Abstract
Motivation
The analysis of longitudinal datasets and construction of gene regulatory networks (GRNs) provide a valuable means to disentangle the complexity of microRNA (miRNA)–mRNA interactions. However, there are no computational tools that can integrate, conduct functional analysis and generate detailed networks from longitudinal miRNA–mRNA datasets.
Results
We present TimiRGeN, an R package that uses time point-based differential expression results to identify miRNA–mRNA interactions influencing signaling pathways of interest. miRNA–mRNA interactions can be visualized in R or exported to PathVisio or Cytoscape. The output can be used for hypothesis generation and directing in vitro or further in silico work such as GRN construction.
Availability and implementation
TimiRGeN is available for download on Bioconductor (https://bioconductor.org/packages/TimiRGeN) and requires R v4.0.2 or newer and BiocManager v3.12 or newer.
Supplementary information
Supplementary data are available at Bioinformatics online.
1 Introduction
MicroRNAs (miRNAs) are single-stranded functional RNAs, around 16–22 nucleotides long which target specific mRNAs for degradation or translational repression; thus affecting protein levels (Selbach et al., 2008). Targeting is achieved by complementary binding between the 3ʹUTR of the target mRNA and a 7–8 nucleotide sequence found on the 5ʹend of the miRNA, known as the seed sequence (Bartel, 2004). There is increased clinical interest in miRNAs for several reasons: (i) miRNAs can be tested in animal models to understand human diseases and conditions. An example is miR-140-5p which is upregulated during chondrogenesis and downregulated during osteoarthritis (Barter et al., 2015; Miyaki et al., 2010). (ii) miRNAs can be secreted via exosomes into surrounding blood, extracellular matrix and urine (Chaturvedi et al., 2015; Chen et al., 2017; Leidinger et al., 2013). Their presence in body fluids provides valuable noninvasive biomarkers to assess the state of difficult to access tissues such as tumors, brain and bone. (iii) Lastly, miRNAs have potential as therapeutic agents as they modulate expression of specific mRNAs (Schwarzenbach et al., 2014).
However, in the laboratory, miRNAs are difficult to study, primarily because a single miRNA can regulate many mRNAs and a single mRNA can be regulated by multiple miRNAs. miRNA–mRNA interactome studies report over 18 000 interactions in HEK293 cells and over 34 000 interactions in human hepatoma cells (Helwak et al., 2013; Moore et al., 2015). A complementary strategy is to use a computational approach. The analysis of longitudinal miRNA–mRNA expression data, construction of gene regulatory networks (GRNs) and subsequent dynamic modeling, is a particularly useful means to gain a better understanding of miRNA–mRNA interactions (Ooi et al., 2018; Proctor et al., 2017; Qin et al., 2015). GRNs are useful tools for integrating multiomic data on mechanistic schematics. Yet, currently there is no computational tool that can handle longitudinal miRNA–mRNA datasets and reduce the volume of data to an extent where GRN construction is possible. This is presented in Table 1.
Table 1.
miRNA–mRNA integration tools
| Tool name | Availability | Time | Funct analysis | Reduction | Updated |
|---|---|---|---|---|---|
| anamiR | Bioc | X | ✓:Kegg, React,+ | ✓ | 2018 |
| DREM2 | Install | ✓ | ✓:GO | X | 2020 |
| MAGIA2 | Online | X | ✓:DAVID | ✓ | 2012 |
| miARMa-seq | Install | ✓ | ✓:GO, Kegg | X | 2019 |
| miRComb | SF | ✓ | ✓:GO, Kegg | ✓ | 2020 |
| miRIntegrator | Bioc | X | ✓:Kegg, React | ✓ | 2016 |
| miRNet | Online | X | ✓:GO, Kegg | X | 2021 |
| miRTarVis+ | Online | X | X | ✓ | 2020 |
| Sigterms | SF | X | ✓:GO | ✓ | 2009 |
| SpidermiR | Bioc | X | X | ✓ | 2020 |
| ToppMiR | Online | X | ✓:GO | ✓ | 2021 |
Note: Comparison of miRNA–mRNA integration tools: several tools are available as R packages that can be downloaded from Bioc (Bioconductor) or SF (SourceForge). Other tools can be installed locally or are available online. Some tools are capable of handling time series datasets. Several can perform funct (functional) analysis, usually utilizing GO, Kegg, React (Reactome), DAVID or others (+) and a few tools can reduce the volume of data. Also shown is when each tool was last updated.
