Abstract
Background & Aims
The epithelial barrier is the host’s first line of defense against damage to the underlying tissue. Upon injury, the epithelium plays a critical role in inflammation. The IκB kinase β (IKKβ)/nuclear factor-κB pathway was shown to be active in the esophageal epithelium of patients with esophageal disease. However, the complex mechanisms by which IKKβ signaling regulates esophageal disease pathogenesis remain unknown. Our prior work has shown that expression of a constitutively active form of IKKβ specifically in esophageal epithelia of mice (IkkβcaEsophageal Epithelial Cell-Knockin (EEC-KI)) is sufficient to cause esophagitis.
Methods
We generated ED-L2/Cre;Rosa26-Ikkβca+/L;Stat3L/L (IkkβcaEEC-KI;Stat3Esophageal Epithelial Cell Knockout (EEC-KO)) mice, in which the ED-L2 promoter activates Cre recombinase in the esophageal epithelium, leading to constitutive activation of IKKβ and loss of Stat3. Esophageal epithelial tissues were collected and analyzed by immunostaining, RNA sequencing, quantitative real-time polymerase chain reaction assays, flow cytometry, and Western blot. IkkβcaEEC-KI mice were treated with neutralizing antibodies against interleukin (IL)23p19 and IL12p40.
Results
Here, we report that IkkβcaEEC-KI mice have increased activation of epithelial Janus kinase 2/STAT3 signaling. Stat3 deletion in IkkβcaEEC-KI mice attenuated the neutrophil infiltration observed in IkkβcaEEC-KI mice and resulted in decreased expression of genes related to immune cell recruitment and activity. Blocking experiments in IkkβcaEEC-KI mice showed that STAT3 activation and subsequent neutrophil recruitment are dependent on IL23 secretion.
Conclusions
Our study establishes a novel interplay between IKKβ and STAT3 signaling in epithelial cells of the esophagus, where IKKβ/IL23/STAT3 signaling controls neutrophil recruitment during the onset of inflammation. GEO accession number: GSE154129.
Keywords: Gene Regulation, Inflammation, Immune Regulation, Transcription Factor
Abbreviations used in this paper: CXCL5, C-X-C motif chemokine ligand 5; ESCC, esophageal squamous cell cancer; EGFP, enhanced green fluorescent protein; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; IKKβ, IκB kinase β; JAK, Janus kinase; IL, interleukin; KEGG, Kyoto Encyclopedia Genes and Genomes; MPO, myeloperoxidase; mRNA, messenger RNA; NF-κB, nuclear factor-κB; PCR, polymerase chain reaction; p-STAT3, phosphorylated signal transducer and activator of transcription 3; STAT3, signal transducer and activator of transcription 3
Graphical abstract
Summary.
We show that IκB kinase β activation induces interleukin 23 expression, leading to the activation of epithelial signal transducer and activator of transcription 3 signaling, which then results in the release of neutrophil chemoattractants and the subsequent recruitment of neutrophils to the esophagus.
Diseases of the esophagus are a significant health burden in the United States and throughout the world.1 Most esophageal diseases arise in the context of inflammation and injury.2, 3, 4 For example, esophageal squamous cell cancer (ESCC) accounts for more than 90% of esophageal cancer and develops progressively from inflammation, hyperplasia, dysplasia, carcinoma in situ, to invasive carcinoma.5,6 Epithelial cells are the first line of defense for the esophagus, which is constantly challenged by exposure to irritants (alcohol, tobacco, food antigens, and gastric acid reflux). In response to injuries, the cross-talk between epithelial cells and immune cells is essential to re-establish homeostasis. Epithelial cells can initiate inflammation through production of cytokines and chemokines, promoting the recruitment and amplification of immune cells. Furthermore, epithelial cells can respond to and amplify signals from infiltrating immune cells. However, the molecular mechanisms by which epithelial cells initiate inflammation and wound repair in the esophagus are not fully understood.
IκB kinase β (IKKβ)/nuclear factor-κB (NF-κB) is a central signaling hub in inflammatory-mediated responses.7 Members of the IKKβ/NF-κB family are pivotal for the maintenance of homeostasis by controlling diverse biological processes, such as inflammation, immunity, cell survival, and cell growth.8 The IKKβ/NF-κB pathway is activated through physiological mechanisms such as local tissue damage or inflammatory cytokine signaling.9,10 In addition to regulating transcription and secretion of cytokines and chemokines, IKKβ signaling also modulates classic inflammatory mediators such as signal transducer and activator of transcription 3 (STAT3).7 STAT3 signaling is activated by the Janus kinases (JAKs) in response to growth factors and cytokines, such as interleukin (IL)6, IL10, IL22, IL23, and epidermal growth factor, and controls diverse biological processes.7,11, 12, 13 Different models of inflammation and cancer have shown the existence of a complex interplay between IKKβ and STAT3 signaling pathways.7,10 This interplay can occur in either an autocrine or paracrine manner, and depends on the cellular context and target gene investigated.14,15 For example, in colitis, IKKβ/NF-κB controls the expression of chemokines that are essential for STAT3 activation in intestinal epithelial cells.16 Both transactivation and repression of target genes have been reported to result from direct interaction or cooperative binding between STAT3 and different NF-κB subunits, varying based on the tissue context and specific output genes studied.17, 18, 19
We recently showed that expression of a constitutively active form of IKKβ in esophageal epithelia of mice is sufficient to cause esophageal inflammation and epithelial hyperplasia in vivo.20 This model permits the unique opportunity to study the interactions between the esophageal epithelium and immune cells during the early stages of inflammation and hyperplasia, which are precursor conditions leading to ESCC development. In this study, we show that epithelial IKKβ signaling modulates the activation of the key inflammatory mediator STAT3 in esophageal epithelia of mice in vivo. The goal of this study was to delineate the complex interplay between IKKβ and STAT3 signaling in the context of esophagitis. Elucidating mechanisms underlying this complex interplay may lead to the identification of new therapeutic targets for esophagitis.
Results
Activation of Epithelial IKKβ Signaling Leads to STAT3 Activation in Esophageal Epithelial Cells
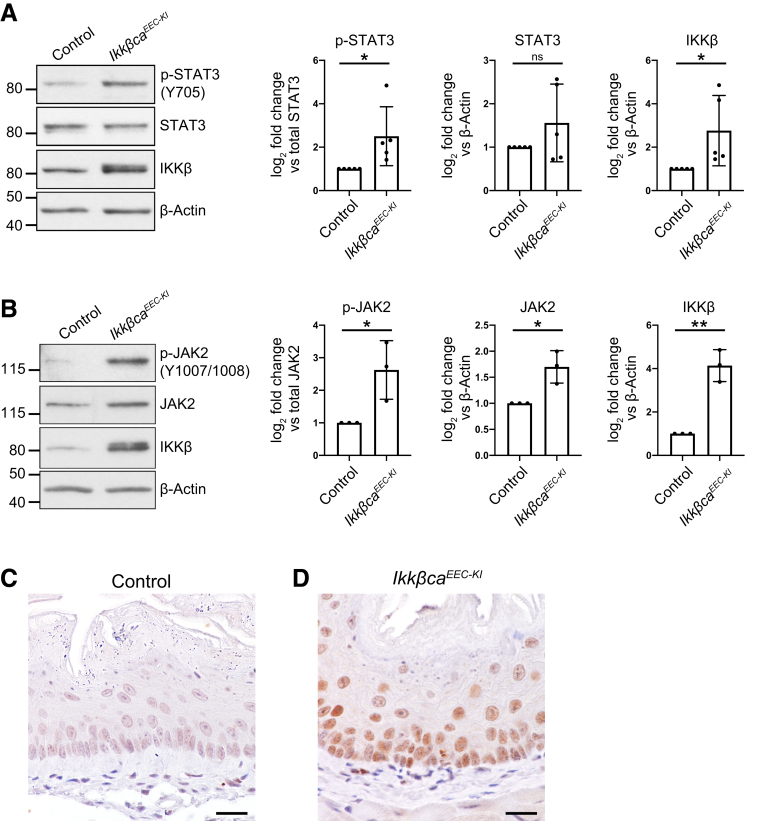
We recently showed that activation of IKKβ in esophageal epithelia of mice using the Epstein–Barr virus ED-L2 promoter leads to esophagitis and epithelial hyperplasia.20 In addition to the IKKβ/NF-κB pathway, STAT3 is another central mediator of inflammatory-mediated responses that is pivotal for the maintenance of homeostasis.12,13 A complex cross-talk between IKKβ/NF-κB and STAT3 signaling has been observed in numerous inflammatory diseases.7,10 We first analyzed STAT3 phosphorylation levels in esophageal epithelia of mice with constitutive activation of IKKβ (ED-L2/Cre;Rosa26-Ikkβca+/loxp (L), further referred to as Ikkβca Esophageal Epithelial Cell-Knockin (EEC-KI)). As shown in Figure 1A, increased STAT3 phosphorylation levels were observed in esophageal epithelia of IkkβcaEEC-KI mice compared with littermate controls. STAT3 is activated primarily through phosphorylation of the Y705 residue by JAK proteins, which are activated to transduce signals after association with membrane-bound receptors.12 Increased activation of JAK2 was observed in esophageal epithelial enrichments from IkkβcaEEC-KI mice compared with control mice by Western blot (Figure 1B). To confirm this STAT3 activation was epithelial in origin, we performed immunohistochemical staining for phosphorylated STAT3 on Y705 (p-STAT3). Strong nuclear staining for p-STAT3 was seen in epithelial cells of IkkβcaEEC-KI mice (Figure 1D) compared with littermate control mice (Figure 1C).
Figure 1.
