Abstract
Background
Channa argus and Channa maculata are the main cultured species of the snakehead fish family, Channidae. The relationship between them is close enough that they can mate; however, their temperature adaptability is quite different.
Results
In this study, we sequenced and assembled the whole genomes of C. argus and C. maculata and obtained chromosome-level genome assemblies of 630.39 and 618.82 Mb, respectively. Contig N50 was 13.20 and 21.73 Mb, and scaffold N50 was 27.66 and 28.37 Mb, with 28,054 and 24,115 coding genes annotated for C. argus and C. maculata, respectively. Our analyses showed that C. argus and C. maculata have 24 and 21 chromosomes, respectively. Three pairs of chromosomes in C. argus correspond to 3 chromosomes in C. maculata, suggesting that 3 chromosomal fusion events occurred in C. maculata. Comparative analysis of their gene families showed that some immune-related genes were unique or expandable to C. maculata, such as genes related to herpes simplex infection. Analysis of the transcriptome differences related to temperature adaptation revealed that the brain and liver of C. argus rapidly produced more differentially expressed genes than C. maculata. Genes in the FoxO signalling pathway were significantly enriched in C. argus during the cooling process (P < 0.05), and the expression of 3 transcription factor genes in this pathway was significantly different between C. argus and C. maculata (P < 0.01).
Conclusions
C. maculata may have higher resistance to certain diseases, whereas C. argus has a faster and stronger response to low-temperature stress and thus has better adaptability to a low-temperature environment. This study provides a high-quality genome research platform for follow-up studies of Channidae and provides important clues regarding differences in the low-temperature adaptations of fish.
Keywords: Channa argus, Channa maculata, genome, transcriptome, low-temperature adaptation
Background
Channa argus (the northern snakehead, NCBI:txid215402, Fishbase ID: 4799) and Channa maculata (the blotched snakehead, NCBI:txid188791, Fishbase ID: 8701) are the main cultured species of the snakehead fish family Channidae [1, 2]. In 2019, the output of the snakehead fish in China reached 460,000 tonnes [3]. Snakehead fish are a serious invasive species in North America owing to their ability to rapidly colonize waterways. These fish have a specialized aerial breathing organ, the suprabranchial chamber, which facilitates aquatic-aerial bimodal breathing. This enables them to migrate short distances over land and makes them a good model for research on bimodal breathing [4, 5].
C. argus is widely distributed from China, India, and Southeast Asia to the far east of Russia, North Korea, Japan, and other major water systems and has high cold resistance. In contrast, C. maculata is distributed in warm water systems in China, the Philippines, Vietnam, Madagascar, the United States, Japan, and other places and has low cold resistance [2]. The ability of fish to adapt to environmental temperature differs as a result of long-term adaptation and evolution, as well as the specific expression of genetic information. The physiological responses of fish to low-temperature exposure have been extensively studied [6], and there is increasing interest in elucidating the mechanisms of fish adaptation to low-temperature environments and tolerance to low-temperature stress at the molecular level [7]. The decreasing costs of high-throughput sequencing and the application of bioinformatics technology have allowed researchers to use omics methods to analyse the molecular mechanisms and signalling pathways of fish responding to low-temperature stress at the overall biological level and explore the functional genes involved in low-temperature tolerance [8].
The cold tolerance of fish is an important economic characteristic of the breed and is related to its growth cycle and extension range [2]. Although the draft genome of C. argus has been previously described [9], to better understand the reasons for the difference in cold tolerance between C. argus and C. maculata, whole-genome sequencing and assembly of both species were carried out in this study. After obtaining chromosome-level genomic sequences of these 2 species, we analysed the transcriptome differences related to temperature adaptation between C. argus and C. maculata, providing clues for research on the low-temperature adaptation of fish.
Data Description
Source of experimental fish and preparation of DNA
A female C. argus and a female C. maculata, which were provided by the Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, were dissected to obtain muscle tissue and immediately frozen in liquid nitrogen for storage. The cetyltrimethylammonium bromide method was used to extract DNA from the muscle tissue, and 1% agarose gel electrophoresis and Qubit 3.0 (Thermo Fisher Scientific, Inc., Waltham, MA, USA) were used to detect the quality and concentration of the extracted DNA.
Before the dissection of the experimental fish, the fish were anaesthetized with ethyl 3-aminobenzoate methanesulfonate. The experimental protocol of this study was approved by the Animal Ethics Committee of the Institute of Hydrobiology, Chinese Academy of Sciences (reference No.: Y81F101).
Illumina sequencing and genome survey
Two 350-bp libraries were constructed using the C. argus and C. maculata muscle tissue DNA, and paired-end 150 bp (PE 150) sequencing was performed on the Illumina NovaSeq 6000 platform (Illumina NovaSeq 6000 Sequencing System, RRID:SCR_016387). The experiments were performed according to the standard protocol provided by Illumina. After the raw data were obtained, 62.90 and 63.90 Gb clean data of C. argus and C. maculata were obtained by routine filtering. Two k-mer distribution maps with k = 19 were constructed on the basis of clean data using jellyfish v2.1.4 (Jellyfish, RRID:SCR_005491) (Supplementary File 1: Figure). Based on the distribution of k-mers in C. argus and C. maculata, it was estimated that the content of repeated sequences was 18.73% and 18.23%, and the heterozygosity was 0.12% and 0.06% using genomescope v1.00 (GenomeScope, RRID:SCR_017014) [10], respectively. A total of 49,571,777,400 and 48,531,014,793 k-mers of C. argus and C. maculata were used for genome length estimation, and the calculated genome lengths were 658.63 and 652.03 Mb (the formula is k-mer number/average k-mer depth), respectively. In addition, according to the sequencing data analysis, the GC contents of C. argus and C. maculata genomes were 40.36% and 40.37%, respectively. These 2 genomes could be assembled directly because of their compact size, low heterozygosity, and complexity.
