Abstract
DECH-box proteins are a subset of DExH/D-box superfamily 2 helicases possessing a conserved Asp-Glu-Cys-His motif in their ATP binding site. The conserved His helps position the Asp and Glu residues, which coordinate the divalent metal cation that connects the protein to ATP and activate the water molecule needed for ATP hydrolysis, but the role of the Cys is still unclear. This study uses site-directed mutants of the model DECH-box helicase encoded by the hepatitis C virus (HCV) to examine the role of the Cys in helicase action. Proteins lacking a Cys unwound DNA less efficiently than wild-type proteins did. For example, at low protein concentrations, a helicase harboring a Gly instead of the DECH-box Cys unwound DNA more slowly than the wild-type helicase did, but at higher protein concentrations, the two proteins unwound DNA at similar rates. All HCV proteins analyzed had similar affinities for ATP and nucleic acids and hydrolyzed ATP in the presence of RNA at similar rates. However, in the absence of RNA, all proteins lacking a DECH-box cysteine hydrolyzed ATP 10–15 times faster with higher Km values, and lower apparent affinities for metal ions, compared to those observed with wild-type proteins. These differences were observed with proteins isolated from HCV genotypes 2a and 1b, suggesting that this role is conserved. These data suggest the helicase needs Cys292 to bind ATP in a state where ATP is not hydrolyzed until RNA binds.
Graphical Abstract

Helicases are motor proteins that use ATP to separate the complementary strands of the DNA double helix, rearrange DNA or RNA secondary structures, or translocate along nucleic acids. Cells need many different helicases to survive and replicate, but the mechanism of action of only a small fraction of these ubiquitous proteins has been studied at the biochemical level. One of the most widely studied helicases is encoded by the hepatitis C virus (HCV).1 The goal of this study is to understand how a conserved cysteine in the HCV helicase ATP binding site helps couple ATP hydrolysis to the ability of the HCV helicase to unwind DNA.
For many years after HCV was discovered,2 there was no model organism or cell culture system available to discover therapies for treating the more than 150 million known HCV patients. Instead, proteins encoded by HCV were intensely studied as targets for structure-based rational drug design in efforts that eventually led to Food and Drug Administration approval of numerous highly effective direct acting antivirals (DAAs).3 The HCV helicase is part of the HCV protein called nonstructural protein 3 (NS3). The N-terminal third of NS3 is a protease, which is a key DAA target that processes the HCV polyprotein4 and cellular proteins needed to mount an antiviral response.5 The remaining C-terminal part of NS3 (NS3h) is a helicase that separates RNA duplexes, DNA duplexes, or DNA:RNA heteroduplexes.6,7 NS3h requires a 3′ single-stranded region to initiate unwinding, and it translocates in a 3′ to 5′ direction on a nucleic acid strand8 while displacing the complementary strand.9,10 NS3h was one of the first helicases to be structurally examined on an atomic level11–16 and one of the first whose movements on DNA and RNA were observed at the single-molecule level.17,18
HCV NS3h is a member of the DExD/H-box family of helicases, which is part of helicase superfamily 2.19 This protein family is named after the amino acid sequence of a conserved motif present in a wide variety of nucleoside triphosphate-fueled proteins, which was originally known as the “Walker B” motif.20 In NS3h, the site-directed mutants lacking the Asp bind a divalent metal needed to activate hydrolysis ~100 times weaker, and those lacking the Glu hydrolyze ATP ~1000 times slower.21 In NS3h structures, Glu291 and Asp291, together with Ser211 of the Walker A motif, appear to interact with water molecules that make up the coordination sphere of the divalent cation bond to the nucleotide phosphates (Figure 1).15,16
Figure 1.
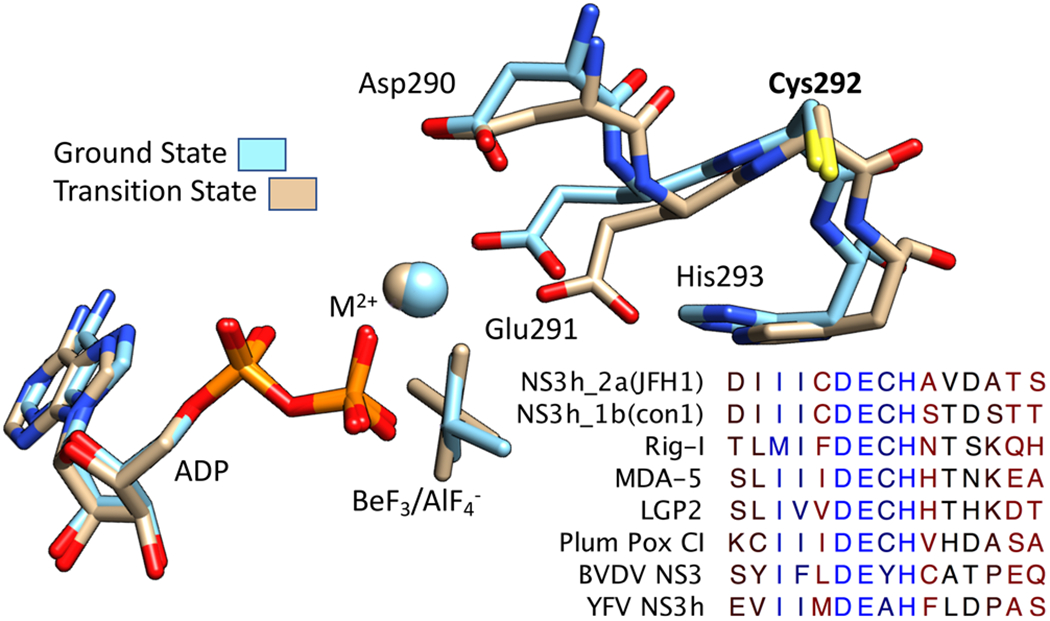
DECH-box motif in RNA helicases. The DECH-box motif in a crystal structure of HCV NS3h bound to the ground state ATP analogue ADP·BeF3 [tan, Protein Data Bank (PDB) entry 3KQN] and transition state analogue ADP·AlF4− (blue, PDB entry 3KQL).15 Also shown are the Mg2+ ion (tan in PDB entry 3KQL) and Mn2+ ions (blue, from PDB entry 3KQL) that connect the DECH box to ADP. Below is an alignment of the amino acids surrounding the DECH box in selected DExD/H-box proteins. Residues are colored on a red–blue gradient based on conservation (blue, conserved; red, variable). RIG-I, MDA5, and LGP2 are cytoplasmic RNA receptors that are key innate immune system components.30 PP C1 is an RNA helicase from plum pox virus, YFV NS3 from yellow fever virus (the namesake flavivirus), and BVDV NS3 from a pestivirus called bovine viral diarrhea virus (flavivirus, pestivirus, and hepacivirus are three genera in the Flaviviridae family).
At least 37 human DExD/H-box proteins have an Ala in the third position and an Asp in the fourth position, and they are therefore called “DEAD-box” proteins.22 A smaller set of proteins possess Cys and His residues in these positions, and they will here be termed DECH-box proteins. Three critical human proteins called RIG-I-like receptors, which initiate the antiviral immune response upon binding viral RNA epitopes, are DECH-box proteins like HCV helicase.23,24 The first RNA helicase isolated from a virus (the plum pox virus CI protein) is also a DECH-box protein.25,26 However, flaviviruses and pestiviruses, which are more closely related to HCV than plum pox virus, encode NS3 proteins with an Ala or a Tyr instead of a Cys (Figure 1).
