Abstract
Hundreds of millions of people worldwide are affected by viral infections each year, and yet, several of them neither have vaccines nor effective treatment during and post-infection. This challenge has been highlighted by the COVID-19 pandemic, showing how viruses can quickly spread and impact society as a whole. Novel interdisciplinary techniques must emerge to provide forward-looking strategies to combat viral infections, as well as possible future pandemics. In the past decade, an interdisciplinary area involving bioengineering, nanotechnology and information and communication technology (ICT) has been developed, known as Molecular Communications. This new emerging area uses elements of classical communication systems to molecular signalling and communication found inside and outside biological systems, characterizing the signalling processes between cells and viruses. In this paper, we provide an extensive and detailed discussion on how molecular communications can be integrated into the viral infectious diseases research, and how possible treatment and vaccines can be developed considering molecules as information carriers. We provide a literature review on molecular communications models for viral infection (intra-body and extra-body), a deep analysis on their effects on immune response, how experimental can be used by the molecular communications community, as well as open issues and future directions.
Keywords: Communicable diseases, infection, molecular communications, virions, virus
I. Introduction
The COVID-19 pandemic shocked the world by demonstrating the severity of the viral infection and how it can disrupt society by impacting human health as well as global economies. As of September 2020, more than 34 million people have contracted the disease resulting in just over a million deaths, with a mortality rate of approximately 4%. During the first months of the pandemic, global stock markets experienced their worst crash since 1987, in the first three months of 2020 the G20 economies fell by 3.4% year-on-year, an estimated 400 million full-time jobs were lost across the world, and income earned by workers globally fell 10%, where all of this effects is equivalent to a loss of over US
 3.5 trillion [1]. As a result, governments around the world have quickly formulated new recovery plans, where for example in the EU, an investment of 750 billion euros is set to bring the continent back to normality within the first half of the decade (this also includes funding for research on COVID-19) [2]. Despite these investments, the world must prepare for not only coping with this new disease and its various effects on the human health, but also seeking for novel technologies that can help minimise, or even block, future pandemics.
3.5 trillion [1]. As a result, governments around the world have quickly formulated new recovery plans, where for example in the EU, an investment of 750 billion euros is set to bring the continent back to normality within the first half of the decade (this also includes funding for research on COVID-19) [2]. Despite these investments, the world must prepare for not only coping with this new disease and its various effects on the human health, but also seeking for novel technologies that can help minimise, or even block, future pandemics.
The SARS-CoV-2 virus itself is likely to remain a challenge for the next couple of years despite the development of vaccines [3]. First, it is challenging to develop a vaccine that is effective for different virus strains and their mutations. Besides, for patients that are infected, the detrimental effect of the virus in the human body can leave lifelong consequences to tissues and organs. To give an example of the difficulty of eradicating viruses, historic virus such as influenza had its first pandemic in the 16th century and is still considered a global health challenge till this present day [4]. Therefore, constant efforts in new robust vaccines, as well as drugs, are continuously being sought and this requires the development of new technologies that focus on the mechanisms of infections, and in particular the virus molecular relationships with the host cells [5].
In the past 10 years, an interdisciplinary research area known as Molecular Communications has been developing, and it bridges the areas of communication engineering and networking, molecular biology, as well as bioengineering [6], [7]. This area focuses on realising radical new technology for subtle sensing and actuation capabilities inside the human body through a network of micro- and nano-sized devices [8], [9]. These devices can use the existing natural signalling of cells and tissues to interact, as well as communication with the human body. The main advantage is the ability to increase the biocompatibility of implantable systems, and one of the ways to realise this is to integrate communication system engineering with systems and synthetic biology [8], [9]. This novel research area can have a central role to combat current and future pandemics, not only for understanding new insights into the viral properties and characteristics, but also for novel treatments [10], [11], [12], [13]. Molecular communications can contribute to (a) the characterisation of the virus propagation within the body, (b) understanding the mechanism used by the virus to enter the human body, or mechanism of expulsion, and (c) elucidating how the airborne virus propagates in the air. Additionally, although not covered in this survey, molecular communications is at the foundations of the Internet of Bio-Nano Things paradigm [8], where communication between engineered cells for viral infection detection/therapy is intercepted, interpreted, and manipulated by bio-cyber interfaces that can transmit data to cloud-based digital healthcare services [14].
However, the literature on molecular communications does not provide a wide range of work that tackles the issue of viral infection as a whole. Even though there are models for molecular communications for bacterial infection [15], [16], there have not been any surveys proposed for molecular communications models of viral infections. To date, molecular communications models for viral infection includes multi-hop transfer of genetic content through diffusion over extracellular channels [17], viral propagation in the air [18], [19], propagation within the respiratory tract [20], as well as interactions with host cells [21]. Even though these models are very interesting and provide a formidable representation of biology through the glasses of a molecular communications researcher, the issue of virus propagation and the infection itself are much more complex. First, models must gather all necessary information about the infection process, which comprises of the replication of viruses and intra-body propagation, going down to the interaction of genes and host cells, as well as the virus spike proteins effects to binding host cell receptors. Secondly, virology is a very active research area and has collected many resources over the years (data and tools) that can benefit molecular communications research. In order to develop research work with a strong societal impact in order to tackle viral infectious diseases, molecular communications researchers are required to bridge the gap between communication theory and experimental biology, and in particular the use of available data.
This paper presents a literature review and analysis of existing models and data for molecular communications. The goals of this paper are as follows: 1) to provide how the infectious disease is currently modelled using molecular communications; 2) to provide a deep analysis on the existing models to provide a direction on how they should be improved, looking from a biological standpoint; 3) to provide initial guidelines on what experimental data can be used and how they should be integrated to molecular communications models, and 4) to identify the main challenges and issues that the community should focus on moving forward. We recognise that molecular communications can support not only the understanding of infectious diseases, but it can also elicit the development of novel technologies for both sensing and actuation in the body based on how viruses propagate, are transmitted and received by host cells.
The paper has the following contributions:
-
•
A literature survey on models of infectious diseases for intra-body and extra-body molecular communications: We present a deep analysis on existing molecular communications models looking at how to address the issue of bridging these models closer to the existing biological literature and data on infectious disease. For the intra-body models, we investigate the virus entry mechanism, the virus spread and the immune system response. For the extra-body models, we look into the transmitter, channel and receiver processes for human-human propagation of infection.
-
•
Analysis of existing open data on viral infections that can be utilised by molecular communications researchers: We collect a variety of data from a number of sources that we believe can be used by the community to gain a better understanding of viral infections. These data vary from the genetic information of a variety of viruses to the molecular structure and the effects on the hosts, e.g., the immune markers and host infection impact. We focus on a selection of viruses that includes the SARS-CoV (1–2), MERS-CoV, Ebola (EBOV), Dengue (DENV), Zika (ZIKV) and hepatitis C (HCV).
-
•
Overview on open issues and challenges: Based on the many opportunities for research in molecular communications and infectious diseases, we provide five different points where we present a deep analysis aiming to highlight what are the main topics to drive future research on molecular communications. We include discussion about: 1) linking experimental data to molecular communications models, 2) novel intra-body viral intervention techniques, 3) emerging technologies for infection theranostics (therapy and diagnostics), 4) bridging molecular communications and bioinformatics tools, and 5) novel molecular communications models.
The paper structure is as follows. Section II presents the background information on infectious disease. Section III presents a literature survey on viral models for infections in intra-body and extra-body settings. Section IV presents a set of experimental data that can be exploited in molecular communications. Section V presents the open issues and challenges for the future of molecular communications research on communicable disease. Finally, Section VI concludes the study.
II. Background Information on Infectious Diseases
In this section, we go through several known communicable viral diseases and provide examples of devastating outbreaks in the 21st century. We select seven viruses that, at the time of writing this paper, do not have a licensed vaccine for treatment or where the intervention mechanisms are only used to alleviate symptoms of the hosts. Our survey focuses on three families of viruses, and this includes Coronavirus, Filovirus, and Flavivirus.
A. Coronavirus
Coronaviridae is a family of viruses that includes SARS-CoV-1, MERS-CoV and SARS-CoV-2. The most severe virus in this family is the SARS-CoV-2. Example properties of SARS-CoV-2 include asymptomatic infection to severe pneumonia and replicates through a variety of cells that exhibit Angiotensin Converting Enzyme 2 (ACE2) expression (a number of these cells are found in the respiratory tract, and, in particular, deep in the alveolar regions). SARS-CoV-1 and MERS-CoV are known to cause severe pneumonia with high replication rates in the respiratory tract. The immune response to the three different viruses is also very different. In the case of SARS-CoV-1 and MERS-CoV, antibodies response at an early stage of the infection process. However, this is not the case for SARS-CoV-2, where the symptoms from the infection process can take up to two weeks.
There are several differences between SARS-CoV-1, MERS-CoV and SARS-CoV-2 in the spreading process. While SARS-CoV-1 and MERS-CoV are known to develop severe pneumonia, they have exhibited limited person-to-person spreading, which is very different from SARS-CoV-2 [22]. Even though promising solutions for vaccines targeting SARS-CoV-2 are in the testing phase, their efficacy is yet unknown or unpredictable. Besides vaccines, the existing immune-based treatment (e.g., plasma transfusion) is only found to have temporary effects [23]. On top of that, there are other several unknowns about how SARS-CoV-2 affects different organs, indicated by clinical data [24], [25].
B. Filovirus
Filoviridae family of viruses comes from the thread-like structure of the virus that also contains many curvy branches [26]. The most common, or well known, is the EBOV (zaire ebolavirus). Currently, there is only one FDA approved Ebola vaccine (approved in 2019 [27]) that has a successful performance of around 70% to 100% efficiency, and this is the rVSV-ZEBOV vaccine [28]. This vaccine acts as the glycoprotein duplicate. Once expressed in the host, it will activate the immune system response. The vaccine was used for clinical trials in West-Africa, to cope with the local 2016 pandemic. However, research is still on-going to analyse the vaccine response for virus genome mutation.
C. Flavivirus
Flaviviridae family of viruses is mainly characterised by the yellow complexion found on the hosts after infection (hence yellow fever), and by the transmission mode of arthropod vectors (mainly ticks and mosquitoes). We analyse three types of virus in this family, and they include DENV, ZIKV and HCV.
The single positive-stranded RNA DENV is mosquito-borne and mostly found in countries in the centre global hemisphere, where the warm temperature is an ideal location for mosquito’s habitation. The virus has not always been found to transmit through mosquitoes. Many years ago, the transmission mode was sylvatic, meaning contraction from wild animal contacts. Over the past 20 years, dengue fever has increased dramatically, affecting more than 390 million people each year. The main molecular characteristic of this virus is the genomic RNA surrounded by numerous protein layers.
ZIKV is transmitted and similarly affects the host cells to the DENV since both share a distinct genetic component. However, the fever from ZIKV infection is more potent and is found to impact developing fetus in pregnant women, and can lead to microcephaly. This virus is relatively new. Hence, no vaccine is available. Similarly to DENV, there are seven non-structural proteins, three structural proteins and a positive single-stranded RNA genome. However, the main difference is the mechanism in which the host cells reacts to the genome upon infection, where the infected cells are found to progress into the swollen stages leading to cell death. This is only possible through the virus gene expression and the inability of the host cell to protect itself against virus binding through the concentration of the IFITM3 protein.
Lastly, HCV is also a single-stranded RNA virus. Infected hosts can have symptoms that include occasional fever, dark urine, abdominal pain and yellow-tinged skin. To date, the virus has infected nearly 71 million people worldwide. Even though simpler in structure, there is still no vaccine for ECV because of its numerous genotypes derived from their protein structure, or even proper medical intervention techniques. These efforts would help minimise damaging effects to the liver, the organ most damaged by this virus in infected patients [29].
D. Viruses Structural Data
In Table I, we collect the main quantitative information about the viruses discussed in this section. In order to obtain an accurate representation of the virus propagation and relationship with the host, we analyse the viral concentrations in specimens, the characteristics of virions and the virus detection for each type of virus. Different basal, or steady-state concentrations, are provided for different settings where specimens were collected. This will help to figure out initial parameter values in reaction-diffusion models that account for virus interactions with the host. Next, we present the characteristics of the viruses, which state the structural dimensions (useful for quantifying the diffusion profile), genetic profile (useful for transcription of gene information between the virus and host cells), as well as types of proteins on the virus surfaces (useful for accounting the reception of the virus by a cell by means of ligand biding). Lastly, we provide many detection mechanisms that were used, and are useful, for measuring the viral concentration on specimens.
TABLE I. Available Quantitative Data to Support Molecular Communications Modelling.
| Type | Viral concentrations in specimens | Characteristics of virion | Detection | Ref. |
|---|---|---|---|---|
| SARS-CoV (1–2) | SARS-CoV: Nasopharyngeal aspirates – mean values of  RNA copies/mL, RNA copies/mL,
 copies/mL, and copies/mL, and
 copies/mL on days 5, 10, and 15, respectively (collected from 14 patients) copies/mL on days 5, 10, and 15, respectively (collected from 14 patients)Stool – a mean value of  copies/mL on day 10 (collected from 1 patient) copies/mL on day 10 (collected from 1 patient)SARS-CoV-2: Lower respiratory specimens (sputum samples) – 641  copies/mL, with a median of copies/mL, with a median of
 (collected from 80 patients) (collected from 80 patients)Posterior oropharyngeal saliva – a median of  copies/mL copies/mLUpper respiratory specimens (nasal and throat swabs) – cycle threshold (Ct) values 19–40 (collected from 14 patients); Ct values are inversely related to viral RNA copy number, with Ct of 30.76, 27.67, 24.56, and 21.48 corresponding to  , ,
 , ,
 , and , and
 copies/mL copies/mL |
Enveloped, spherical, 60–140 nm in diameter with spikes of about 9–12 nm Genome: 30 kb positive-sense, single-stranded RNA RNA Transcript: 5’ cap, 3’ poly-A tail Proteome: 10 proteins |
Viral RNA and protein quantification; Serologic detection of antibodies against viruses |
[30], [31], [32], [33], [34], [35] |
| MERS-CoV | Lower and upper respiratory specimens (sputum or tracheal aspirate, and throat-swab) – a median of
 RNA copies/mL in the severe group and RNA copies/mL in the severe group and
 copies/mL in the mild group (depending on whether oxygen supplementation was used) (P = 0.06) (collected from 17 patients) copies/mL in the mild group (depending on whether oxygen supplementation was used) (P = 0.06) (collected from 17 patients) |
Around 100 nm in diameter Genome: (+)ssrNA, 30 kb (20–33 kb) enveloped, positive-sense, single-stranded RNA virus Proteome: 1a, 1b, S, e, M, N, Assorted OrFs RNA Transcript: 5’-capped |
Viral RNA and protein quantification; Serologic detection of antibodies against viruses |
[36], [35], [37], [38] |
| EBOV | Serum – up to 1010 RNA copies/mL in samples from the patients who died; up to 108 within the first 8 days after the onset of symptoms; Viremia classified as low, intermediate, or high according to the viral load (  , ,
 , and , and
 ) GEQ/mL (genome equivalent per mL), respectively) (collected from 632 patients) ) GEQ/mL (genome equivalent per mL), respectively) (collected from 632 patients) |
Filamentous, 970 nm long, 80 nm in diameter, enveloped Genome: approximately 19 kilobase negative-sense, single-stranded RNA Proteome: 7 sequentially arranged proteins RNA Transcript: 5’ methyl cap, 3’ poly-A tail |
Viral RNA and protein quantification; Serologic detection of antibodies against viruses |
[39], [40], [35] |
| DENV | Serum – 103–1011 RNA copies/mL (collected from 228 patients, whose data were stratified by infecting dengue serotype (DENV-1, DENV-2, or DENV-3) and by immune status and clinical manifestation (primary infection dengue fever, secondary infection dengue fever, or secondary infection dengue hemorrhagic fever) | Around 50 nm in diameter, icosahedral, enveloped Genome: 10.7 kilobase positive-sense, single-stranded RNA Proteome: single polyprotein, co- & post-translationally cleaved into 11 mature proteins RNA Transcript: 5’ methyl cap, no 3’ poly-A tail |
Viral RNA and protein quantification; Serologic detection of antibodies against viruses | [41], [42], [43], [35] |
| ZIKV | Plasma –
 RNA copies/mL RNA copies/mLUrine –  RNA copies/mL RNA copies/mLSemen –  RNA copies/mL (collected 2 weeks after symptom onset from a 32-year-old man) and up to RNA copies/mL (collected 2 weeks after symptom onset from a 32-year-old man) and up to
 RNA copies/mL measured (collected from 12 men in French Guiana with ZIKV infection) RNA copies/mL measured (collected from 12 men in French Guiana with ZIKV infection)Plasma, urine, and saliva – ZIKV RNA found in the saliva of only the woman; ZIKV remained detectable in their plasma for about 2 weeks, while the urine of the man was consistently positive until day 14 and that of the woman until day 25; The greater virus load in the urine observed in the man (collected from a 69-year-old man and 66-year-old-woman) |
Enveloped, spherical, 50 nm in diameter with an electron dense core of 30 nm Genome: 10.8 kb positive-sense, single-stranded RNA Proteome: single polyprotein, co- & post-translationally cleaved into 11 mature proteins RNA Transcript: 5’ methyl cap, no 3’ poly-A tail |
Viral RNA and protein quantification; Serologic detection of antibodies against viruses |
[44], [45], [46], [35] |
| HCV | Serum –
 RNA copies/mL (collected from 38 chronically infected patients – 19 infected by genotype 1 isolates (1a and 1b), 13 by nongenotype 1 isolates (including genotypes 2 a/c, 3a, and 4), and 6 coinfected by genotype 1 and 6 isolates) RNA copies/mL (collected from 38 chronically infected patients – 19 infected by genotype 1 isolates (1a and 1b), 13 by nongenotype 1 isolates (including genotypes 2 a/c, 3a, and 4), and 6 coinfected by genotype 1 and 6 isolates) |
Around 50 nm in diameter, icosahedral, enveloped Genome: 9.6 kilobase positive-sense, single-stranded RNA Proteome: single polyprotein, co- & post-translationally cleaved into 10 mature proteins RNA Transcript: 5’ internal ribosomal entry site (IRES), no 3’ poly-A tail |
Viral RNA quantification; Serologic detection of antibodies against HCV |
[47], [35] |
III. Molecular Communications for Viral Infections
In this section, we present the literature review on the intra-body- and extra-body molecular communications models.