Many existing tools (Table 1) have particular strengths, but none satisfy the criteria necessary to bridge longitudinal multiomic data and GRN creation. anamiR, miRIntegrator, MAGIA2, Sigterms and SpidermiR have substantial miRNA–mRNA integration capabilities but cannot handle longitudinal datasets (Bisognin et al., 2012; Cava et al., 2017; Creighton et al., 2008; Diaz et al., 2017; Wang et al., 2019). Web-based tools such as miRNet, miRTarVis+ and ToppmiR have excellent visualization capabilities but also cannot analyze longitudinal datasets (Fan and Xia, 2018; L’Yi et al., 2017; Wu et al., 2014). DREM2 and miARMa-seq handle longitudinal datasets, but do not reduce the volume of data enough for GRN generation (Andrés-León et al., 2016; Schulz et al., 2012). miRComb can use longitudinal data to generate miRNA–mRNA interactions networks, but the networks lack detail on upstream or downstream information, making the output insufficient for GRN generation (Vila-Casadesús et al., 2016). Furthermore, several tools have not been actively maintained so their usability may be diminished.
There is clearly a need for a tool that can integrate, functionally analyze and generate detailed networks from longitudinal miRNA–mRNA datasets, which can then be used to identify GRNs. Here, we present the R/Bioconductor package TimiRGeN, which uses differential expression (DE) data as input to generate small miRNA–mRNA interaction networks. Results from TimiRGeN can be exported to Cytoscape or PathVisio for further bioinformatic analysis (Kutmon et al., 2015; Smoot et al., 2011). The TimiRGeN package thereby provides a much-needed means to generate hypotheses from longitudinal multiomic datasets. To demonstrate the capabilities of the package several datasets were analyzed (see Section 2), including a comprehensive RNAseq time series miRNA–mRNA folic acid (FA)-induced mouse kidney injury dataset (Fig. 1) (Craciun et al., 2016; Pellegrini et al., 2016).
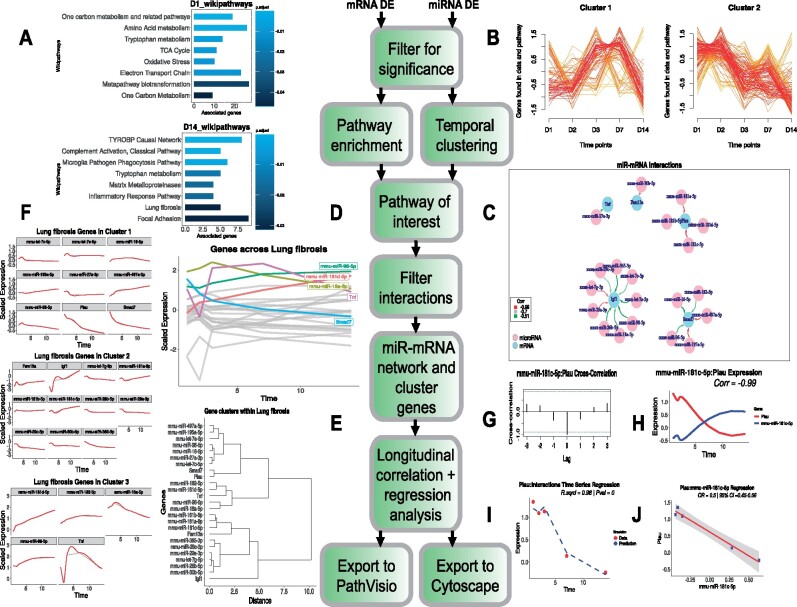
Fig. 1.
