Keywords: Bmal1, cell cycle, fat accumulation, hepatocyte proliferation, liver regeneration
Abstract
After partial hepatectomy (PH), the majority of remnant hepatocytes synchronously enter and rhythmically progress through the cell cycle for three major rounds to regain lost liver mass. Whether and how the circadian clock core component Bmal1 modulates this process remains elusive. We performed PH on Bmal1+/+ and hepatocyte-specific Bmal1 knockout (Bmal1hep−/−) mice and compared the initiation and progression of the hepatocyte cell cycle. After PH, Bmal1+/+ hepatocytes exhibited three major waves of nuclear DNA synthesis. In contrast, in Bmal1hep−/− hepatocytes, the first wave of nuclear DNA synthesis was delayed by 12 h, and the third such wave was lost. Following PH, Bmal1+/+ hepatocytes underwent three major waves of mitosis, whereas Bmal1hep−/− hepatocytes fully abolished mitotic oscillation. These Bmal1-dependent disruptions in the rhythmicity of hepatocyte cell cycle after PH were accompanied by suppressed expression peaks of a group of cell cycle components and regulators and dysregulated activation patterns of mitogenic signaling molecules c-Met and epidermal growth factor receptor. Moreover, Bmal1+/+ hepatocytes rhythmically accumulated fat as they expanded following PH, whereas this phenomenon was largely inhibited in Bmal1hep−/− hepatocytes. In addition, during late stages of liver regrowth, Bmal1 absence in hepatocytes caused the activation of redox sensor Nrf2, suggesting an oxidative stress state in regenerated liver tissue. Collectively, we demonstrated that during liver regeneration, Bmal1 partially modulates the oscillation of S-phase progression, fully controls the rhythmicity of M-phase advancement, and largely governs fluctuations in fat metabolism in replicating hepatocytes, as well as eventually determines the redox state of regenerated livers.
NEW & NOTEWORTHY We demonstrated that Bmal1 centrally controls the synchronicity and rhythmicity of the cell cycle and lipid accumulation in replicating hepatocytes during liver regeneration. Bmal1 plays these roles, at least in part, by ensuring formation of the expression peaks of cell cycle components and regulators, as well as the timing and levels of activation of mitogenic signaling molecules.
INTRODUCTION
After liver resection or acute liver injury, remnant hepatocytes synchronously enter the cell cycle and continuously undergo three major waves of replication to restore the original liver mass (1). This process is driven by the highly coordinated activation of a variety of extrahepatic and intrahepatic signals and pathways, which orchestrate the timely and dynamic expression of cell cycle regulators and components (2). However, how the synchronicity and rhythmicity of cell cycle progression is regulated remains elusive.
The core component of the mammalian circadian clock is composed of Bmal1 and Clock. These two transcription factors form a heterodimer, operate a set of transcriptional/translational feedback loops, generate diurnal outputs of the cellular transcriptome, and ultimately govern various rhythmic biological processes (3, 4). Of note, the circadian clock interacts with the cell cycle to regulate tissue renewal and repair (5). Several lines of evidence, including our previous work, support the notion that the circadian clock controls the timing of hepatocyte mitosis during liver regeneration (1, 6, 7). However, it remains unclear whether and how Bmal1 plays such a role. We aimed to determine how loss of Bmal1 function affects the initiation and progression of the cell cycle in replicating hepatocytes during the entire period of liver regrowth after partial hepatectomy (PH).
MATERIALS AND METHODS
Animal Care and Use
Hepatocyte-specific Bmal1 knockout mice were generated by crossing Bmal1flox/flox mice (Stock No. 009100, The Jackson Laboratory, Bar Harbor, ME) with Albumin-Cre mice (Stock No. 003574, The Jackson Laboratory). Mice were housed in plastic cages at 22 ± 1°C on a 12-h light/12-h dark cycle with lights on from 6:00 AM to 6:00 PM. Standard rodent chow and water were provided ad libitum throughout the feeding period. Male mice (3–4 mo old) were subjected to standard two-thirds PH with intact gall bladders following a procedure described previously (8, 9). The surgery was performed in the morning between 10:00 AM and 12:00 PM, with consistent surgical time on all mice (6). BrdU was injected (100 mg/kg ip) 1 h before euthanization at each time point. The isolated livers were weighed. Liver pieces were fixed in 10% neutral buffered formalin, embedded in optimal cutting temperature compound (Tissue Tek, Torrance, CA), or frozen in liquid nitrogen for various subsequent analyses. All animal experiments were conducted in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals. Protocols for the care and use of animals were approved by the Indiana University-Purdue University Indianapolis Animal Care and Use Committee.
Histology and Immunohistochemistry
Formalin-fixed and paraffin-embedded liver sections were subjected to Ki-67, BrdU, or p-histone H3 immunostaining to visualize and count proliferating hepatocytes. Primary antibodies against Ki-67 (No. RM-9106, Thermo Fisher Scientific, Waltham, MA), BrdU (No. 5292, Cell Signaling Technology, Danvers, MA), or p-histone H3 (No. 9764, Cell Signaling Technology) were used for immunostaining. Ki67-, BrdU-, or p-histone H3-positive hepatocytes were counted in five randomly chosen microscopic fields per section at ×200 magnification using Image-Pro Plus software (Media Cybernetics, Rockville, MD). Oil Red O (Sigma-Aldrich Co., St. Louis, MO) staining was performed on frozen liver sections to visualize intracellular lipid droplets. Stained areas were analyzed in five randomly chosen microscope fields at ×200 magnification per section using Image-Pro Plus software.
Western Blot Analysis
Liver homogenates (10 or 30 µg) were separated by polyacrylamide gel electrophoresis under reducing conditions. Proteins from the gels were electrophoretically transferred onto polyvinylidene difluoride (PVDF) membranes. Antibodies against cyclin D1 (No. 2922), cyclin B1 (No. 4138), and glyceraldehyde 3-phosphate dehydrogenase (GADPH) (No. 5174) were purchased from Cell Signaling Technology; Bmal1 (SC-373955), Wee1 (SC-9037), p-Cdc2 p34 (Tyr15) (SC-7898), and Cdc2 p34 (SC-54) from Santa Cruz Biotechnology (Dallas, TX); cyclin A2 (ab32386), c-Met (ab47431), p-cMet (ab5662), and NQO1 (ab80588) from Abcam (Cambridge, MA); and epidermal growth factor receptor (EGFR) (No. 1114-1) and p-EGFR (No. 1139) from Epitomics (Burlingame, CA). Immune complexes were detected using an enhanced chemiluminescence system (Pierce, Rockford, IL).