A. Intra-Body Molecular Communications Models
1). Virus Entry Mechanisms:
The molecular communications paradigm gives us a clear understanding of how the virus acts and distributes within the body over time. In the context of communications, the virions are considered as information carriers, which propagate messages (genome) from the location of transmission until the location of the reception, which can be the host cells in specific organs or tissues. The information conveyed by the virions is the infection action.
In theory, a single virion is enough to enter the body and initiate a viral infection, provided that the host cells are accessible to the viral binding process. Besides the accessible cells being susceptible to infection – they must also express the receptors to which the virus binds, and permissive to infection – which means they must contain protein and machinery necessary for virus replication [48].
If the virus enters the host through the respiratory tract, gastrointestinal tract, genital tract or optical tract, the main barrier between the virus and internal environment of the body is the epithelial cells – the layer of cells that line the outer surfaces of organs and blood vessels and the inner surfaces of cavities (Fig. 1a). The epithelial cells of the respiratory tract are targeted by SARS-CoV (1–2) and MERS-CoV viruses as the most common portal of entry. Unlike SARS-CoV (1–2), which exclusively infects and releases through the apical route,1 MERS-CoV can spread through either side of human bronchial epithelial cells. SARS-CoV (1–2) and MERS-CoV viruses contained in larger droplets are deposited in the upper respiratory tract (the nose, nasal passages, sinuses, pharynx, and larynx), while smaller aerosolised particles or liquids are transferred into the lower respiratory tract (the trachea, bronchi, and lungs). EBOV targets the epithelial cells as a final attack though, after infecting fibroblasts of any type (especially fibroblastic reticular cells), mononuclear phagocytes (with dendritic cells more affected than monocytes or macrophages) and endothelial cells. On the other hand, if the virus is delivered through penetration of the skin (e.g., DENV- or ZIKV infection from a mosquito bite), wounds or transplantation of an infected organ (e.g., HCV infected organ), the epithelium is bypassed.
Fig. 1.
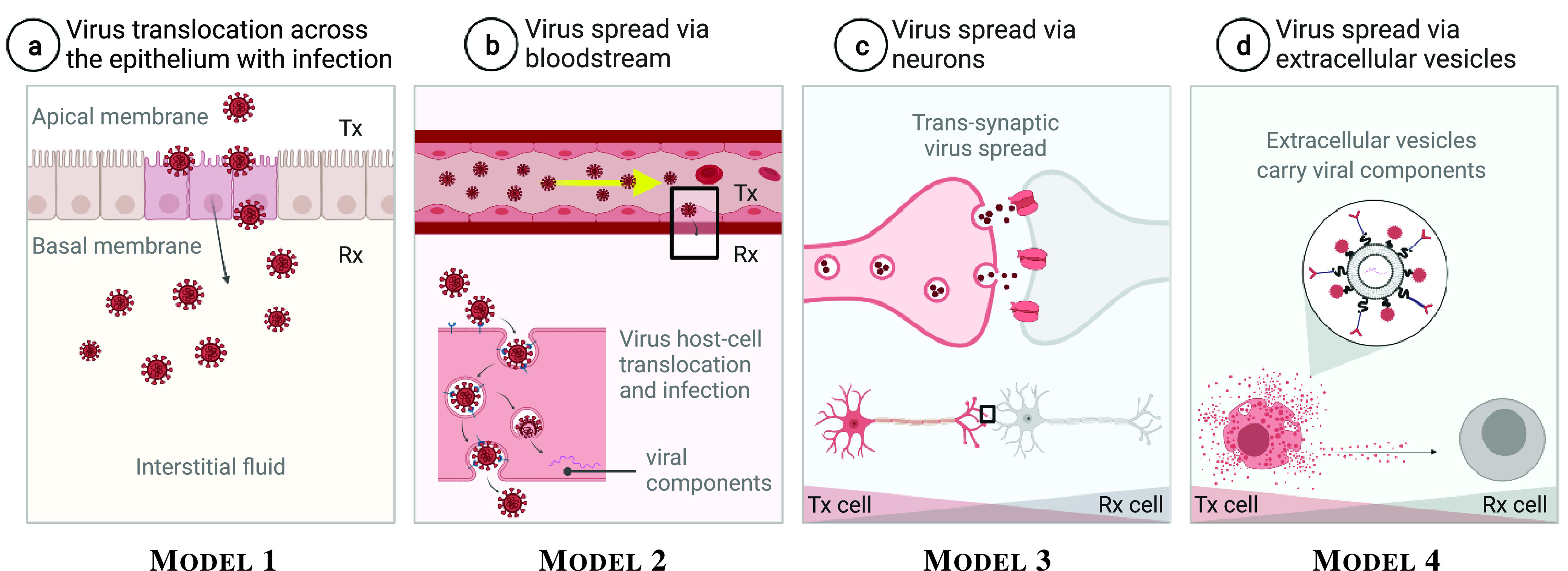
Molecular communications channels of viral intra-body spread. After (a) translocation across the epithelium, the virus spread throughout the body utilising (b) the circulatory system, (c) nervous network, and (d) cell-released EVs that carry viral components as their cargo and deliver to other cells, eventually causing systemic infection.
Viruses have evolved strategies to translocate across the epithelial barrier and act as pathogens. They can enter and infect or cross epithelial cells through the following three modes [49]: 1) Endocytosis and transcytosis (without infection), 2) Polarised surface entry and infection by fusion, and 3) Endocytosis and endosomal fusion with infection.
Endocytosis and transcytosis (without infection) are, respectively, entry- and intracellular transport mechanisms for specific viruses, such as poliovirus, reovirus and human immunodeficiency virus 1 (HIV-1), performed by specific lymphoid areas of the gastrointestinal tract covered by specialised epithelial cells known as M cells. During endocytosis, which is initiated at clathrin- and caveolin-coated pits and vesicles, or lipid raft microdomains, the host cell engulfs the virus. During transcytosis, the host cell transports the virus through its cytosol and eventually eject the virus at the opposite side of the membrane. Polarised surface entry and infection by fusion is an entry mechanism for enveloped viruses, including SARS-CoV (1–2) and MERS-CoV, whose genome is surrounded by a capsid and a membrane [50]. The virus fuses either to the apical membrane or the basal membrane of the epithelial cell and transfers the genome into the cytoplasm. Lastly, endocytosis and endosomal fusion with infection is an entry mechanism for both enveloped and naked (the genome is surrounded only by a capsid) viruses, including EBOV. Other examples include influenza virus types A and C, bovine coronavirus, hepatitis A (HAV), vesicular stomatitis virus (VSV), primary herpes simplex virus (HSV), human cytomegalovirus (HCMV), adeno-associated virus (AAV)-2, simian virus 40 (SV40), measles virus, Semliki Forest virus (SFV), Sindbis virus, Jamestown Canyon (JC) polyomavirus, parvovirus, and the minor group of human rhinoviruses (HRV). These viruses internalise and retain in transport vesicles. To gain access to the cytoplasm, their genome has to leave the vesicle by which it was taken up, usually by penetrating the host cell cytosol through fusion from an endosome.
We refer to modelling viral translocation across the epithelial barrier as Model 1. Despite all differences in the mechanisms involved in this model, the transfer process always starts with the virions binding to the target receptors. Upon binding, the virions become fused with- or internalised into the host cell cytosol. Recycling/negative feedback mechanisms regulate the number of surface bonds between the virions and the receptors. This leads to the following chemical kinetic model representing the viral load/concentration at the extracellular space
 , the viral load at the epithelial host cell membrane (host cell-bound virions)
, the viral load at the epithelial host cell membrane (host cell-bound virions)
 , and the viral load in the host cell cytosol (fused or internalised virions)
, and the viral load in the host cell cytosol (fused or internalised virions)
 , respectively [51]:
, respectively [51]:
 |
where
 is the ratio of the volume of the considered medium containing
is the ratio of the volume of the considered medium containing
 and the host cell volume,
and the host cell volume,
 is the initial viral load at the extracellular space,
is the initial viral load at the extracellular space,
 is a constant rate of viral clearance per virion by mechanisms such as immune elimination (corresponding to a virion half-life
is a constant rate of viral clearance per virion by mechanisms such as immune elimination (corresponding to a virion half-life
 ),
),
 is the total number of the viruses that can be bound,
is the total number of the viruses that can be bound,
 (
(
 ) is the total number of occupied receptors per unit volume at the membrane,
) is the total number of occupied receptors per unit volume at the membrane,
 is the total number of unoccupied receptors per unit volume at the membrane,
is the total number of unoccupied receptors per unit volume at the membrane,
 is the rate defined through the maximal binding rate
is the rate defined through the maximal binding rate
 measured when none of the viruses is bound to the membrane, and
measured when none of the viruses is bound to the membrane, and
 is the virus fusion or internalisation rate [51].
is the virus fusion or internalisation rate [51].
 can have steady-state values considering different cases listed in Table I, where viral concentration values are available. Even with the other model variables that can be obtained from the literature, values for
can have steady-state values considering different cases listed in Table I, where viral concentration values are available. Even with the other model variables that can be obtained from the literature, values for
 set the range where the solutions of the equations converge.
set the range where the solutions of the equations converge.
The presented model can be considered accurate if the data of the concentration of viral ligands interaction with the host cell receptors are available.
2). Virus Spread:
After translocation across the epithelial barrier, the virus infects and replicates at the site of infection, causing localised infections, and/or initiates infection through one organ and then spreads to other sites, causing systemic infections [48].
A straightforward way to describe the viral load (
 ) dynamics in a localised infection is to use the target cell-limited model [52]. This model neglects intracellular processes and takes into account uninfected susceptible target cells (
) dynamics in a localised infection is to use the target cell-limited model [52]. This model neglects intracellular processes and takes into account uninfected susceptible target cells (
 ) and infected virus-producing cells (
) and infected virus-producing cells (
 ) within an observed organ. The basic model is formulated by the following system of nonlinear ordinary differential equations (ODEs) [53]:
) within an observed organ. The basic model is formulated by the following system of nonlinear ordinary differential equations (ODEs) [53]:
 |
where
 |
The target cells become infected cells which produce virus with production rate
 ,
,
 is a constant infectivity rate,
is a constant infectivity rate,
 is a constant rate of death in infected cells [corresponding to an infected cell half-life of
is a constant rate of death in infected cells [corresponding to an infected cell half-life of
 ],
],
 is a constant rate of uninfected target cells production, and
is a constant rate of uninfected target cells production, and
 is a constant rate of uninfected target cells death [corresponding to a target cell half-life of
is a constant rate of uninfected target cells death [corresponding to a target cell half-life of
 ]. This model can be applied to analytically describe the local spread of any family of viruses. Apart from the viruses considered in Section II, rhinovirus and papillomavirus are examples of viruses that cause only a localised infection. Rhinovirus infects the epithelial cells of the upper respiratory tract and replicates there, whereas papillomavirus infects the skin and replicates in the epidermis.
]. This model can be applied to analytically describe the local spread of any family of viruses. Apart from the viruses considered in Section II, rhinovirus and papillomavirus are examples of viruses that cause only a localised infection. Rhinovirus infects the epithelial cells of the upper respiratory tract and replicates there, whereas papillomavirus infects the skin and replicates in the epidermis.
Describing the viral load dynamics in a systemic infection is more challenging since the virus spreads to other organs using mechanisms like the bloodstream (hematogeneous spread), neurons (neurotropic spread), or extracellular vesicles.
Viruses can enter the bloodstream either directly through inoculation into an animal or insect bites (e.g., DENV and ZIKV), or through the release of virions produced at the entry site into the interstitial fluid (e.g., coronaviruses) [48]. This fluid can be taken up by lymphatic vessels that lead back to lymph nodes. Although immune system cells filter the interstitial fluid within the lymph nodes, some virions escape immune cells and continue within the interstitial fluid, which is eventually returned to the bloodstream. The virus takes advantage of the blood distribution network for the propagation of the virions from a location they are injected into the blood flow to a targeted site within reach of the cardiovascular system. Advection and diffusion are the mass transport phenomena in the cardiovascular system [54]. As a result of advection, the virions are transported by the flow of the blood at different velocities in different locations of the cardiovascular system. As a result of diffusion, the virions are transported from a region of higher concentration to a region of lower concentration. This pattern of motion follows the Brownian motion spread in the blood.
To leave the circulatory system and infect other sites in the body, the virions need to penetrate the blood vessel walls made of the endothelial cells (Fig. 1b). We refer to modelling viral translocation across the endothelial barrier as Model 2. Viruses enter and then infect or cross endothelial cells by endocytosis at the apical (luminal) membrane. When infecting the endothelial cell, the virions penetrate the host cell cytosol by fusion from endosomes. However, if the virus crosses the endothelial cell, the virions are transported via intracellular trafficking and ejected from the basolateral (abluminal) membrane into the extracellular space. This leads to the following chemical kinetic model [55]:
 |
where
 ,
,
 ,
,
 ,
,
 and
and
 represent the viral load at the extracellular space (luminal side), the viral load at the endothelial host cell membrane (host cell-bound virions), the viral load in the host cell endosomes, the viral load in the host cell cytosol that penetrated the endosomes, and the viral load at the extracellular space (abluminal side), at point
represent the viral load at the extracellular space (luminal side), the viral load at the endothelial host cell membrane (host cell-bound virions), the viral load in the host cell endosomes, the viral load in the host cell cytosol that penetrated the endosomes, and the viral load at the extracellular space (abluminal side), at point
 and time
and time
 ,
,
 (
(
 is the set of points over the endothelial host cell membrane).
is the set of points over the endothelial host cell membrane).
 and
and
 ,
,
 1, 2, 3 are forward and backward reaction rates in ms−1 and
1, 2, 3 are forward and backward reaction rates in ms−1 and
 , respectively, and
, respectively, and
 is the endosome penetration rate in
is the endosome penetration rate in
 .
.
The virion transport mechanism across the blood vessel walls imposes a boundary condition for advection-diffusion in the vessel. Concentrations for different types of viruses in plasma and serum from Table I can be used here as initial values. These values do not take into account the dispersion in the blood and shall be considered in entering points to the blood vessels. This mechanism is modelled by a continuous-time Markov chain framework leading to the following general homogeneous boundary condition [55]:
 |
where
 is the diffusion coefficient in
is the diffusion coefficient in
 of the virions in the blood,
of the virions in the blood,
 ,
,
 is the gradient operator, (
is the gradient operator, (
 ) is the inner multiplication operator, and
) is the inner multiplication operator, and
 is the surface normal at
is the surface normal at
 pointing towards the vessel luminal side. The virion advection-diffusion in the observed blood vessel can then be modelled by the Fick’s second law:
pointing towards the vessel luminal side. The virion advection-diffusion in the observed blood vessel can then be modelled by the Fick’s second law:
 |
subject to the boundary condition (12). The release rate of the virus at point
 is given by the source term
is given by the source term
 (virion
(virion
 ),
),
 is the Laplace operator, and
is the Laplace operator, and
 is a constant rate of viral clearance in the blood. The blood is assumed to have a laminar flow in the axial direction with uniform velocity profile
is a constant rate of viral clearance in the blood. The blood is assumed to have a laminar flow in the axial direction with uniform velocity profile
 ms−1, where
ms−1, where
 is the axial unit vector. The fundamental characteristic function for advection-diffusion called the concentration Green’s function is analytically derived in terms of a convergent infinite series [55]. The obtained concentration Green’s function is coupled to the boundary condition given in (12) and provides a useful tool for prediction of the viral load in blood vessels.
is the axial unit vector. The fundamental characteristic function for advection-diffusion called the concentration Green’s function is analytically derived in terms of a convergent infinite series [55]. The obtained concentration Green’s function is coupled to the boundary condition given in (12) and provides a useful tool for prediction of the viral load in blood vessels.