Nanopore sequencing and initial assembly
Two Oxford Nanopore (Oxford Nanopore Technologies, RRID:SCR_003756) long-read libraries were constructed using C. argus and C. maculata muscle tissue DNA and sequenced on the Nanopore platform. The process was performed using the Ligation Sequencing Kit (catalog number SQK-LSK109; Oxford Nanopore Technologies, Oxford, UK). After filtering out low-quality reads and removing the adapters, 118.24 and 101.34 Gb of clean data were obtained for C. argus and C. maculata, respectively. The total sequencing depth was 187.57× and 163.76×, the N50 reads were 38.83 and 40.48 kb, and the average read length was 26.59 and 28.11 kb, respectively. Using Canu v1.9 (Canu, RRID:SCR_015880) [11], the clean data were corrected and then assembled using WTDBG v1.2.8 (WTDBG, RRID:SCR_017225) [12], then corrected again with the Nanopore and Illumina sequencing data using Racon (Racon, RRID:SCR_017642) [13] and Pilon v1.23 (Pilon, RRID:SCR_014731) [14], respectively. Finally, the initial assembled genome sequence of C. argus and C. maculata had a total length of 630.38 and 618.82 Mb, and contig N50 of 21.50 and 23.25 Mb, respectively. Using BWA (BWA, RRID:SCR_010910) [15] to align the Illumina sequencing data with the initial assembled genome, the matching rates were 98.17% and 98.34% (Supplementary File 2: Table). BUSCO v4.0.6 (BUSCO, RRID:SCR_015008) [16] was used to evaluate the integrity of 3,354 conserved core genes in the initial assembled genome, accounting for 93.65% and 97.14%, respectively (Supplementary File 3: Table), indicating that the initial assemblies were useful.
Super assembly based on Hi-C technology
After fixing and cross-linking the C. argus and C. maculata muscle tissues with formaldehyde, 2 Hi-C libraries of 300–700 bp were constructed according to the methodology described by Rao et al. [17]. After the libraries were qualified, high-throughput sequencing was performed using an Illumina NovaSeq 6000 with PE150. Raw data were filtered to remove low-quality reads and adapters, and 102.43 and 103.13 Gb clean data for C. argus and C. maculata, respectively, were obtained. After aligning the clean data with the initial genome assembly, using HiC-Pro v2.11.1 (HiC-Pro, RRID:SCR_017643) [18] to filter the alignment results, 146,400,814 and 151,732,929 valid interaction pairs were obtained. Based on the valid interaction pairs, the initial genome assemblies were further assembled using LACHESIS (LACHESIS, RRID:SCR_017644) [19], including grouping, sorting, and orientation of the initial assembled sequences. Finally, the genome sequences with total lengths of 619.41 and 616.63 Mb were attached to the 24 and 21 chromosomes [20], accounting for 98.26% and 99.65% (619,407,135/630,381,055 and 616,629,265/618,815,250), and the numbers of corresponding sequences were 293 and 227, respectively (Table 1).
Table 1:
Summary: statistics of the reference genome assemblies of C. argus and C. maculata
| Species | Assembly | Contig No. | Contig length (bp) | Scaffold No. | Scaffold length (bp) |
|---|---|---|---|---|---|
| C. argus | N50 | 15 | 13,290,021 | 11 | 27,662,632 |
| N90 | 60 | 1,903,525 | 22 | 13,584,876 | |
| Max | 28,029,688 | 50,138,606 | |||
| Total | 607 | 630,381,055 | 521 | 630,389,655 | |
| Anchored to chromosomes | 293 | 619,407,135 (98.26%) | |||
| C. maculata | N50 | 13 | 21,727,292 | 9 | 28,367,461 |
| N90 | 44 | 2,420,044 | 19 | 21,794,094 | |
| Max | 26,519,478 | 49,937,344 | |||
| Total | 338 | 618,815,250 | 254 | 618,823,650 | |
| Anchored to chromosomes | 227 | 616,629,265 (99.65%) |
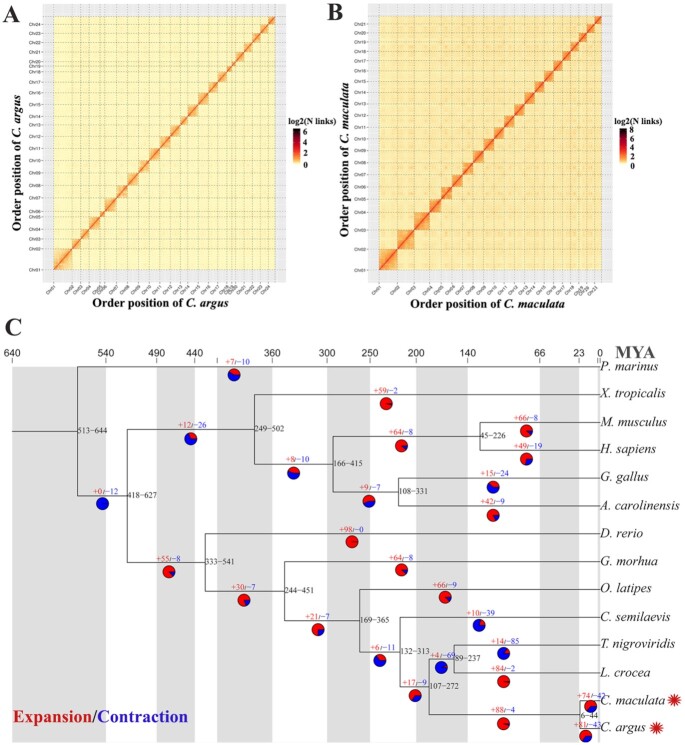
Chromosome-level genomes were cut into 100-kb bins of equal length, and the number of Hi-C read pairs covering any 2 bins was used as the signal of the interaction between the 2 bins. Two heat maps were drawn to evaluate assembly quality (Fig. 1A and B). The image signal distinguished the 24 and 21 chromosome groups, and the intensity of the interaction at the diagonal position on each chromosome was higher than that at the off-diagonal position, indicating that the assembly effect of the chromosomes was strong.
