Abstract
A simple, disposable, and integrated electronic-tube cap (E-tube cap) for DNA detection at the point-of-care was designed, fabricated, and tested. The E-tube cap contains a 3D printed electrode substrate for DNA extraction and label-free pH sensing detection. One Flinders Technology Associates (Whatman FTA) membrane was incorporated into the 3D printed electrode substrate for the isolation, concentration, and purification of DNA. The E-tube cap with captured DNA by the membrane was inserted directly into a reaction tube for loop-mediated isothermal amplification (LAMP). The isothermal amplification process was monitored in real-time by a 3D printed electrochemical electrode coated with pH-sensitive material (carbon/iridium oxide layer). The pH sensing electrode showed an excellent linear response within the pH range of 6-9 with a slope of −31.32 ± 0.5 mV/pH at room temperature. The utility of the integrated E-tube cap was demonstrated by detecting the presence of lambda DNA spiked in saliva samples with a sensitivity of 100 copies per mL sample within 30 minutes. Such a simple, rapid, and affordable diagnostic device is particularly suitable for point-of-care molecular diagnostics of infectious diseases.
Keywords: 3D-printing, point-of-care molecular diagnostics, pH-sensitive electrode, electrochemical detection, loop-mediated, isothermal amplification (LAMP)
Graphical abstract

1. Introduction
To combat infectious diseases, it is of utmost importance to develop a simple, affordable, and sensitive diagnostic platform for the rapid detection of pathogens. Nucleic acid tests provide a powerful tool to perform such disease diagnostics due to their high sensitivity and specificity (Kadimisetty et al. 2018; Lv et al. 2020; Toumazou et al. 2013; Wang et al. 2013; Xie et al. 2014; Zeng et al. 2019, 2021; Zhao et al. 2010). Among them, real-time fluorescence polymerase chain reaction (PCR) method is one of the most widely used technologies for pathogen nucleic acid detection (Estrada-Peña et al., 2017). However, it typically requires delicate optical components and complex thermocycler, which restricts its use in centralized clinical laboratories. As a promising alternative to the PCR method, isothermal amplification technology, such as loop-mediated isothermal amplification (LAMP), recombinase polymerase amplification (RPA), has emerged as an attractive point-of-care molecular diagnostic method because of its simplicity, rapidity, and cost-effectiveness (Kadimisetty et al., 2018; Liu et al. 2020; Stobiecka and Chałupa, 2015; Wang et al., 2013; Xie et al., 2014; Zhao et al., 2010).
Fluorescence-based detection is the most commonly used technology in nucleic acid amplification tests, but it relies on relatively expensive fluorescence dye and complex fluorescence optical components. As an alternative to fluorescent-based optical sensing detection, several attempts have been made to detect nucleic acids through an electrochemical method, enabling miniaturization and simplification of the detection system. In particular, the combination of isothermal amplification or isothermal signal amplification with electrochemical detection provides a simple, affordable, and portable method for nucleic acid-based molecular diagnostics (Fu et al., 2013; Liu et al., 2015). Several groups have exploited the electrochemical detection of DNA amplification by using DNA-intercalative electroactive molecules (e.g. methylene blue) (Hsieh et al., 2012; Luo et al., 2014). However, it suffers from low signal resolution due to the weak binding affinity of electroactive molecules to amplicon products. Alternatively, pH potentiometric measurement is simpler and label-free by taking advantage of the generation of hydrogen ions (H+) during the nucleic acid amplification process. For example, ion-sensitive field-effect transistors (ISLET) and portable pH meters have, respectively, been used for nucleic acid amplification detection (Toumazou et al., 2013; Xie et al., 2014). However, they are not ideal for point-of-care testing due to the need for either expensive semiconductor fabrication technology or bulky equipment. In addition, all of these methods are lacking in nucleic acid sample preparation ability.
In recent years, there has been a major effort to integrate DNA sample preparation processes and amplification detection into an inexpensive portable device for point-of-care tests. Such a simple, low-cost, and integrated diagnostic system is particularly suitable for point-of-care diagnostics at home, in small clinics, and in doctor's offices. For example, Park et al. reported an integrated rotary microfluidic system for DNA extraction, LAMP assay, and lateral flow strip-based detection (Park et al., 2017). Liu et al. described an isothermal amplification microfluidic reactor that combined on-chip nucleic acid extraction and real-time fluorescence LAMP assay for HIV detection with a fluorescence detector (Liu et al. 2011). However, the systems described before based on LAMP still rely on either a separate lateral flow strip or a complex fluorescence detector.
