ABSTRACT
SARS-CoV-2 has had a disproportionate impact on nonhospital health care settings, such as long-term-care facilities (LTCFs). The communal nature of these facilities, paired with the high-risk profile of residents, has resulted in thousands of infections and deaths and a high case fatality rate. To detect presymptomatic infections and identify infected workers, we performed weekly surveillance testing of staff at two LTCFs, which revealed a large outbreak at one of the sites. We collected serum from staff members throughout the study and evaluated it for binding and neutralization to measure seroprevalence, seroconversion, and type and functionality of antibodies. At the site with very few incident infections, we detected that over 40% of the staff had preexisting SARS-CoV-2 neutralizing antibodies, suggesting prior exposure. At the outbreak site, we saw rapid seroconversion following infection. Neutralizing antibody levels were stable for many weeks following infection, suggesting a durable, long-lived response. Receptor-binding domain antibodies and neutralizing antibodies were strongly correlated. The site with high seroprevalence among staff had two unique introductions of SARS-CoV-2 into the facility through seronegative infected staff during the period of study, but these did not result in workplace spread or outbreaks. Together, our results suggest that a high seroprevalence rate among staff can contribute to immunity within a workplace and protect against subsequent infection and spread within a facility.
IMPORTANCE Long-term care facilities (LTCFs) have been disproportionately impacted by COVID-19 due to their communal nature and high-risk profile of residents. LTCF staff have the ability to introduce SARS-CoV-2 into the facility, where it can spread, causing outbreaks. We tested staff weekly at two LTCFs and collected blood throughout the study to measure SARS-CoV-2 antibodies. One site had a large outbreak and infected individuals rapidly generated antibodies after infection. At the other site, almost half the staff already had antibodies, suggesting prior infection. The majority of these antibodies bind to the receptor-binding domain of the SARS-CoV-2 spike protein and are potently neutralizing and stable for many months. The non-outbreak site had two unique introductions of SARS-CoV-2 into the facility, but these did not result in workplace spread or outbreaks. Our results reveal that high seroprevalence among staff can contribute to immunity and protect against subsequent infection and spread within a facility.
KEYWORDS: adaptive immunity, coronavirus, neutralizing antibodies, surveillance studies
INTRODUCTION
The emergence of SARS-CoV-2 and the resultant COVID-19 pandemic have threatened health care systems across the world (1, 2). Long-term care facilities (LTCFs) are a significant venue for SARS-CoV-2 transmission and outbreaks, and LTCF resident deaths account for almost half of all U.S. COVID-19 deaths to date (3, 4). This is due to many factors, including the communal nature of LTCFs and the high-risk health profile of residents (5, 6). LTCF staff have the potential to introduce the virus into the facilities, where it can spread among staff, residents, and be exported back into the community. Additionally, staff at these facilities tend to resist vaccination (7–10). We therefore began weekly SARS-CoV-2 surveillance testing of staff at LTCFs and observed significant facility-associated outbreaks (11). In parallel with surveillance testing, we collected blood to determine seroprevalence, monitor seroconversion, and characterize antibody responses in these populations.
Generation of specific, neutralizing and long-lived antibodies is a key component of adaptive immunity. Studies conducted after the SARS and MERS epidemics of 2003 and 2012, respectively, revealed that the majority of recovered individuals generated antibodies; however, it is unclear whether this immunity was sufficient to provide protection against reinfection (12, 13). Many studies have sought to define the antibody response following SARS-CoV-2 infection (14, 15). These include studies on hospitalized COVID-19 patients (16–19), asymptomatic individuals (20, 21), and retrospective serological studies (22–25). The vast majority of infected individuals seroconvert and generate IgA, IgM, and IgG-specific antibodies within 3 weeks of infection (15). Age, sex, hospitalization, severity of infection, and other factors have all been shown to modulate the level, kinetics, and durability of the antibody response following infection (21, 26–32). Recent work has revealed that up to 7 months after infection, absolute binding antibody levels might decline but neutralizing antibodies appear to be long lived and persist at stable levels (33–39).
We therefore sought to characterize the antibody responses to SARS-CoV-2 in staff at two LTCFs by sampling serum at regular time intervals, both during and post outbreak. Using these samples, we measured antibody binding to two commonly used SARS-CoV-2 antigens, full-length spike and receptor-binding domain (RBD), along with neutralization of live SARS-CoV-2 virus. Our data clearly demonstrate the development of SARS-CoV-2 binding and neutralizing antibodies at approximately 1 to 2 weeks postinfection, during the period of observation for the outbreak facility. Our data also reveal that the facility with high seroprevalence did not have any outbreaks during the study period, despite the introduction of the virus into the facility on two independent occasions. These results suggest that high seroprevalence (>40%) and levels of neutralizing antibodies can contribute to outbreak resistance through community immunity. Additionally, we find that up to 4 months postinfection, neutralizing antibody levels are stable and durable.
RESULTS
SARS-CoV-2 surveillance testing.
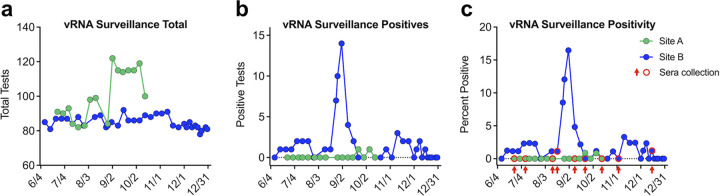
We performed nasal surveillance testing for SARS-CoV-2 viral RNA of staff at two long-term-care facilities over a 4 to 6 month period (Site A, July to Oct 2020; Site B, June to Dec, 2020; Fig. 1). Samples were collected at the workplace. Site A previously experienced a large outbreak in June immediately before our surveillance testing began, with 26 staff and 47 residents testing positive; however, this was before robust surveillance testing efforts were in place, so it is possible far more individuals were infected, whereas at Site B no symptomatic or asymptomatic cases had been diagnosed prior to our surveillance testing. Staff were tested at least once per week, approximately 180 unique individuals at each site participated in testing, with an average of 100 staff at site A and 85 staff at site B testing weekly (Fig. 1a). Positive tests and percent positivity varied by facility, with site A only experiencing two positive tests (from two different staff members) throughout their entire 17-week testing period (Fig. 1b and c). Site B experienced a large outbreak with over 15% of staff testing positive at its peak, and 34 unique staff testing positive throughout the 18-week study (Fig. 1b and c). We collected serum samples from staff at both sites every 3 to 5 weeks, spanning the 5-month surveillance period, including a time point immediately prior to and immediately following an outbreak in early September at site B (Fig. 1c).
FIG 1.
SARS-CoV-2 vRNA surveillance testing at two LTCFs. (a) Total number of staff tested weekly as part of vRNA nasal surveillance testing. (b) Number of positive vRNA tests recorded each week at sites A and B. (c) vRNA positivity expressed as percent positive at each site. Timing of sera collections relative to surveillance testing are indicated by red circles and arrows.
SARS-CoV-2 antibody binding and specificity.