The observations that Cys is less conserved and points away from ATP in the binding site (Figure 1) suggest that Cys292 might not be essential for HCV helicase action. However, some prior site-directed mutagenesis experiments targeting the Cys292 residue have suggested otherwise.27–29 Here, we perform a more detailed biochemical analysis of HCV NS3 proteins lacking Cys292 and show how the conserved Cys is needed to couple helicase-catalyzed ATP hydrolysis to nucleic acid binding and the ability of the protein to unwind DNA.
MATERIALS AND METHODS
Materials.
The genotype 2a(JHF1) “NS3h” proteins studied here lack the NS3 protease region (amino acids 1—165). The previously described plasmid pET24a-Hel_2a(JFH1)31 was used to express wild-type NS3h_2a(JFH1) and as a template for site-directed mutagenesis using the QuikChange II XL kit (Agilent Technologies). The following oligonucleotides (Integrated DNA Technologies, Coralville, IA) were used to introduce a thymine to guanine mutation needed to encode NS3h_2a(C292G): C292G(+), CAT CAT CAT ATG CGA TGA AGG CCA CGC TGT GGA TGC TAC; C292G(−), 5′-GTA GCA TCC ACA GCG TGG CCT TCA TCG CAT ATG ATG ATG-3′. Desired mutagenized plasmids were identified by determining the DNA sequence of the entire NS3h open reading frame (Genewiz, South Plainfield, NJ). Proteins were expressed in BL21(DE3) cells and purified as described by Hanson et al.32
Genotype 1b33 NS3h proteins (amino acids 166–632) were expressed in BL21(DE3) cells using a modified pET32a vector encoding a C-terminal polyhistidine tag. Cys to Ser substituted variants were produced by site-directed mutagenesis as described above for genotype 2a. Proteins, expressed in soluble form at 25 °C, were first purified from clarified lysates, postsonication, by immobilized metal affinity chromatography (Roche) in Tris-buffered saline (pH 8) and then sized by high-performance liquid chromatography using a G3000SWXL column (Tosoh) in the Tris-HCl mobile phase. Monomeric fractions were pooled and exchanged into storage buffer [25 mM Tris-HCl (pH 8), 1 mM EDTA, 1 mM dithiothreitol (DTT), and 30% glycerol] and then stored frozen.
DNA Unwinding Assays.
Molecular beacon-based helicase assays (MBHAs)34 were performed at 23 °C and contained 25 mM MOPS (pH 6.5), 12.5 nM partially duplex DNA substrate, 1.25 mM MgCl2, 0.025 mM DTT, 5 μg/mL bovine serum albumin (BSA), 0.01% (v/v) Tween 20, and the indicated concentrations of NS3h. After assays had been performed as described by Hanson et al.,32 the observed Cy5 fluorescence (Fobs) after ATP addition was fit to a first-order rate equation:
| (1) |
where F0 is the fluorescence before ATP addition, k is a first-order rate constant, t is the time (seconds), and A is the reaction amplitude.
Initial reaction rates were calculated by multiplying k by A and used to calculate a maximal velocity (Vmax) and a helicase concentration yielding half-Vmax (K0.5) by fitting data using nonlinear regression to
| (2) |
DNA Binding Assay.
The affinity of NS3h for a single-stranded oligonucleotide was measured using the fluorescence polarization assay described by Mukherjee et al.35 Assays (50 μL) contained 25 mM MOPS (pH 6.5), 5 nM Cy5-labeled 15-nucleotide poly(dT) oligonucleotide, 1.25 mM MgCl2, 5 μg/mL BSA, 0.025 mM DTT, 0.005% (v/v) Tween 20, and varying concentrations of NS3h. The fluorescence polarization was measured using a TECAN M1000 PRO instrument at 25 °C, exciting at 635 nm (5 nm bandwidth) and measuring the total fluorescence intensity, parallel and perpendicular polarized light at 667 nm (20 nm bandwidth). G factors were calculated from wells with Cy5-dT15 alone. The observed polarization (Pobs) was fit to total DNA concentration [D]T and protein concentration [E]T and the polarization of free DNA (PF) to estimate a dissociation constant (KD) and the polarization of bound DNA (PB):
where
| (3) |
ATP Hydrolysis Assays.
Helicase-catalyzed ATP hydrolysis was monitored with a colorimetric assay based on formation of a molybdate:phosphate:malachite green complex, as described by Sweeney et al.36 Reactions (30 μL) were initiated by adding 3 μL of ATP of a 10× MgATP2− solution to 27 μL of assembled reagents, such that the final concentrations were 25 mM MOPS (pH 6.5), 1.25 mM MgCl2, 5 μg/mL BSA, 0.05% (v/v) Tween 20, 0.03 mM DTT, 10% DMSO, and 100 nM NS3h in the absence of RNA or 5 nM HCV helicase in the presence of RNA. After 15 min at 23 °C, reactions were terminated by adding 200 μL of a freshly prepared malachite green reagent [3 volumes of 0.045% (w/v) malachite green, 1 volume of 4.2% ammonium molybdate in 4 N HCl, and 0.05 volume of 20% Tween 20], followed immediately by adding 30 μL of 35% sodium citrate. After 30 min, A630 was read, and the net amount of phosphate produced was calculated from a phosphate standard curve after subtracting values obtained from control reactions lacking helicase. All assays were conducted under conditions where rates were linear with time and enzyme concentration and performed at least in triplicate.
To estimate the concentration of RNA needed to stimulate ATP hydrolysis to a half-maximum rate (KRNA), observed rates were divided by enzyme concentration and fit to RNA concentration ([RNA]) and a first-order rate constant describing enzyme turnover in the presence of a saturating concentration of RNA and ATP (kfast):
| (4) |
To estimate the concentration of ATP needed to stimulate ATP hydrolysis to the half-maximum level in the absence of RNA (Km), observed rates were divided by enzyme concentration and fit to ATP concentration ([ATP]) and a first-order rate constant describing enzyme turnover in the absence of RNA (kslow):
| (5) |
To estimate the concentration of ATP needed to stimulate ATP hydrolysis to the half-maximum level in the presence of RNA (Km*), observed rates were divided by enzyme concentration and fit to ATP concentration ([ATP]) and a first-order rate constant describing enzyme turnover in the presence of saturating concentrations of RNA and ATP (kfast):
| (6) |
To estimate the concentration of metal ions needed to stimulate ATP hydrolysis to the half-maximum level in the absence of RNA (Ka), observed rates were divided by enzyme concentration and fit to metal ion concentration ([M]) and a first-order rate constant describing enzyme turnover in the presence of a saturating concentration of ATP (kslow):
| (7) |
To estimate the concentration of metal ion needed to stimulate ATP hydrolysis to the half-maximum level in the presence of RNA (Ka*), observed rates were divided by enzyme concentration and fit to metal ion concentration ([M]) and a first-order rate constant describing enzyme turnover in the presence of saturating concentrations of RNA and ATP (kfast):
| (8) |
RESULTS
To study the role of Cys292 in HCV helicase action, we started with a truncated C-terminally His-tagged protein isolated from the genotype 2a infectious clone JFH1.37 This NS3h_2a(JHF1) protein expresses at high levels in Escherichia coli and is more active than NS3h isolated from other HCV strains.38 Attempts to express NS3h_2a(JHF1) with the conservative substitution of Ser for Cys292 failed, but a plasmid encoding genotype 2a(JHF1) HCV helicase harboring a C292G substitution expressed a soluble protein, which will here be termed NS3h_2a(C292G). Both wild-type and mutant proteins were purified to apparent homogeneity using immobilized metal affinity chromatography followed by gel-filtration and ion-exchange chromatography.32 Purification of NS3h_2a(C292G) yielded ~5 mg of helicase from 2 L of culture, which was approximately half the yield observed with the same bacteria expressing the wild-type NS3h_2a(JHF1) protein. Analysis of NS3h_2a(C292G) revealed that the protein retained DNA helicase activity in a molecular beacon-based helicase assay (MBHA)32,34 (Figure 2A). In MBHAs, the wild-type (Figure 2B) and C292G (Figure 2C) proteins separated the duplex to form hairpin molecules leading to the subsequent decrease in the observed fluorescence. In both cases, the rates of the fluorescence decrease were dependent on protein concentration, and rates could be fit to eq 2 to estimate a maximal unwinding rate (Vmax) and a protein concentration yielding a half-maximal unwinding rate (K0.5).