Quantitative Real-Time Polymerase Chain Reaction
Total RNA was isolated from frozen liver tissue using TRIzol reagent according to the manufacturer’s protocol (Invitrogen, Carlsbad, CA). cDNA was synthesized from total RNA (1 µg) of each sample using a Verso cDNA Kit (Thermo Fisher Scientific), diluted fourfold with water, and subjected to real-time quantitative PCR (RT-qPCR) to quantify mRNA levels. TaqMan Universal PCR Master Mix, PCR primers, and TaqMan MGB probes specific for mouse Bmal1 (Mm01269610_m1), Dbp (Mm00497539_m1), cyclin A2 (Mm00438063_m1), cyclin B1 (Mm03053893_gh), Aurka (Mm01248177_m1), Aurkb, (Mm01718146_gl), Prc1 (Mm01320564_m1), Wee1 (Mm00494175_m1), Per1 (Mm00501813_m1), Per2 (Mm00478099_m1), Elovl6 (Mm00851223_s1), Fads1 (Mm00507605_m1), Fads2 (Mm00517221_m1), Srebf1 (Mm00550338_m1), Ppara (Mm00440939_m1), Pparg (Mm00440940_m1), Cd36 (Mm00432403_m1), Fasn (Mm00662319_m1), Cry1 (Mm00514392_m1), Cry2 (Mm00546062_m1), and Ppia (Mm02342430_gl) were purchased from Applied Biosystems (Foster City, CA). RT-qPCR amplification reactions were carried out using the ABI Prism 7900 sequence detection system (Applied Biosystems) with an initial hold step (50°C for 2 min followed by 95°C for 10 min) and 40 cycles of a two-step PCR thermocycling protocol (92°C for 15 s and 60°C for 1 min). The comparative threshold cycle (CT) method was used for relative quantification of the amount of mRNA in each sample normalized to Ppia transcript levels.
Statistical Analysis
Data are shown as means ± standard deviation (SD). Statistical analyses were performed using one-way analysis of variance. Comparisons of means were determined using post hoc analysis. Significant differences were defined as P < 0.05.
RESULTS
Hepatocyte-Specific Bmal1 Knockout Disrupts S-Phase Progression and Abolishes Mitotic Waves in Replicating Hepatocytes in Regenerating Livers
For simplicity, we define Bmal1+/+;Albumin-Cre+ mice as Bmal1+/+ mice and Bmal1flox/flox;Albumin-Cre+ mice as hepatocyte-specific Bmal1 knockout (Bmal1hep−/−) mice.
We performed PH on Bmal1+/+ and Bmal1hep−/− mice and subsequently conducted the following assessments at various time points within a 7-day period. We first quantified Ki67-positive hepatocytes in regenerated liver tissue. Because Ki67 is expressed in all phases of the cell cycle, we used this marker to estimate the total number of cycling hepatocytes. We observed Bmal1-dependent alterations in this parameter at multiple time points after PH (Fig. 1). These data indicated that Bmal1 deficiency in hepatocytes affects PH-induced hepatocyte proliferation.
Figure 1.
Numbers of cycling hepatocytes after partial hepatectomy (PH). Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to PH and euthanized at the indicated time points. Ki-67 immunostaining was performed with liver sections. A: representative liver sections showing Ki67-positive hepatocytes. B: Ki67-positive hepatocytes were counted at ×200 magnification in five randomly chosen fields per section. The results are shown as mean numbers per field ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01).
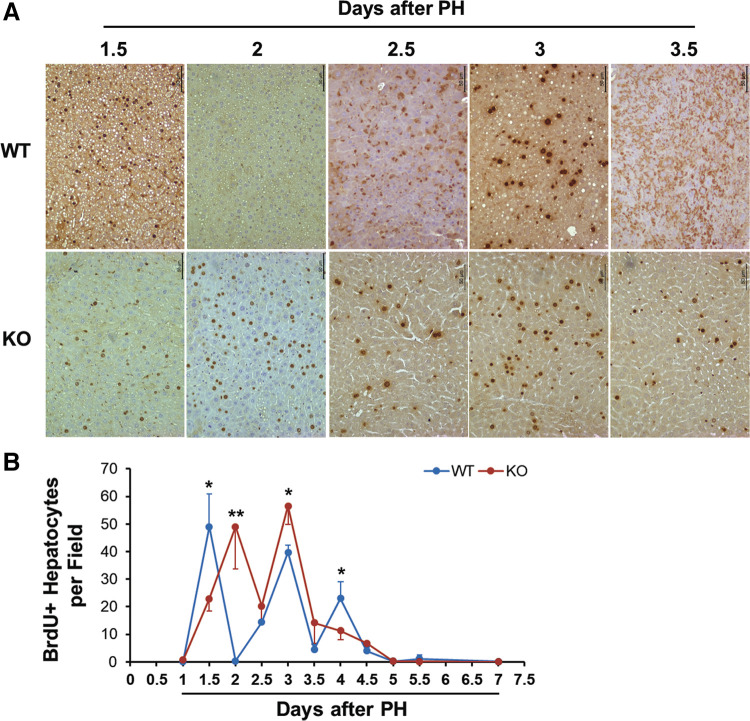
To gain insight into the S-phase progression of proliferating hepatocytes, we injected bromodeoxyuridine (BrdU) into mice 1 h before each time point and quantified BrdU-positive (S-phase) hepatocytes (Fig. 2). As expected, Bmal1+/+ regenerating livers underwent three major waves of DNA synthesis, which consecutively occurred and progressively weakened at days 1.5, 3, and 4 post-PH. In contrast, in Bmal1hep−/− regenerating livers, the first wave of DNA synthesis was delayed by 12 h, the second wave occurred in a timely manner, and the third wave disappeared. Cycling Bmal1hep−/− hepatocytes exhibited dysregulated S-phase progression following PH.
Figure 2.