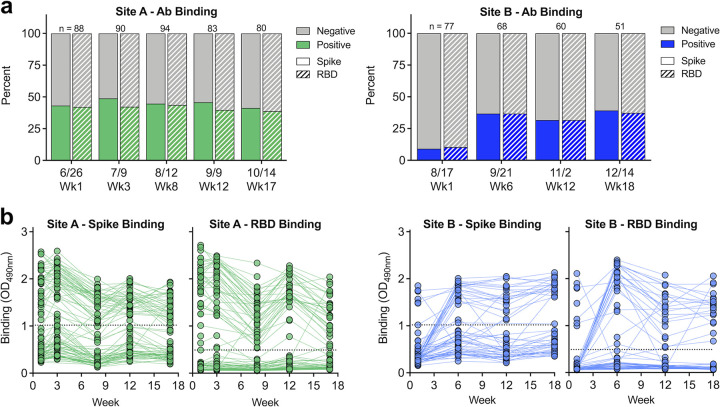
Sera from staff at both sites were evaluated for the presence and levels of polyclonal antibodies capable of binding to recombinant spike and receptor-binding-domain (RBD) proteins (Fig. 2a). At site A, spike and RBD binding seropositivity were approximately 40 to 50% over the 17-week study, with high agreement between the two antigens. Conversely, at site B, binding seropositivity at the start of the study, immediately prior to the large outbreak, was low (∼12%), but rapidly rose to ∼35% post outbreak (Fig. 2a). At site A, spike and RBD antibody binding levels gradually declined over the first 8 weeks, suggesting recent infection and progression from an acute to convalescent stage (Fig. 2b). At site B, binding levels quickly increased immediately following the outbreak and were stable over the following weeks (Fig. 2b).
FIG 2.
SARS-CoV-2 polyclonal antibodies bind spike and RBD. (a) Polyclonal immune sera from sites A and B were evaluated for their ability to bind recombinant spike (solid) and RBD (dash) protein; n indicates the number of samples tested each week. (b) Level of spike and RBD binding as determined by absorbance reading. Dashed line represents Youden cutoffs.
SARS-CoV-2 serum neutralization.
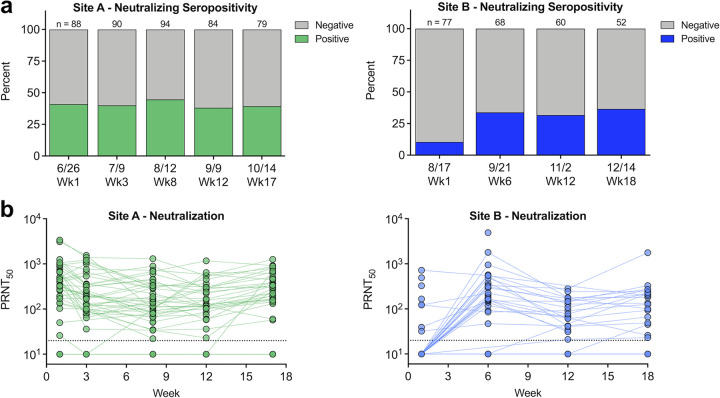
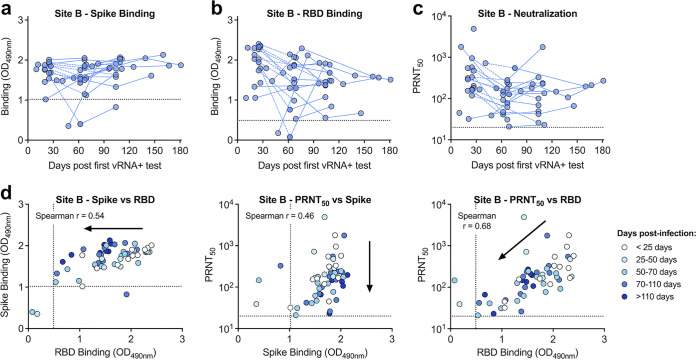
Sera were next evaluated for their ability to neutralize live SARS-CoV-2 virus using a standard plaque reduction neutralization test (PRNT), and their 50% neutralization titers (PRNT50) were calculated (Fig. 3). In agreement with antibody binding results, site A had 40 to 50% neutralizing seropositivity, which was maintained throughout the study, whereas neutralizing seropositivity at site B rapidly increased from 10% to 35% between the first sample and subsequent weeks (Fig. 3a). At site A, neutralizing titers were highly stable over the 17-week study, whereas at site B, neutralizing titers rose as individuals became infected, decreased following the acute response, and were stable during convalescence (Fig. 3b). Neutralizing antibody levels of individuals at site B who were infected prior to the beginning of the study were highly stable over the 18-week study, suggesting they were infected weeks/months prior (Fig. 3b). When analyzed using a more stringent PRNT80 cutoff, we saw slightly lower percent seropositivity (Fig. S1a in the supplemental material) and lower overall neutralization levels; however, the trends match the PRNT50 results (Fig. S1b).
FIG 3.
Polyclonal antibodies neutralize SARS-CoV-2 virus. (a) Polyclonal immune sera from sites A and B were evaluated for the ability to neutralize SARS-CoV-2 virus; n indicates the number of samples tested each week. (b) Neutralizing antibody levels over time. PRNT50 represents the serum dilution factor required to neutralize 50% of virus. The dashed line represents limit of detection (20). Nonneutralizing samples are graphed at half the limit of detection (10).
Relationship between SARS-CoV-2 polyclonal antibody binding and neutralization.
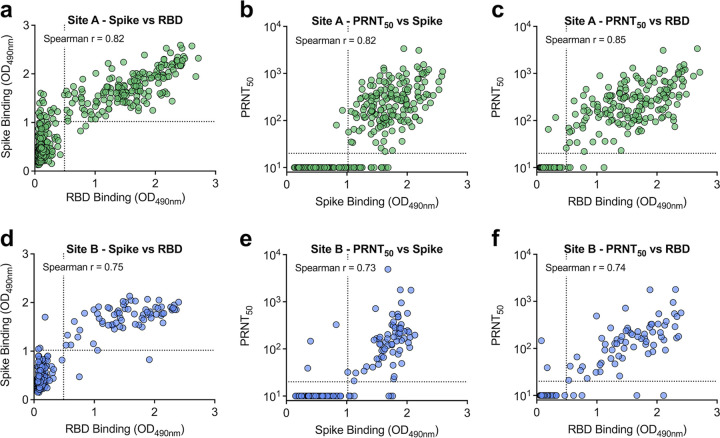
To better understand the relationship between binding and functionally neutralizing antibodies, spike and RBD binding levels and neutralizing titers were compared (Fig. 4). At both sites, spike and RBD levels were highly positively correlated (P < 0.001, Spearman r > 0.7), suggesting the majority of spike antibodies bind within the RBD (Fig. 4a and d). At site A, there was a small population (3.9%) of samples with spike binding antibodies that were negative for RBD (Fig. 4a). Both spike and RBD antibody binding levels were highly correlated with neutralizing titers (P < 0.001, Spearman r > 0.7) (Fig. 4b, c, e, and f); however, at both sites, RBD-binding antibodies were more strongly correlated with neutralization (Fig. 4c and f). Additionally, we compared a 50% neutralization cutoff value with an 80% cutoff and found that both titers were highly correlated (Spearman r > 0.9) (Fig. 2a and d). When comparing the PRNT80 with spike and RBD binding levels, we saw strong correlations (Spearman r > 0.6); however, they were less strongly correlated than when using a PRNT50 neutralization titer (Fig. S2b, c, e, and f).
FIG 4.
Spike binding, RBD binding, and neutralizing antibody levels are highly correlated. Samples from site A (a to c) and site B (d to f) were graphed by spike and RBD binding levels (a and d), spike binding and neutralization titers (b and e), and RBD binding and neutralization titers (c and f). Spike and RBD dashed lines represent Youden cutoffs. PRNT50 represents the serum dilution factor required to neutralize 50% of virus. PRNT50 dashed line represents limit of detection (20). Nonneutralizing samples are graphed at half the limit of detection (10). Two-tailed, nonparametic Spearman correlation is noted in the graphs.
Kinetics of SARS-CoV-2 antibody levels postinfection.