Figure 2.
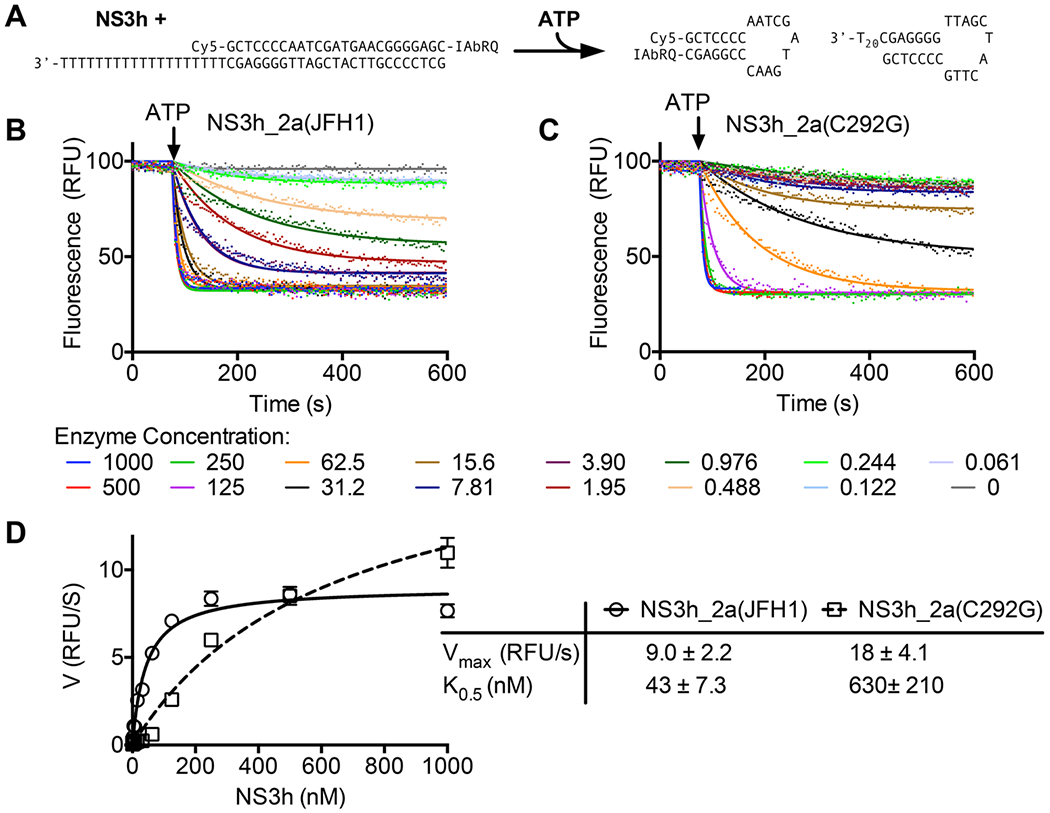
Effect of the C292G substitution on helicase-catalyzed DNA unwinding. (A) Assay setup in which NS3h displaces a molecular beacon annealed to a complementary strand with a 3′ single-stranded DNA tail. The beacon and its complement form hairpins upon separation, leading to a decrease in the observed fluorescence. (B) MBHAs performed with various concentrations (nanomolar) of NS3_2a(JFH1). (C) MBHAs performed with various concentrations of NS3_2a(C292G). In panels B and C, data obtained after ATP addition (points) were fit to eq 1 (solid lines) yielding a rate constant (kobs) and amplitude (A). Initial velocities were calculated by multiplying these two values.32 (D) Initial velocities obtained with NS3_2a(JFH1) (○) and NS3_2a(C292G) (□) fit to eq 2 (Materials and Methods) using the Vmax and K0.5 in the table (right). Uncertainties in the table represent 95% confidence intervals for the nonlinear regression analyses performed with GraphPad Prism.
Although high concentrations of either protein separated DNA at similar rates, careful analysis of the MBHAs revealed a clear effect of the C292G substitution. Unwinding was observed with the wild-type protein even at very low protein concentrations. For example, with 0.488 nM (peach line) or 0.976 nM (dark green line) NS3h_2a(JHF1), there was a clear decrease after ATP addition (Figure 2B). Similar rates of unwinding occurred with only 31.2 nM (black line) or 62.5 nM (orange line) NS3h_2a(C292G) (Figure 2C). These differences were reflected in the K0.5 value that was 15 times lower for the wild-type enzyme. The Vmax values for the two proteins in MBHAs were more similar. In fact, NS3h_2a(C292G) appeared to unwind DNA faster than the wild type did at the highest protein concentration tested (Figure 2D).
The 14-fold difference in K0.5 values mentioned above suggests that NS3_2a(C292G) might bind its DNA substrate less tightly than NS3_2a(JFH1) does. Two different assays were therefore performed to examine the affinity of the proteins for DNA and RNA. In the first, the fluorescence polarization of a Cy5-labeled oligonucleotide was monitored in the absence of protein and in the presence of various concentrations of each protein.35 At each concentration tested, similar polarization values were observed with both NS3_2a(JFH1) and NS3_2a(C292G). Both proteins appeared to bind the oligonucleotide stoichiometrically with a high affinity in this assay with dissociation constants similar to those reported previously (Figure 3A).15,35 The same assays were performed with the DNA used for the MBHAs, and again no difference between the two proteins was observed (data not shown). It is well-established that the binding of ATP to HCV helicase weakens the affinity of the protein for the nucleic acid.21,40–42 It is therefore possible that the different K0.5 values seen in MBHAs might result from differences in the affinity of the helicase for the nucleic acid in the presence of ATP. To examine the effect of the C292G substitution on the ability of HCV helicase to bind RNA in the presence of ATP, assays were performed to monitor helicase-catalyzed ATP hydrolysis in the presence of various concentrations of RNA. Poly(U) RNA was used because it is the nucleic acid that stimulates HCV helicase-catalyzed ATP hydrolysis to the fastest known rate.43,44 The concentration of RNA yielding 50% maximal helicase-catalyzed ATP hydrolysis (KRNA) can be used to estimate the RNA affinity in these experiments. Such an analysis revealed that approximately twice as much RNA was needed to stimulate ATP hydrolysis catalyzed by NS3_2a(C292G) than by NS3_2a(JFH1), suggesting that the substitution weakens nucleic acid binding by ~50%. A smaller difference was observed between the maximal rate of ATP hydrolysis catalyzed by each protein, with the wild-type protein hydrolyzing ATP with a turnover number that was 19% faster than that of the C292G protein (Figure 3B).
Figure 3.