Numbers of hepatocytes undergoing DNA synthesis after partial hepatectomy (PH). Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to PH and euthanized at the indicated time points. One hour before mice were euthanized, BrdU was injected (100 mg/kg ip). Liver sections were subjected to BrdU immunostaining. A: representative liver sections showing BrdU-positive hepatocytes. B: BrdU-positive hepatocytes were counted at ×200 magnification in 5 randomly chosen fields per section. The data are shown as the mean numbers per field ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01). BrdU, bromodeoxyuridine.
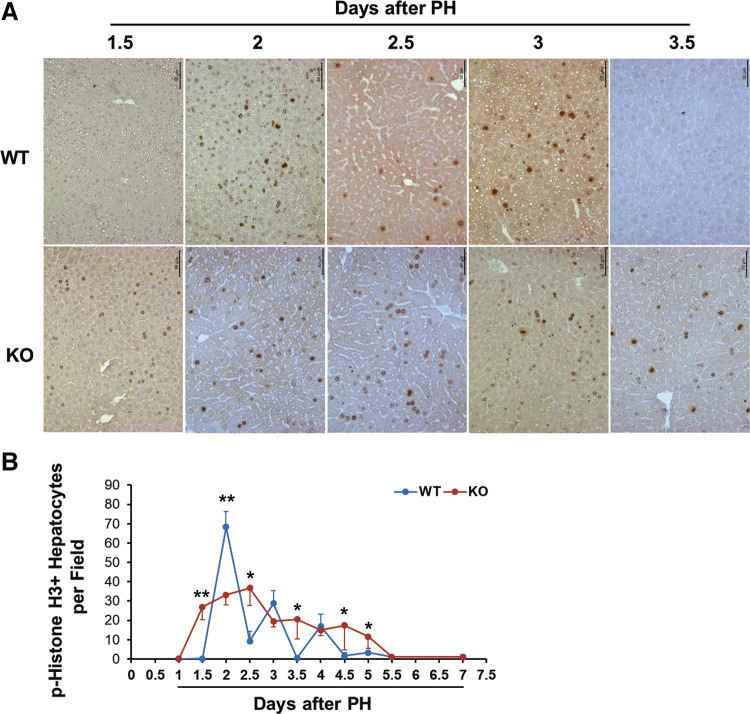
We next examined the M-phase progression of replicating hepatocytes by quantifying hepatocytes positive for p-histone H3, an M-phase cell cycle marker (Fig. 3). As anticipated, Bmal1+/+ regenerating hepatocytes displayed three major waves of mitosis, which were sequentially formed and gradually reduced at days 2, 3, and 4 after PH. In sharp contrast, Bmal1hep−/− regenerating hepatocytes completely lost the oscillation in M-phase progression. Even on day 1.5 following PH, when Bmal1+/+ cycling hepatocytes were undergoing the first S phase (Fig. 2), a significant number of cycling Bmal1hep−/− hepatocytes had already entered the M phase. Thus, Bmal1 is essential for the formation of hepatocyte mitotic waves during PH-induced liver regrowth.
Figure 3.
Numbers of hepatocytes undergoing mitosis after partial hepatectomy (PH). Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to PH and euthanized at the indicated time points. Liver sections were immunostained with anti-p-Histone H3 antibody. A: representative liver sections show p-Histone H3-positive hepatocytes. B: p-Histone H3-positive hepatocytes were counted at ×200 magnification in five randomly chosen fields per section. The data are shown as means per field ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01).
Hepatocyte-Specific Bmal1 Knockout Results in Reduced Fat Accumulation in Regenerating Livers
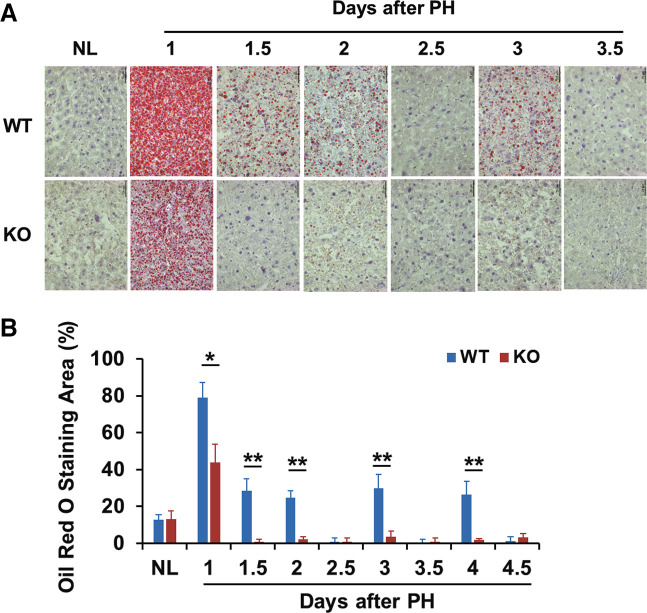
It is known that after PH, replicating hepatocytes dynamically accumulate fat, which is coupled to cell cycle progression (1). We conducted Oil Red O staining and measured the staining areas in the two genotypes of regenerating livers (Fig. 4). As a result, Bmal1+/+ regenerating livers showed massive fat deposits at days 1, 1.5, 3, and 4 following PH. In contrast, Bmal1hep−/− regenerating livers displayed reduced fat staining at day 1 post-PH and largely diminished fat content thereafter. Therefore, regenerating hepatocytes require Bmal1 to dynamically govern lipid metabolism.
Figure 4.
Fat accumulation in regenerating livers. Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3-4-mo-old male mice were subjected to partial hepatectomy (PH) and euthanized at the indicated time points. Frozen liver sections were prepared and used for Oil red O staining. A: representative liver sections show Oil Red O staining. B: the stained areas were quantified in five randomly chosen microscopic fields per liver section (×200 magnification) with Image-Pro software. Data are presented as the means of percent stained areas ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01).
Hepatocyte-Specific Bmal1 Knockout Leads to Suppression of Expression Peaks in a Subset of Cell Cycle Components and Regulators in Regenerating Livers
We found that in Bmal1+/+ mice subjected to PH, hepatic Bmal1 mRNA expression fluctuated approximately fivefold (Fig. 5A). To evaluate Bmal1 transcriptional activity, we quantified transcript levels of hepatic Dbp, a direct target gene of Bmal1 (10). As a result, in Bmal1+/+ regenerating livers, Dbp mRNA exhibited oscillatory expression with an approximate fourfold magnitude, which was inversely correlated with Bmal1 mRNA levels, as anticipated, whereas in Bmal1hep−/− regenerating livers, Dbp mRNA expression lost such oscillation (Fig. 5A). Bmal1 is a suppressor of Per1, Per2, Cry1, and Cry2, which are additional components of the circadian clock (11). The transcript levels of Per1, Per2, Cry1, and Cry2 fluctuated as liver regrowth advanced in Bmal1+/+ mice. Bmal1 deficiency increased mRNA expression of Per2 and Cry1, but mildly affected Per1 and Cry2, in regenerating livers (Fig. 5A). These data indicated that the circadian clock was fully operating during hepatic regeneration.
