Abstract
The purpose of this study is to investigate the robustness of a commonly used convolutional neural network for image segmentation with respect to nearly unnoticeable adversarial perturbations, and suggest new methods to make these networks more robust to such perturbations. In this retrospective study, the accuracy of brain tumor segmentation was studied in subjects with low- and high-grade gliomas. Two representative UNets were implemented to segment four different MR series (T1-weighted, post-contrast T1-weighted, T2-weighted, and T2-weighted FLAIR) into four pixelwise labels (Gd-enhancing tumor, peritumoral edema, necrotic and non-enhancing tumor, and background). We developed attack strategies based on the fast gradient sign method (FGSM), iterative FGSM (i-FGSM), and targeted iterative FGSM (ti-FGSM) to produce effective but imperceptible attacks. Additionally, we explored the effectiveness of distillation and adversarial training via data augmentation to counteract these adversarial attacks. Robustness was measured by comparing the Dice coefficients for the attacks using Wilcoxon signed-rank tests. The experimental results show that attacks based on FGSM, i-FGSM, and ti-FGSM were effective in reducing the quality of image segmentation by up to 65% in the Dice coefficient. For attack defenses, distillation performed significantly better than adversarial training approaches. However, all defense approaches performed worse compared to unperturbed test images. Therefore, segmentation networks can be adversely affected by targeted attacks that introduce visually minor (and potentially undetectable) modifications to existing images. With an increasing interest in applying deep learning techniques to medical imaging data, it is important to quantify the ramifications of adversarial inputs (either intentional or unintentional).
Keywords: Deep learning segmentation, Robustness, Adversarial attacks, Defenses
Introduction
Machine learning algorithms have become increasingly popular in medical imaging [1–3], where highly functional algorithms have been trained to recognize patterns in image data sets and perform clinically relevant tasks such as tumor segmentation and disease diagnosis. In particular, approaches based on deep learning have recently drawn widespread attention [4]. However, an oft-repeated criticism of deep learning is that it uses a “black-box” approach, giving rise to decision-making processes that are uninterpretable even by domain experts and deep learning researchers [5, 6]. Furthermore, it has been demonstrated that neural networks can be tricked into misclassifying images when they are perturbed by negligible amounts of specific types of noise [7].
As more progress has been made by deep learning in applications to medical diagnosis and medical reimbursement decisions, concerns have arisen that adversarial examples may be utilized for fraud [8]. In Paschali et al. [9], the robustness of deep learning algorithms is defined as the performance gap created by introducing adversarial examples to the test data. Experimental results on skin lesion classification and whole-brain segmentation with state-of-the-art neural networks demonstrate that although different deep learning models have comparable performance on clean data, their robustness may vary (with a performance drop of as much as 37% for whole-brain segmentation), revealing the potential vulnerability of deep learning models in medical imaging.
The vast majority of existing robustness theory on deep learning focuses on classification tasks, where the goal of an adversarial attack is to generate a small perturbation to an image that causes a highly accurate neural network to misclassify the perturbed image. In contrast, image segmentation tasks are more complex. Although image segmentation may be viewed as a classification procedure operating on the individual pixels of an image, adversarial attacks typically consist of applying a global perturbation to the entire image that simultaneously changes the classes of all pixels in an adversarial manner. Additional work is needed to study which global perturbations might lead to specific types of segmentation errors, particularly for convolutional neural networks (CNNs), which are nearly ubiquitous in deep learning applications for medical image processing. We mention the recent paper [10], which proposed an adaptive segmentation mask attack (ASMA) to generate targeted adversarial examples for the task of segmentation. Similar to our method, ASMA aims at finding a small additive perturbation to change the prediction of deep learning models. However, our method differs from ASMA in that our general approach is to maximize the difference between predicted and true labels, whereas the goal of ASMA is to force the prediction of models to a specific target output, in which the optimization problem is more difficult to solve. Furthermore, unlike Ozbulak et al. [10], our paper also presents defense techniques to robustify neural networks.
In this paper, we investigate the vulnerability of deep learning algorithms in the context of medical image segmentation and propose methods that may be adopted during training to make deep learning algorithms more robust. Our objective is to study the hypothesis that adversarial attack strategies developed for neural network classifiers using visually subtle perturbations to input images [7, 11] can be adapted to the task of medical image segmentation. Our second contribution is to investigate methods for defending against such adversarial attacks. It has been shown in recent studies that adversarial training [7, 12] and defensive distillation [13] can increase the robustness of neural networks when applied to classification tasks for standard computer vision data sets such as MNIST and ImageNet [14]. In this paper, we will show that these methods are also effective for medical imaging segmentation. Both of our defense strategies, based on adversarial training and defensive distillation, show significantly improved robustness with respect to adversarial attacks.
Materials and Methods
Data from the The Cancer Imaging Archive (TCIA) and the Medical Image Computing & Computer Assisted Intervention (MICCAI) Brain Tumor Segmentation (BraTS) 2017 challenge [15–19] were used for this IRB-exempt study. These publicly available, retrospective data sets from multi-institutional studies consist of magnetic resonance (MR) images of the brain from 283 subjects with either low-grade glioma or glioblastoma multiforme. Each data set includes four different MR series: (a) T1-weighted (T1), (b) post-contrast T1-weighted (T1Gd), (c) T2-weighted (T2), and (d) T2-weighted fluid attenuated inversion recovery (FLAIR). Segmentation volumes, manually segmented by expert neuroradiologists, are provided with the following pixelwise labels: (i) Gd-enhancing tumor (ET-label 4), (ii) peritumoral edema (ED-label 2), (iii) necrotic and non-enhancing tumor (NCR/NET-label 1), and (iv) background (label 0). We reserved 20% of the data for testing. We used two representative CNN architectures, the 3D-UNet model [20], which has demonstrated good performance for segmentation tasks on this data set, and the classic 2D-UNet model [21] for medical image segmentation.
