Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused the pandemic of coronavirus disease 2019 (COVID-19). Great international efforts have been put into the development of prophylactic vaccines and neutralizing antibodies. However, the knowledge about the B cell immune response induced by the SARS-CoV-2 virus is still limited. Here, we report a comprehensive characterization of the dynamics of immunoglobin heavy chain (IGH) repertoire in COVID-19 patients. By using next-generation sequencing technology, we examined the temporal changes in the landscape of the patient’s immunological status and found dramatic changes in the IGH within the patient’s immune system after the onset of COVID-19 symptoms. Although different patients have distinct immune responses to SARS-CoV-2 infection, by employing clonotype overlap, lineage expansion, and clonotype network analyses, we observed a higher clonotype overlap and substantial lineage expansion of B cell clones 2–3 weeks after the onset of illness, which is of great importance to B-cell immune responses. Meanwhile, for preferences of V gene usage during SARS-CoV-2 infection, IGHV3-74 and IGHV4-34, and IGHV4-39 in COVID-19 patients were more abundant than those of healthy controls. Overall, we present an immunological resource for SARS-CoV-2 that could promote both therapeutic development as well as mechanistic research.
Abbreviations: SARS-CoV-2, Severe acute respiratory syndrome coronavirus 2; COVID-19, coronavirus disease 2019; PBMC, peripheral blood mononuclear cells; BCR, B-cell receptor; IGH, immunoglobin heavy chain; CDR3, complementarity determining region 3; SHM, somatic hypermutation
Keywords: COVID-19, SARS-CoV-2, B-cell receptor repertoire, Clonal expansion
1. Introduction
The current outbreak of SARS-CoV-2 infection is threatening global public health [1]. The scale of the humanitarian and economic impact of the COVID-19 is driving intense efforts to develop vaccines and neutralizing antibodies (NAb) against COVID-19. Therefore, understanding the principles of the B-cell responses during SARS-CoV-2 infection is of substantial importance for anti-viral vaccine and NAb development.
The B-cell receptors (BCRs) are immunoglobin molecules located on B-cell surfaces to recognize and bind foreign antigens. Upon encountering their specific antigen, B-cells become activated, proliferate, and may differentiate into produce short-lived effective antibody-secreting plasma cells or long-lived plasma cells and memory cells. At the same time, BCRs undergo a process of affinity maturation, which repeats cycles of somatic hypermutation of BCRs and subsequent clonal selection leads to increased binding affinity. Comparison of BCR sequences among individuals is of great interest because repertoires may have similar features if individuals are exposed to the same pathogen, giving rise to convergent antibodies.
Antibody specificity is largely determined by the IGH gene sequence used by each B-cell [2]. The recent developments in high-throughput sequencing have made it feasible to character IGH repertoire in large numbers of samples and it is increasingly being applied to gain insights into the humoral responses in healthy individuals and a wide range of diseases. This technic has also led to advances in our understanding of how the antibody repertoire changes in response to perturbation arising from initial viral infection, viral evolution, and vaccination. BCR repertoire analysis has been applied, for example, to influenza virus [3], [4], human immunodeficiency virus [5], varicella-zoster virus [6], and dengue virus [7]. However, the dynamics of antibody response elicited by SARS-CoV-2 infection remain to be determined.
To understand how B-cell immune repertoire changes over time during SARS-CoV-2 infection, we obtained IGH repertoires from the peripheral blood samples which were collected multiple times from five COVID-19 patients. We classified sequences into clones by lineage clustering analysis and tracked changes in the following characteristics: unique number of CDR3, Shannon index, the number of high-frequency clones, cumulative frequency of the top 100 clones, and V, J-gene segment usage at a different course of time. We also conducted clonotype overlap, lineage expansion, and CDR3 sequence network structure to explore the similarity and varying trends of IGH repertoire status at different time points. Overall, during the high degree of heterogeneity in B-cell clonal dynamics among patients, key common patterns were observed, i.e. the higher clonotype overlap and substantial lineage expansion of COVID-19 patients after 2–3 weeks of onset of illness.
2. Materials and methods
2.1. Study approval and samples
This study has been approved by the Research Ethics Committee of The Fifth Medical Center of PLA General Hospital, Beijing, China. (approval number: 2020034D), written informed consent was obtained from all patients. Five patients with newly diagnosed COVID-19 and three healthy donors were enrolled. According to the Diagnosis and Treatment Protocol for Novel Coronavirus Pneumonia (Trial Version 4, Released by the National Health Commission & State Administration of Traditional Chinese Medicine on January 27, 2020), as blood samples were tested positive for the SARS-Cov-2 virus, all patients were diagnosed as COVID-19 patients, with the 85-year-old patient developed severe symptoms, and the other four patients aged between 15 and 45 developed mild symptoms. Blood samples were collected at multiple points during hospitalization. Detailed clinical information is shown in Table 1 . The laboratory characteristics were listed in Supplementary Table 1.
Table 1.
Summary of the clinical information of the 5 convalescent patients.
| Category | P1 | P2 | P3 | P4 | P5 |
|---|---|---|---|---|---|
| Gender | F | M | M | F | F |
| Age | 41 | 15 | 45 | 85 | 44 |
| Severity | Mild | Mild | Mild | Severe | Mild |
| Symptom onset date | 01/18/2020 | 01/24/2020 | 01/24/2020 | 02/08/2020 | 01/21/2020 |
| Hospitalization | 01/27/2020 | 01/27/2020 | 01/27/2020 | 02/10/2020 | 01/29/2020 |
| Diagnosis date of COVID-19 (+) | 01/27/2020 | 01/27/2020 | 01/27/2020 | 02/09/2020 | 01/27/2020 |
| Blood sampling date | 01/31/2020 | 01/31/2020 | 01/31/2020 | 02/11/2020 | 01/31/2020 |
| 02/03/2020 | 02/03/2020 | 02/03/2020 | 02/14/2020 | 02/03/2020 | |
| 02/06/2020 | 02/06/2020 | 02/06/2020 | 02/17/2020 | 02/06/2020 | |
| 02/09/2020 | 02/09/2020 | 02/09/2020 | 02/20/2020 | 02/09/2020 | |
| 02/12/2020 | 02/12/2020 | 02/12/2020 | 02/23/2020 | 02/12/2020 | |
| 02/15/2020 | 02/15/2020 | 02/15/2020 | 02/15/2020 | ||
| Turn negative date | 02/01/2020 | 02/05/2020 | 02/06/2020 | 02/19/2020 | 02/08/2020 |
| Discharge date | 02/15/2020 | 02/15/2020 | 02/15/2020 | 02/25/2020 | 02/18/2020 |
2.2. Library preparation and repertoire sequencing
Human peripheral blood mononuclear cells (PBMCs) were isolated from blood samples of five COVID-19 patients as well as three healthy donors by density gradient centrifugation in Ficoll-Paque Plus buffer (GE Healthcare, USA). Total RNA was extracted from approximately 1 × 105 PBMCs of each sample using Trizol Reagent (Invitrogen, USA). cDNA was synthesized with SuperScript II reverse transcriptase (Invitrogen, USA) and Oligo-dN6 random primer. The variable domain of IGH was amplified by using six V segment-specific primers designed from the FR1 region and one J segment-specific primer designed from the FR4 region [8]. The PCR program began with an initial denaturation at 95 °C for 15 min, followed by 28 cycles of denaturation at 94 °C for 30 s, annealing of primer to DNA at 60 °C for 90 s, and extension by 2 × multiplex PCR master mix (QIAGEN, USA) at 72 °C for 30 s. Following the cycling, the reaction was at 72 °C for 5 min, with a final step at 12 °C to hold the terminated reaction. The amplified PCR products were then purified by the same volume of the Ampure-XP beads. Subsequently, the purified PCR products were end-repaired and ligated with a sequencing adapter. The sequencing libraries were quantified using Qubit ssDNA HS (High Sensitivity) Assay Kit (Invitrogen, USA) and sequenced by MGISEQ-2000 (MGI, China) platform with 400 bp single-end reads.