Figure 5.
mRNA expression of a subset of circadian clock components and cell cycle components and regulators in regenerating liver. Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to partial hepatectomy (PH) and euthanized at the indicated time points. Total liver RNA samples were prepared and used for quantifying mRNA levels of the genes indicated by RT-qPCR. A: mRNA expression of a subset of circadian clock components. B: mRNA expression of a subset of cell cycle components and regulators. Data are expressed as the mean fold change relative to mRNA levels pre-PH in WT mice ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01; ***P < 0.001).
To examine cell cycle progression in cycling hepatocytes at the molecular level, we assessed the expression of a subset of genes encoding cell cycle components and regulators in regenerating livers (Fig. 5B). In partially hepatectomized Bmal1+/+ mice, hepatic mRNA expression of cyclin A2, cyclin B1, aurora kinase A (Aurka), aurora kinase B (Aurkb), and protein regulator of cytokinesis 1 (Prc1) consistently exhibited peak expression at day 2 post-PH. In the absence of Bmal1, the formation of these peaks was completely abrogated. Thus, Bmal1 is required for these genes to reach peaks in terms of their mRNA expression in regrowing livers. Exceptionally, the formation of the expression peak of the Wee1 gene, which encodes a kinase controlling cell mitosis, was not dependent on Bmal1 (Fig. 5B). At the protein level, cyclin D1 was persistently expressed from day 1 to day 7 after PH in both liver genotypes. Overall, Bmal1hep−/− livers expressed more cyclin D1 at most time points post-PH than Bmal1+/+ control livers (Fig. 6). Notably, the expression of cyclin A2, cyclin B1, and p-Cdc2 was oscillatory and most abundant on days 2 and 4 following PH in Bmal1+/+ regenerating livers, whereas these oscillations were largely repressed in Bmal1hep−/− regenerating livers (Fig. 6). Taken together, these data demonstrate that Bmal1 exhibits rhythmic activity and determines the formation of expression peaks of a group of cell cycle components and regulators as liver regrowth advances.
Figure 6.
Protein expression of a subset of cell cycle components and mitogenic signaling molecules in regenerating liver. Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to partial hepatectomy (PH) and euthanized at the indicated time points. Liver lysates prepared from 5 mice per time point per genotype were pooled with equal amount of proteins from each preparation. Western blotting was performed using antibodies against the proteins indicated. Glyceraldehyde 3-phosphate dehydrogenase (GADPH) was used as a loading control. Relative densitometry was normalized against GAPDH. NL, normal liver.
Hepatocyte-Specific Bmal1 Knockout Causes Dysregulated Activation of Mitogenic Signaling Molecules in Regenerating Livers
cMet and epidermal growth factor receptor (EGFR) are the two most critical mitogenic signaling molecules involved in liver regeneration (12). We found that in Bmal1+/+ mice, relative to the pre-PH state, hepatic c-Met displayed increased activity (phosphorylation) during the first 3 days following PH. In Bmal1hep−/− mice, compared with Bmal1+/+ mice, hepatic c-Met exhibited markedly (2.6-fold) higher basal activity pre-PH and an altered activation pattern post-PH (Fig. 6). In Bmal1+/+ regrowing livers, EGFR was activated most prominently at day 3, whereas in Bmal1hep−/− regrowing livers, this molecule was activated at multiple time points (days 1.5, 3, and 5) and post-PH (Fig. 6). Together, we demonstrated that Bmal1 determines the basal activity of c-Met in homeostatic livers and the timing of c-Met and EGFR activation in regenerating livers.
Hepatocyte-Specific Bmal1 Knockout Disrupts the Expression Patterns of Genes Involved in Lipid Metabolism in Regenerating Livers
We examined the expression patterns of a group of genes known to participate in lipid metabolism in regrowing livers (Fig. 7). Rev-erbα is a state-dependent regulator of liver energy metabolism and serves to buffer against metabolic challenge (13). Bmal1+/+ mice showed persistently suppressed, whereas Bmal1hep−/− mice displayed further suppressed, mRNA expression of hepatic Rev-erbα following PH. Peroxisome proliferator-activated receptor-γ (Pparg) promotes fatty acid uptake and triglyceride synthesis and storage (14). Cd36 is a free fatty acid transporter responsible for the uptake of fatty acid (15). Fatty acid desaturase 1 (Fads1) is involved in long-chain polyunsaturated fatty acid metabolism (16). During the first 1.5 days after PH, hepatic expression of the genes encoding these three proteins was inhibited due to the absence of Bmal1. Pparα increases fatty acid uptake, esterification, and trafficking, as well as lipid oxidation (14, 17). Fatty acid synthase (Fasn) catalyzes fatty acid synthesis (18). Three days after PH, without Bmal1, the transcript levels of hepatic Pparα, Pparg, and Fasn were all downregulated. Sterol regulatory element-binding transcription factor 1 (SREBF1) induces lipogenesis in the liver (19). The expression of Srebf1, as well as Fads2, did not show a Bmal1-dependent difference in regenerating livers. Elovl6 encodes elongation of very long chain fatty acids protein 6 and is a target gene of Srebf1. The lack of Bmal1 caused a transient upregulation of hepatic Elovl6 mRNA expression on day 2 after PH. Taken together, these results demonstrated that Bmal1 is required for maintaining PH-dependent expression patterns of a subset of genes critical for lipid metabolism.
Figure 7.
mRNA expression of a group of genes involved in lipid metabolism in regenerating liver. Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to partial hepatectomy (PH) and euthanized at the indicated time points. Total liver RNA samples were prepared and used for quantifying mRNA levels of the genes indicated by RT-qPCR. Data are expressed as the mean fold change relative to mRNA levels pre-PH in WT mice ± SD (n = 5 mice/time point/genotype; *P < 0.05; **P < 0.01; ***P < 0.001).