Activation of epithelial IKKβ signaling leads to increased epithelial STAT3 activation in mouse esophagus. (A and B) Western blot determining (A) STAT3 or (B) JAK2 phosphorylation levels in esophageal epithelia of mice with constitutive IKKβ activation. IkkβcaEEC-KI mice have increased levels of (A) p-STAT3 (Y705), (B) phospho-JAK2 (Y1007/8), and (A and B) IKKβ compared with littermate controls. β-actin was used as a loading control. (A and B) Right: Densitometric analysis of Western blot. p-STAT3 and phospho-JAK2 values were normalized to total protein expression, and total protein values were normalized to loading control. Fold change was calculated based on experimental control. (A) n = 5 mice per experimental group, ∗P < .05. (B) n = 3 mice per experimental group, ∗P < .05, ∗∗P < .01. Bar graphs represent means ± SD. All statistics were determined by a 2-tailed Student t test. (C and D) Immunohistochemical staining for p-STAT3 (Y705) in esophageal sections of (C) control and (D) IkkβcaEEC-KI mice. Immunostaining shows increased nuclear expression of p-STAT3 (Y705) in esophageal epithelial cells from (D) IkkβcaEEC-KI mice compared with (C) controls. n = 6 mice per experimental group. Scale bar: 50 μm.
STAT3Loss Attenuates the Phenotype ofIKKβcaEEC-KI Mice
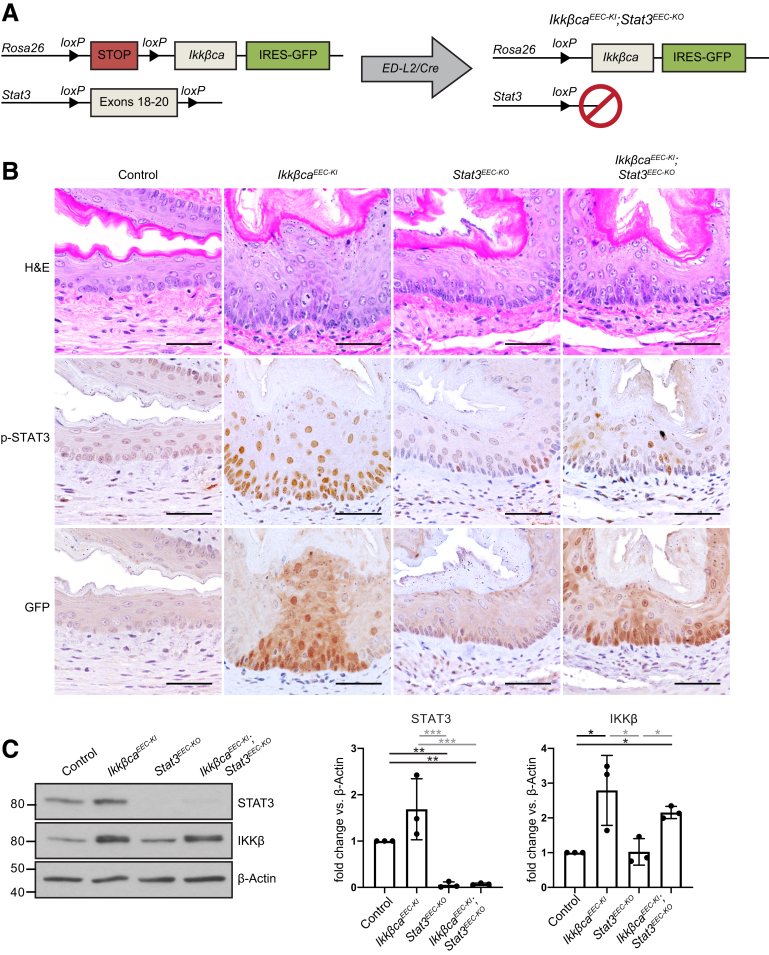
To determine the role of epithelial STAT3 signaling after IKKβ activation, we generated mice with both IKKβ constitutive activation and Stat3 deletion in esophageal epithelial cells. To produce specific loss of Stat3 in esophageal epithelial cells, Stat3 floxed mice21 were crossed with ED-L2/Cre mice (Figure 2A). Mice expressing both the Rosa26-STOPFloxed (FL)-Ikkβca construct and floxed Stat3 were crossed with ED-L2/Cre–expressing mice, generating mice with both activation of IKKβ and loss of Stat3 in esophageal epithelial cells (ED-L2/Cre;Rosa26-Ikkβca+/L;Stat3 Loxp/LoxpL/L, further referred to as IkkβcaEEC-KI;Stat3EEC-KO) (Figure 2A). Mice with loss of Stat3 in esophageal epithelial cells (ED-L2/Cre;Stat3L/L, further referred to as Stat3EEC-KO) were of normal size and weight and had no overt phenotype compared with controls (Figure 2B). IkkβcaEEC-KI;Stat3EEC-KO mice had an attenuated phenotype compared with IkkβcaEEC-KI mice (Figure 2B), with less apparent esophageal epithelial hyperplasia and immune cell infiltration. As shown in Figure 2A, an enhanced green fluorescent protein (EGFP) reporter gene was expressed downstream of the Rosa26-STOPFL-Ikkβca construct in IkkβcaEEC-KI mice22; thus, immunohistochemical staining for GFP was used to detect the expression of the Ikkβca transgene in mice. Serial sections of GFP and H&E stain confirmed that changes in inflammation and hyperplasia occurred in epithelial areas expressing the Ikkβca transgene, with attenuation seen in IkkβcaEEC-KI;Stat3EEC-KO mice compared with IkkβcaEEC-KI (Figure 2B). Immunohistochemical staining for p-STAT3 further showed that increased STAT3 activation was seen in IkkβcaEEC-KI mice and that the attenuated phenotype observed in IkkβcaEEC-KI;Stat3EEC-KO mice occurred in epithelial areas expressing the Ikkβca construct with no to little STAT3 activation (Figure 2B). Loss of Stat3 and increased expression of IKKβ in esophageal epithelia of mice were confirmed by Western blot (Figure 2C).
Figure 2.
Loss of epithelial Stat3 attenuates the phenotype of IkkβcaEEC-KImice. (A) Schematic representation of the generation of IkkβcaEEC-KI;Stat3EEC-KO mice. ED-L2/Cre mice were crossed with Rosa26-STOPFL-Ikkβca-EGFP mice and with Stat3 floxed mice. (B) H&E staining of esophageal sections show that basal cell hyperplasia and immune cell infiltration are attenuated in IkkβcaEEC-KI;Stat3EEC-KO mice when compared with IkkβcaEEC-KI mice. Stat3EEC-KO mice show no significant histologic changes compared with controls. Serial sections were used to confirm expression of GFP and p-STAT3 (Y705), along with H&E staining. Immunostaining shows GFP expression in esophageal epithelial cells of IkkβcaEEC-KI and IkkβcaEEC-KI;Stat3EEC-KO mice compared with control and Stat3EEC-KO mice with no GFP expression. Increased nuclear expression of p-STAT3 (Y705) is seen in esophageal epithelial cells from IkkβcaEEC-KI mice compared with controls, IkkβcaEEC-KI;Stat3EEC-KO and Stat3EEC-KI mice. n = 6 mice per experimental group. Scale bar: 100 μm. (C) Expression levels of IKKβ and STAT3 in protein lysates enriched for esophageal epithelial cells by immunoblot. Immunoblot analyses confirm increased IKKβ expression levels in IkkβcaEEC-KI and IkkβcaEEC-KI;Stat3EEC-KO mice and decreased STAT3 expression in Stat3EEC-KO and IkkβcaEEC-KI;Stat3EEC-KO mice. β-actin is used as a loading control. Right: Densitometric analysis of Western blot. All values were normalized to loading control. Fold change was calculated based on experimental control. n = 3 mice per experimental group. ∗P < .05, ∗∗P < .01, and ∗∗∗P < .001. Bar graphs represent means ± SD. All statistics were determined by 1-way analysis of variance (ANOVA) with the post hoc Tukey multiple comparison test. IRES, Internal Ribosome Entry Site; loxP, locus of X-over P1.
IKKβ and STAT3 Control Both Similar and Distinct Processes in Esophageal Epithelial Cells
Both the IKKβ/NF-κB and STAT3 pathways can converge to regulate transcription of a number of common genes, although these are known to vary based on tissue type, cell type, and disease context. To identify potential targets of both STAT3 and IKKβ in esophageal epithelia, we performed messenger RNA (mRNA) sequencing on esophageal epithelial enrichments from IkkβcaEEC-KI mice, Stat3EEC-KO mice, IkkβcaEEC-KI;Stat3EEC-KO mice, and controls. To compare all of these groups, differential gene expression was determined in each group compared with control mice, resulting in 3 separate pairwise comparisons (Figure 3A). Overall, 271 genes across all comparisons were significantly differentially regulated (false-discovery rate–adjusted P value < .05, |log2 fold change| ≥ 1). IkkβcaEEC-KI mice accounted for 216 of these genes (Figure 3A). Of these 216 genes, 153 were not differentially regulated in IkkβcaEEC-KI;Stat3EEC-KO mice compared with controls (Figure 3B), indicating that they are activated by IKKβ and also are dependent on STAT3 because the loss of Stat3 abolished differential expression. As seen in Figure 3C, the log2 fold changes of these 153 differentially expressed genes showed inverse expression patterns in IkkβcaEEC-KI;Stat3EEC-KO mice compared with IkkβcaEEC-KI mice. The remaining 63 genes were differentially expressed in both comparisons of IkkβcaEEC-KI mice and IkkβcaEEC-KI;Stat3EEC-KO mice against controls (Figure 3B). Thus, these 63 genes are dependent on IKKβ but not dependent on STAT3 because they remained differentially expressed even in mice lacking Stat3 expression. Expression of these 63 genes was directionally consistent across both groups as either up-regulated or down-regulated (Figure 3C). Loss of Stat3 alone from esophageal epithelial cells did little to change the mouse transcriptome, resulting in only 5 total significantly regulated genes (Figure 3A). Lists of differentially expressed genes from IkkβcaEEC-KI mice, Stat3EEC-KO mice, and IkkβcaEEC-KI;Stat3EEC-KO mice compared with controls were compiled and used for Gene Ontology pathway enrichment analysis (Figure 3D, Supplementary Tables 1–3). IkkβcaEEC-KI mice showed increased gene regulation in 18 pathways related to cytokine signaling, immune cell recruitment, and immune cell activation when compared with control. This effect was either partially or completely attenuated in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 3D), which suggests that cooperative IKKβ and STAT3 signaling in the esophageal epithelium is an important regulator of immune cell activation and recruitment to the esophagus.
Figure 3.