At site B, many individuals became infected and seroconverted during the course of the study. Therefore, in these individuals, we calculated the days postinfection (first positive vRNA nasal test) relative to seroconversion and levels of antibody binding and neutralization (Fig. 5). Spike binding antibody levels were high within 30 days of a positive PCR test and remained high throughout the monitoring period (Fig. 5a). RBD binding levels were more variable and dynamic, with some individuals generating RBD-specific antibodies within 10 days following infection, whereas one individual took over 60 days to seroconvert (Fig. 5b). Neutralizing antibody titers were also variable and dynamic across individuals, though most individuals generated high levels within a month following infection (Fig. 5c, Fig. S3a). In 85% of individuals, RBD-binding and neutralizing antibody levels decreased during the first 2 to 3 months following infection and then stabilized (Fig. 5b and c, dashed lines). When comparing the relationship between binding and neutralizing antibodies stratified by timing postinfection, we again saw RBD and neutralizing antibody levels generally decrease ∼30 days postinfection, whereas spike antibodies were highly stable (Fig. 5d, Fig. S3b and c). Additionally, binding and neutralizing antibodies were highly correlated regardless of timing postinfection (P < 0.001).
FIG 5.
Trends in binding and neutralizing antibody levels vary over time. Individuals at site B who were infected during the course of the surveillance study were sampled up to 180 days postinfection. (a) Spike binding; (b) RBD binding; and (c) neutralizing antibody levels are graphed by days post first vRNA positive test. (d) Samples are stratified by days postinfection, and graphed by spike binding, RBD binding, and neutralization titers. Arrows show trend of data over time. Spike and RBD dashed lines represent Youden cutoffs. PRNT50 represents the serum dilution factor required to neutralize 50% of virus. PRNT50 dashed line represents limit of detection (20). Nonneutralizing samples are graphed at half the limit of detection (10). Two-tailed, nonparametic Spearman correlation is noted in the graphs.
Phylogenetic analyses reveal lack of workplace SARS-CoV-2 spread.
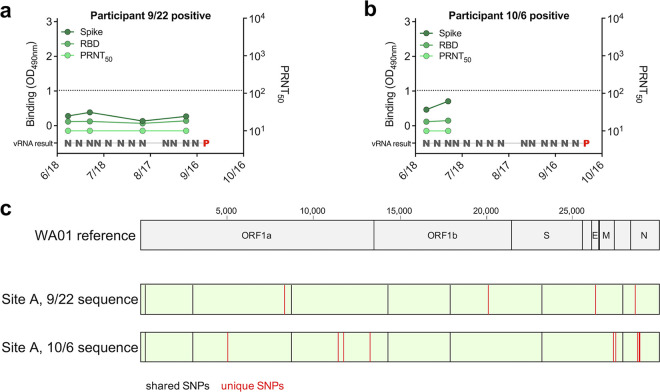
While site A did not experience any outbreaks (infections in more than 3 individuals) during our surveillance testing, two individual staff members tested positive during our study (Fig. 1b). These infections did not result in outbreaks or spread to other staff (Fig. 2a and 3a). The two individuals who tested positive for SARS-CoV-2 vRNA (2 weeks apart on 22 September and 6 October) provided serum samples in the weeks preceding their infections. Both individuals lacked detectable binding or neutralizing antibodies prior to infection and were thus immunologically naive (Fig. 6a and b). To determine if the two viruses were genetically related, and therefore likely acquired from one another, viral genomes from the cases were sequenced. Both viruses contained shared single nucleotide polymorphisms (SNPs) relative to a reference strain (WA01); however, they also contained 13 unique SNPs that strongly distinguish one from the other (Fig. 6c), suggesting two independent infections.
FIG 6.
Two seronegative individuals at site A became vRNA positive with unique strains. (a and b) Spike binding, RBD binding, and neutralizing antibody levels relative to timing of surveillance vRNA testing indicated these two individuals were seronegative prior to infection (N, SARS-CoV-2 negative; P, SARS-CoV-2 positive). (c) Viral RNA from positive surveillance testing was deep sequenced and the consensus sequence was compared to the WA01 SARS-CoV-2 reference sequence. Single nucleotide polymorphisms (SNPs) shared between both site A sequences relative to the reference are shown as black lines. Unique SNPs between the site A sequences are shown as red lines.
DISCUSSION
Weekly surveillance testing revealed facility-specific SARS-CoV-2 infection rates. Site B experienced a large outbreak with 34 of the staff members who participated in surveillance testing positive, whereas site A only had two positive tests out of greater than 1,600 samples total. The high infection rate of staff at site B matches the incidence rates in staff at other LTCFs during outbreaks (11, 40–42), highlighting how quickly the virus can spread among long-term care facility staff. Conversely, the low incidence of SARS-CoV-2 infection among staff at site A, paired with the high seroprevalence, suggests prior exposure and protection. Interestingly, at site A far more staff had antibodies than had previously tested positive for vRNA, suggesting a high fraction of asymptomatic infections, as has been documented in other facilities (41, 43, 44).
Two individuals at site A were vRNA positive for SARS-CoV-2 during the monitoring period. Full genome analysis of RNA recovered from these individuals revealed a significant number of genetic differences between the two isolates, suggesting they were acquired independently outside work as two unique instances of community transmission. Our finding that staff at site A had high preexisting seroprevalence (>40%) prior to intensive monitoring suggests this facility experienced a prior outbreak and had a level of immunity that limited spread of the virus from the two positive staff members (45). It is possible that other control measures and policies instituted at the time of monitoring, such as negative pressure isolation space (46), surveillance and monitoring systems, quarantine of positive staff (40, 47–49), environmental cleaning (50, 51), and other measures (52) additionally contributed to protection against outbreaks. It is notable that at both sites, seroprevalence reached a maximum of 40% during the study period, suggesting this might correspond to a level of naturally acquired immunity when coupled with other preventative measures.
Seroconversion and antibody levels were measured and characterized using three measures, i.e., binding to spike, binding to RBD, and neutralization of live SARS-CoV-2 virus. We found that immediately following infection, antibody levels peaked during the acute phase and then gradually decreased during convalescence. Neutralizing antibody levels were highly stable for at least 4 months postinfection, consistent with results reported by others (33, 37, 39). It is possible there is cross-reactivity between polyclonal antibodies from other human coronaviruses (NL63, OC43, or 229E) and SARS-CoV-2, especially when using low serum dilutions in binding assays; however, others have shown little cross-neutralization between these viruses (53–55). Antibodies that bound to spike antigen were detected earlier and more consistently than antibodies binding to RBD; however, RBD-binding antibody levels correlated most strongly with neutralizing titers, a result reported in other studies (56–60). Within our cohort, there were only four samples (0.57%) that neutralized SARS-CoV-2 but did not bind RBD. These likely neutralized through a mechanism other than blocking receptor interactions (61–63). Since RBD-binding antibodies can be detected using high-throughput platforms such as enzyme-linked immunosorbent assays (ELISAs), whereas live-virus neutralization assays require BSL3 facilities, are lower throughput, and take longer, our observations that RBD-binding antibodies are strongly correlated with neutralization suggest the more convenient binding assay may, in some circumstances, serve as a substitute for functional antiviral assays (64–68).
SARS-CoV-2 infection results in an immune response that includes the development of neutralizing antibodies (14, 15). These antibodies provide some degree of protection against reinfection with SARS-CoV-2; however, their persistence and durability are unknown, and human correlates of antibody-based protection are lacking (69–71). SARS-CoV-2 outbreaks at LTCFs can lead to high levels of seroprevalence that can limit spread within facilities (72). Without complete herd immunity, there are still nonimmune naive individuals who can become infected and spread the virus, possibly leading to secondary outbreaks (73). In our study, we observed that 40% seroprevalence in one facility, coupled with enhanced environmental controls, afforded apparent protection against subsequent outbreaks compared to a facility with low levels of preexisting seroconverted workers. Due to the high risk of infection of vulnerable individuals, staff and residents in LTCFs were among the highest priority for vaccination (74). Even in settings like LTCFs that have moderate levels of immunity acquired from natural infection, it is critical that individuals, including those previously infected, get vaccinated to further increase levels of immunity. This immunity is already drastically reducing the burden of SARS-CoV-2 in many LTCFs (75–77). Immunity, paired with additional infection control measures, will continue to reduce the incidence and prevalence of SARS-CoV-2 infection and mortality in these vulnerable facilities.