Hepatocyte-Specific Bmal1 Knockout Causes Nrf2 Activation during Late Stages of Liver Regrowth
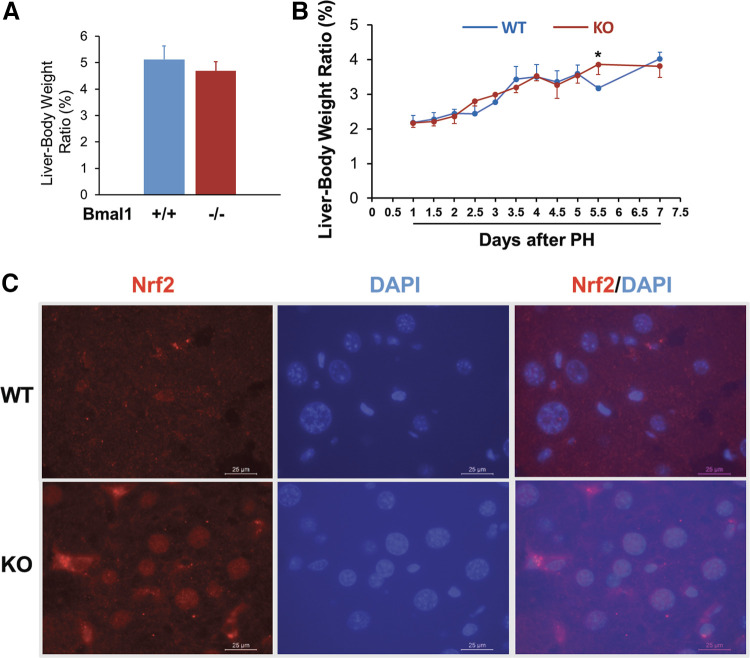
We calculated liver-to-body weight ratios to estimate gross regrowth of resected livers. The liver-to-body weight ratios were not significantly different between Bmal1+/+ and Bmal1hep−/− mice pre-PH (Fig. 8A), consistent with others’ report (20). Bmal1hep−/− mice regained more liver mass than Bmal1+/+ mice by day 5.5 post-PH. Overall, the two genotype groups of mice displayed comparable liver mass recovery within the 7-day period post-PH (Fig. 8B). Notably, at day 3.5 and thereafter following PH, Bmal1hep−/− livers persistently expressed higher levels of NAD(P)H:quinone oxidoreductase 1 (NQO1) than Bmal1+/+ livers (Fig. 6). NQO1 is a direct target of the transcription factor nuclear factor erythroid 2-related factor 2 (Nrf2), a redox sensor (21). In response to oxidative stress, Nrf2 translocates into the nucleus and transactivates a battery of cellular defense genes (22). Thus, we examined the intracellular distribution of Nrf2 in hepatocytes at day 7 post-PH. As a result, we observed overt nuclear translocation of Nrf2 in Bmal1hep−/− hepatocytes compared with Bmal1+/+ hepatocytes (Fig. 8C). These results suggested that absence of Bmal1 in hepatocytes caused oxidative stress as liver regrowth progressed to the later stages, persisting in finally regenerated livers.
Figure 8.
A: liver-to-body weight ratios before partial hepatectomy (PH). B: liver regrowth patterns after PH. Bmal1+/+ (WT) and hepatocyte-specific Bmal1 knockout (KO) 3- to 4-mo-old male mice were subjected to PH and euthanized at the indicated time points. Liver and body weights were recorded. Liver-to-body weight ratio was used as a liver regrowth index. The results are presented as mean liver-to-body weight ratio ± SD (n = 5 mice/time point/genotype; *P < 0.05). C: Nrf2 immunofluorescence analysis was performed on liver sections prepared from formalin-fixed and paraffin-embedded liver tissues collected from mice euthanized at day 7 post-PH. Representative liver sections show Nrf2 (red, cytosol or nucleus) and DAPI (blue, nucleus).
DISCUSSION
Our studies demonstrated that Bmal1 governs the formation of waves of hepatocyte replication by ensuring timely activation of mitogenic signaling and peak expression of cell cycle regulators and components during liver regeneration. Bmal1 loss of function in hepatocytes caused early entry to the cell cycle, disrupted synchronicity of S-phase progression, and, most strikingly, fully lost rhythmicity of mitosis in cycling hepatocytes in regenerating livers. As early as 36 h post-PH, a significant number of hepatocytes deficient for Bmal1 had already been in the M phase, whereas most hepatocytes sufficient for Bmal1 were only in the first S phase. This initial early entry of hepatocytes into the cell cycle may be due to increased basal activity of c-Met in Bmal1hep−/− livers. Here, for the first time, we linked Bmal1 to c-Met activity in the liver under homeostatic conditions. This finding will prompt us and others to delineate how this occurs because dysregulation of the circadian clock is associated with tumor incidence (23). The absence of Bmal1 impaired S-phase progression but did not completely eliminate the formation of DNA synthesis peaks in cycling hepatocytes. This indicated the existence of other agent(s) that more critically control the rhythmicity of DNA synthesis in this setting. Without Bmal1, replicating hepatocytes were unable to form any waves of mitosis in regenerating livers. Thus, these cells rely fully on Bmal1 for timely division. Bmal1 plays an essential role in regulating cyclin A2 and the Cdc2/cyclin B1 pathway. It is well known that the timely accumulation and degradation of cyclin A is required for M-phase progression (24, 25) and that the activity of the Cdc2/cyclin B complex modulates G2/M transition in dividing cells (26). We observed that the expression of cyclin A2, cyclin B1, and p-Cdc2 similarly exhibited large magnitudes of oscillation in Bmal1+/+ regenerating livers, whereas these oscillations largely disappeared owing to the absence of Bmal1. It was proposed by others that Wee1 is a direct target gene of Bmal1, and the Bmal1-Clock/Wee1/Cdc2 pathway determines the timing of hepatocyte division in liver regeneration (6). However, we found that the peak formation of Wee1 mRNA expression was not affected by the loss of function of Bmal1 in regenerating hepatocytes, which does not support this theory. It is known that c-Met and EGFR together mediate mitogenic signaling essential for hepatocyte proliferation, because their combined loss completely abolishes liver regeneration (12). We found that Bmal1 is required to orchestrate the timed activation of c-Met and EGFR as liver regrowth advances. Aurka, Aurkb, and Prc1 are critical regulators of cell division (27, 28). We showed that Bmal1 is essential for reaching peaks in their expression. Collectively, we propose that Bmal1 centrally coordinates the timing and levels of the activity of mitogenic signaling molecules and the expression of cell cycle regulators and components, thus driving the synchronicity and rhythmicity of the initiation and progression of the cell cycle of cycling hepatocytes during liver regeneration. Of note, a very recent report shows that in Bmal1 knockout mice, the skin and liver still exhibit 24-h oscillations of the transcriptome and proteome without daily light (29). The report proposed that erythroblast transformation-specific (ETS) transcription factor family and the redox system may support the oscillations independent of Bmal1. Thus, it is highly likely that these factors may partially compensate for the loss of Bmal1 during liver regeneration, which needs to be further investigated.