Herein, an integrated electronic-tube cap (E-tube cap) system has been designed, fabricated, and tested for DNA detection in saliva samples. The E-tube cap system integrates multiple functions, including DNA extraction from raw saliva samples, LAMP reaction, and label-free electrochemical detection. One Flinders Technology Associates (Whatman FTA) membrane was incorporated into the 3D-printed electrode substrate of the E-tube cap for DNA capture, concentration, and purification. The 3D printed electrode substrate was coated with carbon paste and iridium oxide for pH sensing of LAMP amplification. The performance of the integrated E-tube cap system was validated by detecting lambda DNA in spiked saliva samples.
2. Methods
2.1. Material and Reagents
Potassium chloride, potassium hydroxide, ammonium sulfate, Tween 20, Whatman FTA membrane, and FTA purification reagent were purchased from Sigma-Aldrich Co. Bst 2.0 DNA Polymerase (8,000 U/mL), magnesium sulfate, and deoxyribonucleoside 5′-triphosphates (dNTP) were obtained from New England BioLabs. EvaGreen® dye was purchased from Biotium Inc. LAMP primers of lambda DNA were synthesized by Integrated DNA Technologies according to previous literature (Goto et al., 2009). The homemade non-buffered LAMP reaction solution was prepared by mixing 10 μL 1M (NH4)2SO4, 1 μL Tween 20, 25 μL 1 M KCl, 4 μL 1M KOH, 140 μL dNTP, 40 μL Bst 2.0 DNA Polymerase, and 270 μL ddH2O. LAMP primers mix was prepared by mixing 40 μL 100 μM FIP/BIP (Forward Inner Primer/ Backward Inner Primer), 20 μL 100 μM LF/LB (Forward Loop primer/ Backward Loop primer), and 5 μL 100 μM F3/B3 (Forward outer primer/ Backward outer primer) (Satoh et al., 2013; Zhao et al., 2010). The non-buffered LAMP reaction solution and LAMP primer mix were stored at −20 °C. Real-time fluorescence LAMP reactions were monitored on the CFX96 Touch™ Real-Time PCR detection system (Bio-Rad, USA).
2.2. Design and Fabrication of E-Tube Cap System
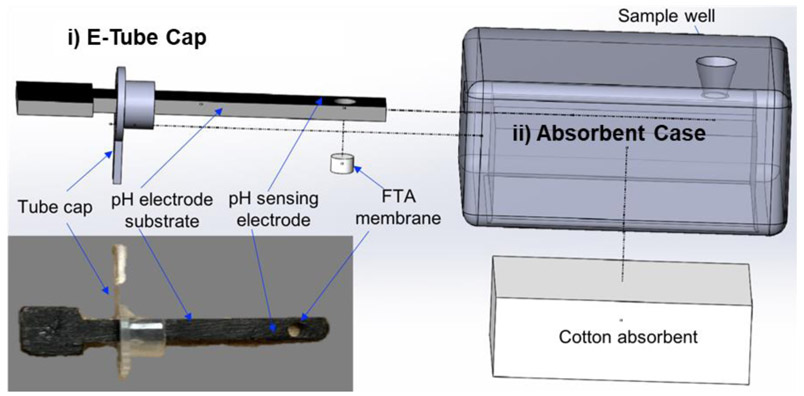
The E-tube cap system (Figure 1) mainly consists of: i) an E-tube cap for nucleic acid extraction and electrochemical detection, and ii) a capillary-driven fluidic absorbent case for waste collection. The E-tube cap (inset of Figure 1) contains i) one 3D printed electrode substrate coating with pH sensing material (carbon/iridium oxide layer), ii) one PCR tube cap and iii) one 2.0 mm-diameter FTA membrane incorporated into 3D printed electrode substrate. Figure S1A illustrated the fabrication process of the pH sensing electrode. Briefly, the pH sensing electrode substrate (Figure S1A(i)) was first designed by using Solidworks and printed by Form 2 3D printer (Formlabs Form 2, Formlabs Inc.) with a high-resolution stereolithographic laser as described previously (Kadimisetty et al., 2018). Next, both sides of the electrode substrate were coated with carbon paste (BQ226, Dupont Inc.) to serve as an electrode conductive layer (Figure S1A(ii)). Then, to generate a pH sensing electrode, one side of the electrode holder was partially coated with an iridium oxide layer (IrOx, Sigma-Aldrich Co.) (Figure S1A(ii)) to serve as pH sensing material (Figure S1B(ii)). Another side without IrOx coating serves as the reference electrode (Figure S1B(i)). The electrode surfaces coated with different material were observed and measured by scanning electron microscope (SEM, FEI Nova NanoSEM 450) with energy-dispersive X-ray spectroscopy (EDS) microanalysis equipment (Oxford Microanalysis X-Max 80) and its electrochemical impedance spectroscopy (EIS) was analyzed by a potentiostat (CHI-600, CHI Inc.). The 3D printed absorbent case consists of: i) one 3D printed case with sample well, electrode entrance, and absorbent chamber, and ii) one cotton absorbent which provides capillary force and collects the waste during DNA extraction.