Pipeline of the TimiRGeN R package: The FA miRNA–mRNA data are input and filtered for significantly expressed genes for each time point. From here, one of two methods can be used to find WikiPathways of interest. (A) Time-dependent pathway enrichment to find enriched pathways at each time point. The enriched pathways are ranked in descending order of adjusted P-values on bar plots. Results from day 1 and day 14 are shown. Or (B) temporal clustering where global trends of the pathways over time are clustered. Two clusters are shown here. Each line is a pathway and the color represents how well a pathway fits into a cluster. Ranking from highest to lowest are: red, orange, yellow. miRNA–mRNA interactions within a selected signaling pathway can be predicted by filtration of miRNA–mRNA pairs using databases and correlation. (C) Filtered miRNA–mRNA pairs can be viewed in R. Nodes are pink for miRNAs or blue for mRNAs and edges are color coded by correlation over time. (D) Behavior of genes within the miRNA–mRNA interaction network can be viewed across the time course and genes which pass a threshold (>1.5 in this example) are highlighted. (E) The genes can also be hierarchically clustered to identify trends. (F) Expression changes within the clusters can be plotted. These line plots include a gray line (data points) and a red line (smooth spline). (G) A selected miRNA–mRNA pair (mmu-miR-181c-5p and Plau) can be analyzed using cross-correlation analysis. (H) The selected mRNA (red) and miRNA (blue) can also be displayed over the time course. The data are scaled and interpolated over a spline and the correlation is displayed. (I) Regression analysis can be performed on a selected miRNA or mRNA. Plau was selected, so its expression over time is predicted based on the chosen miRNAs that target it. In this example mmu-miR-181c-5p is selected to predict the behavior of Plau. Expression values of Plau are displayed as red dots and the predicted expression of Plau is displayed as a dashed blue line. R2 and P-value are shown. (J) Regression can also be performed between a miRNA–mRNA pair. The OR (odds-ratio) between the two time series can be calculated, along with the 95% CI (confidence intervals). Correlation, R2, P-value, OR and CI are rounded to 2 decimal places. Network data can be exported to PathVisio or Cytoscape
2 Materials and methods
FA data from GSE61328 (miRNA) and GSE65267 (mRNA) were downloaded using the fastqc-dump function from SRA toolkit and fastq files were checked with FastQC (Andrews, 2010; Leinonen et al., 2011). Cutadapt removed adapter sequences from miRNA fastq files, and then the trimmed fastq files were processed with mir2deep (mapper, quantifier and miRDeep2 functions) to produce mature miRNA data which could be imported into R (Friedlander et al., 2012; Martin, 2011). Salmon quant aligned and quantified the mRNA fastq files, and tximport imported the output of Salmon into R (Patro et al., 2017; Soneson et al., 2015). Mouse transcriptome GRCm38.cdna.all was indexed for miRNA processing with Bowtie build and mRNA processing with Salmon index (Cunningham et al., 2019; Langmead and Salzberg, 2012). In R, limma was used for DE analysis (Ritchie et al., 2015). The makeContrasts function performed time point-based DE. The zero time point was contrasted against each subsequent time point (1, 2, 3, 7 and 14 days after FA injection). Results were analyzed with the TimiRGeN R package. For the FA kidney injury dataset, the combined mode of analysis found the ‘Lung fibrosis’ WikiPathway (WP3632) to be consistently enriched during days 3, 7 and 14 of the time course. The ‘Lung fibrosis’ pathway was analyzed for potential miRNA–mRNA interactions. Twenty interactions were kept because they were found in at least two databases and had Pearson correlations lower than −0.5. Results were exported to create a dynamic miRNA integrated Lung fibrosis signaling pathway in PathVisio. CellDesigner was then used to create an SBML formatted GRN (Funahashi et al., 2008). A second mouse kidney injury dataset generated by Unilateral Ureter Obstruction (UUO) was downloaded from GSE118340 (miRNA) and GSE118339 (mRNA) (Pavkovic et al., 2017). UUO and FA datasets were processed and analyzed using the same methods. A 10 time point longitudinal miRNA–mRNA breast cancer dataset was downloaded and processed as is described in Supplementary Data. This dataset underwent two separate analysis with TimiRGeN. Once where DESeq2 was used for pairwise DE and a second time where DESeq2 performed whole time course DE with the LRT method (Baran-Gale et al., 2016; Love et al., 2014). A microarray hypoxia dataset was downloaded from GSE47534 and also put through TimiRGeN analysis (Camps et al., 2014). The lumi and AgiMicroRna packages were used for processing and limma for pairwise DE (Du et al., 2008; López-Romero, 2011). Microarray platforms GPL6884 and GPL8227 were downloaded and gene IDs extracted to create a list of probes for enrichment analysis. Scripts and data for reproducibility are linked to in Supplementary Data.
3 Results
3.1 Time point and miRNA specific analysis
Pairwise miRNA and mRNA DE data (Log2FC and adjusted P-values) from each time point can be used as input for TimiRGeN. The tool works on RNAseq and microarray data, and it has two modes of analysis. The combined mode analyses miRNA and mRNA data from the same time point together, and here each gene from a time point can be filtered for significance independent of all other time points. The separate mode analyses miRNA and mRNA data independent of each other. Separate mode analysis allows for a miRNA or mRNA from a time point to be filtered for significance independent of all other time points and gene types (miRNA or mRNA). TimiRGeN uses WikiPathways for functional analysis, and most are curated by either entrez gene IDs or ensemble gene IDs so TimiRGeN provides both for the user. Neither of these annotation types can distinguish between −3p or −5p miRNAs, thus TimiRGeN also provides adjusted IDs, in case a miRNA–mRNA interaction network is generated with both the −3p and −5p versions of a miRNA.
3.2 Filtering data with time-based functional analysis
TimiRGeN offers two functional analysis methods: time-dependent pathway enrichment and temporal pathway clustering analysis. Both use the rWikiPathways package an API for the WikiPathways database to find signaling pathways of interest (Slenter et al., 2018).