Figure 1:
Genome assembly and evolutionary analysis of C. argus and C. maculata. Genome-wide Hi-C heat maps of C. argus (A) and C. maculata (B). Chr 1–24 and Chr 1–21 refer to chromosome 1–24 and chromosome 1–21. (C) Evolutionary tree including C. argus and C. maculata. The black number at each branch represents the divergence time supported by 95% of the highest posterior density (HPD). The top of the tree is absolute age, separated by the shadow of each geological period. The number on the branch shows the number of expanded (red) and contracted (blue) gene families for each clade. The 2 red asterisks indicate C. argus and C. maculata.
Annotation of repetitive sequences, coding genes, and non-coding RNA
Using LTR_FINDER (LTR_Finder, RRID:SCR_015247) [21] and RepeatScout v1.0.5 (RepeatScout, RRID:SCR_014653) [22], 2 repetitive sequence databases of the genomes were constructed on the basis of the principles of structure prediction and de novo prediction. PASTEClassifier (PASTEClassifier, RRID:SCR_017645) [23] was used to classify the databases, which were then merged with the Repbase (Repbase, RRID:SCR_021169) database [24] as the final repetitive sequence databases. RepeatMasker v4.0.9 (RepeatMasker, RRID:SCR_012954) [25] was then used to predict the repetitive sequences of the genomes based on the constructed repetitive sequence database, with repetitive sequences of 117.49 and 118.99 Mb obtained from C. argus and C. maculata, respectively (Supplementary File 4: Table).
Genscan (GENSCAN, RRID:SCR_013362) [26], Augustus v2.4 (Augustus, RRID:SCR_008417) [27], GlimmerHMM v3.0.4 (GlimmerHMM, RRID:SCR_002654) [28], GeneID v1.4 (Entrez Gene, RRID:SCR_002473) [29], and SNAP v2006-07-28 (SNAP—SNP Annotation and Proxy Search, RRID:SCR_002127) [30] were used for de novo prediction of coding genes. GeMoMa v1.3.1 (GeMoMa, RRID:SCR_017646) [31, 32] was used for predictions based on homologous species. Hisat v2.0.4 (HISAT2, RRID:SCR_015530) [33] and Stringtie v1.2.3 (StringTie, RRID:SCR_016323) [34] were used to assemble transcripts with reference sequences, and TransDecoder v2.0 (TransDecoder, RRID:SCR_017647) [35] and GeneMarkS-T v5.1 (GeneMarkS-T, RRID:SCR_017648) [36] were used to perform gene prediction. PASA v2.0.2 (PASA, RRID:SCR_014656) [37] was used to predict unigene sequences based on transcriptome data without reference sequences. EVM v1.1.1 (EVidenceModeler, RRID:SCR_014659) [38] was used to integrate the prediction results obtained from the above methods and was modified with PASA v2.0.2. Finally, 28,054 and 24,115 coding genes in C. argus and C. maculata were predicted (Supplementary File 5: Table). The number of genes supported by the 3 prediction methods ab initio, homology, and RNAseq was 20,544 and 19,990, accounting for 73.23% (20,544/28,054) and 82.89% (19,990/24,115) for C. argus and C. maculata, respectively.
Different strategies have been used to predict different non-coding RNAs based on the structural characteristics of different non-coding RNAs. Using the Rfam (Rfam, RRID:SCR_007891) database [39], BLASTN (BLASTN, RRID:SCR_001598) was used to perform genome-wide alignment to identify microRNAs and ribosomal RNAs. Transfer RNAs (tRNAs) were identified using tRNAscan-SE v2.0 (tRNAscan-SE, RRID:SCR_010835) [40]. Finally, a total of 554 and 247 microRNAs, 1,136 and 633 ribosomal RNAs, and 4,172 and 1,784 tRNAs were predicted in C. argus and C. maculata, respectively (Supplementary File 6: Table).
Whole-genome evolutionary analysis
The genome data of 14 vertebrate species spanning amphibians to mammals with different evolutionary relationships (including C. argus and C. maculata) were compared. Using Orthofinder v2.3.7 (OrthoFinder, RRID:SCR_017118) [41], the protein sequences of these 14 species were classified into families, and the PANTHER v15 (PANTHER, RRID:SCR_004869) database [42] was used to annotate the obtained gene families. A total of 30,269 families were obtained, of which 1,023 were single-copy gene families. A total of 858 families were unique to C. argus, and 46 families were unique to C. maculata (Supplementary File 7: Table). Using the 1,023 single-copy gene families and IQ-TREE v1.6.11 (IQ-TREE, RRID:SCR_017254) [43], an evolutionary tree was constructed using the maximum likelihood (ML) method with the number of bootstraps set to 1,000 and the outgroup set to Petromyzon marinus. PAML v4.9i (PAML, RRID:SCR_014932) [44] was used to calculate the divergence time, and MCMCtreeR v1.1 (PAML, RRID:SCR_014932) [45] was used for the evolutionary tree display (Fig. 1C). The genetic relationship between C. argus and C. maculata, belonging to Perciformes, was the closest, and the differentiation time was 6–44 million years ago (MYA).