Figure 1.
Schematic illustration of the electronic-tube cap (E-tube cap) system consisting of: i) E-tube cap with pH sensing electrode, FTA membrane and tube cap, and ii) 3D printed absorbent case with cotton absorbent. The left bottom insert is a photo of the assembled E-tube cap.
2.3. Electrochemical pH Sensing Detection of E-Tube Cap
To monitor the electrochemical signal, the pH sensing side (IrOx/carbon paste coating) of the 3D printed electrode substrate was connected to the working electrode port of the multi-channel potentiostat (1040C 8-channel potentiostat, CH Instruments, Inc.), and the reference electrode side (carbon paste coating) was connected to the reference electrode port. The pH response of the pH sensing electrode was calibrated by adding sodium hydroxide (NaOH) at room temperature (~20°C) (Xie et al., 2014). The potential-time program of the potentiostat was applied to record the real-time voltage response of the pH sensing electrode. To determine the effect of the solution temperature on the pH measurement of the electrode, the pH response of the pH sensing electrode was measured at five different temperatures (20 °C, 30 °C, 40 °C, 50 °C, and 63 °C). To test the stability of the electrode, the electrode was soaked into the solution which pH is 6.5 for 24 hours. All tests of the pH sensing electrodes were replicated three times.
2.4. Real-time Electrochemical Monitoring of LAMP Amplification by E-Tube Cap
In this study, lambda DNA (#SD0021, Thermo Inc.) was used as a model analyte to evaluate the performance of our E-tube cap. The LAMP reaction system includes 7.5 μL non-buffered LAMP reaction solution, 3.6 μL ddH2O, 0.8 μL LAMP primers mix, 0.9 μL phenol red (114529, Sigma-Aldrich Co.), and 1.2 μL 100 mM MgSO4 (Sigma-Aldrich Co.). For fluorescence detection, 0.5 μL Eva green (Sigma-Aldrich Co.) was added to the LAMP reaction system. All LAMP reactions in the presence/absence of 1 μL target (ddH2O for a negative control) were incubated at 63 °C. For comparison, the real-time fluorescence LAMP assays of various DNA concentrations were tested on the CLX96 Touch™ real-time PCR detection system.
To heat the E-tube cap system for isothermal amplification, one heating block (MyBlock Mini Dry Bath, Universal Medical, Inc.) was used to incubate the E-tube cap system at 63 °C. To monitor the electrochemical signals in real-time, the E-tube cap with pH sensing electrode was directly inserted into the reaction tubes containing different concentration DNA templates for pH sensing detection of isothermal amplification, and the electrochemical signals were monitored in real-time by using 4-Channel Potentiometer/Potentiostat (Model 1400 Series, CHI). The pH monitoring tests during the DNA amplification process were replicated three times.
