Abstract
Tumor heterogeneity is a major obstacle to the success of cancer treatment. An accurate understanding and recognition of tumor heterogeneity is critical in the clinical management of cancer patients. Here, we utilized single-cell RNA sequencing (scRNA-seq) to uncover the intra- and intertumoral heterogeneity of liver metastases from a patient with metastatic uveal melanoma. The two metastases analyzed were largely infiltrated by noncancerous cells with significant variability in the proportion of different cell types. Analysis of copy-number variations (CNVs) showed gain of 8q and loss of 6q in both tumors, but loss of Chromosome 3 was only detected in one of the tumors. Single-nucleotide polymorphism (SNP) array revealed a uniparental isodisomy 3 in the tumor with two copies of Chromosome 3, indicating a regain of Chromosome 3 during the development of the metastatic disease. In addition, both tumors harbored subclones with additional CNVs. Pathway enrichment analysis of differentially expressed genes revealed that cancer cells in the metastasis with isodisomy 3 showed up-regulation in epithelial–mesenchymal transition and myogenesis related genes. In contrast, up-regulation in interferon signaling was observed in the metastasis with monosomy 3 and increased T-cell infiltrate. This study highlights the complexity and heterogeneity of different metastases within an individual case of uveal melanoma.
Keywords: choroidal melanoma
INTRODUCTION
Uveal melanoma (UM) is the second most common subtype of melanoma. Although the primary UM can be controlled by either surgery or radiation brachytherapy, distant metastases eventually develop in up to 50% of UM patients (Krantz et al. 2017). The liver is the initial metastatic site in the overwhelming majority of cases (95%) (Willson et al. 2001). There is no consensus on the treatment for metastatic UM, and the median overall survival is ∼1 yr (Khoja et al. 2019; Rantala et al. 2019). Recent results suggest that treatment with tebentafusp or combination immunotherapy using nivolumab and ipilimumab provides some clinical benefit (Middleton et al. 2020; Pelster et al. 2021). However, more effective treatments for metastatic UM are still urgently needed. Once metastatic disease develops, 80% and 92% of the patients die within 1 and 2 yr, respectively (Diener-West et al. 2005; Jovanovic et al. 2013). The molecular subtypes of UM have been defined based on their gene expression profile (GEP), chromosome copy-number variations (CNVs), and key driving mutations (Onken et al. 2004; Royer-Bertrand et al. 2016; Robertson et al. 2017; Jager et al. 2018; Beasley et al. 2020). Two prognostically favorable subtypes, which have either a low (Group A) or an intermediate (Group B) metastatic risk, are associated with disomy 3 and Class 1 GEP, possess mutations in EIF1AX (Group A) or SF3B1 (Group B), and harbor two (Group A) or more (Group B) copies of Chromosome 8q. In contrast, two prognostically unfavorable subtypes, which are associated with a high metastatic risk, show monosomy 3 and Class 2 GEP, inactivation of BAP1, and an extra copy of Chromosome 8q (Group C) or multiple copies of Chromosome 8q (Group D). Group D is also associated with significant inflammation and immune infiltrate.
It has been well recognized that tumor heterogeneity poses a significant challenge in the management of cancer patients (Bedard et al. 2013). The heterogeneity of liver metastases in UM is underinvestigated because of both rarity of the disease and the fact that surgical removal is not a standard practice for UM patients with liver metastasis. Nonetheless, Krishna and colleagues demonstrated the intersegmental heterogeneity of the immune signatures in UM metastases using bulk analysis (Krishna et al. 2020). In the past decade, single-cell RNA sequencing (scRNA-seq) has been developed as a powerful tool to identify and study tumor heterogeneity (Kolodziejczyk et al. 2015). Here, we characterize the genomic profile of two concurrent liver metastases from a patient with UM using scRNA-seq and copy-number alteration analysis. We further compare the molecular results to the clinical characteristics to exemplify the heterogeneity of metastatic UM tumors.
RESULTS
Clinical Presentation
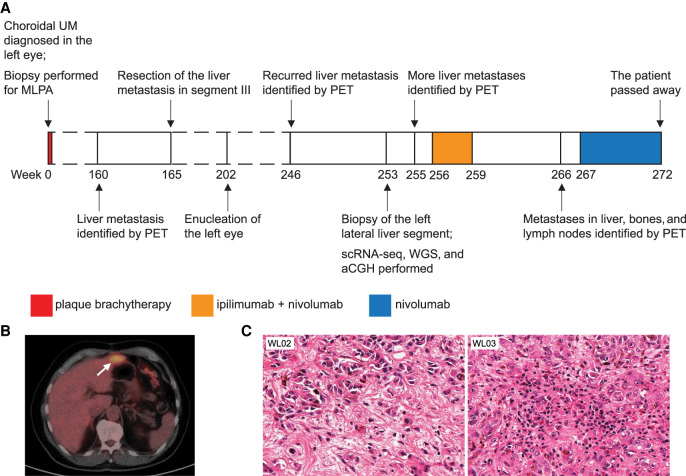
A 69-yr-old male was diagnosed with choroidal UM in his left eye in 2014 (labeled as week 0) and was enrolled in our study prior to Iodine125 plaque insertion (Fig. 1A). The patient was followed up clinically with direct eye examination for local surveillance and six-monthly positron emission tomography (PET) scans to monitor for metastatic disease. An isolated liver lesion (56 mm) in segment III was identified on imaging during his follow-up. The patient underwent a partial liver resection of this lesion at week 165. Histopathology confirmed metastatic UM. Postoperatively, he had no other identified metastases and recommenced active surveillance. At week 202, he underwent enucleation of the left eye because of local recurrence. At week 246, an intensely 18F-fluorodeoxyglucose (FDG)-avid liver lesion was identified in the left lateral liver segment by PET (Fig. 1B). At week 253, the patient underwent a second operation, during which several variably pigmented abnormal nodules were identified macroscopically, ranging from <1 to 17 mm in maximum diameter. Two separate metastatic lesions were resected, providing the specimens described here. After surgery, a restaging PET scan at week 255 indicated further liver metastases in segment VIII and segment VI/VII, which were not previously evident. The patient was commenced on combination immunotherapy with ipilimumab and nivolumab at week 256, but only completed two doses because of the development of grade III immune-related hepatitis. Liver toxicity improved but did not completely dissipate with high-dose steroids; therefore, mycophenolate was added. A restaging scan at week 266 showed marked hepatomegaly and upper abdominal disease in addition to two new bone metastases. At week 267, therapy continued with single-agent nivolumab only for three more cycles, and the patient died soon after from advanced metastatic UM.