2.3. Basic data processing
The single-end 400 bp reads of the samples from the five COVID-19 patients sequenced at the MGISEQ-2000 platform were analyzed by IMonitor [9] software. Briefly, the raw reads were filtered and trimmed by SOAPnuke [10] to remove low-quality reads and adapter sequences. Reference germline gene segments were collected from the IMGT database [11]. BLAST [12] program was used to align all clean reads to a reference, and then a second alignment procedure was executed to improve the alignment accuracy. Subsequently, structure analysis was performed to deduce the structure of BCR molecular, including variable (V), diversity (D), and joining (J) segments usage, the nucleotide and amino acid sequence of the CDR3 region, and the deletions and insertions at rearrangement sites. Finally, several characters reflecting the status of the immune repertoire were counted, such as V-J pairing, V/J usage, CDR3 sequence frequency, and CDR3 length distribution.
2.4. Lineage identification and effective data normalization
We define sequences belonging to the same clonal lineage as those sharing the same V and J germline gene combination, having similar CDR3 length, and at least 90% nucleotide sequence identity in the CDR3 region. According to this definition, we first grouped sequence with identical V and J germline gene origination and same CDR3 length, following clustering those groups by CD-HIT (version 4.6) [13] with the following parameters: -c 0.9, -G 1, -b 20, -d 0 and -n 9. To eliminate the impact of the difference in data volume on clonal diversity, we normalized each of the IGH clones to a unanimous number. Firstly, we counted an effective data size (with an “in-frame” tag in the IMonitor result) for each sample and removed some samples with significantly fewer effective data than others. Secondly, normalized data with the same size were extracted randomly from each sample by an in-house Perl program.
2.5. Comparative analysis with neutralizing antibodies
A total of 15 CDR3 amino acid sequences of antibodies with the potential of neutralizing coronavirus were collected from recently published studies. A BLAST database was created based on those CDR3s, and blastp (Version 2.6.0+) was used to align all CDR3 amino acid sequences in current study to the BLAST database. The CDR3 amino acid sequences have an identity score greater than 50 with any sequence in the database were extracted as candidate sequences similar to potential neutralizing antibodies. Multiple sequence alignment was conducted for those similar sequences by using Clustal Omega (Version 1.2.4), and then maximum likelihood phylogenetic tree were generated by MEGA (Version 10.0.5-1) with 500 bootstrap.
2.6. Feature parameters analysis and comparison of usage frequency of V/J germline gene segments
There are several commonly used parameters reflecting the features of the BCR repertoire, including unique CDR3 number, Shannon index, high-frequency clone number, and cumulate frequency of the top 100 clones. we are using an in-house Perl script to count those parameters base on normalized data from each sample. The Shannon index is frequently used as a measure of repertoire diversity, therefore we calculated the Shannon index of observed clones from normalized data using the following function:
where N is the total number of unique CDR3, and p(i) is the frequency of a single CDR3 in the BCR repertoire.
After data normalization, gene usage frequency of V and J segments were counted and a k-means hierarchical clustering process was executed by R package “hclust” to compare V/J segment usage similarity both within and among individuals.
2.7. Overlap analysis of BCR repertoire
Repertoire overlap analysis is a similarity measure to evaluate the correlation of overlapping clonotypes between samples. The Jaccard index, also known as intersection over union and the Jaccard similarity coefficient, is commonly used to gauging the similarity and diversity of different repertoires. To explore the impact of onset time on the status of the BCR repertoire, we counted the Jaccard index to assess the association between the repertoire at different onset times. The index is calculated as follows:
where |A | denotes the total number of unique CDR3 in sample A, |A ∩ B| represents the intersection of the unique CDR3 number in samples A and B.
We calculate pairwise overlap rate as Jaccard index multiplied by 100, and divided the pairwise overlap rate into 3 clusters: the first cluster is “2–3 week adjacents” including data of time points 12–13 days versus 15–16 days and 15–16 days versus 18–19 days after onset; the second cluster is “other adjacents” which contains two adjacent time points except “2–3 week adjacents“; the third cluster is ”non-adjacent“ which contains all of the paired non-adjacent time points. A two-sample Wilcoxon test (Wilcoxon test in R) was used to compare the differences between clusters.
2.8. Lineage expansion analysis
A lineage expansion analysis was performed based on the normalized data. Firstly, a “join samples” process was executed to join lineage information at all time points together, inside each patient, then the normalized lineage frequency was computed as the geometric mean of lineage frequencies that comprise a given joint lineage in intersected time points. if the lineage is missing. Its frequency is set to 1e-9. Only the most abundant CDR3 sequence was selected as a representative clonotype in each lineage. Secondly, an R program originally generated by VDJtools [14] with some modifications was used to display the expansion and contraction of lineages.
2.9. CDR3 sequence network generating
The network is generated based on the BCR CDR3 amino acid similarity, each vertex in the network represents a CDR3 amino acid sequence, the size of the point represents the abundance of the clone, any two vertices are connected with an edge if they meet the following three criteria, including the same length, the same V and J gene usage, and only one amino substitution. The diagram was plotted by the “igraph” R package (https://igraph.org/redirect.html).
2.10. Parameters of the network calculation
The parameters of the network, including vertices, cluster number, and diameter of the largest cluster, are calculated using “V”, “clusters”, “diameter” functions in the “igraph” package. The Gini coefficient is used to measure the inequality in distribution, and we calculated the Gini coefficient of clone frequency distribution and cluster clone number distribution using the following function:
where y(i) is the frequency of a single clone; and n is the total unique clone number in the calculation of clone frequency Gini coefficient; y(i) is the clone number of a single cluster, and n is the total cluster number in the calculation of cluster Gini coefficient.
3. Result
3.1. Basic clinical characteristics of COVID-19 patients
Among the five patients diagnosed with COVID-19, the 85-year-old patient developed severe symptoms, and the other four patients aged between 15 and 45 developed mild symptoms (Table 1). Blood samples were collected on days 2–5 after diagnosis with 2–3 days intervals until they were tested negative for SARS-CoV-2 on days 17–19. All of them were discharged from the hospital upon recovery. Notably, 3 of the 5 patients are from the same family, that is, Patient 2 is the son of Patient 1 and Patient 3.