2.5. Detection of Lambda DNA Spiked in Saliva Sample by E-Tube Cap
In the saliva sample testing, lambda DNA was added into de-identified saliva samples at various concentrations (i.e., 102, 103, and 104 copies/mL) to mimic clinical samples. To enrich the DNA and remove potential inhibitors from saliva samples, 1 mL of saliva samples spiked with lambda DNA was first introduced in the sample well of the absorbent case with the E-tube cap. When the saliva samples flowed through the FTA membrane of the E-tube cap due to the capillary force provided by cotton absorbent, the DNA was captured and enriched on the FTA membrane. Next, 100 uL of Whatman FTA purification reagent (Sigma-Aldrich Co.) was added to the sample well to remove the potential inhibitors on the FTA membrane. Then, the E-tube cap was pulled out from the absorbent case and inserted into one LAMP reaction tube containing all chemicals required for the LAMP reaction except for DNA templates. The assembled E-tube cap reaction system was incubated in a heating block and the captured DNA by the FTA membrane directly served as a target template for isothermal amplification.
3. Results and Discussion
3.1. Basic Characterization of pH Sensing Electrode
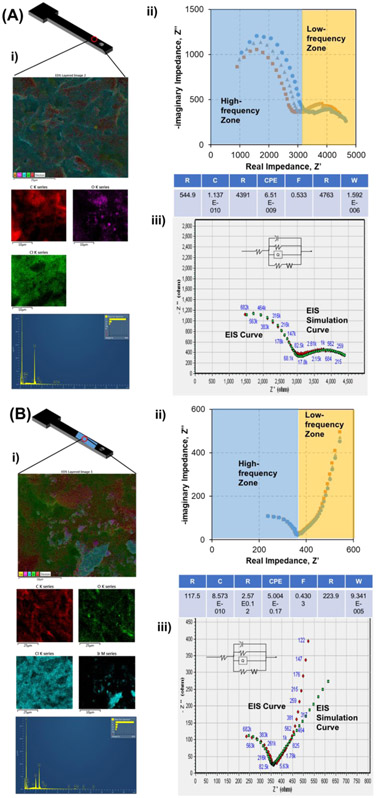
Since the polymer-based resin was used to print the electrode substrate, the 3D printed electrode substrate itself was not electrically conductive. To generate the electrochemical electrodes, the electrically conductive carbon paste (CP) layer and IrOx/CP layer were, respectively, coated on different sides of the electrode substrate to serve as the reference electrode and pH sensing electrode (Castellon et al., 2011; Xu et al., 2016). The electrical resistance of different materials was summarized in Table S1. Figure S2 showed the SEM images of the 3D printed electrode substrate, carbon paste coating, and IrOx/CP coating. To further distinguish the difference of various electrode materials, the energy-dispersive X-ray spectroscopy was used to identify the elemental compositions on the surface of both CP reference electrode and IrOx/CP pH sensing electrode (Zhao et al., 2005). As shown in Figure 2A(i) and 2B(i), there was ~9.65% iridium element on the IrOx/CP electrode, while not on the carbon paste reference electrode, which demonstrated that the iridium oxide was successfully coated on the electrode surface.
Figure 2.
(A) Energy dispersive spectroscopy (EDS) image /element map (i), electrochemical impedance spectroscopy (EIS) curves (ii), and EIS simulation (iii) of the carbon paste (CP) electrode. (B) Energy dispersive spectroscopy (EDS) image /element map (i), EIS (ii), and EIS simulation (iii) of the IrOx/CP electrode.
Next, electrochemical impedance spectroscopy (EIS) was used to study the electrochemical behavior of the CP and IrOx/CP electrodes. As shown in Figure 2A(ii) and 2B(ii), the EIS results showed that there was a perturbative characterization of the dynamics of an electrochemical process on the surface of the electrodes (Suni, 2008). The high-frequency part and the low-frequency part of the EIS spectra were influenced by the ability of the charge transfer and mass transfer, respectively (Suni, 2008). For the high-frequency part of the EIS spectra, the diameter of the CP electrode’s EIS is much smaller than that of the IrOx/CP electrode as shown by the EIS simulation results (Rin-MCP, 544.9 Ω >> Rin-MCI, 117.5 Ω) (Figure 2A(ii) and 2B(ii)), which demonstrated that the IrOx/CP coating has better charge transferability than the CP coating. In the low-frequency area, the slope of the IrOx/CP electrode (pH sensing electrode) was much higher than that of the CP electrode (reference electrode), which indicated better mass transferability (Figure 2A(ii), (iii) and 2B(ii), (iii)). Therefore, the IrOx, as a widely used pH sensing material (Ges et al., 2007; Xu et al., 2016), not only provides pH sensing capability, but also improves the mass transfer and charge transfer properties in our E-tube cap system.