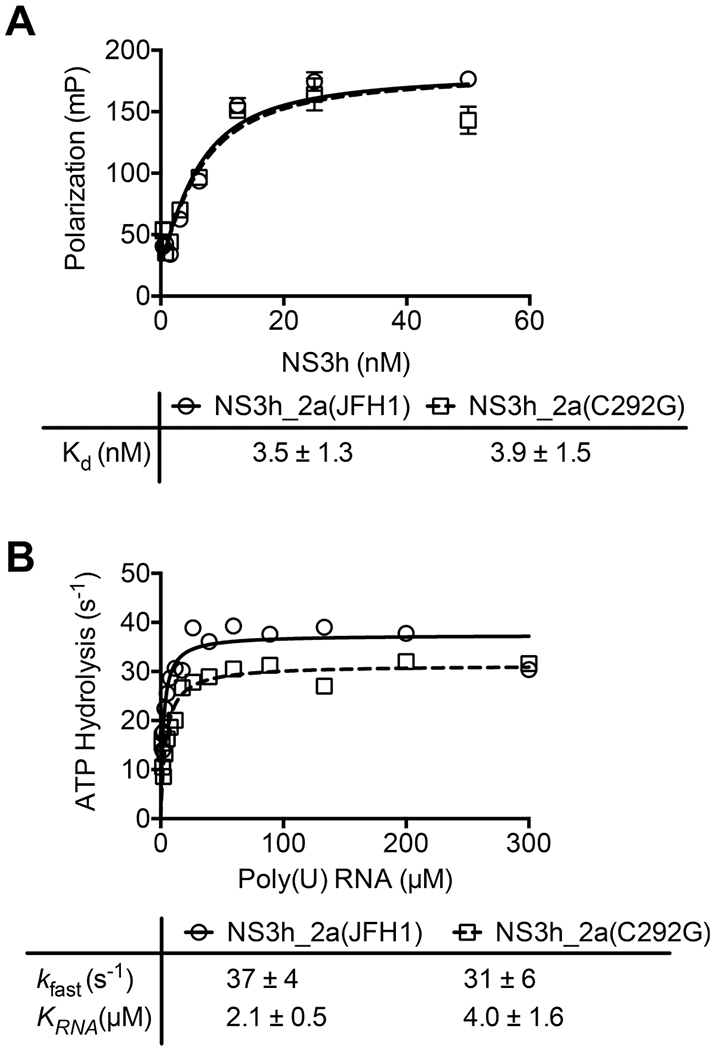
Effect of the C292G substitution on the affinity of NS3h for DNA and RNA. (A) The florescence polarization of Cy5′-dT15 (5 nM) was monitored in solutions containing the indicated concentrations of either NS3_2a(JFH1) (○) or NS3_2a(C292G) (□).35 Averages of three replicates are shown (points). Error bars are standard deviations. Data were fit to a quadratic binding equation (eq 3) with the Kd values listed in the table. (B) Turnover rates of ATP hydrolysis (micromoles of ATP cleaved per second per micromole of enzyme) observed in reaction mixtures containing either helicase (5 nM) and the indicated poly(U) RNA concentrations (in micromoles of nucleotides per liter). The initial ATP concentration in each reaction was 10 mM, and hydrolysis was monitored by measuring the concentration of orthophosphate liberated by the enzyme.36,39 Data were fit to eq 4 with the constants listed in the table. Uncertainties in the tables represent 95% confidence intervals for the nonlinear regression analyses performed with GraphPad Prism.
These data suggest that NS3_2a(JFH1) and NS3_2a(C292G) cleave ATP at similar rates when the enzyme is saturated with ATP (Figure 3). However, they do not rule out the possibility that Cys292 is needed for ATP binding or if different affinities for ATP account for the observed differences in helicase-catalyzed DNA unwinding (Figure 2). Additional assays monitoring rates of ATP hydrolysis at various ATP concentrations were therefore performed to examine if the C292G substitution affects the ability of the helicase to bind ATP. One set of assays was performed in the absence of RNA (Figure 4A), and a second set was performed in the presence of a saturating poly(U) concentration (Figure 4B). Major differences between the enzymes were observed in the absence but not in the presence of RNA.
Figure 4.
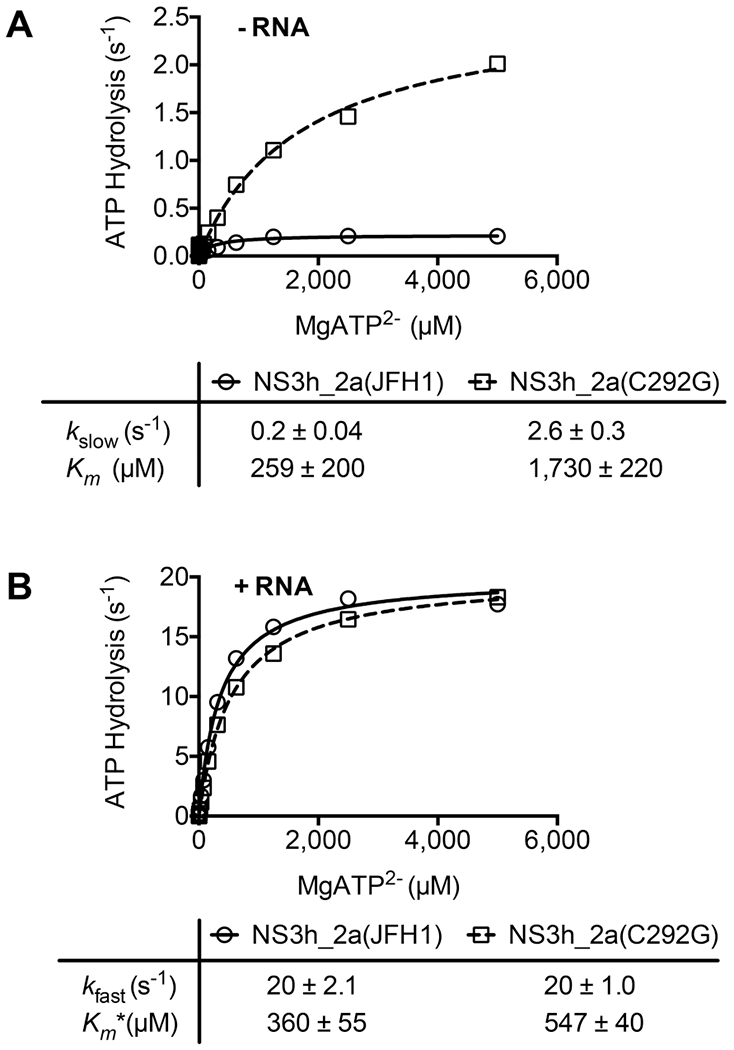
Effect of the C292G substitution on the kinetics of NS3h-catalyzed ATP hydrolysis. Helicase-catalyzed ATP hydrolysis was monitored, as described in the legend of Figure 3B, in reaction mixtures containing various amounts of MgATP2− and additional 1 mM MgCl2. (A) Rates of ATP hydrolysis observed in reaction mixtures containing either helicase (100 nM) in the absence of RNA. (B) Rates of ATP hydrolysis observed in reaction mixtures containing either helicase (5 nM) and 300 μM poly(U) RNA. Data were fit to (A) eq 5 or (B) eq 6 with the indicated values for kslow, Km, Kfast, and Km*. Uncertainties in the tables represent 95% confidence intervals for the nonlinear regression analyses performed with GraphPad Prism.
In the absence of nucleic acids, HCV helicase hydrolyzes ATP 10–100 times more slowly than when DNA or RNA is present. However, unlike related proteins, HCV NS3h still hydrolyzes ATP relatively fast in the absence of nucleic acids, with reported turnover numbers as high as 2 s−1.40,43–47 In the absence of nucleic acids, NS3_2a(C292G) cleaved ATP faster than the wild-type protein did, and at the highest ATP concentration tested, this difference exceeded 10-fold. The apparent Km for ATP was also higher for NS3_2a(C292G). Whereas the wild type was saturated with ATP at 1 mM, rates of hydrolysis catalyzed by NS3_2a(C292G) were still not saturated at the highest ATP concentration tested (5 mM), suggesting a minimal 7-fold difference in Km. The kcat in the absence of a nucleic acid (kslow) was 13 times higher for NS3_2a(C292G) than for NS3_2a(JFH1) (Figure 3A).