Figure 1.
The patient's clinical information. (A) Treatment history of the patient. (B) An intensely 18F-fluorodeoxyglucose (FDG)-avid liver lesion (white arrows) was identified in the left lateral segment by positron emission tomography (PET) in week 246. The total lesion glycolysis was equal to 84 standardized uptake value (SUV) × cm3. (C) Hemotoxylin and eosin (H&E) staining of two liver metastases WL02 and WL03. Tumor cells in WL02 (left) had a spindled to epithelioid appearance with a relatively high number of admixed stromal cells in an oedematous stroma, whereas tumor cells in WL03 (right) had an epithelioid appearance and the stroma was rich in lymphocytes at the periphery of the tumor (H&E, original magnification, both ×350). (UM) Uveal melanoma, (MLPA) multiplex litigation-dependent probe amplification, (scRNA-seq) single-cell RNA sequencing, (WGS) whole-genome sequencing, (aCGH) array comparative genomic hybridization.
Histopathological Characteristics
The enucleated left eye at week 202 contained a large choroidal melanoma (14 × 6 mm), predominantly composed of epithelioid cells according to Callender classification. The tumor was partly necrotic with evidence of radiation-induced tumor reduction/necrosis affecting ∼40% of the total tumor volume. There was also evidence of trans-scleral invasion (to conjunctival stroma). The optic nerve margin was uninvolved.
Representative samples for the two metastatic nodules resected from the left lateral liver segment at week 253 were harvested for molecular studies and labeled as WL02 and WL03. Histopathological examination revealed that the tumor cells in WL02 were composed of a combination of spindle and epithelioid cells, whereas the tumor cells in WL03 had a purely epithelioid appearance (Fig. 1C). Both WL02 and WL03 consisted of tumor cells showing enlarged pleomorphic round to oval nuclei, often prominent nucleoli, and moderately abundant eosinophilic cytoplasm with varying melanin pigmentation. A variable number of admixed lymphocytes, macrophages, and stromal cells (including hepatic stellate cells) were present in the milieu surrounding these nodules, with the WL02 sample showing widespread areas with an oedematous stroma containing a high proportion of stromal cells and macrophages, whereas WL03 contained areas rich in lymphocytes, particularly at the interface between the tumor and native liver tissue.
Genomic Analysis
A tissue sample of the primary tumor was analyzed at the time of radiation plaque insertion by multiplex ligation-dependent probe amplification (MLPA) (Supplemental Fig. 1A). The tumor was suggested to be of high metastatic risk based on the cytogenetic findings including loss of Chromosome 1p, loss of Chromosome 3, and gain of Chromosome 8q.
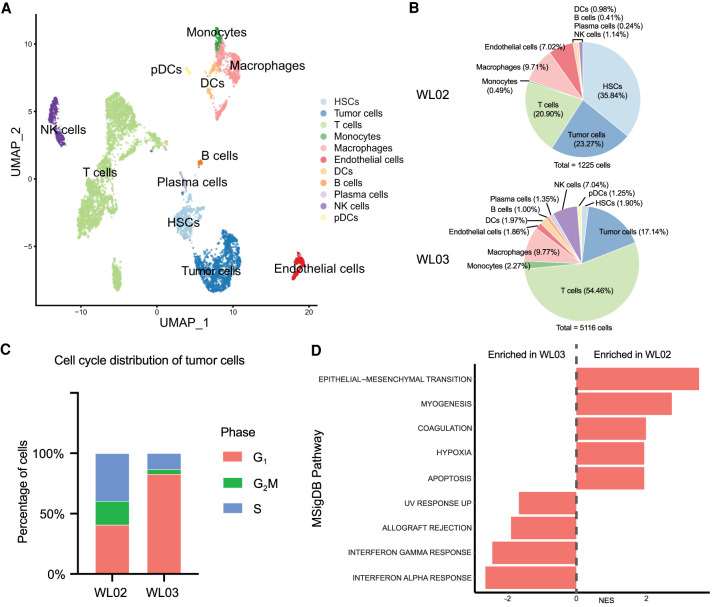
WL02 and WL03 metastatic nodules were analyzed using droplet-based scRNA-seq. After quality control, a total of 1225 and 5116 high-quality single cells were recovered from WL02 and WL03, respectively. Based on the expression of a panel of canonical marker genes, all the cells were manually annotated into 11 groups (Fig. 2A). Both tumors were heavily infiltrated by nonneoplastic cells (76.73% vs. 82.86%) including hepatic stellate cells (HSCs) and immune cells, with tumor cells constituting 17%–23% of the cells sequenced (Fig. 2B). The two tumors differed in the proportion of nontumor cell types in their microenvironment, with WL03 presenting a large T-cell infiltrate (54%). WL02 was enriched in HSCs (36%) and endothelial cells (7%) compared to WL03, which only had a low percentage of both cell types (1.9% of each). Both tumors contained a similar number of macrophages (9.71% vs. 9.77%). Tumor cells in WL02 were more proliferative as evidenced by high proportion of cells in S and G2M phases, whereas WL03 manifested quiescent characteristic with a high proportion of tumor cells in G1 phase (Fig. 2C).