3.2. Electrochemical Characterization of pH Sensing Electrode
According to previous literature (Jogezai and Shabbir, 2018; Xie et al., 2014), the pH value of the LAMP reaction changed from 8.8 to ~ 6.8 when the reaction occurred in a non-buffered solution. Here, we evaluated the pH sensing response characteristics of our pH sensing electrode by using solutions with various pH values ranging from 6 to 9. The linearity relationship between the potential and the pH was described as the Nernst Equation (Ges et al., 2007; Xu et al., 2016) (Eq. 1).
| (1) |
Where E (T) is the potential of the E-tube cap; E°(T) is the constant standard potential (mV) at temperature T (Kelvin); R is the molar gas constant (8.3144 J mol−1 K−1): F is the Faraday constant (96,485 C mol−1); T is the temperature (Kelvin); n is the charge of the ion; The entire term (2.303 RT/nF) is the slope of the Nernst Equation. As shown in Figure S3A, the potential of the pH sensing electrode decreased with the increase of the pH value. As shown in Figure S3B, the pH sensing electrode showed a sub-Nernstian response with a slope of −31.32 ± 0.5 mV/pH at room temperature, and excellent linear response within the pH range of 6–9 with a high R2 value (0.9999). The sub-Nernstian response of our pH sensing electrode may be attributed to the combination of H+ ion exchange and electronic transfer involving one or two electrons (De Marco and Pejcic, 2000; Manjakkal et al., 2015).
According to the Nernst Equation, the theoretical slope (2.303RT/nF, Eq.1) of pH measurement can be affected by the measured temperature of the solutions. Next, we determined the effect of the different measurement temperatures (from 20 °C to 63 °C) on the pH value. As shown in Figure S4, the influence factor of the temperature was 0.0628 pH/°C, which is close to the theoretical value, ~0.06 pH/°C. However, since our electrochemical LAMP reaction occurred at a constant temperature (e.g., 63 °C), there is no substantial effect of temperature variation on the pH monitoring during electrochemical LAMP assay.
In addition, we determined the stability, reproducibility, and interference of the pH sensing electrode. As shown in Figure S5A, the potential of the pH sensing electrode decreased only less than 5 mV in the 24 hours, which demonstrated that our pH sensing electrode has an excellent stability. In addition, the pH sensing electrode showed a high reproducibility and reliability at different pH solution ranging from 6 to 9 (Figure S5B). Since there are different ions in LAMP reaction system and saliva samples, we elvalated the interferences of different ions (Na+, SO42−, NO3−, Cl−, and K+) on the pH sensing electrode. As shown in Figure S5C, there is a less than 2 mV potential change caused by other non-target ions, showing high selectivy for hydrogen ion detection of our pH sensing electrode. It should pointed out that the concentrations of these non-target ions in the LAMP reaction system do not change and maintain constant during LAMP amplification. Thus, their interferences on the pH sensing electrode are nearly ignorable in our real-time electrochemical LAMP assay. Together, our pH sensing electrode showes an excellent stability, reproducibility and selectivity and is suitable for real-time, in situ monitoring of pH in LAMP reaction system.