In contrast to the rates of ATP hydrolysis observed in the absence of RNA, only minor differences were observed between reactions catalyzed by the two enzymes in the presence of a saturating concentration of poly(U) RNA. Turnover numbers in the presence of RNA (kfast) were the same for each protein. The only differences were reflected in a Km* that was 50% higher in reactions catalyzed by NS3_2a(C292G) than in the same reactions catalyzed by NS3_2a(JFH1) (Figure 4B).
Because both the Asp and Glu in the DECH box help coordinate the divalent metal cation needed to assist NS3h with ATP hydrolysis,15,21 it is possible that the C292G substitution might influence the interaction of divalent metal cations and NS3h. To examine this possibility, rates of ATP hydrolysis were monitored in the presence of various concentrations of Mg2+ and Mn2+. Prior work with wild-type NS3h proteins has shown that both metals stimulate ATP hydrolysis in the presence of RNA (or DNA) but only Mg2+ stimulates ATP hydrolysis in the absence of nucleic acids.48 Saturating concentrations of ATP (10 mM) were used in each assay, so that maximal turnover rates (kfast) would be similar to those seen in experiments in which the proteins were titrated with RNA (Figure 3) or ATP (Figure 4). In the absence of a nucleic acid, Mg2+ stimulated ATP hydrolysis catalyzed by both NS3_2a(JFH1) and NS3_2a(C292G) but to vastly different extents. The turnover rate (kslow) with NS3_2a(C292G) was 19 times faster than with NS3_2a(JFH1). The constant describing the apparent affinity of the activating metal cation (Ka) was also higher with NS3_2a(C292G), suggesting that NS3_2a(C292G) bound Mg2+ 25 times weaker than the wild type did (Figure 5A). In contrast, neither protein was stimulated by Mn2+ in the absence of RNA, suggesting that the amino acid substitution did not alter the metal ion specificity (Figure 5B).
Figure 5.
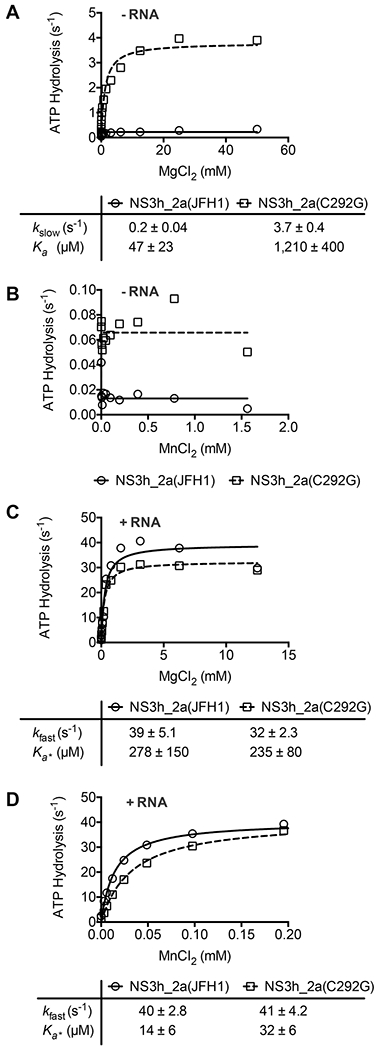
Role of Cys292 in the interaction of HCV helicase with divalent metal cations. Rates of NS3_2a(JFH1)- and NS3_2a(C292G)-catalyzed ATP hydrolysis were monitored in reaction mixtures containing 10 mM ATP and various concentrations of (A and C) MgCl2 or (B and D) MnCl2. Reaction mixtures for panels A and B contained 100 nM enzyme, and reaction mixtures for panels C and D contained either enzyme (5 nM). Rates are expressed as micromolar ATP cleaved per micromolar enzyme per second, and data were fit to (A) eq 7 or (C and D) eq 8 with the indicated values. Uncertainties represent 95% confidence intervals for the fit. Data in panel B did not fit eq 7 because no stimulation by MnCl2 was observed in the absence of RNA.
Only minor differences were observed between the two proteins in the presence of a saturating concentration of poly(U) RNA (300 μM). Mg2+ stimulated ATP hydrolysis catalyzed by both proteins to the same extent and with the same apparent affinity (Figure 5C). Mn2+ also stimulated both proteins to the same extent, but less Mn2+ was required to stimulate the wild type, suggesting that NS3_2a(C292G) bound Mn2+ half as tightly as NS3_2a(JFH1) did (Figure 5D).
The experiments described above were performed with the helicase from the HCV genotype 2a(JFH1) strain because Mukherjee et al. observed that the JFH1 helicase enzyme was more active than the analogous proteins isolated from other HCV genotypes.38 Others have confirmed this observation and have shown that NS3h_2a(JFH1) assumes a conformation not seen with HCV helicases isolated from other genotypes, suggesting that a structural difference accounts for its unusually robust properties.49 To understand if the roles for Cys292 revealed above are retained in other HCV genotypes, its role in HCV genotype 1b was explored. Unlike with genotype 2a(JFH1), a genotype 1b site-directed mutant could be isolated in which Cys292 was substituted with a Ser (C292S), which is a structurally more conservative substitution than the Gly used in NS3h_2a(JFH1). NS3h_1b(C292S) was compared with wild-type NS3h_1b(con1) and a variety of other genotype 1b helicase constructs in which other cysteines were replaced with serine. In addition, a variety of double mutants were constructed in the genotype 1b background in which the C292S substitution was combined with other cysteine mutations (Figure 6A). Prior work showed that NS3h_1b(con1) unwinds DNA more slowly than NS3h_2a(JFH1), but the two proteins hydrolyze ATP at similar rates in the presence of RNA (see Figure 7 of ref 38). In the absence of RNA, NS3h_1b(con1) catalyzed ATP hydrolysis faster than NS3h_2a(JFH1) catalyzed ATP hydrolysis under the same conditions. Despite these differences, replacement of Cys292 led to the same phenotype (Figure 6B); all proteins harboring C292S hydrolyzed ATP faster than the wild type did (or proteins lacking C292S), and the Km for ATP was ~10 times higher for the C292S proteins (Figure 6C).
Figure 6.
