Abstract
The Puma lineage within the family Felidae consists of 3 species that last shared a common ancestor around 4.9 million years ago. Whole-genome sequences of 2 species from the lineage were previously reported: the cheetah (Acinonyx jubatus) and the mountain lion (Puma concolor). The present report describes a whole-genome assembly of the remaining species, the jaguarundi (Puma yagouaroundi). We sequenced the genome of a male jaguarundi with 10X Genomics linked reads and assembled the whole-genome sequence. The assembled genome contains a series of scaffolds that reach the length of chromosome arms and is similar in scaffold contiguity to the genome assemblies of cheetah and puma, with a contig N50 = 100.2 kbp and a scaffold N50 = 49.27 Mbp. We assessed the assembled sequence of the jaguarundi genome using BUSCO, aligned reads of the sequenced individual and another published female jaguarundi to the assembled genome, annotated protein-coding genes, repeats, genomic variants and their effects with respect to the protein-coding genes, and analyzed differences of the 2 jaguarundis from the reference mitochondrial genome. The jaguarundi genome assembly and its annotation were compared in quality, variants, and features to the previously reported genome assemblies of puma and cheetah. Computational analyzes used in the study were implemented in transparent and reproducible way to allow their further reuse and modification.
Keywords: Puma yagouaroundi, 10X Genomics Chromium, Felidae, whole genome assembly, genome annotation, linked reads
The jaguarundi, Puma yagouaroundi, is one of the 41 species that constitute the Felidae, one of the 16 families of the mammalian order Carnivora (Jackson and Nowell 1996; Sunquist and Sunquist 2009; Kitchener et al. 2017). The species ranges from the southwestern United States, through Central America and much of South America, reaching as far as northern Argentina. Within this range, jaguarundis inhabit a wide variety of habitats, from semi-arid and grassland areas to dense dry and wet forests (de Oliveira 1998; Espinosa et al. 2017). Jaguarundis display differences in craniodental morphology as well as 2 primary pelage colors (gray or dark versus reddish) across these heterogeneous habitats, which has led to the definition of as many as 8 subspecies (Allen 1919; Cabrera 1958). Ecological modeling of the pelage variants has shown that they are significantly associated with particular habitats, with gray or dark morphs occurring at a high frequency in moist and dense forests whereas reddish morphs are more common in dry, open habitats such as deserts (da Silva et al. 2016). However, range-wide analyses of partial and whole mitochondrial genomes showed no association between haplotype clusters and subspecies (Ruiz-García and Pinedo-Castro 2013; Ruiz-García et al. 2018). On the basis of these findings, the Cat Classification Task Force of the IUCN Cat Specialist Group concluded that the jaguarundi represents a monotypic species (Kitchener et al. 2017).
Historically, the jaguarundi was included in the genus Herpailurus, but recently, phylogenetic and phylogenomic studies have positioned it among the Puma lineage of the Felidae, which also includes the mountain lion (also called cougar or puma), Puma concolor, and the cheetah, Acinonyx jubatus (Johnson et al. 2006; O’Brien and Johnson 2007; O’Brien et al. 2008; Li et al. 2016). These molecular studies have firmly placed jaguarundis as sister to the mountain lion and dating analyses have suggested that the 2 species diverged approximately 4–7 million years ago. Members of the Puma lineage demonstrate contrasting morphologies and unique adaptations to lifestyle. In striking contrast to mountain lions, which have an average body weight ranging between 34 and 72 kg, jaguarundis only average around 3–7 kg (Sunquist and Sunquist 2009).
Karyotypic analysis of the jaguarundi shows a diploid chromosome complement of 2n = 38, which corresponds to the mostly conserved diploid number found in other species of the Felidae (Wurster-Hill and Gray 1973; Eroğlu 2017; Graphodatsky et al. 2020). To date, de novo genome assemblies have been generated for more than a dozen felid species (NCBI:txid 9681), including the cheetah and mountain lion (Dobrynin et al. 2015; Saremi et al. 2019). Here, we present the first de novo draft assembly of a male jaguarundi, which will facilitate further research on the adaptations and other biological features of this species in relation to other members of the Puma lineage and Felidae.
Methods
Biological Materials
A tissue biopsy of a breeding male jaguarundi originally from Mexico and housed at the Rotterdam Zoo, Netherlands, was collected by Mitchell Bush in 1981. A primary fibroblast cell line (HYA-1) was subsequently initiated from the skin biopsy at the Laboratory of Genomic Diversity, National Cancer Institute (Frederick, MD, United States) and stored frozen there from 1981. Freshly harvested fibroblast cells were utilized for genomic DNA.
Nucleic Acid Library Preparation
Cultured primary fibroblast cells at passage 10 and 80% confluency were harvested by removing growth medium, washing cells twice with DPBS (Dulbecco’s Phosphate Buffered Saline), lifting cells off with 0.25% Trypsin-EDTA, and collecting cells with alpha-MEM with 15% fetal bovine serum (FBS) and centrifuging at 300 g for 10 minutes. The cell pellet was then washed in DPBS and centrifuged at 300 g for 10 minutes. To obtain high molecular weight DNA, lysis was performed by suspending freshly harvested cells in a lysis buffer (0.05 M Tris-HCl, pH 7.5, 0.066 M EDTA, and 0.1 M NaCl) with adding sodium dodecyl sulfate (SDS) to a final concentration of 1% and Proteinase K to a final concentration of 0.75 mg/ml, followed by incubation at 56°Ϲ for 2–4 hours. To remove RNA contamination, RNase A at a final concentration 10 mg/ml was added and the solution was further incubated 1 hour at 56°Ϲ. During incubation, the sample tube was gently rocked several times by hand.