MATERIALS AND METHODS
Human specimens.
This study was reviewed and approved by the Colorado State University IRB under protocol number 20-10057H. Participants were consented and enrolled in our study and promptly informed of all test results. Staff represented all job classifications, including those in direct patient care roles (nurses, physical therapists, etc.) and nondirect patient care roles (custodial, administrative, etc.).
SARS-CoV-2 vRNA surveillance testing.
Nasal swabs were collected, processed, and tested for viral RNA as described previously (11). Briefly, swabs were collected by trained personnel and placed in tubes containing viral transport medium. RNA was extracted and quantitative reverse transcriptase PCR (qRT-PCR) was performed using the CDC 2019-nCoV primers and probes (78) or the Thermo Fisher Scientific TaqPath COVID-19 combo kit, under the U.S. Food and Drug Administration (FDA) Emergency Use Authorization (EUA).
Serum collection and processing.
Whole blood was collected in BD Vacutainer blood collection tubes (catalog number 368660). Samples were incubated for 30 to 60 min at room temperature to ensure clot formation, spun at 1,300 × g for 10 min at 25°C with gradual acceleration and deceleration, after which sera were aliquoted and stored at –20°C. Prior to use, sera were heat inactivated at 56°C for 30 min and then stored at 4°C.
Spike and RBD binding assays.
RBD and spike ELISAs were modified from Amanat et al. (79). Clear, flat-bottom immune 96-well plates were coated at 2 μg/ml with SARS-CoV-2 protein (Sino) and incubated overnight at 4°C. Samples were diluted 1:50 in diluent (1% milk powder, Tween, phosphate-buffered saline [PBS]) and added to plates for 2 h at room temperature after 1 h of blocking (PBS, milk powder, Tween). Positive controls included convalescent COVID-19 patient serum (gift of Raymond Goodrich) and monoclonal antibody CR3022 (Absolute Antibody). Charcoal-inactivated pooled human serum collected in 2015 was used as a negative control (Jackson Immuno Research). Plates were washed 3× and then anti-human IgG-horseradish peroxidase (HRP) (reacts with the heavy chains of human IgG and with light chains common to most human immunoglobulins) diluted 1:3,000 (PBS, 1% milk, Tween) was added for 1 h. Plates were washed 3× and then indicator was added and incubated for 10 min (SigmaFast OPD, Sigma). Reactions were stopped with 3 M HCl and plates were read at 490 nm with a Multiskan Spectrum spectrophotometer. Raw absorbance values were not corrected or normalized to any controls.
The cutoffs for classifying ELISA results as positive/negative were based on the average optical density (OD) values across two replicates. For each binding assay, the OD cutoff was specified as that which maximizes concordance with the SARS-CoV-2 neutralization assay results, specifically that maximizing the sum of the percent positive agreement (PPA) and the percent negative agreement (PNA), akin to Youden’s index. The resulting empirical PPA and PNA were 98% and 97% for the RBD binding assay, respectively, and 99% and 92% for the spike binding assay, respectively.
SARS-CoV-2 neutralization assay.
Vero cells were plated 1 day prior to infection. Heat-inactivated sera were serially diluted in Dulbecco’s modified Eagle medium (DMEM) containing 1% fetal bovine serum (FBS) mixed with ∼50 PFU SARS-CoV-2 (2019-nCoV/USA-WA1/2020 strain), and incubated for 1 h at 37°C. Virus-antibody mixture was added to cells, incubated for 1 h at 37°C, then overlaid with tragacanth medium. Cells were incubated for 2 days at 37°C, then fixed and stained with 30% ethanol and 0.1% crystal violet. Plaques were counted manually. Negatives controls included charcoal-inactivated pooled human serum collected in 2015 (Jackson Immuno Research) and dilution medium with no sera added. Immune sera from SARS-CoV-2 experimentally infected cats were used as positive controls (kindly gifted by Angela Bosco-Lauth).
SARS-CoV-2 whole-genome sequencing.
Sequencing was performed as previously described (11). Briefly, cDNA was generated using SuperScript IV, PCR amplification was performed with ARTIC tiled primers and Q5 High-Fidelity polymerase. PCR products were purified and libraries were prepared using KAPA HyperPrep kit and unique index primers. Libraries were sequenced on the Illumina MiSeq V2 using 2 × 250 paired-end reads. Sequencing data were processed, quality checked, and consensus sequences were determined.
ACKNOWLEDGMENTS
This work was supported by the Boettcher Foundation and funds donated by the Colorado State University Colleges of Health and Human Sciences, Veterinary Medicine and Biomedical Sciences, Natural Sciences, and Walter Scott, Jr. College of Engineering, the Colorado State University Columbine Health Systems Center for Healthy Aging and the CSU One Health Institute.
The authors also gratefully acknowledge the CSU Veterinary Diagnostic Laboratory for diagnostic support and the participation of the workers in the facilities who participated in this study, without whom it could not have been completed.
Footnotes
Supplemental material is available online only.
Contributor Information
Gregory D. Ebel, Email: gregory.ebel@colostate.edu.
S. Wesley Long, Houston Methodist Hospital.
REFERENCES
- 1.Blumenthal D, Fowler EJ, Abrams M, Collins SR. 2020. Covid-19—implications for the health care system. N Engl J Med 383:1483–1488. doi: 10.1056/NEJMsb2021088. [DOI] [PubMed] [Google Scholar]
- 2.Ferrara P, Albano L. 2020. COVID-19 and healthcare systems: what should we do next? Public Health 185:1–2. doi: 10.1016/j.puhe.2020.05.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Centers for Medicare & Medicaid Services. 2020. The nursing home COVID-19 public file. https://data.cms.gov/stories/s/COVID-19-Nursing-Home-Data/bkwz-xpvg/. [PubMed]
- 4.New York Times. 2020. More than 40% of U.S. coronavirus deaths are linked to nursing homes. https://www.nytimes.com/interactive/2020/us/coronavirus-nursing-homes.html.