Figure 2.
Aggregate analysis of 6341 single cells from two liver metastases. (A) Uniform manifold approximation and projection (UMAP) plot of 6341 single cells, colored by cell types that were manually assigned to cell clusters. (B) Pie chart of each liver metastasis showing the percentages of annotated cell types. (C) Cell cycle distribution of tumor cells. Percentage of tumor cells in different phases of the cell cycle was estimated in WL02 (left) and WL03 (right) using CellCycleScoring function in Seurat. (D) Gene set enrichment analysis (GSEA) pathways enriched by differentially expressed genes of tumor cells. Differentially enriched biological pathways are displayed ordered by the normalized enrichment score (NES). (DC) Plasmacytoid dendritic cell, (NK) natural killer, (HSC) hematopoietic stem cell.
Comparison of tumor cells in WL02 and WL03 by gene set enrichment analysis (GSEA) indicated that genes involved in epithelial–mesenchymal transition (EMT), myogenesis, coagulation, and hypoxia were significantly enriched in WL02 (Fig. 2D). Conversely, genes involved in immune-related pathways such as interferon-γ and -α response were significantly enriched in WL03, which was consistent with the high abundance of T-cell infiltrate observed in this tumor (Figs. 1C, 2B).
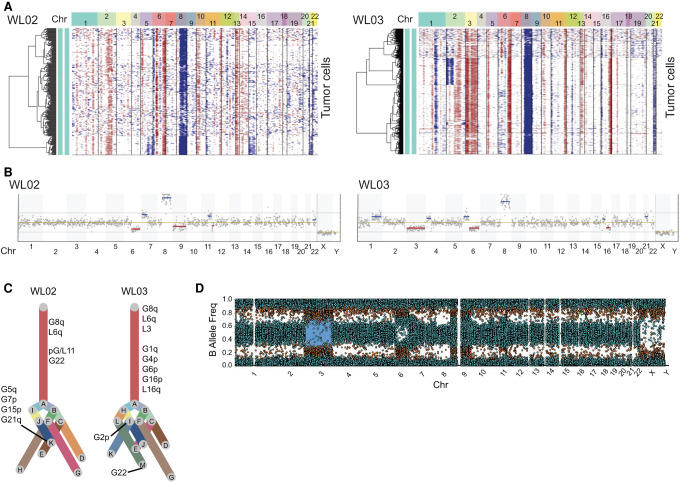
We examined CNVs in the tumor cells from these two metastases using inferCNV (Fig. 3A). Hierarchical clustering indicated gain of 8q and loss of 6q in both tumors consistent with the primary tumor genotype, but loss of Chromosome 3 was only apparent in WL03. The main CNV profile identified by inferCNV, including gain of 8q and loss of 6q in both tumors, and loss of Chromosome 3 in WL03, was confirmed by low-pass whole-genome sequencing (LP-WGS) (Fig. 3B) and by array comparative genomic hybridization (aCGH) (Supplemental Fig. 1B). Targeted sequencing revealed a BAP1 E182V mutation in both WL02 and WL03.
Figure 3.
Copy-number variation (CNV) analysis of two liver metastases. (A) Hierarchical clustering from inferCNV analysis showing CNV changes by chromosomes (columns) for individual tumor cells (rows). Blue indicates copy gains and red indicates copy losses. (B) Low-pass whole-genome sequencing (LP-WGS) analysis of two liver metastases. CNVs are denoted by horizontal lines indicating total or partial chromosomal losses (red) or gains (blue). (C) Clonality trees of each of tumor cells in the two liver metastases. The branches are scaled according to percentage of cells in the calculated subclone containing the indicated CNVs. (D) Single-nucleotide polymorphism (SNP) array-based karyotyping of WL02. (Chr) Chromosome.
In addition, inferCNV revealed the development of subclones within each tumor: A subclone (clone K) carrying gains of 5q, 15p, and 21 was found within WL02, and a subclone (clone I) carrying gain of 2p was found within WL03 (Fig. 3C). These analyses also indicated several distinct CNVs between these two tumors with gain of 7p, and partial gain and loss of 11q uniquely observed in WL02. Conversely, gains of 1q, 4p, 6p, and 16p, and loss of 16q were found in WL03, but were not evident in WL02.