Genomic DNA extraction was performed by the standard phenol-chloroform method followed by precipitation using isopropanol (Sambrook et al. 1989). The precipitated genomic DNA was transferred into 80% ethanol for 24 hours, shipped to Baltimore and then resuspended in 1X TE buffer, pH 8.0 for 4 weeks at 4°Ϲ. Throughout the entire extraction process, the DNA sample was manipulated gently to preserve high molecular weight. Quality and concentration of extracted genomic DNA were measured with a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, United States) and 0.5% agarose gel electrophoresis.
Chromium sequencing libraries were created at the Genetics Resources Core Facility (GRCF), Johns Hopkins University School of Medicine (Baltimore, United States) with Chromium Genome Reagents Kits Version 2, a 10X Genomics Chromium Controller instrument (10X Genomics, United States) and DNA diluted to 1.2 ng/µL. Prior to DNA library construction, DNA optical density was assessed with a Qubit Fluorometer (Thermo Fisher Scientific, United States). Gel Bead-In-Emulsions (GEMs) were used to prepare genomic DNA libraries, which were then nicktranslated using bead-specific unique molecular identifiers (UMIs; Chromium Genome Reagents Kit Version 2 User Guide). Genomic DNA size and concentration were calculated using an Agilent 2100 Bioanalyzer DNA 1000 chip (Agilent Technologies).
DNA Sequencing and Genome Assembly
The Chromium libraries were sequenced at the Genetic Resources Core Facility (GRCF), Johns Hopkins University School of Medicine (Baltimore, United States). The sequencing was performed at 53-fold average sequence coverage on 2 lanes of a single flowcell on the Illumina NovaSeq 6000 (San Diego, United States). Sequenced paired-end reads were assembled into 2 scaffold-level pseudohaplotype sequences using Supernova version 2.1.1 (Weisenfeld et al. 2017).
Scaffolds from the first pseudohaplotype sequence of the Supernova assembly were scanned for duplicates and gap-only scaffolds using the JDGA (Jaguarundi Draft Genome Assembly) package (Tamazian 2021). The filtered scaffolds were submitted to NCBI GenBank and the report by the NCBI Contamination Screen pipeline was received. The report listed scaffolds and scaffold regions recommended for removal from the assembly. After manual review of the scaffolds and the regions, the submitted genome assembly was processed by excluding the reported scaffolds and hard-masking (that is, filling with Ns) the reported regions using the JDGA package. The resulting assembly was accepted to NCBI GenBank (accession GCA_014898765.1) and used in the downstream analysis.
Computational Analysis Framework
The genome assembly and analysis presented in this study involve multiple interconnected computational activities. To ensure transparency and reproducibility of our study, we arranged most of the activities in 2 pipelines implemented in Snakemake (Köster and Rahmann 2012). The first pipeline includes read alignment and variant calling and was launched on a multiprocessor server with 64 CPUs, 512 gigabytes RAM, and a 2 terabyte SSD hard drive. The second pipeline covers analysis of the genome and its annotation and can be launched on a regular laptop.
The Snakemake pipelines contain scripts in Python and R that process the genome sequence and annotation data. The core routines of these scripts are arranged in the JDGA package (Tamazian 2021), which accompanies this paper. R scripts of the Snakemake pipelines use routines from packages GenomicRanges (Lawrence et al. 2013) and tidyverse (Wickham et al. 2019). In the following subsections, we refer explicitly to the JDGA package and other external programs that we used to analyze the data. The external programs and their versions are given in Table 1.
Table 1.
Software and databases used for the assembly and analysis of the jaguarundi genome
| Analysis | Software or Database | Version |
|---|---|---|
| Genome assembly | Supernova | 2.1.1 |
| Assembly assessment | QUAST | 5.0.2 |
| BUSCO | 4.1.4 | |
| OrthoDB’s mammalian_odb10 dataset | Creation date: 2020-09-10, number of BUSCOs: 9226, number of species: 24 | |
| BlobTools2 | 2.3.3 | |
| UpSetR | 1.4.0 | |
| Reference-based repeat annotation | RepeatMasker | 4.0.8 (run with blastp version 2.0MP-WashU) |
| RepBase Update database | 20181026 | |
| Dfam database | Consensus 20181026 | |
| De novo repeat annotation | WindowMasker | 1.0.0 (from package blast 2.2.31) |
| Annotation of protein-coding genes | BUSCO | 4.0.4 |
| OrthoDB’s carnivora_odb10 dataset | Creation date: 2019-11-20, number of BUSCOs: 14502, number of species: 12 | |
| BLAT | 35 | |
| AUGUSTUS | 3.3.3 | |
| BEDTools | 2.27.1 | |
| Read mapping | BWA | 0.7.17-r1188 |
| Samtools | 1.11 (using HTSlib 1.11) | |
| Variant calling | FreeBayes | 1.3.4 |
| Variant effect prediction | SnpEff | 5.0 |
| Mitochondrial genome analysis | NCBI BLAST | 2.9.0+ (from package blast 2.9.0) |
| Circos | 0.69–8 | |
| LAST | 1060 | |
| Auxiliary tools | Snakemake | 5.10.0 and 5.32.0 |
| GNU Parallel | 20150122 | |
| R | 3.6.3 | |
| GenomicRanges | 1.38.0 | |
| tidyverse | 1.3.0 |
The citations are given in the text. The table does not list routines from the JDGA package.