Viruses rarely enter into neurons directly to evoke the neurotropic viral spread (Fig. 1c). We refer to modelling neurotropic viral spread as Model 3. Viruses first replicate locally and then infect nerves associated with the tissue [48]. Thus, viruses first infect neurons of the peripheral nervous system, and then gain access to the central nervous system. There is emerging speculations that the central nervous system may be involved during SARS-CoV-2 infection, where neuron-to-neuron transmission route is used to spread the virus [56]. Other examples of neuroinvasive viruses include several herpes viruses (e.g., herpes simplex virus) and poliovirus, which is weakly neuroinvasive, and rabies virus, which requires tissue trauma to become neuroinvasive. The literature is, however, very sparse concerning biological models on the neurotropic viral spread. Therefore, we advocate for the consideration of detailed biological models where the characterisation of viral spread throughout the nervous system is considered in more details. This includes addressing secondary mechanisms evolved by some viruses to help them replicate and spread (e.g., binding to a host cell protein called dynein which then transports viral capsid to the neural nucleus for replication).
Extracellular vesicles (EVs) are exchanged between all cells and emerge as the novel, yet obscure cell-to-cell communication mediators. EVs vary in size (50–5000 nm) and contain and transport transmembrane proteins in their lipid bilayer, as well as the cytosol molecular components from the parental cell. The latter includes functional proteins, lipids, and genetic materials [e.g., messenger RNA (mRNA), non-coding RNA (ncRNA), and DNA] [57]. EVs can also transfer functionally active cargo and have the ability to participate in biological reactions associated with viral dissemination – the evidence exists for HCV, HIV, and Epstein-Barr virus (EBV) – and immune response (Fig. 1d) [58].
EVs and viruses share common features in their size, structure, biogenesis and uptake [59]. EVs either favour viral infections or limit them, by prompting viral spread or modulating the immune response, respectively. When leveraging viral infection, virus-associated EVs deploy mechanisms such as the delivery of (a) proteins that make the cell more susceptible to infection, (b) viral receptors to cells that are devoid of these receptors thus allowing cells to be infected, (c) nucleic acids that improve and sustain the production of a virus, and (d) molecules that eliminate the host protein relevant for an antiviral response [58], [59]. On the other hand, different mechanisms can be activated by the EVs released by infected cells to prompt an immune response against viruses. The most important mechanisms are the spreading of viral antigens via EVs, and the transfer of cytosolic proteins and nucleic acids involved in antiviral responses. Nonetheless, it is still unclear what cell conditions and virus types release EVs that favour or fight infection.
For initial molecular communications system modelling, it seems that viral components hijack the EV secretory routes to exit infected cells and use EV endocytic routes to enter uninfected and immune system cells [58]. We refer to modelling EV-based viral spread as Model 4. Each infected cell in this model serves as the transmitter, actively interacting with other cells [60]. The transmitting cell either 1) produces EVs (specifically, exosomes) through its intracellular machinery and releases them upon the fusion of intermediate vesicle-containing endosome compartments, referred to as multivesicular bodies, with the plasma membrane, or 2) involves vertical trafficking of molecular cargo to the plasma membrane, a redistribution of membrane lipids, and the use of contractile machinery at the surface to allow for vesicle pinching (specifically, microvesicles) [61]. This corresponds to EVs moving from the intracellular space to the extracellular space (the propagation medium). The aspects of EV release yet need to be theoretically investigated addressing infection factors.
The extracellular matrix (ECM) is the interstitial channel through which EVs are exchanged between the infected virus-producing transmitting cells and the target uninfected receiving cells. The ECM is a 3D molecular network composed of macromolecules. To reach the targeted cell, both the virions and virus-associated EVs should navigate around these macromolecules and diffuse inside and outside other cells in the ECM. The Langevin stochastic differential equation (SDE) can potentially be utilised as a channel modelling tool [6]. Since EVs propagate within the ECM based on a drifted random walk, the Langevin SDE needs to contain contributions from the Brownian stochastic force and the drift velocity of the interstitial fluid. Besides, the Langevin SDE needs to be modified to address (a) the losses or clearances of EVs via uptake from other cells and/or degradation through enzymatic attacks, and (b) the anisotropic EV diffusion affected by the ECM properties, i.e., volume fraction and tortuosity. The volume fraction defines the percentage of the total ECM volume accessible to the virus-bearing EVs. The tortuosity describes the average hindrance of a medium relative to an obstacle-free medium. Hindrance results in an effective diffusion that is decreased compared with the free diffusion coefficient of EVs.
The receiving cell takes up EVs once they bind to the cell-membrane utilising one of the three mechanisms: 1) juxtacrine signalling—where EVs elicit transduction via intracellular signalling pathways, 2) fusion—where EVs fuse with the cellular membrane and transfer cargo (i.e., virus-associated components) into the cytoplasm, and 3) endocytosis—where EVs internalise and retain in transport vesicles. Non-linear EV-uptake associated with these various mechanisms have been initially investigated in terms of EV-based drug delivery, utilising the Volterra series and multi-dimensional Fourier analysis [51]. The ability to receive viral loads and react accordingly can serve as the performance indicator to reconstruct the information sent by the transmitting cell.
3). Immune System Response:
Similar to the virus analysis, the molecular communications paradigm can give us a clear understanding of how the immune system acts and develops within the body over time. Without going into detailed elaboration, we identify the following two systems:
-
•
The Cytokines-based molecular communications system, which represents cells like macrophages, T-helper cells, natural killer cells, neutrophils, dendritic cells, mast cells, monocytes, B cells and T cells, all serving as transceivers; we refer to modelling cytokines-based molecular communications system as Model 5, and
-
•
The Antibody-based molecular communications system, which represents cells like plasma B cells and T cells serving as transmitters, and the virus serving as the receiver; we refer to modelling antibody-based molecular communications system as Model 6.
The immune system is a complex network of cells and proteins that defends the body against pathogen infection. Two subsystems compose the immune system: the innate immune system and the adaptive immune system. The innate immune system is referred to as non-specific as it provides a general defence against harmful germs and substances. The adaptive immune system is referred to as specific as it makes and uses antibodies to fight certain germs that the body has been previously exposed to. The immune system thus works to eradicate the virus. Considering a detailed role of the immune system, i.e., the additional mechanisms in fighting a viral infection
 from the innate immune response (IIR) and adaptive immune response (AIR), ODEs (4)–(6) can be extended [62]:
from the innate immune response (IIR) and adaptive immune response (AIR), ODEs (4)–(6) can be extended [62]:
 |
Additional effects are also included: two populations of infected cells – infected but not yet virus-producing cells (
 ) with the duration of latent eclipse phase of
) with the duration of latent eclipse phase of
 , and infected and virus-producing cells (
, and infected and virus-producing cells (
 ), as well as dead cell (
), as well as dead cell (
 ) replacement by new susceptible cells at a constant rate
) replacement by new susceptible cells at a constant rate
 . The IIR (
. The IIR (
 ) frees the virus at a constant rate
) frees the virus at a constant rate
 and dies at a constant rate
and dies at a constant rate
 ;
;
 is the strength of innate response. The AIR (
is the strength of innate response. The AIR (
 ) is activated proportional to the free viral load at a constant rate
) is activated proportional to the free viral load at a constant rate
 . Activation is followed by clonal expansion at a constant rate
. Activation is followed by clonal expansion at a constant rate
 . The AIR neutralises the virus with a constant rate
. The AIR neutralises the virus with a constant rate
 .
.
 and
and
 in (18) and (19) represent concentrations of cytokines and antibodies, respectively. Cytokines are peptides secreted by immune cells (predominantly macrophages, dendritic cell and T-helper cells) to orchestrate an immune response or an attack on the invading pathogen (Fig. 2c). Cytokines spread through the body and attach to surface receptors of other immune cells. The receptors then signal the cell to help fight the infection. Cytokines are divided into four categories – interleukins, interferons, chemokines and tumour necrosis factors – which can be pro-inflammatory or anti-inflammatory, thus promoting or inhibiting the proliferation and functions of other immune cells. Antibodies are unique proteins encoded by millions of genes which are made and mutated in the human body. They are secreted by immune cells (predominantly plasma B cells differentiated from B cells) to neutralise the pathogen (Fig. 2d). The antibody neutralises the pathogen by recognising a unique molecule of the pathogen, called an antigen, via the fragment antigen-binding (Fab) variable region.
in (18) and (19) represent concentrations of cytokines and antibodies, respectively. Cytokines are peptides secreted by immune cells (predominantly macrophages, dendritic cell and T-helper cells) to orchestrate an immune response or an attack on the invading pathogen (Fig. 2c). Cytokines spread through the body and attach to surface receptors of other immune cells. The receptors then signal the cell to help fight the infection. Cytokines are divided into four categories – interleukins, interferons, chemokines and tumour necrosis factors – which can be pro-inflammatory or anti-inflammatory, thus promoting or inhibiting the proliferation and functions of other immune cells. Antibodies are unique proteins encoded by millions of genes which are made and mutated in the human body. They are secreted by immune cells (predominantly plasma B cells differentiated from B cells) to neutralise the pathogen (Fig. 2d). The antibody neutralises the pathogen by recognising a unique molecule of the pathogen, called an antigen, via the fragment antigen-binding (Fab) variable region.
Fig. 2.
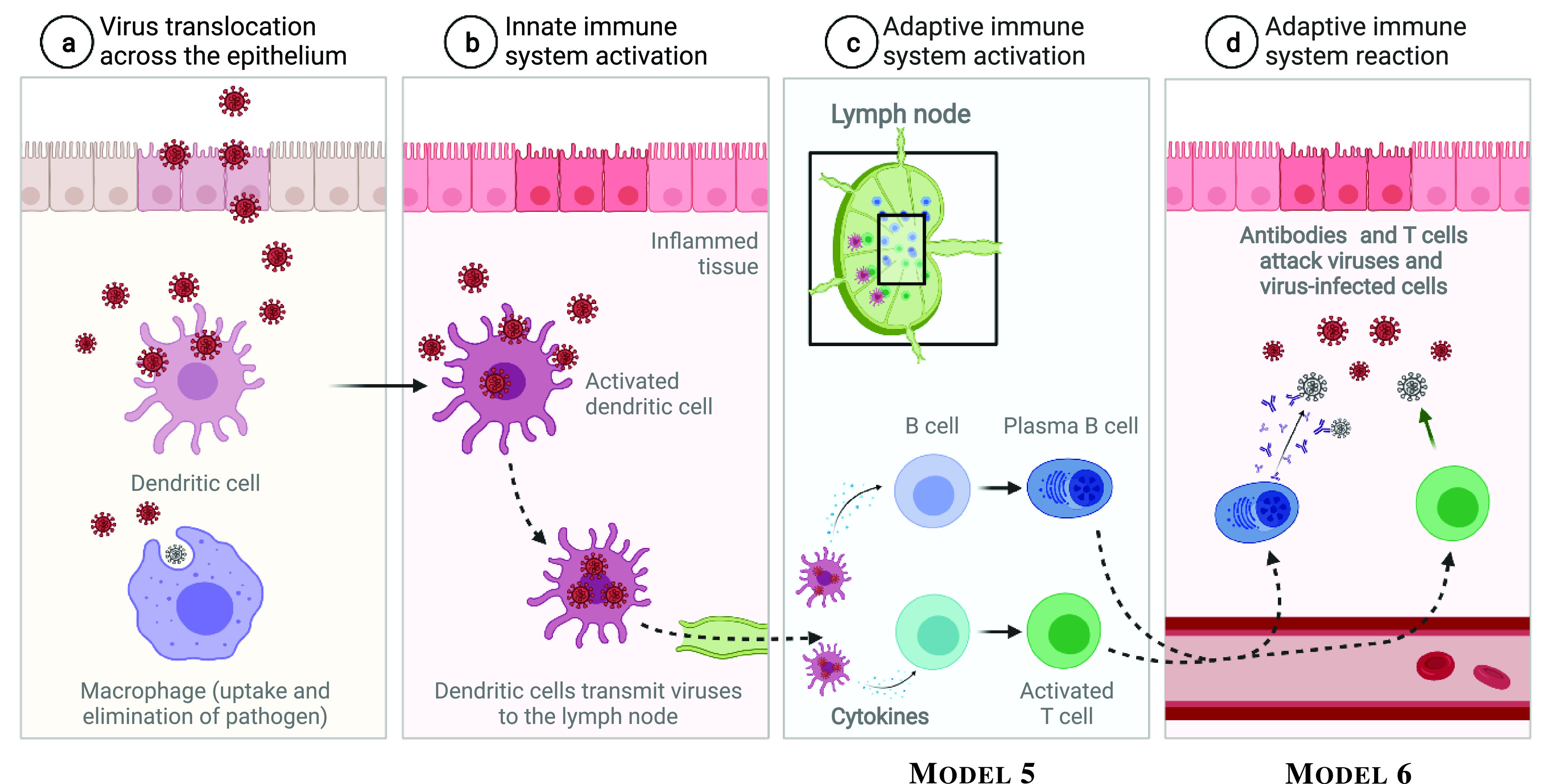
Activation of innate and adaptive immune systems. The cytokine-based- and antibody-based molecular communications systems are shown in Phase (c) and (d), respectively.
In the context of communications, the cytokines and antibodies are thus considered as information carriers, which propagate messages from the location of transmission until the location of the reception. The information conveyed by the cytokines and antibodies is infection reaction, as a response to infection action. For both Model 5 and Model 6, solutions to
 and
and
 should consider the types of specific immune markers and antibodies per virus as described in Table IV. These values can be used as initial reference values to lead the parameter fitting in the model for predicting the time progression of the innate immune response and the adaptive immune response. Since this model is relatively new, the integration of these values may lead to the development of further modelling that is not covered in this paper.
should consider the types of specific immune markers and antibodies per virus as described in Table IV. These values can be used as initial reference values to lead the parameter fitting in the model for predicting the time progression of the innate immune response and the adaptive immune response. Since this model is relatively new, the integration of these values may lead to the development of further modelling that is not covered in this paper.
TABLE IV. Viral Infection Impact and Immune Markers for Entry Points.
| Type | Infection impact | Specific immune markers | Specific antibodies | Ref. |
|---|---|---|---|---|
| SARS-CoV (1–2) | Initiates by the attachment of ACE2 glycoprotein to host receptors | SARS-CoV-2: Lower CD4, CD8, NK cell counts, PD-1, Tim-3 on T cells, Phenotype change in monocytes, IP-10, MCP-3, IL-1RA, IL-6, IL-8, IL-10, IL-2R, IL-1
 , IL-4, IL-18, GM-CSF, IL-2 and IFN- , IL-4, IL-18, GM-CSF, IL-2 and IFN-

|
Neutralising IgG antibodies that target internal N protein or external S glycoprotein (RBD) | [74], [75] |
| MERS-CoV | Initiates by the attachment of a glycoprotein to host DPP4 receptor | IFN-
 , IL-12, IL-17, IFN- , IL-12, IL-17, IFN-

|
RBD of S protein | [76] |
| EBOV | Initiates by the attachment of glycoprotein to host receptors | IL-1
 (high), TNF- (high), TNF-
 (high), IL-6 (high), IL-1RA, sTNF-R (high), SIL-6R, MIP-1 (high), IL-6 (high), IL-1RA, sTNF-R (high), SIL-6R, MIP-1
 , MIP-1 , MIP-1
 , Neopterin, SAA, NO2-, Cortisol, IL-10 , Neopterin, SAA, NO2-, Cortisol, IL-10 |
Epitopes on the glycoprotein (RBD) | [77], [78] |
| DENV | Initiates by glycoprotein E protein binding to DC-SIGN, a C-type lectin | IFN-
 , IL-6, IL-15, TNF- , IL-6, IL-15, TNF-
 , IL-1RA, CXCL8, CCL3, CCL4, CCL5, CCL11, M-CSF, G-CSF, ICAM-1, L-Selectin, P-Selectin, CD40, CD40L, Fas, FasL, TRAIL-R2, TRAIL-R3, RANK-L, C3a, IL-2R, TNF-R1 , IL-1RA, CXCL8, CCL3, CCL4, CCL5, CCL11, M-CSF, G-CSF, ICAM-1, L-Selectin, P-Selectin, CD40, CD40L, Fas, FasL, TRAIL-R2, TRAIL-R3, RANK-L, C3a, IL-2R, TNF-R1 |
Envelope glycoprotein (non-neutralising antibodies may cause ADE) | [79], [80] |
| ZIKV | Initiates by the attachment of the viral glycoprotein E protein to host receptors | CXCL10, ST2/IL-1R4 | IgM binds ZIKV pre-membrane and envelope glycoprotein | [79], [81] |
| HCV | Initiates by E2 protein interacting with cell surface heparan sulfate proteoglycans | CD127+CD8+ T cells, CD127-CD8+ T cells, CD161+CD8+ T cells, PD-1, Regulatory CD4+ and CD8 T cells, IL-10, CXCR3, CXCR6 | IgG binds HCV E1E2 envelope glycoprotein | [82], [83], [84] |
B. Extra-Body Molecular Communications Models
The airborne spread of infection is the main mechanism of human-human transmission of viruses. Once viruses are excreted into the air, they propagate towards another person that inhales them into its lungs. This mechanism allows the virus infection spreading to local or pandemic levels, which can occur in a matter of days. There are other modes of human-human transmission of viruses, including human contact transmission, or sexual transmission, but we do not explore these modes of transmission in this paper. Our objective in this section is to explore and analyse the airborne virus molecular communications system. It is comprised of the human excretion system as the virus transmitter, the propagation of the virus in the air as the channel, and the human respiratory system as the receiver.