Proliferating hepatocytes accumulate fat (steatosis), which is a naturally occurring event, as the liver is regenerating. Excessive lipids provide cellular components for newly regenerated hepatocytes and act as signaling molecules to promote liver regrowth (30). Our previous work demonstrated that cycling hepatocytes rhythmically accumulate fat, which is closely linked to hepatocyte mitosis after PH (1). Here, we revealed that without Bmal1 in hepatocytes, initial fat deposits were reduced, and subsequent fat accumulation was lost as liver regrowth progressed. Thus, proliferating hepatocytes largely rely on Bmal1 to accumulate lipids. How the steatosis is formed in regenerating liver remains unclear. Our data suggest that Bmal1 coordinates the expression of genes involved in lipid uptake, synthesis, storage, and oxidation. The consequence of loss of such coordination is the reduction or prevention of lipid accumulation in regenerating liver. Of note, a report showed that liver-specific Bmal1 knockout mice display an increased steatosis phenotype with suppressed de novo lipogenesis and fatty acid oxidation in alcohol liver disease (31). Therefore, our findings revealed a liver regeneration-dependent role of Bmal1 in lipid metabolism.
Liver regeneration requires highly coordinated and complex interactions between different liver cell types (32). For instance, after PH, Kupffer cells produce proinflammatory cytokines such as IL-6 to prime hepatocytes to respond to hepatocyte growth factor, which is elaborated by hepatic stellate cells (HSCs), stimulating hepatocytes to enter the cell cycle first. In turn, replicating hepatocytes release growth factors such as platelet-derived growth factor to drive nonparenchymal cell proliferation, which occurs 24 h later than that of hepatocytes. It is known that the circadian clock operates in liver nonparenchymal cells and regulates their functions (33–35). Bmal1 controls glycolysis, cell cycle progression, and fibrotic phenotypes in HSCs (34). Bmal1 modulates inflammatory cytokine production, glycolysis, and polarization of Kupffer cells, but the loss of Bmal1 in these cells does not affect pathogenesis of alcoholic liver disease (31, 35, 36). Bmal1 is also expressed in cholangiocytes and regulates their hyperplasia in cholestasis (33, 37). These pieces of evidence support important roles of Bmal1 in liver nonparenchymal cells in homeostasis and disease. Here we demonstrate that the loss of function of Bmal1 disrupts the cell cycle progression and lipid metabolism in hepatocytes. Whether and how liver nonparenchymal cells are affected in this setting needs to be investigated.
Our studies showed that although hepatocytes deficient in Bmal1 lose cell cycle rhythm following PH, they are still able to expand, eventually recovering the lost liver mass. However, the regenerated livers are in an oxidative stress state, which is manifested by Nrf2 activation during the later stage of liver regrowth. It has been reported that Bmal1 directly regulates Nrf2 expression and activity in β-cells (38), macrophages (39), lung (40), and kidney (41). The lack of Bmal1 causes diminished Nrf2 activity and oxidative stress in these cells and organs. It has also been shown that in hepatocytes, Nrf2 activation directly upregulates the expression of the clock repressor gene Cry2, thereby repressing Bmal1/Clock-regulated E-box transcription (42). These reports outline the integration of the circadian clock with redox balance in various cell types. NQO1 expression is solely controlled by Nrf2 during liver injury and regeneration, and serves as a reliable index of Nrf2 activity (21, 43). We found that compared with Bmal1+/+ mice, hepatic Nrf2 activity was slightly lower pre-PH, largely comparable during the first three days after PH, and persistently higher 3.5 days post-PH in hepatocyte-specific Bmal1 knockout mice. These observations suggested that oxidative stress in regenerated livers is not directly and fundamentally caused by the lack of Bmal1 in hepatocytes. Instead, it was a new event that occurred following disruption of the hepatocyte cell cycle. Therefore, it is highly likely that this is a consequence of the arrhythmic cell cycle in Bmal1-deficient proliferating hepatocytes.
In summary, we demonstrated that after PH, fluctuating Bmal1 activity partially drives synchronous and rhythmic S-phase progression and fully controls synchronous and rhythmic M-phase progression of cycling hepatocytes, which eventually restores the redox balance in regenerated livers. Mechanistically, Bmal1 ensures the formation of the expression peaks of a group of cell cycle components and regulators, coordinates the timely activation of mitogenic signaling molecules, and modulates waves of fat accumulation as liver regeneration progresses.
GRANTS
This research was supported by National Institutes of Health Grant 1R01DK117076.
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the authors.
AUTHOR CONTRIBUTIONS
H.J. and G.D. conceived and designed research; H.J., V.G., J.A.Y., and J.L. performed experiments; H.J., J.A.Y., and J.L. analyzed data; H.J. interpreted results of experiments; H.J., V.G., and J.A.Y. prepared figures; H.J. and G.D. drafted manuscript; G.D. edited and revised manuscript; H.J. and G.D. approved final version of manuscript.