Assembly Evaluation
Statistics of the raw jaguarundi reads and the assembled jaguarundi genome sequence were reported by Supernova during the genome assembly process. Features of the filtered jaguarundi genome assembly, which was released on NCBI GenBank, were evaluated using QUAST version 5.0.2 (Gurevich et al. 2013) and the JDGA package. The distributions of the assembly features were visualized by the snail plot produced with the BlobTools2 program version 2.3.3 (Challis et al. 2020). Joint distribution of GC content and length for scaffolds shorter than 1 Mbp was visualized in a separate figure using the JDGA package and the R script from the Snakemake pipeline.
To estimate the completeness of the jaguarundi genome assembly, we launched BUSCO version 4.1.4 (Seppey et al. 2019) in the genome mode with the reference set of single-copy mammalian orthologs (dataset mammalian_odb10; 9,226 genes) from the OrthoDB database (Kriventseva et al. 2019). Based on identified single-copy BUSCO genes, we arranged fragments of the jaguarundi scaffolds in blocks on chromosome sequences of the domestic cat genome assembly Felis_catus_9.0 (Buckley et al. 2020) and visualized the blocks. Each block contained at least 2 consecutive BUSCO single-copy genes, which were positioned on a scaffold in the same order as on a domestic cat chromosome. Summation of the BUSCO results, the block construction and visualization were implemented in the JDGA package.
To compare the jaguarundi genome assembly with the genome assemblies of cheetah and puma, we obtained snail plots and performed the BUSCO analyses for their NCBI RefSeq assemblies (accessions GCF_003709585.1 and GCF_003327715.1) in the same way as for the jaguarundi genome assembly. Joint distribution of the BUSCO genes by categories in the 3 genome assemblies was visualized in an UpSet plot (Lex et al. 2014) produced using the UpSetR package (Conway et al. 2017).
Read Alignment
Reads of the sequenced male jaguarundi (individual HYA-1) were aligned to our NCBI GenBank assembly of the jaguarundi genome (accession GCA_014898765.1) with the added jaguarundi mitochondrial genome assembly (Li et al. 2016) from NCBI RefSeq (accession NC_028311.1). We also aligned whole-genome sequencing reads of a female jaguarundi (individual HJA5) presented in Li, et al. (2019) to the same reference genome. The alignment was performed using the MEM algorithm implemented in BWA version 0.7.17-r1188 (Li 2013). BWA was launched with options -K 100000000 -R -Y, which fixed the number of input bases in a batch, added read group headers to generated alignments, and specified using soft clipping CIGAR operations for supplementary alignments.
Read alignments were converted from the SAM format to the binary BAM format using the view tool from the Samtools package (Li et al. 2009). The alignments were filtered using the Samtools view program launched with options -f3 -F2316, which kept primary alignments of paired reads mapped in proper pairs and removed unmapped reads, alignments of reads without mapped pairs, secondary and supplementary alignments. Duplicate alignments were removed using the Samtools markdup tool launched with option -r. Samtools version 1.11 was used to process the read alignments.
We aligned whole-genome sequencing reads of 2 male cheetahs downloaded from the NCBI RefSeq Sequence Read Archive (SRA) to the NCBI RefSeq assembly of the cheetah genome (accession GCF_003709585.1). For puma, we aligned whole-genome sequencing reads of 2 male individuals to the NCBI RefSeq assembly of the puma genome (accession GCF_003327715.1). The puma reads and genome assembly were obtained from Saremi et al. (2019). Both cheetah and puma genome assemblies included mitochondrial genomes from NCBI RefSeq (accessions NC_005212.1 and NC_016470.1, respectively). Read alignment and postprocessing for cheetah and puma were performed using BWA and Samtools in the same way as for jaguarundi. The list of whole-genome sequencing read datasets with their accessions in the NCBI SRA database is given in Supplementary Table 1.
Genome Annotation
Repeats
The annotation of known genomic repeats in the jaguarundi genome assembly was performed by NCBI using RepeatMasker version 4.0.8 (Smit, Hubley and Green, 2013–2015) with the combined database of RepBase Update version 20181026 (Jurka et al. 2005) and Dfam Consensus (Wheeler et al. 2013). RepeatMasker was launched with options -engine wublast -species “puma yagouaroundi” -s -no_is -cutoff 255, which specified running with the Washington University BLAST as the search engine and using Puma yagouaroundi as the source species of the query sequence in the sensitive mode without checking for bacterial insertion elements and with the cutoff score for masking repeats set to 255.
For database-free annotation of repeats in the jaguarundi genome assembly, we launched WindowMasker version 1.0.0 (Morgulis et al. 2006) with its default options. We compared repeat annotations for jaguarundi by RepeatMasker and WindowMasker using the R script from the Snakemake pipeline.
RepeatMasker annotations for the cheetah and puma genome assemblies were provided by NCBI. For cheetah, RepeatMasker version 4.0.6 was launched with the RepBase Update version 20150807 database and with options -engine wublast -species “Acinonyx jubatus” -s -no_is -cutoff 255 -frag 20000. For puma, RepeatMasker version 4.0.8 was launched with the same database as for jaguarundi and with options -engine wublast -species “Puma concolor” -s -no_is -cutoff 255 -frag 20000.