1). Transmitter:
We consider the human as the source of virus transmission in the air. The infected humans excrete the virus with a particular concentration rate and velocity via the respiratory system. The respiratory system is composed of the nasal/oral cavity, pharynx, larynx, trachea and lungs, which are comprised of the bronchus and alveolus. Excretion of the virus starts from the alveolus, and propagation to the bronchus towards the nasal/oral cavity.
Recent works in the literature have been reporting several models that describe the release of particles or droplets in the air by the respiratory system. For example, the model presented in [19] explores the release of droplets by breath, sneeze and cough. The authors consider a rate model together with an event profiler to condition the rates of droplet release based on the three modes of transmission. The authors are interested in the steady-state derivations where these three modes converge to an averaged exhalation process. We believe that steady-state models do provide attractive mathematical solutions; however, they do not consider the generation process of the droplets and phenomena that influence the fate of the droplets apart from diffusion properties. In a similar goal but with a different approach, another initial transmitter model that analyses the air cloud produced by events driven by exhalation processes was proposed [63]. It withstands the same issues with the previous model, where the release rates neglect different phenomena that influence the droplets, and in this particular case, the cloud and its characteristics. The work developed in [64] showed that the droplets evolve inside a turbulent jet transitioning shortly to a puff. Ejected droplets are surrounded by a dynamically evolving air volume that is coupled to the droplet trajectory. While the major interest has been paid to static or averaged conditions of droplets, we argue that the literature fails to address (a) how the generation of droplets by the human body is coupled with existing modelling efforts and (b) how the conditions of the human body of the infected person impact the exhalation of the droplets in the air. Therefore, we advocate for the consideration of biological models, or variables adjusted from them, where the characterisation of the droplets is thoroughly considered while having a more detail process of how they are generated.
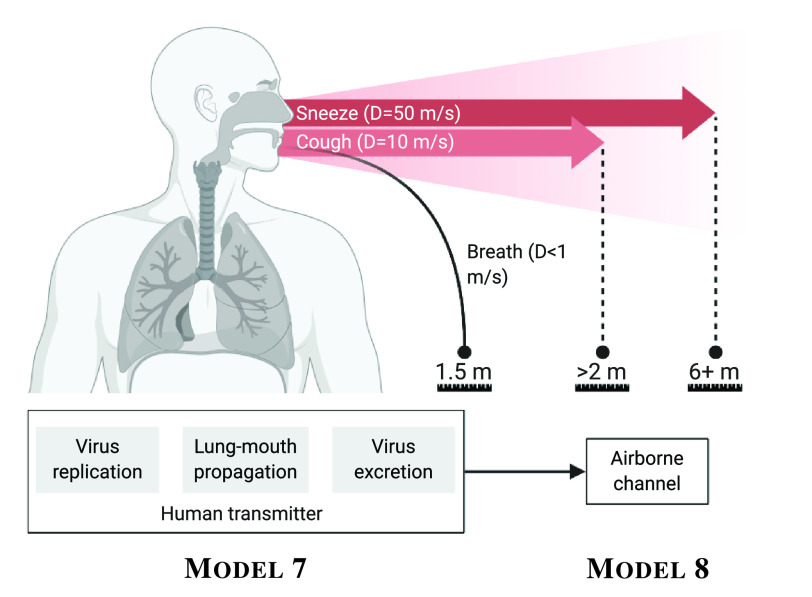
In this paper, we concentrate on the analysis of three different modes of transmission models for the virus excretion processes, which include breathing, coughing and sneezing, as depicted in Fig. 3. These modes dictate the initial properties of the virus propagation in the air, which alters the outcome of its range of propagation and the velocity that the virus diffuses. In [18], the authors compiled these modes of virus excretion under one umbrella and referred to them as exhaled breathing. However, more details should be provided as to how these different modes of transmission impact virus propagation. We explain more in the following, where we provide an initial molecular communications model of this transmission process which we refer to as Model 7.
Fig. 3.
Molecular communications model of a human transmitter of airborne viruses. The system is comprised of the virus replication, lung-mouth propagation and virus excretion phases. They dictate the rate and strength by which the virus is released to the environment that leads to different range in propagation distance. The sneeze, cough and breath are three different transmission modes for virus excretion.
We consider that the concentration of droplets
 released by the human transmitter is modelled as a convection process of virus concentration
released by the human transmitter is modelled as a convection process of virus concentration
 in the lungs and enters the nasal/oral cavity with rate
in the lungs and enters the nasal/oral cavity with rate
 , representing the virus excretion process. We define this process as follows:
, representing the virus excretion process. We define this process as follows:
 |
and,
 |
 is the initial concentration of virus in the nasal/oral cavity,
is the initial concentration of virus in the nasal/oral cavity,
 , is the rate of virus replication in the lungs,
, is the rate of virus replication in the lungs,
 is the propagation of virus from the lungs to the nasal/oral cavity, and
is the propagation of virus from the lungs to the nasal/oral cavity, and
 is the convolution operator. We do not explore this model in detail, since the detailed version can be found in [19], [64]. However, we acknowledge that
is the convolution operator. We do not explore this model in detail, since the detailed version can be found in [19], [64]. However, we acknowledge that
 is directly linked with the transmission modes that we discussed earlier (breathing, coughing and sneezing). Typically, these modes are always considered to affect the velocity of propagation of droplets. We argue that they are fundamental in the characterisation of virus conversion to droplets as well as the rate of released droplets themselves. Many approaches do consider
is directly linked with the transmission modes that we discussed earlier (breathing, coughing and sneezing). Typically, these modes are always considered to affect the velocity of propagation of droplets. We argue that they are fundamental in the characterisation of virus conversion to droplets as well as the rate of released droplets themselves. Many approaches do consider
 in the steady-state, but we like to draw attention by the community that it can have non-linear relationships with the infected humans, so it should also be associated with different disease stages over time that is bounded by transition probabilities of stage changes (severe stage to mild, and vice versa). Moreover, as shown in both (20) and (21), that non-linearities do exist, for example, the relationship
in the steady-state, but we like to draw attention by the community that it can have non-linear relationships with the infected humans, so it should also be associated with different disease stages over time that is bounded by transition probabilities of stage changes (severe stage to mild, and vice versa). Moreover, as shown in both (20) and (21), that non-linearities do exist, for example, the relationship
 is added to the model but is currently non-existing in the literature. The initial virus concentration
is added to the model but is currently non-existing in the literature. The initial virus concentration
 can be directly obtained from values in Table I. Also
can be directly obtained from values in Table I. Also
 and
and
 can be obtained from [65], [66].
can be obtained from [65], [66].
2). Channel:
Droplets travel in the air following diffusion properties bounded by airflow properties. For example, the modes of transmission discussed above can impact on different types of turbulent flows that lead to a puff scenario, further resulting in airflow forces that are weakened and gravitational forces on the droplet to get stronger. These droplets tend to travel a few meters away from the human excretion point (transmitter), which takes around several seconds, and can reach the human receiving points (e.g., nose or mouth) up to 6 meters of distance. Molecular communications channel models can be used to model these effects of droplet concentration release in the air, and possibly be used to characterise the number of delivered droplets to the human receiving points. In this section, we review existing models and analyse a general channel model.
We summarise the literature on existing droplet propagation models with certain properties of the droplet and airborne viral particles in Table II. Even though they are not entirely classified as pure molecular communications channels, they present not only the physical modelling of droplet propagation but also the effects of droplet propagation and thus can be considered by the community for characterising Molecular Communication Systems. We analyse the literature in terms of the completeness of the physics that govern the droplet propagation. First, we look at their modes of propagation, either air-based or molecular-based. These modes dictate the way these models are constructed. Then, we classify each type of medium that can be utilised for modelling these molecular communications approaches. The types of species include airflow behaviour with average droplet concentration (transient air, air cloud) to more focused on the characterisation of the number of droplets (single, distribution, concentration and concentration/rate). We also analyse the airflow properties that are mostly secreted from a person as the transmitting point, this includes turbulent flow and puff flow. Turbulent flow accounts for the advection-diffusion of particles that are influenced by a force, in this case, the air turbulence and flow created from the transmitting point. The puff flow can be regarded as the Brownian diffusion in the air and can be influenced by gravity. Lastly, we analyse the properties of the environment that affects the state of droplets once they are excreted and this includes evaporation and crystallisation. Since the majority of the droplets is comprised of water, it is subject to effects from temperature change that can result in evaporation, as well as the quantity of salt in the droplet that lead to its crystallisation. From Table II, we observe that all species comply with the puff and turbulent flow ([19], [67], [69], [70]). However, we do recognise that environmental effects on the droplets are not fully explored for molecular communications models. The environmental effects have a significant impact on the propagation of the droplets, as it can either impact (a) on the flow behaviour in space, or (b) on the rate of virus reception by the receiving organ (e.g., nose or lung). Besides future investigation in environmental effects on the droplet propagation, there are also needs for further investigation on the effects of jet streams that affect the viral propagation behaviour. This includes understanding the aerodynamic airflow within confined and open areas and how this affects the flow of the viral particle propagation.
TABLE II. Literature Review Summary on Channel Models for Virus Air Propagation.
We now describe the propagation model of airborne droplets, which we refer to as Model 8. We assume the source is located at
 and emits droplets with rate
and emits droplets with rate
 . Based on the Fick’s second law of diffusion, we consider a droplet concentration varying over time with
. Based on the Fick’s second law of diffusion, we consider a droplet concentration varying over time with
 |
where
 is the mass, and
is the mass, and
 is the droplet degradation loss derived from environmental effects. There are many approaches to derive
is the droplet degradation loss derived from environmental effects. There are many approaches to derive
 , and this largely depends on the environment. For example, the authors in [19] focus on expanding the term to include Fick’s advection and diffusion effects on the flow. We understand that those terms represent the turbulent flow and puff flow, respectively. On the other hand, [67] presents a more complex model based on the aerodynamics of the airflow, which precisely addresses the turbulent and jet flows that drive droplet propagation. The authors present the validation of their model using experimental data.
, and this largely depends on the environment. For example, the authors in [19] focus on expanding the term to include Fick’s advection and diffusion effects on the flow. We understand that those terms represent the turbulent flow and puff flow, respectively. On the other hand, [67] presents a more complex model based on the aerodynamics of the airflow, which precisely addresses the turbulent and jet flows that drive droplet propagation. The authors present the validation of their model using experimental data.
We show in (22) that
 influences directly the propagation of droplets from environmental effects such as evaporation or crystallisation. This is an interesting effect where the temperature, water content, and the salt crystals concentration in the droplet jointly impact diffusion and the rates or concentration when it reaches the receiver. Even though this phenomenon has been explored before [68], [69], the authors of [67] investigated this effect, where they derived a model that couples aerodynamic properties and environmental effects on the droplet propagation behaviour. Their model was also validated based on imaging experiments using an ultrasonic levitator to capture transient dynamics of evaporation and precipitation of the evaporating droplet.
influences directly the propagation of droplets from environmental effects such as evaporation or crystallisation. This is an interesting effect where the temperature, water content, and the salt crystals concentration in the droplet jointly impact diffusion and the rates or concentration when it reaches the receiver. Even though this phenomenon has been explored before [68], [69], the authors of [67] investigated this effect, where they derived a model that couples aerodynamic properties and environmental effects on the droplet propagation behaviour. Their model was also validated based on imaging experiments using an ultrasonic levitator to capture transient dynamics of evaporation and precipitation of the evaporating droplet.
One of the main advantages of modelling virus propagation using the molecular communications paradigm is the derivation of particle propagation rates, which can be used to study the viral entry mechanism into a human receiver. In [19], the authors utilised the solution in (22), by breaking down diffusion components into three dimensions
 , and algebraically developing the diffusive and mass matrices based on wind flow. To derive a closed-form solution, the authors considered the steady-state response of the breathing process, where they obtain a closed-form Green’s function. The obtained closed-form solution for the droplet concentration is represented as
, and algebraically developing the diffusive and mass matrices based on wind flow. To derive a closed-form solution, the authors considered the steady-state response of the breathing process, where they obtain a closed-form Green’s function. The obtained closed-form solution for the droplet concentration is represented as
 |
where
 is the turbulence indicator due to wind sources,
is the turbulence indicator due to wind sources,
 is the height of a person’s mouth to the ground, and
is the height of a person’s mouth to the ground, and
 is the flow velocity in the
is the flow velocity in the
 dimension.
dimension.
These models are attractive from the propagation and system theory point of view. However, the authors do not explore practical results in terms of infection. During exposure to an infected human transmitter, several variables dictate the fate of the human receiver, possibly leading to another infection, and this can include the time of exposure as well as the distance between the person emitting the droplets and the human receiver. The authors in [63] present an analysis on this end-to-end scenario, where the probabilities of infection at the human receiver are evaluated in terms of distance, time of exposure, and coughing angle. As droplets propagation is bounded by the angle of release, it is also important to incorporate spatial analysis on the infection probability, such as the coughing angle.
3). Receiver:
In the human receiver, the airborne droplets are absorbed by the nasal-mouth cavity and propagate along the respiratory tract until they reach the alveoli. During this path, the virus undergoes replication, penetrating deep into the epithelial tissues, and then into the circulatory system to infect other organs and systems. In this section, we focus on the high-level modelling and analysis focused on the infection process through the human respiratory tract virus propagation.
Modelling the human receiver using a systems theory approach can be increasingly complex since there are two main factors that, individually, comprise of several steps. First, there is the entrance of the droplets containing viruses into the human body and residing in the lungs. Secondly, there is the infection process of internal progression. The model for the viral entry can be found in [20], which accounts for virus dispersion along the respiratory tract impacted by factors such as the respiratory rate, viral exposure levels (i.e., the quantity of virus that is inhaled), and the virus particle size. The authors developed a computational model to account for the changes in the virus propagation as it enters and propagates in the respiratory tract. For the virus infection process, [20] also considers the impact of the immune system and how it influences the overall concentration of the virus within the respiratory tract. The model in [63] focuses on a high-level generic approach based on probability of infection calculated from the probability of reception of droplets containing the virus. Both models are different in terms of biological details or probabilistic infection estimation solutions. We not only support their integration, and creating a scenario where the probability of infection is dependent on increased biology realism, but also for these models to contribute towards modelling the progression of infection and state of infectivity. Understanding the end-to-end propagation to the infectivity process is a crucial contribution for researchers, because this can lead to insights of molecular interactions at the cellular level and the impact of the infection on subsystems of the body, which in turn can lead to precision medicine in clinical decisions for treatments and patient recovery guidelines.