3.2.1 Time-dependent pathway enrichment method
Overrepresentation analysis from clusterProfiler is applied to time series data (Yu et al., 2012). Hypergeometric tests are performed to contrast the number of genes found in common between each time point (after filtering for significantly differentially expressed genes) and each species specific WikiPathway. This produces a list of enriched pathways for each time point (Fig. 1A). Alternatively, if the separate mode of analysis is applied, enrichment analysis is performed for each time point per gene type. The background/universe used to perform overrepresentation analysis can be adjusted by the user e.g. probes in a microarray or all known genes within a cell type.
3.2.2 Temporal pathway clustering method
Temporal pathway clustering (Fig. 1B) utilizes Mfuzz (Kumar and Futschik, 2007). Supervised soft clusters are created based on temporal patterns which stem from the number of genes found in each time point (after filtering for significance) and each species specific WikiPathway. This will show global trends within the dataset. Pathways are assigned fitness scores for each cluster, from 0 to 1, and these can be filtered to find highly correlating pathways in clusters of interest. If the separate mode is used, temporal pathway clustering is performed for each gene type individually.
3.3 Filter miRNA–mRNA interactions from a signaling pathway of interest
After a signaling pathway has been selected for further analysis, the TimiRGeN pipeline will extract each mRNA that is found in common between the selected pathway and the input mRNA data. Each of these mRNAs are assumed to be potential targets of every miRNA in the input data. This results in a miRNA–mRNA interaction matrix which can be used to filter out miRNA–mRNA interactions that are not likely to occur by using correlations and miRNA–mRNA interaction databases TargetScan, miRDB and miRTarBase (Agarwal et al., 2015; Chen and Wang, 2020; Huang et al., 2020). Correlations are calculated for changes over time (Log2fc or average expression) between a given miRNA and a given mRNA. The default method is Pearson, but users can also select between Spearman or Kendall. Since miRNAs negatively regulate mRNAs, highly negative correlation values from miRNA–mRNA pairs could be used to identify miRNA–mRNA interactions that are likely regulate the selected pathway. Users can define a correlation threshold to filter for miRNA–mRNA interactions. The default setting for maximum correlation is −0.5. Three miRNA target databases are also usable to filter for miRNA–mRNA interactions. This includes two predictive target databases (TargetScan and miRDB) and one functional database (miRTarBase) which has had all functional support labeled as ‘weak’ removed. Predictive databases TargetScan and miRDB were selected because, although they have differences in their prediction methods, they share usage of 3ʹUTR-seed site complementarity and seed site conservation to predict miRNA–mRNA interactions (Peterson et al., 2014). Comparisons between different miRNA–mRNA prediction methods find that 3ʹUTR-seed site complementarity identify the most true positive miRNA–mRNA interactions (Mazière and Enright, 2007; Zhang and Verbeek, 2010). Interactions found or not found in the three databases will be represented as 1 or 0, respectively. Users have the option to choose which combination of databases they wish to mine information from and they can choose the number of databases which an interaction needs to be mined from to be filtered. The default setting for the minimum number of databases needed to filter a miRNA–mRNA interaction is 1. Once correlation and databases have been used to filter for miRNA–mRNA interactions which may be affecting the signaling pathway of interest, they can be displayed in an internal R network (Fig. 1C). Resulting genes found in the miRNA–mRNA interaction network can be viewed over the time course. Here, genes that pass a user-defined threshold can be highlighted (Fig. 1D). The genes can also be sorted into hierarchical clusters shown by a dendrogram, from which clusters can be plotted to show the behavior of the genes (Fig. 1E and F). A heatmap, which is compatible with the dendrogram, can also be generated (Supplementary Figure S1).