3.3. Real-time Electrochemical Monitoring of LAMP amplification by E-Tube Cap
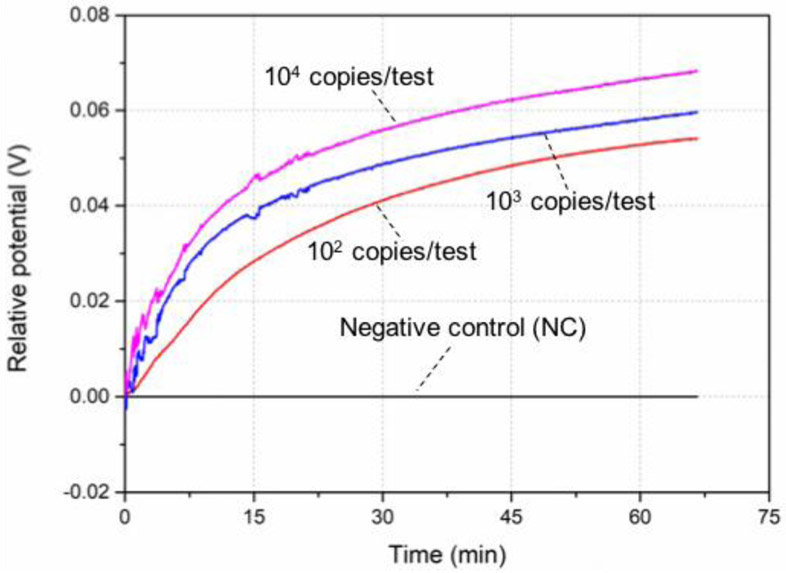
In the LAMP process, the DNA amplification is caused by the binding of the complementary primer to its target sequence. Successive incorporation of nucleotides leads to the release of hydrogen ions, which results in the pH decrease of non-buffered LAMP reaction solution (Toumazou et al., 2013). To minimize the effect of the potential-drifting during the LAMP process, the potentials of all pH sensing electrodes were normalized by subtracting the potential of the negative sample, which was set as the baseline. The higher the concentration of DNA samples, the higher the potential change (Figure 3). As shown in Figure 3, we can consistently detect 102 copies of DNA per test. For comparison purpose, we also tested the DNA samples by using real-time fluorescence LAMP and endpoint colorimetric LAMP assay. As shown in Figure S6 and S7A, both endpoint LAMP detection and real-time fluorescence LAMP assay are able to detect 102 copies of DNA per test. In the real-time fluorescence LAMP assay, the threshold time was defined as the reaction time when the fluorescence intensity increases 20% above the baseline level (Liu et al. 2011). Figure S7B depicted the threshold time (min) as a function of initial DNA concentration on a semi-log plot. Therefore, our pH sensing electrode system shows a comparable sensitivity to the conventional fluorescence LAMP assay.
Figure 3.
Real-time relative potential of the different concentration samples (0 (negative control), 102, 103, and 104 copies/test) during LAMP reaction. The electrochemical LAMP reaction system contains 7.5 μL non-buffered LAMP reaction solution, 3.6 μL ddH2O, 0.8 μL LAMP primers mix, and 1.2 μL 100 mM MgSO4 with/without 1 μL target DNA. The incubation temperation of LAMP assay is 63 °C.
3.4. Detection of Lambda DNA Spiked in Saliva Samples by E-Tube Cap System
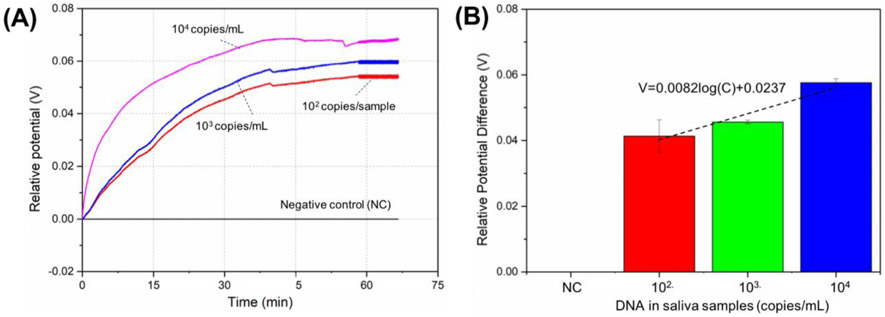
To further validate the feasibility of our E-tube cap system for human sample testing, we tested lambda DNA spiked in saliva samples with various concentrations, including 0 (negative control), 102, 103, and 104 copies per mL saliva sample. Figure 4A showed the real-time potential monitoring curves of the LAMP assay for the detection of different concentration samples in our E-tube cap system. When target DNA was present in saliva samples, the potentials of the pH sensing electrodes significantly increase. The higher the target DNA concentration in the saliva sample was, the higher the potential of the pH sensing electrode was. With the increase of the LAMP reaction time, the potential of the positive samples significantly increased in the first 30 minutes and tended to reach a plateau after approximately 30 minutes. Thus, in the point-of-care testing, real-time detection could be terminated in less than 30 min, which reduces the analysis time.
Figure 4.