Repeat annotations for the jaguarundi, cheetah, and puma genome assemblies were summarized by the R script from the Snakemake pipeline.
Protein-Coding Genes
We annotated protein-coding genes in the jaguarundi genome assembly in 2 steps. First, we launched BUSCO version 4.0.4 (Seppey et al. 2019) in the genome mode with the reference set of carnivore genes (dataset carnivora_odb10; 14,502 genes) from the OrthoDB database (Kriventseva et al. 2019). Complete single-copy genes identified by BUSCO formed the first part of the annotated gene set.
Second, regions of the genes identified in the first step were hard-masked in the jaguarundi assembly scaffolds. CDS sequences from the domestic cat genome assembly Felis_catus_9.0 (Buckley et al. 2020) were aligned to the hard-masked scaffolds using BLAT version 35 (Kent 2002). The obtained alignments and the RepeatMasker repeats were passed to the AUGUSTUS gene prediction tool version 3.3.3 (Stanke et al. 2006) as hints for reference-free gene prediction. AUGUSTUS was launched in the parallel mode using the GNU Parallel program (Tange 2011).
Regions of genes predicted by AUGUSTUS were intersected with the BUSCO-identified genes using BEDTools version 2.27.1 (Quinlan and Hall 2010). We then formed a second part of the annotated gene set from the AUGUSTUS-predicted genes that did not overlap with any BUSCO-predicted genes.
Chromosome X Scaffolds
We identified scaffolds from the jaguarundi X chromosome using blocks of X-linked BUSCO genes. We assumed that a scaffold belonged to the X chromosome if more than half of it was covered by the blocks located on domestic cat X chromosome. The procedure to identify the scaffolds was implemented in the R script from the Snakemake pipeline.
Genomic Variants
Genomic variants were obtained from the filtered read alignments using FreeBayes version 1.3.4 (Garrison and Marth 2012). Option--standard-filters, which enabled stringent filters for base and mapping qualities, was specified for the FreeBayes genotyping. FreeBayes was launched in the parallel mode using the wrapper from the JDGA package, which split a reference genome into segments at gaps of 10 kbp or longer and filtered out variants with quality less than 30. Haploid variants were called in X chromosome scaffolds for the male jaguarundi (HYA-1) and on the mitochondrial genome for both jaguarundis (HYA-1 and HJA5). Both jaguarundis were genotyped separately from each other. Genotypes of genomic variants from the 2 jaguarundis were compared to each other using routines from the JDGA package and the R script from the Snakemake pipeline.
Effects of the identified genomic variants were predicted with respect to the annotated protein-coding genes using SnpEff version 5.0 (Cingolani et al. 2012). SnpEff was launched with options -no-downstream -no-intergenic -no-intron -no-upstream -no-utr, which disabled reporting most of the effects for variants outside coding regions.
Genomic variants for the 2 cheetah individuals and the 2 puma individuals were called in the same way as for the male jaguarundi. For jaguarundi and cheetah, genome sizes were estimated by multiplying the total numbers of biallelic heterozygous diploid SNPs and biallelic alternative haploid SNPs with the mean distances between heterozygous SNPs reported by Supernova. SNPs from the individuals used to generate the genome assemblies of the jaguarundi and cheetah were used for this analysis. To estimate genome-wide heterozygosity, we excluded the mitochondrial genome and scaffolds from the X chromosome. Heterozygosity was estimated as the ratio of the number of biallelic heterozygous SNPs in the selected scaffolds and the total size of these scaffolds.
Statistics of the identified genomic variants, their predicted effects, estimated heterozygosity and transition-transversion ratios were obtained using the JDGA package and the R scripts from the Snakemake pipeline.
Mitochondrial Genome Analysis
We estimated coverage of the jaguarundi mitochondrial genome by sequenced reads of the 2 jaguarundis using their filtered read alignments. The coverage was calculated in non-overlapping 100 bp windows using the bedcov tool from Samtools (Li et al. 2009). The window coordinates were produced by the makewindows tool from BEDTools (Quinlan and Hall 2010). The read coverage for both jaguarundis was visualized in the circular plot produced by Circos version 0.69–8 (Krzywinski et al. 2009). The Circos plot also included the mitochondrial genome annotation obtained from NCBI RefSeq, the putative insertion-deletion variants for the 2 jaguarundis, and regions of the mitochondrial genome investigated in previous studies (Ruiz-García and Pinedo-Castro 2013; Ruiz-García et al. 2018). These mitochondrial genome regions were obtained by aligning sequences from NCBI GenBank to the jaguarundi mitochondrial genome using the LAST alignment program version 1060 (Kiełbasa et al. 2011).
Results
For the male jaguarundi HYA-1, the Illumina NovaSeq 6000 produced 918.05 million barcoded paired-end reads from 2 lanes of a flowcell. Read barcodes identified DNA molecules from the Chromium sequencing library and less than 5% of the reads were not barcoded. The length of an untrimmed single read was 151 base pairs (bp). Statistics reported by Supernova for the sequenced reads and the assembled genome sequence are given in Supplementary Table 2.