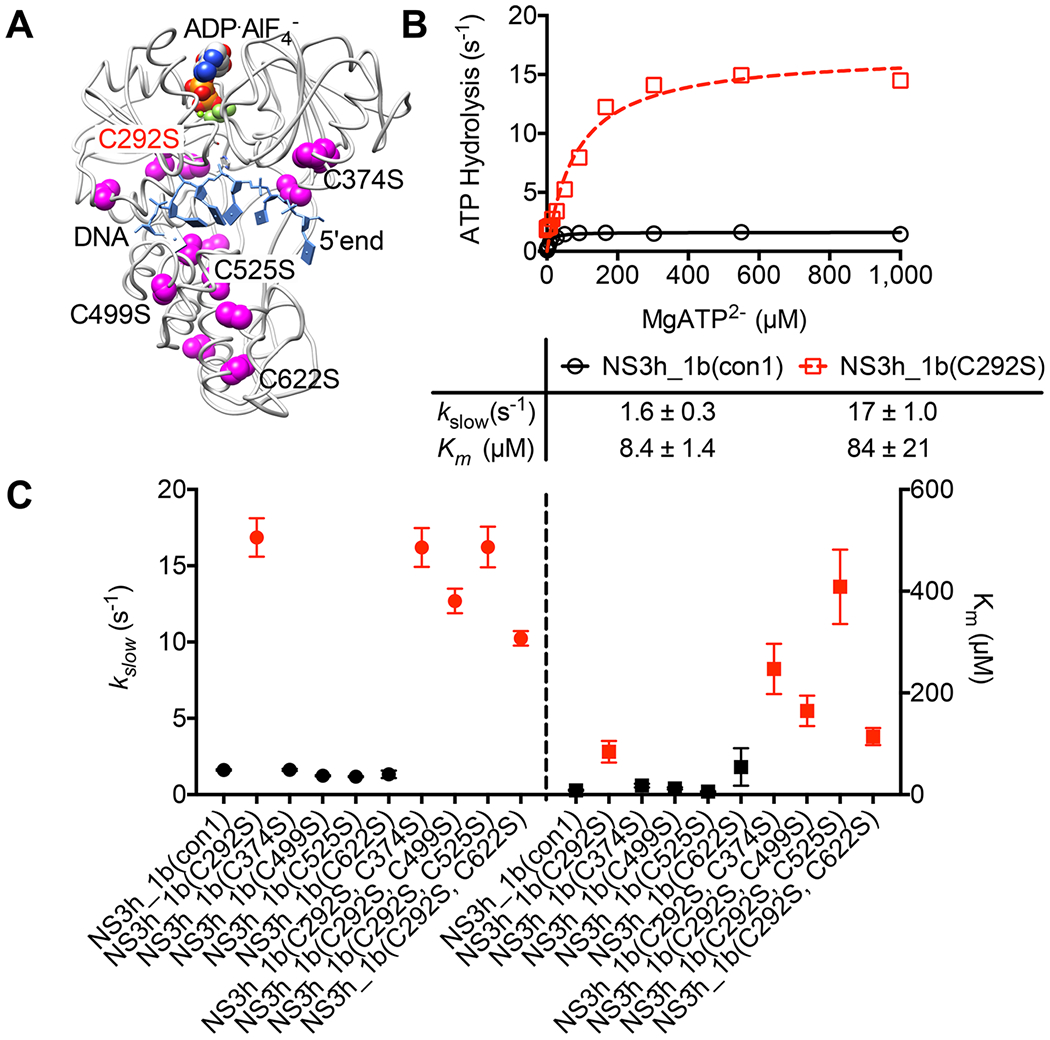
Effect of an NS3 C292S substitution in the HCV genotype 1b genetic background. (A) All cysteine residues (magenta) highlighted in a structure of the genotype 1b NS3h protein bound to DNA (blue) and the nonhydrolyzable ATP analogue ADP·AlF4 (PDB entry 3KQL).15 Residues targeted for site-directed mutagenesis are labeled. (B) Comparison of rates of helicase-catalyzed ATP hydrolysis at various ATP concentrations as described in Figure 4. (C) Comparison of kslow values (left axis) and Km values (right axis) obtained with each protein listed on the x-axis in assays monitoring ATP hydrolysis. In B and C, red points designate proteins harboring a C292S substitution, and in C, and error bars represent 95% confidence intervals for the fit.
DISCUSSION
The data presented above clarify the role of the Cys in the DECH-box motif conserved in a subset of superfamily 2 helicases. The helicase needs the cysteine to stabilize an ATP binding mode that is not ideal for hydrolysis, unless RNA is present. In addition to what was learned about the HCV helicase, the findings might be relevant to other clinically important DECH-box enzymes such as the RIG-like receptors, which are key proteins that modulate the cellular response to viruses.24 NS3h proteins in which Cys292 was replaced with other amino acids have been studied before, but prior results had led to conclusions ranging from the idea that Cys292 is indispensable for helicase activity to the idea that Cys292 is not needed for helicase action.27–29 Our results show how such conclusions could be made on the basis of how assays were set up. They also reveal a more precise role for Cys292 in coupling ATP hydrolysis to unwinding. These results might aid the rational design of therapeutics that modulate the activity of DECH-box proteins.
The role of Cys292 was first studied 20 years ago using NS3h subcloned from the original HCV isolate, which is now called genotype 1a.27 NS3h_1a(C292S) retained an ability to bind RNA but lacked any detectable ability to separate duplex RNA or hydrolyze ATP, suggesting that Cys292 is critical for helicase action.27 However, another study at the time showed that a different NS3h_1a(C292S) protein unwound RNA and cleaved ATP almost as well as the same amount of wild-type protein, suggesting that Cys292 was not needed for activity.28 In a third study that focused on a genotype 1b HCV isolate, NS3h_1b(C292A) was shown to cleave ATP only 54% as fast as the wild type and unwind DNA 18% more slowly than the wild type, again suggesting a minor role for Cys292.29 Our more detailed analysis using a wider variety of assay conditions shows how one might conclude that Cys292 is dispensable while others might conclude it is critical for ATP hydrolysis or unwinding. For example, much smaller differences were seen in assays performed at high enzyme concentrations than at low concentrations (Figure 2), and very large effects on ATP hydrolysis were observed only in the absence of RNA (Figures 3–6).
The statistical significance of each observed difference between proteins was evaluated by examining all nonlinear regression results with the standard tools provided in GraphPad Prism (version 6). All pairs of data sets were tested to determine if the best fit parameters differ between data sets using both an exact sum of squares F-test and Akaike’s information criteria. In all cases, except for the polarization assay used to determine the Kd for helicase and DNA (Figure 3A), the null hypothesis that one curve could fit both data sets was rejected (P < 0.001).
Despite the statistical significance, care should be taken when interpreting how the observed differences might impact helicase function because a 10-fold change in an equilibrium constant alters ΔG° by only 1.4 kcal/mol (i.e., the energy of a hydrogen bond). The differences observed here on that scale are the apparent Km, Ka, and the turnover rate of ATP hydrolysis in the absence of RNA (Figures 4–6). On this basis, our data imply that loss of Cys292 leads to a protein that hydrolyzes ATP faster, and binds MgATP2− weaker, in the absence of nucleic acids. This might explain why at low concentrations, the C292G protein unwinds DNA poorly compared to the wild type. In other words, an enzyme without the DECH-box Cys is less processive. ATP binding weakens the affinity of NS3h helicase for DNA (or RNA),40 and in such a state, the helicase can translocate along the nucleic acid. ATP cleavage returns the helicase to a tightly bound conformation with respect to the DNA.41 If a mutant protein more rapidly hydrolyzes ATP after RNA dissociates, then the protein might have less time to translocate and unwind fewer nucleotides in each step. Such a hypothesis could be tested by measuring the number of base pairs unwound per ATP hydrolyzed using transient state or single-molecule assays.
Another issue that should be addressed in future studies is exactly which step in helicase-catalyzed ATP cleavage is modulated by the DECH box. Helicase-catalyzed ATP cleavage occurs in at least three steps: substrate binding, a chemical step, and release of the two products, ADP and inorganic phosphate. Because all ATPase assays here were performed under steady state conditions, all rates are set by the rate-limiting step. Wang et al. determined that the rate of phosphate release is the same as that of each unwinding step catalyzed by NS3,50 so it is likely that the loss of Cys292 is structurally perturbing the release of phosphate rather than ATP binding or hydrolysis. Such an idea could be tested using transient state assays to measure the kinetics of phosphate release.51
Although the structural basis for the role of the DECH-box cysteine awaits a high-resolution structure of one of the mutants described here, some ideas can be gleaned from data obtained with wild-type NS3.15,16 When the various NS3h structures in the Protein Data Bank are aligned along their peptide backbones, clear movement of the DECH-box residues can be seen upon the binding of ATP analogues (Figure 1). Interestingly, the position of Cys292 differs the least, suggesting that the residue acts as a pivot. For example, in a structure of NS3h bound to the ground state ATP analogue ADP·BeF3 (blue, Figure 1), Glu291 directly contacts that divalent metal ion. However, in a structure with the transition state analogue ADP·AlF4− (tan, Figure 1), Glu291 interacts with water molecules that make up the coordination sphere of the magnesium ion. Thus, we speculate that in the absence of RNA, Cys292 helps lock the protein in a conformation that binds ATP but does not proceed to the transition state. In other words, Cys292 makes for an ATP binding mode that is not ideal for hydrolysis, unless RNA is present. Such an idea is supported by the fact that the Km of ATP is higher with C292G/C292S proteins, and the C292G mutation decreases NS3h’s affinity for Mg2+ by 2 orders of magnitude.