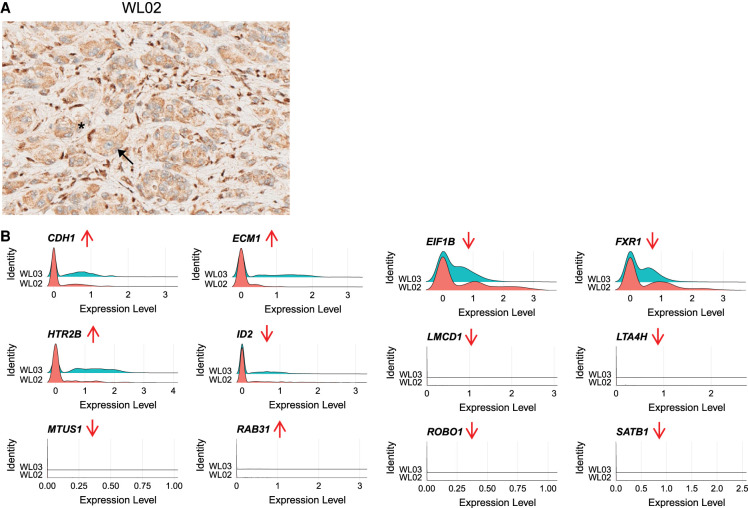
Overall, the major discordance between the two tumors was the lack of monosomy 3 in WL02. SNP array analysis indicated uniparental isodisomy of Chromosome 3 in WL02 (Fig. 3D). Isodisomy 3 in WL02 did not reconstitute the expression of BAP1, indicated by the lack of mRNA expression (Supplemental Fig. 2A), further validated by the lack of BAP1 expression in the corresponding formalin-fixed paraffin-embedded (FFPE) blocks (Fig. 4A). However, GEP of a few genes was slightly shifted in tumor cells from WL02 compared with WL03. These included expression of CDH1, ECM1, EIF1B, FXR1, and HTR2B, which are associated with UM risk classification, suggesting a gene expression profile trend toward a Class 1 GEP subtype (Fig. 4B).
Figure 4.
Analysis of gene expression in two liver metastases. (A) Immunohistochemical staining with BAP1 antibody in WL02; note the loss of nuclear staining in tumor cells (arrow) in the presence of positive internal control (retained nuclear staining) in endothelial and stromal cells (asterisk) (BAP-1 IHC, original magnification ×320). (B) Ridge plots of the 15-gene panel in tumor cells between WL02 and WL03. Red arrows indicate the direction of change in a Class 2 tumor.
DISCUSSION
The results of the presented study demonstrated a significant inter- and intratumor heterogeneity in two spatially separated liver metastases of UM at the single-cell level. Tumor cells in WL02 showed migratory properties with an enrichment of genes involved in EMT, myogenesis, coagulation, and hypoxia. These cells were also found to be in an active proliferative state. Altogether this suggests a high migratory and invasive propensity, which was further supported by the evidence of widespread metastases detected in other liver segments, lymph node, and bone in subsequent PET scans.
Our analysis shows that cells from a primary tumor with monosomy 3 could regain a copy of this chromosome in the form of isodisomy 3 in all tumor cells in one of its metastatic deposits, and not the other. The lack of BAP1 expression in the presence of two copies of Chromosome 3 in this metastatic deposit suggests duplication of the chromosome harboring an inactivating mutation in BAP1 allele. Loss of BAP1 expression is associated with a Class 2 GEP (Harbour et al. 2010; Robertson et al. 2017), and a recent study demonstrated that the reintroduction of a wild-type BAP1 copy in vitro could shift the GEP toward a Class 1 subtype (Karlsson et al. 2020). Conversely, it can be speculated that isodisomy 3 without a functional BAP1 in WL02 could still partially reverse the GEP to a Class 1 subtype, if there were other factors on Chromosome 3 that were involved in the regulation of the GEP. It is unclear how Chromosome 3 duplication may benefit disease progression once metastatic disease is stablished. However, it could represent a lack of pressure to maintain a Class 2 GEP at the late stage of the disease. Notably, isodisomy 3 as well as losses of 1p, 6q and gains of 1q, 8q have been reported to be acquired during the metastatic stage (Rodrigues et al. 2019), suggesting that restoration of heterozygosity may provide a selective advantage at the metastatic stage.
Loss of Chromosome 9p has been observed in a quarter of UM (Harbour 2012); similarly, our analysis showed a loss of Chromosome 9 in WL02 (Fig. 3B). Chromosome 9p loss has been associated with loss of CDKDN2, resulting in a higher proliferative state of tumor cells as observed in WL02, implied by the cell cycle scoring analysis. Interestingly, we also found that WL02 harbored the loss of Chromosome 9, concurrently having a higher level of 8q gain, which is consistent with the proposed evolutive pattern of metastatic UM (Shain et al. 2019).
In a recent study, Pandiani et al. (2021) used scRNA-seq to study intratumoral heterogeneity in primary UM and demonstrated the critical role of HES6 in enhancing growth and motile ability of primary UM. Our data showed increased expression of HES6 in both liver metastases (Supplemental Fig. 2B). However, the tumor (WL02) with more proliferative and motile characteristics had a lower HES6 expression, suggesting that this metastatic tumor might have relied on alternative signaling to promote proliferation and migration.
A recent study in cutaneous melanoma demonstrated that CNVs can influence the local immune composition, and Chromosome 7 gain was attributed to a low-leukocyte infiltrate in the tumor microenvironment (Mitra et al. 2020). In agreement with this study, WL02 harboring a gain of 7p showed a lower T-cell density compared to WL03. Concomitantly, WL02 had a large proportion of HSCs infiltrate. As demonstrated in a previous study (Yu et al. 2004), HSCs can suppress T-cell infiltration, suggesting that HSCs may have also affected the reduced T-cell composition in this sample. These observations signify a potential differential sensitivity in response to immune checkpoint inhibitors in these two separate tumors.