On the basis of the phylogenetic tree showing divergence time and the results of gene family clustering, the number of ancestral gene family members of each branch was estimated using CAFE v4.2 (CAFE, RRID:SCR_005983) [46], so as to predict the expansion and contraction of the gene family relative to its ancestors (P < 0.05) (Supplementary File 8: Table). The results showed that there were 81 expanded gene families, including 606 genes, and 43 contracted gene families, including 95 genes in C. argus, while C. maculata contained 74 expanded gene families, including 721 genes, and 42 contracted gene families, including 8 genes. GO and KEGG enrichment analyses were performed using clusterProfile v3.5.1 (clusterProfiler, RRID:SCR_016884) (Fig. 2A, Supplementary File 9: Figure). The results showed that there were specific immune pathway–related genes in C. maculata compared to C. argus, such as the genes involved in the intestinal immune network for lgA production and the genes related to the herpes simplex infection pathway. In addition, the members of the herpes simplex infection gene family in C. maculata showed significant expansion (P < 0.05).
Figure 2:

Comparative analysis of the C. argus and the C. maculata genomes. (A) KEGG enrichment analysis of the unique, expansion, and contraction gene families. The ordinate is KEGG terms, the abscissa is the number of genes in the pathway, and the colour represents the corresponding P-value. Left, enrichment result for C. argus; right, enrichment result for C. maculata, same asterisks indicate same terms. (B) There was a high collinearity between the 2 species. Chr 2 and 3 of C. argus correspond to Chr 2 of C. maculata, Chr 4 and 5 of C. argus correspond to Chr 3 of C. maculata, Chr 18 and 19 of C. argus correspond to Chr 16 of C. maculata.
Using BLASTP (BLASTP, RRID:SCR_001010) to compare the gene protein sequences of these 2 species, the genes in all collinearity blocks were obtained, and the collinearity map of the coding genes of C. argus and C. maculata was drawn using MCScanX [47] (Fig. 2B). Chr 2 and 3 of C. argus correspond to Chr 2 of C. maculata, Chr 4 and 5 of C. argus correspond to Chr 3 of C. maculata, and Chr 18 and 19 of C. argus correspond to Chr 16 of C. maculata. Using the 24 chromosomes of C. argus as a reference, the Hi-C data of C. argus and C. maculata were mapped to it, and the mapping results confirmed the structural differences (Fig. 3).
Figure 3:
Verification of chromosome structure differences between C. argus and C. maculata genomes. (A) Complete collinearity map. (B) Partial collinearity map showing only the chromosomes with structural differences. (C) C. argus chromosomes were set as the reference sequence to which the Hi-C data of C. argus and C. maculata were mapped.
Low-temperature stress and transcriptome sequencing
A total of 180 C. argus and C. maculata specimens (aged 2 months), weighing 86 ± 17 and 56 ± 9 g, were placed in 2 barrels of 700-L capacity, 90 in each barrel. One group was monitored for observation and statistical mortality, while the other was used to collect materials. The fish were kept at 31°C for 2 weeks. Subsequently, the circulating water-cooling device was connected, and the temperature gradually decreased (Supplementary File 10: Table). During this process, the status and mortality of C. argus and C. maculata were recorded daily (Supplementary File 10: Table), and a cumulative mortality map was drawn (Fig. 4). C. argus began to die at 7°C, and 34 died at 7–2°C, with a mortality rate of 38% (34/90). No deaths occurred in the following 3 days. C. maculata began to die at 8°C, peaking at 7°C, and all specimens died at 8–4°C, with a mortality rate of 100% (Fig. 4A). Three C. argus and C. maculata were randomly selected before cooling (31°C), and brain and liver tissues were collected from each fish. During the cooling period, samples were collected after the temperature was maintained at 16°C for 24 h, and again at 10°C, 8°C, 6°C, and 4°C. Sampling took place before 8:00 (before cooling) on the day, with brain and liver tissue from 3 specimens for each species collected at each time point.
Figure 4:

Low-temperature experiment and transcriptome sequencing of C. argus and C. maculata. (A) Cumulative mortality of C. argus and C. maculata during cooling. Abscissa represents temperature and ordinate represents cumulative mortality. (B) Principal component analysis (PCA) of expression genes in brain and liver at different temperatures. Coordinates are the first 3 principal components PC1, PC2, and PC3 of PCA, and the scale value represents the contribution of the sample to the principal component.
After completion of the low-temperature stress, 72 tissue samples (6 time points, 3 C. argus and 3 C. maculata, 2 tissues per fish) were collected for transcriptome sequencing (PE 150). The sequencing platform was an Illumina NovaSeq 6000, and each sample produced ≥6 Gb of clean data.
Statistical analysis of transcriptional sequencing data and expressed genes
The data obtained from each tissue are shown in Supplementary File 11: Table. Using hisat2 (HISAT2, RRID:SCR_015530) [48], clean reads from each tissue were aligned with the genomes of C. argus and C. maculata. After the initial treatment of the gene count matrix by rlogTransformation of DEseq2 (DESeq2, RRID:SCR_015687) [49], the gene expression density map of the normalized genes showed that the gene expression in brain and liver tissues of C. argus and C. maculata was negative binomial (Supplementary File 12: Figure A and Supplementary File 13: Figure A).