Our analysis on the viral impact on the human receiver model is depicted in Fig. 4, which we refer to as Model 9. There are three main blocks in the model: the droplets entrance rate, the mouth-lung propagation, and the virus replication and deposition rates. The general model of virus propagation is based on a governing equation based on the Fick’s second law for advection-diffusion investigated in [20]. This model accounts for the concentration of the virus in a particular branch of the respiratory tract
 , and can be represented as
, and can be represented as
 |
where
 is the generation number (i = 0, 1, 2, 3,
is the generation number (i = 0, 1, 2, 3,
 , 23) of the lung branches,
, 23) of the lung branches,
 is the virus deposition rate,
is the virus deposition rate,
 is the virus replication rate,
is the virus replication rate,
 is the diffusion coefficient, and
is the diffusion coefficient, and
 is the direction of the virus propagation (downward or upward in the respiratory tract). The
is the direction of the virus propagation (downward or upward in the respiratory tract). The
 represents the droplet entrance rate. Fig. 4d and Fig. 4e depicts the replication and deposition of the virus, respectively. The authors in [20] also explored the dynamics change in the viral pleomorphic size changes during the propagation. The advection-diffusion component is used to model the breathing process to understand the airflow into the lung. Even though they concentrate on COVID-19, it is clear that their proposed model can be applied to other virus types that propagate through the respiratory tract. For the future, such models may need not only experimental validation, but further integration with infection development itself. As shown in Fig. 4c, the changes in the virus infection process for SARS-CoV-2 (e.g., from moderate to severe), can present changes in the volume of mucus present in the alveolus and the entire respiratory tract. This infection process can not only produce changes in the breathing rhythms and cough rate, but it can also change the advection-diffusion process within the lung generation. As the virus penetrate areas with a high quantity of mucus, the propagation medium changes enough to make the advection property of virus flow to dramatically reduced. Moreover, the deposition and replication rates changes within the mucus areas. This, in turn, can affect the binding process of the virus, which is different for pure diffusion in air pockets compared to a space with mucus. One interesting development could be the investigation of stochastic diffusion coefficients, deposition rates and replication rates of the virus, and how this evolves as the mucus production increases using a multi-medium molecular communications model. The initial virus concentration
represents the droplet entrance rate. Fig. 4d and Fig. 4e depicts the replication and deposition of the virus, respectively. The authors in [20] also explored the dynamics change in the viral pleomorphic size changes during the propagation. The advection-diffusion component is used to model the breathing process to understand the airflow into the lung. Even though they concentrate on COVID-19, it is clear that their proposed model can be applied to other virus types that propagate through the respiratory tract. For the future, such models may need not only experimental validation, but further integration with infection development itself. As shown in Fig. 4c, the changes in the virus infection process for SARS-CoV-2 (e.g., from moderate to severe), can present changes in the volume of mucus present in the alveolus and the entire respiratory tract. This infection process can not only produce changes in the breathing rhythms and cough rate, but it can also change the advection-diffusion process within the lung generation. As the virus penetrate areas with a high quantity of mucus, the propagation medium changes enough to make the advection property of virus flow to dramatically reduced. Moreover, the deposition and replication rates changes within the mucus areas. This, in turn, can affect the binding process of the virus, which is different for pure diffusion in air pockets compared to a space with mucus. One interesting development could be the investigation of stochastic diffusion coefficients, deposition rates and replication rates of the virus, and how this evolves as the mucus production increases using a multi-medium molecular communications model. The initial virus concentration
 can be directly obtained from values in Table I. Also
can be directly obtained from values in Table I. Also
 ,
,
 and
and
 can be obtained from [65], [66].
can be obtained from [65], [66].
Fig. 4.
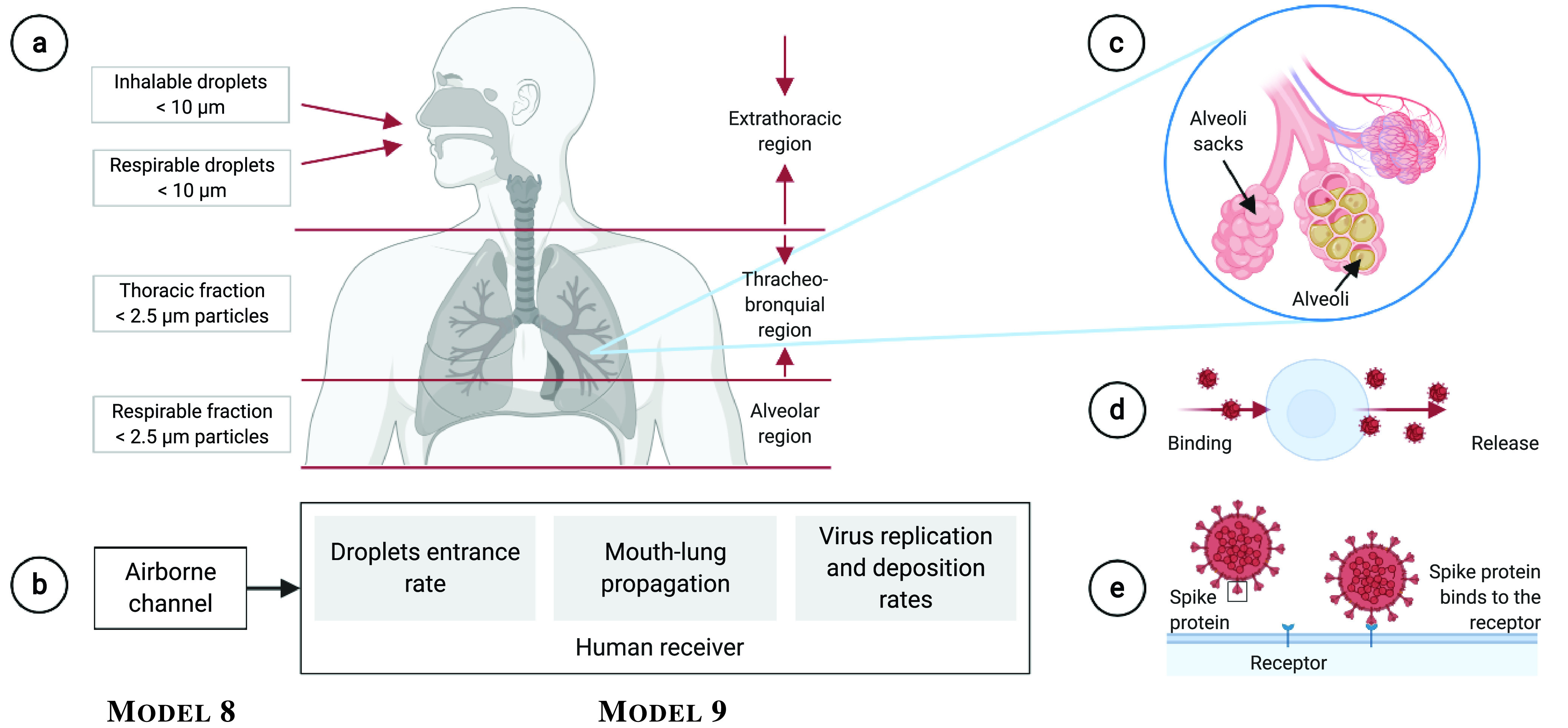
Molecular communications model of the human receiver of airborne viruses. a) According to the different regions in the respiratory tract, the size of the particles propagating downwards is smaller; b) The human receiver model is comprised of droplets entrance rate, mouth-lung propagation, virus replication and deposition rates; c) Alveoli sack and alveoli with moderate and severe mucus presence due to infection progression; d) The virus duplication process; e) The virus deposition process.
IV. Leveraging Experimental Data for Molecular Communications
The molecular communications community has been abstracting cellular signalling for more than a decade of active research, replicating the characterisation of the functions of wired and wireless networking and computing systems. However, with only a few experimental demonstration systems reported to date [71], [72], the molecular communications field generally lacks validation aspects. We anticipate similar challenges to arise when building molecular communications models of the virus intra-body and extra-body propagation, which are critically needed to understand virus dynamics and unveil new insights that will increase our understanding of virus pathogenesis and enable spread and infection patterns to be more predictable in vivo. To characterise the viral dynamics and evolution, one of the initial generic computational modellings demonstrated the necessity to integrate biophysical models and infection properties [20]. In this section, we discuss many sources of open data that can be used to be incorporated into the models presented in the previous section, and that can inspire the molecular communications community to produce new models. The link between the presented models and the experimental data is summarised in Table III, where we present what experimental data should be used for each model and where to find such data.
TABLE III. Summary of the Link Between the Presented Models and Experimental Data.
| Model | Experimental Data | Availability |
|---|---|---|
| 1 | Airflow profile, diameter and length of each airway, virus concentration profiles | Table I, [65] |
| 2 | Blood velocity profile, diameter and length of each vessel generation, virus concentration profiles | Table I, [73] |
| 3 | Virus proliferation rates | None |
| 4 | Virus proliferation rates | None |
| 5 | Cytokine and antibody concentration levels | Table IV |
| 6 | Cytokine and antibody concentration levels | Table IV |
| 7 | Airflow profile, diameter and length of each airway, ACE2 receptor biding rate | Table I, [65], [66] |
| 8 | Airflow profile, diameter and length of each airway | Table I, [65] |
| 9 | Airflow profile, diameter and length of each airway, blood velocity profile, diameter and length of each vessel, genome, proteome, RNA transcripts, ACE2 receptor biding rate | Table I, [65], [73], [66] |
Biophysical models present real physiological parameters associated with the physical space where the virus propagates through. These parameters are typically available in the literature to be readily used by the molecular communications community for in silico modelling. The examples include the analysis of entry and spread of SARS-CoV (1–2) and MERS-CoV in the respiratory system, where Model 1 and Model 7–9 require values for the airflow profile, and diameter and length of each airway generation available from [65]. For the analysis of entry and spread of DENV, ZIKV and HCV in the circulatory system, Model 2 AND Model 9 require values for the blood velocity profile, diameter and length of each vessel generation available from [73].
The basic set of infection properties includes viral exposure levels (in different specimens), virion characteristics, and how viruses interact with cells and the immune system. We summarise relevant data for SARS-CoV (1–2), MERS-CoV, EBOV, DENV, ZIKV and HCV in Table I and Table IV. Apart from exposure levels and virion characteristics (including their genome, proteome and RNA transcripts), Table I identifies detection methods, whereas Table IV outlines the infection impact and immune markers for each of the considered viruses. This information is necessary for the effects on the receiver communication models, or Model 9, where the interactions between virus-host cells are found.
Based on the available data, at least the concentration profiles of each of these viruses given in Table I, SARS-CoV (1–2), MERS-CoV and ZIKV translocation across the epithelium, or Model 1, and EBOV, DENV, ZIKV and HCV spread via the bloodstream, or Model 2, can be initially analysed. One way of doing such analyses is to associate the available concentration profiles given in Table I with
 given in (1) of Model 1 and
given in (1) of Model 1 and
 given in (7) of Model 2, respectively, and assume initial values for other relevant concentration profiles occurring in (1)–(3) and (7)–(11). In addition, the parametric values for the virus interaction with the host cells and immune system cells, e.g., forward- and backward reaction rates, are still unavailable and need to be assumed. Obtaining these values from the experiments is challenging but can lead to accurate predictions of viral dynamics.
given in (7) of Model 2, respectively, and assume initial values for other relevant concentration profiles occurring in (1)–(3) and (7)–(11). In addition, the parametric values for the virus interaction with the host cells and immune system cells, e.g., forward- and backward reaction rates, are still unavailable and need to be assumed. Obtaining these values from the experiments is challenging but can lead to accurate predictions of viral dynamics.
More advanced modelling approaches also operate with the host cell surface receptor distribution, host cell distribution, replication and deposition rates, and immune-response-relevant parameters. For example, in the case of SARS-CoV-2, the virions use the ACE2 receptor to bind to and enter host cells (Table IV), which is important for Model 7 and Model 9. The density of ACE2-expressing host cells is modelled in the literature to follow the Gaussian distribution (e.g.,
 (Copies/mL) [66]). Spatial distributions additionally complicate the molecular communications modelling since ACE2-expressing host cells are not spread evenly, thus creating a heterogeneous concentration distribution across the respiratory system [20]. Other parameters relevant for the target-cell model (presented in Section III-A) that facilitate binding by SARS-CoV-2 virus in the respiratory systems can be found in [85], [86]. The overview of the infection impact of other considered viruses is also given in Table IV.
(Copies/mL) [66]). Spatial distributions additionally complicate the molecular communications modelling since ACE2-expressing host cells are not spread evenly, thus creating a heterogeneous concentration distribution across the respiratory system [20]. Other parameters relevant for the target-cell model (presented in Section III-A) that facilitate binding by SARS-CoV-2 virus in the respiratory systems can be found in [85], [86]. The overview of the infection impact of other considered viruses is also given in Table IV.
In the case of virus proliferation via neurons and EVs, no relevant data for Model 3 and Model 4 have been collected in the tables. We thus advocate for the need to conduct relevant experiments and back up the corresponding molecular communications models that are yet to be developed. As of the immune system reaction via cytokines and antibodies, we list the relevant immune markers and antibodies for each of the considered viruses in Table IV. The corresponding concentration levels are typically available in the literature, e.g., cytokine levels in SARS-CoV [87], [88], [89], and can be used to support Model 5 and Model 6.
The airborne virus propagation models through droplets based on Model 8 need a set of data to characterise diffusion properties when a human transmitter sneezes, coughs or talks. The availability of this data is a critical issue as we found very limited resources that can be used. However, existing testbeds can be used to generate data that can be used by the molecular communications researchers. A particular testbed that is appropriate for this scenario is the Tabletop Molecular Communication
[72], where the authors presented the release of isopropyl alcohol molecules through an electronic-activated spray. These airborne molecules flow towards an alcohol sensor through the airflow produced by a fan. The authors demonstrate the successful encoding, transmission and reception of information encoded using molecules concentration. This testbed can be used to study the effects of droplet propagation, i.e., coupling it with Model 8. The usage of fans can provide modifications to the airflow that drives the propagation of the droplets, and hence air jet streams found in different indoor or outdoor scenarios. This would help estimate parameters for velocity and turbulence measures, such as
 and
and
 in (23). The production of droplets is not clear, especially when trying to emulate events of sneeze, cough and speech. However, we believe the testbed can be extended to include different versions of the electronic-activated spray that can modulate the release rate of molecules, for example. In that way, parameter
in (23). The production of droplets is not clear, especially when trying to emulate events of sneeze, cough and speech. However, we believe the testbed can be extended to include different versions of the electronic-activated spray that can modulate the release rate of molecules, for example. In that way, parameter
 in (22) would be associated with a proper virus release rate profile in each emulated event. Lastly, the propagation of the droplets itself can be used to determine other model parameters including the droplet degradation loss derived from environmental effects, which is
in (22) would be associated with a proper virus release rate profile in each emulated event. Lastly, the propagation of the droplets itself can be used to determine other model parameters including the droplet degradation loss derived from environmental effects, which is
 in (22). The value of
in (22). The value of
 can be broken down into different components as shown in Table II, where the testbed can be used to estimate
can be broken down into different components as shown in Table II, where the testbed can be used to estimate
 .
.
Other sources of open data, specifically for COVID-19, include studies presented in [90] and [91]. They offer a wide range of information for genetic sequences for both viruses and hosts, genetic expressions, protein, biochemistry and even imaging. Apart from the latter, all others can be integrated into all models discussed in this paper. On the other hand, the position paper given in [92] summarises a large number of studies on the formation of virus-laden aerosol particles and their spread. This paper presents many interesting effects to aerosol propagation that were not discussed here, including ventilation systems and the effectiveness of the usage of masks. Lastly, the dataset presented in [93] explores the usage of laser particle counter to measure the emission of aerosol particles during speaking, singing and shouting activities.
V. Open Issues and Challenges
In this section, we explore the open issues and challenges derived from the discussion presented in our survey so far. Our vision is that molecular communications aligns itself further with the area of bioengineering so that models of methods can be verified by experiments, working in close collaboration with relevant experts. We explore linking of experimental data to molecular communications models, novel intra-body viral intervention techniques, emerging technologies for infection theranostics, bridging molecular communications and bioinformatics, and novel molecular communications models.
A. Linking Experimental Data to Molecular Communications Models
As discussed, molecular communications needs to provide further integration of its models with experimental data, either already available or through novel experiments. The validation of existing models will be of major benefit to the community as it helps calibrate the works already developed, to be directly applicable to the interpretation or prediction of in vitro or even in vivo phenomena. Without validation, molecular communications models are likely prevented from expanding its usage to other areas of potential applications. There are already some interesting works of how molecular communications and experimental data can be integrated into different scenarios and for different applications. For example, models with experimental data can be found for calcium signalling [94], [95], bacteria communication [15], [96], [97], neural communication [98], and macro particle diffusion [72], [99]. Other works have made great usage of experimental data, such as data extracted from open source databases and integrated into their Molecular Communication models, and examples include works in [100], [101].
The reader must consider referring to all the data provided in Section IV and the references provided for data acquisition for new models of infection using the molecular communications paradigm. Specifically, transmitter designs would benefit from the process of genetic information encoding with data from Table I. Propagation of viruses can be based on the diffusion information in different medium provided in the same table. Lastly, the receiver design directly benefits from the information provided in both tables. If the receivers are based on electronic technology, the reader is referred to Table I, but if the receivers are actual biological-based devices (e.g., lung cells), the reader is referred to Table IV. However, the data provided here are not sufficient to derive all the necessary variables for required non-linear models with complex behaviour, and for that, the reader should consider them as a guiding basis for the correct understanding of the communication parts involved in molecular communications for infections diseases.