3.4 Longitudinal miRNA–mRNA pair analysis
The TimiRGeN R package has a suite of longitudinal analysis approaches for analyzing predicted miRNA–mRNA interacting pairs. This includes several correlation- and regression-based methods which are commonly used to analyze longitudinal datasets (Ding and Bar-Joseph, 2020). Cross-correlation analysis is a useful method to determine similarity between two time series (Fig. 1G). If the time series is of sufficient length, the metric can be used to identify delays and further filter for miRNA–mRNA interacting pairs with interesting dynamics (Jung et al., 2011; Lakshmipathy et al., 2007). miRNA–mRNA pairs can also be plotted in a time series line plot. This plot can be scaled and interpolated over a spline (Fig. 1H). Two types of regression analysis can also be performed. First, a linear model is generated from a selected gene (mRNA or miRNA) and any number of its predicted binding partners. The combination of miRNA–mRNA interactions is left for the user to define. The longitudinal behavior of the selected gene is predicted based on the binding partners used in the linear model. The predicted simulation and the gene data are plotted along with the R2 value and P-value (Fig. 1I). This type of regression prediction is useful in cases where a mRNA is targeted by multiple miRNAs or if a miRNA targets multiple mRNAs. Next, a linear model can be created from a single miRNA–mRNA pair. The odds-ratio is calculated from the regression coefficient. This measures the likelihood of one gene influencing the behavior of another gene and has previously been used as a metric to determine miRNA–mRNA relationships (Jayaswal et al., 2009). 95% confidence intervals are calculated which give a range where there is a 95% certainty of the mean of the data being within the range (Fig. 1J) (Szumilas, 2010). Selecting a miRNA–mRNA pair to investigate can be made easier by plotting a heatmap which orders the interacting pairs by descending correlation (Supplementary Figure S2). Statistics generated from correlation and regression analyses may be overestimations if too few time points are found within the input data. Thus, the tool will generate an error if fewer than three time points are detected and warnings if fewer than five time points are detected.
3.5 Output of the TimiRGeN package and exportation of data from R
TimiRGeN is an open-ended tool that exports to networking software PathVisio and Cytoscape for further in silico analysis. The TimiRGeN R package produces two data files for upload onto PathVisio. A file which includes a single result type, e.g. Log2FC, from each time point and gene IDs. This can be uploaded into PathVisio to show how the genes in a signaling pathway of interest change over the time course. Also a file which contains all filtered miRNAs can be uploaded into PathVisio. The second file requires the user to install the MAPPbuilder app in PathVisio (Kutmon et al., 2015). With this, changes over time in a miRNA integrated signaling network of interest can be visualized to show how the miRNAs may be influencing the signaling pathway. This type of display is ideal for bottom-up GRN construction (Supplementary Figure S3). Filtered miRNA–mRNA interactions can also be exported to Cytoscape for improved visualization and alternative analysis via Cytoscape apps (Smoot et al., 2011). The enhanced graphics of Cytoscape are especially useful to visualize large numbers of miRNA–mRNA interactions (Supplementary Figure S4).
3.6 Data from nonpairwise DE
The FA kidney injury dataset had pairwise DE performed using the zero time point as the denominator. This type of pairwise analysis is recommended for time series datasets with <8 time points; however, longer time series datasets may be more suitable for DE without using the pairwise approach e.g. over a cubic spline, maSigPro or the LRT method with DESeq2 (Conesa et al., 2006; Spies et al., 2019). In these cases, users are recommended to filter out significantly differentially expressed genes from averaged count or expression data, and to use this as input for TimiRGeN. Pathway enrichment can be used to identify the most enriched pathways from the whole time course or temporal clustering can first cluster genes based on temporal behavior. From here, genes can be sorted based on clusters, and then pathway enrichment can be used to identify enriched pathways from each temporal cluster. An alternative pipeline is shown in Supplementary Figure S5 and this is explained in Section 5 of the vignette.
3.7 Datasets with multiple interventions
More complex datasets may include interventions other than time. In these cases, TimiRGeN should be used for each individual time series and then the results can be contrasted between different interventions. This requires the same signaling pathway to be explored in each time series. As an example, the ‘Lung fibrosis’ pathway was analyzed in the FA and UUO datasets. A pipeline is shown in Supplementary Figure S6 and Section 6 of the vignette provides detail.
3.8 Hypothesis generation with TimiRGeN
To demonstrate the tools ability to generate biologically relevant hypotheses, the FA mouse kidney injury dataset was analyzed with TimiRGeN (Fig. 1). Findings from the analysis were used to hypothesize how of FA can induce fibrosis. A GRN was constructed to formalize the hypotheses (Fig. 2). Investigation of these results can be used to ratify the miRNA–mRNA interactions predicted by TimiRGeN and make a stronger case for experimental validation. FA injection is known to cause acute injury conditions in the kidneys, resulting in a reversible chronic kidney disease (CKD) like condition (Craciun et al., 2016; Pellegrini et al., 2016). During the 14 day time course, a number of different processes occur, such as inflammatory response, scar tissue forming, wound healing, cytokine activity (Leask and Abraham, 2004). TimiRGeN analysis highlights several of these processes and GRNs were generated to represent how miRNAs may be influencing fibrosis factors (Fig. 2) and scar tissue forming by collagen synthesis (Supplementary Figures S7–S10). The GRN presented in Figure 2 indicates that Igf1 acts as a miRNA sponge. Many of the presented miRNA-Igf1 interactions have been reported, including miR-18a, miR-98, miR-365 and miR-26b (Hu et al., 2013; Liu et al., 2016, 2017; Sun et al., 2019). let-7c-5p has been reported to target Igf1, and TimiRGeN predicted other let-7 family genes let-7e-5p and let-7g-5p also target Igf1 (Liu et al., 2018). Finally, miR29 family members are predicted to target Igf1, and research indicates that Igf1 is a miR-29 family sponge (Gao et al., 2016). It is unknown why Igf1 may be a miRNA sponge, but Igf1 is known to induce collagen production, which contributes to kidney fibrosis and CKD (Hung et al., 2013). Exploration of Igf1 as a miRNA sponge in kidney injury conditions could be beneficial for therapeutics for CKD. Overall, this case study highlights that the TimiRGeN R package can be used to identify biologically relevant miRNA–mRNA interactions from potentially tens-of-thousands of possible miRNA–mRNA interactions. The ability to reduce the volume of big multiomic data is an important feature of TimiRGeN and one which could lead to making miRNA research easier and faster for users. Further analysis on a breast cancer dataset is also found in Supplementary Figures S11–S16.