During the process of COVID-19 diagnosis, several clinical indicators were detected by routine blood examination. At least 3 of the 5 patients showed pronounced lymphopenia. The other 2 have too few tested indicators and can’t be reasonably counted. Along with the progression of COVID-19, the percentage of T lymphocytes went through a process of decreasing initially and then rebouncing. The phenomenon of lymphopenia in COVID-19 patients in our results is highly consistent with previous studies. It is well established that lymphopenia is a major immunological abnormality that occurs in COVID-19 patients [15], [16], [17], [18]. However, we found that the percentage of B lymphocytes showed an opposite trend to T cells: among these patients, the percentage of B cells in Patients 3, Patient 4, and Patient 5 reached the highest value until 14th, 6th and 17th day after the onset of the disease, respectively (Supplementary Table 1).
3.2. IGH repertoire altered substantially upon infection with SARS-CoV-2
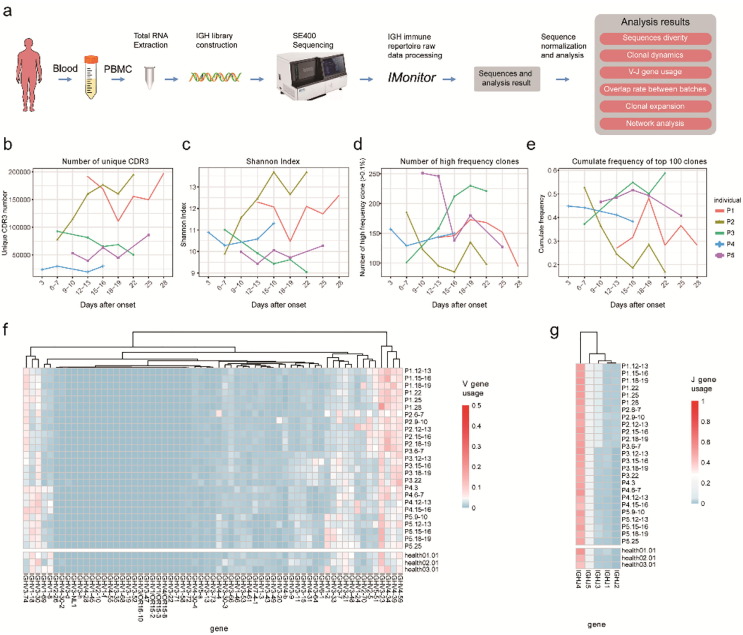
We performed longitudinal IGH repertoire sequencing to examine the global effects of SARS-CoV-2 infection on the B-cell compartment (Fig. 1 a). To enable direct comparisons among time points of each patient, datasets of all time points of each patient were sub-sampled to a unanimous number of clonotypes. We investigated the diversity of the IGH repertoires through four parameters, which are unique CDR3 number, Shannon index, high-frequency clone number (defined as frequency > 0.1%), and cumulate frequency of top 100 clones with the highest frequency. These parameters fluctuate substantially throughout time with no common tendencies among patients (Fig. 1 b, e), which may be due to the dramatic change of the predominant clones and lineages.
Fig. 1.
Schematic diagram of the experimental process and overall dynamic changes of the BCR repertoires.
Next, we thought to test whether there are preferences of V and J gene segments during SARS-CoV-2 infection. By using a hierarchical clustering strategy, we compared the V and J gene usage frequency of COVID-19 patients with that of healthy donors. We found that the most frequently observed V segments were IGHV3-23, IGHV3-30, IGHV4-34, IGHV4-39, and IGHV4-59 (Fig. 1 f, g), which is similar to previous studies [19], [20]. Additionally, the germline gene usage of IGHV3-74, IGHV4-34, and IGHV4-39 in COVID-19 patients were more abundant than those of in healthy controls. Preferred J gene segments showed a high degree of consistency among all time points in both patients and healthy donors (Fig. 1 g), with the most frequent J segment is IGHJ4, which is consistent with the previous report [21]. For each individual, the frequency of distinct gene segments usage over the infection time course was fairly stable, shown as high correlations in the heatmap of Fig. 1 f and g, suggesting that the overall V, J segment useage is not markedly altered by SARS-CoV-2. This phenomenon may reflect that the human immune system is highly variable between individuals but stays relatively stable over time in a given person [22].
a. Schematics of the experimental design and bioinformatics workflow. (b–e) Unique CDR3 numbers, Shannon index, high frequency (>0.1%) clone numbers, and the cumulative frequency of the top 100 clones of the BCR repertoire. Each dotted line represents the dynamic tendency of a single sample over time after onset. (f–g) The heat map shows the gene usage proportion inside each sample for the V/J genes listed at the bottom of the figure, each row corresponds to an individual, each column corresponds to a V/J germline gene, the color in each tile corresponds to the frequency of V/J gene in the repertoire, the deeper the color, the higher the frequency.
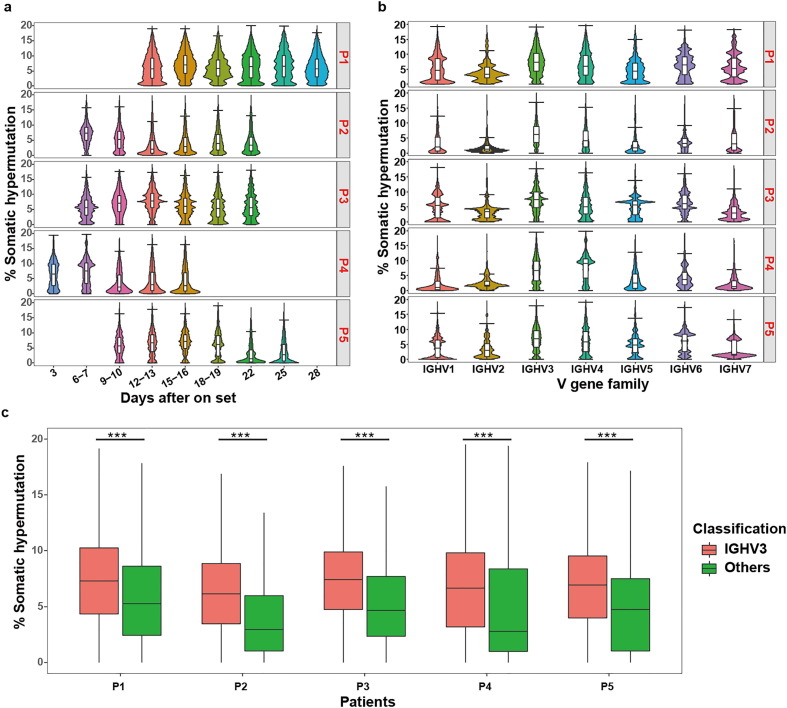
To study SARS-CoV-2-specific humoral responses as days post-symptom onset, we also explored our datasets for the somatic hypermutation (SHM) from the time series dimension. We calculated the somatic hypermutation rate as the proportion of mismatched bases of the heavy-chain variable regions by comparing DNA sequences of the heavy-chain variable regions (V, D, and J regions) to germline sequences. We found that among almost all 5 COVID-19 patients, the somatic hypermutation rate of IGH repertoire shows a slight upward in the early stages of onset and then a rapid decrease, while the time point of SHM rate decline is varied in different patients (Fig. 2 a). It is worth noting that the time point of SHM rate decline in Patient 4 was much earlier than that of other patients. Even at the second week after the symptom onset, we could begin to see the SHM of Patient 4 decreased significantly. In addition, further investigations on the SHM rate of different V gene families revealed skewing of higher SHM in IGHV3 gene family (Fig. 2 b). Notably, after extracting the clones of the IGHV3 gene family separately, we found that this part of the clones had significantly higher SHM rate than other gene families, in all patients (Fig. 2 c).