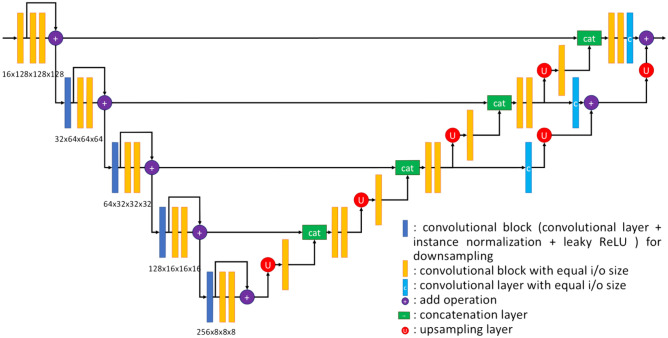
This 3D-UNet model contains 28 convolutional blocks, which include 3D convolution, instance normalization, and leaky ReLU layers. To make the network more efficient, it also has residual connections [22]. The network architecture is illustrated in Fig. 1.
Fig. 1.
The 3D-UNet architecture
The encoder module contains 15 convolutional blocks with residual connections. In the convolutional layers of these blocks, the size of all kernels is 3 × 3 × 3. The stride is set to 2 if we want the output size of the convolutional layers to be reduced to half the input size; otherwise, the stride is set to 1. Since the kernel size is odd, the zero-padding strategy is different. We add 1 column in the left, 2 columns in the right, 1 row at the top, and 2 rows at the bottom. There is a 3D dropout layer with dropout rate of 0.3 between the two orange convolutional blocks to make the training process faster and improve the generalization performance of the model.
In the decoder module, after each upsampling layer, there is one convolution block with kernel size of 3 × 3 × 3 and stride of 1. After each concatenation layer, there are two convolution blocks. The first convolution has the same structure as the convolution block after upsampling layers. The kernel size in the second one is 1 × 1 × 1. In the residual links, there are three convolutional layers (colored blue). The kernel size for all these convolutions is 1 × 1 × 1. The number of kernels in each convolution is the same as the number of labels in the ground truth. After the decoder, there is a softmax layer to calculate the probability of each label. 2D-UNets are constructed in a similar manner, except 3D convolutions are replaced by 2D convolutions.
Attacks
We focused on first-order attacks, which construct perturbations based on the gradient of a loss function evaluated on input images for a given trained network. We also studied targeted attacks, which encourage the result of a perturbation to fall into a specific category. For example, given an input tumor image, the goal might be to construct an adversary that results in an output that moves the tumor label to a certain (incorrect) position. We developed attack strategies based on the fast gradient sign method (FGSM) [7], iterative FGSM (i-FGSM) [11], and targeted iterative FGSM (ti-FGSM) [11].
FGSM perturbs an image according to the equation
| 1 |
where is the magnitude of perturbation on individual pixels, and loss(•) is the loss function with respect to input image X and its corresponding ground truth label . In this work, we adopt the soft Dice loss, which will be explained later.
i-FGSM, which is expected to have a higher success rate than FGSM for generating incorrectly classified images, consists of applying FGSM for multiple iterations:
| 2 |
where , and is the magnitude of the perturbation to individual pixels in each iteration.
The ti-FGSM perturbations are defined by the equation:
| 3 |
where , corresponding to iteratively minimizing the loss between the output label and the target label.
A key idea in our approach was to replace the cross-entropy loss used for usual classification tasks with the Dice coefficient loss in the FGSM algorithm. The Dice coefficient loss is equal to (1 – Dice coefficient) of the model segmented image and the true output, where the Dice coefficient is a metric which assesses the spatial overlap of two image segmentations [23]. For , we chose 5% of the maximum pixel magnitude of the input image. We chose and the number of steps , such that of the maximum pixel magnitude of the input image. Typically, we normalized input image pixel values to be within [0,1], so our was 0.05 and . We chose this perturbation level at 5% of the maximum pixel magnitude so that the perturbed images were not too different from the original images and indistinguishable to the human eye.
Defenses
For defenses, we first explored a method based on distillation [13]. The key idea is to retrain a neural network on a data set using vectors of soft labels that are obtained from an initial training stage of the neural network. The classification function of the “distilled” neural network, which predicts soft label vectors from inputs, is a continuous function that is smoother over the domain of input variables than the original network, thus is less sensitive to small input variations.
Specifically, there are two independent networks with the same hyperparameters in this method, which we call and , respectively. Firstly, is trained by the original data set. For classification problems, the label of each input is generally represented as a one-hot vector. A temperature parameter , which controls the desired level of smoothness of the distilled network, is introduced to the activation function of the output layer during training. After the training process is completed, the network will make predictions on all data points in the training data set. The outputs of are considered as soft labels and replace the original one-hot vectors in the training data set. Then, , the distilled network, will be trained on the new data set. Since the labels are changed, the loss function for training will also be defined differently.
Let be the model used for distillation, where is the input image. For example, in the 3D-UNet, the output of is a 4D array, and for each pixel of the input 3D image , we have an array of soft labels
| 4 |
where is the element with index in the 4D matrix before the activation function is applied, and is the number of classes in the data set. The temperature is a constant. If , the function above is the usual softmax.
Let be the number of rows, columns, and channels of , respectively. Given the one-hot truth matrix with dimensions , the soft Dice coefficient is calculated as
| 5 |
where is a small positive real number. Unlike the Dice coefficient, which is obtained after thresholding the prediction to convert it to a binary mask for each label class, the soft Dice coefficient is differentiable, which is essential for backward propagation. Note that by default, we set the normal pixels (i.e., the background of the image and the part of the tissue that does not have any disease) to class 0 ( here), and these pixels are ignored when calculating . Then, the loss function is defined according to the soft Dice coefficient
| 6 |
Let be the distilled network, which has the same architecture as . When training , the only difference is that instead of the one-hot matrix , we use the output from the first trained network . We define
| 7 |
and define the loss function of the distilled network as .
For adversarial training [7], the goal is to determine a model with trainable parameters that minimizes the population risk:
| 8 |
where is the set of allowed perturbations, is the data distribution, and is the loss function. In practice, the set is often defined to be the -ball of radius , meaning that each pixel can be perturbed by at most . To minimize the expectation above, a natural strategy is to perform gradient descent on the adversarial loss function. It may be shown that the gradient of the adversarial loss function at is identical to the gradient of the usual loss function evaluated at the “worst-case” point in the neighborhood of [12]. Identifying this worst-case point is not computationally feasible, so a popular alternative is to use an adversarial attack (e.g., FGSM), and then train the model by evaluating the gradient at the adversarial example. In practice, we often use the adversarial objective function based on FGSM as an effective regularizer [7]:
| 9 |
where is the weight factor. This method works well with , although other values may exist which achieve better performance. The goal of adversarial training is to minimize over the training data set.