The transcripts per million (TPM) of each gene were calculated, and the genes of TPM > 1 in all samples were counted. Based on this, we drew the box line and cluster diagrams, and a PCA map of tissue expression to analyse the overall expression of the genes and the correlations between tissues (Supplementary File 12: Figure B, C, Supplementary File 13: Figure B, C, and Fig. 4B). The box line diagram showed that the number of genes detected in the brain tissue of C. argus and C. maculata was obviously higher than that in the liver (Supplementary File 12: Figure B and Supplementary File 13: Figure B). PCA and cluster analysis showed that the difference between the brain and liver in C. argus and C. maculata was the most significant variable (∼75%) in gene expression, and the change in temperature was the second largest variable in PCA, accounting for 4%–6% of the total variable.
Differential expression analysis of genes
The number of differentially expressed genes (DEGs) with log fold-change ≥1 at each time point was counted, with the gene expression level at the control temperature (31°C) set as the baseline control (Fig. 5). As the temperature decreased, the number of DEGs in the brain and liver of C. argus increased rapidly. At 16°C, genes in the brain and liver were obviously upregulated and downregulated. At 4°C, the number of DEGs in the brain began to decrease, while the opposite response was observed in the liver. In C. maculata, the number of DEGs in the brain increased considerably at 10°C. At 8°C, the number of DEGs in the brain of C. maculata suddenly decreased to the same level as that observed at 16°C, which may be related to the phenotypic characteristics of death and massive shock that occurred at 8°C (Supplementary File 10: Table).
Figure 5:
Number of DEGs in brain and liver of C. argus (A) and C. maculata (B) during cooling. The abscissa represents temperature and the ordinate represents the number of genes. Red indicates up-regulated genes and blue indicates down-regulated genes. FC: fold change.
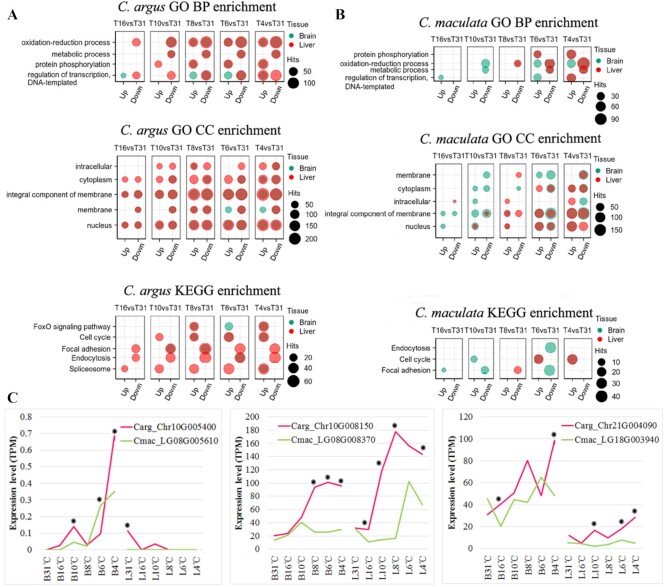
DEG enrichment at each time point was assessed by GO and KEGG analyses, and the top 5 significantly enriched items (P < 0.05) were selected for illustration. It was found that the functions of DEGs were mainly involved in oxidation-reduction processes, metabolic processes, protein phosphorylation, and the pathways mainly involved the FoxO signalling pathway, cell cycle, focal adhesions, and so forth (Fig. 6A and B). We noticed that the FoxO signalling pathway only appeared in the top 5 items for C. argus. The FoxO signalling pathway is a transcription factor–related signalling pathway (Fig. 6A). We selected all 88 genes enriched in the FoxO signalling pathway in C. argus, and iTAK [50] predicted that 10 of these were transcription factors. Based on the collinear relationship of genes between C. argus and C. maculata, we identified 10 corresponding genes in C. maculatus (Supplementary File 14: Table). Transcriptome data were used to analyse the expression changes of 10 transcription factor genes during the cooling process, and it was found that 3 showed significant differences between C. argus and C. maculata (P < 0.01) (Fig. 6C). It is speculated that these genes may be involved in the regulation of cold tolerance traits in C. argus.
Figure 6:
GO and KEGG enrichment analysis of DEGs. The genes with noticeable differences between C. argus and C. maculata were selected for display. (A) Enrichment results for C. argus (green for brain, red for liver). The area of the circle indicates the number of genes. (B) Enrichment results for C. maculata. (C) The expression of 3 transcription factor genes in C. argus and C. maculata. The abscissa represents the tissue samples at different temperatures, and the ordinate represents the expression quantity. The asterisk indicates a significant difference (P < 0.01).
Conclusion
In this study, we sequenced the whole genome of 2 Channidae fish, C. argus and C. maculata, and assembled genome sequences at the chromosome level, which can provide a high-quality genome research platform for follow-up research. The contig N50 was 13.20 and 21.73 Mb, and the scaffold N50 was 27.66 and 28.37 Mb for C. argus and C. maculata, respectively. Compared with the previously published draft genome of C. argus, which had a contig N50 of 81.4 kb and a scaffold N50 of 4.5 Mb [9], the quality of the genomes obtained in this study represents a substantial improvement.
