. albicans is required for some alkaline responses. Further, our results also indicate that a second pH response pathway must exist in C. albicans.
MATERIALS AND METHODS
Strains and plasmids.
The C. albicans strains used in this study are derivatives of CAI4 and are described in Table 1. Creation of the rim101−/rim101− and rim20−/rim20− strains was described previously (27). They were referred to as hrm101−/hrm101− and enx3−/enx3−, respectively. For complementation and suppression studies (see below for details), the appropriate strains were subjected to transformation and selection for histidine prototrophy. Plasmids were maintained and amplified with the bacterial strain DH5α.
TABLE 1.
C. albicans strains
| Name | Genotype | Reference |
|---|---|---|
| BWP17 | ura3Δ::λimm434his1::hisGarg4::hisG | 27 |
| ura3Δ::λimm434 his1::hisG arg4::hisG | ||
| DAY2 | ura3Δ::λimm434his1::hisGarg4::hisGrim101::ARG4 | 27 |
| ura3Δ::λimm434 his1::hisG arg4::hisG RIM101 | ||
| DAY5 | ura3Δ::λimm434his1::hisGarg4::hisGrim101::ARG4 | 27 |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim101::URA3 | ||
| DAY23 | ura3Δ::λimm434his1::hisGarg4::hisGrim20::URA3 | 27 |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim20::ARG4 | ||
| DAY44 | ura3Δ::λimm434his1::hisGarg4::hisGrim101::ARG4pRIM101::HIS1 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim101::URA3 RIM101 | ||
| DAY61 | ura3Δ::λimm434his1::hisGarg4::hisGrim8::URA3 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim8::ARG4 | ||
| DAY62 | ura3Δ::λimm434his1::hisGarg4::hisGrim8::URA3pRIM101-405::HIS1 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim8::ARG4 RIM101 | ||
| DAY90 | ura3Δ::λimm434his1::hisGarg4::hisGpRIM101-405::HIS1 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG RIM101 | ||
| DAY93 | ura3Δ::λimm434his1::hisGarg4::hisGrim20::URA3pRIM101-405::HIS1 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim20::ARG4 RIM101 | ||
| DAY96 | ura3Δ::λimm434his1::hisGarg4::hisGrim101::ARG4pRIM101-405::HIS1 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim101::URA3 | ||
| DAY111 | ura3Δ::λimm434his1::hisGarg4::hisGrim20::ARG4pRIM101 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim20::URA3 RIM101 | ||
| DAY114 | ura3Δ::λimm434his1::hisGarg4::hisGrim8::ARG4pRIM101 | This study |
| ura3Δ::λimm434 his1::hisG arg4::hisG rim8::URA3 RIM101 |
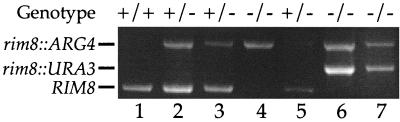
The rim8−/rim8− mutant (DAY61) was generated as follows. Strain BWP17 was subjected to consecutive rounds of transformation with rim8::ARG4 and rim8::URA3 using primers RIM8-5DR-2 and RIM8-3DR-2 as described previously (27). This deletes sequences from −77 to +1620, removing residues 1 to 398 of the predicted protein. Correct integration was demonstrated by PCR with the primers rim8-3-2 and rim8-5-2, which flank the site of integration (Fig. 1).
FIG. 1.
Confirmation of the rim8−/rim8− mutant. The figure shows a gel following PCR with flanking detection primers of genomic DNA of wild type (sample 1), rim8::ARG4 transformants (samples 2 to 5), and rim8::URA3 transformants (samples 6 and 7). Note integration at both RIM8 copies in one case (sample 5) following tranformation with rim8::ARG4.
Plasmid pDDB61 was constructed as follows. Full-length RIM101 was PCR amplified with primers seq4c and gc3cloneSalI (Table 2) and cloned into pGEM-T (Promega) to generate plasmid pDDB52. pDDB52 was digested with SalI, and the RIM101-containing fragment was cloned into the SalI site of pGEM-HIS1 (27), generating pDDB61.
TABLE 2.
Synthetic oligonucleotides
| Primer name | Sequence (5′–3′) |
|---|---|
| RIM8-5DR-2 | CTCGGTTTGGGTGGAATCAAAGTTAAACTTAAACTTGTCCAGTGATGATTTGGTTGCTTGGTTTTCCCAGTCACGACGTT |
| RIM8-3DR-2 | GAAAAGTGTATCAACCATTCTCATTCTTTCTTTTTTTTTTTTTCTCTCTACGAACAGAAATGTGGAATTGTGAGCGGATA |
| rim8-5-2 | ATCGGTATTGATATAGGTAGATC |
| rim8-3-2 | AAAACTAAATTAGTCTCCACCAC |
| carim8-5 | CTCGGTTTGGGTGGAATCAAAG |
| carim8-3 | CAACCATTTACGAAAGATCCAGG |
| RIM8-500clone | GGGTCGACCCTTTGACCTAAAAAGGCCCAGAC |
| RIM8 seq3c | GTTCCTGGACAAATCGTCATCC |
| rim20dn5 | TGAAACAATATTTGTTACAGGCTAAC |
| rim20dn3 | CCAATTGTTCTGCCGTTAAC |
| PHR2 5′ | GGGGAATTCCACACATTCGATCGCTATG |
| PHR2 3′ | GGGGAATTCGGAGAGTACACTCGTGAG |
| PRA1 5′ | GGGGAATTCGGCAACAATATCTCGTTGG |
| PRA1 3′ | GGGGAATTCGCCTGAACTTAACAATTAACAGTGG |
| TEF1-5′ | ATAGTCATAATCAATCATGGGT |
| TEF1-3′ | CTTACATAATATTCAACTAGC |
| PHR1-1 | ATTAGAGTCGCCATTGCCGATTATTTC |
| PHR1-2 | CCTGGACATTGCAAAGTCTTGCTGGCG |
| gc3cloneSalI | GGGTCGACGATANAAAAGTAAAATCGGTATAATTGC |
| RIM101 seq4c | CCATGGTTCCAAGAAATCACC |
Plasmid pDDB71, which contains the RIM101-405 allele, was generated as follows. pDDB61 was digested with EcoRI, filled in with Klenow polymerase, and religated to create pDDB71. This generates a RIM101 gene with a stop codon following amino acid N405. pDDB61 and pDDB71 were digested with PpuMI for transformations to target integration to the RIM101 or rim101− locus.