- 5.Ouslander JG, Grabowski DC. 2020. COVID-19 in nursing homes: calming the perfect storm. J Am Geriatr Soc 68:2153–2162. doi: 10.1111/jgs.16784. [DOI] [PubMed] [Google Scholar]
- 6.Lai CC, Wang JH, Ko WC, Yen MY, Lu MC, Lee CM, Hsueh PR, Society of Taiwan Long-term Care Infection Prevention and Control. 2020. COVID-19 in long-term care facilities: an upcoming threat that cannot be ignored. J Microbiol Immunol Infect 53:444–446. doi: 10.1016/j.jmii.2020.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Daugherty JD, Blake SC, Grosholz JM, Omer SB, Polivka-West L, Howard DH. 2015. Influenza vaccination rates and beliefs about vaccination among nursing home employees. Am J Infect Control 43:100–106. doi: 10.1016/j.ajic.2014.08.021. [DOI] [PubMed] [Google Scholar]
- 8.Tamburrano A, Mellucci C, Galletti C, Vitale D, Vallone D, Barbara A, Sguera A, Zega M, Damiani G, Laurenti P. 2019. Improving nursing staff attitudes toward vaccinations through academic detailing: the HProImmune questionnaire as a tool for medical management. Int J Environ Res Public Health 16:2006. doi: 10.3390/ijerph16112006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Berry SD, Johnson KS, Myles L, Herndon L, Montoya A, Fashaw S, Gifford D. 2021. Lessons learned from frontline skilled nursing facility staff regarding COVID-19 vaccine hesitancy. J Am Geriatr Soc 69:1140–1146. doi: 10.1111/jgs.17136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Harrison J, Berry S, Mor V, Gifford D. 2021. “Somebody like me”: understanding COVID-19 vaccine hesitancy among staff in skilled nursing facilities. J Am Med Dir Assoc 22:1133–1137. doi: 10.1016/j.jamda.2021.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gallichotte EN, Quicke KM, Sexton NR, Fitzmeyer E, Young MC, Janich AJ, Dobos K, Pabilonia KL, Gahm G, Carlton EJ, Ebel GD, Ehrhart N. 2020. Longitudinal surveillance for SARS-CoV-2 among staff in six Colorado long-term care facilities: epidemiologic, virologic and sequence analysis. medRxiv. doi: 10.1101/2020.06.08.20125989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.de Wit E, van Doremalen N, Falzarano D, Munster VJ. 2016. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol 14:523–534. doi: 10.1038/nrmicro.2016.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hsueh PR, Huang LM, Chen PJ, Kao CL, Yang PC. 2004. Chronological evolution of IgM, IgA, IgG and neutralisation antibodies after infection with SARS-associated coronavirus. Clin Microbiol Infect 10:1062–1066. doi: 10.1111/j.1469-0691.2004.01009.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mackey K, Arkhipova-Jenkins I, Armstrong C, Gean E, Anderson J, Paynter RA, Helfand M. 2021. Antibody response following SARS-CoV-2 infection and Implications for immunity: a Rapid Living Review. Agency for Healthcare Research and Quality, Rockville, MD. [PubMed] [Google Scholar]
- 15.Carvalho T, Krammer F, Iwasaki A. 2021. The first 12 months of COVID-19: a timeline of immunological insights. Nat Rev Immunol 21:245–256. doi: 10.1038/s41577-021-00522-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wolfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Muller MA, Niemeyer D, Jones TC, Vollmar P, Rothe C, Hoelscher M, Bleicker T, Brunink S, Schneider J, Ehmann R, Zwirglmaier K, Drosten C, Wendtner C. 2020. Virological assessment of hospitalized patients with COVID-2019. Nature 581:465–469. doi: 10.1038/s41586-020-2196-x. [DOI] [PubMed] [Google Scholar]
- 17.Pierce CA, Preston-Hurlburt P, Dai Y, Aschner CB, Cheshenko N, Galen B, Garforth SJ, Herrera NG, Jangra RK, Morano NC, Orner E, Sy S, Chandran K, Dziura J, Almo SC, Ring A, Keller MJ, Herold KC, Herold BC. 2020. Immune responses to SARS-CoV-2 infection in hospitalized pediatric and adult patients. Sci Transl Med 12:eabd5487. doi: 10.1126/scitranslmed.abd5487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Suthar MS, Zimmerman MG, Kauffman RC, Mantus G, Linderman SL, Hudson WH, Vanderheiden A, Nyhoff L, Davis CW, Adekunle O, Affer M, Sherman M, Reynolds S, Verkerke HP, Alter DN, Guarner J, Bryksin J, Horwath MC, Arthur CM, Saakadze N, Smith GH, Edupuganti S, Scherer EM, Hellmeister K, Cheng A, Morales JA, Neish AS, Stowell SR, Frank F, Ortlund E, Anderson EJ, Menachery VD, Rouphael N, Mehta AK, Stephens DS, Ahmed R, Roback JD, Wrammert J. 2020. Rapid generation of neutralizing antibody responses in COVID-19 patients. Cell Rep Med 1:100040. doi: 10.1016/j.xcrm.2020.100040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Okba NMA, Muller MA, Li W, Wang C, GeurtsvanKessel CH, Corman VM, Lamers MM, Sikkema RS, de Bruin E, Chandler FD, Yazdanpanah Y, Hingrat QL, Descamps D, Houhou-Fidouh N, Reusken C, Bosch BJ, Drosten C, Koopmans MPG, Haagmans BL. 2020. Severe acute respiratory syndrome coronavirus 2-specific antibody responses in coronavirus disease patients. Emerg Infect Dis 26:1478–1488. doi: 10.3201/eid2607.200841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yang Y, Wang X, Du RH, Zhang W, Si HR, Zhu Y, Shen XR, Li Q, Li B, Men D, Zhou YN, Wang H, Tong XL, Zhang XE, Shi ZL, Zhou P. 2021. Serological investigation of asymptomatic cases of SARS-CoV-2 infection reveals weak and declining antibody responses. Emerg Microbes Infect 10:905–912. doi: 10.1080/22221751.2021.1919032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Long QX, Tang XJ, Shi QL, Li Q, Deng HJ, Yuan J, Hu JL, Xu W, Zhang Y, Lv FJ, Su K, Zhang F, Gong J, Wu B, Liu XM, Li JJ, Qiu JF, Chen J, Huang AL. 2020. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat Med 26:1200–1204. doi: 10.1038/s41591-020-0965-6. [DOI] [PubMed] [Google Scholar]
- 22.Hanrath AT, Schim van der Loeff I, Lendrem DW, Baker KF, Price DA, McDowall P, McDowall K, Cook S, Towns P, Schwab U, Evans A, Dixon J, Collins J, Burton-Fanning S, Saunders D, Harwood J, Samuel J, Schmid ML, Pareja-Cebrian L, Hunter E, Murphy E, Taha Y, Payne BAI, Duncan CJA. 2021. SARS-CoV-2 testing of 11,884 healthcare workers at an acute NHS hospital trust in England: a retrospective analysis. Front Med (Lausanne) 8:636160. doi: 10.3389/fmed.2021.636160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Germain N, Herwegh S, Hatzfeld AS, Bocket L, Prevost B, Danze PM, Marchetti P. 2021. Retrospective study of COVID-19 seroprevalence among tissue donors at the onset of the outbreak before implementation of strict lockdown measures in France. Cell Tissue Bank 1–8. doi: 10.1007/s10561-021-09901-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Comelli A, Foca E, Sansone E, Tomasi C, Albini E, Quiros-Roldan E, Tomasoni LR, Sala E, Bonfanti C, Caccuri F, Caruso A, Palma GD, Castelli F. 2021. Serological response to SARS-CoV-2 in health care workers employed in a large tertiary hospital in Lombardy, Northern Italy. Microorganisms 9:488. doi: 10.3390/microorganisms9030488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tang H, Tian JB, Dong JW, Tang XT, Yan ZY, Zhao YY, Xiong F, Sun X, Song CX, Xiang CG, Tu C, Lei CT, Liu J, Su H, Huang J, Qiu Y, Miao XP, Zhang C. 2020. Serologic detection of SARS-CoV-2 infections in hemodialysis centers: a multicenter retrospective study in Wuhan, China. Am J Kidney Dis 76:490–499. doi: 10.1053/j.ajkd.2020.