Genome comparison analysis revealed that C. maculata contains genes involved in the intestinal immune network for lgA production and the herpes simplex infection pathway that are not present in C. argus. In addition, members of the herpes simplex infection gene family also have a significant expansion in C. maculata. Compared with C. argus, C. maculata may have higher resistance to disease, especially herpes simplex infection.
There are 3 pairs of chromosomes in C. argus that correspond to 3 chromosomes in C. maculata. The median number of chromosomes in fish with known chromosome number is 24 [51–53]. Therefore, we speculate that these chromosomes in C. maculata fused, while those in C. argus did not.
This study carried out transcriptome analysis to analyse why the cold tolerance of C. argus is better than that of C. maculata. It is found that both C. argus and C. maculata had obvious up-regulation and down-regulation responses in oxidation-reduction processes, metabolic processes, protein phosphorylation, and other pathways, representing the core molecular response to low-temperature exposure. However, a key difference was that the brain and liver of C. argus quickly produced more DEGs, indicating that the response of C. argus to low temperature was faster and stronger than that of C. maculata. In many organisms, transcriptional regulation is a direct response to cold environments. Cold-adapted fish rely on special strategies to acclimate to cold conditions, such as protein biosynthesis, energy metabolism, immune system, lipid metabolism, and signalling pathways, and these strategies have been proven to be species-specific [54]. The FoxO transcription factor–related signalling pathway was significantly enriched in C. argus (P < 0.05) (Fig. 6A). The FoxO family of transcription factors regulates the expression of genes involved in cellular physiological events including apoptosis, cell cycle control, glucose metabolism, oxidative stress resistance, and longevity [55]. Three genes in this pathway showed significant differential expression between C. argus and C. maculata (Fig. 6C), and their function in low-temperature adaptation requires further accurate verification and analysis.
Data Availability
Genome, annotation files, and raw sequences for genome assembly including Illumina, Nanopore, and Hi-C reads of C. argus were deposited in the NCBI and can be accessed with accession No. PRJNA731586, and the corresponding data of C. maculata can be accessed with accession No. PRJNA730430. The transcriptome data related to temperature adaptation of C. argus and C. maculata can be accessed with accession No. PRJNA732763. Supporting data and materials are available in the GigaDB database [56], with individual datasets for C. argus [57] and C. maculata [58].
Additional Files
Supplementary File 1: Figure. k-mer distribution of reads of C. argus (A) and C. maculata (B). k-mers (k = 19) were extracted from the paired-end library with an insert size of 350 bp. The peak 19-mer depths were 76 (A) and 75 (B), respectively.
Supplementary File 2: Table. Mapping rates of the Illumina sequencing data.
Supplementary File 3:Table. Integrity of 3.354 conserved core genes.
Supplementary File 4: Table. Annotation of repetitive sequences.
Supplementary File 5: Table. Annotation of coding genes.
Supplementary File 6: Table. Annotation of non-coding RNA.
Supplementary File 7: Table. Single-copy genes and specific genes in C. argus and C. maculata.
Supplementary File 8: Table. Gene family statistics for expansion and contraction.
Supplementary File 9: Figure. GO enrichment analysis of genes in expansion/contraction families. (A, B) Results for C. argus. (C, D) Results for C. maculata. The abscissa represents the GO terms, and the ordinate represents the number and percentage of genes. Ten GO terms with the most significant enrichment were selected and displayed.
Supplementary File 10: Table. Status and mortality of C. argus and C. maculata during cooling.
Supplementary File 11: Table. Statistical data of transcriptome sequencing.
Supplementary File 12: Figure. Preliminary analysis of C. argus sequencing data. (A) Gene expression in the brain and liver showed a negative binomial distribution. The abscissa represents the log2 value of the amount of gene expression, and the ordinate represents the percentage. (B) The box line diagram shows that the number of genes detected in the brain was higher than that in the liver. The abscissa represents the tissue and the ordinate represents the number of genes. (C) Cluster diagram of brain and liver at different temperatures. Different font colours indicate different temperatures.
Supplementary File 13: Figure. Preliminary analysis of C. maculata sequencing data. (A) Gene expression in the brain and liver showed a negative binomial distribution. The abscissa represents the log2 value of the amount of gene expression, and the ordinate represents the percentage. (B) The box line diagram shows that the number of genes detected in the brain was higher than that in the liver. The abscissa represents the tissue and the ordinate represents the number of genes. (C) Cluster diagram of brain and liver at different temperatures. Different font colours indicate different temperatures.
Supplementary File 14: Table. Ten transcription factor genes in the FoxO signalling pathway in C. argus and C. maculata.
Abbreviations
BLAST: Basic Local Alignment Search Tool; BWA: Burrows-Wheeler Aligner; DEGs: differentially expressed genes; Gb: gigabase pairs; GC: guanine cytosine; GO: gene ontogeny; kb: kilobase pairs; KEGG: Kyoto Encyclopedia of Genes and Genomes; Mb: megabase pairs; ML: maximum likelihood; MYA: million years ago; NCBI: National Center for Biotechnology Information; PCA: principal component analysis; PE: paired end; SRA: Sequence Read Archive; TPM: transcripts per million.
Competing Interests
The authors declare that they have no competing interests.
Funding
This work was supported by the National Key Research & Development Program of China(2018YFD0901201) and the State of Key Laboratory of Freshwater Ecology and Biotechnology (2019FBZ05).