Transcriptomic analysis identifies pathways differentially regulated by STAT3 downstream of activation of IKKβ. (A) RNA sequencing was performed on samples enriched for esophageal epithelial cells from mice. The number of differentially expressed genes (DEGs) from IkkβcaEEC-KI, Stat3EEC-KO, and IkkβcaEEC-KI;Stat3EEC-KO mice as compared with control are shown. n = 3 mice per experimental group. (B) Venn diagrams showing overlap of DEGs from each comparison. (C) Heat map representing the log2 fold change for all DEGs identified from RNA sequencing across all comparisons. (D) DEGs from each comparison were used to identify enrichment of functional pathways by Gene Ontology analysis. A heatmap of the top enriched pathways is shown. Ctrl, control; ERK, extracellular signal-regulated kinase; FDR, false-discovery rate, log2FC, log2 fold change.
STAT3Regulates the Esophageal Recruitment of Neutrophils Downstream of IKKβ Activation
As seen in Figure 3, our RNA sequencing analyses indicate global changes in immune cell recruitment in the esophagus of IkkβcaEEC-KI mice and IkkβcaEEC-KI;Stat3EEC-KO mice. We next sought to determine the precise composition of immune cells enriched within the esophagus. We developed a flow cytometry–based immunophenotyping panel to analyze the live, CD45+ fraction of esophageal cells for T cells (CD3+), helper T cells (CD3+, CD4+), regulatory T cells (CD3+ CD4+ Forkhead Box P3 (FoxP3) +), cytotoxic T lymphocytes (CD3+, CD8+), B cells (CD19+), macrophages (F4/80+), dendritic cells (CD11c+), NK cells (CD49b+ Natural killer (NK) 1.1+), eosinophils (CD11b+ Sialic acid binding immunoglobulin-like lectin (Siglec)-F+), and neutrophils (CD11b+ Lymphocyte antigen 6 complex locus G (Ly6G)hi). Of these subsets, neutrophils were enriched significantly in the esophagus of IkkβcaEEC-KI mice compared with controls (Figure 4A and B). This response was attenuated in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4A and B). No enrichment of neutrophils was detected in Stat3EEC-KO mice (Figure 4A and B). Immunostaining for myeloperoxidase (MPO), a neutrophil marker, confirmed the presence of neutrophils in both the lamina propria and epithelium of the esophagus of IkkβcaEEC-KI mice (Figure 4C). In IkkβcaEEC-KI;Stat3EEC-KO mice, fewer numbers of intraepithelial neutrophils were detected, and no neutrophils were found in esophageal tissue sections of control or Stat3EEC-KO mice (Figure 4C). GFP and p-STAT3 immunostaining along with MPO confirmed that recruitment of neutrophils occurred in epithelial areas with GFP expression and increased STAT3 activation in IkkβcaEEC-KI mice and that decreased neutrophil infiltration was observed in epithelial areas with increased GFP and decreased STAT3 activation in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4C). Neutrophils additionally were enriched in peripheral blood and spleens of IkkβcaEEC-KI mice compared with controls by flow cytometry (Figure 4D and E), a change that was attenuated in spleens of IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4E). Further supporting the observation of neutrophil recruitment after activation of epithelial IKKβ signaling, increased mRNA expression levels of the proinflammatory genes Lcn2 and S100a8 were detected in esophageal epithelial enrichments from IkkβcaEEC-KI mice (Figure 4F and G). Both the expression of Lcn2 and S100a8 indicate the accumulation of intraepithelial neutrophils,23,24 and increased expression of both transcripts were reduced in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4F and G). Furthermore, expression of the potent neutrophil chemoattractant Cxcl525 also was up-regulated in esophageal epithelial enrichments from IkkβcaEEC-KI mice and was reduced in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4H). Overall, this indicates that STAT3 is a key player downstream of IKKβ in the induction of a proinflammatory response and neutrophil recruitment.
Figure 4.
Loss of epithelial Stat3 reduces the recruitment of neutrophils in the esophagus of IKKβcaEEC-KImice. (A and B) Flow cytometry was used to quantify immune cells isolated from mouse esophagi. (A) Live CD45+/CD11b+/Ly6Ghi cells were identified as neutrophils, as seen by representative contour plots. (B) The percentage of CD11b+/Ly6Ghi neutrophils per total CD45+ cells show that loss of Stat3 significantly diminishes the presence of neutrophils in the esophagus of IkkβcaEEC-KI mice. n = 6 mice per experimental group. ∗∗∗P < .001, ∗∗∗∗P < .0001. Bar graphs represent means ± SD. Statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. (C) Representative immunofluorescence staining for MPO is shown in mouse esophageal sections. Infiltration of MPO-positive cells into the esophagus is diminished in IkkβcaEEC-KI;Stat3EEC-KO mice compared with IkkβcaEEC-KI mice. Serial sections were used to examine MPO, GFP, and p-STAT3 expression. Dotted white line indicates the location of the basement membrane. n = 3 mice per experimental group. Scale bars: 100 μm. (D and E) Bar graphs showing the fraction of neutrophils per total CD45+ cells isolated from mouse (D) peripheral blood and (E) spleen. n = 6 mice per experimental group. Bar graphs represent means ± SD. Statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. ∗P < .05, ∗∗P < .01, and ∗∗∗∗P < .0001. (F–H) Using quantitative PCR, mRNA expression levels of the neutrophil-expressed genes (F) Lcn2 and (G) S100a8, and of the neutrophil chemoattractant gene (H) Cxcl5 in enriched mouse esophageal epithelia were decreased in IkkβcaEEC-KI;Stat3EEC-KO mice compared with IkkβcaEEC-KI mice. Bar graphs represent means ± SD. Statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. (F) n = 3 mice per experimental group, ∗P < .05, ∗∗P < .01, and ∗∗∗∗P < .0001. (G) n = 3 mice per experimental group, ∗∗∗P < .001 and ∗∗∗∗P < .0001. (H) n = 4 mice per experimental group, ∗P < .05, ∗∗∗P < .001, and ∗∗∗∗P < .0001. DAPI, 4’,6-diamidino-2-phenylindole.
IKKβ Activates STAT3 Through Increased Expression/Secretion of IL23
We next sought to determine whether STAT3 activation occurred in a cell-autonomous or non–cell-autonomous manner downstream of activated IKKβ signaling. When performing immunohistochemistry on serial sections for GFP to detect expression of the Ikkβca transgene in control and IkkβcaEEC-KI mice, we observed that GFP was detected in distinct clonal populations of esophageal epithelial cells in IkkβcaEEC-KI mice (Figure 5B), but was absent in control mice (Figure 5A). Interestingly, GFP staining did not co-localize with p-STAT3 staining in IkkβcaEEC-KI mice other than in a few cells (Figure 5B and D). This suggests mostly non–cell-autonomous activation of STAT3 by epithelial IKKβ signaling, with the possibility of some cell-autonomous activation to a lesser extent.
Figure 5.
Increased IL23 expression/secretion regulates epithelial STAT3 activation and the recruitment of neutrophils to the esophagus. (A–D) Immunohistochemical staining for (A and B) GFP or (C and D) p-STAT3 (Y705) in esophageal serial sections of (A and C) control and (B and D) IKKβcaEEC-KI mice. Immunostaining shows GFP expression (dotted lines) specifically in esophageal epithelial cells of (B) IkkβcaEEC-KI mice, compared with (A) control mice with no GFP expression. Increased nuclear expression of p-STAT3 (Y705) is seen in esophageal epithelial cells from (D) IkkβcaEEC-KI mice compared with (C) controls. (D) In addition, increased intensity of epithelial p-STAT3 is seen outside of GFP-positive regions (black arrowheads) in IkkβcaEEC-KI mice. n = 4 mice per experimental group. Scale bar: 50 μm. (E) Heat map representing the log2 fold change for the JAK-STAT pathway–associated differentially expressed genes from RNA sequencing. (F) Table summarizing the log2 fold change for Il23a expression levels from RNA sequencing. (G) By quantitative PCR, mRNA expression levels of the cytokine Il23a in mouse esophageal mucosa are increased in IkkβcaEEC-KI and IkkβcaEEC-KI;Stat3EEC-KO mice compared with control mice. n = 3 mice per experimental group. Bar graphs represent means ± SD. Statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. ∗∗P < .01 and ∗∗∗P < .001. (H) IkkβcaEEC-KI and control mice were treated with either an IgG1 control or an IL23p19 neutralizing antibody every Monday and Thursday for 2 weeks and analyzed by immunoblot for p-STAT3 (Y705) levels. Blocking IL23p19 decreases p-STAT3 (Y705) levels in Ikkβca mice compared with IgG treatment. Increased IKKβ expression was observed in IkkβcaEEC-KI mice compared with controls, and GAPDH was used as a loading control. Right: Densitometric analysis of Western blot. p-STAT3 values were normalized to total protein expression, and total protein values were normalized to loading control. Fold change was calculated based on experimental control. n = 3 mice per experimental group. ∗P < .05, ∗∗P < .01, ∗∗∗P < .001, and ∗∗∗∗P < .0001. Bar graphs represent means ± SD. All statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. (I) IkkβcaEEC-KI and control mice were treated with either an IgG2a control or an IL12p40 neutralizing antibody every Monday and Thursday for 2 weeks and analyzed by immunoblot for p-STAT3 (Y705) levels. Blocking IL12p40 decreases p-STAT3 (Y705) levels in Ikkβca mice compared with IgG treatment. Increased IKKβ expression was observed in IkkβcaEEC-KI mice compared with controls, and GAPDH was used as a loading control. Right: Densitometric analysis of Western blot. p-STAT3 values were normalized to total protein expression, and total protein values were normalized to loading control. Fold change was calculated based on experimental control. n = 3 mice per experimental group. ∗P < .05, ∗∗P < .01, ∗∗∗P < .001, and ∗∗∗∗P < .0001. Bar graphs represent means ± SD. All statistics were determined by 1-way analysis of variance with the post hoc Tukey multiple comparison test. (J and K) Flow cytometry was used to quantify neutrophils isolated from mouse esophagi. (J) Live CD45+/CD11b+/Ly6Ghi cells were identified as neutrophils, as seen by representative contour plots. (K) The total counts of CD11b+/Ly6Ghi neutrophils show that IL23 p19 neutralization significantly diminishes the presence of neutrophils in the esophagus of IkkβcaEEC-KI mice. Because the cell counts were not normally distributed, these were log-transformed for statistical analyses. n = 5 mice per experimental group. Bar graphs represent means ± SD. Statistics were determined by 1-way analysis of variance with the Sidak multiple comparison test. ∗P < .05, ∗∗P < .01. Ctrl, control; FDR, false-discovery rate; log2FC, log2 fold change.