Fig. 2.
IGH repertoire SHM rate overview of COVID-19 patients. a. Violin diagram depicting the distribution of SHM rate at various time points after symptom onset. b. Violin diagram depicting the distribution of SHM rate in various germline gene families. c. Boxplot diagram comparing the difference of SHM rate between IGHV3 and other gene families. The comparison was performed using paired Wilcoxon test. *** P < 0.001.
3.3. Comparation with the neutralizing antibody sequence reveals a subset of CDR3s similar to an anti-SARS-CoV-2 single-domain antibody
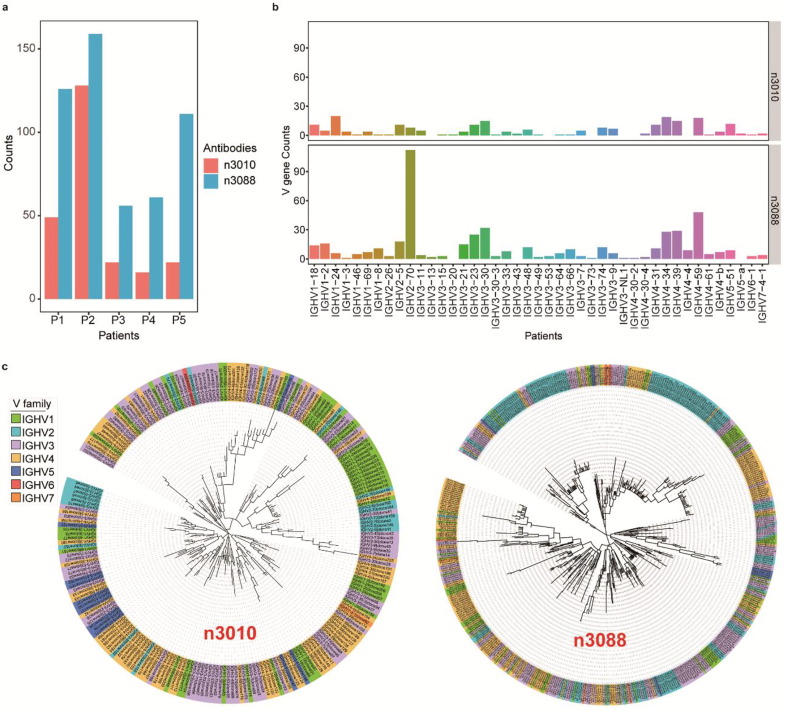
Neutralizing antibodies are responsible for defending cells from pathogens. Upon triggering by infections, the body can produce neutralizing antibodies as part of its immune response. To examine the potential of our IGH repertoire to provide basic support for the development of neutralizing antibodies, we collected 15 CDR3 amino acid sequences of antibodies with neutralizing potential from two recently published studies, and conducted comparison analysis with our datasets. Comparing with our datasets, , we found that Dr. Wu’s [23] study had fewer but higher similar CDR3s than Dr. Cao’s [24] study (Supplementary Fig 1). Specifically, examing the sequences of antibody BD-494 and BD-501 from Dr. Cao’s study, we found 14 and 1 clone, respecitively, having identical CDR3 amino acid sequence (100% identity score reported by blastp) in our dataset, (Supplementary Table 2). However, there is no clone in our data has exactly the same CDR3 amino acid sequence as the antibody reported in the Wu’s study, the highest sequence similarity is 93.75%. Notably, we detected a total of 226 and 497 unique CDR3s amino acid sequences similar to the anti-SARS-CoV-2 single-domain antibody n3010 and n3088 isolated by Wu et al, respectively. Among them, Patient 2 showed the highest enrichment of CDR3s similar to those antibodies (Fig. 3 a). Further analysis to explore the usage patterns of germline V genes shows that clones with CDR3 amino acid sequences similar to potential neutralizing antibodies has a clear preferential usage of Ig heavy chain variable region 3 (IGHV3) and 4 (IGHV4) subfamily genes. It is worth noting that there is a branch also had a bias toward IGHV1-24 in clones similar to n3010, and a branch that is biased towards IGHV2-70 in clones similar to n3088. (Fig. 3 b). Next, we performed phylogenetic sequence analysis on two subsets of clones similar to antibody n3010 and n3088 respectively. Phylogenetic analysis revealed that the CDR3s belonging to the IGHV1 and IGHV2 gene families clustered into the unique clusters respectively, while the CDR3s of the IGHV3 and IGHV4 gene families were fused together (Fig. 3 c).
Fig. 3.
Features of CDR3s similar to anti-SARS-CoV-2 single-domain antibody. a. Bar graph shows the number of CDR3s similar to anti-SARS-CoV-2 antibody n3010 and n3088 in the IGH repertoire of 5 COVID-19 patients. b. Bar graph depicting the abundance of IGHV gene usage of two subsets of clones similar to antibody n3010 and n3088 extracted from comparative analysis with neutralizing antibodies. c. Phylogenetic tree analysis showing the 226 and 497 unique CDR3s that were similar to antibodies n3010 and n3088.
3.4. Repertoire overlap analysis revealed higher similarity of immunological status between the adjacent time points in 2–3 weeks since symptom onset
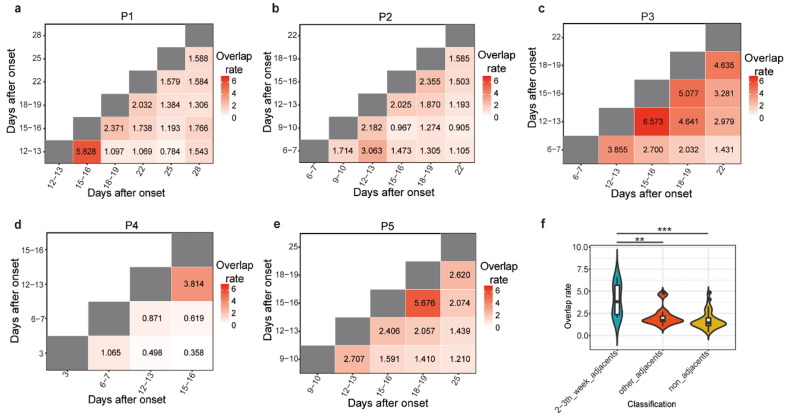
To better understand the potential mechanism behind changes in IGH repertoire diversity, we examined how the composition of the clonotype changed during the infection. We evaluated the level of clonotype overlap between any two-time points of each patient by using an all-versus-all pairwise overlap analysis. In general, the overlap rate between different time points ranges from 0.36% to 6.57% (average 2.13%) in each individual (Fig. 4 a-e). As expected, samples from adjacent sampling batches share more clones than those from other batches. A notable rule that shows in almost every patient is that the overlap rate of adjacent batches peaks at around 2–3 weeks. We found that “2-3 week adjacents” cluster shared significantly more clones than “other adjacents” (p = 7.78 × 10−3) and “non-adjacent” (p = 4.52 × 10−5). (Fig. 4 f). This indicates that, compared with other time points, the composition of the IGH immune repertoire within individuals is more similar during 2–3 weeks after the onset. The 2–3 weeks after onset is likely a critical stage of B-cell immune response.