Plasmid pRIM8-URA3 was constructed as follows. A fragment of C. albicans RIM8 was amplified by PCR with primers carim8-5 and carim8-3 and cloned into pGEM-T, generating plasmid pBW113. pBW113 was digested with SacII and SpeI, and the RIM8-containing fragment was cloned into SacII/SpeI-digested pRSARG4ΔSpeI (27), generating plasmid pRIM8-ARG4. Plasmid pGEMT-URA3 (27) was digested with SalI and SphI, and the URA3-containing fragment was cloned into SalI/SphI-digested pBR322, generating plasmid pBRURA3. pBRURA3 was digested with ClaI/SpeI, and the URA3-containing fragment was cloned into ClaI/SpeI-digested pRIM8-ARG4, generating plasmid pRIM8-URA3.
Plasmid pRS424ARG4-URA3-BH1 was constructed as follows. Plasmid pGEMT-URA3 was digested with SacII and SpeI, and the URA3-containing fragment was cloned into the SacII/SpeI site of pRSARG4ΔSpeI, generating plasmid pRS314ARG4-URA3. An Asp718-BamHI linker was inserted into the KpnI site of pRS314ARG4-URA3, generating plasmid pRS314ARG4-URA3-BH1. pRS314ARG4-URA3-BH1 was digested with BamHI and SacII, and the ARG4-URA3-containing fragment was cloned into BamHI/SacII-digested pRS424, generating plasmid pRS424ARG4-URA3-BH1.
Media and growth conditions.
C. albicans was routinely grown in YPD plus uridine (2% Bacto Peptone, 1% yeast extract, 2% dextrose, and 80 μg of uridine per ml). Selection following transformation was done on synthetic medium (6.7% yeast nitrogen base plus ammonium sulfate and without amino acids, 2% dextrose, 80 μg of uridine per ml except when selecting for URA3, and supplemented with the necessary auxotrophic requirements of the cells) (1). TC199 medium (Gibco BRL) was buffered at either pH 4.0 or pH 8.0 with 150 mM HEPES and supplemented with 80 μg of uridine per ml. Cell densities were determined by light scattering at 600 nm.
For filamentation and Northern blot analyses, strains were grown overnight in YPD plus uridine at 30°C. The following day, cells were pelleted, washed with ddH2O, and diluted 40 to 100× into buffered TC199 or serum (germ tube medium; Remel) prewarmed to 38°C.
Retrieval of RIM101, RIM20, and RIM8.
C. albicans RIM101 was cloned by plasmid insertion-retrieval. pRS424-ARG4-URA3-BH1 was digested with BglII and integrated into rim101::ARG4 of DAY2. Genomic DNA was purified, digested with NcoI, ligated at 16°C for 5 days, and recovered in Escherichia coli by electroporation. Appropriate plasmids were purified on a Qiagen column and sequenced.
The 5′ end of RIM20 was cloned by integration of PpuMI-digested pRS424-ARG4-URA3-BH1 into DAY18. Genomic DNA was purified, digested with BamHI and BglII, and religated, and the vector with 5′ flanking DNA was recovered in DH5α. Primers for sequencing were generated to sequences from contig 3-3609 which contains preliminary sequence data including RIM20. The 3′ end of RIM20 was sequenced following PCR amplification with primers rim20dn5 and rim20dn3.
The 5′ end of the RIM8 sequence was obtained through searches of the C. albicans genomic database with S. cerevisiae RIM8 sequence (W. Xu and A. P. Mitchell, unpublished data). Ambiguous regions were sequenced following PCR amplification of RIM8 with primers RIM8-500 clone and seq3c. The 3′ end of C. albicans RIM8 was cloned and sequenced by plasmid retrieval similar to that for RIM101, except that pRIM8-URA3 was digested with BglII to target integration and genomic DNA was digested with SacI prior to ligation and introduction into DH5α.
Northern blot analyses.
Following 4 h of incubation at 38°C in TC199 medium, cells were harvested by vacuum filtration. RNA was purified either directly or from frozen cell pellets (1). Fifteen micrograms of total RNA was dried, resuspended in sample buffer, and separated by 1.2% formaldehyde gel electrophoresis. RNA was transferred to nylon membranes by capillary action and cross-linked. PCR-amplified PHR2, PHR1, PRA1, RIM101, and TEF1 from C. albicans genomic DNA were used to make probes (see Table 2 for primers used). Probes were made by using the High Prime kit (Boehringer Mannheim) and purified by using the QiaQuick PCR purification kit. Blots were analyzed with a phosphorimager and quantitated with IQMac v1.2.
Filamentation.
Following 4 h of incubation at 38°C in TC199 medium plus uridine or 2 h in serum plus uridine at 38°C, cells were fixed with 2 volumes of 100% ethanol. Cells were pelleted and resuspended in phosphate-buffered saline prior to microscopic analysis. At least 200 cells were counted for each sample.
Nucleotide sequence accession numbers.
The GenBank accession numbers for RIM101, RIM20, and RIM8 are AF173841, AF173843, and AF173842, respectively.
RESULTS
Cloning and sequencing of RIM101, RIM8, and RIM20.