We identified 521 duplicate scaffolds and 20 scaffolds that consisted of gap characters in the first pseudohaplotype produced by Supernova. The NCBI Contamination Screen reported 7 contamination-derived scaffolds (5 fragments of the jaguarundi mitochondrial genome, a fragment of the human mitochondrial genome, and the phi X 174 bacteriophage genome) and 25 regions of primer or adapter sequences in 21 scaffolds. We excluded the duplicate, gap-only, and contamination-derived scaffolds and hard-masked the primer or adapter sequences with their flanks. The total size of the contamination-derived scaffolds was 37,732 bp and the hard masking changed 1,014 bp in the scaffold sequences. The filtered genome assembly was released on NCBI GenBank and its statistics are given in Supplementary Table 3.
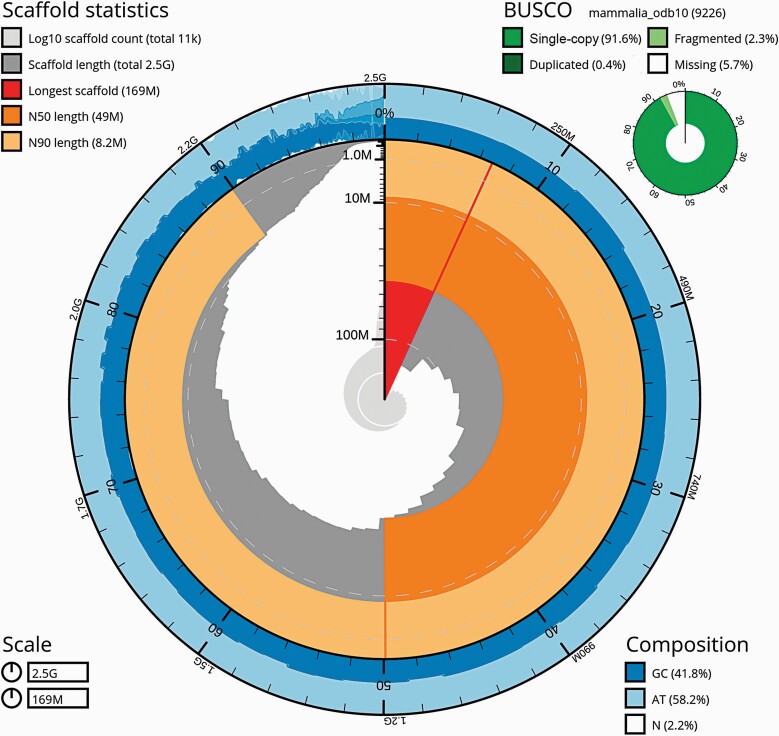
The jaguarundi genome assembly is composed of 10,947 scaffolds, with a total length of 2.47 Gbp and a scaffold N50 of 49.3 Mbp. The scaffold L90 value is 61, which means that 90% of the assembly length is represented by less than 1% of the scaffolds. The distribution of scaffold lengths and GC content is visualized by the snail plot for the whole assembly (Figure 1) and by the scatter plot for scaffolds shorter than 1 Mbp (Supplementary Table 4 and Supplementary Figure 1). The distribution visualized by the snail plot corresponds to a scaffold-level assembly that contains a series of large scaffolds that are chromosome arm in length. The scatter plot shows no bias between length and GC content of the short scaffolds.
Figure 1.
Summary of the jaguarundi genome assembly. The assembly scaffolds are shown by dark-gray bars arranged on the inner side of the circle. The circle radius is the length of the longest scaffold in the assembly. The scaffold bars follow each other clockwise according to the scaffold lengths starting from the longest scaffold, which is represented by the red sector at the center of the circle. The bar widths decrease with the scaffold size resulting in higher density of smaller scaffolds while turning clockwise. The light-gray spiral at the center of the plot shows the cumulative scaffold count for the corresponding point on the circle. Both scaffold lengths (dark gray) and cumulative counts (light gray) are shown on a log scale. The orange circular stripe denotes the area which corresponds to scaffold lengths greater than the N50 scaffold length. The light-orange circular stripe marks the area between the N50 and N90 scaffold lengths. The outer ring of the circle visualizes GC and gap contents of the scaffolds. The wave-like pattern in the outer ring section between 90% and 0% marks is an overplotting artifact caused by multiple small scaffolds corresponding to that region. The summary of OrthoDB’s mammalia_odb10 dataset genes identified in the assembly by BUSCO is given in the upper right corner of the figure. The gap percentage of the assembly and the percentages of GC and AT base pairs in the assembly sequence without gaps are given in the lower right corner of the figure.
More than 91% of mammalian benchmarking universal single-copy orthologs (BUSCOs) were identified in a complete single copy in the assembled jaguarundi genome (Figure 1 and Supplementary Table 5). Blocks formed by consecutive complete single-copy BUSCOs between jaguarundi and domestic cat are shown in Figure 2 and listed in Supplementary Table 6. Scaffolds that contain at least one block constitute more than 96% of the jaguarundi genome assembly.
Figure 2.
Fragments of scaffolds from the jaguarundi genome assembly arranged on domestic cat chromosomes. The fragments are shown by blue blocks with the black outline. Short fragments are shown by black bars without the blue fill. White blocks show domestic cat chromosomes with putative centromere positions as given in the Felis_catus_9.0 assembly of the domestic cat genome. The fragments were formed by consecutive BUSCO genes from OrthoDB’s mammalia_odb10 dataset. The order of the BUSCO genes in the fragments was preserved between jaguarundi scaffolds and domestic cat chromosomes. Each block contains at least 2 BUSCO genes.