(A) Real-time relative potential monitoring of LAMP amplification of saliva samples containing 0 (negative control), 102, 103 and 104 copies DNA/mL saliva sample in our integrated E-tube cap system. The electrochemical LAMP reaction system contains 7.5 μL non-buffered LAMP reaction solution, 3.6 μL ddH2O, 0.8 μL LAMP primers mix, and 1.2 μL 100 mM MgSO4. The incubation temperation of LAMP assay is 63 °C. (B) Relative potential change after 30-minute incubation as function of log of initial concentration of DNA in saliva samples (0, 102, 103 and 104 copies/mL) (n=3).
Figure 4B depicts the potential change after 30-minute LAMP incubation as a function of target DNA concentration in saliva (C) on a semi-log plot. In the range 102 < C < 104 copies DNA/mL, the voltage change decreases linearly as a function of log (C), which was correlated with the formula V= 0.0237 + 0.0082 log(C). In addition, the existence of the FTA membrane on the pH sensing electrode did not obviously interfere with LAMP amplification and pH sensing detection. The results clearly indicate that as few as 100 copies/mL in saliva samples can be detected with our E-tube cap system. In our E-tube cap system, the amplification results were monitored in real-time with a disposable and compact pH sensing electrode integrated with the FTA membrane, which eliminates the need for complex DNA sample preparation and enables a simple, low-cost, and integrated nucleic acid detection. To the best of our knowledge, this is the first demonstration of a compact pH sensing system integrating nucleic acid sample preparation, isothermal amplification, and label-free electrochemical DNA detection.
4. Conclusion
In this study, an integrated and compact E-tube cap system has been designed, fabricated, and tested for DNA sample extraction, isothermal amplification, and label-free pH sensing detection. The E-tube cap was incorporated with: i) a 3D printed electrode substrate that was coated with IrOx/carbon paste for pH sensing during the LAMP amplification reaction, and ii) a flow-through FTA membrane for DNA enrichment and purification from a raw saliva sample. Unlike previously reported electrochemical DNA detection approaches (Toumazou et al. 2013, Xie et al. 2014), our E-tube cap system provides an integrated and multi-functional DNA sensing platform with DNA extraction, isothermal amplification, and electrochemical detection. The E-tube cap system has been successfully demonstrated to detect lambda DNA in saliva samples with the sensitivity of 100 copies/mL. Besides, the fabrication of the pH sensing electrodes by 3D printing and coating processes allows for simple, rapid, and low-cost production. Such a simple, affordable, and sensitive diagnostic system has great potential to be used as a point-of-care diagnostic device for the rapid detection of various infectious diseases, including the recently emerged COVID-19 disease.
Supplementary Material
Highlights.
A simple and integrated electronic-tube cap (E-tube cap) was developed for DNA detection in saliva sample at the point-of-care
A multifunctional 3D printed electrode is able to perform DNA extraction, isothermal amplification, and real-time electrochemical detection
Real-time, quantitative electrochemical detection of nucleic acid without need of electrochemical label
Simple and disposable pH sensing electrode showed an excellent linear response within the pH range of 6-9 with a slope of −31.32 ± 0.5 mV/pH
A detection sensitivity of 100 copies /mL has been achieved for lambda DNA detection in saliva sample within 30 minutes.
Acknowledgment
The work was supported, in part, by National Institutes of Health R01EB023607, R61AI154642, and R01CA214072 as well as the Penn Center for AIDS Research (P30 AI 045008).