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Klein SL, Pekosz A, Park HS, Ursin RL, Shapiro JR, Benner SE, Littlefield K, Kumar S, Naik HM, Betenbaugh MJ, Shrestha R, Wu AA, Hughes RM, Burgess I, Caturegli P, Laeyendecker O, Quinn TC, Sullivan D, Shoham S, Redd AD, Bloch EM, Casadevall A, Tobian AA. 2020. Sex, age, and hospitalization drive antibody responses in a COVID-19 convalescent plasma donor population. J Clin Invest 130:6141–6150. doi: 10.1172/JCI142004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Takahashi T, Ellingson MK, Wong P, Israelow B, Lucas C, Klein J, Silva J, Mao T, Oh JE, Tokuyama M, Lu P, Venkataraman A, Park A, Liu F, Meir A, Sun J, Wang EY, Casanovas-Massana A, Wyllie AL, Vogels CBF, Earnest R, Lapidus S, Ott IM, Moore AJ, Yale IRT, Shaw A, Fournier JB, Odio CD, Farhadian S, Dela Cruz C, Grubaugh ND, Schulz WL, Ring AM, Ko AI, Omer SB, Iwasaki A, Yale IMPACT Research Team. 2020. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 588:315–320. doi: 10.1038/s41586-020-2700-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yang HS, Costa V, Racine-Brzostek SE, Acker KP, Yee J, Chen Z, Karbaschi M, Zuk R, Rand S, Sukhu A, Klasse PJ, Cushing MM, Chadburn A, Zhao Z. 2021. Association of age with SARS-CoV-2 antibody response. JAMA Netw Open 4:e214302. doi: 10.1001/jamanetworkopen.2021.4302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zeng F, Dai C, Cai P, Wang J, Xu L, Li J, Hu G, Wang Z, Zheng F, Wang L. 2020. A comparison study of SARS-CoV-2 IgG antibody between male and female COVID-19 patients: a possible reason underlying different outcome between sex. J Med Virol 92:2050–2054. doi: 10.1002/jmv.25989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chen X, Pan Z, Yue S, Yu F, Zhang J, Yang Y, Li R, Liu B, Yang X, Gao L, Li Z, Lin Y, Huang Q, Xu L, Tang J, Hu L, Zhao J, Liu P, Zhang G, Chen Y, Deng K, Ye L. 2020. Disease severity dictates SARS-CoV-2-specific neutralizing antibody responses in COVID-19. Signal Transduct Target Ther 5:180. doi: 10.1038/s41392-020-00301-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hansen CB, Jarlhelt I, Pérez-Alós L, Hummelshøj Landsy L, Loftager M, Rosbjerg A, Helgstrand C, Bjelke JR, Egebjerg T, Jardine JG, Sværke Jørgensen C, Iversen K, Bayarri-Olmos R, Garred P, Skjoedt M-O. 2021. SARS-CoV-2 antibody responses are correlated to disease severity in COVID-19 convalescent individuals. J Immunol 206:109–117. doi: 10.4049/jimmunol.2000898. [DOI] [PubMed] [Google Scholar]
- 32.Luo YR, Chakraborty I, Yun C, Wu AHB, Lynch KL. 2020. Kinetics of SARS-CoV-2 antibody avidity maturation and association with disease severity. Clin Infect Dis ciaa1389. doi: 10.1093/cid/ciaa1389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Betton M, Livrozet M, Planas D, Fayol A, Monel B, Vedie B, Bruel T, Tartour E, Robillard N, Manuguerra JC, Blanchard A, Ghosn J, Visseaux B, Pere H, Lebeaux D, Schwartz O, Veyer D, Hulot JS, French COVID cohort study group. 2021. Sera neutralizing activities against SARS-CoV-2 and multiple variants six month after hospitalization for COVID-19. Clin Infect Dis ciab308. doi: 10.1093/cid/ciab308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Iyer AS, Jones FK, Nodoushani A, Kelly M, Becker M, Slater D, Mills R, Teng E, Kamruzzaman M, Garcia-Beltran WF, Astudillo M, Yang D, Miller TE, Oliver E, Fischinger S, Atyeo C, Iafrate AJ, Calderwood SB, Lauer SA, Yu J, Li Z, Feldman J, Hauser BM, Caradonna TM, Branda JA, Turbett SE, LaRocque RC, Mellon G, Barouch DH, Schmidt AG, Azman AS, Alter G, Ryan ET, Harris JB, Charles RC. 2020. Dynamics and significance of the antibody response to SARS-CoV-2 infection. medRxiv. doi: 10.1101/2020.07.18.20155374. [DOI] [Google Scholar]
- 35.Crawford KHD, Dingens AS, Eguia R, Wolf CR, Wilcox N, Logue JK, Shuey K, Casto AM, Fiala B, Wrenn S, Pettie D, King NP, Greninger AL, Chu HY, Bloom JD. 2021. Dynamics of neutralizing antibody titers in the months after severe acute respiratory syndrome coronavirus 2 infection. J Infect Dis 223:197–205. doi: 10.1093/infdis/jiaa618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rodda LB, Netland J, Shehata L, Pruner KB, Morawski PA, Thouvenel CD, Takehara KK, Eggenberger J, Hemann EA, Waterman HR, Fahning ML, Chen Y, Hale M, Rathe J, Stokes C, Wrenn S, Fiala B, Carter L, Hamerman JA, King NP, Gale M, Jr., Campbell DJ, Rawlings DJ, Pepper M. 2021. Functional SARS-CoV-2-specific immune memory persists after mild COVID-19. Cell 184:169–183. doi: 10.1016/j.cell.2020.11.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ripperger TJ, Uhrlaub JL, Watanabe M, Wong R, Castaneda Y, Pizzato HA, Thompson MR, Bradshaw C, Weinkauf CC, Bime C, Erickson HL, Knox K, Bixby B, Parthasarathy S, Chaudhary S, Natt B, Cristan E, Aini TE, Rischard F, Campion J, Chopra M, Insel M, Sam A, Knepler JL, Capaldi AP, Spier CM, Dake MD, Edwards T, Kaplan ME, Scott SJ, Hypes C, Mosier J, Harris DT, LaFleur BJ, Sprissler R, Nikolich-Zugich J, Bhattacharya D. 2020. Orthogonal SARS-CoV-2 serological assays enable surveillance of low-prevalence communities and reveal durable humoral immunity. Immunity 53:925–933. doi: 10.1016/j.immuni.2020.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Isho B, Abe KT, Zuo M, Jamal AJ, Rathod B, Wang JH, Li Z, Chao G, Rojas OL, Bang YM, Pu A, Christie-Holmes N, Gervais C, Ceccarelli D, Samavarchi-Tehrani P, Guvenc F, Budylowski P, Li A, Paterson A, Yue FY, Marin LM, Caldwell L, Wrana JL, Colwill K, Sicheri F, Mubareka S, Gray-Owen SD, Drews SJ, Siqueira WL, Barrios-Rodiles M, Ostrowski M, Rini JM, Durocher Y, McGeer AJ, Gommerman JL, Gingras AC. 2020. Persistence of serum and saliva antibody responses to SARS-CoV-2 spike antigens in COVID-19 patients. Sci Immunol 5:eabe5511. doi: 10.1126/sciimmunol.abe5511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wajnberg A, Amanat F, Firpo A, Altman DR, Bailey MJ, Mansour M, McMahon M, Meade P, Mendu DR, Muellers K, Stadlbauer D, Stone K, Strohmeier S, Simon V, Aberg J, Reich DL, Krammer F, Cordon-Cardo C. 2020. Robust neutralizing antibodies to SARS-CoV-2 infection persist for months. Science 370:1227–1230. doi: 10.1126/science.abd7728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Taylor J, Carter RJ, Lehnertz N, Kazazian L, Sullivan M, Wang X, Garfin J, Diekman S, Plumb M, Bennet ME, Hale T, Vallabhaneni S, Namugenyi S, Carpenter D, Turner-Harper D, Booth M, Coursey EJ, Martin K, McMahon M, Beaudoin A, Lifson A, Holzbauer S, Reddy SC, Jernigan JA, Lynfield R, Minnesota Long-Term Care COVID-19 Response Group. 