Based on the BUSCO blocks, we identified 13 scaffolds that belonged to the X chromosome of the jaguarundi genome. Since the sequenced jaguarundi individual was male, no genomic variants should be present on its X chromosome except for the pseudoautosomal regions. As expected, we observed low numbers of variants on the identified X chromosome scaffolds. The total length of the X chromosome scaffolds and the numbers of variants in them are given in Supplementary Table 7.
RepeatMasker identified 43% of the jaguarundi genome assembly as genomic repeats, which is consistent with the percentage of RepeatMasker repeats in the domestic cat genome (Buckley et al. 2020). Using WindowMasker, we identified 296 Mbp of putative genomic repeats outside the ones annotated by RepeatMasker. Summaries of RepeatMasker and WindowMasker repeats in the assembled jaguarundi genome are given in Supplementary Tables 8 and 9.
We obtained the twofold set of 20,287 annotated protein-coding genes in the jaguarundi genome assembly (Supplementary Table 10). Of these genes, 12,612 were identified using BUSCO and have annotated homologs from the OrthoDB dataset of carnivore single copy orthologs. The remaining 7,675 genes were annotated using the reference-free approach implemented in AUGUSTUS. The total number of the annotated genes is close to the 19,748 protein-coding genes identified in the domestic cat genome assembly (Buckley et al. 2020).
We identified 6.17 million genomic variants, including 35,148 alterations in coding regions, in our draft jaguarundi genome assembly (Supplementary Tables 11 and 12). As expected for the reference genome individual, more than 97.5% of the identified variants were biallelic and heterozygous. In another jaguarundi individual (HJA5) we identified 6.19 million genomic variants, of which 44.3% were biallelic and had the alternative homozygous genotype (Supplementary Table 11). Comparison of genotypes between the 2 jaguarundis at the biallelic heterozygous loci of the reference individual (Supplementary Table 13) confirmed high genetic variability of the species as reported in previous studies (Ruiz-García et al. 2018). Variant effect annotation showed that most effects were synonymous and missense mutations caused by SNPs (Supplementary Table 14). Despite sufficient coverage of the mitochondrial genome by aligned reads, only one putative insertion-deletion variant was identified in the 2 jaguarundis (Supplementary Figure 2). The identified variant was located outside the mitochondrial genome regions analyzed in previous studies (Ruiz-García and Pinedo-Castro 2013; Ruiz-García et al. 2018).
We demonstrated that the numbers of identified genomic variants were consistent with the mean distances between heterozygous SNPs reported by Supernova for jaguarundi and cheetah (Supplementary Table 15). The ranking of the 6 individuals from the 3 species by genome-wide heterozygosity based on the identified genomic variants was consistent with results reported in Dobrynin et al. (2015) and Saremi et al. (2019): the jaguarundi was the most heterozygous species followed by the puma and the cheetah (Supplementary Table 16). Transition-transversion ratios for the 3 species (Supplementary Table 17) were consistent with the reported values for the puma (Ochoa et al. 2019) and the domestic cat (Buckley et al. 2020).
We obtained assembly statistics (Supplementary Figures 3 and 4, Supplementary Table 2), identified BUSCO genes (Supplementary Table 18), constructed BUSCO gene blocks (Supplementary Figures 5 and 6, Supplementary Table 6) and annotated repeats and genomic variants (Supplementary Tables 19–22) for the cheetah and puma genome assemblies. Intersections of the BUSCO gene categories between the 3 genome assemblies are shown in Figure 3.
Figure 3.
Intersections of BUSCO genes from 4 categories: single copy (S), missing (M), fragmented (F), and duplicated (D) in the genome assemblies of jaguarundi (PYA), cheetah (AJU), and puma (PCO).
Discussion
In this paper we present the scaffold-level assembly of the jaguarundi (Puma yagouaroundi) genome. The presented assembly contains a series of chromosome arm scale scaffolds and is consistent with available genome assemblies of other 2 species from the Puma lineage: the cheetah (Acinonyx jubatus) and the mountain lion (Puma concolor). We assessed the assemblies and provide annotated genes, repeats, and genomic variants obtained in a transparent and reproducible way. While the puma genome assembly shows the highest scaffold N50 value, the genome assemblies of the 3 Puma lineage species show similar characteristics that indicate comparable genome assembly levels. Repeat content in the 3 assemblies is relatively conserved and contributes to about 40% of the genome. To demonstrate the high contiguity of the assembled jaguarundi genome, we present BUSCO results and show that features of the annotated genome elements are consistent with each other and with similar features of the cheetah and puma genome assemblies. The estimated heterozygosity among the 3 species is concordant with previous research on the population structure and demographic history of the Puma lineage species (Dobrynin et al. 2015; Saremi et al. 2019).
Adding the first genome assembly of the jaguarundi to the assemblies of the cheetah and puma genomes provides new opportunities for studies of population structure and conservation genomics, research in mammalian evolution and adaptations, and improving gene annotations in the Felidae family. The presented dataset and the related computational framework will be a valuable resource for intra- and interspecific studies in the Puma lineage and the Felidae more generally.
Supplementary Material
Acknowledgments
Authors are grateful to Mitchell Bush for collecting the sample for the cell line. Authors are thankful to Mary Thompson (NCI-Frederick) for establishing the cell line.
Funding
This work was supported by Russian Foundation for Basic Research, RFBR project 20-34-70055. A.K. and A.Z. were supported by the Government of the Russian Federation through the ITMO Fellowship and Professorship Program. DNA extraction and quality control for the jaguarundi sample were funded by a grant from the Russian Science Foundation (No. 19-14-00034 to P.L.P., N.A.S., S.K. and A.S.G.).