B. Novel Intra-Body Viral Intervention Techniques
Even with the existing efforts in designing vaccination and drugs for viral infections treatment, there is still a need for novel interventions requiring robust intra-body solutions [3]. The main reasons are twofold: 1) the effectiveness of vaccines is not always optimal, and 2) there are consequences to tissues and organs during the infection that might require repairing procedures [3]. Stem cell-based treatment is a novel intra-body solution that has been argued as a leading technology for future intervention. This technology is used in different ways, e.g., can be used for drug delivery, and as regenerative therapy (studies have shown that stem cells can modulate the immune system for patients suffering from SARS-CoV-2 infection). Particular stem cells can be derived from many tissues types, including umbilical cord bone marrow, trabecular bone, synovial membrane, and fetal tissues such as lung, pancreas, spleen, liver, etc. [102], [103], [104]. By interacting with the media through chemical agents, stem cells can eliminate existing pathogenic behaviours and repair the tissue or organ.
More specifically, Mesenchymal stem cell-based approaches have been proposed for interventions in many viral infections, including hepatitis [3], [105], and SARS-CoV [106], [107]. Mesenchymal stem cells are adherent, Fibroblast-like cells with the ability of self-renewal and differentiation into multiple cell lineages such as Osteoblasts, Chondrocytes, Adipocytes, and Hepatocytes. Looking at specific lung-damaging infections, Mesenchymal stem cells can secrete IL-10, hepatocyte growth factor, Keratinocytes growth factor, and VEGF to alleviate Acute Respiratory Distress Syndrome (ARDS), regenerate and repair lung damage and resist Fibrosis [106], [108], [109]. This opens new possibilities for the development of novel molecular communications solutions that look at communications between the Mesenchymal stem cells signalling and its process of repairing damaged tissue from the communication process. Similar approaches have been proposed in the community, where Exosome Vesicles (EVs) were used to model the interactions between stem cells and Glioblastoma [51], [60]. Even though EVs are also suggested for the treatment of lung damage [106], we recognise that different types of signalling information carriers can be used to create a multi-carrier, or multi-molecules communication system, serving as ways to increase the overall treatment capacity.
C. Emerging Technologies for Infection Theranostics
In the intersection of fields such as bioengineering, material sciences, and medical sciences lies the development of innovative technology that can lead to efficient treatment or diagnosis of infectious diseases, herein referred to as infection theranostics. Molecular communications can play a role in these emerging technologies by integrating communications properties of transmission, propagation and reception. For example, microfluidic-based organ-on-a-chip devices provide experimental models of transmission, propagation and reception of molecules from cell-cell, tissue-tissue and even organ-organ, with or without other external molecular agents [110]. Another example is the airborne viral detection from biosensors that can be either integrated into all-in-one devices [18] or integrated biosensors with proposed emerging infrastructures for 6G, such as the Intelligent Reflecting Surface [111]. In this case, virus macro-scale propagation and its reception, through either binding-ligand proteins or electrically charged droplets, can include modelling expertise from molecular communications researchers to develop such infrastructures. In the following, we dive deeper into this topic using the above-mentioned examples.
Microfluidic-based organ-on-a-chip are alternative experimental models compared to conventional in vitro and animal models, since they capture many tissue structures that are found in human organs [112]. The presented literature investigates how organ-on-a-chip can be used to study virus-host interactions, viral therapy-resistance evolution, and development of new antiviral therapeutics, as well as underlying pathogenesis. This can be applied to different organs of the human body; for example, infection models have been applied to liver chip [113], gut chips [114], [115], neural chips [116] and lung chips [117], [118]. Molecular communications models of microfluidic-based organ-on-a-chip can be used to develop future infection assays to study virus-induced diseases in real-time and at high resolution. Molecular communications can aid in inferring methods of disease communication and progression by analysing how cellular molecular functions operate in both healthy as well as infectious states. This can be further extended to design novel molecular modulation mechanisms are used either to augment cellular communication or to understand effects from external molecular signals, such as a viral drug. Within the molecular communications community, there are a number of existing research on microfluidics modelling and experiments [119], [120], and this can be extended to utilise organ-on-a-chip devices for viral theranostics.
Molecular communications models can be used to design the sensitivity of the binding process for airborne viral detection technology. The key principle is to couple the modelling of air particle flow (as discussed in Section III-B) with the receiver design, which in this case is an airborne viral biosensor. These biosensors can be built in a number of ways, including ligand-binding protein receptors [111], electrically charged particles [18], CMOS-coupled immunological assays, and even through Polymerase Chain Reaction (PCR) technology [121]. The receiver design for biosensor technology based on a realistic propagation model of viral particles is missing. For example, in an outdoor space, the propagation of the airborne viral particles can undergo stochastic propagation patterns due to the changes in airflow directions and turbulence. Therefore, the design of the biosensor receptor structure can incorporate molecular communications model to help enhance the design of the sensors that is appropriate for the specific environment. However, this will require considerable experimental work where molecular propagation flow can be studied with fluorescent technology, similar to the works proposed in [67], [122], [123], in order to characterise the propagation patterns.
D. Bridging Molecular Communications and Bioinformatics Tools
In this paper, we mostly investigated the Molecular Communication models of viral infection, but in Section IV, we explored the link between these models to currently available experimental data for viral-host genetic interactions. There exist many interesting works that cover the analysis of these interactions coming from a bioinformatics perspective, which can analyse a large number of protein interactions and how they affect cells activity and fate [124], [125], [126].
Bioinformatics is a very advanced area in the analysis of genomics, as previously noted, with several tools and data available for researchers who desire to investigate the genetic relationships of viruses and host cells. Based on the genomic sequencing, these tools can provide the assembly of fragmented sequenced data, phylogenetic analysis of taxonomic groups, identification of genetic structures, identification of domains, functions and metabolic pathways, as well as data sharing capabilities [124]. Researchers in the area are hoping that, by having open-shared data and tools access policy, they facilitate the discovery of targeted genes and what leads to cell behaviour [127]. Some works focus on the discovery of new drugs or even vaccines [125], [126]. Even though some exciting works related to the topic of genomics are presented in the molecular communications field [17], [128], [129], the total and practical integration of molecular communications and bioinformatics tools are far from being explored. More works linking genomics with cell behaviour can solve existing issues, including the characterisation of natural cell signalling modulation techniques, sources of noise and interference, encoding/decoding of molecular information, and synthetic molecular communications. The bioinformatics existing tools already provide the information on the cell genetics and, sometimes, linkage to cell behaviour. Seamless integration with molecular communications leads to studies such as on the interaction of viral DNA content [17] and techniques for linking DNA exchange between bacteria for bacteria-based molecular communications [128]. Looking at the various models and systems presented in this paper, we advocate that the viral genetic interactions with host cells can provide novel understandings on how infected cells propagate the genetic content to other cells from an information and communication theory approach, while at the same time also considering the evolution of the genetic content, which can provide novel mechanisms for inferring infection propagation inside the body.
E. Novel Molecular Communications Models
New advancements in molecular communications modelling are needed, from this point in time forward, to help understand the virus propagation both intra-body and extra-body and the end-to-end communication system. For intra-body, the link between virus replication and tissue response has yet to be thoroughly investigated. For example, the interactions of the virus with neural communications [13], [130], or calcium signalling in different tissues or organs (especially the epithelium) [131], [132], can provide further understanding between the virus and hosts interactions using molecular signals as infection information carriers. As we know from the literature, the human immune response is triggered after the human body recognises the presence of infectious agents, foreign to the body itself [133]. The immune response is yet to be explored using concepts from molecular communications, similar to the models initially discussed in Section II, and to tie this to other Molecular Communication models (e.g., propagation within the circulatory system), in order to create an end-to-end system model. The key benefit of these studies is the ability to accurately capture the effects of viral propagation communication on the immune response communication, which would create computational models that can benefit vaccine designers in the future if the various sub-communication systems and interactions are well understood. These new models could be used to analyse new techniques to modulate the immune system response, coming from a regenerative medical intervention, based on the precise calculation of infections stages derived from virus-host interactions, in order to lead to an end-to-end diagnostic and therapeutic strategy for vaccine development.
For extra-body models, where models have recently been introduced on airborne droplet propagation, there are further developments required. First, transmitter models for extra-body molecular communications should consider in more detail the conditions of the human transmitter, i.e., the level of infection and condition of the respiratory system. This understanding can provide new relationships between the virus release rates from the host, and the analysis of viral reception concentration by another host. In the receiver, as explored in Section III-A, the need to couple models of the propagation of the respiratory tract, with actual rates derived from the airborne droplet propagation models are needed. This will contribute towards and end-to-end model that considers the Molecular Communication airflow propagation coupled with environmental effects on the droplets, and to link this with the intra-body propagation of the virus into the lungs. Even though an end-to-end analysis of these systems can lead to increasingly complex models, they can be useful in accurately predicting infection spreading patterns, and how this can impact on people with different health conditions, in order to personalise and classify their risk levels. Such accurate modelling can prevent total lockdowns for the entire society, where people in different categories can be allowed into society provided they take certain protective measures.
VI. Conclusion
Molecular communications can play a significant role in viral infectious disease research, by considering the detail characterisation of the transmission, reception and propagation of viruses inside and outside the human body. We provided an extensive review of the existing literature on the topic, by analysing the existing models for intra-body and extra-body molecular communications. For intra-body models, we explored the viral translocation across the epithelial and endothelial barriers, neurotropic viral spread, EV-based viral spread, cytokine-based- and antibody-based molecular communications. In extra-body models, we analysed models for the transmission process of viruses spelt from a human transmitter, the airborne droplet and virus propagation coupled with many environmental effects including turbulent flow, puff flow, droplet evaporation and droplet crystallisation, and finally, the reception process of viruses in the human receiver. Besides, we reviewed models for the virus entrance mechanism focusing on the virus concentration in the respiratory tract. We showed how the available experimental data can be integrated into molecular communications models, and what are the open issues and future directions. We are looking forward to exciting new research that can come as an output of the interdisciplinary works using molecular communications for developing new methods for treatment of infection as well as vaccination methods. Based on the analysis provided in the paper, we are confident that fantastic novel research can emerge and help in the fight against the current and future pandemics.
Biographies
Michael Taynnan Barros (Member, IEEE) received the Ph.D. degree in telecommunication software from the Waterford Institute of Technology in 2016. He is a Marie Skłodowska Curie Individual Fellowship (MSCA-IF) with the BioMediTech Institute, Tampere University, Finland, and an Assistant Professor (Lecturer) with the School of Computer Science and Electronic Engineering, University of Essex, U.K. He has authored or coauthored over 70 research papers in various international flagship journals and conferences in the areas of wireless communications, molecular and nanoscale communications, as well as bionanoscience. His research interests include Internet of Bio-Nano Things, molecular communications, bionanoscience, and 6G communications. He received the CONNECT Prof. Tom Brazil Excellence in Research Award in 2020.
Mladen Veletić received the B.Sc. and M.Sc. degrees in electronics and telecommunications from the Faculty of Electrical Engineering, University of Banja Luka (UNIBL), Bosnia and Herzegovina, in 2010 and 2012, respectively, and the Ph.D. degree in telecommunications from the Department of Electronic Systems, Norwegian University of Science and Technology, Norway, and the Faculty of Electrical Engineering, UNIBL. From 2011 to 2017, he worked as a Senior Teaching and Research Assistant with UNIBL. He is currently a Postdoctoral Research Scientist with the Intervention Center, Oslo University Hospital. His research interests include molecular and nano-neural communications, wireless communications, and positioning in cellular networks. He was awarded a Gold Plaque from the UNIBL for his achievements throughout the undergraduate education.
Masamitsu Kanada studied cell biology and fluorescence live cell imaging with Tsukuba University, Japan. After completing his Ph.D., he worked for a pharmaceutical company developing cancer therapies. During his postdoctoral training, he studied intravital microscopy and whole-body preclinical imaging in the laboratories of Susumu Terakawa at Hamamatsu University School of Medicine and Christopher Contag at Stanford University. He is an Assistant Professor with the Department of Pharmacology and Toxicology. His research focuses on addressing questions in cancer biology using multidisciplinary approaches.
Massimiliano Pierobon (Member, IEEE) received the Ph.D. degree in electrical and computer engineering from the Georgia Institute of Technology, Atlanta, GA, USA, in 2013. Since August 2013, he has been an Assistant Professor with the Department of Computer Science and Engineering, University of Nebraska–Lincoln (UNL), NE, USA, where he also holds a courtesy appointment with the Department of Biochemistry. His research interests are in molecular communication theory, nanonetworks, intrabody networks, communication engineering applied to synthetic biology, and the Internet of Bio-Nano Things. His selected honors include the 2011 Georgia Tech BWN Lab Researcher of the Year Award, the 2013 IEEE Communications Letters Exemplary Reviewer Award, the UNL CSE Upper and Graduate Level Teaching Award in 2016 and 2017, the 2017 IEEE INFOCOM Best Paper Runner-Up Award, and the Best In-Session Presentation Award. He is currently the PI of the NSF Project WetComm: Foundations of Wet Communication Theory, and a Co-PI of the NSF Project Redox-Enabled Bio-Electronics for Molecular Communication and Memory (RE-BIONICS), and has been the lead PIs of the NSF Project TelePathy: Telecommunication Systems Modeling and Engineering of Cell Communication Pathways. He has been the Co-Editor-in-Chief of Nano Communication Networks (Elsevier) since July 2017, and was an Associate Editor of IEEE Transactions on Communications from 2013 to 2020.
Seppo Vainio is a PI in Biocenter Oulu and Infotech Oulu research organization. He has published aver 130 original research arcticles and reviews. His Hirch index is 29 and his work has been sited more then 10 000 times. He has served as an invited presentations to peer-reviewed, internationally established conferences. Healso serves as referee in many international journals such as Nature, Cell, Development, PNAS, JASN. Nal. He has also organized international research congresses such as a symposia on Development, regeneration & Aging, Oulu 7–9. 2012 and served as session chair as in the interdisciplinary Argumenta Finnish Cultural Foundation funded symposia in Oulu 7–9. 2012, The Gordon Research Conference on mammary Gland Biology, June 1–6, (2008), Il Ciocco Hotel and Resort Lucca (Barga) and International Symposium for Gonad and Brain Sex Differentiation, Fukuoka, Japan (2008); The 1st Kumamoto University Global COE International Symposium on Cell Fate Regulation, January 16th, (2008), Fourth Swedish-Finnish Connective Tissue Meeting, Friday 13 April to Saturday April 14, (2007), Lund- Sweden World congress on nephrology, April, 21–25, (2007), Rio de Janeiro, Brazil; ELSO — the European Life Scientist Organization meeting in (2007), Dresden, 1–4 September; Fourth course on “Genetics a From Developmental Genes to Dysmorphology. He had had substantial research funding from academy of Finland, European union and he is PI in the Academy of Finland Centers of excellence and a Coordinator of EU project “Wnt and stem cells” (QLK3-CT-2001- 01275)(2001–2004) 7. International Prizes &Awards &Academy memberships: Centre of Excellence of Academy of Finland. He has hosted FiDIpr Susan Quaggin in his lab and also worked as visiting distinguished Prof. of Ulm Univ., 2013-, Prof. Pool grant 2012–2013, Biocenter Oulu. As awards he has selected as a discovery of the year 2011, Physiologist of the year in Finland (1999), Discovery of the year in Biocenter Oulu (1999, 2004, 2009, 2010 rank position). He has granted such as “Induction of kidney tubule formation, U.S. patents No 09/937,735, Wnt signalling in reproductive organs, U.S. patent No 00246/232001. Process for producing active Wnt proteins in prokaryotes. Finnish Patent application FI 20040842”.