Fig. 4.
Overview of BCR repertoire overlap changes among various time points after onset. (a–e) Each sub-graph is a heatmap describe an all-vs-all intersection between two-time points in a single patient, the value in each tile represents the clonotype overlap rate between two-time points as shown on the X-axis and Y-axis, the color in each tile corresponds to the clonotype overlap rate between two-time points, the deeper the color, the higher the overlap rate. f. The comparison of clonotype overlap rate among various time point categories. The comparison was performed using paired Wilcoxon test. ** P < 0.01, *** P < 0.001.
3.5. Tracking analysis revealed evidence of lineage expansion of IGH repertoire during SARS-CoV-2 infection
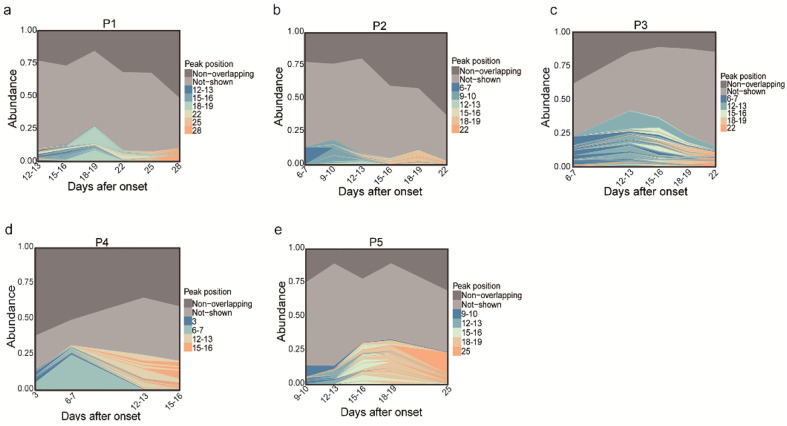
Next, we investigated the relative degree of clonal expansion. To measure the alterations in lineage frequency over time, we first defined clonotypes as a lineage if they meet 3 criteria: sharing the same VH and JH germline gene segments, having the same nucleotide length in the CDR3 region, and at least 90% nucleotide identity in the CDR3 region. Using this strategy, we computed the geometric mean of linage frequencies of all samples and performed a lineage tracking analysis using VDJtools. Here, we defined the highly expanded lineages as the 100 most abundant lineages (top 100 lineages). We observed an evident lineage expansion process over time. As shown in Fig. 5 a–e, the cumulative frequencies of the top 100 lineages of almost all patients increased after the second or third weeks following the onset of illness, with the cumulative frequencies of the top 100 lineages peaked at day 18–19 in Patient 1 and Patient 5, day 12–13 in Patient 3, day 6–7 in Patient 4 and both peaked at day 9–10 and day 18–19 in Patient 2. After that, this part of lineages began to decrease gradually. This result indicates that during the 2–3 weeks of disease progression, the patient’s IGH repertoire experienced a substantial lineage expansion.
Fig. 5.
BCR lineage expands and contracts over time after COVID-19 onset. Each sub-graph shows the detailed profiles for lineage expansion/constriction in a single patient. The top 100 lineages are colored by the peak position of their abundance profile, other lineages in each BCR repertoire are divided into two groups: non-overlapping (“Non-overlapping”) lineages are marked in dark gray, and the remaining lineages (“Not-shown”) are marked in light gray.
3.6. Network features delineate the evolution of the BCR repertoire landscape
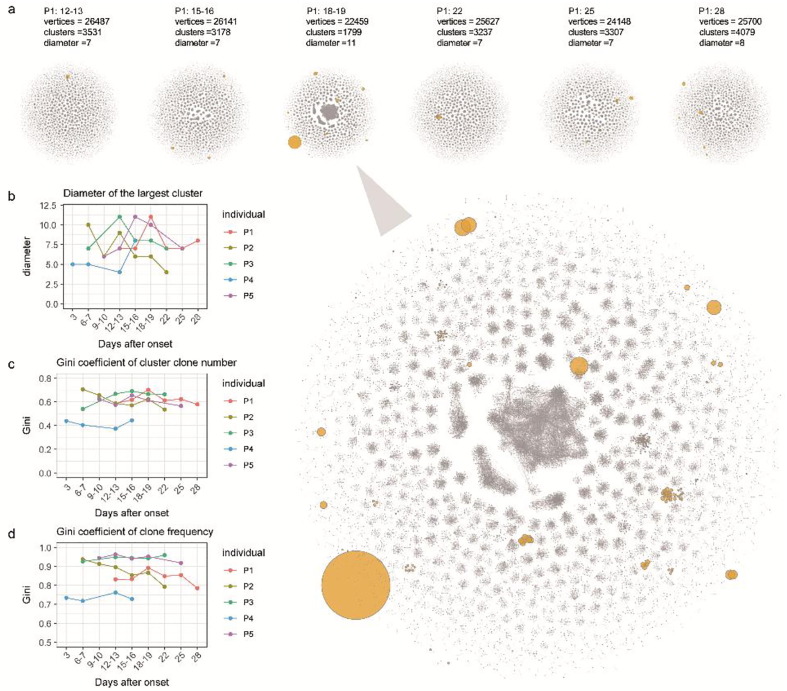
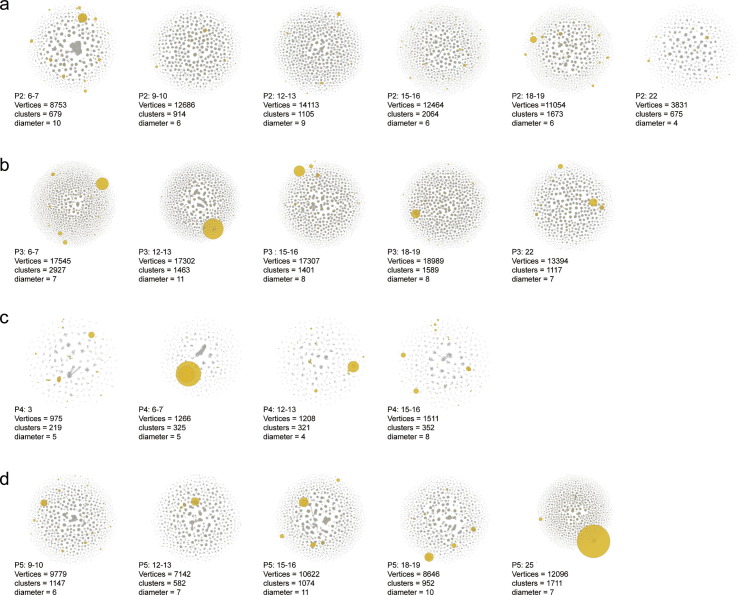
BCRs with similar CDR3 amino acid sequences are supposed to be gathered in clusters based on antigen-driven SHM. To investigate the degree of clone convergence during the disease, we build the BCR repertoire CDR3 sequence network for each individual, using data of different times after the onset of the disease based on a specific approach [25]. Networks of all the 5 patients after symptom onset have been plotted in Supplementary Fig. 2 and the network of the 41-year-old female Patient 1 at different time points is demonstrated in Fig. 6 a since its relatively high number of CDR3 clones (vertices). Here, a clear clonal clustering phenomenon could be seen from the network of this patient on days 18–19 (Fig. 6 a). Subsequently, three network-related parameters, namely diameter of the largest cluster, Gini coefficient of cluster clone number, and Gini coefficient of clone frequency, were counted and summarized in Fig. 6 b–d. There was a similar trend that first goes up and peaks around 2–3 weeks and falls afterward in these three parameters of almost all patients. Each parameter of the network has its implication, firstly, the diameter parameter exhibit in Fig. 6 b is the length of the longest geodesic (edges between any two reachable vertices) of a network. The result indicates a higher evolution level of BCR clones in the repertoire from 2 to 3 weeks. Secondly, the Gini coefficient of cluster clone number which represents the level of clonal clustering also peaked around 2–3 weeks. Thirdly, the Gini coefficient of clone frequency which indicates the skewed expansion of several clones was very much synchronized with the former parameter, despite they were relatively independent. Moreover, the most abundant clones usually don’t appear in the largest cluster (Fig. 6 d, Supplementary Fig. 2), suggesting the formation of big clusters may be due to the rapid iteration of naïve B-cells to generate affinity maturation, while the clonal expansion may result from the proliferate of the effector cells. Nevertheless, both the cluster clone number increment and clonal expansion are supposed to be triggered by the reaction to counteract the SARS-CoV-2 infection.