Due to memory limitations, we implemented the minimization in an iterative way. For each batch of training data, we first generated adversarial examples based on the current model, and then performed forward and backward propagation using these adversarial examples. Then, the model was updated according to the original batch of data.
Recent work [24–26] has suggested that data augmentation, which introduces artificially generated images to the training set by adding random transformations to training images (e.g., random noise from uniform or Gaussian distributions on pixel magnitudes [24, 26], or random rotations to the input image [25]), can produce more robust networks. However, previous literature also suggests that data augmentation may have limited benefits for adversarial robustness. We compared our distillation strategy to the performance of the more straightforward data augmentation technique.
Measuring Robustness
To study the effects of adversarial attacks, we used fixed values of . For FGSM, we chose to be 0.05, which corresponds to 5% of the maximum intensity of the image. For i-FGSM, we chose and the number of iterates to be 10. For ti-FGSM, the target was all labels in the image equal to 1 (i.e., necrotic and non-enhancing tumor). To study the effects of network defenses, we used a range of values from 0 to 0.010, in increments of 0.001.
We evaluated the robustness of the attacked and defended networks by quantifying the effect of adversarial perturbations. The overall robustness of a classifier was obtained by comparing the average Dice coefficient of the segmented, adversarially perturbed test images with the average Dice coefficient of segmented, non-perturbed images. Wilcoxon signed-rank tests were used to determine whether the proposed perturbation strategies for inputs resulted in significantly different Dice coefficients of segmented outputs. Similarly, the peak signal-to-noise ratio (PSNR), Structural Similarity Index (SSIM), and root mean squared error (RMSE) of the perturbed input images were compared to the ground truth images for each type of attack.
Results
Analysis was performed on a data set containing 283 subjects. Demographic data is not available for this data set; however, for 163 of the subjects, age (60.3 ± 12.1 years) and overall survival (423 ± 350 days) were available. All experiments were conducted on one Titan XP GPU with 12 GB memory.
For data augmentation, we applied uniform perturbations of radius of 0.01 in infinity norm to the input data. For each batch of input data, we first trained the model with the perturbed images and then trained the model with the clean images, in order to obtain a fair comparison to adversarial training.
For data preprocessing, we applied N4 bias field correction [27] and global standardization. The 3D images were resized to 128 × 128 × 128 to match the input shape of the 3D-UNet. For training, we used a batch size of 1 and 100 epochs. The Adam optimizer was used with a learning rate of 1e-4. The training process in the distillation method is divided into two parts that share the same hyperparameters: When the temperature is high, more iterations are required, so we increased the number of epochs to 400 and the learning rate to 5e-4. To allow improved generalization, data augmentation was performed on the 3D images, which included rotation within the axial slices, flips, and matrix transposes.
In the training process of the 2D-UNet, the 3D images were converted into 128 axial 2D slices. We used a batch size of 64 and 400 epochs. The same Adam optimizer was used with a different learning rate of 1e-5 throughout all experiments. The same data augmentation methods were applied. Although only slice-wise prediction was allowed for the 2D-UNet, the predictions were combined for each input 3D volume to be evaluated.
Adversarial Attacks
We first present the results for the 3D-UNet. Example adversarial attacks are shown in Fig. 2, where we see that all three adversaries successfully inject errors into the segmented images, with minimal visual disturbance to the input images. This verifies that small adversarial perturbations to the input image can indeed have a substantial impact on the resulting segmentation.
Fig. 2.

Top row: selected axial slices of input 3D images. Bottom row: predicted segmentation for the three adversarial approaches (FGSM, i-FGSM, ti-FGSM) compared to the unperturbed input (far left)
The average Dice coefficients (mean ± standard deviation) of the predicted output with respect to the ground-truth masks and the PSNR, SSIM, and RMSE of the input images are shown in Table 1.
Table 1.
Segmentation results for three different attacks: fast gradient sign method (FGSM), iterative FGSM (i-FGSM), and targeted iterative FGSM (ti-FGSM), quantified via the Dice coefficient of the output segmentation and the PSNR, SSIM, and RMSE of the perturbed input images. For the Dice coefficient measurements, an asterisk (*) indicates statistically significant differences relative to “No attack” at the level , corrected for multiple comparisons
| Attack type | Dice coef — tumor core | Dice coef — enhancing tumor | Dice coef — whole tumor | Input PSNR | Input SSIM | Input RMSE |
|---|---|---|---|---|---|---|
| No attack | 0.821 ± 0.042 | 0.668 ± 0.253 | 0.748 ± 0.043 | - | - | - |
| FGSM | 0.561 ± 0.077* | 0.370 ± 0.239* | 0.549 ± 0.078* | 27.69 ± 0.28 | 0.646 ± 0.043 | 0.041 ± 0.001 |
| i-FGSM | 0.342 ± 0.087* | 0.230 ± 0.162* | 0.393 ± 0.090* | 27.93 ± 0.13 | 0.470 ± 0.015 | 0.040 ± 0.001 |
| ti-FGSM | 0.461 ± 0.085* | 0.365 ± 0.225* | 0.486 ± 0.082* | 29.48 ± 0.34 | 0.735 ± 0.064 | 0.034 ± 0.001 |
A Wilcoxon signed-rank test was used to compare the Dice coefficient to the ground truth data for each attack type (). A Bonferroni correction was applied to correct for multiple comparisons. The attacks were highly successful, since all variants of FGSM resulted in a significantly lower Dice coefficient. Compared to the “No attack” condition, the attacks reduced the Dice coefficient by 30.5%, 58.3%, and 43.8% in the tumor core; 44.6%, 65.6%, and 45.4% in the enhancing tumor; and 26.7%, 47.5%, and 35.0% in the whole tumor for FGSM, i-FGSM, and ti-FGSM, respectively. Despite visually subtle changes, the image quality metrics PSNR and SSIM suggest measurable differences between input images, while RMSE differences are low.