REFERENCES
- 1.Zou Y, Bao Q, Kumar S, Hu M, Wang GY, Dai G. Four waves of hepatocyte proliferation linked with three waves of hepatic fat accumulation during partial hepatectomy-induced liver regeneration. PLoS One 7: e30675, 2012. doi: 10.1371/journal.pone.0030675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Michalopoulos GK, Bhushan B. Liver regeneration: biological and pathological mechanisms and implications. Nat Rev Gastroenterol Hepatol 18: 40–55, 2021. doi: 10.1038/s41575-020-0342-4. [DOI] [PubMed] [Google Scholar]
- 3.Ayyar VS, Sukumaran S. Circadian rhythms: influence on physiology, pharmacology, and therapeutic interventions. J Pharmacokinet Pharmacodyn 48: 321–338, 2021. doi: 10.1007/s10928-021-09751-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.de Assis LVM, Oster H. The circadian clock and metabolic homeostasis: entangled networks. Cell Mol Life Sci 78: 4563–4587, 2021. doi: 10.1007/s00018-021-03800-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ruby CL, Major RJ, Hinrichsen RD. Regulation of tissue regeneration by the circadian clock. Eur J Neurosci 53: 3576–3597, 2021. doi: 10.1111/ejn.15244. [DOI] [PubMed] [Google Scholar]
- 6.Matsuo T, Yamaguchi S, Mitsui S, Emi A, Shimoda F, Okamura H. Control mechanism of the circadian clock for timing of cell division in vivo. Science 302: 255–259, 2003. doi: 10.1126/science.1086271. [DOI] [PubMed] [Google Scholar]
- 7.Souto M, Llanos JM. The circadian optimal time for hepatectomy in the study of liver regeneration. Chronobiol Int 2: 169–175, 1985. doi: 10.3109/07420528509055556. [DOI] [PubMed] [Google Scholar]
- 8.Dai G, He L, Bu P, Wan YJ. Pregnane X receptor is essential for normal progression of liver regeneration. Hepatology 47: 1277–1287, 2008. doi: 10.1002/hep.22129. [DOI] [PubMed] [Google Scholar]
- 9.Greene AK, Puder M. Partial hepatectomy in the mouse: technique and perioperative management. J Invest Surg 16: 99–102, 2003. doi: 10.1080/713829590. [DOI] [PubMed] [Google Scholar]
- 10.Zhang YK, Yeager RL, Klaassen CD. Circadian expression profiles of drug-processing genes and transcription factors in mouse liver. Drug Metab Dispos 37: 106–115, 2009. doi: 10.1124/dmd.108.024174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Guan D, Lazar MA. Interconnections between circadian clocks and metabolism. J Clin Invest 131: e148278, 2021. doi: 10.1172/JCI148278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Paranjpe S, Bowen WC, Mars WM, Orr A, Haynes MM, DeFrances MC, Liu S, Tseng GC, Tsagianni A, Michalopoulos GK. Combined systemic elimination of MET and epidermal growth factor receptor signaling completely abolishes liver regeneration and leads to liver decompensation. Hepatology 64: 1711–1724, 2016. doi: 10.1002/hep.28721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hunter AL, Pelekanou CE, Adamson A, Downton P, Barron NJ, Cornfield T, Poolman TM, Humphreys N, Cunningham PS, Hodson L, Loudon ASI, Iqbal M, Bechtold DA, Ray DW. Nuclear receptor REVERBα is a state-dependent regulator of liver energy metabolism. Proc Natl Acad Sci USA 117: 25869–25879, 2020. doi: 10.1073/pnas.2005330117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Montaigne D, Butruille L, Staels B. PPAR control of metabolism and cardiovascular functions. Nat Rev Cardiol 2021. doi: 10.1038/s41569-021-00569-6. [DOI] [PubMed] [Google Scholar]
- 15.Bonen A, Campbell SE, Benton CR, Chabowski A, Coort SL, Han XX, Koonen DP, Glatz JF, Luiken JJ. Regulation of fatty acid transport by fatty acid translocase/CD36. Proc Nutr Soc 63: 245–249, 2004. doi: 10.1079/PNS2004331. [DOI] [PubMed] [Google Scholar]
- 16.Lattka E, Illig T, Heinrich J, Koletzko B. FADS gene cluster polymorphisms: important modulators of fatty acid levels and their impact on atopic diseases. J Nutrigenet Nutrigenomics 2: 119–128, 2009. doi: 10.1159/000235559. [DOI] [PubMed] [Google Scholar]
- 17.Berger J, Moller DE. The mechanisms of action of PPARs. Annu Rev Med 53: 409–435, 2002. doi: 10.1146/annurev.med.53.082901.104018. [DOI] [PubMed] [Google Scholar]
- 18.Kuhajda FP, Pizer ES, Li JN, Mani NS, Frehywot GL, Townsend CA. Synthesis and antitumor activity of an inhibitor of fatty acid synthase. Proc Natl Acad Sci USA 97: 3450–3454, 2000. doi: 10.1073/pnas.050582897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Shimano H, Yahagi N, Amemiya-Kudo M, Hasty AH, Osuga J, Tamura Y, Shionoiri F, Iizuka Y, Ohashi K, Harada K, Gotoda T, Ishibashi S, Yamada N. Sterol regulatory element-binding protein-1 as a key transcription factor for nutritional induction of lipogenic enzyme genes. J Biol Chem 274: 35832–35839, 1999. doi: 10.1074/jbc.274.50.35832. [DOI] [PubMed] [Google Scholar]
- 20.Pati P, Valcin JA, Zhang D, Neder TH, Millender-Swain T, Allan JM, Sedaka R, Jin C, Becker BK, Pollock DM, Bailey SM, Pollock JS. Liver circadian clock disruption alters perivascular adipose tissue gene expression and aortic function in mice. Am J Physiol Regul Integr Comp Physiol 320: R960–R971, 2021. doi: 10.1152/ajpregu.00128.2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Aleksunes LM, Slitt AL, Maher JM, Dieter MZ, Knight TR, Goedken M, Cherrington NJ, Chan JY, Klaassen CD, Manautou JE. Nuclear factor-E2-related factor 2 expression in liver is critical for induction of NAD(P)H:quinone oxidoreductase 1 during cholestasis. Cell Stress Chaperones 11: 356–363, 2006. doi: 10.1379/csc-217.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Taguchi K, Yamamoto M. The KEAP1-NRF2 system as a molecular target of cancer treatment. Cancers (Basel) 13: 46, 2020. doi: 10.3390/cancers13010046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Allada R, Bass J. Circadian mechanisms in medicine. N Engl J Med 384: 550–561, 2021. doi: 10.1056/NEJMra1802337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Parry DH, O'Farrell PH. The schedule of destruction of three mitotic cyclins can dictate the timing of events during exit from mitosis. Curr Biol 11: 671–683, 2001. doi: 10.1016/S0960-9822(01)00204-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sullivan M, Morgan DO. Finishing mitosis, one step at a time. Nat Rev Mol Cell Biol 8: 894–903, 2007. doi: 10.1038/nrm2276. [DOI] [PubMed] [Google Scholar]