The expression of immune checkpoint proteins in T cells in both tumors was consistent with the previous studies (Supplemental Fig. 2C; Durante et al. 2020; Karlsson et al. 2020). T cells did not express HAVCR2 (TIM-3) or PDCD1 (PD-1). The latter could explain why nivolumab, after discontinuation of combination immunotherapy, was ineffective in this patient. On the other hand, LAG3 was strongly expressed in T cells in WL03, but not in T cells in WL02. This heterogeneity among UM metastatic deposits may have future therapeutic implications as LAG3 blockade is pursued as promising treatment for cancer (Atkinson et al. 2020).
In summary, this case study highlights the complexity of UM liver metastases, broadening the spectrum of transcriptome landscape in such tumors at a single-cell level. More importantly, it emphasizes the significance of an accurate assessment of inter- and intratumoral heterogeneity for delivering the most specific and personalized therapeutic intervention at the right time for patients.
METHODS
Patient and Sample Collection
Resected tumors were triaged by pathologist immediately after surgery and immersed into RPMI 1640 supplemented with 20% fetal bovine serum, 10,000 U/mL penicillin, and 10,000 µg/mL streptomycin (ThermoFisher Scientific) at 4°C to maximize the viability of cells. A fraction of each tumor was freshly subjected to single-cell dissociation, whereas the rest was processed as FFPE blocks for histopathology analysis.
Preparation of Single-Cell Suspension
Tumors obtained from the surgery were freshly subjected to single-cell dissociation using Human Tumor Dissociation Kit (Miltenyi Biotec) according to the manufacturer's instructions. Briefly, the tumors were washed with plain RPMI (ThermoFisher Scientific) supplemented with penicillin and streptomycin (ThermoFisher Scientific) and processed into pieces (1-mm3) using scalpels with the recommended volume of enzyme mix as per instructions. The tumor pieces and enzyme mix were then transferred into gentleMACS C tubes (Miltenyi Biotec), and then the tubes were loaded onto the gentleMACS Octo Dissociator with Heaters (Miltenyi Biotec) with the program according to the manufacturer's instructions. After incubation, digested tissues were filtered through Falcon 70-µm strainers (Corning) and washed. Cell suspension was further filtered through 40-µm strainers to remove cell clumps. The viability was assessed by ReadyProbe Cell Viability Imaging Kit (ThermoFisher Scientific) to ensure the viability was >90%.
Single-Cell RNA Sequencing and Analysis
Single-cell RNA sequencing was performed using Chromium Single Cell 3’ Reagent Kits v3 (10× Genomics) according to the manufacturer's protocol. WL02 was aimed with a targeted capture of 2000 cells because of the limited tissue obtained from the surgery, whereas WL03 was aimed with a targeted capture of 9000 cells. Gel Beads-in-emulsion was generated using Chromium Single Cell B Chip Kit. Libraries were sequenced using the Illumina NovaSeq 6000 and S2 flow cells (100-cycle kit) with a read one length of 28 cycles and a read two length of 94 cycles.
Raw data was processed as described previously (Denisenko et al. 2020). Briefly, BCL files were demultiplexed and converted into FASTQ using bcl2fastq utility of Illumina BaseSpace Sequence Hub. FASTQ files were processed using Cell Ranger 3.0.2 with refdata-cellranger-GRCh38-3.0.0 reference. Raw gene-barcode matrices from Cell Ranger output were used for downstream processing. Cells were distinguished from background noise using EmptyDrops (Lun et al. 2019). Outlier cells with a high ratio of number of detected UMI to genes (>3 median absolute deviations from median) were removed using Scater (McCarthy et al. 2017). The preprocessed data from these two tumors was then integrated in R (version 3.6.2) (C Core Team 2013) with the SCTransform workflow using the Seurat toolkit (version 3.2.3) (Stuart et al. 2019). Quality control was performed by retaining cells with nCount_RNA > 400, nFeature_RNA > 100 but < 8000, and mitochondrial content <10%. CellCycleScoring( ) function was used to predict the phases of each cell according to Seurat tutorial. In SCTransform( ) function, percent of mitochondrial content was used in “vars.to.regress” argument to regress out variation. In FindIntegrationAnchors( ) function, WL02 was used as a reference data set as it contained fewer doublets. The first 30 principal components were used in RunUMAP( ) and FindNeighbors( ) functions, whereas the resolution parameter was set to 2 in FindCluster( ) function. For other parameters unspecified above, default values were used in the SCTransform workflow.
Clusters were visualized in a two-dimensional unified manifold approximation and projection (UMAP) and were annotated using canonical marker genes based on the literature (MacParland et al. 2018; Durante et al. 2020; Sharma et al. 2020; Zhao et al. 2020). The marker genes were as listed below: tumor cells (MLANA, MITF, DCT), hepatic stellate cells (ACTA2, COL1A1, COL1A2, TAGLN, COL3A1, RBP1, SPARC), endothelial cells (PECAM1, VWF), NK cells (KLRF1), T cells (CD3D), B cells (CD19, CD79A, MS4A1), plasma cells (IGHG1, MZB1, SDC1, CD79A), plasmacytoid dendritic cells (LILR4, GZMB), dendritic cells (CLEC9A, CD1C, CD1E), monocytes (S100A8, S100A9, S100A12), and macrophages (CD163, IL10, C1QA, C1QB, C1QC).