Data Availability
We have deposited the primary data underlying the presented analyses as follows:
Raw and filtered read alignments, jaguarundi gene annotation, genomic variants, and auxiliary datasets: Dryad, https://doi.org/10.5061/dryad.wstqjq2mf
The JDGA package: GitLab repository https://gitlab.com/gtamazian/jdga
The Snakemake pipelines: GitLab repository https://gitlab.com/gtamazian/jdga_pipelines
Raw sequence reads: NCBI SRA accession SRX8453956
The jaguarundi genome sequence assembly: NCBI GenBank accession GCA_014898765.1 (PumYag) under NCBI BioSample accession SAMN14930875 and NCBI BioProject accession PRJNA633021 (Puma yagouaroundi).
References
- Allen JA. 1919. Notes on the synonymy and nomenclature of the smaller spotted cats of tropical America. Bull Am Mus Nat Hist. 41:341–419. [Google Scholar]
- Buckley RM, Davis BW, Brashear WA, Farias FHG, Kuroki K, Graves T, Hillier LW, Kremitzki M, Li G, Middleton RP, et al. 2020. A new domestic cat genome assembly based on long sequence reads empowers feline genomic medicine and identifies a novel gene for dwarfism. PLoS Genet. 16:e1008926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cabrera A. 1958. Catalogo de los Mamiferos de America del Sur. Ciencias Zoologicas. 4:1–308. [Google Scholar]
- Challis R, Richards E, Rajan J, Cochrane G, Blaxter M. 2020. BlobToolKit - interactive quality assessment of genome assemblies. G3 (Bethesda). 10:1361–1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM. 2012. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 6:80–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conway JR, Lex A, Gehlenborg N. 2017. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics. 33:2938–2940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva LG, de Oliveira TG, Kasper CB, Cherem JJ, Moraes EA Jr, Paviolo A, Eizirik E. 2016. Biogeography of polymorphic phenotypes: mapping and ecological modelling of coat colour variants in an elusive Neotropical cat, the jaguarundi (Puma yagouaroundi). J Zool. 299:295–303. [Google Scholar]
- de Oliveira TG. 1998. Herpailurus yagouaroundi. Mamm Species. 578:1–6. doi: 10.2307/3504500. [DOI] [Google Scholar]
- Dobrynin P, Liu S, Tamazian G, Xiong Z, Yurchenko AA, Krasheninnikova K, Kliver S, Schmidt-Küntzel A, Koepfli KP, Johnson W, et al. 2015. Genomic legacy of the African cheetah, Acinonyx jubatus. Genome Biol. 16:277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eroğlu HE. 2017. The comparison of the Felidae species with karyotype symmetry/asymmetry index. Punjab Univ J Zool. 32:229–235. [Google Scholar]
- Espinosa CC, Trigo TC, Tirelli FP, da Silva LG, Eizirik E, Queirolo D, Mazim FD, Peters FB, Favarini MO, de Freitas TRO. 2017. Geographic distribution modeling of the margay (Leopardus wiedii) and jaguarundi (Puma yagouaroundi): a comparative assessment. J Mammal. 99:252–262. [Google Scholar]
- Garrison E, Marth G. 2012. Haplotype-based variant detection from short-read sequencing. arXiv preprint arXiv:1207.3907. [Google Scholar]
- Graphodatsky A, Perelman P, O’Brien SJ. 2020. Atlas of mammalian chromosomes. Hoboken, NJ: John Wiley & Sons, Inc. [Google Scholar]
- Gurevich A, Saveliev V, Vyahhi N, Tesler G. 2013. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 29:1072–1075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson P, Nowell K. 1996. Wild cats: status survey and conservation action plan. Gland, Switzerland: IUCN. [Google Scholar]
- Johnson WE, Eizirik E, Pecon-Slattery J, Murphy WJ, Antunes A, Teeling E, O’Brien SJ. 2006. The late Miocene radiation of modern Felidae: a genetic assessment. Science. 311:73–77. [DOI] [PubMed] [Google Scholar]
- Jurka J, Kapitonov VV, Pavlicek A, Klonowski P, Kohany O, Walichiewicz J. 2005. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet Genome Res. 110:462–467. [DOI] [PubMed] [Google Scholar]
- Kent WJ. 2002. BLAT–the BLAST-like alignment tool. Genome Res. 12:656–664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiełbasa SM, Wan R, Sato K, Horton P, Frith MC. 2011. Adaptive seeds tame genomic sequence comparison. Genome Res. 21:487–493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitchener AC, Breitenmoser-Würsten C, Eizirik E, Gentry A, Werdelin L, Wilting A, Yamaguchi N, Abramov AV, Christiansen P, Driscoll C. 2017. A revised taxonomy of the Felidae: the final report of the Cat Classification Task Force of the IUCN Cat Specialist Group. Cat News Special Issue; 11, 1–80. [Google Scholar]
- Köster J, Rahmann S. 2012. Snakemake–a scalable bioinformatics workflow engine. Bioinformatics. 28:2520–2522. [DOI] [PubMed] [Google Scholar]
- Kriventseva EV, Kuznetsov D, Tegenfeldt F, Manni M, Dias R, Simão FA, Zdobnov EM. 2019. OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res. 47:D807–D811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA. 2009. Circos: an information aesthetic for comparative genomics. Genome Res. 19:1639–1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, Carey VJ. 2013. Software for computing and annotating genomic ranges. PLoS Comput Biol. 9:e1003118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lex A, Gehlenborg N, Strobelt H, Vuillemot R, Pfister H. 2014. UpSet: visualization of intersecting sets. IEEE Trans Vis Comput Graph. 20:1983–1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H. 2013. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:1207.3907. [Google Scholar]