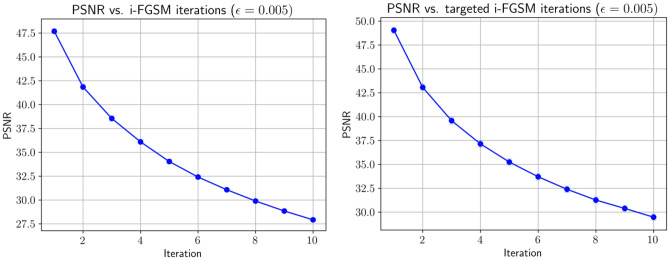
In Fig. 3, we show plots of the average Dice coefficient vs. number of iterations in i-FGSM and ti-FGSM. As expected, with an increasing number of iterations, we see a steadily decreasing Dice coefficient—indicating that with more steps, the adversaries become stronger, causing the segmentation output to worsen. The effects of i-FGSM iterations on image input quality are shown in Fig. 4. The decrease in PSNR is expected; however, note that the average PSNR is still reasonably large, implying that the quality of the perturbed images is relatively high. Additionally, as seen in Fig. 2, the effects are barely discernible, suggesting that PSNR (and the other image quality metrics of SSIM and RMSE) are sensitive to FGSM attacks.
Fig. 3.
Plots of the Dice coefficient vs. number of iterations for all study data using i-FGSM and ti-FGSM. Error bars are also shown. As the number of iterations increases, the adversaries become stronger, causing the segmentation output to worsen
Fig. 4.
Average PSNR vs. number of iterations in i-FGSM and ti-FGSM
Defense via Distillation
The prediction performance of distilled 3D neural networks for different training temperatures is shown in Fig. 5. In each plot, the robustness of the neural network clearly increases with . For , the gains are 0.14, 0.27, and 0.22, respectively, compared to the network without distilled training () at the worst attack case. This indicates that distillation is indeed effective in defending against the proposed adversarial attacks.
Fig. 5.
Performance of distillation in the 3D-UNet. Dice coefficients of a “Whole Tumor,” b “Tumor Core,” and c “Enhancing Tumor” vs. FGSM with different
It is also notable that the improvement appears to saturate when the temperature exceeds a certain threshold. For example, gains in robustness for temperatures over 100 in Fig. 5a are negligible. This phenomenon is not observed in Fig. 5b, c, because the threshold for the temperature is higher than in (a).
Moreover, we observe that increasing the temperature makes neural networks more robust, while maintaining a test accuracy that is comparable to the original model. This corroborates previous findings on non-medical image data [13]. Defensive distillation also has the potential to improve testing accuracy [13]: This phenomenon is more obvious in Fig. 5c, in which all distilled networks outperform the original model when is equal to 0. The main drawback of using a larger temperature is slower convergence during training, leading to a higher computational workload. This may impose practical constraints on the magnitude of that can be used while training.
Similar plots in Fig. 6 can be obtained for the 2D-UNet. Although the 2D-UNet has a different architecture, it is also sensitive to subtle perturbations, and the distillation method can also improve robustness. It is notable the 3D-UNet outperformed the 2D-UNet, potentially due to its ability to utilize the similarity between continuous axial slices using 3D convolutions, whereas the 2D-UNet deals with individual slices during both the training and prediction processes.
Fig. 6.
Performance of distillation in the 2D-UNet. Dice coefficients of a “Whole Tumor,” b “Tumor Core,” and c “Enhancing Tumor” vs. FGSM with different
Adversarial Training
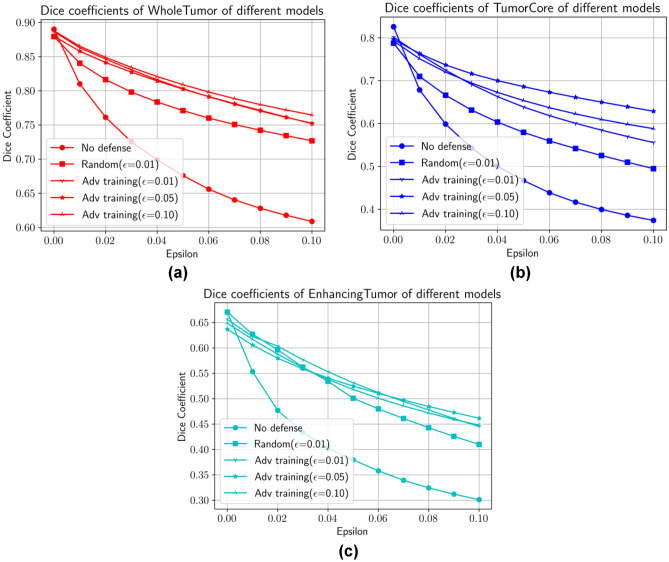
We use the same hyperparameters as in the distillation method. Figure 7 shows the Dice coefficients of different models by using adversarial training with different values of . For all categories, adversarial training is seen to enhance the robustness of neural networks. For comparison, we also plot the curve (marked with stars) corresponding to data augmentation with random perturbations of radius of 0.01 in infinity norm [24–26]. This leads to better robustness than the original neural network; however, the starred curve lies below all other curves, indicating that more sophisticated defenses will make the trained networks more robust.
Fig. 7.
Performance of adversarial training in the 3D-UNet. Dice coefficients of a “Whole Tumor,” b “Tumor Core,” and c “Enhancing Tumor” vs. FGSM with different
Similar to defensive distillation, different values of used in adversarial training only have moderate effects on the test accuracy, which may be seen by comparing the curves in each category when is 0. However, when we evaluate the performance of each model across all categories, the increase of in the training process does not ensure improved robustness. Moreover, the training process may diverge for large values of , making the choice of crucial. Similar observations can be found in Fig. 8 for the 2D-UNet.
Fig. 8.
Performance of adversarial training in the 2D-UNet. Dice coefficients of a “Whole Tumor,” b “Tumor Core,” and c “Enhancing Tumor” vs. FGSM with different
Figure 9 shows an example of adversarial images of different 3D UNet models and the corresponding predicted labels. The leftmost column contains the original image and its true label. Note that all models perform well on the unperturbed images, since the Dice coefficient for label = 4 (enhancing tumor) is around 0.70. Next, we apply FGSM with to generate adversarial images, which are shown in the middle row.
Fig. 9.
Top row: true labels and predicted segmentations of each 3D UNet model given the original input image. Middle row: original input image and adversarial examples for each model generated by FGSM with . Bottom row: true labels and predicted images of each model given their corresponding adversarial examples. Models starting from the 2nd row: adversarial training with , adversarial training with , adversarial training with , model with no defense, distillation with , distillation with , distillation with , and distillation with
We can see that the perturbations are nearly imperceptible to the human eye. However, the Dice coefficients in the 5th and 6th columns (model with no defense, and distilled model with ) drop down significantly in the 3rd row, while the others remain almost the same.