Fig. 2.
miRNAs influencing antifibrosis factor Tnfa and profibrosis factor Igf1: This GRN shows how FA may be downregulating let-7c-5p, let-7e-5p, let-7g-5p, miR-18a-5p, miR-26b-5p, miR-29a-3p, miR-29c-3p, miR-365-3p and miR-98-5p, which are all predicted to target profibrosis factor Igf1. Also this GRN indicates how FA may upregulate miR-27a-3p, which is predicted to target antifibrosis factor Tnfa. Reduction of Tnfa will increasing levels of profibrosis factor Tgfb
4 Conclusion
As recognized in Bar-Joseph et al. (2012), generation of complex transcriptomic datasets will continue, so computational biologists will need more sophisticated and up-to-date software to analyze these datasets (Bar-Joseph et al., 2012). We have presented a novel R/Bioconductor package which aims to help researchers find direction when working with large longitudinal multiomic datasets. Overall, TimiRGeN is a useful new tool which could become a part of miRNA–mRNA data analysis pipelines.
Funding
K.P., I.M.C. and D.Y. are supported by the Dunhill Medical Trust [R476/0516]. D.P.S. is supported by Novo Nordisk Fonden Challenge Programme: Harnessing the Power of Big Data to Address the Societal Challenge of Aging [NNF17OC0027812]. C.P., D.Y. and S.C. by the MRC and Versus Arthritis as part of the Medical Research Council Versus Arthritis Centre for Integrated Research into Musculoskeletal Ageing (CIMA) [MR/R502182/1].
Conflict of Interest: none declared.
Supplementary Material
Contributor Information
K Patel, Campus for Ageing and Vitality, Biosciences Institute, Newcastle University, Newcastle upon Tyne NE4 5PL, UK.
S Chandrasegaran, Campus for Ageing and Vitality, Biosciences Institute, Newcastle University, Newcastle upon Tyne NE4 5PL, UK.
I M Clark, School of Biological Sciences, University of East Anglia, Norwich NR4 7TJ, UK.
C J Proctor, Campus for Ageing and Vitality, Biosciences Institute, Newcastle University, Newcastle upon Tyne NE4 5PL, UK.
D A Young, Life Science Centre, Biosciences Institute, Newcastle University, Newcastle upon Tyne NE1 4EP, UK.
D P Shanley, Campus for Ageing and Vitality, Biosciences Institute, Newcastle University, Newcastle upon Tyne NE4 5PL, UK.