- 26.Fisher D, Krasinska L, Coudreuse D, Novák B. Phosphorylation network dynamics in the control of cell cycle transitions. J Cell Sci 125: 4703–4711, 2012. doi: 10.1242/jcs.106351. [DOI] [PubMed] [Google Scholar]
- 27.Fulcher LJ, Sapkota GP. Mitotic kinase anchoring proteins: the navigators of cell division. Cell Cycle 19: 505–524, 2020. doi: 10.1080/15384101.2020.1728014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jiang W, Jimenez G, Wells NJ, Hope TJ, Wahl GM, Hunter T, Fukunaga R. PRC1: a human mitotic spindle-associated CDK substrate protein required for cytokinesis. Mol Cell 2: 877–885, 1998. doi: 10.1016/s1097-2765(00)80302-0. [DOI] [PubMed] [Google Scholar]
- 29.Ray S, Valekunja UK, Stangherlin A, Howell SA, Snijders AP, Damodaran G, Reddy AB. Circadian rhythms in the absence of the clock gene Bmal1. Science 367: 800–806, 2020. doi: 10.1126/science.aaw7365. [DOI] [PubMed] [Google Scholar]
- 30.Delgado-Coello B, Briones-Orta MA, Macías-Silva M, Mas-Oliva J. Cholesterol: recapitulation of its active role during liver regeneration. Liver Int 31: 1271–1284, 2011. doi: 10.1111/j.1478-3231.2011.02542.x. [DOI] [PubMed] [Google Scholar]
- 31.Zhang D, Tong X, Nelson BB, Jin E, Sit J, Charney N, Yang M, Omary MB, Yin L. The hepatic BMAL1/AKT/lipogenesis axis protects against alcoholic liver disease in mice via promoting PPARα pathway. Hepatology 68: 883–896, 2018. doi: 10.1002/hep.29878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Michalopoulos GK. Liver regeneration. J Cell Physiol 213: 286–300, 2007. doi: 10.1002/jcp.21172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Renzi A, Glaser S, Demorrow S, Mancinelli R, Meng F, Franchitto A, Venter J, White M, Francis H, Han Y, Alvaro D, Gaudio E, Carpino G, Ueno Y, Onori P, Alpini G. Melatonin inhibits cholangiocyte hyperplasia in cholestatic rats by interaction with MT1 but not MT2 melatonin receptors. Am J Physiol Gastrointest Liver Physiol 301: G634–G643, 2011. [Erratum in Am J Physiol Gastrointest Liver Physiol 301: G943, 2011]. doi: 10.1152/ajpgi.00206.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Xu L, Yang TY, Zhou YW, Wu MF, Shen J, Cheng JL, Liu QX, Cao SY, Wang JQ, Zhang L. Bmal1 inhibits phenotypic transformation of hepatic stellate cells in liver fibrosis via IDH1/alpha-KG-mediated glycolysis. Acta Pharmacol Sin 2021. [Erratum in Acta Pharmacol Sin 2021]. doi: 10.1038/s41401-021-00658-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhou Y, Wu M, Xu L, Cheng J, Shen J, Yang T, Zhang L. Bmal1 regulates macrophage polarize through glycolytic pathway in alcoholic liver disease. Front Pharmacol 12: 640521, 2021. doi: 10.3389/fphar.2021.640521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Scheiermann C, Kunisaki Y, Frenette PS. Circadian control of the immune system. Nat Rev Immunol 13: 190–198, 2013. doi: 10.1038/nri3386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Han Y, Onori P, Meng F, DeMorrow S, Venter J, Francis H, Franchitto A, Ray D, Kennedy L, Greene J, Renzi A, Mancinelli R, Gaudio E, Glaser S, Alpini G. Prolonged exposure of cholestatic rats to complete dark inhibits biliary hyperplasia and liver fibrosis. Am J Physiol Gastrointest Liver Physiol 307: G894–G904, 2014. doi: 10.1152/ajpgi.00288.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lee J, Moulik M, Fang Z, Saha P, Zou F, Xu Y, Nelson DL, Ma K, Moore DD, Yechoor VK. Bmal1 and β-cell clock are required for adaptation to circadian disruption, and their loss of function leads to oxidative stress-induced β-cell failure in mice. Mol Cell Biol 33: 2327–2338, 2013. doi: 10.1128/MCB.01421-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Early JO, Menon D, Wyse CA, Cervantes-Silva MP, Zaslona Z, Carroll RG, Palsson-McDermott EM, Angiari S, Ryan DG, Corcoran SE, Timmons G, Geiger SS, Fitzpatrick DJ, O'Connell D, Xavier RJ, Hokamp K, O'Neill LAJ, Curtis AM. Circadian clock protein BMAL1 regulates IL-1β in macrophages via NRF2. Proc Natl Acad Sci USA 115: E8460–E8468, 2018. doi: 10.1073/pnas.1800431115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pekovic-Vaughan V, Gibbs J, Yoshitane H, Yang N, Pathiranage D, Guo B, Sagami A, Taguchi K, Bechtold D, Loudon A, Yamamoto M, Chan J, van der Horst GT, Fukada Y, Meng QJ. The circadian clock regulates rhythmic activation of the NRF2/glutathione-mediated antioxidant defense pathway to modulate pulmonary fibrosis. Genes Dev 28: 548–560, 2014. doi: 10.1101/gad.237081.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sun Q, Zeng C, Du L, Dong C. Mechanism of circadian regulation of the NRF2/ARE pathway in renal ischemia-reperfusion. Exp Ther Med 21: 190, 2021. doi: 10.3892/etm.2021.9622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wible RS, Ramanathan C, Sutter CH, Olesen KM, Kensler TW, Liu AC, Sutter TR. NRF2 regulates core and stabilizing circadian clock loops, coupling redox and timekeeping in Mus musculus. eLife 7: e31656, 2018. doi: 10.7554/eLife.31656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zou Y, Hu M, Lee J, Nambiar SM, Garcia V, Bao Q, Chan JY, Dai G. Nrf2 is essential for timely M phase entry of replicating hepatocytes during liver regeneration. Am J Physiol Gastrointest Liver Physiol 308: G262–G268, 2015. doi: 10.1152/ajpgi.00332.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]