GSEA
To obtain differentially expressed genes in tumor cells between WL02 and WL03, FindMarkers( ) function from Seurat package (version 3.2.3) was used (Stuart et al. 2019). GSEA software (version 4.1.0, http://www.gsea-msigdb.org/gsea/downloads.jsp) for Linux was used to calculate hallmark gene set enrichment (h.all.v7.2.symbols.gmt) using GSEAPreanked and Log2 fold change. Hallmark normalized enrichment score (NES) was plotted against enriched pathways using R (version 4.0.4).
CNV Analysis
InferCNV (version 1.7.1) was used to predict the copy-number variations from scRNA-seq data (InferCNV of the Trinity CTAT Project: https://github.com/broadinstitute/InferCNV). To keep high-quality T cells as normal reference cells, only T cells that had no expression of PRAME, HTR2B, MLANA, and MITF were used. A value of 0.1 was used for the “cutoff” argument and default values were used in other unspecified arguments. Intratumor evolutionary trees were generated using UPyhloplot2 (version 2.3; https://github.com/harbourlab/uphyloplot2) as described in Kurtenbach et al. (2021). A phylogenetic tree was manually annotated using the output of UPhyloplot2 and “.pred_cnv_regions.dat” file from InferCNV in Adobe Illustrator (version 25.2.3).
Shallow LP-WGS was used to detect CNVs from WL02 and WL03. DNA was extracted from FFPE tissue using the AllPrep DNA/RNA Mini Kit (QIAGEN). An amount of 100 ng of DNA was digested using the MseI restriction enzyme (New England Biolabs) and 200-bp libraries constructed using the NEBNext Fast DNA Library Prep Set for Ion Torrent (New England Biolabs) to manufacturer's specifications. Resultant DNA was templated and loaded into an Ion 540 chip (ThermoFisher Scientific) at 60 pM using the Ion Chef (Thermofisher Scientific). Libraries were sequenced on an Ion S5 (ThermoFisher Scientific) for 520 flows. Data was uploaded and analyzed using Ion Reporter (version 5.14).
CGH and SNP arrays were performed in the same FFPE-derived DNA samples. For CGH assay, the test DNA and sex mismatched human reference DNA were differentially labeled with fluorophores. Equal quantities of the two DNA samples were mixed and hybridized onto a SurePrint G3 Human CGH Microarray, 8 × 60K (Agilent) and digitally scanned to capture and quantify the relative fluorescence intensities. The ratio of the fluorescence intensities was analyzed by CytoGenomics software (Agilent) for further analysis and detection of potential CNVs. For SNP array the Infinium HD assay was performed utilizing Illumina iScan and HumanCytoSNP FFPE-12 BeadChip array according to the manufacturer's protocol (Illumina). The BeadChips were stained and then imaged, using a BeadArray Reader (Illumina). Image data were analyzed with GenomeStudio (Illumina).
ADDITIONAL INFORMATION
Database Deposition and Access
The scRNA-seq data generated in this study has been deposited in Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under accession code GSE176029.
Ethics Statement
This study was approved by the Human Research Ethics Committee protocols from Edith Cowan University (No. 11543) and Sir Charles Gairdner Hospital (No. 2013-246) in compliance with the Declaration of Helsinki. Experiments were performed in accordance with institutional and national guidelines and regulations. The patient provided signed informed consent for the collection of specimens, detailed analyses of the derived genetic material, and access to clinical data.
Acknowledgments
We are grateful to the patient who donated the samples in this research. We thank Dr. Bryon Jaques at Sir Charles Gairdner Hospital who performed the surgery to collect the tumor samples, Professor Helen Rizos at Macquarie University who shared the methods of tumor dissociation, Dr. Zeyad Al-Ogaili at Fiona Stanley Hospital who calculated the total lesion glycolysis of the PET scan, and Dr. Cleo Robinson and Dr. Karen Woodward from PathWest for CGH and SNP array analyses. We also thank Mr. Michael A. Durante and Professor J. William Harbour at Bascom Palmer Eye Institute for the discussion of this project.
Author Contributions
W.L., A.B.B., N.M.A., and E.S.G. wrote the manuscript. W.L. prepared the libraries for scRNA-seq. M.J. carried out the Novaseq sequencing. L.C. prepared the libraries and performed the LP-WGS. W.L., A.B.B., and E.D. carried out the bioinformatic analysis. L.W. recruited the patient and reviewed the clinical history. N.M.A. and B.A.W. performed the histopathological analysis. A.R.R.F. and E.S.G. designed and supervised the study. All authors have read and approved the final manuscript.
Funding
This project was supported by the Cancer Research Trust and Cancer Council Western Australia (CCWA) through the “Enabling advanced single cell cancer genomics in WA” grant to A.R.R.F, and a Collaborative CCWA grant to W.L., N.M.A., and E.D. Genomic data was generated at the Australian Cancer Research Foundation Centre for Advanced Cancer Genomics. Analysis was made possible with computational resources provided by the Pawsey Supercomputing Centre with funding from the Australian Government and the Government of Western Australia. A.R.R.F. is supported by an Australian National Health and Medical Research Council (NHMRC) Fellowship APP1154524. L.W. is supported by an NHMRC Scholarship APP1190643. A.B.B. is supported by an Edith Cowan University Postgraduate Scholarship and a CCWA PhD Top-up Scholarship. E.S.G. is supported by a CCWA Fellowship.