Fig. 6.
BCR repertoire network evolves after onset. (a) The network diagrams show how CDR3s are clustered with similar clones, each vertex represents a CDR3 amino acid sequence, the size of the point represents the abundance of the clone, any two vertices meet three conditions, including same length, same V, and J gene usage, and only one amino substitution are connected with an edge. Six batches of network plot from Patient 1 were lined up from left to right, the 18-19th day after onset were zoomed in. (b-d) cluster number, the diameter of the biggest cluster, and the average degree of the network, each dotted line represents the dynamic tendency of an individual over time after the onset of the disease.
4. Discussion
To obtain a more detailed understanding of the dynamics and sophistication of B-cell immune response in COVID-19 patients, we applied a comprehensive analysis of IGH repertoire during the disease process. Given the advantages of longitudinal data in profiling the fast repertoire dynamics through time in individuals responding to an immune challenge [19], [26], we collected a set of longitudinal immune repertoire data from 5 COVID-19 patients and conducted a detailed analysis of their B-cell immune repertoires. The simultaneous profiling of clonotype overlap, lineage expansion, and clonotype network enabled us to find that a period of 2–3 weeks after symptom onset is of great importance to B-cell immune response. In short, these results reveal a clear dynamic landscape of BCR repertoire in COVID-19 patients, as well as the characteristics of the immune repertoire at different time points after symptom onset.
V, J gene usage analysis revealed that each individual’s frequency of different V and J gene segment usage stays relatively stable over time (Fig. 1 f). Similar results have also been reported by Chen Wang et. al [6] suggesting that V, J segment use is not markedly altered by SARS-CoV-2 infection. However, the overall use of V components among patients shows a differentiated preference. As Petter Brodin mentioned [22], this may reflect that the human immune system is highly variable between individuals but relatively stable over time within a given person. We observed a preference of the use of V gene segments inconsistent with the single-cell BCR repertoire research by Wen et al [27], but consistent with Jacob's [28] bulk BCR repertoire study. For example, Wen's group reported that IGHV3-23 was over-represented in patients compared with healthy controls, while both our and Jacob's result showed that IGHV3-23 is highly represented both in patients and healthy controls. Additionally, we and Jacob's group both found that IGHV4-34 is more frequently used in COVID-19 patients compared with healthy donors. We believe that, given the advantages of large amounts of sequencing data, bulk BCR sequencing may be more suitable for BCR gene usage frequency assessment. Nevertheless, with only five patients, the power of our study is almost certainly constrained by the small sample size. Larger samples and more sequencing data need to be collected to help further understand these differences.
We found from the longitudinal immune repertoire data that all patients' BCR repertoire composition in 2–3 weeks of onset is different from that in other periods. There are significantly more common clones between adjacent time points and evident lineage expansion during 2–3 weeks. Additionally, network analysis revealed that each patient’s average degree and diameter indicators peaked around 2–3 weeks after onset. This timeline of repertoire change is consistent with an antigen (virus) induced B-cell differentiation and maturation. The maturation process is on par with other viruses, such as influenza viruses [29], and a similar phenomenon and timeline has also been observed in patients with the Middle East respiratory syndrome (MERS) [30]. We speculated that the higher clonotype overlap and substantial lineage expansion of COVID-19 patients after 2–3 weeks of illness may be associated with strong neutralizing antibody responses, accompanied by a large expansion of virus-specific B-cell clones and finally causing B-cell clone convergence.
Many studies have shown that the monoclonal antibodies targeting viral surface antigens have therapeutic and preventive effects on infectious diseases such as HIV, Ebola, and MERS [31], [32], [33]. Their safety and effectiveness in patients have been confirmed in many clinical trials [34], [35]. With the efforts of scientists from various countries, recent positive progress has been made in the development of anti-SARS-CoV-2 drugs [36], [35], [37], [38], [39], [23]. We compared the IGH repertoire of COVID-19 patients in this study with the 15 CDR3s of antibodies with neutralizing potential collected in two recently published studies. The results revealed that there is a clear preference for sequence homology between clones in this study and antibodies with neutralizing potential. Only the “n3010” antibody from the study of Wu et al has similar sequences to almost all the IGH repertoire in our study, which may reflects the huge diversity of the antibody immune repertoire and the huge differences in the neutralizing activity of antibodies produced by different patients.
Patient 4 deserves our attention. According to the results of clone expansion analysis, we can see that the peak time of clone expansion of this patient is much earlier than others. Notably, Patient 4 is the only severe case in this study, and the age is as high as 85 years old. We hypothesized that the earlier peak of clone expansion time point may be assoicated with the severity of clinical symptoms and the age of patients.
In general, based on high-throughput immune repertoire sequencing data, this study provides a comprehensive B cell deep immune profiling of COVID-19 patients. Conclusions from this research will help understand the immune response of COVID-19 patients against the SARS-CoV-2 virus and will provide the necessary basic knowledge for the development of vaccines and neutralizing antibodies.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Acknowledgments
We sincerely thank the support provided by China National GeneBank. This study was supported by CAMS Research Units of Adaptive Evolution and Control of Emerging Viruses (2018RU009). William J. Liu is supported by the Excellent Young Scientist Program of the National Natural Science Foundation of China (NSFC, 81822040) and Beijing New-star Plan of Science and Technology (Z181100006218080). The study was supported by Guangdong Provincial Key Laboratory of Genome Read and Write (No. 2017B030301011), Shenzhen Science and Technology Innovation Fund (JCYJ20170412152916724 and JCYJ20170817145305022), Zhongnanshan Medical Foundation of Guangdong Province (ZNSA-2021004) and the COVID-19 Prevention and Treatment Fund from Shantou Science and Technology Agency. We thank Dr. Sunil Kumar Sahu at BGI-Shenzhen for his linguistic assistance in this manuscript preparation.