2020. Minnesota long-term care, serial testing for SARS-CoV-2 and virus whole genome sequencing inform infection risk at two skilled nursing facilities with COVID-19 outbreaks—Minnesota, April–June 2020. MMWR Morb Mortal Wkly Rep 69:1288–1295. doi: 10.15585/mmwr.mm6937a3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Karmarkar E, Blanco I, Amornkul P, DuBois A, Deng X, Moonan PK, Rubenstein BL, Miller DA, Kennedy I, Yu J, Dauterman J, Ongpin M, Hathaway W, Hoo L, Trammell S, Ememu E, Yu G, Khwaja Z, Lu W, Talai N, Jain S, Louie J, Philip S, Federman S, Masinde G, Wadford DA, Bobba N, Stoltey J, Smith A, Epson E, Chiu C, Bennett A, Vasquez AM, Williams T. 2020. Timely intervention and control of a novel coronavirus (COVID-19) outbreak at a large skilled nursing facility—San Francisco, California, 2020. Infect Control Hosp Epidemiol 1–8. doi: 10.1017/ice.2020.1375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Harris DA, Archbald-Pannone L, Kaur J, Cattell-Gordon D, Rheuban KS, Ombres RL, Albero K, Steele R, Bell TD, Mutter JB. 2021. Rapid telehealth-centered response to COVID-19 outbreaks in postacute and long-term care facilities. Telemed J E Health 27:102–106. doi: 10.1089/tmj.2020.0236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Feaster M, Goh YY. 2020. High proportion of asymptomatic SARS-CoV-2 infections in 9 long-term care facilities, Pasadena, California, USA, April 2020. Emerg Infect Dis 26:2416–2419. doi: 10.3201/eid2610.202694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Goldberg SA, Lennerz J, Klompas M, Mark E, Pierce VM, Thompson RW, Pu CT, Ritterhouse LL, Dighe A, Rosenberg ES, Grabowski DC. 2021. Presymptomatic transmission of severe acute respiratory syndrome coronavirus 2 among residents and staff at a skilled nursing facility: results of real-time polymerase chain reaction and serologic testing. Clin Infect Dis 72:686–689. doi: 10.1093/cid/ciaa991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Reilly TA. 2021. Development of herd immunity to COVID-19 in Louisiana nursing homes. J Am Med Dir Assoc 22:256–257. doi: 10.1016/j.jamda.2020.11.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Miller SL, Mukherjee D, Wilson J, Clements N, Steiner C. 2021. Implementing a negative pressure isolation space within a skilled nursing facility to control SARS-CoV-2 transmission. Am J Infect Control 49:438–446. doi: 10.1016/j.ajic.2020.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Garibaldi PMM, Ferreira NN, Moraes GR, Moura JC, Esposito DLA, Volpe GJ, Calado RT, Fonseca BAL, Borges MC. 2021. Efficacy of COVID-19 outbreak management in a skilled nursing facility based on serial testing for early detection and control. Braz J Infect Dis 25:101570. doi: 10.1016/j.bjid.2021.101570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Smith DRM, Duval A, Pouwels KB, Guillemot D, Fernandes J, Huynh BT, Temime L, Opatowski L, AP-HP/Universities/Inserm COVID-19 research collaboration . 2020. Optimizing COVID-19 surveillance in long-term care facilities: a modelling study. BMC Med 18:386. doi: 10.1186/s12916-020-01866-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Danis K, Fonteneau L, Georges S, Daniau C, Bernard-Stoecklin S, Domegan L, O'Donnell J, Hauge SH, Dequeker S, Vandael E, Van der Heyden J, Renard F, Sierra NB, Ricchizzi E, Schweickert B, Schmidt N, Abu Sin M, Eckmanns T, Paiva JA, Schneider E, ECDC Public Health Emergency Team. 2020. High impact of COVID-19 in long-term care facilities, suggestion for monitoring in the EU/EEA, May 2020. Euro Surveill 25:2000956. doi: 10.2807/1560-7917.ES.2020.25.22.2000956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kraay ANM, Hayashi MAL, Berendes DM, Sobolik JS, Leon JS, Lopman BA. 2021. Risk for fomite-mediated transmission of SARS-CoV-2 in child daycares, schools, nursing homes, and offices. Emerg Infect Dis 27:1229–1231. doi: 10.3201/eid2704.203631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Mouchtouri VA, Koureas M, Kyritsi M, Vontas A, Kourentis L, Sapounas S, Rigakos G, Petinaki E, Tsiodras S, Hadjichristodoulou C. 2020. Environmental contamination of SARS-CoV-2 on surfaces, air-conditioner and ventilation systems. Int J Hyg Environ Health 230:113599. doi: 10.1016/j.ijheh.2020.113599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Rios P, Radhakrishnan A, Williams C, Ramkissoon N, Pham B, Cormack GV, Grossman MR, Muller MP, Straus SE, Tricco AC. 2020. Preventing the transmission of COVID-19 and other coronaviruses in older adults aged 60 years and above living in long-term care: a rapid review. Syst Rev 9:218. doi: 10.1186/s13643-020-01486-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chan KH, Cheng VC, Woo PC, Lau SK, Poon LL, Guan Y, Seto WH, Yuen KY, Peiris JS. 2005. Serological responses in patients with severe acute respiratory syndrome coronavirus infection and cross-reactivity with human coronaviruses 229E, OC43, and NL63. Clin Diagn Lab Immunol 12:1317–1321. doi: 10.1128/CDLI.12.11.1317-1321.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Jiang S, Hillyer C, Du L. 2020. Neutralizing antibodies against SARS-CoV-2 and other human coronaviruses. Trends Immunol 41:355–359. doi: 10.1016/j.it.2020.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Poston D, Weisblum Y, Wise H, Templeton K, Jenks S, Hatziioannou T, Bieniasz P. 2020. Absence of SARS-CoV-2 neutralizing activity in pre-pandemic sera from individuals with recent seasonal coronavirus infection. Clin Infect Dis ciaa1803. doi: 10.1093/cid/ciaa1803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Freeman J, Conklin J, Oramus D, Cycon K. 2021. Serology testing demonstrates that antibodies to SARS-CoV-2 S1-RBD correlate with neutralization of virus infection of Vero E6 cells. J Appl Lab Med jfab027. doi: 10.1093/jalm/jfab027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Padoan A, Bonfante F, Cosma C, Chiara CD, Sciacovelli L, Pagliari M, Bortolami A, Costenaro P, Musso G, Basso D, Giaquinto C, Plebani M. 2021. Analytical and clinical performances of a SARS-CoV-2 S-RBD IgG assay: comparison with neutralization titers. Clin Chem Lab Med 59:1444–1452. doi: 10.1515/cclm-2021-0313. [DOI] [PubMed] [Google Scholar]
- 58.Premkumar L, Segovia-Chumbez B, Jadi R, Martinez DR, Raut R, Markmann A, Cornaby C, Bartelt L, Weiss S, Park Y, Edwards CE, Weimer E, Scherer EM, Rouphael N, Edupuganti S, Weiskopf D, Tse LV, Hou YJ, Margolis D, Sette A, Collins MH, Schmitz J, Baric RS, de Silva AM. 2020. The receptor binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci Immunol 5:eabc8413. doi: 10.1126/sciimmunol.abc8413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Rockstroh A, Wolf J, Fertey J, Kalbitz S, Schroth S, Lubbert C, Ulbert S, Borte S. 2021. Correlation of humoral immune responses to different SARS-CoV-2 antigens with virus neutralizing antibodies and symptomatic severity in a German COVID-19 cohort. Emerg Microbes Infect 10:774–781. doi: 10.1080/22221751.2021.1913973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Laterza R, Schirinzi A, Bruno R, Genco R, Contino R, Ostuni A, Di Serio F. 2021. SARS-CoV-2 antibodies: comparison of three high-throughput immunoassays versus the neutralization test. Eur J Clin Invest 5:e13573. doi: 10.1111/eci.13573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rogers TF, Zhao F, Huang D, Beutler N, Burns A, He WT, Limbo O, Smith C, Song G, Woehl J, Yang L, Abbott RK, Callaghan S, Garcia E, Hurtado J, Parren M, Peng L, Ramirez S, Ricketts J, Ricciardi