To identify potential pH response regulators in C. albicans, we searched the partial Candida genomic database and identified short sequences with homology to S. cerevisiae RIM101, RIM20, and RIM8. To test the role of these sequences in the pH response, we generated insertion-deletion mutants with a PCR-product-directed gene disruption technique (27). Promising results from these studies, described below, prompted us to clone the remainder of these genes by plasmid integration and retrieval (see Materials and Methods).
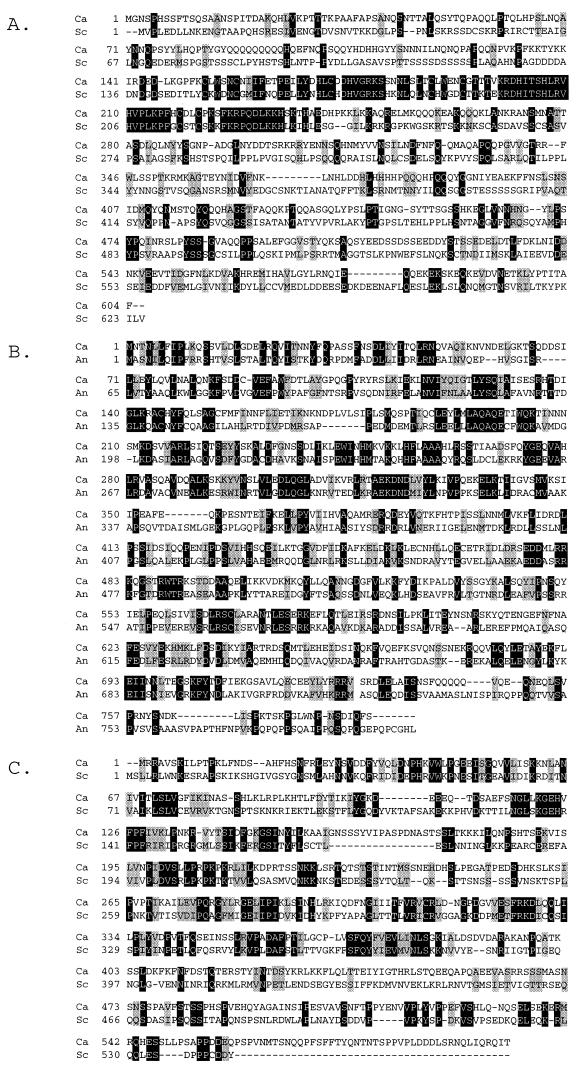
C. albicans RIM101 specifies a protein of 604 amino acids. The Rim101p/PacC family has limited sequence identity over the entire protein sequence; C. albicans and S. cerevisiae Rim101p are approximately 20% identical. However, they do share three structural features (Fig. 2A). First, the zinc finger domain of the Rim101p/PacC family is highly conserved; C. albicans Rim101p has 57% identity and 87% similarity to the other homologs. Second, all members of this family have a 50- to 90-residue D/E-rich C-terminal domain; C. albicans Rim101p has an 84-amino-acid C-terminal region that is 32% D/E residues. Third, these proteins are similar in size, ranging from 585 to 667 amino acids, with the zinc fingers positioned approximately 50 to 150 amino acids from the N terminus. C. albicans Rim101p also has several Q-rich regions, a feature shared with Rim101p from Y. lipolytica (10). This level of structural homology indicates that the sequence we identified specifies a Rim101 homolog.
FIG. 2.
ClustalW alignments of C. albicans Rim101p (A), Rim20p (B), and Rim8p (C) with the S. cerevisiae homolog (A and C) and the A. nidulans homolog (B). Alignments were done by using the server at http://dot.imgen.bcm.tmc.edu:9331/multi-align/Options/clustalw.html. C. albicans RIM101, RIM20, and RIM8 sequences have been deposited in GenBank (see Materials and Methods). S. cerevisiae sequences were obtained from the S. cerevisiae database at http://www-genome.stanford.edu/Saccharomyces/. The A. nidulans sequence was obtained from the GenBank database at http://www.ncbi.nlm.nih.gov/. Ca, C. albicans; Sc, S. cerevisiae; An, A. nidulans.
C. albicans RIM20 specifies a protein of 785 amino acids that is approximately 30% identical and 50% similar to Rim20p/PalA from S. cerevisiae and A. nidulans (Fig. 2B). Unlike the Rim101p/PacC family, the Rim20p/PalA family has homology over the entire molecule. C. albicans Rim20p has homology with a number of signal transduction proteins from higher eukaryotes, including Xp95 from Xenopus laevis, Alix/AIP1 from mice, and a putative Caenorhabditis elegans protein. The level of homology between Rim20p/PalA and C. albicans Rim20p suggests that the sequence we identified specifies a Rim20p homolog.
C. albicans RIM8 specifies a protein of 602 amino acids that is approximately 28% identical and 40% similar to Rim8p/PalF from S. cerevisiae and A. nidulans (Fig. 2C). The Rim8p/PalF family also has homology over the entire molecule. Therefore, the C. albicans RIM8 gene specifies a Rim8p homolog.
RIM101 is required for alkaline-induced filamentation.
We considered the hypothesis that C. albicans RIM101 is required for alkaline responses. To test this possibility, alkaline-induced filamentation was analyzed in RIM101/RIM101 (BWP17), RIM101/rim101− (DAY2), and rim101−/rim101− (DAY5) cells. These strains all grew in the yeast form at acidic pH (Fig. 3A and C and Table 3). RIM101/RIM101 and RIM101/rim101− cells grew primarily in the filament form at alkaline pH (Fig. 3B and Table 3): approximately 65 to 80% of the RIM101/RIM101 and RIM101/rim101− cells produced filaments after 4 h at alkaline pH. However, rim101−/rim101− cells did not produce filaments at alkaline pH after 4 h (Fig. 3D and Table 3) or 36 h (data not shown). Thus, RIM101 is required for alkaline-induced filamentation.