Our lab’s interest in Cys292 started because we were trying to develop antiviral agents that inhibit HCV helicase by targeting critical cysteines. That project was inspired by the fact that the cysteine modifier ebselen is a potent antiviral and helicase inhibitor.38 The mutants described herein were originally constructed to find ebselen-resistant HCV NS3h variants. All proteins studied here are still as sensitive as the wild-type protein to ebselen and its analogues, but this observation does not rule out the possibility that specific antivirals targeting Cys292 might be discovered. For example, Kyono et al. have speculated that Cys292 might serve as the target of antiviral mercuric reagents.52 Although numerous effective direct acting antivirals for HCV are available, most of the world’s HCV patients cannot afford their high cost. New drug targets, like the ATP binding site of NS3,53 might also be needed if HCV evolves to be resistant to present therapies before it can be eradicated.
Funding
This work was supported by National Institutes of Health Grant R01 AI088001 (to D.N.F.).
Footnotes
The authors declare no competing financial interest.
REFERENCES
- (1).Frick DN (2007) The hepatitis C virus NS3 protein: a model RNA helicase and potential drug target. Curr. Issues Mol. Biol 9, 1–20. [PMC free article] [PubMed] [Google Scholar]
- (2).Choo QL, Kuo G, Weiner AJ, Overby LR, Bradley DW, and Houghton M (1989) Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science 244, 359–362. [DOI] [PubMed] [Google Scholar]
- (3).Scheel TK, and Rice CM (2013) Understanding the hepatitis C virus life cycle paves the way for highly effective therapies. Nat. Med 19, 837–849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (4).Grakoui A, McCourt DW, Wychowski C, Feinstone SM, and Rice CM (1993) Characterization of the hepatitis C virus-encoded serine proteinase: determination of proteinase-dependent polyprotein cleavage sites. J. Virol 67, 2832–2843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (5).Li K, Foy E, Ferreon JC, Nakamura M, Ferreon AC, Ikeda M, Ray SC, Gale M, and Lemon SM (2005) Immune evasion by hepatitis C virus NS3/4A protease-mediated cleavage of the Toll-like receptor 3 adaptor protein TRIF. Proc. Natl. Acad. Sci. U. S. A 102, 2992–2997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (6).Kim DW, Gwack Y, Han JH, and Choe J (1995) C-terminal domain of the hepatitis C virus NS3 protein contains an RNA helicase activity. Biochem. Biophys. Res. Commun 215, 160–166. [DOI] [PubMed] [Google Scholar]
- (7).Gwack Y, Kim DW, Han JH, and Choe J (1996) Characterization of RNA binding activity and RNA helicase activity of the hepatitis C virus NS3 protein. Biochem. Biophys. Res. Commun 225, 654–659. [DOI] [PubMed] [Google Scholar]
- (8).Lin CT, Tritschler F, Lee KS, Gu M, Rice CM, and Ha T (2017) Single-molecule imaging reveals the translocation and DNA looping dynamics of hepatitis C virus NS3 helicase. Protein Sci. 26, 1391–1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (9).Morris PD, and Raney KD (1999) DNA helicases displace streptavidin from biotin-labeled oligonucleotides. Biochemistry 38, 5164–5171. [DOI] [PubMed] [Google Scholar]
- (10).Beran RK, Bruno MM, Bowers HA, Jankowsky E, and Pyle AM (2006) Robust translocation along a molecular monorail: the NS3 helicase from hepatitis C virus traverses unusually large disruptions in its track. J. Mol. Biol 358, 974–982. [DOI] [PubMed] [Google Scholar]
- (11).Yao N, Hesson T, Cable M, Hong Z, Kwong AD, Le HV, and Weber PC (1997) Structure of the hepatitis C virus RNA helicase domain. Nat. Struct. Biol 4, 463–467. [DOI] [PubMed] [Google Scholar]
- (12).Kim JL, Morgenstern KA, Griffith JP, Dwyer MD, Thomson JA, Murcko MA, Lin C, and Caron PR (1998) Hepatitis C virus NS3 RNA helicase domain with a bound oligonucleotide: the crystal structure provides insights into the mode of unwinding. Structure 6, 89–100. [DOI] [PubMed] [Google Scholar]
- (13).Cho HS, Ha NC, Kang LW, Chung KM, Back SH, Jang SK, and Oh BH (1998) Crystal structure of RNA helicase from genotype 1b hepatitis C virus. A feasible mechanism of unwinding duplex RNA. J. Biol. Chem 273, 15045–15052. [DOI] [PubMed] [Google Scholar]
- (14).Mackintosh SG, Lu JZ, Jordan JB, Harrison MK, Sikora B, Sharma SD, Cameron CE, Raney KD, and Sakon J (2006) Structural and biological identification of residues on the surface of NS3 helicase required for optimal replication of the hepatitis C virus. J. Biol. Chem 281, 3528–3535. [DOI] [PubMed] [Google Scholar]
- (15).Gu M, and Rice CM (2010) Three conformational snapshots of the hepatitis C virus NS3 helicase reveal a ratchet translocation mechanism. Proc. Natl. Acad. Sci. U. S. A 107, 521–528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (16).Appleby TC, Anderson R, Fedorova O, Pyle AM, Wang R, Liu X, Brendza KM, and Somoza JR (2011) Visualizing ATP-dependent RNA translocation by the NS3 helicase from HCV. J. Mol. Biol 405, 1139–1153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (17).Dumont S, Cheng W, Serebrov V, Beran RK, Tinoco I, Pyle AM, and Bustamante C (2006) RNA translocation and unwinding mechanism of HCV NS3 helicase and its coordination by ATP. Nature 439, 105–108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (18).Frick DN (2006) Step-by-step progress toward understanding the hepatitis C virus RNA helicase. Hepatology 43, 1392–1395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Iyer LM, Leipe DD, Koonin EV, and Aravind L (2004) Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol 146, 11–31. [DOI] [PubMed] [Google Scholar]
- (20).Walker JE, Saraste M, Runswick MJ, and Gay NJ (1982) Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (21).Frick DN, Banik S, and Rypma RS (2007) Role of divalent metal cations in ATP hydrolysis catalyzed by the hepatitis C virus NS3 helicase: magnesium provides a bridge for ATP to fuel unwinding. J. Mol. Biol 365, 1017–1032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (22).Linder P, and Jankowsky E (2011) From unwinding to clamping - the DEAD box RNA helicase family. Nat. Rev. Mol. Cell Biol 12, 505–516. [DOI] [PubMed] [Google Scholar]
- (23).Ranji A, and Boris-Lawrie K (2010) RNA helicases: emerging roles in viral replication and the host innate response. RNA Biol. 7, 775–787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (24).Corby MJ, Stoneman MR, Biener G, Paprocki JD, Kolli R, Raicu V, and Frick DN (2017) Quantitative microspectro-scopic imaging reveals viral and cellular RNA helicase interactions in live cells. J. Biol. Chem 292, 11165–11177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Laín S, Riechmann JL, and García JA (1990) RNA helicase: a novel activity associated with a protein encoded by a positive strand RNA virus. Nucleic Acids Res. 18, 7003–7006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (26).Laín S, Martín MT, Riechmann JL, and García JA (1991) Novel catalytic activity associated with positive-strand RNA virus infection: nucleic acid-stimulated ATPase activity of the plum pox potyvirus helicaselike protein. J. Virol 65, 1–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).Kim DW, Kim J, Gwack Y, Han JH, and Choe J (1997) Mutational analysis of the hepatitis C virus RNA helicase. J. Virol 71, 9400–9409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Wardell AD, Errington W, Ciaramella G, Merson J, and McGarvey MJ (1999) Characterization and mutational analysis of the helicase and NTPase activities of hepatitis C virus full-length NS3 protein. J. Gen. Virol 80, 701–709. [DOI] [PubMed] [Google Scholar]