Footnotes
Declaration of interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Castellon J, Nguyen HN, Agnel S, Toureille A, Frechette M, Savoie S, Krivda A, Schmidt LE, 2011. IEEE Trans. Dielectr. Electr. Insul 18(3), 651–658. [Google Scholar]
- De Marco R, Pejcic B, 2000. Anal. Chem 72(4), 669–679. [DOI] [PubMed] [Google Scholar]
- Estrada-Peña A, Roura X, Sainz A, Miró G, Solano-Gallego L, 2017. Ticks Tick. Borne. Dis 8(4), 443–452. [DOI] [PubMed] [Google Scholar]
- Fu L, Tang D, Zhuang J, Lai W, Que X, Chen G, 2013. Biosens. Bioelectron 47, 106–112. [DOI] [PubMed] [Google Scholar]
- Ges IA, Ivanov BL, Werdich AA, Baudenbacher FJ, 2007. Biosens. Bioelectron 22(7), 1303–1310. [DOI] [PubMed] [Google Scholar]
- Goto M, Honda E, Ogura A, Nomoto A, Hanaki KI, 2009. Biotechniques 46(3), 167–172. [DOI] [PubMed] [Google Scholar]
- Hsieh K, Patterson AS, Ferguson BS, Plaxco KW, Soh HT, 2012. Angew. Chemie - Int. Ed 51(20), 4896–4900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jogezai N, Shabbir MI, 2018. Anal. Methods 10(35), 4233–4241. [Google Scholar]
- Kadimisetty K, Song J, Doto AM, Hwang Y, Peng J, Mauk MG, Bushman FD, Gross R, Jarvis JN, Liu C, 2018. Biosens. Bioelectron 109, 156–163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu B, Chen J, Wei Q, Zhang B, Zhang L, Tang D, 2015. Biosens. Bioelectron 69, 241–248. [DOI] [PubMed] [Google Scholar]
- Liu C, Geva E, Mauk M, Qiu X, Abrams WR, Malamud D, Curtis K, Owen SM, Bau HH, 2011. Analyst 136(10), 2069–2076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Lu B, Tang Y, Du Y, Li B, 2020. Electrochem. commun 111, 106665. [Google Scholar]
- Luo J, Fang X, Ye D, Li H, Chen H, Zhang S, Kong J, 2014. Biosens. Bioelectron 60, 84–91. [DOI] [PubMed] [Google Scholar]
- Lv S, Zhang K, Zhu L, Tang D, 2019. Anal. Chem 92(1), 1470–1476. [DOI] [PubMed] [Google Scholar]
- Manjakkal L, Cvejin K, Bajac B, Kulawik J, Zaraska K, Szwagierczak D, 2015. Electroanalysis 27(3), 770–781. [Google Scholar]
- Park BH, Oh SJ, Jung JH, Choi G, Seo JH, Kim DH, Lee EY, Seo TS, 2017. Biosens. Bioelectron 91, 334–340. [DOI] [PubMed] [Google Scholar]
- Satoh T, Matsumoto K, Fujii T, Sato O, Gemma N, Onuki M, Saito H, Aoki D, Hirai Y, Yoshikawa H, 2013. J. Virol. Methods 188(1–2), 83–93. [DOI] [PubMed] [Google Scholar]
- Stobiecka M, Chałupa A, 2015.Chem. Pap 69 (1), 62–76. [Google Scholar]
- Suni II, 2008. TrAC - Trends Anal. Chem 27(7), 604–611. [Google Scholar]
- Toumazou C, Shepherd LM, Reed SC, Chen GI, Patel A, Garner DM, Wang CJA, Ou CP, Amin-Desai K, Athanasiou P, Bai H, Brizido IMQ, Caldwell B, Coomber-Alford D, Georgiou P, Jordan KS, Joyce JC, La Mura M, Morley D, Sathyavruthan S, Temelso S, Thomas RE, Zhang L, 2013. Nat. Methods 10(7), 641. [DOI] [PubMed] [Google Scholar]
- Wang Q, Zhou Y, Li S, Zhuo C, Xu S, Huang L, Yang L, Liao K, 2013. PLoS One 8(7). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xie S, Yuan Y, Song Y, Zhuo Y, Li T, Chai Y, Yuan R, 2014. Chem. Commun 50(100), 15932–15935. [DOI] [PubMed] [Google Scholar]
- Xu Z, Dong Q, Otieno B, Liu Y, Williams I, Cai D, Li Y, Lei Y, Li B, 2016. Sensors Actuators, B Chem. 237, 1108–1119. [Google Scholar]
- Zeng R, Luo Z, Su L, Zhang L, Tang D, Niessner R, Knopp D, 2019. Anal. Chem 91(3), 2447–2454. [DOI] [PubMed] [Google Scholar]
- Zeng R, Wang W, Chen M, Wan Q, Wang C, Knopp D, Tang D, 2021. Nano Energy 82, 105711. [Google Scholar]
- Zhao P, Liang Q, Li Y, 2005. Appl. Surf. Sci 252(5), 1596–1607. [Google Scholar]
- Zhao X, Li Y, Wang L, You L, Xu Z, Li L, He X, Liu Y, Wang J, Yang L, 2010. Mol. Biol. Rep 37(5), 2183–2188. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.