Ilangko Balasingham (Senior Member, IEEE) received the M.Sc. and Ph.D. degrees in signal processing from the Department of Electronic Systems, Norwegian University of Science and Technology (NTNU), Trondheim, Norway, in 1993 and 1998, respectively. He performed his master’s degree thesis with the Department of Electrical and Computer Engineering, University of California at Santa Barbara, Santa Barbara, USA. From 1998 to 2002, he worked as a Research Engineer developing image and video streaming solutions for mobile handheld devices with Fast Search & Transfer ASA, Oslo, Norway, which is now part of Microsoft Inc. Since 2002, he has been with the Intervention Center, Oslo University Hospital, Oslo, where he heads the Wireless Sensor Network Research Group. He was appointed a Professor of Medical Signal Processing and Communications with NTNU in 2006. For the academic year 2016/2017, he was a Professor by courtesy with the Frontier Institute, Nagoya Institute of Technology, Japan. His research interests include super robust short-range communications for both in-body and on-body sensors, body area sensor network, microwave short-range sensing of vital signs, short-range localization and tracking mobile sensors, and nanoscale communication networks. He has authored or coauthored over 280 journal and conference papers, seven book chapters, 42 abstracts, six patents, and 20 articles in popular press. He has given 21 invited/keynotes at the international conferences. In addition, he is active in organizing conferences (the Steering Committee Member of ACM NANOCOM since 2018; the General Chair of the 2019 IEEE International Symposium of Medical ICT and the 2012 Body Area Networks (BODYNETS) conference; and the TPC Chair of the 2015 ACM NANOCOM) and editorial board (an Area Editor of Nano Communication Networks (Elsevier) since 2013 and the Specialty Chief Editor of Frontiers in Communications and Networks since 2020).
Sasitharan Balasubramaniam (Senior Member, IEEE) received the B.E. degree in electrical and electronic engineering from The University of Queensland, Brisbane, QLD, Australia in 1998, the M.E. degree in computer and communication engineering from the Queensland University of Technology, Brisbane, in 1999, and the Ph.D. degree from The University of Queensland, in 2005. He was previously the Director of the Research of the Walton Institute, Waterford Institute of Technology, Waterford, Ireland. He is currently an Associate Professor at the Department of Computer Science and Engineering at the University of Nebraska, Lincoln, USA. He has authored or coauthored more than 100 articles in various journals and conferences and actively participates in a number of technical program committees for various conferences. His current research interests include molecular communications, the Internet of (Bio) NanoThings, and terahertz wireless communications. Dr. Balasubramaniam is on the Steering Board Committee of ACM NanoCom, which he co-founded. In 2018, he was the IEEE Nanotechnology Council Distinguished Lecturer. He previously served as an Associate Editor for the IEEE INTERNET OF THINGS JOURNAL. He currently serves as an Associate Editor for the IEEE TRANSACTIONS ON MOLECULAR, BIOLOGICAL AND MULTI-SCALE COMMUNICATIONS, IEEE TRANSACTIONS ON MOBILE COMPUTING, and Nano Communication Networks (Elsevier).
Funding Statement
This work was supported by the European Union’s Horizon 2020 Research and Innovation Programme through the Marie Skłodowska-Curie Grant under Agreement 839553. The work of Massimiliano Pierobon was supported by the U.S. National Science Foundation under Grant CCF-1816969.
Footnotes
The apical membrane faces the external (luminal) compartment and contains proteins that determine secretion and absorption, whereas the basolateral domain faces the internal (systemic) compartment (tissues and blood).
References
- [1].Jones L., Brown D., and Palumbo D., Coronavirus: A Visual Guide to the Economic Impact, BBC News, London, U.K., 2020. [Google Scholar]
- [2].Stevis-Gridneff M., A 750 Billion Virus Recovery Plan Thrusts Europe Into a New Frontier, New York Times, New York, NY, USA, 2020. [Google Scholar]
- [3].Saxena V., “Mesenchymal stem cell based therapeutic intervention against viral hepatitis: A perspective,” J. Stem. Cells Genet., vol. 1, no. 1, pp. 1–2, 2017. [Google Scholar]
- [4].Thompson T., Annals of Influenza or Epidemic Catarrhal Fever in Great Britain From 1510 to 1837, vol. 21. London, U.K.: Sydenham Soc., 1852. [Google Scholar]
- [5].Roldão A., Mellado M. C. M., Castilho L. R., Carrondo M. J., and Alves P. M., “Virus-like particles in vaccine development,” Exp. Rev. vaccines, vol. 9, no. 10, pp. 1149–1176, 2010. [DOI] [PubMed] [Google Scholar]
- [6].Akyildiz I. F., Pierobon M., and Balasubramaniam S., “An information theoretic framework to analyze molecular communication systems based on statistical mechanics,” Proc. IEEE, vol. 107, no. 7, pp. 1230–1255, Jul. 2019. [Google Scholar]
- [7].Nakano T., “Molecular communication: A 10 year retrospective,” IEEE Trans. Mol. Biol. Multi Scale Commun., vol. 3, no. 2, pp. 71–78, Jun. 2017. [Google Scholar]
- [8].Akyildiz I. F., Pierobon M., Balasubramaniam S., and Koucheryavy Y., “The Internet of Bio-Nano Things,” IEEE Commun. Mag., vol. 53, no. 3, pp. 32–40, Mar. 2015. [Google Scholar]
- [9].Akyildiz I. F., Brunetti F., and Blázquez C., “NanoNetworks: A new communication paradigm,” Comput. Netw., vol. 52, no. 12, pp. 2260–2279, 2008. [Google Scholar]
- [10].Felicetti L., Femminella M., Reali G., and Liò P., “Applications of molecular communications to medicine: A survey,” Nano Commun. Netw., vol. 7, pp. 27–45, Mar. 2016. [Google Scholar]
- [11].Atakan B., Akan O. B., and Balasubramaniam S., “Body area nanonetworks with molecular communications in nanomedicine,” IEEE Commun. Mag., vol. 50, no. 1, pp. 28–34, Jan. 2012. [Google Scholar]
- [12].Barros M. T., Silva W., and Regis C. D. M., “The multi-scale impact of the Alzheimer’s disease on the topology diversity of astrocytes molecular communications nanonetworks,” IEEE Access, vol. 6, pp. 78904–78917, 2018. [Google Scholar]
- [13].Veletić M. and Balasingham I., “Synaptic communication engineering for future cognitive brain–machine interfaces,” Proc. IEEE, vol. 107, no. 7, pp. 1425–1441, Jul. 2019. [Google Scholar]
- [14].Balasubramaniam S. and Kangasharju J., “Realizing the Internet of Nano Things: Challenges, solutions, and applications,” Computer, vol. 46, no. 2, pp. 62–68, 2012. [Google Scholar]
- [15].Martins D. P., Leetanasaksakul K., Barros M. T., Thamchaipenet A., Donnelly W., and Balasubramaniam S., “Molecular communications pulse-based jamming model for bacterial biofilm suppression,” IEEE Trans. Nanobiosci., vol. 17, no. 4, pp. 533–542, Oct. 2018. [DOI] [PubMed] [Google Scholar]
- [16].Martins D. P., Barros M. T., and Balasubramaniam S., “Using competing bacterial communication to disassemble biofilms,” in Proc. 3rd ACM Int. Conf. Nanoscale Comput. Commun., 2016, pp. 1–6. [Google Scholar]
- [17].Walsh F. and Balasubramaniam S., “Reliability and delay analysis of multihop virus-based nanonetworks,” IEEE Trans. Nanotechnol., vol. 12, no. 5, pp. 674–684, Sep. 2013. [Google Scholar]
- [18].Khalid M., Amin O., Ahmed S., Shihada B., and Alouini M.-S., “Communication through breath: Aerosol transmission,” IEEE Commun. Mag., vol. 57, no. 2, pp. 33–39, Feb. 2019. [Google Scholar]
- [19].Khalid M., Amin O., Ahmed S., Shihada B., and Alouini M.-S., “Modeling of viral aerosol transmission and detection,” IEEE Trans. Commun., vol. 68, no. 8, pp. 4859–4873, Aug. 2020. [Google Scholar]
- [20].Vimalajeewa D., Balasubramaniam S., Berry D. P., and Barry G.. (2020). In Silico Modeling of Virus Particle Propagation and Infectivity Along the Respiratory Tract: A Case Study for SARS-CoV-2. [Online]. Available: https://www.biorxiv.org/content/10.1101/2020.08.20.259242v1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Martins D. P., Barros M. T., Pierobon M., Lio M. K. P., and Balasubramaniam S., “Computational models for trapping Ebola virus using engineered bacteria,” IEEE/ACM Trans. Comput. Biol. Bioinformat., vol. 15, no. 6, pp. 2017–2027, Nov./Dec. 2018. [DOI] [PubMed] [Google Scholar]
- [22].Sariol A. and Perlman S., “Lessons for COVID-19 immunity from other coronavirus infections,” Immunity, vol. 53, no. 2, pp. 248–263, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Florindo H. F.et al. , “Immune-mediated approaches against COVID-19,” Nat. Nanotechnol., vol. 15, pp. 630–645, Jul. 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Lebeau G.et al. , “Deciphering SARS-CoV-2 virologic and immunologic features,” Int. J. Mol. Sci., vol. 21, no. 16, p. 5932, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Wadman M., Couzin-Frankel J., Kaiser J., and Matacic C., “A rampage through the body,” Science, vol. 368, no. 6489, pp. 356–360, 2020. [DOI] [PubMed] [Google Scholar]
- [26].Ansari A. A., “Clinical features and pathobiology of Ebolavirus infection,” J. Autoimmunity, vol. 55, pp. 1–9, Dec. 2014. [DOI] [PubMed] [Google Scholar]
- [27].Food U.et al. , “First FDA-approved vaccine for the prevention of Ebola virus disease, marking a critical milestone in public health preparedness and response,” Press Announc, 2019. Accessed: Apr. 16, 2021. [Online] Available: https://www.fda.gov/news-events/press-announcements/first-fda-approved-vaccine-prevention-ebola-virus-disease-marking-critical-milestone-public-health
- [28].Henao-Restrepo A. M.et al. , “Efficacy and effectiveness of an rVSV-vectored vaccine in preventing Ebola virus disease: Final results from the guinea ring vaccination, open-label, cluster-randomised trial (Ebola Ça Suffit!),” Lancet, vol. 389, no. 10068, pp. 505–518, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Yu C. I. and Chiang B.-L., “A new insight into hepatitis C vaccine development,” J. Biomed. Biotechnol., vol. 2010, Jun. 2010, Art. no. 548280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [30].Peiris J.et al. , “Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: A prospective study,” Lancet, vol. 361, no. 9371, pp. 1767–1772, 2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Hung I. F. N.et al. , “Viral loads in clinical specimens and SARS manifestations,” Emerg. Infectious Diseases, vol. 10, no. 9, pp. 1550–1557, 2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Pan Y., Zhang D., Yang P., Poon L. L. M., and Wang Q., “Viral load of SARS-CoV-2 in clinical samples,” Lancet Infectious Diseases, vol. 20, no. 4, pp. 411–412, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].To K. K.-W.et al. , “Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study,” Lancet Infectious Diseases, vol. 20, no. 5, pp. 565–574, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Zou L.et al. , “SARS-CoV-2 viral load in upper respiratory specimens of infected patients,” New England J. Med., vol. 382, no. 12, pp. 1177–1179, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Pickett B. E.et al. , “ViPR: An open bioinformatics database and analysis resource for virology research,” Nucl. Acids Res., vol. 40, no. D1, pp. D593–D598, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Oh M.-D.et al. , “Viral load kinetics of MERS coronavirus infection,” New England J. Med., vol. 375, no. 13, pp. 1303–1305, 2016. [DOI] [PubMed] [Google Scholar]
- [37].Fehr A. R. and Perlman S., “Coronaviruses: An overview of their replication and pathogenesis,” in Coronaviruses. Heidelberg, Germany: Springer, 2015, pp. 1–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Al Hajjar S., Memish Z. A., and McIntosh K., “Middle east respiratory syndrome coronavirus (MERS-CoV): A perpetual challenge,” Ann. Saudi Med., vol. 33, no. 5, pp. 427–436, 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Towner J. S.et al. , “Rapid diagnosis of Ebola hemorrhagic fever by reverse transcription-PCR in an outbreak setting and assessment of patient viral load as a predictor of outcome,” J. Virol., vol. 78, no. 8, pp. 4330–4341, 2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].de La Vega M.-A.et al. , “Ebola viral load at diagnosis associates with patient outcome and outbreak evolution,” J. Clin. Invest., vol. 125, no. 12, pp. 4421–4428, Dec. 2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Ben-Shachar R., Schmidler S., and Koelle K., “Drivers of inter-individual variation in Dengue viral load dynamics,” PLoS Comput. Biol., vol. 12, no. 11, pp. 1–26, Nov. 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Wang W.-K.et al. , “High levels of plasma Dengue viral load during defervescence in patients with Dengue Hemorrhagic Fever: Implications for pathogenesis,” Virology, vol. 305, no. 2, pp. 330–338, 2003. [DOI] [PubMed] [Google Scholar]
- [43].Wang W. K.et al. , “Slower rates of clearance of viral load and virus-containing immune complexes in patients with Dengue hemorrhagic fever,” Clin. Infectious Diseases, vol. 43, no. 8, pp. 1023–1030, 2006. [DOI] [PubMed] [Google Scholar]
- [44].Mansuy J. M.et al. , “Zika virus: High infectious viral load in semen, a new sexually transmitted pathogen?” Lancet Infectious Diseases, vol. 16, no. 4, p. 405, 2016. [DOI] [PubMed] [Google Scholar]
- [45].de Laval F., Matheus S., Labrousse T., Enfissi A., Rousset D., and Briolant S., “Kinetics of Zika viral load in semen,” New England J. Med., vol. 377, no. 7, pp. 697–699, 2017. [DOI] [PubMed] [Google Scholar]
- [46].Fourcade C.et al. , “Viral load kinetics of Zika virus in plasma, urine and saliva in a couple returning from Martinique, French West Indies,” J. Clin. Virol., vol. 82, pp. 1–4, Sep. 2016. [DOI] [PubMed] [Google Scholar]
- [47].Lerat H.et al. , “In vivo tropism of hepatitis C virus genomic sequences in hematopoietic cells: Influence of viral load, viral genotype, and cell phenotype,” Blood, vol. 91, no. 10, pp. 3841–3849, 1998. [PubMed] [Google Scholar]
- [48].Louten J., “Virus transmission and epidemiology,” Essential Human Virol., vol. 2016, pp. 71–92, May 2016. [Google Scholar]
- [49].Bomsel M. and Alfsen A., “Entry of viruses through the epithelial barrier: Pathogenic trickery,” Nat. Rev. Mol. Cell Biol., vol. 4, no. 1, pp. 57–68, 2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Cong Y. and Ren X., “Coronavirus entry and release in polarized epithelial cells: A review,” Rev. Med. Virol., vol. 24, no. 5, pp. 308–315, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [51].Veletić M., Barros M. T., Balasingham I., and Balasubramaniam S., “A molecular communication model of exosome-mediated brain drug delivery,” in Proc. 6th Annu. ACM Int. Conf. Nanoscale Comput. Commun. (NANOCOM), 2019, p. 22. [Google Scholar]
- [52].Zitzmann C. and Kaderali L., “Mathematical analysis of viral replication dynamics and antiviral treatment strategies: From basic models to age-based multi-scale modeling,” Front. Microbiol., vol. 9, p. 1546, Jul. 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [53].Bonhoeffer S., May R. M., Shaw G. M., and Nowak M. A., “Virus dynamics and drug therapy,” Proc. Nat. Acad. Sci. USA, vol. 94, no. 13, pp. 6971–6976, 1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [54].Chahibi Y., Pierobon M., Song S. O., and Akyildiz I. F., “A molecular communication system model for particulate drug delivery systems,” IEEE Trans. Biomed. Eng., vol. 60, no. 12, pp. 3468–3483, Dec. 2013. [DOI] [PubMed] [Google Scholar]
- [55].Arjmandi Y., Zoofaghar M., Rouzegar S. V., Veletić M., and Balasingham I., “On mathematical analysis of active drug transport coupled with flow-induced diffusion in blood vessels,” IEEE Trans. Nanobiosci., vol. 20, no. 1, pp. 105–115, Jan. 2021. [DOI] [PubMed] [Google Scholar]