JAK/STAT signaling is activated by secreted factors, including cytokines and growth factors.12 To determine if IKKβ activates STAT3 through the regulation of secreted factors, we looked back at our RNA sequencing data to establish if known STAT3 activators were up-regulated in IkkβcaEEC-KI mice compared with controls. Kyoto Encyclopedia Genes and Genomes (KEGG) pathway analyses showed enrichment of the JAK/STAT signaling pathway in IkkβcaEEC-KI mice compared with controls, along with the key genes responsible for this enrichment (Figure 5E). Only 2 of these genes, Il23a and Il7r, also were up-regulated significantly in IkkβcaEEC-KI;Stat3EEC-KO mice compared with controls (Figure 5E). Notably, the IL23 signaling cascade is known to activate STAT3 signaling.26,27 We found a 2.4 log2 fold change in mRNA expression levels of Il23a in IkkβcaEEC-KI mice compared with controls, an increase unaffected by the loss of Stat3 (Figure 5F). We confirmed these differences in Il23a expression levels by quantitative polymerase chain reaction (PCR). As shown in Figure 5G, Il23a was up-regulated in esophageal mucosa from IkkβcaEEC-KI mice and in IkkβcaEEC-KI;Stat3EEC-KO mice compared with controls. IL23 is a heterodimeric cytokine composed of the IL23p19 subunit (encoded by Il23a) and the IL12p40 subunit (encoded by Il12b).27 Although IL23p19 is specific for IL23, IL12p40 is shared between IL23 and IL12.28 Interestingly, we previously reported increased Il12b mRNA expression levels in IkkβcaEEC-KI mice compared with controls before the onset of inflammation.20 To show that IL23 was required for STAT3 activation downstream of IKKβ activation, we treated IkkβcaEEC-KI and control mice with a neutralizing antibody against IL23p19, IL12p40, or IgG controls. Increased STAT3 phosphorylation was detected by Western blot in IkkβcaEEC-KI mice treated with IgG compared with controls treated with IgG (Figure 5H and I). IL23p19 and IL12p40 neutralization attenuated STAT3 phosphorylation levels in IkkβcaEEC-KI mice compared with IkkβcaEEC-KI mice treated with IgG controls by 40% and 33%, respectively (Figure 5H and I). To show that IL23 operates in the same signaling axis upstream of STAT3 and downstream of epithelial IKKβ activation, we next treated IkkβcaEEC-KI and control mice with in vivo neutralizing anti-IL23p19 or IgG1 antibodies and determined neutrophil counts by flow cytometry. As shown in Figure 5J and K, neutrophil recruitment was increased significantly in the esophagus of IkkβcaEEC-KI mice treated with IgG1 compared with controls treated with IgG1. IL23p19 neutralization significantly decreased this enrichment of neutrophils in the esophageal mucosa of IkkβcaEEC-KI mice compared with IkkβcaEEC-KI mice treated with IgG1 (Figure 5J and K). Taken together, this shows that IL23 secretion promotes the activation of STAT3 in esophageal epithelial cells and the subsequent recruitment of neutrophils to the esophagus downstream of epithelial IKKβ activation in vivo.
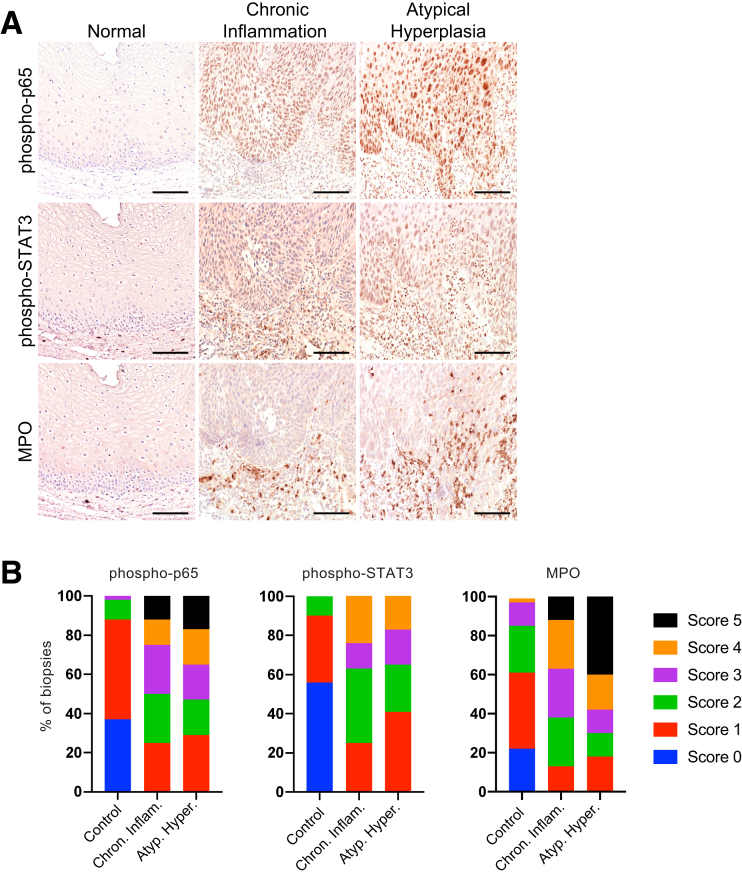
Inflammation is strongly linked to the development of esophageal cancer.6 ESCC develops sequentially, progressing through stages of inflammation, hyperplasia, dysplasia, carcinoma in situ, to invasive carcinoma.5,6 Given that our mice with constitutive activation of IKKβ develop inflammation, hyperplasia, neutrophilia, and have increased epithelial STAT3 signaling, we sought to determine if our findings could be relevant to understanding the development of precursor lesions leading to ESCC, an urgently needed area of research. Using human tissue microarrays, we examined Phosphorylated (P)-p65 NF-κB, p-STAT3, and MPO expression levels in human esophageal biopsy specimens from patients with healthy esophagus, chronic inflammation, and hyperplasia. As shown in Figure 6, P-p65 NF-κB, p-STAT3, and MPO all increased progressively from normal esophagus to chronic inflammation to hyperplasia. Thus, the IKKβ/NF-κB-STAT3–neutrophil axis observed in mice may additionally play an important role in the onset of human disease.
Figure 6.
P-p65 NF-κB, p-STAT3, and MPO expression increases during progression from normal esophagus to chronic inflammation to epithelial hyperplasia in human patients. (A) Representative immunohistochemistry panels from normal esophagus, chronic inflammation, and atypical hyperplasia. Expression of P-p65 NF-κB, p-STAT3, and MPO increased in chronic inflammation and atypical hyperplasia compared with normal esophagus. Scale bar: 100 μm. (B) 100% stacked bar chart indicates the scoring of P-p65 NF-κB, p-STAT3, and MPO levels by pathologic diagnosis for human patients with normal esophagus, chronic inflammation, and hyperplasia, on a scale from 0 to 5. Normal esophagus, n = 41; chronic inflammation, n = 8; hyperplasia, n = 17. Atyp. Hyper., atypical hyperplasia; Chron. Inflam., chronic inflammation.
Discussion
Communication between epithelial cells and immune cells is crucial in the initiation of the inflammatory response and occurs through complex signaling networks. Within the esophagus, epithelial cells sense injury and initiate inflammation through the production of inflammatory molecules that attract and communicate with immune cells. The IKKβ/NF-κB pathway is central to directing the inflammatory response, but the function of this pathway depends greatly on the context, tissue, and cell type studied.29 Recently, we showed that expression of an active form of IKKβ specifically in esophageal epithelia of mice leads to the development of esophagitis and hyperplasia.20 In this study, we show that activation of epithelial IKKβ signaling regulates the recruitment of neutrophils through the activation of epithelial STAT3 signaling in the esophagus.
A functional interplay between IKKβ/NF-κB and STAT3 signaling has been described in numerous inflammatory conditions and cancers.7,10,30,31 Different types of interactions between NF-κB and STAT3 have been reported, including physical association, increased transcription or secretion of activators of the other pathway, overlapping sets of target genes, and competition or cooperation at gene promoters.7,10 Given the complexity of the interaction between these 2 pathways, we were interested in understanding the mechanism after IKKβ activation in the context of esophagitis. We found that in esophageal epithelial cells, STAT3 activity in the presence of IKKβ activation was mostly due to paracrine signaling mediated by the release of secreted factors, specifically through the expression/secretion of IL23.
How does epithelial IKKβ signaling regulate IL23? The epithelial IKKβ/NF-κB pathway promotes immune cell infiltration into the esophagus,18 and the cellular source of IL23 most commonly is recognized to consist of dendritic cells, macrophages, and monocytes.32 Interestingly, our RNA sequencing analyses showed increased mRNA expression levels of the dendritic cell marker Itgax (CD11c) in IkkβcaEEC-KI mice, raising the possibility that activation of epithelial IKKβ signaling could lead to the recruitment of dendritic cells, which subsequently may secrete IL23. Alternatively, recent studies also have reported direct Il23a expression by epithelial cells in multiple contexts.33, 34, 35 The NF-κB subunit RelA associates with the Il23a promoter and is necessary for Il23a transcription,36 suggesting that increased epithelial IKKβ/NF-κB signaling could produce IL23 directly in esophageal epithelial cells. Because of this complex interplay between cell types, precisely how epithelial IKKβ signaling up-regulates esophageal IL23 remains to be determined. Our findings also show that IL23 blockade in IkkβcaEEC-KI mice prevents the activation of STAT3 and reduces subsequent neutrophilia, which may be owing to a similarly complex set of direct and indirect interactions. Because infiltrating immune cells introduce a host of additional cytokines into a tissue, many of which activate STAT3, the epithelial STAT3 activation seen in IkkβcaEEC-KI mice may have an indirect source.37, 38, 39 In colorectal adenocarcinoma, IL23 stimulates T-helper 17 cells to produce cytokines IL6, IL17A, and IL22, which then either directly or indirectly stimulate epithelial cells.39 Because the role for IL23 to stimulate T-helper 17 cells is well established,40, 41, 42 this may be occurring in our system as well. Alternatively, IL23 itself can directly activate JAK2/STAT3 signaling after association with its receptor, IL23R.43 Although the function of IL23R is best established in immune cells, epithelial expression of IL23R also has been observed in the intestine, where it contributes to intestinal homeostasis44; thus, esophageal epithelial signaling through IL23/IL23R interactions also may be possible. Given the complexity of this interplay between IKKβ, IL23, and STAT3, expanding the knowledge of the IL23 signaling cascade in our model is an area of future investigation.