A Wilcoxon signed-rank test was used to compare the Dice coefficient to the ground truth data for each attack type (), and a Bonferroni correction was applied to correct for multiple comparisons. A summary of the performance of these models on the testing data set can be found in Tables 2 and 3 for 3D-UNets and 2D-UNets, respectively. Notably, although these defensive models achieve better performance on adversarial examples, they still perform worse than the models applied to unperturbed images.
Table 2.
Results of different 3D-UNet models quantified using the Dice coefficient of label 4 (enhancing tumor) when attacked by FGSM with equal to 0, 0.05, and 0.1. An asterisk (*) indicates statistically significant differences at , corrected for multiple comparisons
| Segmentation type | Defense type | Dice coefficient | Dice coefficient difference P-value | ||||
|---|---|---|---|---|---|---|---|
| Whole tumor | No defense | 0.890 ± 0.052 | 0.676 ± 0.189 | 0.609 ± 0.216 | - | - | - |
| Distillation (T = 20) | 0.893 ± 0.053 | 0.741 ± 0.162* | 0.686 ± 0.180* | 0.2287 | 6.628e-4 | 0.0013 | |
| Distillation (T = 100) | 0.891 ± 0.057 | 0.807 ± 0.123* | 0.753 ± 0.149* | 0.3714 | 9.011e-9 | 9.970e-8 | |
| Distillation (T = 500) | 0.896 ± 0.056* | 0.814 ± 0.153* | 0.759 ± 0.174* | 0.0224 | 1.729e-8 | 3.916e-7 | |
| Distillation (T = 5000) | 0.885 ± 0.080 | 0.809 ± 0.161* | 0.758 ± 0.182* | 0.8706 | 4.296e-8 | 1.678e-7 | |
| Adversarial training—0.01 | 0.887 ± 0.075 | 0.803 ± 0.136* | 0.752 ± 0.158* | 0.5646 | 2.176e-8 | 3.755e-7 | |
| Adversarial training—0.05 | 0.880 ± 0.061* | 0.803 ± 0.107* | 0.752 ± 0.124* | 0.0457 | 1.991e-7 | 1.007e-6 | |
| Adversarial training—0.1 | 0.888 ± 0.053 | 0.809 ± 0.128* | 0.765 ± 0.143* | 0.3464 | 6.176e-9 | 7.119e-9 | |
| Tumor core | No defense | 0.826 ± 0.142 | 0.467 ± 0.242 | 0.374 ± 0.233 | - | - | - |
| Distillation (T = 20) | 0.820 ± 0.174 | 0.603 ± 0.266* | 0.521 ± 0.256* | 0.2137 | 1.982e-6 | 1.950e-5 | |
| Distillation (T = 100) | 0.801 ± 0.174* | 0.633 ± 0.259* | 0.543 ± 0.268* | 0.0197 | 2.411e-6 | 9.801e-6 | |
| Distillation (T = 500) | 0.823 ± 0.165 | 0.661 ± 0.279* | 0.574 ± 0.289* | 0.9968 | 1.828e-7 | 7.289e-6 | |
| Distillation (T = 5000) | 0.810 ± 0.175 | 0.713 ± 0.236* | 0.640 ± 0.247* | 0.5486 | 3.158e-9 | 3.478e-9 | |
| Adversarial training—0.01 | 0.801 ± 0.181 | 0.639 ± 0.244* | 0.556 ± 0.255* | 0.2137 | 4.699e-8 | 1.334e-6 | |
| Adversarial training—0.05 | 0.796 ± 0.171* | 0.686 ± 0.230* | 0.629 ± 0.237* | 0.0019 | 4.216e-9 | 7.464e-9 | |
| Adversarial training—0.1 | 0.793 ± 0.165* | 0.654 ± 0.232* | 0.588 ± 0.241* | 6.628e-4 | 2.478e-9 | 1.250e-8 | |
| Enhancing tumor | No defense | 0.670 ± 0.295 | 0.380 ± 0.267 | 0.301 ± 0.242 | - | - | - |
| Distillation (T = 20) | 0.672 ± 0.286 | 0.506 ± 0.274* | 0.425 ± 0.261* | 0.0902 | 1.062e-6 | 5.837e-6 | |
| Distillation (T = 100) | 0.693 ± 0.238* | 0.517 ± 0.289* | 0.425 ± 0.283* | 0.0016 | 0.0012 | 0.0035 | |
| Distillation (T = 500) | 0.704 ± 0.261 | 0.561 ± 0.298* | 0.471 ± 0.294* | 0.3224 | 7.785e-8 | 3.351e-6 | |
| Distillation (T = 5000) | 0.660 ± 0.286 | 0.583 ± 0.285* | 0.522 ± 0.275* | 0.1436 | 2.345e-8 | 1.068e-8 | |
| Adversarial training—0.01 | 0.637 ± 0.289 | 0.525 ± 0.282* | 0.462 ± 0.269* | 0.6259 | 4.602e-8 | 3.648e-6 | |
| Adversarial training—0.05 | 0.656 ± 0.304* | 0.531 ± 0.283* | 0.445 ± 0.266* | 2.979e-5 | 1.155e-9 | 2.365e-9 | |
| Adversarial training—0.1 | 0.649 ± 0.283* | 0.518 ± 0.283* | 0.448 ± 0.269* | 0.0057 | 5.064e-9 | 7.594e-8 | |
Table 3.