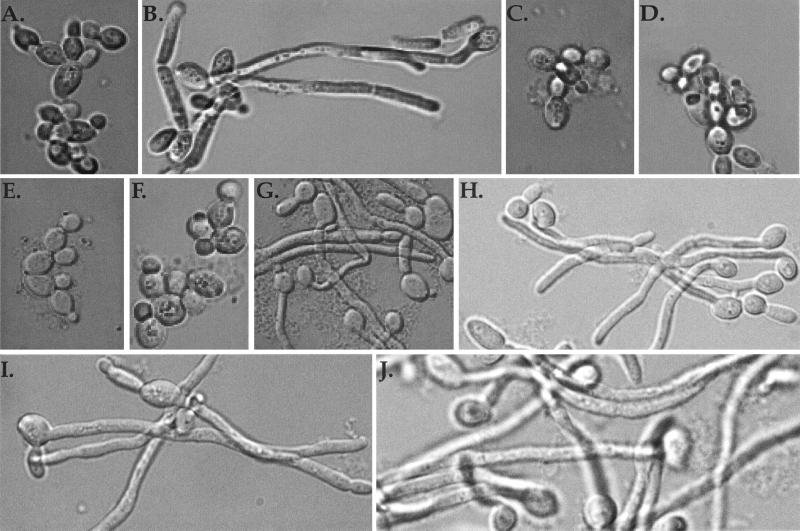
FIG. 3.
Morphology of strains grown at acidic and alkaline pH. The figure shows wild-type cells (A and B) and rim101−/rim101− (C and D), rim20−/rim20− (E), rim8−/rim8− (F), rim101−/rim101−, pRIM101 (G), rim101−/rim101−, pRIM101-405 (H), rim20−/rim20− pRIM101-405 (I), and rim8−/rim8− pRIM101-405 (J) cells grown at acidic (A and C) or alkaline (B and D to J) pH.
TABLE 3.
Filamentation of RIM101 pathway mutantsa
| Strain | Genotype | Plasmidc | pH | % Filamentation |
|---|---|---|---|---|
| BWP17b | RIM101/RIM101 | 4 | <0.25 | |
| BWP17 | RIM101/RIM101 | 8 | 65 | |
| DAY2 | RIM101/rim101− | 4 | <0.5 | |
| DAY2 | RIM101/rim101− | 8 | 80 | |
| DAY5 | rim101−/rim101− | 4 | <0.25 | |
| DAY5 | rim101−/rim101− | 8 | <0.25 | |
| DAY44 | rim101−/rim101− | pRIM101 | 4 | 1 |
| DAY44 | rim101−/rim101− | pRIM101 | 8 | 90 |
| DAY96 | rim101−/rim101− | pRIM101-405 | 4 | <0.5 |
| DAY96 | rim101−/rim101− | pRIM101-405 | 8 | 70 |
| DAY90 | RIM101/RIM101 | pRIM101-405 | 4 | <0.5 |
| DAY90 | RIM101/RIM101 | pRIM101-405 | 8 | 70 |
| DAY23 | rim20−/rim20− | 4 | <0.25 | |
| DAY23 | rim20−/rim20− | 8 | <0.25 | |
| DAY93 | rim20−/rim20− | pRIM101-405 | 4 | 5 |
| DAY93 | rim20−/rim20− | pRIM101-405 | 8 | 70 |
| DAY111 | rim20−/rim20− | pRIM101 | 4 | <0.5 |
| DAY111 | rim20−/rim20− | pRIM101 | 8 | <0.5 |
| DAY61 | rim8−/rim8− | 4 | <0.25 | |
| DAY61 | rim8−/rim8− | 8 | <0.25 | |
| DAY62 | rim8−/rim8− | pRIM101-405 | 4 | 5 |
| DAY62 | rim8−/rim8− | pRIM101-405 | 8 | 70 |
| DAY114 | rim8−/rim8− | pRIM101 | 4 | <0.5 |
| DAY114 | rim8−/rim8− | pRIM101 | 8 | <0.5 |
| BWP17 | RIM101/RIM101 | (Serum) | 90 | |
| DAY5 | rim101−/rim101− | (Serum) | 90 | |
| DAY23 | rim20−/rim20− | (Serum) | 90 | |
| DAY61 | rim8−/rim8− | (Serum) | 80 |
Filamentation of strains in TC199 buffered at pH 4 or pH 8 for 4 h or incubated in serum for 2 h. Percent filamentation is rounded to the nearest 5%. Standard deviations range from 15% for samples that filament >65% to <5% for samples that filament ≤5%.
This strain is wild type for RIM20 and RIM8 as well.
Refers to the relevant genotype of the plasmid integrated into the Candida genome. pRIM101 is pDDB52, and pRIM101-405 is pDDB71.
To determine if the rim101−/rim101− filamentation defect is due to a growth defect, the growth rates of RIM101/RIM101 and rim101−/rim101− cells were measured. These strains grew at comparable rates at both acidic and alkaline pH (Table 4). These results were confirmed by growth tests on plates (data not shown). Therefore, the alkaline-induced filamentation defect of the rim101−/rim101− mutant is not due to an inability to grow at alkaline pH. RIM101 is distinct from other genes important for alkaline responses as it is essential for the response but dispensable for growth (21).
TABLE 4.