Our study further shows that activation of epithelial STAT3 signaling by IKKβ contributes to neutrophil infiltration into the esophagus. Neutrophils are primary responders to tissue injury or infection and are key potentiators of the inflammatory response.45 After an injury, tissue-produced chemokines promote neutrophil migration from the circulation into damaged tissue, where they stimulate epithelial repair and secrete cytotoxic effectors to eliminate invading pathogens.46 In cases of severe inflammation, critically high numbers of neutrophils can be found adjacent to epithelium that has been ulcerated or eroded.47 The potent neutrophil chemoattractant and activator, C-X-C motif chemokine ligand 5 (CXCL5), is a chemokine produced by epithelial cells in many organs.48,49 In a mouse model of chronic esophageal inflammation, epithelial NF-κB/CXCL5 signaling and neutrophil infiltration both were increased.50 Epithelial STAT3 also has been shown to contribute to maximal CXCL5 induction in the lung, leading to neutrophil recruitment.51 In this study, we show strong induction of Cxcl5 mRNA transcripts in IkkβcaEEC-KI mice, as well as milder induction in IkkβcaEEC-KI;Stat3EEC-KO mice (Figure 4H), indicating that epithelial IKKβ is sufficient to promote CXCL5, with epithelial STAT3 required for maximal production. Further blockade experiments will be needed to show the role of CXCL5 in neutrophil recruitment downstream of IKKβ and STAT3 in our model.
Our study also provides important insights into the development of precursor lesions leading to ESCC, which are characterized by the presence of chronic inflammation and hyperplasia.52 Here, we show that phosphorylated p65 NF-κB, p-STAT3, and the neutrophil marker MPO increase progressively from normal esophagus to esophageal chronic inflammation to hyperplasia in human patients. This supports recent findings from a study reporting substantial neutrophilia and dysregulated NF-κB and JAK/STAT signaling in precursor lesions (chronic inflammation and hyperplasia) of ESCC in mice.52 Interestingly, this increase of neutrophils also was observed along the different stages of ESCC pathogenesis in human patients.52 Moreover, we previously showed an increase in p65 NF-κB signaling and neutrophil recruitment in the early stages of chronic inflammation in a mouse model of ESCC.50 An important limitation to our scoring analyses of patient esophageal biopsy specimens is the limited availability of patient samples with precursor lesions of ESCC. Because most ESCC diagnoses occur at a late disease stage, few patients with early precursor lesions are diagnosed in the clinic, and thus few biopsy specimens are collected.6 High levels of neutrophilic infiltration also can be observed in many other esophageal diseases, including infectious esophagitis caused by candidiasis,53,54 nonerosive reflux disease,55,56 and severe cases of gastroesophageal reflux disease.57, 58, 59 Because our results show that cooperative IKKβ and STAT3 signaling regulates neutrophil trafficking to the esophagus, our study underscores the importance of evaluating the interplay between IKKβ and STAT3 in these different etiologies of esophagitis with tissue neutrophilia.
Another implication of our data is that it provides insights for the development of therapeutics to specifically target esophageal neutrophilia and inflammation. As such, IKKβ and STAT3 theoretically could be therapeutic targets. However, because a careful balance of IKKβ and STAT3 activation is required for the maintenance of normal epithelial and immune homeostasis,60,61 targeting IKKβ or STAT3 might not be ideal. To avoid potential toxicity, it is worth considering targeting specific cytokines/chemokines regulated by IKKβ and/or STAT3. Based on our findings, IL23 could be a good therapeutic target worth exploring for the treatment of chronic esophagitis, hyperplasia, and esophageal neutrophilia. The recent discovery of the importance of IL23 in the pathogenesis of numerous inflammatory diseases, such as psoriasis, inflammatory bowel diseases, multiple sclerosis, and rheumatoid arthritis has led to the emergence of novel biologic agents targeting IL23.62 The results from clinical trials have been impressive, offering promises for the treatment of other diseases in which IL23 plays a central role. In addition, JAK2 also could be a therapeutic target worth exploring because the emergence of a new class of therapeutics targeting JAKs showed promises for therapeutic interventions for rheumatoid arthritis, psoriatic arthritis, and inflammatory bowel disease.60
In sum, we propose a model in which epithelial IKKβ signaling activates epithelial STAT3 through IL23 and other secreted factors, with subsequent infiltration of neutrophils into the esophagus through the epithelial expression of neutrophil chemoattractants (graphical abstract).
Materials and Methods
All authors had access to the study data and reviewed and approved the final manuscript.
Generation of ED-L2/Cre;Rosa26-Ikkβca+/L;Stat3L/L Mice
All animal studies were approved by the Institutional Animal Care and Use Committee at Northwestern University. ED-L2/Cre;Rosa26-Ikkβca+/L;Stat3L/L (IkkβcaEEC-KI;Stat3EEC-KO) mice were generated by crossing Rosa26-STOPFL-Ikkβca mice (Jackson Laboratory, Bar Harbor, ME)22 and Stat3L/L mice (Jackson Laboratory)21 with mice harboring Cre recombinase under the control of the Epstein-Barr Virus (EBV)-ED-L2 promoter.63 An EGFP reporter gene is expressed downstream of the IKKβca construct in Rosa26-STOPFL-Ikkβca mice.22 Four groups of mice were analyzed: Rosa26-STOPFLIkkβcaSFL (control) mice, EBV-ED-L2/Cre;Rosa26-Ikkβca+/L (IkkβcaEEC-KI) mice, EBV-ED-L2/Cre;Rosa26-Ikkβca+/L;Stat3L/L (IkkβcaEEC-KI;Stat3EEC-KO) mice, and EBV-ED-L2/Cre;Stat3L/L (Stat3EEC-KO) mice. All mice used for experiments were maintained on a pure c57BL/6 background, were 6 weeks of age, and were sex-matched. Both male and female experimental groups were used. Mice were examined grossly and processed for histology. Briefly, tissue was fixed in zinc formalin (Sigma Aldrich, St. Louis, MO) for 4 days and embedded in paraffin, and 4-μm sections were applied to charged plus slides. Slides were stained with H&E, and images were captured on a Nikon Eclipse Ci-E microscope with a Nikon DS-Ri2 camera and NIS Elements software (Melville, NY). The following numbers of age-matched mice were examined histologically: 6 controls, 6 IkkβcaEEC-KI mice, 6 IkkβcaEEC-KI;Stat3EEC-KO mice, and 6 Stat3EEC-KO mice.
Immunohistochemistry and Immunofluorescence
For paraffin-embedded esophageal sections, heat antigen retrieval (2100 Antigen Retriever; Electron Microscopy Sciences, Hatfield, PA) was performed and slides were incubated with the following antibodies: 1:50 rabbit anti–phospho-Stat3 (Tyr705) (9145; Cell Signaling, Danvers, MA), 1:50 goat anti-GFP (ab6673; Abcam, Cambridge, MA), and 1:50 rabbit anti-MPO (RB-373-A0; Thermo Fisher Scientific, Pittsburgh, PA). Species-specific secondary antibodies were added, and detection was performed as previously described.64 For fluorescent labeling, Alexa Fluor 488 (A32814; Thermo Fisher Scientific) was used. 4′,6-diamidino-2-phenylindole was used as a nuclear stain.
Western Blot
Mouse esophageal mucosa were harvested in Triton lysis buffer (1% Triton X-100, 50 mmol/L Tris-HCl pH 7.5, 100 mmol/L NaCl, 5 mmol/L EDTA, 40 mmol/L β-glycerophosphate, 5% glycerol, 50 mmol/L NaF) plus protease (Pierce, Rockford, IL) and phosphatase inhibitors (Sigma-Aldrich). The protein concentration was determined using the Pierce BCA protein assay (Thermo Fisher Scientific). Proteins were separated on NuPage 4%–12% Bis-Tris gels (Thermo Fisher Scientific) and transferred onto polyvinylidene difluoride membrane (EMD Millipore, Billerica, MA). After blocking, membranes were incubated overnight at 4°C with the following antibodies: 1:500 rabbit anti–phospho-Jak2 (Tyr1007/1008) (3771; Cell Signaling), 1:2000 rabbit anti-Jak2 (3230; Cell Signaling), 1:4000 rabbit anti–phospho-Stat3 (Tyr705) (9145; Cell Signaling), 1:2000 rabbit anti–phospho-Stat3 (Ser727) (9134; Cell Signaling), 1:4000 rabbit anti-Stat3 (9132; Cell Signaling), 1:6000 rabbit anti-IKKβ (8943; Cell Signaling), 1:10,000 mouse anti–β-actin (A5441; Sigma-Aldrich), and 1:10,000 rabbit anti–glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (G9545; Sigma-Aldrich). Quantitation of bands was performed with FIJI (version 2.0.0).65 The mean gray value was measured for each protein of interest. All values were normalized to loading control (GAPDH or β-actin) or to total protein in the case of phosphorylated proteins. Control bands were adjusted to 1 and fold change was calculated.
RNA Analyses
Mouse esophagi were harvested and the esophageal mucosa was mechanically separated from whole esophagus; tissue fractions were flash-frozen and stored at -80°C. After homogenization, RNA was extracted and purified using the RNeasy kit (Qiagen, Germantown, MD). Reverse transcription was performed with the Maxima First-Strand complementary DNA Synthesis for Reverse-Transcription quantitative PCR kit (Thermo Fisher Scientific). Quantitative real-time PCR was performed using TaqMan Universal Master Mix (Thermo Fisher Scientific). TATA box binding protein or GAPDH genes were used as the internal control. The RT2 Profiler PCR Array Mouse Cytokines and Chemokines (Qiagen) was used to examine Il23a expression in mouse esophageal mucosa.