- Li G, Davis BW, Eizirik E, Murphy WJ. 2016. Phylogenomic evidence for ancient hybridization in the genomes of living cats (Felidae). Genome Res. 26:1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li G, Figueiró HV, Eizirik E, Murphy WJ. 2019. Recombination-aware phylogenomics reveals the structured genomic landscape of hybridizing cat species. Mol Biol Evol. 36:2111–2126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R; 1000 Genome Project Data Processing Subgroup . 2009. The sequence alignment/map format and SAMtools. Bioinformatics. 25:2078–2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morgulis A, Gertz EM, Schäffer AA, Agarwala R. 2006. WindowMasker: window-based masker for sequenced genomes. Bioinformatics. 22:134–141. [DOI] [PubMed] [Google Scholar]
- O’Brien SJ, Johnson WE. 2007. The evolution of cats. Sci Am. 297:68–75. [PubMed] [Google Scholar]
- O’Brien SJ, Johnson W, Driscoll C, Pontius J, Pecon-Slattery J, Menotti-Raymond M. 2008. State of cat genomics. Trends Genet. 24:268–279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ochoa A, Onorato DP, Fitak RR, Roelke-Parker ME, Culver M. 2019. De novo assembly and annotation from parental and F1 puma genomes of the florida panther genetic restoration program. G3 (Bethesda). 9:3531–3536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinlan AR, Hall IM. 2010. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 26:841–842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz-García M, Pinedo-Castro M. 2013. Population genetics and phylogeography analyses of the jaguarundi (Puma yagouarundi) by means of three mitochondrial markers: the first molecular population study of this species. In: Ruiz-Garcia M, Shostell JM, editors. Molecular population genetics, evolutionary biology, and biological conservation of neotropical carnivores. New York: Nova Publishers. [Google Scholar]
- Ruiz-García M, Pinedo-Castro M, Shostell JM. 2018. Mitogenomics of the jaguarundi (Puma yagouaroundi, Felidae, Carnivora): disagreement between morphological subspecies and molecular data. Mamm Biol. 93:153–168. [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. 1989. Molecular cloning: a laboratory manual. New York: Cold Spring Harbor Laboratory Press. [Google Scholar]
- Saremi NF, Supple MA, Byrne A, Cahill JA, Coutinho LL, Dalén L, Figueiró HV, Johnson WE, Milne HJ, O’Brien SJ, et al. 2019. Puma genomes from North and South America provide insights into the genomic consequences of inbreeding. Nat Commun. 10:4769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seppey M, Manni M, Zdobnov EM. 2019. BUSCO: assessing genome assembly and annotation completeness. Methods Mol Biol. 1962:227–245. [DOI] [PubMed] [Google Scholar]
- Smit AFA, Hubley R, Green P. 2013–2015. RepeatMasker Open-4.0. [Google Scholar]
- Stanke M, Schöffmann O, Morgenstern B, Waack S. 2006. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinf. 7:62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sunquist ME, Sunquist FC. 2009. Family Felidae (cats). In: Wilson DE, Mittermeier RA, Cavallini P, editors. Handbook of the mammals of the world, Vol. 1. Barcelona, Spain: Lynx Edicions. p. 54–169. [Google Scholar]
- Tamazian G. 2021. JDGA package [Internet]. [cited 2021 April 12]. Available from: https://gitlab.com/gtamazian/jdga.
- Tange O. 2011. GNU Parallel - the command-line power tool. login: the USENIX Magazine. 36:42–47. [Google Scholar]
- Weisenfeld NI, Kumar V, Shah P, Church DM, Jaffe DB. 2017. Direct determination of diploid genome sequences. Genome Res. 27:757–767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler TJ, Clements J, Eddy SR, Hubley R, Jones TA, Jurka J, Smit AF, Finn RD. 2013. Dfam: a database of repetitive DNA based on profile hidden Markov models. Nucleic Acids Res. 41:D70–D82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickham H, Averick M, Bryan J, Chang W, McGowan LDA, François R, Grolemund G, Hayes A, Henry L, Hester J. 2019. Welcome to the Tidyverse. Journal of Open Source Software. 4:1686. [Google Scholar]
- Wurster-Hill DH, Gray CW. 1973. Giemsa banding patterns in the chromosomes of twelve species of cats (Felidae). Cytogenet Cell Genet. 12:388–397. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
We have deposited the primary data underlying the presented analyses as follows:
Raw and filtered read alignments, jaguarundi gene annotation, genomic variants, and auxiliary datasets: Dryad, https://doi.org/10.5061/dryad.wstqjq2mf
The JDGA package: GitLab repository https://gitlab.com/gtamazian/jdga
The Snakemake pipelines: GitLab repository https://gitlab.com/gtamazian/jdga_pipelines
Raw sequence reads: NCBI SRA accession SRX8453956
The jaguarundi genome sequence assembly: NCBI GenBank accession GCA_014898765.1 (PumYag) under NCBI BioSample accession SAMN14930875 and NCBI BioProject accession PRJNA633021 (Puma yagouaroundi).