Growth rates
| Strain | Genotype | pH | Doubling timea (h) |
|---|---|---|---|
| BWP17 | Wild type | 4 | 1.95 |
| BWP17 | Wild type | 8 | 3.1 |
| DAY5 | rim101−/rim101− | 4 | 1.9 |
| DAY5 | rim101−/rim101− | 8 | 2.3 |
| DAY23 | rim20−/rim20− | 4 | 2.05 |
| DAY23 | rim20−/rim20− | 8 | 2.45 |
| DAY61 | rim8−/rim8− | 4 | 2.05 |
| DAY61 | rim8−/rim8− | 8 | 2.65 |
Average doubling times of strains grown in TC199 medium buffered at pH 4 or pH 8 from two experiments were determined by measurements of the opitcal density at 600 nm (OD600) (values vary by ≤25%). OD600 was determined following shift to fresh TC199 medium and monitored continually over 4 h. These results include the samples used for the Northern blot analyses in Fig. 4 and 5.
Filamentation can be induced by a number of conditions including incubation in serum (16). To determine if RIM101 is required for filamentation under all conditions, the ability of rim101−/rim101− cells to filament in serum was determined. Both RIM101/RIM101 and homozygous rim101−/rim101− cells produce filaments at comparable levels in serum (Table 3). Thus, RIM101 is not essential for filamentation under all conditions.
Because C. albicans lacks a sexual cycle, complementation or suppression tests are required to demonstrate that a defined mutation confers a particular phenotype. Using the remaining his1 marker in our strains, we were able to introduce full-length RIM101 without having to recycle markers. Integration of plasmid pRIM101 into the rim101−/rim101− mutant (DAY44) restored alkaline-induced filamentation ability (Fig. 3G and Table 3). Therefore, the alkaline-induced filamentation defect of the rim101−/rim101− strain is due to loss of Rim101p function.
RIM101 is required for alkaline response gene expression.
C. albicans responds to alkaline pH through changes in morphology and gene expression. Therefore, the expression of two alkaline-induced genes, PHR1 and PRA1, was analyzed in RIM101/RIM101, RIM101/rim101−, and rim101−/rim101− cells. These strains all had no appreciable expression of either PHR1 or PRA1 at acidic pH (Fig. 4, samples 1 to 3). RIM101/RIM101 and RIM101/rim101− cells expressed both PHR1 and PRA1 at alkaline pH (Fig. 4, samples 7 and 8). However, rim101−/rim101− cells did not express either PHR1 or PRA1 at alkaline pH (Fig. 4, sample 9). When these Northern blots were quantitated and normalized for TEF1 mRNA, we found that RIM101/RIM101 and RIM101/rim101− cells express >10-fold-more PHR1 and PRA1 mRNA than the rim101−/rim101− cells (Fig. 5, samples 7 to 9). Complementation of the rim101−/rim101− mutant restored substantial expression of both PHR1 and PRA1 at alkaline pH (Fig. 4 and 5, sample 10). These results demonstrate that RIM101 is required for the expression of these two alkaline-induced genes.
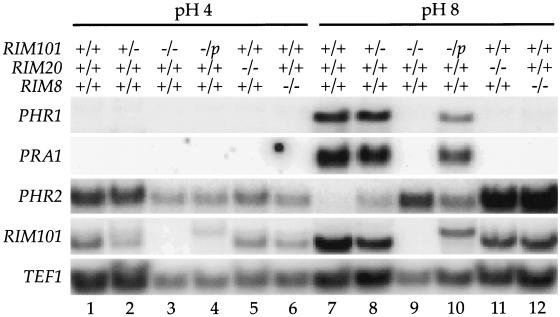
FIG. 4.
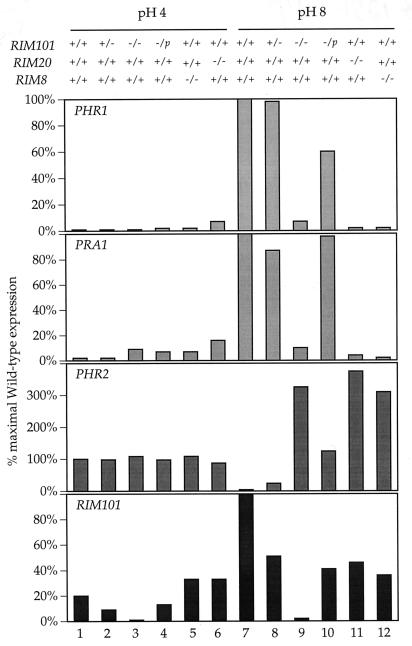
Northern blot analyses of strains grown at acidic and alkaline pH. RNA was prepared from RIM101/RIM101 (samples 1 and 7), RIM101/rim101− (samples 2 and 8), rim101−/rim101− (samples 3 and 9), rim101−/rim101− pRIM101 (samples 4 and 10), rim20−/rim20− (samples 5 and 11), and rim8−/rim8− (samples 6 and 12) cells grown at acidic (samples 1 to 6) or alkaline (samples 7 to 12) pH. Northern blots were visualized with a phosphorimager.
FIG. 5.
Quantitation of Northern blots from Fig. 4. Samples were normalized for loading with the TEF1 signal. Note that the scale of PHR2 has been expanded to permit comparison of maximal wild-type and mutant levels.
PHR1 is required for growth at alkaline pH (21). We observed that rim101−/rim101− cells do not express PHR1 but are able to grow at alkaline pH. Thus, we hypothesized that PHR2, a PHR1 homolog, may be expressed in rim101−/rim101− cells. RIM101/RIM101 cells did not express PHR2 at alkaline pH (Fig. 4, sample 7). However, we found that rim101−/rim101− cells overexpressed PHR2 to levels three- to fourfold higher than maximal wild-type levels (Fig. 4 and 5, samples 1 and 9). Complementation of the rim101−/rim101− mutant did reduce PHR2 expression, although not to wild-type levels (Fig. 5, samples 7, 9, and 10). Thus, PHR2 becomes an alkaline response gene in the absence of Rim101p. This result argues that a RIM101-independent pathway promotes elevated PHR2 expression at alkaline pH.
RIM20 and RIM8 are required for alkaline pH responses.