Results of different 2D-UNet models quantified using the Dice coefficient of label 4 (enhancing tumor) when attacked by FGSM with equal to 0, 0.05, and 0.1. An asterisk (*) indicates statistically significant differences at , corrected for multiple comparisons
| Segmentation type | Defense type | Dice coefficient | Dice coefficient difference P-value | ||||
|---|---|---|---|---|---|---|---|
| Whole tumor | No defense | 0.804 ± 0.131 | 0.607 ± 0.226 | 0.549 ± 0.236 | - | - | - |
| Distillation (T = 20) | 0.831 ± 0.110 | 0.726 ± 0.185* | 0.686 ± 0.201* | 0.3568 | 0.0073 | 0.0040 | |
| Distillation (T = 100) | 0.814 ± 0.132 | 0.718 ± 0.196* | 0.684 ± 0.207* | 0.5371 | 0.0132 | 0.0067 | |
| Distillation (T = 500) | 0.828 ± 0.113 | 0.731 ± 0.180* | 0.689 ± 0.198* | 0.3385 | 0.0057 | 0.0039 | |
| Distillation (T = 5000) | 0.818 ± 0.130 | 0.760 ± 0.158* | 0.729 ± 0.171* | 0.5609 | 5.014e-4 | 1.844e-4 | |
| Adversarial training—0.01 | 0.816 ± 0.135 | 0.748 ± 0.168* | 0.709 ± 0.190* | 0.4259 | 0.0015 | 7.943e-4 | |
| Adversarial training—0.05 | 0.809 ± 0.127 | 0.757 ± 0.151* | 0.735 ± 0.164* | 0.8300 | 0.0011 | 1.657e-4 | |
| Adversarial training—0.1 | 0.805 ± 0.131 | 0.749 ± 0.160* | 0.726 ± 0.173* | 0.9929 | 0.0014 | 3.701e-4 | |
| Tumor core | No defense | 0.701 ± 0.167 | 0.481 ± 0.231 | 0.428 ± 0.231 | - | - | - |
| Distillation (T = 20) | 0.723 ± 0.156 | 0.602 ± 0.207* | 0.557 ± 0.214* | 0.5914 | 0.0153 | 0.0113 | |
| Distillation (T = 100) | 0.716 ± 0.161 | 0.611 ± 0.204* | 0.572 ± 0.211* | 0.7071 | 0.0085 | 0.0043 | |
| Distillation (T = 500) | 0.706 ± 0.176 | 0.613 ± 0.205* | 0.573 ± 0.209* | 0.7678 | 0.0102 | 0.0048 | |
| Distillation (T = 5000) | 0.690 ± 0.183 | 0.631 ± 0.190* | 0.600 ± 0.192* | 0.8300 | 0.0053 | 0.0011 | |
| Adversarial training—0.01 | 0.707 ± 0.180 | 0.625 ± 0.200* | 0.584 ± 0.210* | 0.7815 | 0.0057 | 0.0026 | |
| Adversarial training—0.05 | 0.695 ± 0.176 | 0.623 ± 0.196* | 0.601 ± 0.200* | 0.9216 | 0.0067 | 9.328e-4 | |
| Adversarial training—0.1 | 0.695 ± 0.173 | 0.621 ± 0.192* | 0.593 ± 0.199* | 0.8933 | 0.0079 | 0.0012 | |
| Enhancing tumor | No defense | 0.629 ± 0.278 | 0.444 ± 0.250 | 0.376 ± 0.233 | - | - | - |
| Distillation (T = 20) | 0.649 ± 0.266 | 0.535 ± 0.268* | 0.481 ± 0.259* | 0.8022 | 0.0432 | 0.0259 | |
| Distillation (T = 100) | 0.646 ± 0.276 | 0.554 ± 0.275* | 0.500 ± 0.266* | 0.7406 | 0.0178 | 0.0126 | |
| Distillation (T = 500) | 0.580 ± 0.283 | 0.509 ± 0.260 | 0.460 ± 0.249* | 0.2137 | 0.1329 | 0.0750 | |
| Distillation (T = 5000) | 0.567 ± 0.275 | 0.507 ± 0.259 | 0.457 ± 0.248 | 0.0961 | 0.1423 | 0.0892 | |
| Adversarial training—0.01 | 0.620 ± 0.279 | 0.483 ± 0.278 | 0.421 ± 0.276 | 0.7406 | 0.2448 | 0.2377 | |
| Adversarial training—0.05 | 0.611 ± 0.277 | 0.488 ± 0.269 | 0.430 ± 0.266 | 0.5609 | 0.2137 | 0.1739 | |
| Adversarial training—0.1 | 0.593 ± 0.285 | 0.488 ± 0.266 | 0.432 ± 0.258 | 0.3164 | 0.2341 | 0.1977 | |
Distillation vs. Adversarial Training
Based on the results in Tables 2 and 3, for enhancing tumor segmentation, the defensive distillation method yields more robust performance than adversarial training for . However, it is not necessarily true that defensive distillation will always outperform adversarial training in terms of a one-step attack. With a more careful choice of , the performance of adversarial training may exceed that of distillation; however, it may be more difficult to find the optimal choice of , compared to tuning the temperature to obtain better performance.
Discussion
We have demonstrated the vulnerability of deep learning algorithms for image segmentation tasks to adversarial perturbations. Adversarial attacks create imperceptible visual differences to the input data, yet have profound effects on the segmented output. Furthermore, we have developed methods for easily constructing adversarial perturbations using generalizations of FGSM, and have similarly studied defense mechanisms based on distillation and adversarial training. We have illustrated the effectiveness of our methods on magnetic resonance images from the BraTS data set.
In this work, we have mainly focused on one-step adversarial attacks that are visually imperceptible. Integrating more sophisticated adversaries during training is likely to make the networks more robust, and constitutes part of our future work. Furthermore, recent work [11] shows that adversarial training may result in label “leak” if the original task is difficult, such as classification tasks on the ImageNet data set. Label leak occurs when a model is trained using adversarial attacks generated by FGSM and again evaluated using images with FGSM perturbations, producing higher accuracy on adversarial examples than on clean images. A potential explanation is that the gradient added to the original image in adversarial training contains extra information from the label, making classification of adversarial examples easier if a neural network uses that information. We plan to investigate whether label leak also occurs for segmentation and classification tasks in medical imaging. Lastly, we plan to evaluate the effectiveness of different defense techniques beyond standard white-box attacks on the trained model. For instance, we are interested in examining whether a defense strategy is effective against black-box adversarial examples or transferred adversarial attacks [28, 29].