Authors’ Contributions
K.C. and Y.W. conceived and designed the experiments. M.O., R.H., and B.G. performed the experiments. C.Y., Q.L., J.Z., and L.L. analysed the genome and transcriptome data. R.H., M.O., and Y.L. drafted the manuscript. Y.W. and Z.Z. provided advice on manuscript writing. All authors reviewed and declare that they have no conflict of interest.
Supplementary Material
Nicolas Rohner -- 6/18/2021 Reviewed
Nansheng (Jack) Chen -- 7/7/2021 Reviewed
Contributor Information
Mi Ou, Key Laboratory of Tropical and Subtropical Fishery Resources Application and Cultivation, Ministry of Agriculture, Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Guangzhou 510380, China.
Rong Huang, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Cheng Yang, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Bin Gui, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Qing Luo, Key Laboratory of Tropical and Subtropical Fishery Resources Application and Cultivation, Ministry of Agriculture, Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Guangzhou 510380, China.
Jian Zhao, Key Laboratory of Tropical and Subtropical Fishery Resources Application and Cultivation, Ministry of Agriculture, Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Guangzhou 510380, China.
Yongming Li, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Lanjie Liao, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Zuoyan Zhu, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China.
Yaping Wang, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China; Innovative Academy of Seed Design, Chinese Academy of Sciences, Beijing 100101, China.
Kunci Chen, Key Laboratory of Tropical and Subtropical Fishery Resources Application and Cultivation, Ministry of Agriculture, Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Guangzhou 510380, China.
References
- 1. Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Shanghai Fisheries University et al.. Freshwater Fishes of Guangdong. Guangzhou: Guangdong Science and Technology Press; 1990:511–4. [Google Scholar]
- 2. Ou M, Zhao J, Luo Q, et al. Characteristics of hybrids derived from Channa argus ♀ × Channa maculata ♂. Aquaculture. 2018;492:349–56. [Google Scholar]
- 3. Fishery Administration of Ministry of Agriculture and Rural Afairs, National Fisheries Technology Extension Center, and China Society of fisheries. China Fisheries Statistical Yearbook. 2020. China Agriculture Press. [Google Scholar]
- 4. Ishimatsu A, Itazawa Y. Ventilation of the air-breathing organ in the snakehead Channa argus. Japan J Ichthyol. 1981;28(3):276–82. [Google Scholar]
- 5. Jiang Y, Feng S, Xu J, et al. Comparative transcriptome analysis between aquatic and aerial breathing organs of Channa argus to reveal the genetic basis underlying bimodal respiration. Mar Genomics. 2016;29:89–96. [DOI] [PubMed] [Google Scholar]
- 6. Chen L, Devries AL, Cheng CH. Evolution of antifreeze glycoprotein gene from a trypsinogen gene in Antarctic notothenioid fish. Proc Natl Acad Sci U S A. 1997;94(8):3811–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Beers JM, Jayasundara N. Antarctic notothenioid fish: what are the future consequences of ‘losses’ and ‘gains’ acquired during long-term evolution at cold and stable temperatures?. J Exp Biol. 2015;218(12):1834–45. [DOI] [PubMed] [Google Scholar]
- 8. Volkoff H, Rønnestad I. Effects of temperature on feeding and digestive processes in fish. Temperature. 2020;7(4):307–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Xu J, Bian C, Chen K, et al. Draft genome of the northern snakehead, Channa argus. Gigascience. 2017;6(4):doi: 10.1093/gigascience/gix011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Ranallo-Benavidez TR, Jaron KS, Schatz MC. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat Commun. 2020;11(1):1432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Koren S, Walenz BP, Berlin K, et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27(5):722–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Ruan J, Li H. Fast and accurate long-read assembly with wtdbg2. Nat Methods. 2020;17(2):155–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Vaser R, Ivan S, Nagarajan N, et al. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 2017;27(5):737–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Walker BJ, Abeel T, Shea T, et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One. 2014;9(11):e112963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25(14):1754–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Simão FA, Waterhouse RM, Ioannidis P, et al. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 2015;31(19):3210–2. [DOI] [PubMed] [Google Scholar]
- 17. Rao SS, Huntley MH, Durand NC, et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159(7):1665–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Servant N, Varoquaux N, Lajoie BR, et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 2015;16:259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Burton JN, Adey A, Patwardhan RP, et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat Biotechnol. 2013;31(12):1119–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Zhang C, Ni L, Yang X, et al. Comparison on karyotype of Minxiangli (Channa maculata ♀ × C. argus ♂) and its parents. J Shanghai Fish Univ. 2005;14(2):103–7. [Google Scholar]
- 21. Xu Z, Wang H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 2007;35(Web Server):W265–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Price AL, Jones NC, Pevzner PA. De novo identification of repeat families in large genomes. Bioinformatics. 2005;21(Suppl 1):i351–8. [DOI] [PubMed] [Google Scholar]
- 23. Hoede C, Arnoux S, Moisset M, et al. PASTEC: an automatic transposable element classification tool. PLoS One. 2014;9(5):e91929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Jurka J, Kapitonov VV, Pavlicek A, et al. Repbase update, a database of eukaryotic repetitive elements. Cytogenet Genome Res. 2005;110(1-4):462–7. [DOI] [PubMed] [Google Scholar]
- 25. Tarailo-Graovac M, Chen N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics. 2009;4(4):10. [DOI] [PubMed] [Google Scholar]