- [56].Huang J., Zheng M., Tang X., Chen Y., Tong A., and Zhou L., “Potential of SARS-CoV-2 to cause CNS infection: Biologic fundamental and clinical experience,” Front. Neurol., vol. 11, p. 659, Jun. 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Kalluri R. and LeBleu V. S., “The biology, function, and biomedical applications of exosomes,” Science, vol. 367, no. 6478, 2020, Art. no. eaau6977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [58].Urbanelli L.et al. , “The role of extracellular vesicles in viral infection and transmission,” Vaccines, vol. 7, no. 3, p. 102, 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Nolte-’t Hoen E., Cremer T., Gallo R. C., and Margolis L. B., “Extracellular vesicles and viruses: Are they close relatives?” Proc. Nat. Acad. Sci. USA, vol. 113, no. 33, pp. 9155–9161, 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [60].Veletić M., Barros M. T., Arjmandi H., Balasubramaniam S., and Balasingham I., “Modeling of modulated exosome release from differentiated induced neural stem cells for targeted drug delivery,” IEEE Trans. Nanobiosci., vol. 19, no. 3, pp. 357–367, Jul. 2020. [DOI] [PubMed] [Google Scholar]
- [61].Tricarico C., Clancy J., and D’Souza-Schorey C., “Biology and biogenesis of shed microvesicles,” Small GTPases, vol. 8, no. 4, pp. 220–232, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [62].Handel A., Longini I. M., and Antia R., “Towards a quantitative understanding of the within-host dynamics of influenza A infections,” J. Roy. Soc. Interface, vol. 7, no. 42, pp. 35–47, 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [63].Gulec F. and Atakan B., “A molecular communication perspective on airborne pathogen transmission and reception via droplets generated by coughing and sneezing,” 2020. [Online]. Available: http://arXiv:2007.07598 [Google Scholar]
- [64].Bourouiba L., Dehandschoewercker E., and Bush J. W., “Violent expiratory events: On coughing and sneezing,” J. Fluid Mech., vol. 745, pp. 537–563, Mar. 2014. [Google Scholar]
- [65].Balásházy I., Hofmann W., and Martonen T. B., “Inspiratory particle deposition in airway bifurcation models,” J. Aerosol Sci., vol. 22, no. 1, pp. 15–30, 1991. [Google Scholar]
- [66].Xu H.et al. , “High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa,” Int. J. Oral Sci., vol. 12, no. 1, p. 8, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [67].Chaudhuri S., Basu S., Kabi P., Unni V. R., and Saha A., “Modeling the role of respiratory droplets in COVID-19 type pandemics,” Phys. Fluids, vol. 32, no. 6, 2020, Art. no. 063309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [68].Chen C. and Zhao B., “Some questions on dispersion of human exhaled droplets in ventilation room: Answers from numerical investigation,” Indoor Air, vol. 20, no. 2, pp. 95–111, 2010. [DOI] [PubMed] [Google Scholar]
- [69].Han Z., Weng W., and Huang Q., “Characterizations of particle size distribution of the droplets exhaled by sneeze,” J. Roy. Soc. Interface, vol. 10, no. 88, 2013, Art. no. 20130560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [70].Mui K., Wong L., Wu C., and Lai A. C., “Numerical modeling of exhaled droplet nuclei dispersion and mixing in indoor environments,” J. Hazardous Mater., vol. 167, nos. 1–3, pp. 736–744, 2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [71].Jamali V., Ahmadzadeh A., Wicke W., Noel A., and Schober R., “Channel modeling for diffusive molecular communication—A tutorial review,” Proc. IEEE, vol. 107, no. 7, pp. 1256–1301, Jul. 2019. [Google Scholar]
- [72].Farsad N., Guo W., and Eckford A. W., “Tabletop molecular communication: Text messages through chemical signals,” PLoS ONE, vol. 8, no. 12, 2013, Art. no. e82935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [73].Olufsen M. S., Peskin C. S., Kim W. Y., Pedersen E. M., Nadim A., and Larsen J., “Numerical simulation and experimental validation of blood flow in arteries with structured-tree outflow conditions,” Ann. Biomed. Eng., vol. 28, no. 11, pp. 1281–1299, 2000. [DOI] [PubMed] [Google Scholar]
- [74].Vabret N.et al. , “Immunology of COVID-19: Current state of the science,” Immunity, vol. 52, no. 6, pp. 910–941, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [75].Tay M. Z., Poh C. M., Rsnia L., MacAry P. A., and Ng L. F. P., “The trinity of COVID-19: Immunity, inflammation and intervention,” Nat. Rev. Immunol., vol. 20, no. 6, pp. 363–374, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [76].Faure E.et al. , “Distinct immune response in two MERS-CoV-infected patients: Can we go from bench to bedside?” PLoS ONE, vol. 9, no. 2, 2014, Art. no. e88716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [77].Baize S.et al. , “Inflammatory responses in Ebola virus-infected patients,” Clin. Exp. Immunol., vol. 128, no. 1, pp. 163–168, 2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [78].Levine M. M., “Monoclonal antibody therapy for Ebola virus disease,” New England J. Med., vol. 381, no. 24, pp. 2365–2366, 2019. [DOI] [PubMed] [Google Scholar]
- [79].Fares-Gusmao R., Rocha B. C., Sippert E., Lanteri M. C., Áñez G., and Rios M., “Differential pattern of soluble immune markers in asymptomatic Dengue, West Nile and Zika virus infections,” Sci. Rep., vol. 9, no. 1, 2019, Art. no. 17172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [80].Guzman M. G.et al. , “Dengue: A continuing global threat,” Nat. Rev. Microbiol., vol. 8, no. 12, pp. S7–S16, 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [81].Collins M. H.et al. , “Human antibody response to Zika targets type-specific quaternary structure epitopes,” JCI Insight, vol. 4, no. 8, 2019, Art. no. e124588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [82].Klenerman P. and Thimme R., “T cell responses in hepatitis C: The good, the bad and the unconventional,” Gut, vol. 61, no. 8, pp. 1226–1234, 2012. [DOI] [PubMed] [Google Scholar]
- [83].Hetta H., Mehta M., and Shata M., “Gut immune response in the presence of hepatitis C virus infection,” World J. Immunol., vol. 4, no. 2, pp. 52–62, 2014. [Google Scholar]
- [84].Swann R. E.et al. , “Broad anti-hepatitis C virus (HCV) antibody responses are associated with improved clinical disease parameters in chronic HCV infection,” J. Virol., vol. 90, no. 9, pp. 4530–4543, 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [85].Vargas E. A. H. and Velasco-Hernandez J. X., “In-host modelling of COVID-19 kinetics in humans,” Annu. Rev. Control, vol. 50, pp. 448–456, Sep. 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [86].Wolfel R.et al. , “Virological assessment of hospitalized patients with COVID-2019,” Nature, vol. 581, no. 7809, pp. 465–469, 2020. [DOI] [PubMed] [Google Scholar]
- [87].Zhang Y.et al. , “Analysis of serum cytokines in patients with severe acute respiratory syndrome,” Infection Immunity, vol. 72, no. 8, pp. 4410–4415, 2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [88].Cheung C. Y.et al. , “Cytokine responses in severe acute respiratory syndrome coronavirus-infected macrophages in vitro: Possible relevance to pathogenesis,” J. Virol., vol. 79, no. 12, pp. 7819–7826, 2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [89].Jiang Y.et al. , “Characterization of cytokine/chemokine profiles of severe acute respiratory syndrome,” Amer. J. Respir. Crit. Care Med., vol. 171, no. 8, pp. 850–857, 2005. [DOI] [PubMed] [Google Scholar]
- [90].European Comission. (2021). COVID-19 Data Portal. [Online]. Available: https://www.covid19dataportal.org
- [91].Semantic Scholar. (2021). CORD-19 Open Research Dataset. [Online]. Available: https://www.semanticscholar.org/cord19
- [92].G. fur Aerosolforschung. Position Paper of the Gesellschaft fur Aerosolforschung on Understanding the Role of Aerosol Particles in SARS-CoV-2 Infection. Accessed: Dec. 2020. [Online]. Available: https://doi.org/10.5281/zenodo.4350494
- [93].Marbe D., Kriegel M., Lange J., Schumann L., Hartmann A., and Fleischer M.. Data to: Aerosol Emission of Child Voices During Speaking, Singing and Shouting. Accessed: Sep. 2020. [Online]. Available: https://doi.org/10.5281/zenodo.4035990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- [94].Barros M. T., Doan P., Kandhavelu M., Jennings B., and Balasubramaniam S., “Engineering calcium signaling of astrocytes for neural-molecular computing logic gates,” 2020. [Online]. Available: http://arXiv:2007.06646 [DOI] [PMC free article] [PubMed] [Google Scholar]
- [95].Nakano T.et al. , “Roles of remote and contact forces in epithelial cell structure formation,” Biophys. J., vol. 118, no. 6, pp. 1466–1478, Mar. 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [96].Grebenstein L.et al. , “A molecular communication testbed based on proton pumping bacteria: Methods and data,” IEEE Trans. Mol. Biol. Multi-Scale Commun., vol. 5, no. 1, pp. 56–62, Oct. 2019. [Google Scholar]
- [97].Liu Y.et al. , “Using a redox modality to connect synthetic biology to electronics: Hydrogel-based chemo-electro signal transduction for molecular communication,” Adv. Healthcare Mater., vol. 6, no. 1, 2017, Art. no. 1600908. [DOI] [PubMed] [Google Scholar]
- [98].Balasubramaniam S.et al. , “Development of artificial neuronal networks for molecular communication,” Nano Commun. Netw., vol. 2, nos. 2–3, pp. 150–160, 2011. [Google Scholar]
- [99].McGuiness D. T., Giannoukos S., Marshall A., and Taylor S., “Experimental results on the open-air transmission of macro-molecular communication using membrane inlet mass spectrometry,” IEEE Commun. Lett., vol. 22, no. 12, pp. 2567–2570, Dec. 2018. [Google Scholar]
- [100].Felicetti L., Femminella M., Reali G., Gresele P., Malvestiti M., and Daigle J. N., “Modeling CD40-based molecular communications in blood vessels,” IEEE Trans. Nanobiosci., vol. 13, no. 3, pp. 230–243, Sep. 2014. [DOI] [PubMed] [Google Scholar]
- [101].Vishveshwara S., Ghosh A., and Hansia P., “Intra and inter-molecular communications through protein structure network,” Current Protein Peptide Sci., vol. 10, no. 2, pp. 146–160, 2009. [DOI] [PubMed] [Google Scholar]
- [102].Lee O. K., Kuo T. K., Chen W.-M., Lee K.-D., Hsieh S.-L., and Chen T.-H., “Isolation of multipotent mesenchymal stem cells from umbilical cord blood,” Blood, vol. 103, no. 5, pp. 1669–1675, 2004. [DOI] [PubMed] [Google Scholar]
- [103].Sottile V., Halleux C., Bassilana F., Keller H., and Seuwen K., “Stem cell characteristics of human trabecular bone-derived cells,” Bone, vol. 30, no. 5, pp. 699–704, 2002. [DOI] [PubMed] [Google Scholar]
- [104].De Bari C., Dell’Accio F., Tylzanowski P., and Luyten F. P., “Multipotent mesenchymal stem cells from adult human synovial membrane,” Arthritis Rheumatism, vol. 44, no. 8, pp. 1928–1942, 2001. [DOI] [PubMed] [Google Scholar]
- [105].Sato Y.et al. , “Human mesenchymal stem cells xenografted directly to rat liver are differentiated into human hepatocytes without fusion,” Blood, vol. 106, no. 2, pp. 756–763, 2005. [DOI] [PubMed] [Google Scholar]
- [106].Gupta S., Krishnakumar V., Sharma Y., Dinda A. K., and Mohanty S., “Mesenchymal stem cell derived exosomes: A nano platform for therapeutics and drug delivery in combating COVID-19,” Stem Cell Rev. Rep., vol. 17, no. 1, pp. 33–43, 2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [107].Islam M. T.et al. , “A perspective on emerging therapeutic interventions for COVID-19,” Front. Public Health, vol. 8, p. 281, Jul. 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [108].Cruz T. and Rojas M., “Preclinical evidence for the role of stem/stromal cells in targeting ards,” in Stem Cell-Based Therapy for Lung Disease. Cham, Switzerland: Springer, 2019, pp. 199–217. [Google Scholar]
- [109].Zanoni M., Cortesi M., Zamagni A., and Tesei A., “The role of mesenchymal stem cells in radiation-induced lung fibrosis,” Int. J. Mol. Sci., vol. 20, no. 16, p. 3876, 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [110].Perestrelo A. R., Águas A. C., Rainer A., and Forte G., “Microfluidic organ/body-on-a-chip devices at the convergence of biology and microengineering,” Sensors, vol. 15, no. 12, pp. 31142–31170, 2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [111].Šiljak H.et al. , “Evolving intelligent reflector surface towards 6G for public health: Application in airborne virus detection,” 2020. [Online]. Available: http://arXiv:2009.02224 [Google Scholar]
- [112].Tang H.et al. , “Human organs-on-chips for virology,” Trends Microbiol., vol. 28, no. 11, pp. 934–946, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [113].Ringehan M., McKeating J. A., and Protzer U., “Viral hepatitis and liver cancer,” Philos. Trans. Roy. Soc. B Biol. Sci., vol. 372, no. 1732, 2017, Art. no. 20160274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [114].Villenave R.et al. , “Human gut-on-a-chip supports polarized infection of coxsackie B1 virus in vitro,” PLoS ONE, vol. 12, no. 2, 2017, Art. no. e0169412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [115].Cifuente J. O.et al. , “Molecular determinants of disease in coxsackievirus B1 murine infection,” J. Med. Virol., vol. 83, no. 9, pp. 1571–1581, 2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [116].Johnson B. N.et al. , “3D printed nervous system on a chip,” Lab Chip, vol. 16, no. 8, pp. 1393–1400, 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [117].Benam K. H.et al. , “Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro,” Nat. Methods, vol. 13, no. 2, pp. 151–157, 2016. [DOI] [PubMed] [Google Scholar]
- [118].Si L.et al. (2019). Discovery of Influenza Drug Resistance Mutations and Host Therapeutic Targets Using a Human Airway Chip. [Online]. Available: https://www.biorxiv.org/content/10.1101/685552v1.full [Google Scholar]
- [119].Deng Y., Pierobon M., and Nallanathan A., “A microfluidic feed forward loop pulse generator for molecular communication,” in Proc. GLOBECOM IEEE Global Commun. Conf., 2017, pp. 1–7. [Google Scholar]
- [120].Hamidović M., Haselmayr W., Grimmer A., Wille R., and Springer A., “Passive droplet control in microfluidic networks: A survey and new perspectives on their practical realization,” Nano Commun. Netw., vol. 19, pp. 33–46, Mar. 2019. [Google Scholar]
- [121].Fronczek C. F. and Yoon J.-Y., “Biosensors for monitoring airborne pathogens,” J. Lab. Autom., vol. 20, no. 4, pp. 390–410, 2015. [DOI] [PubMed] [Google Scholar]
- [122].Grebenstein L.et al. , “Biological optical-to-chemical signal conversion interface: A small-scale modulator for molecular communications,” IEEE Trans. Nanobiosci., vol. 18, no. 1, pp. 31–42, Jan. 2019. [DOI] [PubMed] [Google Scholar]
- [123].McGuiness D. T., Giannoukos S., Marshall A., and Taylor S., “Parameter analysis in macro-scale molecular communications using advection-diffusion,” IEEE Access, vol. 6, pp. 46706–46717, 2018. [Google Scholar]
- [124].Roumpeka D. D., Wallace R. J., Escalettes F., Fotheringham I., and Watson M., “A review of bioinformatics tools for bio-prospecting from metagenomic sequence data,” Front. Genet., vol. 8, p. 23, Mar. 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [125].Soria-Guerra R. E., Nieto-Gomez R., Govea-Alonso D. O., and Rosales-Mendoza S., “An overview of bioinformatics tools for epitope prediction: Implications on vaccine development,” J. Biomed. Informat., vol. 53, pp. 405–414, Feb. 2015. [DOI] [PubMed] [Google Scholar]
- [126].Garrido C.et al. , “Evaluation of eight different bioinformatics tools to predict viral tropism in different human immunodeficiency virus type 1 subtypes,” J. Clin. Microbiol., vol. 46, no. 3, pp. 887–891, 2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [127].Wang Q.et al. , “Xander: Employing a novel method for efficient gene-targeted metagenomic assembly,” Microbiome, vol. 3, no. 1, p. 32, 2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [128].Balasubramaniam S. and Liò P., “Multi-hop conjugation based bacteria nanonetworks,” IEEE Trans. Nanobiosci., vol. 12, no. 1, pp. 47–59, Mar. 2013. [DOI] [PubMed] [Google Scholar]
- [129].Bilgin B. A., Dinc E., and Akan O. B., “DNA-based molecular communications,” IEEE Access, vol. 6, pp. 73119–73129, 2018. [Google Scholar]
- [130].Veletić M. and Balasingham I., “An information theory of neuro-transmission in multiple-access synaptic channels,” IEEE Trans. Commun., vol. 68, no. 2, pp. 841–853, Feb. 2020. [Google Scholar]
- [131].Barros M. T., Balasubramaniam S., and Jennings B., “Comparative end-to-end analysis of Ca2+-signaling-based molecular communication in biological tissues,” IEEE Trans. Commun., vol. 63, no. 12, pp. 5128–5142, Dec. 2015. [Google Scholar]
- [132].Barros M. T., “CA2+-signaling-based molecular communication systems: Design and future research directions,” Nano Commun. Netw., vol. 11, pp. 103–113, Mar. 2017. [Google Scholar]
- [133].Sompayrac L. M., How the Immune System Works. Hoboken, NJ, USA: Wiley, 2019. [Google Scholar]