References
- Agarwal V. et al. (2015) Predicting effective microRNA target sites in mammalian mRNAs. eLife, 4, e05005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrés-León E. et al. (2016) miARma-Seq: a comprehensive tool for miRNA, mRNA and circRNA analysis. Sci. Rep., 6, 25749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrews S. (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- Baran-Gale J. et al. (2016) An integrative transcriptomics approach identifies miR-503 as a candidate master regulator of the estrogen response in MCF-7 breast cancer cells. RNA, 22, 1592–1603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bar-Joseph Z. et al. (2012) Studying and modelling dynamic biological processes using time-series gene expression data. Nat. Rev. Genet., 13, 552–564. [DOI] [PubMed] [Google Scholar]
- Bartel D.P. (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell, 116, 281–297. [DOI] [PubMed] [Google Scholar]
- Barter M.J. et al. (2015) Genome-wide microRNA and gene analysis of mesenchymal stem cell chondrogenesis identifies an essential role and multiple targets for miR-140-5p. Stem Cells, 33, 3266–3280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bisognin A. et al. (2012) MAGIA2: from miRNA and genes expression data integrative analysis to microRNA-transcription factor mixed regulatory circuits. Nucleic Acids Res., 40, W13–W21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camps C. et al. (2014) Integrated analysis of microRNA and mRNA expression and association with HIF binding reveals the complexity of microRNA expression regulation under hypoxia. Mol. Cancer, 13, 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cava C. et al. (2017) SpidermiR: an R/bioconductor package for integrative analysis with miRNA data. Int. J. Mol. Sci., 18, 274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaturvedi P. et al. (2015) Differential miRNA expression in cells and matrix vesicles in vascular smooth muscle cells from rats with kidney disease. PLoS One, 10, e0131589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y., Wang X. (2020) miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res., 48, D127–D131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C. et al. (2017) Urinary miR-21 as a potential biomarker of hypertensive kidney injury and fibrosis. Sci. Rep., 7, 1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craciun F.L. et al. (2016) RNA sequencing identifies novel translational biomarkers of kidney fibrosis. J. Am. Soc. Nephrol., 27, 1702–1713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conesa A. et al. (2006) maSigPro: a method to identify significantly differential expression profiles in time-course microarray experiments. Bioinformatics, 22, 1096–1102. [DOI] [PubMed] [Google Scholar]
- Creighton C.J. et al. (2008) A bioinformatics tool for linking gene expression profiling results with public databases of microRNA target predictions. RNA, 14, 2290–2296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham F. et al. (2019) Ensembl 2019. Nucleic Acids Res., 47, D745–D751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz D. et al. (2017) MicroRNA-augmented pathways (mirAP) and their applications to pathway analysis and disease subtyping. Pac. Symp. Biocomput., 390–401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding J., Bar-Joseph Z. (2020) Analysis of time-series regulatory networks. Curr. Opin. Syst. Biol., 21, 16–24. [Google Scholar]
- Du P. et al. (2008) lumi: a pipeline for processing Illumina microarray. Bioinformatics, 24, 1547–1548. [DOI] [PubMed] [Google Scholar]
- Fan Y., Xia J. (2018) miRNet-functional analysis and visual exploration of miRNA-target interactions in a network context. In: Stechow L.V., Delgado S.A. (Eds.) Computational Cell Biology. Methods in Molecular Biology. Vol. 1819, Humana Press, New York, NY, pp. 215–233. [DOI] [PubMed] [Google Scholar]
- Friedlander M.R. et al. (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res., 40, 37–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Funahashi A. et al. (2008) CellDesigner 3.5: a versatile modeling tool for biochemical networks. Proc. IEEE, 96, 1254–1265. [Google Scholar]
- Gao S. et al. (2016) IGF1 3ʹUTR functions as a ceRNA in promoting angiogenesis by sponging miR-29 family in osteosarcoma. J. Mol. Histol., 47, 135–143. [DOI] [PubMed] [Google Scholar]
- Helwak A. et al. (2013) Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell, 153, 654–665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu Y. et al. (2013) MicroRNA-98 induces an Alzheimer’s disease-like disturbance by targeting insulin-like growth factor 1. Neurosci. Bull., 29, 745–751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H. et al. (2020) miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res., 48, D148–D154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hung C.F. et al. (2013) Role of IGF-1 pathway in lung fibroblast activation. Respir. Res., 14, 102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayaswal V. et al. (2009) Identification of microRNAs with regulatory potential using a matched microRNA-mRNA time-course data. Nucleic Acids Res., 37, e60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung D.E. et al. (2011) Differentially expressed microRNAs in pancreatic cancer stem cells. Pancreas, 40, 1180–1187. [DOI] [PubMed] [Google Scholar]
- Kumar L., Futschik M.E. (2007) Mfuzz: a software package for soft clustering of microarray data. Bioinformation, 2, 5–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kutmon M. et al. (2015) PathVisio 3: an extendable pathway analysis toolbox. PLoS Comput. Biol., 11, e1004085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lakshmipathy U. et al. (2007) MicroRNA expression pattern of undifferentiated and differentiated human embryonic stem cells. Stem Cells Dev., 16, 1003–1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B., Salzberg S.L. (2012) Fast gapped-read alignment with Bowtie2. Nat. Methods, 9, 357–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leask A., Abraham D.J. (2004) TGF-B signaling and the fibrotic response. FASEB J., 18, 816–827. [DOI] [PubMed] [Google Scholar]