This work is not without limitations. First, we have focused on two basic UNets for medical imaging segmentation, a 3D-UNet model and a 2D-UNet. It would be interesting to see if other network structures also lead to similar trends with respect to adversarial attacks and defenses—perhaps specific network structures could be designed to increase robustness to certain types of attacks. However, both UNet models have been widely applied and studied for many medical imaging segmentation problems, and are thus useful baselines for comparison. Second, we have mainly studied adversarial attacks based on FGSM, since they generate adversarial perturbations in a fast, simple way. However, one could similarly adapt other attack methods such as Deepfool [30], JSMA [31], and DAG [32] from classification to segmentation tasks. These methods could lead to more effective attacks, particularly for targeted attack strategies, where our results show that iterative FGSM is relatively ineffective. Third, the perturbations we have constructed may not correspond to natural variation in medical images. The study of physics-based perturbations that may be more prevalent in MR images (e.g., motion or other types of image artifacts) will be important to study in future work. Other types of contamination that might feasibly arise include random noise in training or testing images, or incorrect labels that are introduced in a random or adversarial manner. Although we hypothesize that the defense strategies proposed in this paper may also be more robust with respect to such perturbations, their efficacy based on this study is unclear. In practice, it may be necessary to devise other defense strategies that are specific to these types of perturbations. However, this work shows that deep learning segmentation networks applied to medical imaging are susceptible to visually subtle attacks, suggesting that they could be prone to intentional manipulation.
With respect to computational complexity, adversarial training needs to perform forward and backward propagation twice for each batch of data, compared to three forward and two backward processes required for defensive distillation. Therefore, adversarial training is less computationally complex given the same configuration. Moreover, higher values of in distillation require more iterations for convergence, leading to higher computational costs during the training process. Furthermore, adversarial training is generally more interpretable than defensive distillation: we can check that the perturbed images generated during the training process should indeed be segmented in the same way as the unperturbed images, provided the radius of perturbation is sufficiently small. This provides a natural way to bound the magnitude of , whereas it is more difficult to determine the “right” magnitude of to use without cross-validating the distilled model on test data.
Conclusion
In summary, we have shown that segmentation networks can be adversely affected by the use of targeted attacks which utilize visually minor (and potentially undetectable) modifications to existing images. By adding a small perturbation calculated by FGSM to the input MR image of a patient, normal tissue can be regarded as a tumor by the network. With increased interest in applying deep learning techniques to medical imaging data, it is important to understand the ramifications of adversarial inputs (either intentional or unintentional), as these tools may be used in clinical decision-making. We have demonstrated that defensive techniques such as distillation and adversarial training can help combat one-step perturbations added to MR images. As the temperature grows, robustness increases at the cost of computational complexity. Therefore, future studies of how deep learning networks could be both unintentionally (e.g., as a result of artifacts or operator error) or intentionally (e.g., by a bad actor) tricked into misclassifying or mislabeling medical images is a critically important consideration as deep learning approaches move toward routine clinical utilization.
Acknowledgements
Research reported in this publication was supported by the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health under award number R01LM013151, as well as the National Science Foundation under award number DMS-1749857.
Declarations
Conflict of Interest
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.de Bruijne M. Machine learning approaches in medical image analysis: From detection to diagnosis. Medical Image Analysis. 2016;33:94–97. doi: 10.1016/j.media.2016.06.032. [DOI] [PubMed] [Google Scholar]
- 2.Wang S, Summers RM. Machine learning and radiology. Medical Image Analysis. 2012;16:933–951. doi: 10.1016/j.media.2012.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wernick, M., Yang, Y., Brankov, J., Yourganov, G., Strother, S., 2010. Machine Learning in Medical Imaging. IEEE signal processing magazine 27, 25–38. URL: http://ieeexplore.ieee.org/document/5484160/, 10.1109/MSP.2010.936730. [DOI] [PMC free article] [PubMed]
- 4.Lecun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–444. doi: 10.1038/nature14539. [DOI] [PubMed] [Google Scholar]
- 5.Lipton, Z.C., 2016. The Mythos of Model Interpretability, in: arXiv preprint. URL: http://arxiv.org/abs/1606.03490 , arXiv:1606.03490.
- 6.Montavon G, Samek W, Müller KR. Methods for interpreting and understanding deep neural networks. Digital Signal Processing: A Review Journal. 2018;73:1–15. doi: 10.1016/j.dsp.2017.10.011. [DOI] [Google Scholar]
- 7.Goodfellow, I.J., Shlens, J., Szegedy, C., 2014. Explaining and Harnessing Adversarial Examples, in: arXiv preprint. URL: http://arxiv.org/abs/1412.6572 , arXiv:1412.6572.
- 8.Finlayson, S.G., Chung, H.W., Kohane, I.S., Beam, A.L., 2018. Adversarial Attacks Against Medical Deep Learning Systems, in: arXiv preprint. URL: http://arxiv.org/abs/1804.05296 , arXiv:1804.05296.
- 9.Paschali, M., Conjeti, S., Navarro, F., Navab, N., 2018. Generalizability vs. robustness: Investigating medical imaging networks using adversarial examples, in: Proceedings of the Medical Image Computing and Computer-Assisted Intervention, Springer Verlag. pp. 493–501. 10.1007/978-3-030-00928-1_56.
- 10.Ozbulak, U., Van Messem, A., De Neve, W., 2019. Impact of Adversarial Examples on Deep Learning Models for Biomedical Image Segmentation, in: Proceedings of the Medical Image Computing and Computer- Assisted Intervention. URL: http://arxiv.org/abs/1907.13124 , arXiv:1907.13124.
- 11.Kurakin, A., Goodfellow, I., Bengio, S., 2016. Adversarial Machine Learning at Scale, in: International Conference on Learning Representations. URL: http://arxiv.org/abs/1611.01236 , arXiv:1611.01236.
- 12.Madry, A., Makelov, A., Schmidt, L., Tsipras, D., Vladu, A., 2017. Towards Deep Learning Models Resistant to Adversarial Attacks, in: International Conference on Learning Representations. URL: http://arxiv.org/abs/1706.06083 , arXiv:1706.06083.
- 13.Papernot, N., McDaniel, P., Wu, X., Jha, S., Swami, A., 2016b. Distillation as a Defense to Adversarial Perturbations Against Deep Neural Networks, in: Proceedings of the IEEE Symposium on Security and Privacy, Institute of Electrical and Electronics Engineers Inc. pp. 582–597. 10.1109/SP.2016.41.
- 14.Akhtar N, Mian A. Threat of Adversarial Attacks on Deep Learning in Computer Vision: A Survey. IEEE Access. 2018;6:14410–14430. doi: 10.1109/ACCESS.2018.2807385. [DOI] [Google Scholar]
- 15.Bakas, S., 2017. Multimodal Brain Tumor Segmentation Challenge. URL: https://www.med.upenn.edu/sbia/brats2017/data.html .