RNA Sequencing
Esophageal mucosa were harvested from 6-week-old mice. RNA was extracted and purified using the RNeasy Mini kit (Qiagen). Complementary DNA libraries were generated using the Tru-Seq Stranded mRNA sequencing library prep (Illumina, San Diego, CA) following the manufacturer’s instructions. Sequencing was performed using Illumina HiSeq 4000 (Northwestern University NuSeq Core Facility). The quality of DNA reads, in fastq format, was evaluated using FastQC.66 Adapters were trimmed and reads of poor quality or aligning to ribosomal RNA sequences were filtered. The cleaned reads were aligned to the Mus musculus genome (mm10) using STAR.67 Read counts for each gene were calculated using htseq-count68 in conjunction with a gene annotation file for mm10 obtained from the University of California Santa Cruz (http://genome.ucsc.edu). Normalization and differential expression were determined using DESeq2.69 The cut-off value for determining significantly differentially expressed genes was a false-discovery rate–adjusted P value of less than .05. RNA sequencing was deposited in Gene Expression Omnibus (GSE154129) and can be accessed at http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE154129. Gene ontology terms were identified using the Metascape online platform (http://metascape.org).70 KEGG terms were identified using the gseKEGG function within the clusterProfiler R package.71
Flow Cytometry
For flow cytometric analysis of in vivo experiments, mice were killed at 6 weeks of age and spleen, blood, and esophagus were harvested. Forceps were used to separate esophageal mucosa from whole esophageal tissue, and each tissue fraction was minced. Esophageal mucosa was digested in 0.05% trypsin-EDTA (Thermo Fisher Scientific) for 2 minutes, with shaking at 37°C before neutralization with fetal bovine serum (Corning) and filtration through 70-μm nylon filters (Fisher Scientific, Hampton, NH). The remaining esophageal tissue was digested in collagenase V (Sigma-Aldrich) for 15 minutes, with shaking at 37°C before filtration through 70-μm nylon filters. Blood was obtained via cardiac puncture and red blood cells were lysed with ACK lysis buffer (Thermo Fisher Scientific). Whole spleen was harvested and filtered through 70-μm nylon filters, after which red blood cells were lysed with ACK lysis buffer (Thermo Fisher Scientific). Spleen suspensions were filtered again through 70-um nylon filters. Up to 1 million cells were stained per sample. Live/dead cell discrimination was performed using eBioscience (Thermo Fisher Scientific) Fixable Viability Dye eFluor 506 (Thermo Fisher Scientific) according to the manufacturer’s instructions. Cell surface staining for the indicated antibodies was performed for 30 minutes. Intracellular staining was performed using the FoxP3 fixation/permeabilization kit (BioLegend, San Diego, CA) according to the manufacturer’s instructions. T cells were phenotyped as CD45+CD11b–CD3+, T-helper cells as CD45+CD11b–CD3+CD4+CD8–, regulatory T cells as CD45+CD11b–CD3+CD4+CD8–Foxp3+, cytotoxic T lymphocytes as CD45+CD11b–CD3+CD8+CD4–, B cells as CD45+CD19+CD3–, macrophages as CD45+CD19-CD3–F4/80+CD11b+, dendritic cells as CD45+CD19-CD3–CD11c+, neutrophils as CD45+CD19-CD3–CD11b+Ly6Ghi, natural killer cells as CD45+CD19-CD3–CD49b+NK1.1+, eosinophils as CD45+CD19-CD3–SiglecF+, and monocytes as CD45+CD19-CD3–CD115+. All flow cytometric analyses were performed using a BD FACSymphony A5 (BD Biosciences, San Jose, CA) and analyzed using FlowJo software (BD Biosciences). Antibodies for flow cytometry are provided in Table 1.
Table 1.
Antibodies for Flow Cytometry
| Antibody | Clone | Fluorophore | Company | Catalog number |
|---|---|---|---|---|
| CD115 | T38-320 | BV421 | BD Biosciences | 743638 |
| CD117 | 2B8 | BV711 | BD Biosciences | 563160 |
| CD11b | M1/70 | BUV737 | BD Biosciences | 564443 |
| CD11c | HL3 | BUV395 | BD Biosciences | 564080 |
| CD170 | E50-2440 | PE-CF594 | BD Biosciences | 562757 |
| CD19 | 1D3 | APC-Cy7 | BD Biosciences | 557655 |
| CD31 | 390 | AF488 | BD Biosciences | 563607 |
| CD3e | 500A2 | AF700 | BD Biosciences | 557984 |
| CD4 | GK1.5 | BUV496 | BD Biosciences | 564667 |
| CD44 | 1M7 | BV786 | BD Biosciences | 563736 |
| CD45.2 | 104 | BV650 | BD Biosciences | 740490 |
| CD49b | Hma2 | BV605 | BD Biosciences | 740363 |
| CD8a | 53-6.7 | BB700 | BD Biosciences | 566409 |
| F4/80 | T45-2342 | A647 | BD Biosciences | 565854 |
| FoxP3 | FJK-16s | PE | eBioscience | 12-5773-80 |
| Live/Dead | N/A | 506e | eBioscience | 65-0866-14 |
| Ly6G | 1A8 | PE-Cy7 | BD Biosciences | 560601 |
| NK 1.1 | PK136 | BB630 | BD Biosciences | 624294 |
In Vivo Anti-IL23p19 and Anti-IL12p40 Treatment
Starting at 4 weeks of age, IkkβcaEEC-KI mice and littermate controls were administered intraperitoneal injections of IL23p19 neutralizing antibody G23-8 or rat IgG1 isotype control at doses of 100 μg per dose, or of IL12p40 neutralizing antibody C17.8 or rat IgG2a at doses of 200 μg per dose (BioXCell, West Hanover, NH). Mice were treated every Monday and Thursday until they reached 6 weeks of age. The following numbers of mice were injected with neutralizing antibody or control: IgG1, 8 pairs; G23-8, 8 pairs; IgG2a, 3 pairs; and C17.8, 3 pairs.
Tissue Microarray
Tissue microarrays from human biopsy specimens of patients with chronic inflammation (n = 8), hyperplasia (n = 17), and normal esophagus (n = 41) (US Biomax, Inc, Rockville, MD) were stained with P-p65 NF-κB, p-STAT3, and MPO. Staining intensity was scored in a blinded fashion on a scale from 0 to 5 (0, none; 1, weak; 2, mild; 3, moderate; 4, intense; and 5, very intense).
Statistical Analyses
Results are expressed as means ± SD, with statistical significance of differences performed on normalized values between experimental conditions established at 95%. The Student t test was used to indicate statistical difference between pairwise comparisons. One-way analysis of variance was used to analyze the statistical difference between groups followed by the Tukey post hoc test, and the Sidak multiple comparisons test was used to compare preselected groups of means. All statistics were performed using GraphPad Prism version 8.4.1 (GraphPad Software, San Diego, CA).
CRediT Authorship Contributions
Kelsey Nicole Wiles (Conceptualization: Equal; Formal analysis: Lead; Methodology: Lead; Validation: Lead; Writing – original draft: Lead; Writing – review & editing: Supporting)
Cara Maria Alioto (Formal analysis: Supporting; Methodology: Supporting; Validation: Supporting; Writing – review & editing: Supporting)
Nathan Bruce Hodge (Formal analysis: Supporting; Methodology: Supporting; Validation: Supporting; Writing – review & editing: Supporting)
Margarette Helen Clevenger (Formal analysis: Supporting; Methodology: Supporting; Validation: Supporting; Writing – review & editing: Supporting)
Lia Elyse Tsikretsis (Methodology: Supporting)
Frederick T.J. Lin (Formal analysis: Supporting; Methodology: Supporting)
Marie-Pier Tetreault, PhD (Conceptualization: Lead; Formal analysis: Supporting; Methodology: Supporting; Supervision: Lead; Validation: Supporting; Writing – original draft: Supporting; Writing – review & editing: Lead)
Footnotes
Conflicts of interest The authors disclose no conflicts.
Funding This work was supported by National Institutes of HealthNational Institute of Diabetes and Digestive and Kidney Diseases R00 DK094977 and R01 DK116988 (M.P.T.); National Institutes of Health National Institute of General Medical Sciences T32 GM008061 (K.N.W. and N.B.H.); National Institutes of HealthNational Heart, Lung, and Blood Institute F31 HL147413 (M.H.C.); by the Robert H. Lurie Comprehensive Cancer Center (National Institutes of Health National Cancer Institute CCSG P30 CA060553) through the Northwestern University Mouse Histology and Phenotyping Core, the Flow Cytometry Facility, and by the NUSeq Core facility.