- 26. Burge C, Karlin S. Prediction of complete gene structures in human genomic DNA. J Mol Biol. 1997;268(1):78–94. [DOI] [PubMed] [Google Scholar]
- 27. Stanke M, Waack S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics. 2003;19(Suppl 2):ii215–25. [DOI] [PubMed] [Google Scholar]
- 28. Majoros WH, Pertea M, Salzberg SL. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics. 2004;20(16):2878–9. [DOI] [PubMed] [Google Scholar]
- 29. Blanco E, Parra G, Guigó R. Using geneid to identify genes. Curr Protoc Bioinformatics. 2007;4(4):3. [DOI] [PubMed] [Google Scholar]
- 30. Korf I. Gene finding in novel genomes. BMC Bioinformatics. 2004;5(1):59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Keilwagen J, Wenk M, Erickson JL, et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Res. 2016;44(9):e89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Keilwagen J, Hartung F, Paulini M, et al. Combining RNA-seq data and homology-based gene prediction for plants, animals and fungi. BMC Bioinformatics. 2018;19(1):189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Pertea M, Pertea GM, Antonescu CM, et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33(3):290–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Haas BJ, Papanicolaou A. TransDecoder (Find Coding Regions Within Transcripts). http://transdecoder.github.io. Accessed 15 January 2020. [Google Scholar]
- 36. Tang S, Lomsadze A, Borodovsky M. Identification of protein coding regions in RNA transcripts. Nucleic Acids Res. 2015;43(12):e78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Campbell MA, Haas BJ, Hamilton JP, et al. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics. 2006;7(1):327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Haas BJ, Salzberg SL, Zhu W, et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 2008;9(1):R7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Griffiths-Jones S, Moxon S, Marshall M, et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res. 2004;33(Database issue):D121–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25(5):955–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Emms DM, Kelly S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 2019;20(1):238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Mi H, Muruganujan A, Ebert D, et al. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019;47(D1):D419–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Nguyen LT, Schmidt HA, Von Haeseler A, et al. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2015;32(1):268–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Yang Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Bioinformatics. 1997;13(5):555–6. [DOI] [PubMed] [Google Scholar]
- 45. Puttick MN. MCMCtreeR: functions to prepare MCMCtree analyses and visualize posterior ages on trees. Bioinformatics. 2019;35(24):5321–2. [DOI] [PubMed] [Google Scholar]
- 46. Han MV, Thomas GW, Lugo-Martinez J, et al. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol Biol Evol. 2013;30(8):1987–97. [DOI] [PubMed] [Google Scholar]
- 47. Wang Y, Tang H, DeBarry JD, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012;40(7):e49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Kim D, Paggi JM, Park C, et al. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37(8):907–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Zheng Y, Jiao C, Sun H, et al. iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol Plant. 2016;9(12):1667–70. [DOI] [PubMed] [Google Scholar]
- 51. Wang Y, Lu Y, Zhang Y, et al. The draft genome of the grass carp (Ctenopharyngodon idellus) provides genomic insights into its evolution and vegetarian diet adaptation. Nat Genet. 2015;47(6):625–31. [DOI] [PubMed] [Google Scholar]
- 52. Howe K, Clark MD, Torroja CF, et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013;496(7446):498–503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Ren L, Li W, Qin Q, et al. The subgenomes show asymmetric expression of alleles in hybrid lineages of Megalobrama amblycephala × Culter alburnus. Genome Res. 2019;29(11):1805–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Nie M, Tan X, Lu Y, et al. Network of microRNA-transcriptional factor-mRNA in cold response of turbot Scophthalmus maximus. Fish Physiol Biochem. 2019;45(2):583–97. [DOI] [PubMed] [Google Scholar]
- 55. Link W. Introduction to FOXO Biology. Methods Mol Biol. 2019;1890:doi: 10.1007/978-1-4939-8900-3_1. [DOI] [PubMed] [Google Scholar]
- 56. Ou M, Huang R, Yang C, et al. Supporting data for “Chromosome-level genome assemblies of Channa argus and Channa maculata and comparative analysis of their temperature adaptability.”. GigaScience Database. 2021. 10.5524/100925. [DOI] [PMC free article] [PubMed]
- 57. Ou M, Huang R, Yang C, et al. Chromosome-level genome assembly of C. argus. GigaScience Database. 2021. 10.5524/100930. [DOI]
- 58. Ou M, Huang R, Yang C, et al. Chromosome-level genome assembly of C. maculata. GigaScience Database. 2021. 10.5524/100931. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
- Ou M, Huang R, Yang C, et al. Supporting data for “Chromosome-level genome assemblies of Channa argus and Channa maculata and comparative analysis of their temperature adaptability.”. GigaScience Database. 2021. 10.5524/100925. [DOI] [PMC free article] [PubMed]
- Ou M, Huang R, Yang C, et al. Chromosome-level genome assembly of C. argus. GigaScience Database. 2021. 10.5524/100930. [DOI]
- Ou M, Huang R, Yang C, et al. Chromosome-level genome assembly of C. maculata. GigaScience Database. 2021. 10.5524/100931. [DOI] [PMC free article] [PubMed]
Supplementary Materials
Nicolas Rohner -- 6/18/2021 Reviewed
Nansheng (Jack) Chen -- 7/7/2021 Reviewed
Data Availability Statement
Genome, annotation files, and raw sequences for genome assembly including Illumina, Nanopore, and Hi-C reads of C. argus were deposited in the NCBI and can be accessed with accession No. PRJNA731586, and the corresponding data of C. maculata can be accessed with accession No. PRJNA730430. The transcriptome data related to temperature adaptation of C. argus and C. maculata can be accessed with accession No. PRJNA732763. Supporting data and materials are available in the GigaDB database [56], with individual datasets for C. argus [57] and C. maculata [58].