- 16.Bakas, S., Akbari, H., Sotiras, A., Bilello, M., Rozycki, M., Kirby, J.S., Freymann, J.B., Farahani, K., Davatzikos, C., 2017a. Advancing The Cancer Genome Atlas glioma MRI collections with expert seg- mentation labels and radiomic features. Scientific data 4, 170117. URL: http://www.ncbi.nlm.nih.gov/pubmed/28872634, 10.1038/sdata.2017.117. [DOI] [PMC free article] [PubMed]
- 17.Bakas, S., Akbari, H., Sotiras, A., Bilello, M., Rozycki, M., Kirby, J.S., Freymann, J.B., Farahani, K., Davatzikos, C., 2017b. Segmentation Labels and Radiomic Features for the Pre-operative Scans of the TCGA-GBM col- lection. URL: 10.7937/K9/TCIA.2017.KLXWJJ1Q , 10.7937/K9/TCIA.2017.KLXWJJ1Q. [DOI] [PMC free article] [PubMed]
- 18.Bakas, S., Akbari, H., Sotiras, A., Bilello, M., Rozycki, M., Kirby, J.S., Freymann, J.B., Farahani, K., Davatzikos, C., 2017c. Segmentation Labels and Radiomic Features for the Pre-operative Scans of the TCGA-LGG collection [Data Set]. The Cancer Imaging Archive. 10.7937/K9/TCIA.2017.GJQ7R0EF. [DOI] [PMC free article] [PubMed]
- 19.Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, Burren Y, Porz N, Slotboom J, Wiest R, Lanczi L, Gerstner E, Weber MA, Arbel T, Avants BB, Ayache N, Buendia P, Collins DL, Cordier N, Corso JJ, Criminisi A, Das T, Delingette H, Demiralp C, Durst CR, Dojat M, Doyle S, Festa J, Forbes F, Geremia E, Glocker B, Golland P, Guo X, Hamamci A, Iftekharuddin KM, Jena R, John NM, Konukoglu E, Lashkari D, Mariz JA, Meier R, Pereira S, Precup D, Price SJ, Raviv TR, Reza SM, Ryan M, Sarikaya D, Schwartz L, Shin HC, Shotton J, Silva CA, Sousa N, Subbanna NK, Szekely G, Taylor TJ, Thomas OM, Tustison NJ, Unal G, Vasseur F, Wintermark M, Ye DH, Zhao L, Zhao B, Zikic D, Prastawa M, Reyes M, Van Leemput K. The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS) IEEE Transactions on Medical Imaging. 2015;34:1993–2024. doi: 10.1109/TMI.2014.2377694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Çiçek, Ö., Abdulkadir, A., Lienkamp, S.S., Brox, T., Ronneberger, O., 2016. 3D U-net: Learning dense volumetric segmentation from sparse annotation, in: Proceedings of the Medical Image Computing and Computer- Assisted Intervention, Springer Verlag. pp. 424–432. 10.1007/978-3-319-46723-8_49.
- 21.Ronneberger, O., Fischer, P. and Brox, T., 2015, October. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical image computing and computer-assisted intervention (pp. 234–241). Springer, Cham.
- 22.He, K., Zhang, X., Ren, S., Sun, J., 2016. Deep residual learning for image recognition, in: Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, IEEE Computer Society. pp. 770–778. 10.1109/CVPR.2016.90.
- 23.Zou KH, Warfield SK, Bharatha A, Tempany CM, Kaus MR, Haker SJ, Wells WM, Jolesz FA, Kikinis R. Statistical Validation of Image Segmentation Quality Based on a Spatial Overlap Index. Academic Radiology. 2004;11:178–189. doi: 10.1016/S1076-6332(03)00671-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Carlini, N., Wagner, D., 2017. MagNet and “Efficient Defenses Against Adversarial Attacks” are Not Robust to Adversarial Examples, in: arXiv preprint. URL: https://github.com/carlini/MagNet , arXiv:1711.08478v1.
- 25.Engstrom, L., Tran, B., Tsipras, D., Schmidt, L., Madry, A., 2019. Exploring the Landscape of Spatial Robustness. Proceedings of Machine Learning Research 97, 1802–1811. URL: http://arxiv.org/abs/1712.02779 , arXiv:1712.02779.
- 26.Zantedeschi, V., Nicolae, M.I., Rawat, A., 2017. Efficient defenses against adversarial attacks, in: Proceedings of the 10th ACM Workshop on Artificial Intelligence and Security, co-located with CCS 2017, Association for Computing Machinery, Inc. pp. 39–49. 10.1145/3128572.3140449.
- 27.Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, Gee JC. N4ITK: Improved N3 bias correction. IEEE Transactions on Medical Imaging. 2010;29:1310–1320. doi: 10.1109/TMI.2010.2046908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liu, Y., Chen, X., Liu, C., Song, D., 2016. Delving into Transferable Adversarial Examples and Black-box Attacks, in: International Conference on Learning Representations. URL: http://arxiv.org/abs/1611.02770 , arXiv:1611.02770.
- 29.Papernot, N., McDaniel, P., Goodfellow, I., Jha, S., Celik, Z.B., Swami, A., 2017. Practical black-box attacks against machine learning, in: Proceedings of the 2017 ACM Asia Conference on Computer and Communications Security, Association for Computing Machinery, Inc. pp. 506–519. 10.1145/3052973.3053009.
- 30.Moosavi-Dezfooli, S.M., Fawzi, A., Frossard, P., 2016. DeepFool: A Simple and Accurate Method to Fool Deep Neural Networks, in: Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, IEEE Computer Society. pp. 2574–2582. 10.1109/CVPR.2016.282.
- 31.Papernot, N., Mcdaniel, P., Jha, S., Fredrikson, M., Celik, Z.B., Swami, A., 2016a. The limitations of deep learning in adversarial settings, in: Proceedings of the IEEE European Symposium on Security and Privacy, EURO S and P 2016, Institute of Electrical and Electronics Engineers Inc. pp. 372– 387. 10.1109/EuroSP.2016.36.
- 32.Xie, C., Wang, J., Zhang, Z., Zhou, Y., Xie, L., Yuille, A., 2017. Adversarial Examples for Semantic Segmentation and Object Detection, in: Proceedings of the IEEE International Conference on Computer Vision, Institute of Electrical and Electronics Engineers Inc. pp. 1378–1387. 10.1109/ICCV.2017.153.