- Leidinger P. et al. (2013) A blood based 12-miRNA signature of Alzheimer disease patients. Genome Biol., 14, R78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leinonen R. et al. (2011) The sequence read archive. Nucleic Acids Res., 39, D19–D21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H. et al. (2016) MicroRNA-26b is upregulated in a double transgenic mouse model of Alzheimer’s disease and promotes the expression of amyloid-B by targeting insulin-like growth factor 1. Mol. Med. Rep., 13, 2809–2814. [DOI] [PubMed] [Google Scholar]
- Liu C. et al. (2017) miR-18a induces myotubes atrophy by down-regulating IgfI. Int. J. Biochem. Cell Biol., 90, 145–154. [DOI] [PubMed] [Google Scholar]
- Liu G.-X. et al. (2018) Hsa-let-7c controls the committed differentiation of IGF1-treated mesenchymal stem cells derived from dental pulps by targeting IGF-1R via the MAPK pathways. Exp. Mol. Med., 50, 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- López-Romero P. (2011) Pre-processing and differential expression analysis of Agilent microRNA arrays using the AgiMicroRna Bioconductor library. BMC Genomics, 12, 64–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love M.I. et al. (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol., 15, 550–521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- L’Yi S. et al. (2017) miRTarVis+: web-based interactive visual analytics tool for microRNA target predictions. Methods, 124, 78–88. [DOI] [PubMed] [Google Scholar]
- Martin M. (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J., 17, 10–12. [Google Scholar]
- Mazière P., Enright A. (2007) Prediction of microRNA targets. Drug Discov., 12, 452–458. [DOI] [PubMed] [Google Scholar]
- Miyaki S. et al. (2010) MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes Dev., 24, 1173–1185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore M.J. et al. (2015) miRNA-target chimeras reveal miRNA 3-end pairing as a major determinant of Argonaute target specificity. Nat. Commun., 6, 1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ooi C.Y. et al. (2018) Network modeling of microRNA-mRNA interactions in neuroblastoma tumorigenesis identifies miR-204 as a direct inhibitor of MYCN. Cancer Res., 78, 3122–3134. [DOI] [PubMed] [Google Scholar]
- Patro R. et al. (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods, 14, 417–419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavkovic M. et al. (2017) Multi omics analysis of fibrotic kidneys in two mouse models. Sci. Data, 6, 92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pellegrini K.L. et al. (2016) Application of small RNA sequencing to identify microRNAs in acute kidney injury and fibrosis. Toxicol. Appl. Pharmacol., 312, 42–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson S.M. et al. (2014) Common features of microRNA target prediction tools. Front. Genet., 5, 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proctor C.J. et al. (2017) Computer simulation models as a tool to investigate the role of microRNAs in osteoarthritis. PLoS One, 12, e0187568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin S. et al. (2015) Gene regulatory networks by transcription factors and microRNAs in breast cancer. Bioinformatics, 31, 76–83. [DOI] [PubMed] [Google Scholar]
- Ritchie M.E. et al. (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res., 43, e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulz M.H. et al. (2012) DREM 2.0: improved reconstruction of dynamic regulatory networks from time-series expression data. BMC Syst. Biol., 6, 104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarzenbach H. et al. (2014) Clinical relevance of circulating cell-free microRNAs in cancer. Nat. Rev. Clin. Oncol., 11, 145–156. [DOI] [PubMed] [Google Scholar]
- Selbach M. et al. (2008) Widespread changes in protein synthesis induced by microRNAs. Nature, 455, 58–63. [DOI] [PubMed] [Google Scholar]
- Slenter D.N. et al. (2018) WikiPathways: a multifaceted pathway database bridging metabolomics to other omics research. Nucleic Acids Res., 46, D661–D667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smoot M.E. et al. (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics, 27, 431–432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soneson C. et al. (2015) Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research, 4, 1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spies D. et al. (2019) Comparative analysis of differential gene expression tools for RNA sequencing time course data. Brief. Bioinform., 20, 288–298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun W. et al. (2019) miR-365 inhibits duck myoblast proliferation by targeting IGF-I via PI3K/Akt pathway. Biosci. Rep., 39, BSR20190295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szumilas M. (2010) Explaining odds ratios. J. Am. Acad. Child Adolesc. Psychiatry, 19, 227. [PMC free article] [PubMed] [Google Scholar]
- Vila-Casadesús M. et al. (2016) MiRComb: an R package to analyse miRNA-mRNA interactions. Examples across five digestive cancers. PLoS One, 11, e0151127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang T.-T. et al. (2019) anamiR: integrated analysis of MicroRNA and gene expression profiling. BMC Bioinform., 20, 239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C. et al. (2014) ToppMiR: ranking microRNAs and their mRNA targets based on biological functions and context. Nucleic Acids Res., 42, W10–W113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G. et al. (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. Omics, 16, 284–287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Verbeek F.J. (2010) Comparison and integration of target prediction algorithms for microRNA studies. J. Integr. Bioinform., 7, 169–181. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.