- (29).Tai CL, Pan WC, Liaw SH, Yang UC, Hwang LH, and Chen DS (2001) Structure-based mutational analysis of the hepatitis C virus NS3 helicase. J. Virol 75, 8289–8297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (30).Lässig C, and Hopfner KP (2017) Discrimination of cytosolic self and non-self RNA by RIG-I-like receptors. J. Biol. Chem 292, 9000–9009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (31).Belon CA, High YD, Lin TI, Pauwels F, and Frick DN (2010) Mechanism and specificity of a symmetrical benzimida-zolephenylcarboxamide helicase inhibitor. Biochemistry 49, 1822–1832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (32).Hanson AM, Hernandez JJ, Shadrick WR, and Frick DN (2012) Identification and analysis of inhibitors targeting the hepatitis C virus NS3 helicase. Methods Enzymol. 511, 463–483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (33).Kato N, Hijikata M, Ootsuyama Y, Nakagawa M, Ohkoshi S, Sugimura T, and Shimotohno K (1990) Molecular cloning of the human hepatitis C virus genome from Japanese patients with non-A, non-B hepatitis. Proc. Natl. Acad. Sci. U. S. A 87, 9524–9528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (34).Belon CA, and Frick DN (2008) Monitoring helicase activity with molecular beacons. BioTechniques 45, 433–440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (35).Mukherjee S, Hanson AM, Shadrick WR, Ndjomou J, Sweeney NL, Hernandez JJ, Bartczak D, Li K, Frankowski KJ, Heck JA, Arnold LA, Schoenen FJ, and Frick DN (2012) Identification and analysis of hepatitis C virus NS3 helicase inhibitors using nucleic acid binding assays. Nucleic Acids Res. 40, 8607–8621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (36).Sweeney NL, Shadrick WR, Mukherjee S, Li K, Frankowski KJ, Schoenen FJ, and Frick DN (2013) Primuline derivatives that mimic RNA to stimulate hepatitis C virus NS3 helicase-catalyzed ATP hydrolysis. J. Biol. Chem 288, 19949–19957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (37).Wakita T, Pietschmann T, Kato T, Date T, Miyamoto M, Zhao Z, Murthy K, Habermann A, Kräusslich HG, Mizokami M, Bartenschlager R, and Liang TJ (2005) Production of infectious hepatitis C virus in tissue culture from a cloned viral genome. Nat. Med 11, 791–796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (38).Mukherjee S, Weiner WS, Schroeder CE, Simpson DS, Hanson AM, Sweeney NL, Marvin RK, Ndjomou J, Kolli R, Isailovic D, Schoenen FJ, and Frick DN (2014) Ebselen inhibits hepatitis C virus NS3 helicase binding to nucleic acid and prevents viral replication. ACS Chem. Biol 9, 2393–2403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (39).Sweeney NL, Hanson AM, Mukherjee S, Ndjomou J, Geiss BJ, Steel JJ, Frankowski KJ, Li K, Schoenen FJ, and Frick DN (2015) Benzothiazole and pyrrolone flavivirus inhibitors targeting the viral helicase. ACS Infect. Dis 1 , 140–148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (40).Levin MK, Gurjar MM, and Patel SS (2003) ATP binding modulates the nucleic acid affinity of hepatitis C virus helicase. J. Biol. Chem 278, 23311–23316. [DOI] [PubMed] [Google Scholar]
- (41).Levin MK, Gurjar M, and Patel SS (2005) A Brownian motor mechanism of translocation and strand separation by hepatitis C virus helicase. Nat. Struct. Mol. Biol 12, 429–435. [DOI] [PubMed] [Google Scholar]
- (42).Lam AM, Rypma RS, and Frick DN (2004) Enhanced nucleic acid binding to ATP-bound hepatitis C virus NS3 helicase at low pH activates RNA unwinding. Nucleic Acids Res. 32, 4060–4070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (43).Suzich JA, Tamura JK, Palmer-Hill F, Warrener P, Grakoui A, Rice CM, Feinstone SM, and Collett MS (1993) Hepatitis C virus NS3 protein polynucleotide-stimulated nucleoside triphosphatase and comparison with the related pestivirus and flavivirus enzymes. J. Virol 67, 6152–6158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (44).Lam AM, Keeney D, Eckert PQ, and Frick DN (2003) Hepatitis C virus NS3 ATPases/helicases from different genotypes exhibit variations in enzymatic properties. J. Virol 77, 3950–3961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (45).Preugschat F, Averett DR, Clarke BE, and Porter DJ (1996) A steady-state and pre-steady-state kinetic analysis of the NTPase activity associated with the hepatitis C virus NS3 helicase domain. J. Biol. Chem 271, 24449–24457. [DOI] [PubMed] [Google Scholar]
- (46).Porter DJ (1998) A kinetic analysis of the oligonucleotide-modulated ATPase activity of the helicase domain of the NS3 protein from hepatitis C virus. The first cycle of interaction of ATP with the enzyme is unique. J. Biol. Chem 273, 14247–14253. [DOI] [PubMed] [Google Scholar]
- (47).Beran RK, Lindenbach BD, and Pyle AM (2009) The NS4A protein of hepatitis C virus promotes RNA-coupled ATP hydrolysis by the NS3 helicase. J. Virol 83, 3268–3275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (48).Belon CA, and Frick DN (2009) Fuel specificity of the hepatitis C virus NS3 helicase. J. Mol. Biol 388, 851–864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (49).Zhou T, Ren X, Adams RL, and Pyle AM (2018) NS3 from Hepatitis C Virus Strain JFH-1 Is an Unusually Robust Helicase That Is Primed To Bind and Unwind Viral RNA. J. Virol 92, e01253–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (50).Wang Q, Arnold JJ, Uchida A, Raney KD, and Cameron CE (2010) Phosphate release contributes to the rate-limiting step for unwinding by an RNA helicase. Nucleic Acids Res. 38, 1312–1324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (51).Brune M, Hunter JL, Corrie JE, and Webb MR (1994) Direct, real-time measurement of rapid inorganic phosphate release using a novel fluorescent probe and its application to actomyosin subfragment 1 ATPase. Biochemistry 33, 8262–8271. [DOI] [PubMed] [Google Scholar]
- (52).Kyono K, Miyashiro M, and Taguchi I (2003) Characterization of ATPase activity of a hepatitis C virus NS3 helicase domain, and analysis involving mercuric reagents. J. Biochem 134, 505–511. [DOI] [PubMed] [Google Scholar]
- (53).Ndjomou J, Corby MJ, Sweeney NL, Hanson AM, Aydin C, Ali A, Schiffer CA, Li K, Frankowski KJ, Schoenen FJ, and Frick DN (2015) Simultaneously Targeting the NS3 Protease and Helicase Activities for More Effective Hepatitis C Virus Therapy. ACS Chem. Biol 10, 1887–1896. [DOI] [PMC free article] [PubMed] [Google Scholar]