Data availability
The data that support the findings of this study have been deposited into the CNGB Sequence Archive (CNSA: https://db.cngb.org/cnsa/) of CNGBdb with accession number CNP0001106.
Author contributions
HX and LW analyzed the sequencing data and prepared the figures. YZ, PY, CrB, ZW, LY, JY, FM, CqB, NZ, YG, YB, SZ, and CZ collected the specimens, and retrieved and analyzed the clinical data. XL, MW, YZ, ZX, BY, PW, and XL performed the experiments. HX, LW, MW, and XL wrote the draft with the help of all authors. LT, NY, LL, YS, GW, WJL, YZ, PL, and YG provided a critical review of the manuscript. WJL, NY, PY, and GFG conceived the idea and supervised the entire study. All other authors contributed to the work. All authors read and approved the manuscript for submission.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.humimm.2021.10.007.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
Supplementary figure 1.
References
- 1.Zhu N., et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382:727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Xu J.L., Davis M.M. Diversity in the CDR3 region of V(H) is sufficient for most antibody specificities. Immunity. 2000;13(1):37–45. doi: 10.1016/s1074-7613(00)00006-6. [DOI] [PubMed] [Google Scholar]
- 3.Jiang N., He J., Weinstein J.A., Penland L., Sasaki S., He X.-S., Dekker C.L., Zheng N.-Y., Huang M., Sullivan M., Wilson P.C., Greenberg H.B., Davis M.M., Fisher D.S., Quake S.R. Lineage structure of the human antibody repertoire in response to influenza vaccination. Sci. Transl. Med. 2013;5(171) doi: 10.1126/scitranslmed.3004794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Vollmers C., Sit R.V., Weinstein J.A., Dekker C.L., Quake S.R. Genetic measurement of memory B-cell recall using antibody repertoire sequencing. Proc. Natl. Acad. Sci. U. S. A. 2013;110:13463–13468. doi: 10.1073/pnas.1312146110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.K.B. Hoehn, et al., Dynamics of immunoglobulin sequence diversity in HIV-1 infected individuals. (2015). [DOI] [PMC free article] [PubMed]
- 6.Wang C., et al. B-cell repertoire responses to varicella-zoster vaccination in human identical twins. Proc. Natl. Acad. Sci. U. S. A. 2015;112:500–505. doi: 10.1073/pnas.1415875112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Parameswaran P., Liu Y.i., Roskin K., Jackson K.L., Dixit V., Lee J.-Y., Artiles K.L., Zompi S., Vargas M., Simen B., Hanczaruk B., McGowan K., Tariq M., Pourmand N., Koller D., Balmaseda A., Boyd S., Harris E., Fire A. Convergent antibody signatures in human dengue. Cell Host Microbe. 2013;13(6):691–700. doi: 10.1016/j.chom.2013.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.van Dongen J.J.M., Langerak A.W., Brüggemann M., Evans P.A.S., Hummel M., Lavender F.L., Delabesse E., Davi F., Schuuring E., García-Sanz R., van Krieken J.H.J.M., Droese J., González D., Bastard C., White H.E., Spaargaren M., González M., Parreira A., Smith J.L., Morgan G.J., Kneba M., Macintyre E.A. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: Report of the BIOMED-2 concerted action BMH4-CT98-3936. Leukemia. 2003;17(12):2257–2317. doi: 10.1038/sj.leu.2403202. [DOI] [PubMed] [Google Scholar]
- 9.W. Zhang, et al., Imonitor: A robust pipeline for TCR and BCR repertoire analysis, Genetics 201 (2015) 459–472. [DOI] [PMC free article] [PubMed]
- 10.Y. Chen, et al., SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data, Gigascience 7 (2018) 1–6. [DOI] [PMC free article] [PubMed]
- 11.Lefranc M.P., et al. IMGT®, the international ImMunoGeneTics information system®. Nucleic Acids Res. 2009;37:1006–1012. doi: 10.1093/nar/gkn838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 13.Li W., Godzik A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics. 2006;22(13):1658–1659. doi: 10.1093/bioinformatics/btl158. [DOI] [PubMed] [Google Scholar]
- 14.Shugay M., et al. VDJtools: Unifying Post-analysis of T Cell Receptor Repertoires. PLoS Comput. Biol. 2015;11:1–16. doi: 10.1371/journal.pcbi.1004503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Guan W.-J., Ni Z.-y., Hu Y.u., Liang W.-H., Ou C.-Q., He J.-X., Liu L., Shan H., Lei C.-L., Hui D.S.C., Du B., Li L.-J., Zeng G., Yuen K.-Y., Chen R.-C., Tang C.-l., Wang T., Chen P.-Y., Xiang J., Li S.-Y., Wang J.-L., Liang Z.-J., Peng Y.-X., Wei L.i., Liu Y., Hu Y.-H., Peng P., Wang J.-M., Liu J.-Y., Chen Z., Li G., Zheng Z.-J., Qiu S.-Q., Luo J., Ye C.-J., Zhu S.-Y., Zhong N.-S. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020;382(18):1708–1720. doi: 10.1056/NEJMoa2002032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Huang C., Wang Y., Li X., Ren L., Zhao J., Hu Y.i., Zhang L.i., Fan G., Xu J., Gu X., Cheng Z., Yu T., Xia J., Wei Y., Wu W., Xie X., Yin W., Li H., Liu M., Xiao Y., Gao H., Guo L.i., Xie J., Wang G., Jiang R., Gao Z., Jin Q.i., Wang J., Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan. China. Lancet. 2020;395(10223):497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Schultheiß C., Paschold L., Simnica D., Mohme M., Willscher E., von Wenserski L., Scholz R., Wieters I., Dahlke C., Tolosa E., Sedding D.G., Ciesek S., Addo M., Binder M. Next-Generation Sequencing of T and B Cell Receptor Repertoires from COVID-19 Patients Showed Signatures Associated with Severity of Disease. Immunity. 2020;53(2):442–455.e4. doi: 10.1016/j.immuni.2020.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jafarzadeh A., Jafarzadeh S., Nozari P., Mokhtari P., Nemati M. Lymphopenia an important immunological abnormality in patients with COVID-19: Possible mechanisms. Scand. J. Immunol. 2021;93(2) doi: 10.1111/sji.v93.210.1111/sji.12967. [DOI] [PubMed] [Google Scholar]
- 19.Laserson U., et al. High-resolution antibody dynamics of vaccine-induced immune responses. Proc. Natl. Acad. Sci. 2014;111:4928–4933. doi: 10.1073/pnas.1323862111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Y.C. Wu, et al., High-throughput immunoglobulin repertoire analysis distinguishes between human IgM memory and switched memory B-cell populations, Blood 116 (2010) 1070–1078. [DOI] [PMC free article] [PubMed]