Supplementary Material
References
- 1.Peery A.F., Crockett S.D., Murphy C.C., Lund J.L., Dellon E.S., Williams J.L., Jensen E.T., Shaheen N.J., Barritt A.S., Lieber S.R., Kochar B., Barnes E.L., Fan Y.C., Pate V., Galanko J., Baron T.H., Sandler R.S. Burden and cost of gastrointestinal, liver, and pancreatic diseases in the United States: update 2018. Gastroenterology. 2019;156:254–272.e11. doi: 10.1053/j.gastro.2018.08.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bellizzi A.M., Nardone G., Compare D., Bor S., Capanoglu D., Farré R., Neumann H., Neurath M.F., Vieth M., Chen H., Chen X. Tissue resistance in the normal and diseased esophagus. Ann N Y Acad Sci. 2013;1300:200–212. doi: 10.1111/nyas.12248. [DOI] [PubMed] [Google Scholar]
- 3.Dunbar K.B., Agoston A.T., Odze R.D., Huo X., Pham T.H., Cipher D.J., Castell D.O., Genta R.M., Souza R.F., Spechler S.J. Association of acute gastroesophageal reflux disease with esophageal histologic changes. JAMA. 2016;315:2104–2112. doi: 10.1001/jama.2016.5657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lee H.J., Park J.M., Han Y.M., Gil H.K., Kim J., Chang J.Y., Jeong M., Go E.J., Hahm K.B. The role of chronic inflammation in the development of gastrointestinal cancers: reviewing cancer prevention with natural anti-inflammatory intervention. Expert Rev Gastroenterol Hepatol. 2016;10:129–139. doi: 10.1586/17474124.2016.1103179. [DOI] [PubMed] [Google Scholar]
- 5.Chen X.X., Zhong Q., Liu Y., Yan S.M., Chen Z.H., Jin S.Z., Xia T.L., Li R.Y., Zhou A.J., Su Z., Huang Y.H., Huang Q.T., Huang L.Y., Zhang X., Zhao Y.N., Yun J.P., Wu Q.L., Lin D.X., Bai F., Zeng M.S. Genomic comparison of esophageal squamous cell carcinoma and its precursor lesions by multi-region whole-exome sequencing. Nat Commun. 2017;8:524. doi: 10.1038/s41467-017-00650-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Rustgi A.K., El-Serag H.B. Esophageal carcinoma. N Engl J Med. 2014;371:2499–2509. doi: 10.1056/NEJMra1314530. [DOI] [PubMed] [Google Scholar]
- 7.Bollrath J., Greten F.R. IKK/NF-kappaB and STAT3 pathways: central signalling hubs in inflammation-mediated tumour promotion and metastasis. EMBO Rep. 2009;10:1314–1319. doi: 10.1038/embor.2009.243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Israël A. The IKK complex, a central regulator of NF-kappaB activation. Cold Spring Harb Perspect Biol. 2010;2:a000158. doi: 10.1101/cshperspect.a000158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hoesel B., Schmid J.A. The complexity of NF-κB signaling in inflammation and cancer. Mol Cancer. 2013;12:86. doi: 10.1186/1476-4598-12-86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Grivennikov S.I., Karin M. Dangerous liaisons: STAT3 and NF-kappaB collaboration and crosstalk in cancer. Cytokine Growth Factor Rev. 2010;21:11–19. doi: 10.1016/j.cytogfr.2009.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hirano T., Ishihara K., Hibi M. Roles of STAT3 in mediating the cell growth, differentiation and survival signals relayed through the IL-6 family of cytokine receptors. Oncogene. 2000;19:2548–2556. doi: 10.1038/sj.onc.1203551. [DOI] [PubMed] [Google Scholar]
- 12.Levy D.E., Darnell J.E., Jr. Stats: transcriptional control and biological impact. Nat Rev Mol Cell Biol. 2002;3:651–662. doi: 10.1038/nrm909. [DOI] [PubMed] [Google Scholar]
- 13.Yu H., Pardoll D., Jove R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer. 2009;9:798–809. doi: 10.1038/nrc2734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gao S.P., Mark K.G., Leslie K., Pao W., Motoi N., Gerald W.L., Travis W.D., Bornmann W., Veach D., Clarkson B., Bromberg J.F. Mutations in the EGFR kinase domain mediate STAT3 activation via IL-6 production in human lung adenocarcinomas. J Clin Invest. 2007;117:3846–3856. doi: 10.1172/JCI31871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Grivennikov S., Karin M. Autocrine IL-6 signaling: a key event in tumorigenesis? Cancer Cell. 2008;13:7–9. doi: 10.1016/j.ccr.2007.12.020. [DOI] [PubMed] [Google Scholar]
- 16.Eckmann L., Nebelsiek T., Fingerle A.A., Dann S.M., Mages J., Lang R., Robine S., Kagnoff M.F., Schmid R.M., Karin M., Arkan M.C., Greten F.R. Opposing functions of IKKbeta during acute and chronic intestinal inflammation. Proc Natl Acad Sci U S A. 2008;105:15058–15063. doi: 10.1073/pnas.0808216105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee H., Herrmann A., Deng J.H., Kujawski M., Niu G., Li Z., Forman S., Jove R., Pardoll D.M., Yu H. Persistently activated Stat3 maintains constitutive NF-kappaB activity in tumors. Cancer Cell. 2009;15:283–293. doi: 10.1016/j.ccr.2009.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yang J., Liao X., Agarwal M.K., Barnes L., Auron P.E., Stark G.R. Unphosphorylated STAT3 accumulates in response to IL-6 and activates transcription by binding to NFkappaB. Genes Dev. 2007;21:1396–1408. doi: 10.1101/gad.1553707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yu Z., Zhang W., Kone B.C. Signal transducers and activators of transcription 3 (STAT3) inhibits transcription of the inducible nitric oxide synthase gene by interacting with nuclear factor kappaB. Biochem J. 2002;367:97–105. doi: 10.1042/BJ20020588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tétreault M.P., Weinblatt D., Ciolino J.D., Klein-Szanto A.J., Sackey B.K., Twyman-Saint Victor C., Karakasheva T., Teal V., Katz J.P. Esophageal expression of active IκB kinase-β in mice up-regulates tumor necrosis factor and granulocyte-macrophage colony-stimulating factor, promoting inflammation and angiogenesis. Gastroenterology. 2016;150:1609–1619.e11. doi: 10.1053/j.gastro.2016.02.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Moh A., Iwamoto Y., Chai G.X., Zhang S.S., Kano A., Yang D.D., Zhang W., Wang J., Jacoby J.J., Gao B., Flavell R.A., Fu X.Y. Role of STAT3 in liver regeneration: survival, DNA synthesis, inflammatory reaction and liver mass recovery. Lab Invest. 2007;87:1018–1028. doi: 10.1038/labinvest.3700630. [DOI] [PubMed] [Google Scholar]
- 22.Sasaki Y., Derudder E., Hobeika E., Pelanda R., Reth M., Rajewsky K., Schmidt-Supprian M. Canonical NF-kappaB activity, dispensable for B cell development, replaces BAFF-receptor signals and promotes B cell proliferation upon activation. Immunity. 2006;24:729–739. doi: 10.1016/j.immuni.2006.04.005. [DOI] [PubMed] [Google Scholar]
- 23.Moschen A.R., Adolph T.E., Gerner R.R., Wieser V., Tilg H. Lipocalin-2: a master mediator of intestinal and metabolic inflammation. Trends Endocrinol Metab. 2017;28:388–397. doi: 10.1016/j.tem.2017.01.003. [DOI] [PubMed] [Google Scholar]
- 24.Gross S.R., Sin C.G., Barraclough R., Rudland P.S. Joining S100 proteins and migration: for better or for worse, in sickness and in health. Cell Mol Life Sci. 2014;71:1551–1579. doi: 10.1007/s00018-013-1400-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Verbeke H., Geboes K., Van Damme J., Struyf S. The role of CXC chemokines in the transition of chronic inflammation to esophageal and gastric cancer. Biochim Biophys Acta. 2012;1825:117–129. doi: 10.1016/j.bbcan.2011.10.008. [DOI] [PubMed] [Google Scholar]
- 26.Floss D.M., Mrotzek S., Klöcker T., Schröder J., Grötzinger J., Rose-John S., Scheller J. Identification of canonical tyrosine-dependent and non-canonical tyrosine-independent STAT3 activation sites in the intracellular domain of the interleukin 23 receptor. J Biol Chem. 2013;288:19386–19400. doi: 10.1074/jbc.M112.432153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pastor-Fernández G., Mariblanca I.R., Navarro M.N. Decoding IL-23 signaling cascade for new therapeutic opportunities. Cells. 2020;9:2044. doi: 10.3390/cells9092044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Vignali D.A., Kuchroo V.K. IL-12 family cytokines: immunological playmakers. Nat Immunol. 2012;13:722–728. doi: 10.1038/ni.2366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Taniguchi K., Karin M. NF-κB, inflammation, immunity and cancer: coming of age. Nat Rev Immunol. 2018;18:309–324. doi: 10.1038/nri.2017.142. [DOI] [PubMed] [Google Scholar]
- 30.Grivennikov S.I., Karin M. Inflammatory cytokines in cancer: tumour necrosis factor and interleukin 6 take the stage. Ann Rheum Dis. 2011;70(Suppl 1):i104–i108. doi: 10.1136/ard.2010.140145. [DOI] [PubMed] [Google Scholar]
- 31.Bode J.G., Albrecht U., Häussinger D., Heinrich P.C., Schaper F. Hepatic acute phase proteins--regulation by IL-6- and IL-1-type cytokines involving STAT3 and its crosstalk with NF-κB-dependent signaling. Eur J Cell Biol. 2012;91:496–505. doi: 10.1016/j.ejcb.2011.09.008. [DOI] [PubMed] [Google Scholar]
- 32.Oppmann B., Lesley R., Blom B., Timans J.C., Xu Y., Hunte B., Vega F., Yu N., Wang J., Singh K., Zonin F., Vaisberg E., Churakova T., Liu M., Gorman D., Wagner J., Zurawski S., Liu Y., Abrams J.S., Moore K.W., Rennick D., de Waal-Malefyt R., Hannum C., Bazan J.F., Kastelein R.A. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity. 2000;13:715–725. doi: 10.1016/s1074-7613(00)00070-4. [DOI] [PubMed] [Google Scholar]
- 33.Lim K.S., Yong Z.W.E., Wang H., Tan T.Z., Huang R.Y., Yamamoto D., Inaki N., Hazawa M., Wong R.W., Oshima H., Oshima M., Ito Y., Voon D.C. Inflammatory and mitogenic signals drive interleukin 23 subunit alpha (IL23A) secretion independent of IL12B in intestinal epithelial cells. J Biol Chem. 2020;295:6387–6400. doi: 10.1074/jbc.RA120.012943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kopp T., Lenz P., Bello-Fernandez C., Kastelein R.A., Kupper T.S., Stingl G. IL-23 production by cosecretion of endogenous p19 and transgenic p40 in keratin 14/p40 transgenic mice: evidence for enhanced cutaneous immunity. J Immunol. 2003;170:5438–5444. doi: 10.4049/jimmunol.170.11.5438. [DOI] [PubMed] [Google Scholar]