Two other homologs of the RIM101 pathway, RIM20 and RIM8, have been identified in C. albicans. We assayed the requirement of these genes for pH responses by testing the ability of rim20−/rim20− (DAY23) and rim8−/rim8− (DAY61) cells to form filaments at alkaline pH. Like rim101−/rim101− cells, rim20−/rim20− and rim8−/rim8− cells grew in the yeast form at both acidic pH and alkaline pH (Fig. 3E and F and Table 3). RIM20 and RIM8 are not general regulators of filamentation, as rim20−/rim20− and rim8−/rim8− cells were able to form filaments in serum (Table 3). Thus, RIM20 and RIM8 are required for alkaline-induced filamentation.
We also expected that RIM20 and RIM8 would be required for alkaline response gene expression. We found that rim20−/rim20− and rim8−/rim8− cells did not express PHR1 or PRA1 at either acidic or alkaline pH (Fig. 4, samples 11 and 12), thus suggesting that Rim101p, Rim20p, and Rim8p act in the same pathway. We also found that PHR2 became an alkaline response gene in rim20−/rim20− and rim8−/rim8− cells (Fig. 4 and 5, samples 1, 11, and 12). Thus, the pathway responsible for PHR2 alkaline induction must be independent of Rim20p and Rim8p.
RIM101 is an alkaline-induced gene.
Expression of many transcription factors is regulated by the condition to which they respond (23, 24). Therefore, we determined whether RIM101 expression is regulated in response to alkaline pH. RIM101/RIM101 cells expressed RIM101 at both acidic and alkaline pH (Fig. 4, samples 1 and 7). However, RIM101 expression was elevated fivefold at alkaline pH over acidic pH levels (Fig. 4 and 5, samples 1 and 7). Although complemented rim101−/rim101− pRIM101 cells showed the same fivefold increase in expression at alkaline pH as did the wild type (Fig. 5, samples 4 and 10), we observed that these cells expressed a longer RIM101 mRNA than did the wild type (Fig. 4, samples 1, 4, 7, and 10). We infer that this longer message includes neighboring vector sequences following the stop codon. These results demonstrate that RIM101 is an alkaline-induced gene.
Since RIM101 is an alkaline response gene, its expression may also depend on the RIM101 pathway. To address this possibility, rim20−/rim20− and rim8−/rim8− cells were analyzed for expression of RIM101. rim20−/rim20− and rim8−/rim8− cells expressed RIM101 at both acidic and alkaline pH (Fig. 4). However, both mutants failed to increase RIM101 expression at alkaline pH (Fig. 5, samples 5, 6, 11, and 12). Thus, RIM101 is an alkaline-induced gene that requires RIM20 and RIM8 for increased expression at alkaline pH.
Suppression by truncated Rim101p.
S. cerevisiae Rim101p is activated by proteolytic removal of the D/E-rich C-terminal tail following shift to alkaline pH (11, 20). If the C terminus of C. albicans Rim101p is removed in response to alkaline pH, then a RIM101 allele that lacks the C-terminal D/E-rich domain should be functional. To test this possibility, we created the RIM101-405 allele, which has a stop codon following residue 405, introduced it into rim101−/rim101− cells, and assayed for alkaline-induced filamentation. Although rim101−/rim101− cells grew in the yeast form at alkaline pH (Fig. 3D), rim101−/RIM101-405 cells grew in the filament form at alkaline pH (Fig. 3H and Table 3). These results demonstrate that RIM101-405 can complement the alkaline-induced filamentation defect of rim101−/rim101− cells.
If Rim20p and Rim8p act to stimulate processing of Rim101p, then the RIM101-405 allele should suppress the filamentation defects of the rim20−/rim20− and rim8−/rim8− mutants. Therefore, the pRIM101-405 plasmid was introduced into rim20−/rim20− and rim8−/rim8− cells. Unlike rim20−/rim20− and rim8−/rim8− cells, rim20−/rim20− pRIM101-405 and rim8−/rim8− pRIM101-405 cells grew in the filament form at alkaline pH (Fig. 3I and J and Table 3). The rim20−/rim20− and rim8−/rim8− alkaline-induced filamentation defects were not suppressed by introduction of the wild-type pRIM101 allele (Table 3). These results indicate that Rim20p and Rim8p promote the pH response by processing Rim101p.
In rim20−/rim20− pRIM101-405 and rim8−/rim8− pRIM101-405 strains, Rim101p activity is independent of Rim20p and Rim8p. However, these strains filament weakly at acidic pH compared to alkaline pH (Table 3). Therefore, alkaline pH still induces filamentation independent of upstream RIM101 pathway components, provided that Rim101p is expressed in an active form.
DISCUSSION
The ability of C. albicans to adapt to diverse environments is central to its pathogenesis. Here we have used a streamlined genetic approach to dissect the mechanisms that govern the response of C. albicans to external pH. We identified possible homologs of fungal pH regulatory genes in the Candida genomic database and assessed their functional significance through PCR product-directed gene disruption. The homozygous mutants displayed pH response defects, thus prompting us to retrieve and characterize each entire gene. Finally, because we used a triply marked strain, we carried out complementation and suppression analyses without having to recycle markers. Our findings support two main conclusions (Fig. 6). First, the RIM101 pathway is conserved in C. albicans and governs several alkaline responses. Second, both disruption and bypass mutants of the RIM101 pathway demonstrate the existence of an additional alkaline response pathway.
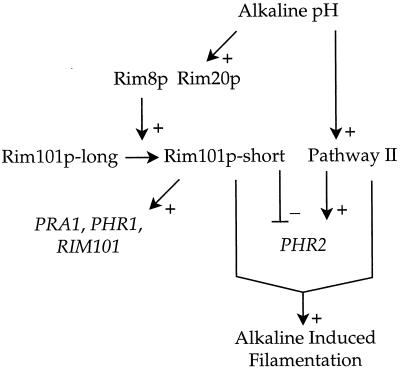
FIG. 6.