Competing Interest Statement
The authors have declared no competing interest.
Supplementary Material
Footnotes
[Supplemental material is available for this article.]
REFERENCES
- Atkinson V, Khattak A, Haydon A, Eastgate M, Roy A, Prithviraj P, Mueller C, Brignone C, Triebel F. 2020. Eftilagimod alpha, a soluble lymphocyte activation gene-3 (LAG-3) protein plus pembrolizumab in patients with metastatic melanoma. J Immunother Cancer 8: e001681. 10.1136/jitc-2020-001681 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beasley AB, Bentel J, Allcock RJ, Vermeulen T, Calapre L, Isaacs T, Ziman MR, Chen FK, Gray ES. 2020. Low-pass whole-genome sequencing as a method of determining copy number variations in uveal melanoma tissue samples. J Mol Diagn 22: 429–434. 10.1016/j.jmoldx.2019.12.005 [DOI] [PubMed] [Google Scholar]
- Bedard PL, Hansen AR, Ratain MJ, Siu LL. 2013. Tumour heterogeneity in the clinic. Nature 501: 355–364. 10.1038/nature12627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denisenko E, Guo BB, Jones M, Hou R, de Kock L, Lassmann T, Poppe D, Clément O, Simmons RK, Lister R, et al. 2020. Systematic assessment of tissue dissociation and storage biases in single-cell and single-nucleus RNA-seq workflows. Genome Biol 21: 130. 10.1186/s13059-020-02048-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diener-West M, Reynolds SM, Agugliaro DJ, Caldwell R, Cumming K, Earle JD, Hawkins BS, Hayman JA, Jaiyesimi I, Jampol LM, et al. 2005. Development of metastatic disease after enrollment in the COMS trials for treatment of choroidal melanoma: Collaborative Ocular Melanoma Study Group Report No. 26. Arch Ophthalmol 123: 1639–1643. 10.1001/archopht.123.12.1639 [DOI] [PubMed] [Google Scholar]
- Durante MA, Rodriguez DA, Kurtenbach S, Kuznetsov JN, Sanchez MI, Decatur CL, Snyder H, Feun LG, Livingstone AS, Harbour JW. 2020. Single-cell analysis reveals new evolutionary complexity in uveal melanoma. Nat Commun 11: 496. 10.1038/s41467-019-14256-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harbour JW. 2012. The genetics of uveal melanoma: an emerging framework for targeted therapy. Pigment Cell Melanoma Res 25: 171–181. 10.1111/j.1755-148X.2012.00979.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA, Council ML, Matatall KA, Helms C, Bowcock AM. 2010. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science 330: 1410–1413. 10.1126/science.1194472 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jager MJ, Brouwer NJ, Esmaeli B. 2018. The Cancer Genome Atlas project: an integrated molecular view of uveal melanoma. Ophthalmology 125: 1139–1142. 10.1016/j.ophtha.2018.03.011 [DOI] [PubMed] [Google Scholar]
- Jovanovic P, Mihajlovic M, Djordjevic-Jocic J, Vlajkovic S, Cekic S, Stefanovic V. 2013. Ocular melanoma: an overview of the current status. Int J Clin Exp Pathol 6: 1230–1244. [PMC free article] [PubMed] [Google Scholar]
- Karlsson J, Nilsson LM, Mitra S, Alsén S, Shelke GV, Sah VR, Forsberg EMV, Stierner U, All-Eriksson C, Einarsdottir B, et al. 2020. Molecular profiling of driver events in metastatic uveal melanoma. Nat Commun 11: 1894. 10.1038/s41467-020-15606-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khoja L, Atenafu E, Suciu S, Leyvraz S, Sato T, Marshall E, Keilholz U, Zimmer L, Patel S, Piperno-Neumann S. 2019. Meta-analysis in metastatic uveal melanoma to determine progression free and overall survival benchmarks: an international rare cancers initiative (IRCI) ocular melanoma study. Ann Oncol 30: 1370–1380. 10.1093/annonc/mdz176 [DOI] [PubMed] [Google Scholar]
- Kolodziejczyk AA, Kim JK, Svensson V, Marioni JC, Teichmann SA. 2015. The technology and biology of single-cell RNA sequencing. Mol Cell 58: 610–620. 10.1016/j.molcel.2015.04.005 [DOI] [PubMed] [Google Scholar]
- Krantz BA, Dave N, Komatsubara KM, Marr BP, Carvajal RD. 2017. Uveal melanoma: epidemiology, etiology, and treatment of primary disease. Clin Ophthalmol 11: 279–289. 10.2147/OPTH.S89591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krishna Y, Acha-Sagredo A, Sabat-Pośpiech D, Kipling N, Clarke K, Figueiredo CR, Kalirai H, Coupland SE. 2020. Transcriptome profiling reveals new insights into the immune microenvironment and upregulation of novel biomarkers in metastatic uveal melanoma. Cancers (Basel) 12: 2832. 10.3390/cancers12102832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurtenbach S, Cruz AM, Rodriguez DA, Durante MA, Harbour JW. 2021. Uphyloplot2: visualizing phylogenetic trees from single-cell RNA-seq data. BMC Genomics 22: 1–3. 10.1186/s12864-021-07739-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lun AT, Riesenfeld S, Andrews T, Gomes T, Marioni JC. 2019. EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol 20: 1–9. 10.1186/s13059-018-1612-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacParland SA, Liu JC, Ma XZ, Innes BT, Bartczak AM, Gage BK, Manuel J, Khuu N, Echeverri J, Linares I, et al. 2018. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun 9: 4383. 10.1038/s41467-018-06318-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy DJ, Campbell KR, Lun AT, Wills QF. 2017. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33: 1179–1186. 10.1093/bioinformatics/btw777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Middleton MR, McAlpine C, Woodcock VK, Corrie P, Infante JR, Steven NM, Evans TRJ, Anthoney A, Shoushtari AN, Hamid O. 2020. Tebentafusp, A TCR/Anti-CD3 bispecific fusion protein targeting gp100, potently activated antitumor immune responses in patients with metastatic melanoma. Clin Cancer Res 26: 5869–5878. 10.1158/1078-0432.CCR-20-1247 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitra A, Andrews MC, Roh W, De Macedo MP, Hudgens CW, Carapeto F, Singh S, Reuben A, Wang F, Mao X, et al. 2020. Spatially resolved analyses link genomic and immune diversity and reveal unfavorable neutrophil activation in melanoma. Nat Commun 11: 1839. 10.1038/s41467-020-15538-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onken MD, Worley LA, Ehlers JP, Harbour JW. 2004. Gene expression profiling in uveal melanoma reveals two molecular classes and predicts metastatic death. Cancer Res 64: 7205–7209. 10.1158/0008-5472.CAN-04-1750 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandiani C, Strub T, Nottet N, Cheli Y, Gambi G, Bille K, Husser C, Dalmasso M, Béranger G, Lassalle S, et al. 2021. Single-cell RNA sequencing reveals intratumoral heterogeneity in primary uveal melanomas and identifies HES6 as a driver of the metastatic disease. Cell Death Differ 28: 1990–2000. 10.1038/s41418-020-00730-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelster MS, Gruschkus SK, Bassett R, Gombos DS, Shephard M, Posada L, Glover MS, Simien R, Diab A, Hwu P. 2021. Nivolumab and ipilimumab in metastatic uveal melanoma: results from a single-arm phase II study. J Clin Oncol 39: 599–607. 10.1200/JCO.20.00605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rantala ES, Hernberg M, Kivelä TT. 2019. Overall survival after treatment for metastatic uveal melanoma: a systematic review and meta-analysis. Melanoma Res 29: 561. 10.1097/CMR.0000000000000575 [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Core Team. 2013. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org [Google Scholar]
- Robertson AG, Shih J, Yau C, Gibb EA, Oba J, Mungall KL, Hess JM, Uzunangelov V, Walter V, Danilova L, et al. 2017. Integrative analysis identifies four molecular and clinical subsets in uveal melanoma. Cancer Cell 32: 204–220 e215. 10.1016/j.ccell.2017.07.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigues M, Mobuchon L, Houy A, Alsafadi S, Baulande S, Mariani O, Marande B, Rais KA, Van der Kooij MK, Kapiteijn E. 2019. Evolutionary routes in metastatic uveal melanomas depend on MBD4 alterations. Clin Cancer Res 25: 5513–5524. 10.1158/1078-0432.CCR-19-1215 [DOI] [PubMed] [Google Scholar]
- Royer-Bertrand B, Torsello M, Rimoldi D, El Zaoui I, Cisarova K, Pescini-Gobert R, Raynaud F, Zografos L, Schalenbourg A, Speiser D, et al. 2016. Comprehensive genetic landscape of uveal melanoma by whole-genome sequencing. Am J Hum Genet 99: 1190–1198. 10.1016/j.ajhg.2016.09.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shain AH, Bagger MM, Yu R, Chang D, Liu S, Vemula S, Weier JF, Wadt K, Heegaard S, Bastian BC. 2019. The genetic evolution of metastatic uveal melanoma. Nat Genet 51: 1123–1130. 10.1038/s41588-019-0440-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma A, Seow JJW, Dutertre CA, Pai R, Blériot C, Mishra A, Wong RMM, Singh GSN, Sudhagar S, Khalilnezhad S, et al. 2020. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183: 377–394 e321. 10.1016/j.cell.2020.08.040 [DOI] [PubMed] [Google Scholar]
- Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM III, Hao Y, Stoeckius M, Smibert P, Satija R. 2019. Comprehensive integration of single-cell data. Cell 177: 1888–1902 e1821. 10.1016/j.cell.2019.05.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willson JK, Albert DM, Diener-West M, McCaffrey L, Moy CS, Scully RE. 2001. Assessment of metastatic disease status at death in 435 patients with large choroidal melanoma in the Collaborative Ocular Melanoma Study (COMS) COMS Report no. 15. Arch Ophthalmol 119: 670–676. 10.1001/archopht.119.5.670 [DOI] [PubMed] [Google Scholar]
- Yu MC, Chen CH, Liang X, Wang L, Gandhi CR, Fung JJ, Lu L, Qian S. 2004. Inhibition of T-cell responses by hepatic stellate cells via B7-H1-mediated T-cell apoptosis in mice. Hepatology 40: 1312–1321. 10.1002/hep.20488 [DOI] [PubMed] [Google Scholar]
- Zhao J, Zhang S, Liu Y, He X, Qu M, Xu G, Wang H, Huang M, Pan J, Liu Z, et al. 2020. Single-cell RNA sequencing reveals the heterogeneity of liver-resident immune cells in human. Cell Discov 6: 22. 10.1038/s41421-020-0157-z [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.