- 35.Hor Y.T., Voon D.C., Koo J.K., Wang H., Lau W.M., Ashktorab H., Chan S.L., Ito Y. A role for RUNX3 in inflammation-induced expression of IL23A in gastric epithelial cells. Cell Rep. 2014;8:50–58. doi: 10.1016/j.celrep.2014.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Mise-Omata S., Kuroda E., Niikura J., Yamashita U., Obata Y., Doi T.S. A proximal kappaB site in the IL-23 p19 promoter is responsible for RelA- and c-Rel-dependent transcription. J Immunol. 2007;179:6596–6603. doi: 10.4049/jimmunol.179.10.6596. [DOI] [PubMed] [Google Scholar]
- 37.Liao Y., Hu X., Guo X., Zhang B., Xu W., Jiang H. Promoting effects of IL-23 on myocardial ischemia and reperfusion are associated with increased expression of IL-17A and upregulation of the JAK2-STAT3 signaling pathway. Mol Med Rep. 2017;16:9309–9316. doi: 10.3892/mmr.2017.7771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tang Q., Li J., Zhu H., Li P., Zou Z., Xiao Y. Hmgb1-IL-23-IL-17-IL-6-Stat3 axis promotes tumor growth in murine models of melanoma. Mediators Inflamm. 2013;2013:713859. doi: 10.1155/2013/713859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Grivennikov S.I., Wang K., Mucida D., Stewart C.A., Schnabl B., Jauch D., Taniguchi K., Yu G.Y., Osterreicher C.H., Hung K.E., Datz C., Feng Y., Fearon E.R., Oukka M., Tessarollo L., Coppola V., Yarovinsky F., Cheroutre H., Eckmann L., Trinchieri G., Karin M. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature. 2012;491:254–258. doi: 10.1038/nature11465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Aggarwal S., Ghilardi N., Xie M.H., de Sauvage F.J., Gurney A.L. Interleukin-23 promotes a distinct CD4 T cell activation state characterized by the production of interleukin-17. J Biol Chem. 2003;278:1910–1914. doi: 10.1074/jbc.M207577200. [DOI] [PubMed] [Google Scholar]
- 41.Langrish C.L., Chen Y., Blumenschein W.M., Mattson J., Basham B., Sedgwick J.D., McClanahan T., Kastelein R.A., Cua D.J. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. J Exp Med. 2005;201:233–240. doi: 10.1084/jem.20041257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Weaver C.T., Hatton R.D., Mangan P.R., Harrington L.E. IL-17 family cytokines and the expanding diversity of effector T cell lineages. Annu Rev Immunol. 2007;25:821–852. doi: 10.1146/annurev.immunol.25.022106.141557. [DOI] [PubMed] [Google Scholar]
- 43.Parham C., Chirica M., Timans J., Vaisberg E., Travis M., Cheung J., Pflanz S., Zhang R., Singh K.P., Vega F., To W., Wagner J., O'Farrell A.M., McClanahan T., Zurawski S., Hannum C., Gorman D., Rennick D.M., Kastelein R.A., de Waal Malefyt R., Moore K.W. A receptor for the heterodimeric cytokine IL-23 is composed of IL-12Rbeta1 and a novel cytokine receptor subunit, IL-23R. J Immunol. 2002;168:5699–5708. doi: 10.4049/jimmunol.168.11.5699. [DOI] [PubMed] [Google Scholar]
- 44.Aden K., Rehman A., Falk-Paulsen M., Secher T., Kuiper J., Tran F., Pfeuffer S., Sheibani-Tezerji R., Breuer A., Luzius A., Jentzsch M., Häsler R., Billmann-Born S., Will O., Lipinski S., Bharti R., Adolph T., Iovanna J.L., Kempster S.L., Blumberg R.S., Schreiber S., Becher B., Chamaillard M., Kaser A., Rosenstiel P. Epithelial IL-23R signaling licenses protective IL-22 responses in intestinal inflammation. Cell Rep. 2016;16:2208–2218. doi: 10.1016/j.celrep.2016.07.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Nathan C. Neutrophils and immunity: challenges and opportunities. Nat Rev Immunol. 2006;6:173–182. doi: 10.1038/nri1785. [DOI] [PubMed] [Google Scholar]
- 46.Kolaczkowska E., Kubes P. Neutrophil recruitment and function in health and inflammation. Nat Rev Immunol. 2013;13:159–175. doi: 10.1038/nri3399. [DOI] [PubMed] [Google Scholar]
- 47.Brazil J.C., Quiros M., Nusrat A., Parkos C.A. Innate immune cell-epithelial crosstalk during wound repair. J Clin Invest. 2019;129:2983–2993. doi: 10.1172/JCI124618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Walz A., Schmutz P., Mueller C., Schnyder-Candrian S. Regulation and function of the CXC chemokine ENA-78 in monocytes and its role in disease. J Leukoc Biol. 1997;62:604–611. doi: 10.1002/jlb.62.5.604. [DOI] [PubMed] [Google Scholar]
- 49.Rajarathnam K., Schnoor M., Richardson R.M., Rajagopal S. How do chemokines navigate neutrophils to the target site: dissecting the structural mechanisms and signaling pathways. Cell Signal. 2019;54:69–80. doi: 10.1016/j.cellsig.2018.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Tetreault M.P., Wang M.L., Yang Y., Travis J., Yu Q.C., Klein-Szanto A.J., Katz J.P. Klf4 overexpression activates epithelial cytokines and inflammation-mediated esophageal squamous cell cancer in mice. Gastroenterology. 2010;139:2124–2134.e9. doi: 10.1053/j.gastro.2010.08.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Traber K.E., Hilliard K.L., Allen E., Wasserman G.A., Yamamoto K., Jones M.R., Mizgerd J.P., Quinton L.J. Induction of STAT3-dependent CXCL5 expression and neutrophil recruitment by Oncostatin-M during pneumonia. Am J Respir Cell Mol Biol. 2015;53:479–488. doi: 10.1165/rcmb.2014-0342OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yao J., Cui Q., Fan W., Ma Y., Chen Y., Liu T., Zhang X., Xi Y., Wang C., Peng L., Luo Y., Lin A., Guo W., Lin L., Lin Y., Tan W., Lin D., Wu C., Wang J. Single-cell transcriptomic analysis in a mouse model deciphers cell transition states in the multistep development of esophageal cancer. Nat Commun. 2020;11:3715. doi: 10.1038/s41467-020-17492-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Alsomali M.I., Arnold M.A., Frankel W.L., Graham R.P., Hart P.A., Lam-Himlin D.M., Naini B.V., Voltaggio L., Arnold C.A. Challenges to "classic" esophageal candidiasis: looks are usually deceiving. Am J Clin Pathol. 2017;147:33–42. doi: 10.1093/ajcp/aqw210. [DOI] [PubMed] [Google Scholar]
- 54.Martin I.W., Atkinson A.E., Liu X., Suriawinata A.A., Lefferts J.A., Lisovsky M. Mucosal inflammation in Candida esophagitis has distinctive features that may be helpful diagnostically. Mod Pathol. 2018;31:1653–1660. doi: 10.1038/s41379-018-0060-4. [DOI] [PubMed] [Google Scholar]
- 55.Yoshida N. Inflammation and oxidative stress in gastroesophageal reflux disease. J Clin Biochem Nutr. 2007;40:13–23. doi: 10.3164/jcbn.40.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Nwokediuko S.C., Ijoma U., Okafor O. Esophageal intraepithelial neutrophil infiltration is common in Nigerian patients with non-erosive reflux disease. Gastroenterology Res. 2011;4:20–25. doi: 10.4021/gr284e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zentilin P., Savarino V., Mastracci L., Spaggiari P., Dulbecco P., Ceppa P., Savarino E., Parodi A., Mansi C., Fiocca R. Reassessment of the diagnostic value of histology in patients with GERD, using multiple biopsy sites and an appropriate control group. Am J Gastroenterol. 2005;100:2299–2306. doi: 10.1111/j.1572-0241.2005.50209.x. [DOI] [PubMed] [Google Scholar]
- 58.Frierson H.F., Jr. Histological criteria for the diagnosis of reflux esophagitis. Pathol Annu. 1992;27:87–104. [PubMed] [Google Scholar]
- 59.Collins B.J., Elliott H., Sloan J.M., McFarland R.J., Love A.H. Oesophageal histology in reflux oesophagitis. J Clin Pathol. 1985;38:1265–1272. doi: 10.1136/jcp.38.11.1265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.O'Shea J.J., Schwartz D.M., Villarino A.V., Gadina M., McInnes I.B., Laurence A. The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu Rev Med. 2015;66:311–328. doi: 10.1146/annurev-med-051113-024537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Egan L.J., Toruner M. NF-kappaB signaling: pros and cons of altering NF-kappaB as a therapeutic approach. Ann N Y Acad Sci. 2006;1072:114–122. doi: 10.1196/annals.1326.009. [DOI] [PubMed] [Google Scholar]
- 62.Frieder J., Kivelevitch D., Haugh I., Watson I., Menter A. Anti-IL-23 and anti-IL-17 biologic agents for the treatment of immune-mediated inflammatory conditions. Clin Pharmacol Ther. 2018;103:88–101. doi: 10.1002/cpt.893. [DOI] [PubMed] [Google Scholar]
- 63.Tetreault M.P., Yang Y., Travis J., Yu Q.C., Klein-Szanto A., Tobias J.W., Katz J.P. Esophageal squamous cell dysplasia and delayed differentiation with deletion of krüppel-like factor 4 in murine esophagus. Gastroenterology. 2010;139:171–181.e9. doi: 10.1053/j.gastro.2010.03.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Yang Y., Goldstein B.G., Nakagawa H., Katz J.P. Krüppel-like factor 5 activates MEK/ERK signaling via EGFR in primary squamous epithelial cells. FASEB J. 2007;21:543–550. doi: 10.1096/fj.06-6694com. [DOI] [PubMed] [Google Scholar]
- 65.Schindelin J, Arganda-Carreras I, Frise E. Fiji: an open-source platform for biological-image analysis. Nature Methods. 2012;9:676–682. doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.FastQC: A Quality Control Tool for High Throughput Sequence Data 2010.
- 67.Dobin A., Davis C.A., Schlesinger F., Drenkow J., Zaleski C., Jha S., Batut P., Chaisson M., Gingeras T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Anders S., Pyl P.T., Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–169. doi: 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Zhou Y., Zhou B., Pache L., Chang M., Khodabakhshi A.H., Tanaseichuk O., Benner C., Chanda S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523. doi: 10.1038/s41467-019-09234-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Yu G., Wang L.G., Han Y., He Q.Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics. 2012;16:284–287. doi: 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.