Model for RIM101-dependent and RIM101-independent control of alkaline responses. Alkaline pH stimulates Rim101p activity through increased expression and proteolytic activation, both of which require Rim8p and Rim20p. Full-length Rim101p-long does not have a known activity. Processed Rim101p-short is required for the alkaline response, which includes activation of alkaline-induced genes, repression of alkaline-repressed genes, and filamentation. Since RIM101 is an alkaline-induced gene, its expression may depend on autoregulation by Rim101p-short. Alkaline pH also stimulates a RIM101-independent pathway. (We have diagrammed one RIM101-independent pathway for simplicity, but there may be several.) This pathway activates PHR2 expression (in the absence of a functional RIM101 pathway) and stimulates filamentation in conjunction with Rim101p.
The RIM101 pathway plays a distinct role in each of three alkaline responses (Fig. 6). First, Rim101p, Rim8p, and Rim20p are positive regulators of the alkaline-induced genes PRA1, PHR1, and RIM101 itself, because loss of RIM101 pathway function blocks induction of these genes at alkaline pH. Second, these proteins are negative regulators of the alkaline-repressed gene PHR2, because loss of RIM101 pathway function permits expression of PHR2 at alkaline pH. Third, these proteins are positive regulators of alkaline-induced filamentation, because loss of RIM101 pathway function prevents filamentation at alkaline pH. The finding that truncated Rim101-405p suppresses the rim8−/rim8− and rim20−/rim20− alkaline-induced filamentation defects suggests that Rim101p is processed by and acts downstream of Rim8p and Rim20p. These results are expected based on findings from Saccharomyces, Aspergillus, and Yarrowia (10, 11, 20, 25, 26).
However, unlike Aspergillus and Yarrowia, C. albicans remains pH responsive in the absence of the RIM101 pathway. For example, the dominant RIM101-405 allele, which complements the filamentation defect of the rim101−/rim101− mutant, promotes filamentation very weakly at acidic pH. Thus, the uncoupling of Rim101p processing from the upstream regulators does not completely bypass the control of filamentation by external pH. In addition, PHR2 becomes an alkaline-induced gene in cells that lack RIM101 pathway function. Thus, both morphological and gene expression responses point to the existence of a RIM101-independent pH response pathway. This pathway has two roles: to stimulate PHR2 expression at alkaline pH, and to act in conjunction with Rim101p to activate filamentation. We have depicted this pathway as a positively acting pathway (Fig. 6), but other explanations are also possible.
Often a particular organism has a unique balance between the contributions of partially redundant pathways. For example, filamentous growth is activated by both the Ste12p/Cph1p and Phd1p/Efg1p pathways in Saccharomyces and C. albicans, and yet Ste12p/Cph1p is the major pathway in Saccharomyces and Phd1p/Efg1p is the major pathway in C. albicans (8, 12–14). Similarly, Rim101p seems to be the major activator of sporulation in Yarrowia but has roles overlapping with those of Mck1p and Ime4p in Saccharomyces (10, 25). Thus, it seems reasonable that an additional pH response pathway exists in these other fungi and that its unique balance with the RIM101 pathway in C. albicans has simplified its detection.
Rim101p may activate filamentation through a functional interaction with the negative regulator Tup1p or the positive regulators Cph1p and Efg1p. It is intriguing that Rim101p and Tup1p together regulate transcription from the Saccharomyces IME1 and DIT1/DIT2 promoters (2, 7, 17). In fact, regulation of these target genes in Saccharomyces is parallel to the regulation of filamentation in C. albicans: they are positively regulated by Rim101p and negatively regulated by Tup1p. Thus, Rim101p and Tup1p may converge on a common set of filamentation-specific promoters in C. albicans.
ACKNOWLEDGMENTS
We thank Teresa Lamb and Brian Enloe for critical reading of the manuscript and all members of the Mitchell lab for numerous helpful discussions. We gratefully acknowledge Stew Scherer for maintaining an accessible and updated Candida genomic database. Dana Davis wishes to thank Debra McWilliam for continued support throughout the course of this work.
This work was supported by a Mycology Scholar Award from the Burroughs Wellcome Fund (to A.P.M.) and by grants T32 AI07161-21 and PO1 AI377194 from the National Institutes of Health.