- 21.B. Shi, et al., The usage of human IGHJ genes follows a particular nonrandom selection: The recombination signal sequence affects the usage of human IGHJ genes, bioRxiv 792085 (2019). doi:10.1101/792085. [DOI] [PMC free article] [PubMed]
- 22.Brodin P., Davis M.M. Human immune system variation. Nature Reviews Immunology. 2017;17(1):21–29. doi: 10.1038/nri.2016.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wu Y., Li C., Xia S., Tian X., Kong Y.u., Wang Z., Gu C., Zhang R., Tu C., Xie Y., Yang Z., Lu L.u., Jiang S., Ying T. Identification of Human Single-Domain Antibodies against SARS-CoV-2. Cell Host Microbe. 2020;27(6):891–898.e5. doi: 10.1016/j.chom.2020.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cao Y., Su B., Guo X., Sun W., Deng Y., Bao L., Zhu Q., Zhang X.u., Zheng Y., Geng C., Chai X., He R., Li X., Lv Q.i., Zhu H., Deng W., Xu Y., Wang Y., Qiao L., Tan Y., Song L., Wang G., Du X., Gao N., Liu J., Xiao J., Su X.-D., Du Z., Feng Y., Qin C., Qin C., Jin R., Xie X.S. Potent Neutralizing Antibodies against SARS-CoV-2 Identified by High-Throughput Single-Cell Sequencing of Convalescent Patients’ B Cells. Cell. 2020;182(1):73–84.e16. doi: 10.1016/j.cell.2020.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bashford-Rogers R.J.M., et al. Network properties derived from deep sequencing of human b-cell receptor repertoires delineate b-cell populations. Genome Res. 2013;23:1874–1884. doi: 10.1101/gr.154815.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.de Bourcy C.F.A., Dekker C.L., Davis M.M., Nicolls M.R., Quake S.R. Dynamics of the human antibody repertoire after B cell depletion in systemic sclerosis. Sci. Immunol. 2017;2(15) doi: 10.1126/sciimmunol.aan8289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wen W., et al. Immune cell profiling of COVID-19 patients in the recovery stage by single-cell sequencing. Cell Discov. 2020;6:31. doi: 10.1038/s41421-020-0168-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.J.D. Galson, et al., Deep sequencing of B cell receptor repertoires from COVID-19 patients reveals strong convergent immune signatures, bioRxiv (2020). doi:10.1101/2020.05.20.106294. [DOI] [PMC free article] [PubMed]
- 29.Lam J.H., Baumgarth N. The Multifaceted B Cell Response to Influenza Virus. J. Immunol. 2019;202:351–359. doi: 10.4049/jimmunol.1801208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Al-Abdely H.M., et al. Middle east respiratory syndrome coronavirus infection dynamics and antibody responses among clinically diverse patients. Saudi Arabia. Emerg. Infect. Dis. 2019;25:753–766. doi: 10.3201/eid2504.181595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Scheid J.F., Mouquet H., Feldhahn N., Seaman M.S., Velinzon K., Pietzsch J., Ott R.G., Anthony R.M., Zebroski H., Hurley A., Phogat A., Chakrabarti B., Li Y., Connors M., Pereyra F., Walker B.D., Wardemann H., Ho D., Wyatt R.T., Mascola J.R., Ravetch J.V., Nussenzweig M.C. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature. 2009;458(7238):636–640. doi: 10.1038/nature07930. [DOI] [PubMed] [Google Scholar]
- 32.Corti D., et al. Protective monotherapy against lethal Ebola virus infection by a potently neutralizing antibody. Science (80-.) 2016;351:1339–1342. doi: 10.1126/science.aad5224. [DOI] [PubMed] [Google Scholar]
- 33.Wang L., Shi W., Chappell J.D., Joyce M.G., Zhang Y.i., Kanekiyo M., Becker M.M., van Doremalen N., Fischer R., Wang N., Corbett K.S., Choe M., Mason R.D., Van Galen J.G., Zhou T., Saunders K.O., Tatti K.M., Haynes L.M., Kwong P.D., Modjarrad K., Kong W.-P., McLellan J.S., Denison M.R., Munster V.J., Mascola J.R., Graham B.S., Gallagher T. Importance of Neutralizing Monoclonal Antibodies Targeting Multiple Antigenic Sites on the Middle East Respiratory Syndrome Coronavirus Spike Glycoprotein To Avoid Neutralization Escape. J. Virol. 2018;92(10) doi: 10.1128/JVI.02002-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Xu J., et al. Antibodies and vaccines against Middle East respiratory syndrome coronavirus. Emerg. Microbes Infect. 2019;8:841–856. doi: 10.1080/22221751.2019.1624482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Caskey M., et al. Antibody 10–1074 suppresses viremia in HIV-1-infected individuals. Nat. Med. 2017;23:185–191. doi: 10.1038/nm.4268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Long Q.-X., Liu B.-Z., Deng H.-J., Wu G.-C., Deng K., Chen Y.-K., Liao P.u., Qiu J.-F., Lin Y., Cai X.-F., Wang D.-Q., Hu Y., Ren J.-H., Tang N.i., Xu Y.-Y., Yu L.-H., Mo Z., Gong F., Zhang X.-L., Tian W.-G., Hu L.i., Zhang X.-X., Xiang J.-L., Du H.-X., Liu H.-W., Lang C.-H., Luo X.-H., Wu S.-B., Cui X.-P., Zhou Z., Zhu M.-M., Wang J., Xue C.-J., Li X.-F., Wang L.i., Li Z.-J., Wang K., Niu C.-C., Yang Q.-J., Tang X.-J., Zhang Y., Liu X.-M., Li J.-J., Zhang D.-C., Zhang F., Liu P., Yuan J., Li Q., Hu J.-L., Chen J., Huang A.-L. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 2020;26(6):845–848. doi: 10.1038/s41591-020-0897-1. [DOI] [PubMed] [Google Scholar]
- 37.Pinto D., Park Y.-J., Beltramello M., Walls A.C., Tortorici M.A., Bianchi S., Jaconi S., Culap K., Zatta F., De Marco A., Peter A., Guarino B., Spreafico R., Cameroni E., Case J.B., Chen R.E., Havenar-Daughton C., Snell G., Telenti A., Virgin H.W., Lanzavecchia A., Diamond M.S., Fink K., Veesler D., Corti D. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature. 2020;583(7815):290–295. doi: 10.1038/s41586-020-2349-y. [DOI] [PubMed] [Google Scholar]
- 38.Wu Y., Wang F., Shen C., Peng W., Li D., Zhao C., Li Z., Li S., Bi Y., Yang Y., Gong Y., Xiao H., Fan Z., Tan S., Wu G., Tan W., Lu X., Fan C., Wang Q., Liu Y., Zhang C., Qi J., Gao G.F., Gao F., Liu L. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science. 2020;368(6496):1274–1278. doi: 10.1126/science:abc2241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.B. Ju, et al., Potent human neutralizing antibodies elicited by SARS-CoV-2 infection, bioRxiv (2020). doi:10.1101/2020.03.21.990770. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data that support the findings of this study have been deposited into the CNGB Sequence Archive (CNSA: https://db.cngb.org/cnsa/) of CNGBdb with accession number CNP0001106.