MJ, Rawlings SA, Wu NC, Yuan M, Smith DM, Nemazee D, Teijaro JR, Voss JE, Wilson IA, Andrabi R, Briney B, Landais E, Sok D, Jardine JG, Burton DR. 2020. Isolation of potent SARS-CoV-2 neutralizing antibodies and protection from disease in a small animal model. Science 369:956–963. doi: 10.1126/science.abc7520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Chi X, Yan R, Zhang J, Zhang G, Zhang Y, Hao M, Zhang Z, Fan P, Dong Y, Yang Y, Chen Z, Guo Y, Zhang J, Li Y, Song X, Chen Y, Xia L, Fu L, Hou L, Xu J, Yu C, Li J, Zhou Q, Chen W. 2020. A neutralizing human antibody binds to the N-terminal domain of the Spike protein of SARS-CoV-2. Science 369:650–655. doi: 10.1126/science.abc6952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Brouwer PJM, Caniels TG, van der Straten K, Snitselaar JL, Aldon Y, Bangaru S, Torres JL, Okba NMA, Claireaux M, Kerster G, Bentlage AEH, van Haaren MM, Guerra D, Burger JA, Schermer EE, Verheul KD, van der Velde N, van der Kooi A, van Schooten J, van Breemen MJ, Bijl TPL, Sliepen K, Aartse A, Derking R, Bontjer I, Kootstra NA, Wiersinga WJ, Vidarsson G, Haagmans BL, Ward AB, de Bree GJ, Sanders RW, van Gils MJ. 2020. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science 369:643–650. doi: 10.1126/science.abc5902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Abe KT, Li Z, Samson R, Samavarchi-Tehrani P, Valcourt EJ, Wood H, Budylowski P, Dupuis AP, 2nd, Girardin RC, Rathod B, Wang JH, Barrios-Rodiles M, Colwill K, McGeer AJ, Mubareka S, Gommerman JL, Durocher Y, Ostrowski M, McDonough KA, Drebot MA, Drews SJ, Rini JM, Gingras AC. 2020. A simple protein-based surrogate neutralization assay for SARS-CoV-2. JCI Insight 5:e142362. doi: 10.1172/jci.insight.142362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Meyer B, Reimerink J, Torriani G, Brouwer F, Godeke GJ, Yerly S, Hoogerwerf M, Vuilleumier N, Kaiser L, Eckerle I, Reusken C. 2020. Validation and clinical evaluation of a SARS-CoV-2 surrogate virus neutralisation test (sVNT). Emerg Microbes Infect 9:2394–2403. doi: 10.1080/22221751.2020.1835448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Putcharoen O, Wacharapluesadee S, Chia WN, Paitoonpong L, Tan CW, Suwanpimolkul G, Jantarabenjakul W, Ruchisrisarod C, Wanthong P, Sophonphan J, Chariyavilaskul P, Wang LF, Hemachudha T. 2021. Early detection of neutralizing antibodies against SARS-CoV-2 in COVID-19 patients in Thailand. PLoS One 16:e0246864. doi: 10.1371/journal.pone.0246864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Tan CW, Chia WN, Qin X, Liu P, Chen MI, Tiu C, Hu Z, Chen VC, Young BE, Sia WR, Tan YJ, Foo R, Yi Y, Lye DC, Anderson DE, Wang LF. 2020. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2-spike protein-protein interaction. Nat Biotechnol 38:1073–1078. doi: 10.1038/s41587-020-0631-z. [DOI] [PubMed] [Google Scholar]
- 68.Valcourt EJ, Manguiat K, Robinson A, Chen JC, Dimitrova K, Philipson C, Lamoureux L, McLachlan E, Schiffman Z, Drebot MA, Wood H. 2021. Evaluation of a commercially-available surrogate virus neutralization test for severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2). Diagn Microbiol Infect Dis 99:115294. doi: 10.1016/j.diagmicrobio.2020.115294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Krammer F. 2021. Correlates of protection from SARS-CoV-2 infection. Lancet 397:1421–1423. doi: 10.1016/S0140-6736(21)00782-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Brown M. 2020. Immune correlates of SARS-CoV-2 protection. Nat Rev Immunol 20:593. doi: 10.1038/s41577-020-00442-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Sui Y, Bekele Y, Berzofsky JA. 2021. Potential SARS-CoV-2 immune correlates of protection in infection and vaccine immunization. Pathogens 10:138. doi: 10.3390/pathogens10020138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Lumley SF, O'Donnell D, Stoesser NE, Matthews PC, Howarth A, Hatch SB, Marsden BD, Cox S, James T, Warren F, Peck LJ, Ritter TG, de Toledo Z, Warren L, Axten D, Cornall RJ, Jones EY, Stuart DI, Screaton G, Ebner D, Hoosdally S, Chand M, Crook DW, O'Donnell AM, Conlon CP, Pouwels KB, Walker AS, Peto TEA, Hopkins S, Walker TM, Jeffery K, Eyre DW, Oxford University Hospitals Staff Testing Group. 2021. Oxford University Hospitals staff testing, antibody status and incidence of SARS-CoV-2 infection in health care workers. N Engl J Med 384:533–540. doi: 10.1056/NEJMoa2034545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Cavanaugh AM, Thoroughman D, Miranda H, Spicer K. 2021. Suspected recurrent SARS-CoV-2 infections among residents of a skilled nursing facility during a second COVID-19 outbreak—Kentucky, July–November 2020. MMWR Morb Mortal Wkly Rep 70:273–277. doi: 10.15585/mmwr.mm7008a3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Dooling K, McClung N, Chamberland M, Marin M, Wallace M, Bell BP, Lee GM, Talbot HK, Romero JR, Oliver SE. 2020. The Advisory Committee on Immunization Practices' interim recommendation for allocating initial supplies of COVID-19 vaccine—United States, 2020. MMWR Morb Mortal Wkly Rep 69:1857–1859. doi: 10.15585/mmwr.mm6949e1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Britton A, Jacobs Slifka KM, Edens C, Nanduri SA, Bart SM, Shang N, Harizaj A, Armstrong J, Xu K, Ehrlich HY, Soda E, Derado G, Verani JR, Schrag SJ, Jernigan JA, Leung VH, Parikh S. 2021. Effectiveness of the Pfizer-BioNTech COVID-19 vaccine among residents of two skilled nursing facilities experiencing COVID-19 outbreaks—Connecticut, December 2020–February 2021. MMWR Morb Mortal Wkly Rep 70:396–401. doi: 10.15585/mmwr.mm7011e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mor V, Gutman R, Yang X, White EM, McConeghy KW, Feifer RA, Blackman CR, Kosar CM, Bardenheier BH, Gravenstein SA. 2021. Short-term impact of nursing home SARS-CoV-2 vaccinations on new infections, hospitalizations, and deaths. J Am Geriatr Soc doi: 10.1111/jgs.17176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.McEllistrem MC, Clancy CJ, Buehrle DJ, Lucas A, Decker BK. 2021. Single dose of a mRNA SARS-CoV-2 vaccine is associated with lower nasopharyngeal viral load among nursing home residents with asymptomatic COVID-19. Clin Infect Dis ciab263. doi: 10.1093/cid/ciab263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Centers for Disease Control. 2020. Research use only 2019-novel coronavirus (2019-nCoV) real-time RT-PCR primers and probes. https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html.
- 79.Amanat F, Stadlbauer D, Strohmeier S, Nguyen THO, Chromikova V, McMahon M, Jiang K, Arunkumar GA, Jurczyszak D, Polanco J, Bermudez-Gonzalez M, Kleiner G, Aydillo T, Miorin L, Fierer DS, Lugo LA, Kojic EM, Stoever J, Liu STH, Cunningham-Rundles C, Felgner PL, Moran T, Garcia-Sastre A, Caplivski D, Cheng AC, Kedzierska K, Vapalahti O, Hepojoki JM, Simon V, Krammer F. 2020. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat Med 26:1033–1036. doi: 10.1038/s41591-020-0913-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material. Download SPECTRUM00224-21_Supp_1_seq10.pdf, PDF file, 0.6 MB (667KB, pdf)