REFERENCES
- 1.Adams A, Gotschling D E, Kaiser C A, Stearns T. Methods in yeast genetics. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1997. [Google Scholar]
- 2.Bogengruber E, Eichberger T, Briza P, Dawes I W, Breitenbach M, Schricker R. Sporulation-specific expression of the yeast DIT1/DIT2 promoter is controlled by a newly identified repressor element and the short form of Rim101p. Eur J Biochem. 1998;258:430–436. doi: 10.1046/j.1432-1327.1998.2580430.x. [DOI] [PubMed] [Google Scholar]
- 3.Braun B R, Johnson A D. Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science. 1997;277:105–109. doi: 10.1126/science.277.5322.105. [DOI] [PubMed] [Google Scholar]
- 4.Chaffin W L, Lopez-Ribot J L, Casanova M, Gozalbo D, Martinez J P. Cell wall and secreted proteins of Candida albicans: identification, function, and expression. Microbiol Mol Biol Rev. 1998;62:130–180. doi: 10.1128/mmbr.62.1.130-180.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Denison S H, Negret-Urtasun S, Mingot J M, Tilburn J, Mayer W A, Goel A, Espeso E A, Penalva M A, Arnst H N. Putative membrane components of signal transduction pathways for ambient pH regulation in Aspergillus and meiosis in Saccharomyces are homologous. Mol Microbiol. 1998;30:259–264. doi: 10.1046/j.1365-2958.1998.01058.x. [DOI] [PubMed] [Google Scholar]
- 6.Denison S H, Orejas M, Arnst H N. Signaling of ambient pH in Aspergillus involves a cysteine protease. J Biol Chem. 1995;270:28519–28522. doi: 10.1074/jbc.270.48.28519. [DOI] [PubMed] [Google Scholar]
- 7.Friesen H, Hepworth S, Segall J. An Ssn6-Tup1-dependent negative regulatory element controls sporulation-specific expression of DIT1 and DIT2 in Saccharomyces cerevisiae. Mol Cell Biol. 1997;17:123–134. doi: 10.1128/mcb.17.1.123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gimeno C J, Fink G R. Induction of pseudohyphal growth by overexpression of PHD1, a Saccharomyces cerevisiae gene related to transcriptional regulators of fungal development. Mol Cell Biol. 1994;14:2100–2112. doi: 10.1128/mcb.14.3.2100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kwon-Chung K J, Bennett J E. Medical mycology. 1st ed. Philadelphia, Pa: Lea & Febiger; 1992. [Google Scholar]
- 10.Lambert M, Blanchin-Roland S, Louedec F L, Lepingle A, Gaillardin C. Genetic analysis of regulatory mutants affecting synthesis of extracellular proteinases in the yeast Yarrowia lipolytica: identification of a RIM101/pacC homolog. Mol Cell Biol. 1997;17:3966–3976. doi: 10.1128/mcb.17.7.3966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li W, Mitchell A P. Proteolytic activation of Rim1p, a positive regulator of yeast sporulation and invasive growth. Genetics. 1997;145:63–73. doi: 10.1093/genetics/145.1.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Liu H, Kohler J, Fink G R. Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science. 1994;266:1723–1726. doi: 10.1126/science.7992058. [DOI] [PubMed] [Google Scholar]
- 13.Liu H, Styles C A, Fink G R. Elements of the yeast pheromone response pathway required for filamentous growth of diploids. Science. 1993;262:1741–1744. doi: 10.1126/science.8259520. [DOI] [PubMed] [Google Scholar]
- 14.Lo H-J, Kohler J R, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink G R. Nonfilamentous C. albicans mutants are avirulent. Cell. 1997;90:939–949. doi: 10.1016/s0092-8674(00)80358-x. [DOI] [PubMed] [Google Scholar]
- 15.Maccheroni W, May G S, Martinez-Rossi N M, Rossi A. The sequence of palF, an environmental pH response gene in Aspergillus nidulans. Gene. 1997;194:163–167. doi: 10.1016/s0378-1119(97)00095-4. [DOI] [PubMed] [Google Scholar]
- 16.Mitchell A P. Dimorphism and virulence in Candida albicans. Curr Opin Microbiol. 1998;1:687–692. doi: 10.1016/s1369-5274(98)80116-1. [DOI] [PubMed] [Google Scholar]
- 17.Mizuno T, Nakazawa N, Remgsamrarn P, Kunoh T, Oshima Y, Harashima S. The Tup1-Ssn6 general repressor is involved in repression of IME1 encoding a transcriptional activator of meiosis in Saccharomyces cerevisiae. Curr Genet. 1998;33:239–247. doi: 10.1007/s002940050332. [DOI] [PubMed] [Google Scholar]
- 18.Muhlschlegel F A, Fonzi W A. PHR2 of Candida albicans encodes a functional homolog of the pH-regulated gene PHR1 with an inverted pattern of pH-dependent expression. Mol Cell Biol. 1997;17:5960–5967. doi: 10.1128/mcb.17.10.5960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Odds F C. Candida and candidosis. 2nd ed. London, United Kingdom: Bailliere Tindall; 1988. [Google Scholar]
- 20.Orejas M, Espeso E A, Tilburn J, Sarkar S, Arnst H N, Penalva M A. Activation of the Aspergillus PacC transcription factor in response to alkaline pH requires proteolysis of the carboxy-terminal moiety. Genes Dev. 1995;9:1622–1632. doi: 10.1101/gad.9.13.1622. [DOI] [PubMed] [Google Scholar]
- 21.Saporito-Irwin S M, Birse C E, Sypherd P S, Fonzi W A. PHR1, a pH-regulated gene of Candida albicans, is required for morphogenesis. Mol Cell Biol. 1995;15:601–613. doi: 10.1128/mcb.15.2.601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sentandreu M, Elorza M V, Sentandreu R, Fonzi W A. Cloning and characterization of PRA1, a gene encoding a novel pH-regulated antigen of Candida albicans. J Bacteriol. 1998;180:282–289. doi: 10.1128/jb.180.2.282-289.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Smith H E, Driscoll S E, Sia R A L, Yuan H E, Mitchell A P. Genetic evidence for transcriptional activation by the yeast IME1 gene product. Genetics. 1993;133:775–784. doi: 10.1093/genetics/133.4.775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Smith H E, Su S Y, Neigeborn L, Driscoll S E, Mitchell A P. Role of IME1 expression in regulation of meiosis in Saccharomyces cerevisiae. Mol Cell Biol. 1990;10:6103–6113. doi: 10.1128/mcb.10.12.6103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Su S S Y, Mitchell A P. Molecular characterization of the yeast meiotic regulatory gene RIM1. Nucleic Acids Res. 1993;21:3789–3797. doi: 10.1093/nar/21.16.3789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tilburn J, Sarkar S, Widdick D A, Espeso E A, Orejas M, Mungroo J, Penalva M A, Arnst H N., Jr The Aspergillus PacC zinc finger transcription factor mediates regulation of both acid- and alkaline-expressed genes by ambient pH. EMBO J. 1995;14:779–790. doi: 10.1002/j.1460-2075.1995.tb07056.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wilson R B, Davis D, Mitchell A P. Rapid hypothesis testing in Candida albicans through gene disruption with short homology regions. J Bacteriol. 1999;181:1868–1874. doi: 10.1128/jb.181.6.1868-1874.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]